Abstract
The recently discovered wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2 was analyzed through a high-throughput experimental design to improve ethanol yields in synthetic media with glucose, xylose, and cellobiose as carbon sources and acetic acid, furfural, formic acid, and NaCl as fermentation inhibitors. After Plackett–Burman (PB) and central composite design (CCD), the optimized condition was used in a fermentation kinetic analysis to compare this yeast's performance with an industrial Saccharomyces cerevisiae strain (JDY-01) genetically engineered to achieve a higher xylose fermentation capacity and fermentation inhibitors tolerance by overexpressing the genes XYL1, XYL2, XKS1, and TAL1. Our results show that furfural and NaCl had no significant effect on sugar consumption by UFFS-CE-3.1.2. Surprisingly, acetic acid negatively affected glucose but not xylose and cellobiose consumption. In contrast, the pH positively affected all the analyzed responses, indicating a cell's preference for alkaline environments. In the CCD, sugar concentration negatively affected the yields of ethanol, xylitol, and cellular biomass. Therefore, fermentation kinetics were carried out with the average concentrations of sugars and fermentation inhibitors and the highest tested pH value (8.0). Although UFFS-CE-3.1.2 fermented glucose efficiently, xylose and cellobiose were mainly used for cellular growth. Interestingly, the genetically engineered strain JDY-01 consumed ~ 30% more xylose and produced ~ 20% more ethanol. Also, while UFFS-CE-3.1.2 only consumed 32% of the acetic acid of the medium, JDY-01 consumed > 60% of it, reducing its toxic effects. Thus, the overexpressed genes played an essential role in the inhibitors' tolerance, and the applied engineering strategy may help improve 2G ethanol production.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Second-generation (2G) biorefineries use lignocellulosic residues as raw material, and this production process depends on the hydrolysis of these biomasses rich in cellulose and hemicellulose. As a result of this hydrolysis, mainly the hexose glucose, the pentose xylose, and the disaccharide cellobiose (consisting of two glucose molecules joined together by a ß-1,4 glycosidic bond) are obtained—cellobiose, though, is present in a proportion of at least five times smaller in relation to the two mentioned monosaccharides [1,2,3,4]. Therefore, the production of 2G ethanol depends on the efficient fermentation of glucose, xylose, and cellobiose. However, the yeast species currently used in the bioethanol production (Saccharomyces cerevisiae) is capable of fermenting only the hexose among these three carbohydrates. To overcome this problem, 2G ethanol depends either on the selection of new species that can metabolize not only glucose but also xylose and cellobiose, or on the construction of genetically modified strains of S. cerevisiae to enable it to ferment these two carbohydrates [5, 6].
Biorefineries, however, can produce more than one product from the same lignocellulosic residue. In fact, in addition to ethanol, more recently, xylitol has also been widely discussed in the literature, given its numerous applications, especially in the pharmaceutical, cosmetics, and food industries [7,8,9]. In this sense, while glucose and cellobiose can be fermented to ethanol by the yeasts used in the process, xylose can be converted either into ethanol or into the so-referred sugar-alcohol.
For xylose fermentation into ethanol, yeast cells are required to be able to express the enzymes xylose reductase (XR), xylitol dehydrogenase (XDH), and xylulokinase (XK). In subsequent reactions, these enzymes will convert xylose to xylitol, xylulose, and, finally, xylulose-5P, which is then destined for the pentose phosphate pathway. Also, it is desirable for XR and XDH to be capable of recycling the same coenzyme in both oxide-reduction reactions (NADH/NAD+), thus avoiding a possible redox imbalance inside the cells, which tends to accumulate xylitol to the detriment of ethanol production [6, 10]. Indeed, this causes some species of yeast to end up only generating xylitol from xylose, which also, as mentioned above, presents itself as an important biotechnological product [11, 12].
However, the fermentative capacity is not the only desirable characteristic for the microorganism to be used in the process. Inhibitor compounds of the fermentative metabolism, such as furfural, formic acid, and predominantly acetic acid, are generated in the pretreatment and hydrolysis stages of the process, which are carried out before fermentation. Due to the presence of these compounds, to guarantee efficiency in the process, it is necessary to select yeasts (prospecting wild species or strains) tolerant to their toxic effects [3, 13, 14] or to genetically engineer S. cerevisiae to increase its tolerance to those inhibitor compounds. Thus, in the context of genetic-engineered yeasts, besides the heterologous expression of XR and XDH enzymes recycling the same coenzyme, and the overexpression of XK, some authors have pointed to the modification of the genes that encode the enzymes of the pentose phosphate pathway (PPP) as a strategy to increase tolerance to the inhibitory compounds [15,16,17,18,19]. In their work with laboratory strains of S. cerevisiae, Hasunuma et al. [20, 21] demonstrated that increasing tolerance to inhibitors through overexpression of PPP enzymes (such as the transaldolase encoded by the TAL1 gene) consequently increased ethanol yields in fermentations carried out under different concentrations of weak acids (acetic acid and formic acid). However, as most of these studies used laboratory strains of S. cerevisiae, there is still uncertainty regarding the reproducibility of the results when conducted in industrial strains or strains derived from them and isolated from industrial environments, which have genomic adaptations different from those found in laboratory yeasts [22, 23]. Indeed, several reports have started to show that the phenotypic consequences of genomic modifications (e.g., gene deletions) can vary considerably between different strain backgrounds [24], an issue that can have significant implications in metabolic engineering strategies for generating optimized industrial yeast strains [25, 26].
Another desirable characteristic of an industrial yeast is its tolerance to the osmotic stress caused by high sugar concentration. This is, in fact, one of the differentials of S. cerevisiae, which is widely recognized for its high fermentative performance even at sugar concentrations as high as 200 g/L [27]. Not that this is a rule for every bioprocess, but normally the higher the sugar concentration, the higher the productivity of the target product, in such a way that the prospection or development of new yeasts includes higher tolerance to a hyperosmotic environment as a targeted trait. Although the industrial fermentation vats are known to select and domesticate xerotolerant yeasts [28], nature also presents niches with reduced water activity where resistant wild strains can be found, and this has recently shown to be quite promising [29].
Last but not least, besides the factors mentioned above, the biorefineries' water footprint has also been identified as a parameter to be optimized [30]. In this sense, the employment of seawater in the fermentation vats is envisaged, partially or totally replacing the use of freshwater. For this, however, fermenting microorganisms must also tolerate high concentrations of salt in addition to the characteristics mentioned above [31,32,33].
In this context, the present work compared the biotechnological potential of two taxonomically distant yeast strains: the recently isolated wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2 [1] and a new genetically modified industrial strain derivative from the widely known S. cerevisiae PE-2 [34,35,36]. After subjecting UFFS-CE-3.1.2 to two subsequent experimental design analyses in synthetic media with the carbohydrates glucose, xylose, and cellobiose, as well as the inhibitors furfural, acetic acid, formic acid, and NaCl, the optimized culture condition was applied to a fermentation kinetics assay to compare the performances of both the wild and genetically engineered strains.
Materials and methods
Yeasts
Two yeasts were used: the wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2, previously isolated from decaying wood samples [1], and the genetically modified strain S. cerevisiae JDY-01. This industrial recombinant strain was constructed based on the efficient industrial fuel ethanol yeast strain PE-2 [36]. This diploid S. cerevisiae strain was initially transformed with the chromosome-integrative plasmid pAUR-XKXDHXR [37] allowing overexpression of xylose reductase (XR, encoded by XYL1) and xylitol dehydrogenase (XDH, encoded by XYL2) from Scheffersomyces stipitis, as well as xylulokinase (XK, encoded by XKS1) from S. cerevisiae [38, 39]. Briefly, plasmid pAUR-XKXDHXR was digested with BsiWI, and then chromosomally integrated into the AUR1 locus of the yeast strain. After 90 min cultivation on rich YP medium (10 g/L yeast extract and 20 g/L peptone, pH5.0) containing 20 g/L glucose, the transformed cells were selected on plates of the same medium containing 20 g/L agar and 0.5 mg/L aureobasidin A (Takara Bio, Kyoto, Japan), producing strain MP-P5 (isogenic to PE-2, but AUR1-C::PPGK1-XKS1-TPGK1, PPGK1-XYL2-TPGK1, PPGK1-XYL1-TPGK1). Yeast transformation was performed by the lithium acetate method [40]. Strain MP-P5 was further improved by promoting the overexpression of the S. cerevisiae TAL1 gene encoding the transaldolase enzyme of the non-oxidative pentose phosphate pathway [41, 42]. The promoter region of this TAL1 gene was modified according to the polymerase chain reaction (PCR)-based gene replacement procedure [43]. The kanMX-PADH1 module from plasmid pFA6a-kanMX6-PADH1 [43] was amplified using primers TAL1-Kanr-F and TAL1-PADH1-R (Table 1), and the PCR product of 2394 bp (flanked by 40 bp of homology to the promoter and start regions of the TAL1 gene) containing the kanMX6 gene and the constitutive promoter of the ADH1 gene was used to transform competent yeast cells. After 2-h cultivation on YP-20 g/L glucose, the transformed yeast cells were plated on the same medium containing 20 g/L agar and 200 mg/L G-418 (Merck Sigma Aldrich Brasil, Barueri, Brazil) and incubated at 28 °C. G-418-resistant isolates were tested for proper genomic integration of the kanMX-PADH1 cassette at the TAL1 locus by diagnostic PCR using 3 primers (V-TAL1-F, V-TAL1-R, and V-kanr-F; Table 1). This set of three primers amplified a 501 bp fragment (primers V-TAL1-F and V-TAL1-R) from a normal TAL1 locus, or yielded a 2,700 bp fragment (primers V-TAL1-F and V-TAL1-R) or a 1479 bp fragment (primers V-kanr-F and V-TAL1-R) if the kanMX-PADH1 module was correctly integrated at the promoter region of the TAL1 gene, producing strain JDY-01.
Culture media and growth conditions
Cells were grown in cotton-plugged Erlenmeyer flasks with 1/5 of their volume filled with culture medium in a shaker with controlled temperature and agitation. Before each experimental design, the yeasts were pre-grown for 48 h at 30 °C and 150 rpm in YP rich media containing 20 g/L of glucose until they reached the exponential growth phase (OD570nm ~ 3,5). Then the cells were inoculated in minimal YNB synthetic media (6.7 g/L of yeast nitrogen base) plus different concentrations of glucose, xylose, cellobiose, acetic acid, furfural, formic acid, and NaCl, according to the experimental designs described below. Likewise, the cultures were carried out in different pH ranges, temperature, and agitation according to the matrices presented in Tables 2 and 3.
Plackett–Burman's experimental design
The Plackett–Burman (PB) experimental design matrix was assembled and analyzed using the Protimiza Experimental Design software [44] with the following variables: temperature, pH, agitation, and concentration of inoculum, sugars, and inhibitors (Table 2). The inoculum of each PB fermentation was prepared with yeasts pre-grown as described above. Then the cells were centrifuged (5000g, 3 min), washed twice with distilled water, and resuspended in the fermentation culture medium to reach an initial cell concentration of 1, 3, or 5 g/L (Table 2). After 48 h fermentation, cells were centrifuged (5000g, 3 min), and supernatants were filtered (0.45 μm) for subsequent quantification of sugar consumption by HPLC as described in the "Analytical methods".
Central composite design
The central composite design (CCD) was also assembled and analyzed with Protimiza software. Pre-grown cells were inoculated at the concentration of 1:100 (v/v—volume of pre-inoculum per volume of medium to be inoculated). Yeasts were cultivated at 30 °C and 150 rpm in YNB medium containing 1.0 g/L of furfural, 0.5 g/L of formic acid, 17.5 g/L of NaCl, and different concentrations of carbohydrates and acetic acid (Table 2). The pH was also analyzed as a variable of this CCD. In Table 3, the column "total sugars" represents the combination of the three carbohydrates analyzed: 40 g/L of glucose, 40 g/L of xylose, and 8 g/L of cellobiose (88 g/L total sugar); 25 g/L glucose, 25 g/L xylose, and 5 g/L cellobiose (55 g/L total sugar); and 10 g/L glucose, 10 g/L xylose, and 2 g/L cellobiose (22 g/L total sugar). The remaining sugars and the concentration of xylitol and ethanol produced were determined by HPLC as described in the "Analytical methods".
Fermentation kinetics
For the fermentation kinetics, YNB media containing 25 g/L of glucose, 25 g/L of xylose, and 5 g/L of cellobiose (55 g/L of total sugars) were inoculated at the concentration of 1:100 (as described above). The inhibitors were added to the culture medium at concentrations of 1.0 g/L of furfural, 0.5 g/L of formic acid, 2.5 g/L of acetic acid, and 17.5 g/L of NaCl, and the pH was adjusted to 8.0. The assays were carried out with agitation of 150 rpm and a temperature of 30 °C. A total of ten samples were collected from each culture during 48 h of incubation to determine cellular growth by turbidity measurements at 570 nm (OD570nm) [45], and sugars, acetic acid, ethanol, and xylitol concentration through HPLC analysis (see below).
Analytical methods
During cell cultures, samples were periodically harvested, centrifuged (5000g, 3 min), and filtered (0.45 μm) for subsequent quantification of carbohydrates, ethanol, xylitol, and acetic acid by high-performance liquid chromatography (HPLC) as described by Tadioto et al. [8]. The analyses were performed using an LCMS-2020 chromatograph (Shimadzu) with a refractive index detector (RID-10, Shimadzu). For all compounds, a column for organic acids (Aminex HPX-87H, Bio-Rad) was used with a flow rate of 0.6 mL/min, using 5 mM of H2SO4 as mobile phase at a temperature of 50 °C. Calibration curves were used with four different concentrations of each analyzed compound.
Quantitative RT-PCR analysis
Quantitative RT-PCR (qRT-PCR) was conducted to verify the overexpression of the TAL1 gene in strain JDY-01 when compared to the expression of this gene in strain MP-P5. The yeast strains were grown in YP-20 g/L glucose medium to mid-log phase, centrifuged (5000g, 4 min at 4 °C), washed with cold distilled water, and according to the manufacturer's protocols, the total RNA of the cell pellets was extracted using the RNeasy® Mini Kit (Qiagen Brazil, São Paulo, Brazil). The total RNA of each sample (1 ug) was reverse transcribed to cDNA using the QuantiTect® Reverse Transcription Kit (Qiagen). The qRT-PCR reactions were performed with the QuantiFast® Sybr Green PCR Kit and the Rotor-Gene® Q equipment (Qiagen) using the primers for the TAL1 gene (primers RT-PCR TAL1-F and RT-PCR TAL1-R, Table 1) and for the actin gene (ACT1) that was selected as the endogenous reference gene (primers RT-PCR ACT1-F and RT-PCR ACT1-R, Table 1). A dissociation curve was generated for each assay to confirm the amplification of only one product. The 2−∆∆CT method [46] was used to calculate the relative expression levels of the TAL1 gene relative to the ACT1 gene, for each yeast strain, in triplicate.
Results and discussion
High-throughput analysis of a recently discovered Wickerhamomyces sp. strain
Although it has been only recently isolated from rotten wood [1], the wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2 has already displayed high biorefinery employment potential when it was submitted to lignocellulosic [1, 3] and pectin-rich hydrolysates [47] even with seawater-based media [48]. Indeed, we have previously shown that the wild strain UFFS-CE-3.1.2 showed 50% higher ethanol productivity in seawater-based papaya hydrolysates than the well-known industrial S. cerevisiae CAT-1 [48], another widely used yeast in the Brazilian fuel ethanol industry [22, 35, 38, 49]. Thus, aiming to reach even higher performances with the yeast Wickerhamomyces sp. UFFS-CE-3.1.2, we designed a widely comprehensive Plackett–Burman (PB) experimental plan to evaluate the effects of 11 variables on glucose, xylose, and cellobiose metabolism by the cells. The data in Table 2 show that UFFS-CE-3.1.2 is able to metabolize the three most abundant sugars in lignocellulosic hydrolysates, however, with a marked advantage for glucose, corroborating our previous studies [1, 3, 47, 48]. Indeed, this preference for glucose metabolism is a prevailing feature in yeasts [50,51,52]. After glucose, cellobiose was the most consumed sugar, although the concentration of this disaccharide was always lower than that of the glucose and xylose, which is usual in lignocellulosic hydrolysates [1,2,3,4].
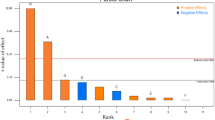
The PB statistical analysis showed that the pH effect (calculated t) was significantly positive for glucose and xylose consumption considering p ≤ 0.05, and for cellobiose consumption considering p ≤ 0.1 (Fig. 1), indicating that higher pH values favor sugar metabolization. Sugar concentration also had a significant effect on the percentual of their consumption, although, in this case, the influence was always negative, meaning that higher sugar concentration led to lower consumption (in percentual terms). Glucose uptake was negatively affected by high glucose and cellobiose titer; cellobiose consumption was impaired by itself and xylose; and the percentual of xylose internalized was only decreased in higher concentrations of this pentose (Fig. 1). In fact, the negative effects of one sugar on another have been described elsewhere [52,53,54], and it was somehow already expected that the higher the sugar concentration, the lower the percentual of sugar consumption by the yeast cells.
Besides the sugar concentration and pH, all the other variables had no consistent influence on the metabolization of the three carbohydrates analyzed; some of them had significant effects on one response but not on the other two (Fig. 1). Surprisingly, this was the case with acetic acid, which is known to impair sugar consumption, especially under low pH values, when it is in its undissociated (liposoluble) form, easily diffusing through the plasma membrane lipid bilayer and reaching the cell cytoplasm. In the cytosol, pH values higher than the external environment provide the dissociation of acetic acid to the ionic form, releasing protons that decrease cytoplasmic pH and consequently may damage the cell and thus produce lower ethanol yield [55,56,57,58].
Moreover, our data showed that this carboxylic acid exerted a significantly negative effect (p ≤ 0.05) only on glucose consumption, although most data in the literature indicate that xylose fermentation is usually more impaired by acetic acid [3, 57, 59]. Nevertheless, considering that acetic acid is probably the most common fermentation inhibitor in second-generation biorefinery processes, we decided to include it as a variable in a next step experimental design (see below). It should also be noted that NaCl had no significant influence on sugar consumption by UFFS-CE-3.1.2.
Since curvature (which represents a favorable trend for the average values of the variables) had a significant positive effect (p ≤ 0.05) on the three responses analyzed (Fig. 1), the variables other than sugars, pH, and acetic acid were kept in their central values in the central composite design (CCD) that followed the PB analysis. In the CCD, glucose, xylose, and cellobiose concentrations were taken together as one variable: total sugar concentration (Table 3). Once again, acetic acid showed no significant effect on xylose and cellobiose consumption (data not shown), even though it significantly impaired ethanol and xylitol yields (Fig. 2a, b). Higher pH values always favored yeast metabolism, and sugar concentration had significant positive effects on ethanol and xylitol but a negative effect on cellular biomass yields (Fig. 2).
Calculated t and p value for Wickerhamomyces sp. UFFS-CE-3.1.2 central composite design (CCD), considering the yields of ethanol (a), xylitol (b), and cellular biomass (c) as responses. Significant effects are labeled with * for p < 0.05 and with ** for p < 0.1. Coefficients of determination: 79.42% (ethanol yield), 96.13% (xylitol yield), 94.52% (biomass yield)
In the next step, UFFS-CE-3.1.2 was subjected to a fermentation kinetic analysis. To this end, except for pH (which has consistently proven to have a beneficial influence at alkaline values), the other analyzed variables were kept in their central values (see "Fermentation kinetics" at "Materials and methods"). The kinetic analysis suggests that glucose was almost entirely fermented into ethanol, while xylose and cellobiose were mostly used as carbon sources for cellular growth (Fig. 3). Although xylose has not entirely been exhausted by the yeast cells, our data show an improvement in the consumption rate of this pentose compared with a previous study [3], indicating that PB and CCD led to an optimization of the UFFS-CE-3.1.2 fermentation performance. Moreover, it is worth noting that this better performance was achieved in the presence of acetic acid (2.5 g/L), formic acid (0.5 g/L), furfural (1.0 g/L), and NaCl (17.5 g/L), which have shown to inhibit sugar metabolism at similar concentration ranges [21, 31, 60, 61].
Fermentation kinetics of Wickerhamomyces sp. UFFS-CE-3.1.2 (closed symbols) and S. cerevisiae JDY-01 (open symbols) in the presence of fermentation inhibitors. During 48 h incubation period, samples were harvested from the media for the quantification of glucose (circle), xylose (triangle), or cellobiose (square) consumption (a), ethanol (cross-hair circle), and xylitol (diamond) production (b), and cellular growth (inverted triangle) and acetic acid concentration (hexagon) in the medium (c). The data are expressed as averages ± standard errors from three completely independent experiments
Only a small amount of the xylose consumed by the cells (~ 5%) was secreted in the media as xylitol (Fig. 3b), indicating that the common redox imbalance is not impairing the first two reactions in xylose metabolization (catalyzed by XR and XDH) by Wickerhamomyces sp. UFFS-CE-3.1.2. In contrast, this also indicates a low potential of this yeast for xylitol production in a multiproduct biorefinery context since previous studies have shown xylitol yields by wild yeasts up to 11 times higher in similar fermentation conditions [8, 62, 63].
Comparing the wild yeast with a genetically engineered industrial strain
Although bioprospection of wild yeasts has shown to be a promising approach for making many biorefinery processes economically viable [64,65,66], genetic engineering has also proven to be a feasible alternative to improve residual biomasses conversion into bioproducts [5, 67, 68]. In this sense, we decided to compare the wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2 fermentation performance with that of a genetically modified industrial S. cerevisiae strain. The recombinant fuel ethanol S. cerevisiae strain JDY-01 is derived from strain PE-2, an efficient fermenting diploid strain used in first-generation bioethanol production in Brazil [36], but unable to ferment xylose [4, 69]. However, in this study, PE-2 was initially transformed with a chromosome-integrative plasmid to overexpress the three enzymes (XR, XDH, and XK) required for xylose utilization, yielding strain MP-P5. This strain was further improved by promoting the overexpression of the S. cerevisiae transaldolase encoded by the TAL1 gene, a rate-limiting enzyme of the non-oxidative pentose phosphate pathway required for improved xylose consumption [41, 42]. Our PCR analysis revealed that, in strain JDY-01, only one of the TAL1 genes had its promoter region replaced by the kanMX-PADH1 module, and thus the other TAL1 gene present in this diploid strain retained its normal promoter.
Interestingly, S. cerevisiae JDY-01 in the presence of the inhibitors (acetic and formic acids, furfural, and NaCl) consumed glucose faster than the wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2, and consumed ~ 30% more xylose, displaying a ~ 20% higher ethanol production, even with no cellobiose utilization (Fig. 3). As expected, this higher ethanol production reflects its lower xylitol production and cellular growth, indicating that the engineered S. cerevisiae strain deviates more sugar (glucose and xylose) to the alcoholic fermentation route than Wickerhamomyces sp. UFFS-CE-3.1.2, thus being more suitable for the 2G ethanol industry. At this point, it is important to remember that the strain JDY-01 lacks a periplasmic ß-glucosidase (or a cellobiose transporter plus an intracellular ß-glucosidase) to allow it to consume and ferment cellobiose, which could ensure an even higher ethanol yield if this recombinant yeast were further modified to ferment this disaccharide.
As it can be claimed that the overexpression of XR, XDH, and XK may be the major reason behind the better fermentative performance of JDY-01, it is worth mentioning that this strain also consumed more xylose and produced more ethanol than its parental strain MP-P5 (data not shown), which overexpresses the same three enzymes. The only difference between these two strains is the strong promoter PADH1 upstream of the TAL1 gene in strain JDY-01. When TAL1 expression was analyzed through qRT-PCR, we observed that this gene was 5.4 times upregulated in JDY-01 compared to MP-P5. Thus, JDY-01 higher transaldolase activity most likely led this strain to increase its xylose consumption capacity even in the presence of fermentation inhibitors. This is corroborated by the acetic acid consumption by JDY-01, since during the 48 h incubation, this strain consumed over 60% of this acid available in the medium (Fig. 3c), probably using it as a carbon source and thus reducing its toxic effects. Significantly, strain UFFS-CE-3.1.2 only consumed 32% of the acetic acid, mostly at the end of the fermentation.
By consuming acetic acid at such a rate, JDY-01 shows to be able to use it as a carbon source. Ending or decreasing the repressive catabolic effect of glucose, acetate may be converted into acetyl-CoA and then stimulate the glyoxylate cycle and gluconeogenesis [60, 70]. By doing this, the strain JDY-01 not only has the toxic effect of acetic acid reduced but also has it used as an energy source [71, 72]. Thus, our results suggest that, as previously hypothesized [15, 20, 21], overexpression of TAL1 may increase yeast tolerance to the so-referred carboxylic acid.
Moreover, it should be noted that the fermentative performance observed for the strain JDY-01 was achieved in the presence of 17.5 g/L of NaCl, which corresponds to approximately half the concentration of this salt in seawater [73]. In this sense, our data indicate that the engineered strain S. cerevisiae JDY-01 could handle 2G ethanol production in a lower water footprint condition, considering, for instance, a situation where freshwater and seawater are used in a ratio of 1:1 (v/v) in the wort.
It is also worth noting that our results not only presented a new engineered S. cerevisiae strain with high potential to be employed in the 2G ethanol industry but also a highly feasible approach to improve xylose fermentation by this yeast species. Considering the genotypic and phenotypic variability among several S. cerevisiae strains [23, 74], it would be interesting to reproduce the same genetic modifications herein analyzed in other genetic backgrounds. With the aim of increasing water security during 2G ethanol production, the marine S. cerevisiae strains that Zaky and coworkers [32, 33, 75, 76] have been isolating and characterizing, for instance, are certainly worth trying.
Conclusion
The newly discovered wild yeast Wickerhamomyces sp. UFFS-CE-3.1.2 has previously shown biotechnological potential when analyzed in seawater-based lignocellulosic and pectin hydrolysates. In the present study, a high-throughput experimental design was used to improve its fermentation performance in the presence of the main sugars and inhibitors found in the biorefinery vats. Although acetic acid, furfural, formic acid, and NaCl showed to play a low (or no) effect on its sugar metabolism, this yeast was unable to ferment xylose when subjected to intermediary concentrations of these inhibitory compounds. In this situation, the strain Wickerhamomyces sp. UFFS-CE-3.1.2 consumed ~ 62% of the xylose available through the respiratory route generating cellular biomass. Nevertheless, the experimental designs that were carried out highly improved the xylose consumption capacity of this yeast (compared to our previous studies) and allowed the cells to fully consume cellobiose.
On the other hand, the genetically engineered strain S. cerevisiae JDY-01 consumed over 83% of xylose from the medium and produced 20% more ethanol than the wild yeast in the presence of the same fermentation inhibitors. Furthermore, this higher fermentative performance was achieved despite lacking cellobiose consumption, which was completely metabolized by the wild yeast. Thus, if JDY-01 is additionally engineered to ferment this disaccharide, it will very likely be able to further increase ethanol yield in a seawater-based biorefinery context.
References
Bazoti SF, Golunski S, Pereira Siqueira D, Scapini T, Barrilli ÉT, Alex Mayer D, Barros KO, Rosa CA, Stambuk BU, Alves SL, Valério A, de Oliveira D, Treichel H (2017) Second-generation ethanol from non-detoxified sugarcane hydrolysate by a rotting wood isolated yeast strain. Bioresour Technol 244:582–587. https://doi.org/10.1016/j.biortech.2017.08.007
Conesa C, Seguí L, Fito P (2018) Hydrolytic performance of Aspergillus niger and Trichoderma reesei cellulases on lignocellulosic industrial pineapple waste intended for bioethanol production. Waste Biomass Valorization 9:1359–1368. https://doi.org/10.1007/S12649-017-9887
Bonatto C, Venturin B, Mayer DA, Bazoti SF, de Oliveira D, Alves SL, Treichel H (2020) Experimental data and modelling of 2G ethanol production by Wickerhamomyces sp. UFFS-CE-3.1.2. Renew Energy 145:2445–2450. https://doi.org/10.1016/j.renene.2019.08.010
Bohn LR, Dresch AP, Cavali M, Vargas ACG, Führ JF, Tironi SP, Fogolari O, Mibielli GM, Junior SLA, Bender JP (2021) Alkaline pretreatment and enzymatic hydrolysis of corn stover for bioethanol production. Res Soc Development 10:e149101118914. https://doi.org/10.33448/rsd-v10i11.18914
AlvesJunior SL, Scapini T, Warken A, Klanovicz N, Procópio DP, Tadioto V, Stambuk BU, Basso TO, Treichel H (2022) Engineered saccharomyces or prospected non-Saccharomyces: is there only one good choice for biorefineries? In: AlvesJunior S, Treichel H, Basso T, Stambuk BU (eds) Yeasts: from nature to bioprocesses. Bentham Science Publishers, Sharjah, pp 243–283
Eliodório KP, Cunha GC, Müller C, Lucaroni AC, Giudici R, Walker GM, Alves SL, Basso TO (2019) Advances in yeast alcoholic fermentations for the production of bioethanol, beer and wine. In: Advances in applied microbiology. Academic Press Inc., pp 61–119
Antunes FAF, Thomé LC, Santos JC, Ingle AP, Costa CB, dos Anjos V, Bell MJV, Rosa CA, Silva SSD (2021) Multi-scale study of the integrated use of the carbohydrate fractions of sugarcane bagasse for ethanol and xylitol production. Renew Energy 163:1343–1355. https://doi.org/10.1016/j.renene.2020.08.020
Tadioto V, Milani LM, Barrilli ÉT, Baptista CW, Bohn L, Dresch A, Harakava R, Fogolari O, Mibielli GM, Bender JP, Treichel H, Stambuk BU, Müller C, Alves SL (2022) Analysis of glucose and xylose metabolism in new indigenous Meyerozyma caribbica strains isolated from corn residues. World J Microbiol Biotechnol 38:35. https://doi.org/10.1007/s11274-021-03221-0
Trichez D, Steindorff AS, Soares CEVF, Formighieri EF, Almeida JRM (2019) Physiological and comparative genomic analysis of new isolated yeasts Spathaspora sp JA1 and Meyerozyma caribbica JA9 reveal insights into xylitol production. FEMS Yeast Res 19:foz034. https://doi.org/10.1093/femsyr/foz034
Stambuk BU, Eleutherio ECA, Florez-Pardo LM, Souto-Maior AM, Bon EPS (2008) Brazilian potential for biomass ethanol: Challenge of using hexose and pentose cofermenting yeast strains. J Sci Ind Res (India) 67:918–926
Cadete RM, Rosa CA (2018) The yeasts of the genus Spathaspora: potential candidates for second-generation biofuel production. Yeast 35:191–199. https://doi.org/10.1002/yea.3279
Carneiro CVGC, Silva FCP, Almeida JRM (2019) Xylitol production: identification and comparison of new producing yeasts. Microorganisms. https://doi.org/10.3390/microorganisms7110484
Moncada J, Tamayo JA, Cardona CA (2014) Integrating first, second, and third generation biorefineries: Incorporating microalgae into the sugarcane biorefinery. Chem Eng Sci 118:126–140. https://doi.org/10.1016/j.ces.2014.07.035
Morales-Oyervides L, Ruiz-Sánchez JP, Oliveira JC, Sousa-Gallagher MJ, Morales-Martínez TK, Albergamo A, Salvo A, Giuffrida D, Dufossé L, Montañez J (2020) Medium design from corncob hydrolyzate for pigment production by Talaromyces atroroseus GH2: Kinetics modeling and pigments characterization. Biochem Eng J 161:107698. https://doi.org/10.1016/j.bej.2020.107698
Fujitomi K, Sanda T, Hasunuma T, Kondo A (2012) Deletion of the PHO13 gene in Saccharomyces cerevisiae improves ethanol production from lignocellulosic hydrolysate in the presence of acetic and formic acids, and furfural. Bioresour Technol 111:161–166. https://doi.org/10.1016/j.biortech.2012.01.161
Kim SR, Skerker JM, Kong II, Kim H, Maurer MJ, Zhang G-C, Peng D, Wei N, Arkin AP, Jin Y-S (2017) Metabolic engineering of a haploid strain derived from a triploid industrial yeast for producing cellulosic ethanol. Metab Eng 40:176–185. https://doi.org/10.1016/j.ymben.2017.02.006
Li B, Liu N, Zhao X (2022) Response mechanisms of Saccharomyces cerevisiae to the stress factors present in lignocellulose hydrolysate and strategies for constructing robust strains. Biotechnol Biofuels Bioproducts 15:28. https://doi.org/10.1186/s13068-022-02127-9
Sanda T, Hasunuma T, Matsuda F, Kondo A (2011) Repeated-batch fermentation of lignocellulosic hydrolysate to ethanol using a hybrid Saccharomyces cerevisiae strain metabolically engineered for tolerance to acetic and formic acids. Bioresour Technol 102:7917–7924. https://doi.org/10.1016/j.biortech.2011.06.028
Menegon YA, Gross J, Jacobus AP (2022) How adaptive laboratory evolution can boost yeast tolerance to lignocellulosic hydrolyses. Curr Genet 1:1–24. https://doi.org/10.1007/s00294-022-01237-z
Hasunuma T, Ismail KSK, Nambu Y, Kondo A (2014) Co-expression of TAL1 and ADH1 in recombinant xylose-fermenting Saccharomyces cerevisiae improves ethanol production from lignocellulosic hydrolysates in the presence of furfural. J Biosci Bioeng 117:165–169. https://doi.org/10.1016/j.jbiosc.2013.07.007
Hasunuma T, Sung K, Sanda T, Yoshimura K, Matsuda F, Kondo A (2011) Efficient fermentation of xylose to ethanol at high formic acid concentrations by metabolically engineered Saccharomyces cerevisiae. Appl Microbiol Biotechnol 90:997–1004. https://doi.org/10.1007/s00253-011-3085-x
Stambuk BU, Dunn B, Alves SL Jr, Duval EH, Sherlock G (2009) Industrial fuel ethanol yeasts contain adaptive copy number changes in genes involved in vitamin B1 and B6 biosynthesis. Genome Res 19:2271–2278. https://doi.org/10.1101/gr.094276.109
Franco-Duarte R, Bigey F, Carreto L, Mendes I, Dequin S, Santos MA, Pais C, Schuller D (2015) Intrastrain genomic and phenotypic variability of the commercial Saccharomyces cerevisiae strain Zymaflore VL1 reveals microevolutionary adaptation to vineyard environments. FEMS Yeast Res 15:fov063. https://doi.org/10.1093/femsyr/fov063
Galardini M, Busby BP, Vieitez C, Dunham AS, Typas A, Beltrao P (2019) The impact of the genetic background on gene deletion phenotypes in Saccharomyces cerevisiae. Mol Syst Biol. https://doi.org/10.15252/msb.20198831
Yi X, Alper HS (2022) Considering strain variation and non-type strains for yeast metabolic engineering applications. Life 12:510. https://doi.org/10.3390/life12040510
Varize CS, Bücker A, Lopes LD, Christofoleti-Furlan RM, Raposo MS, Basso LC, Stambuk BU (2022) Increasing ethanol tolerance and ethanol production in an industrial fuel ethanol Saccharomyces cerevisiae strain. Fermentation 8:470. https://doi.org/10.3390/fermentation8100470
Lopes ML, Paulillo SCL, Godoy A, Cherubin RA, Lorenzi MS, Giometti FHC, Bernardino CDD, de Amorim Neto HB, de Amorim HV (2016) Ethanol production in Brazil: a bridge between science and industry. Braz J Microbiol 47:64–76. https://doi.org/10.1016/j.bjm.2016.10.003
Hong M, Li J, Chen Y (2019) Characterization of tolerance and multi-enzyme activities in non-Saccharomyces yeasts isolated from Vidal blanc icewine fermentation. J Food Biochem 43:e13027. https://doi.org/10.1111/jfbc.13027
Perrusquía-Luévano S, Cano-Herrera MS, Guigón-López C, Avitia-Talamantes MDC, Torres-Torres C, Villalpando I (2019) Microbiology of high-sugar must fermentation by novel yeasts from the chihuahuan desert. FEMS Yeast Res 19:99. https://doi.org/10.1093/femsyr/foy099
Scapini T, Dalastra C, Camargo AF, Kubeneck S, Modkovski TA, Júnior SLA, Treichel H (2022) Seawater-based biorefineries: a strategy to reduce the water footprint in the conversion of lignocellulosic biomass. Bioresour Technol 344:126325. https://doi.org/10.1016/j.biortech.2021.126325
Greetham D, Zaky AS, Du C (2019) Exploring the tolerance of marine yeast to inhibitory compounds for improving bioethanol production. Sustain Energy Fuels 3:1545–1553. https://doi.org/10.1039/c9se00029a
Zaky AS, Greetham D, Tucker GA, Du C (2018) The establishment of a marine focused biorefinery for bioethanol production using seawater and a novel marine yeast strain. Sci Rep 8:12127. https://doi.org/10.1038/s41598-018-30660-x
Zaky AS, French CE, Tucker GA, Du C (2020) Improving the productivity of bioethanol production using marine yeast and seawater-based media. Biomass Bioenergy 139:105615. https://doi.org/10.1016/j.biombioe.2020.105615
Basso TO, Basso TP, LuizAlvesJúnior S, Stambuk BU, Basso LC (2022) Saccharomyces: the 5 Ws and one H. In: LuizAlvesJúnior S, Treichel H, Basso T, Stambuk BU (eds) Yeasts: from nature to bioprocesses. Bentham Science Publishers, Sharjah, pp 73–112. https://doi.org/10.2174/9789815051063122020006
Della-Bianca BE, Basso TO, Stambuk BU, Basso LC, Gombert AK (2013) What do we know about the yeast strains from the Brazilian fuel ethanol industry? Appl Microbiol Biotechnol 97:979–991. https://doi.org/10.1007/s00253-012-4631-x
Basso LC, de Amorim HV, de Oliveira AJ, Lopes ML (2008) Yeast selection for fuel ethanol production in Brazil. FEMS Yeast Res 8:1155–1163. https://doi.org/10.1111/j.1567-1364.2008.00428.x
Matsushika A, Inoue H, Kodaki T, Sawayama S (2009) Ethanol production from xylose in engineered Saccharomyces cerevisiae strains: current state and perspectives. Appl Microbiol Biotechnol 84:37–53. https://doi.org/10.1007/s00253-009-2101-x
Pereira IO, dos Santos ÂA, Gonçalves DL, Purificação M, Guimarães NC, Tramontina R, Coutouné N, Zanella E, Matsushika A, Stambuk BU, Ienczak JL (2021) Comparison of Spathaspora passalidarum and recombinant Saccharomyces cerevisiae for integration of first- and second-generation ethanol production. FEMS Yeast Res 21:foab048. https://doi.org/10.1093/femsyr/foab048
de Sales BB, Scheid B, Gonçalves DL, Knychala MM, Matsushika A, Bon EPS, Stambuk BU (2015) Cloning novel sugar transporters from Scheffersomyces (Pichia) stipitis allowing d-xylose fermentation by recombinant Saccharomyces cerevisiae. Biotechnol Lett 37:1973–1982. https://doi.org/10.1007/S10529-015-1893-2/TABLES/3
Gietz D, Jean AS, Woods RA, Schiestl RH (1992) Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res 20:1425–1425. https://doi.org/10.1093/NAR/20.6.1425
Matsushika A, Goshima T, Fujii T, Inoue H, Sawayama S, Yano S (2012) Characterization of non-oxidative transaldolase and transketolase enzymes in the pentose phosphate pathway with regard to xylose utilization by recombinant Saccharomyces cerevisiae. Enzyme Microb Technol 51:16–25. https://doi.org/10.1016/J.ENZMICTEC.2012.03.008
Walfridsson M, Hallborn J, Penttila M, Keranen S, Hahn-Hagerdal B (1995) Xylose-metabolizing Saccharomyces cerevisiae strains overexpressing the TKL1 and TAL1 genes encoding the pentose phosphate pathway enzymes transketolase and transaldolase. Appl Environ Microbiol 61:4184–4190. https://doi.org/10.1128/AEM.61.12.4184-4190.1995
Petracek ME, Longtine MS (2002) PCR-based engineering of yeast genome. Methods Enzymol 350:445–469. https://doi.org/10.1016/S0076-6879(02)50978-2
Rodrigues MA, Iemma AF (2014) Experimental design and process optimization, 1st edn. CRC Press, Boca Raton
Barrilli ÉT, Tadioto V, Milani LM, Deoti JR, Fogolari O, Müller C, Barros KO, Rosa CA, dos Santos AA, Stambuk BU, Treichel H, Alves SL (2020) Biochemical analysis of cellobiose catabolism in Candida pseudointermedia strains isolated from rotten wood. Arch Microbiol 202:1729–1739. https://doi.org/10.1007/s00203-020-01884-1
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protocols 3:1101–1108. https://doi.org/10.1038/nprot.2008.73
Zanivan J, Bonatto C, Scapini T, Dalastra C, Bazoti SF, Júnior SLA, Fongaro G, Treichel H (2022) Evaluation of bioethanol production from a mixed fruit waste by Wickerhamomyces sp. UFFS-CE-3.1.2. Bioenergy Res 15:175–182. https://doi.org/10.1007/s12155-021-10273-5
Bonatto C, Scapini T, Zanivan J, Dalastra C, Bazoti SF, Alves S, Fongaro G, de Oliveira D, Treichel H (2021) Utilization of seawater and wastewater from shrimp production in the fermentation of papaya residues to ethanol. Bioresour Technol 321:124501. https://doi.org/10.1016/j.biortech.2020.124501
Babrzadeh F, Jalili R, Wang C, Shokralla S, Pierce S, Robinson-Mosher A, Nyren P, Shafer RW, Basso LC, de Amorim HV, de Oliveira AJ, Davis RW, Ronaghi M, Gharizadeh B, Stambuk BU (2012) Whole-genome sequencing of the efficient industrial fuel-ethanol fermentative Saccharomyces cerevisiae strain CAT-1. Mol Genet Genomics 287:485–494. https://doi.org/10.1007/S00438-012-0695-7/FIGURES/3
Kayikci Ö, Nielsen J (2015) Glucose repression in Saccharomyces cerevisiae. FEMS Yeast Res 15:68. https://doi.org/10.1093/FEMSYR/FOV068
Lane S, Xu H, Oh EJ, Kim H, Lesmana A, Jeong D, Zhang G, Tsai CS, Jin YS, Kim SR (2018) Glucose repression can be alleviated by reducing glucose phosphorylation rate in Saccharomyces cerevisiae. Sci Rep 8:2613. https://doi.org/10.1038/s41598-018-20804-4
Gao M, Ploessl D, Shao Z (2019) Enhancing the co-utilization of biomass-derived mixed sugars by yeasts. Front Microbiol 10:3264. https://doi.org/10.3389/FMICB.2018.03264/BIBTEX
Chomvong K, Kordić V, Li X, Bauer S, Gillespie AE, Ha SJ, Oh EJ, Galazka JM, Jin YS, Cate JHD (2014) Overcoming inefficient cellobiose fermentation by cellobiose phosphorylase in the presence of xylose. Biotechnol Biofuels 7:85. https://doi.org/10.1186/1754-6834-7-85/FIGURES/7
Kim SR, Ha S-J, Wei N, Oh EJ, Jin Y-S (2012) Simultaneous co-fermentation of mixed sugars: a promising strategy for producing cellulosic ethanol. Trends Biotechnol 30:274–282. https://doi.org/10.1016/j.tibtech.2012.01.005
Palmqvist E, Hahn-Hägerdal B (2000) Fermentation of lignocellulosic hydrolysates. II: inhibitors and mechanisms of inhibition. Bioresour Technol 74:25–33. https://doi.org/10.1016/S0960-8524(99)00161-3
Sato TK, Liu T, Parreiras LS, Williams DL, Wohlbach DJ, Bice BD, Ong IM, Breuer RJ, Qin L, Busalacchi D, Deshpande S, Daum C, Gasch AP, Hodge DB (2014) Harnessing genetic diversity in Saccharomyces cerevisiae for fermentation of xylose in hydrolysates of alkaline hydrogen peroxide-pretreated biomass. Appl Environ Microbiol 80:540–554. https://doi.org/10.1128/AEM.01885-13/SUPPL_FILE/ZAM999105036SO1.PDF
Casey E, Sedlak M, Ho NWY, Mosier NS (2010) Effect of acetic acid and pH on the cofermentation of glucose and xylose to ethanol by a genetically engineered strain of Saccharomyces cerevisiae. FEMS Yeast Res 10:385–393. https://doi.org/10.1111/J.1567-1364.2010.00623.X
Palma M, Guerreiro JF, Sá-Correia I (2018) Adaptive response and tolerance to acetic acid in Saccharomyces cerevisiae and Zygosaccharomyces bailii: a physiological genomics perspective. Front Microbiol 9:274. https://doi.org/10.3389/FMICB.2018.00274/BIBTEX
Zhang K, Wells P, Liang Y, Love J, Parker DA, Botella C (2019) Effect of diluted hydrolysate as yeast propagation medium on ethanol production. Bioresour Technol 271:1–8. https://doi.org/10.1016/J.BIORTECH.2018.09.080
Cunha JT, Costa CE, Ferraz L, Romaní A, Johansson B, Sá-Correia I, Domingues L (2018) HAA1 and PRS3 overexpression boosts yeast tolerance towards acetic acid improving xylose or glucose consumption: unravelling the underlying mechanisms. Appl Microbiol Biotechnol 102:4589–4600. https://doi.org/10.1007/s00253-018-8955-z
Sakihama Y, Hasunuma T, Kondo A (2015) Improved ethanol production from xylose in the presence of acetic acid by the overexpression of the HAA1 gene in Saccharomyces cerevisiae. J Biosci Bioeng 119:297–302. https://doi.org/10.1016/j.jbiosc.2014.09.004
Bedő S, Fehér A, Khunnonkwao P, Jantama K, Fehér C (2021) Optimized bioconversion of xylose derived from pre-treated crop residues into xylitol by using Candida boidinii. Agronomy 11:79. https://doi.org/10.3390/agronomy11010079
Raj K, Krishnan C (2020) Improved co-production of ethanol and xylitol from low-temperature aqueous ammonia pretreated sugarcane bagasse using two-stage high solids enzymatic hydrolysis and Candida tropicalis. Renew Energy 153:392–403. https://doi.org/10.1016/j.renene.2020.02.042
Radecka D, Mukherjee V, Mateo RQ, Stojiljkovic M, Foulquié-Moreno MR, Thevelein JM (2015) Looking beyond Saccharomyces: the potential of non-conventional yeast species for desirable traits in bioethanol fermentation. FEMS Yeast Res 15:fov053. https://doi.org/10.1093/femsyr/fov053
Mukherjee V, Radecka D, Aerts G, Verstrepen KJ, Lievens B, Thevelein JM (2017) Phenotypic landscape of non-conventional yeast species for different stress tolerance traits desirable in bioethanol fermentation. Biotechnol Biofuels 10:216. https://doi.org/10.1186/S13068-017-0899-5
Geijer C, Ledesma-Amaro R, Tomas-Pejo E (2022) Unraveling the potential of non-conventional yeasts in biotechnology. FEMS Yeast Res 22:foab71. https://doi.org/10.1093/femsyr/foab071
Wang L, Li B, Su RR, Wang SP, Xia ZY, Xie CY, Tang YQ (2022) Screening novel genes by a comprehensive strategy to construct multiple stress-tolerant industrial Saccharomyces cerevisiae with prominent bioethanol production. Biotechnol Biofuels Bioproducts 15:1. https://doi.org/10.1186/S13068-022-02109-X
Sun Y, Kong M, Li X, Li Q, Xue Q, Hou J, Jia Z, Lei Z, Xiao W, Shi S, Cao L (2022) Metabolic and evolutionary engineering of diploid yeast for the production of first- and second-generation ethanol. Front Bioeng Biotechnol 9:1527. https://doi.org/10.3389/fbioe.2021.835928/bibtex
Patiño MA, Ortiz JP, Velásquez M, Stambuk BU (2019) d-Xylose consumption by nonrecombinant Saccharomyces cerevisiae: a review. Yeast 36:541–546. https://doi.org/10.1002/yea.3429
Chen Y, Stabryla L, Wei N (2016) Improved acetic acid resistance in Saccharomyces cerevisiae by overexpression of the WHI2 gene identified through inverse metabolic engineering. Appl Environ Microbiol 82:2156–2166. https://doi.org/10.1128/aem.03718-15
Kolouchová I, Schreiberová O, Sigler K, Masák J, Řezanka T (2015) Biotransformation of volatile fatty acids by oleaginous and non-oleaginous yeast species. FEMS Yeast Res 15:fov076. https://doi.org/10.1093/femsyr/fov076
Gerós H, Cássio F, Leão C (2000) Utilization and transport of acetic acid in Dekkera anomala and their implications on the survival of the yeast in acidic environments. J Food Prot 63:96–101. https://doi.org/10.4315/0362-028X-63.1.96
Zaky AS (2021) Introducing a marine biorefinery system for the integrated production of biofuels, high-value-chemicals, and co-products: a path forward to a sustainable future. Processes 9:1841. https://doi.org/10.3390/pr9101841
Franco-Duarte R, Mendes I, Umek L, Drumonde-Neves J, Zupan B, Schuller D (2014) Computational models reveal genotype-phenotype associations in Saccharomyces cerevisiae. Yeast 31:265–277. https://doi.org/10.1002/yea.3016
Zaky AS, Tucker GA, Daw ZY, Du C (2014) Marine yeast isolation and industrial application. FEMS Yeast Res 14:813–825. https://doi.org/10.1111/1567-1364.12158
Zaky AS, Greetham D, Louis EJ, Tucker GA, Du C (2016) A new isolation and evaluation method for marine-derived yeast spp. with potential applications in industrial biotechnology. J Microbiol Biotechnol 26:1891–1907. https://doi.org/10.4014/jmb.1605.05074
Acknowledgements
This work was supported in part by grants and fellowships from the Brazilian agencies National Council for Scientific and Technological Development (CNPq, Grant numbers 490029/2009-4, 551392/2010-0, 307290/2012-3, 478841/2013-2, 308627/2015-6, 420480/2018-8, 308389/2019-0, and 305258/2018-4); Financier of Studies and Projects (FINEP, Grant number 01.09.0566.00/1421-08); Research and Innovation Funding Agency of the State of Santa Catarina (FAPESC, Grant numbers 17293/2009-6, 749/2016-TO2016TR2188); VTT Brasil—Pesquisa e Desenvolvimento Ltda. (VTT-FAPEU, process no. 096/2014); Coordination for the Improvement of Higher Education Personnel (CAPES/PNPD, no. 88887.352933/2019-00); the Research Promotion Program and the Support Program for Scientific and Technological Initiation from the Federal University of Fronteira Sul (Grant numbers PES-2018-0945, PES-2019-0638, PES-2020-0213, PES-2021-0387); and by the Japanese International Cooperation Agency (JICA).
Author information
Authors and Affiliations
Contributions
VT, CM, AG, LD, and ACL carried out the experimental design assays and the fermentation kinetic analysis. JRD, BRS, and MP engineered the S. cerevisiae strains. OF carried out the HPLC analysis. AM, HT, BUS, and SLAJ participated in designing the study and provided financial support. SLAJ also wrote the manuscript, which was revised and approved by all authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tadioto, V., Deoti, J.R., Müller, C. et al. Prospecting and engineering yeasts for ethanol production under inhibitory conditions: an experimental design analysis. Bioprocess Biosyst Eng 46, 1133–1145 (2023). https://doi.org/10.1007/s00449-022-02812-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-022-02812-x