Abstract
Main conclusion
Light and ABA independently regulated anthocyanin biosynthesis via activation of FaMYB10 expression. FaMYB10 accelerated anthocyanin synthesis of pelargonidin 3-glucoside and cyanidin 3-glucoside during strawberry fruit ripening.
Light is an integral factor in fruit ripening. Ripening in non-climacteric fruit is also effected by the plant hormone abscisic acid (ABA). However, how light and/or ABA regulate fruit ripening processes, such as strawberry color development remains elusive. Results of the present study showed light and ABA regulated strawberry fruit coloration via activation of FaMYB10 expression, an R2R3 MYB transcription factor. Light exposure increased FaMYB10 transcript levels, flavonoid pathway genes, and anthocyanin content. Exogenous ABA promoted FaMYB10 expression, and anthocyanin content, accompanied by increased ABA-responsive transcript levels and flavonoid pathway genes. ABA biosynthesis inhibitor treatment, and RNAi-mediated down-regulation of the ABA biosynthetic gene (9-cis epoxycarotenoid dioxygenase: FaNCED1), and ABA receptor (magnesium chelatase H subunit: FaCHLH/ABAR) showed inverse ABA effects. Furthermore, additive effects were observed in anthocyanin accumulation under combined light and ABA, indicating independent light and ABA signaling pathways. FaMYB10 down-regulation by Agrobacterium-mediated RNA interference (RNAi) in strawberry fruits showed decreased pelargonidin 3-glucoside and cyanidin 3-glucoside levels, accompanied by consistent flavonoid pathway gene expression levels. FaMYB10 over-expression showed opposite FaMYB10 RNAi phenotypes, particularly cyanidin 3-glucoside synthesis by FaMYB10, which was correlated with FaF3′H transcript levels. These data provided evidence that light and ABA promoted FaMYB10 expression, resulting in anthocyanin accumulation via acceleration of flavonoid pathway gene expression. Finally, our results suggested FaMYB10 serves a role as a signal transduction mediator from light and ABA perception to anthocyanin synthesis in strawberry fruit.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fruit ripening is a complex, genetically programmed progression that culminates in notable changes in fresh fruit color, texture, flavor, and aroma. These processes have been extensively studied at biochemical and genetic levels in many fruit crops. Fruits with different ripening mechanisms are divided into two groups, climacteric and non-climacteric. In climacteric fruit, a peak in respiration accompanies ripening, with a concomitant ethylene burst. In non-climacteric fruit, a marked change in respiration has not been observed, and ethylene production remains at very low levels (Alexander and Grierson 2002). Klee and Giovannoni (2011) reviewed the molecular mechanisms of ethylene action in climacteric fruits using tomato as a model, and reported the ethylene surge was required for normal fruit ripening. Furthermore, other studies showed ethylene affected transcription and translation of many ripening-related genes (Deikman 1997; Giovannoni 2004; Gapper et al. 2013). However, identity of the signal and/or signals responsible for the transduction mechanism in non-climacteric fruit ripening remains largely unresolved.
Several studies associated the plant hormone abscisic acid (ABA) with regulation of non-climacteric fruit ripening (Davies et al. 1997; Rodrigo et al. 2003; Zhang et al. 2009a; Symons et al. 2012; Karppinen et al. 2013). Several studies demonstrated ABA integral in various plant roles, including plant growth and development, response to environmental stressors (Finkelstein et al. 2002; Yamaguchi-Shinozaki and Shinozaki 2006; Hirayama and Shinozaki 2007; Nakashima and Yamaguchi-Shinozaki 2013), and fruit sugar metabolism and accumulation (Kobashi et al. 1999; Pan et al. 2005; Rook et al. 2006; Jia et al. 2013a). A key enzyme in ABA biosynthesis, 9-cis epoxycarotenoid dioxygenase (NCED) (Qin and Zeevaart 1999) was shown to ripen several fruits, including avocado (Chernys and Zeevaart 2007), orange (Rodrigo et al. 2006), grape, peach (Zhang et al. 2009a), and tomato (Zhang et al. 2009b). Strawberry (Fragaria × ananassa) is widely applied as a research model to study non-climacteric fruits. It has also been associated with ABA to induce strawberry ripening (Kano and Asahira 1981; Jiang and Joyce 2003; Li et al. 2011). Jia et al. (2011) and Chai et al. (2011) reported down-regulation of NCED1, magnesium chelatase H subunit (CHLH/ABAR), and pyrabactin resistance (PYR1), each ABA receptors, resulted in unpigmented strawberry fruit phenotypes. Furthermore, ABA-treated fruits exhibited accelerated development and coloration. Recently, Jia et al. (2013b) characterized ABI1, a type 2C protein phosphatase involved in ABA signaling, also functioned as a negative regulator of strawberry fruit ripening. However, the signal transduction molecular mechanism from ABA perception to coloration remains elusive.
Anthocyanins are major pigments in many plant groups. This pigment class belongs to a parent class of molecules called flavonoids, which are also important secondary metabolites that serve as key contributors to the antioxidant capacity and nutritional value in strawberry fruit (Lopes-da-Silva et al. 2007; He and Giusti 2010). Anthocyanin biosynthesis is highly regulated by light quantity and quality (Cominelli et al. 2008; Kami et al. 2010), and in strawberry, blue light is most effective in anthocyanin biosynthesis induction. Two cryptochromes (FaCRY1 and FaCRY2) and two phototropins (FaPHOT1 and FaPHOT2) serve as blue light receptors in strawberry. Kadomura-Ishikawa et al. (2013) reported FaPHOT2 in strawberry fruit was in part associated with blue light-induced anthocyanin biosynthesis. However, little is known regarding signal transduction molecular mechanisms from fruit light perception to coloration.
In plants, the flavonoid biosynthetic pathway (Fig. 1) structural genes are largely regulated at the transcriptional level (Cominelli et al. 2008; Miyawaki et al. 2012). In all plant species studied to date, regulation of flavonoid pathway gene expression is through an MYB transcription factor (TF) complex, basic helix–loop–helix (bHLH) TFs, and WD-repeat protein (the MYB-bHLH-WD40 “MBW” complex; Baudry et al. 2004). Koes et al. (2005) proposed a flavonoid pathway gene activation model, with regulator interaction forming transcriptional complexes in conjunction with structural gene promoters. R2R3 MYBs are well-known anthocyanin biosynthesis regulators. For example, Borevitz et al. (2000) showed AtPAP1 gene (AtMYB75, At1g56650) over-expression resulted in anthocyanin accumulation in Arabidopsis. Several phenylpropanoid pathway repressors, and perhaps anthocyanins specifically, are also MYB TFs, including a R2R3 MYB repressor from strawberry FaMYB1 (Aharoni et al. 2001), Arabidopsis AtMYB4 (Jin et al. 2000), Antirrhinum AmMYB308 (Tamagnone et al. 1998), and a one repeat MYB in Arabidopsis AtMYBL2 (Dubos et al. 2008; Matsui et al. 2008). Lin-Wang et al. (2010) reported FaMYB10 over-expression exhibited high strawberry fruit anthocyanin accumulation. Recently, Medina-Puche et al. (2014) reported FaMYB10 down-regulation decreased anthocyanin levels in strawberry fruit. Furthermore, ABA and the plant hormone auxin regulated FaMYB10 expression in strawberry fruit (Daminato et al. 2013; Medina-Puche et al. 2014).
Flavonoid biosynthesis during strawberry fruit developmental and ripening stages. Flavonoid biosynthetic pathway in strawberry plants. Enzymes for each step are shown in bold, and include the following: chalcone synthase (CHS), chalcone isomerase (CHI), flavanone 3-hydroxylase (F3H), flavonoid 3′-hydroxylase (F3′H), dihydroflavonol-4-reductase (DFR), anthocyanidin synthase (ANS), and flavonoid glycosyltransferases (FGTs)
In the present study, we showed FaMYB10 regulated flavonoid pathway gene expression by FaMYB10 gene over-expression or down-regulation in strawberry fruits. Furthermore, we showed that FaMYB10 gene expression was independently regulated by light and ABA.
Materials and methods
Strawberry fruit samples
Octaploid strawberries (Fragaria × ananassa, cv. Sachinoka; plants obtained from Miyoshi & Co. Ltd., Tokyo, Japan) were cultivated under greenhouse conditions (from winter to spring). Strawberry fruits were harvested at all six stages: small green (SG), medium green (MG), big green (BG), white (Wh), turning (Tu), and red (Re) (Fait et al. 2008). Harvested fruits were subsequently sterilized according to Miyawaki et al. (2012), placed on plates, and maintained at 23 °C in a MLR-351 growth chamber (Sanyo, Osaka, Japan).
Light treatment
Harvested Wh stage strawberry fruits were placed in a plant box for white light time course experiments, and maintained at 23 °C under a 16 h light/8 h dark irradiation photocycle for 8 days. Fluorescent lights (FL40SS W, Panasonic, 110 μmol m−2 s−1) provided white light in a growth chamber (MLR-351, Sanyo).
ABA and ABA synthesis inhibitor treatments
Harvested Wh stage strawberry fruits were treated with ABA (Sigma, Osaka, Japan) or ABA synthetic inhibitor (fluridone, Fluka, Osaka, Japan) following Chai et al. (2011) with several modifications. A 200 μl aliquot of 50 μM ABA or 500 μM fluridone was injected into harvested Wh stage strawberry fruits on alternate days (days 0, 2, and 4; a total of three times during the 8 days) using a 1-ml syringe. Distilled water was used as a control. Treated fruits were placed in a plant box, and maintained at 23 °C under a 16 h light/8 h dark photocycle, or a dark only control. Fluorescent lights (FL40SS W, Panasonic, 110 μmol m−2 s−1) provided white light in a growth chamber (MLR-351, Sanyo).
Extraction and determination of anthocyanins
Spectrophotometry was used to measure total anthocyanin content according to Kortstee et al. (2011). Briefly, achenes were removed; each receptacle was cut into pieces (1–2 g each), and soaked in 5–10 volumes of methanol containing 1 % (v/v) hydrochloric acid overnight. Solutions were centrifuged at 3,800g for 10 min, supernatants collected, and centrifuged a second time at 20,400g for 5 min. Supernatants were subsequently transferred to fresh tubes, and absorbance measured at 530 and 657 nm to correct for chlorophyll content. Pelargonidin 3-glucoside (Funakoshi, Tokyo, Japan) was used as the standard.
Cyanidin 3-glucoside and pelargonidin 3-glucoside content calculations
Anthocyanin (cyanidin 3-glucoside and pelargonidin 3-glucoside) calculations were performed following calibration curve construction. Pelargonidin 3-glucoside and cyanidin 3-glucoside were purchased from Funakoshi. De-achened frozen samples were powdered with liquid nitrogen in a mortar for anthocyanin extraction. Anthocyanins were extracted with methanol containing 0.1 % (v/v) hydrochloric acid overnight from 2 g powdered samples. Extracted anthocyanins were centrifuged at 3,800g for 10 min; supernatants collected, centrifuged a second time at 20,400g for 10 min, and filtered through a 0.45-μm filter disc.
Anthocyanin analyses were conducted using an LC-20AD low-pressure gradient system (Simadzu, Kyoto, Japan) equipped with SPD-M20A photodiode array detector and manual injector (10 μl injection volume). A DGU-20A3 degasser was employed for analytical HPLC. HPLC data were analyzed using LC solution software. Separations were performed in a 4.6 mm i.d. × 250 mm i.d., 5 μm, reverse phase C18 analytical column (Nacalai Tesque, Kyoto, Japan) operating at room temperature with a flow rate of 500 μl/min. Anthocyanins were quantified between 280 and 600 nm wavelengths, and monitored at 510 and 520 nm. Elution was performed using the following solvent gradient: 0.1 % trifluoroacetic acid (solvent A) and methanol (solvent B). The gradient was a mixture of 20 % B and 80 % A, up to 25.0 min; and 100 % B, up to 35.0 min. Components were identified by comparisons of their retention times to authentic standard times under the same analysis conditions.
RNA extraction and cDNA synthesis
Achenes were removed from receptacles prior to RNA isolation. Total RNA was isolated using an RNAqueous kit with Plant RNA Isolation Aid (Ambion, Carlsbad, CA, USA). The total RNA obtained was treated with DNase (TURBO DNA-free™ Kit, Ambion) following the manufacturer’s instructions, to remove any genomic DNA contamination. RNA samples were considered DNA free when amplicons corresponding to analyzed DNA genes were not observed, using RNA as a template under quantitative real-time reverse transcription polymerase chain reaction (qRT-PCR) conditions. First-strand cDNA was synthesized from 1 μg total RNA using a Superscript III First-Strand Synthesis System (Invitrogen, Tokyo, Japan) using recommended oligo (dT)20 primers following the manufacturer’s instructions.
qRT-PCR analysis
qRT-PCR was performed using an ABI 7900HT system (Applied Biosystems, Tokyo, Japan), and gene-specific primers (GSPs). We identified two dihydroflavonol-4-reductase (DFR) genes in strawberry (GenBank No. AY695812 and strawberry gene ID: GENE29482, not deposited in GenBank), named AY695812 and GENE29482 as FaDFR1 and FaDFR2, respectively. FaMYB10, FaABI3, FaNCED1, FaCHLH/ABAR, and flavonoid pathway gene primer sequences, with the exception of FaDFR2 were based on previous reports (Hoffmann et al. 2006; Almeida et al. 2007; Salvatierra et al. 2010; Chai et al. 2011; Jia et al. 2011; Miyawaki et al. 2012; Kadomura-Ishikawa et al. 2013). The FaDFR2 gene primer sequences were designed based on GENE29482. Primer sequences are listed in Supplemental Table S1. The glyceraldehyde-3-phosphate dehydrogenase (GAPDH) gene (GenBank No. AB363963) was used as the internal control. PCR assays included the following reaction mixture: 1 ng cDNA, 200 nM each primer, and 5 μl 2× Power SYBR Green PCR Master Mix (Applied Biosystems). PCR reactions were conducted under the following conditions: 95 °C for 10 min; 40 cycles of 95 °C for 15 s, and 60 °C for 1 min. The target quantity in each sample was normalized to a reference control based on Salvatierra et al. (2010) using the comparative (\(2^{{ - \Delta \Delta C_{\text{t}} }}\)) method described by the manufacturer. Efficiency of each qRT-PCR, and product melting curves were also analyzed to ensure a single amplification peak corresponding to a unique molecular species. Furthermore, we confirmed a single PCR product by gel electrophoresis.
Plasmid construction for RNA interference (RNAi), over-expression assays, and Agrobacterium-infiltration
Plasmid construction was conducted following Kadomura-Ishikawa et al. (2013), with the exception of target gene fragment amplification. FaMYB10 (GenBank No. EU155162; Lin-Wang et al. 2010), FaNCED1 (GenBank No. HQ290318; Jia et al. 2011), and FaCHLH/ABAR (GenBank No. GQ201451; Jia et al. 2011) fragments were amplified using sense (s), and anti-sense (a) primers (listed in Supplemental Table S2) using cDNA isolated from receptacles. Amplified products were ligated into a pRI-intron vector derived from pRI 201-AN (Takara, Shiga, Japan), containing a strawberry quinone oxidoreductase gene intron (GenBank No. AY158836). The resulting constructs were named, pRI-myb10-ihpRNAi (pRI-myb10i), pRI-nced1-ihpRNAi (pRI-nced1i), and pRI-chlh/abar-ihpRNA (pRI-chlhi), respectively.
The FaMYB10 open reading frame (ORF) was amplified by receptacle cDNA, and PCR primers were designed from GenBank sequence data (Supplemental Table S2) for over-expression analysis. PCR products were cloned into a pGEM-T Easy Vector system (Promega, Madison, WI, USA). FaMYB10 ORF was cleaved from the pGEM-T Easy Vector, and ligated into the pRI-201-AN vector. The resulting construct was named pRI-MYB10-overexpression (pRI-MYB10-OE). Each construct sequence was verified using a BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems), and an ABI PRISM 310 Genetic Analyzer (Applied Biosystems).
A pRI-intron, pRI-myb10i, pRI-nced1i, pRI-chlhi, or pRI-MYB10-OE construct was introduced into Agrobacterium tumefaciens strain GV2260, and strawberry transformation was conducted using an Agrobacterium-infiltration method following Hoffmann et al. (2006) and Miyawaki et al. (2012). The Agrobacterium suspension harboring each construct was injected into strawberry fruit on day 0, and subsequently on days 2 and 4. pRI-intron-infiltrated fruit were used as experimental controls. These treated fruits were placed in a plant box, and maintained at 23 °C under a 16 h light/8 h dark irradiation photocycle for 7 days. Fluorescent lights (FL40SS W, Panasonic, 110 μmol m−2 s−1) provided white light in a growth chamber (MLR-351, Sanyo).
Statistical analyses
Data were expressed as mean ± standard deviation (SD), using IBM SPSS Statistics software v 19 for analyses.
Results
Light effects on FaMYB10 expression in strawberry fruit
In a recent study, we showed light, particularly blue light induced coloration in strawberry fruit, and one of the blue light photoreceptors, FaPHOT2, was involved in this process (Miyawaki et al. 2012; Kadomura-Ishikawa et al. 2013). In the present study, we initially assessed light effects on anthocyanin accumulation by measuring anthocyanin content in light-treated fruits. Wh stage harvested strawberry fruits were treated to a 16 h light/8 h dark cycle for 8 days. Harvested fruits without light treatment were used as a dark control. Anthocyanin content in light-treated fruit significantly increased compared to the dark control at days 4 (P < 0.01), 6 (P < 0.01), and 8 (P < 0.001) (Fig. 2a).
Light effects on FaMYB10 expression in strawberry fruits. Harvested Wh stage strawberry fruits were exposed to 16 h light (110 μmol m−2 s−1), followed by 8 h dark. The photocycle continued for 8 days. Dark control fruits were maintained in the same growth chamber in the absence of light for 8 days. a Anthocyanin content was measured for light (white bars) and dark control fruits (black bars) (n = 5). FaCHS (b) and FaMYB10 (c) were detected using qRT-PCR. Relative expression levels for each gene were normalized to FaGAPDH levels. In addition, normalized expression levels for each gene with light treatment at day 0 were set to 1. Data are expressed as mean ± SD (n = 3–5). Asterisks indicate significant differences between light-treated and dark control samples (Student’s t test; *P < 0.05, **P < 0.01, and ***P < 0.001)
We subsequently performed qRT-PCR analysis of the flavonoid pathway gene (FaCHS) and FaMYB10 in light-treated, and dark control fruits. FaCHS in light-treated fruits maintained higher expression levels compared to dark control fruits throughout the experimental period (0–8 days). FaCHS transcript levels increased to the highest levels in light-treated fruit on day 4, but remained significantly lower in the dark control (P < 0.01). At day 6, light-treated fruit showed a decrease, and the dark control exhibited an increase; however, FaCHS transcript levels remained significantly higher (P < 0.05). Finally, at day 8, light-treated and dark control fruits showed decreased expression levels; however, significant differences were not detected (Fig. 2b). Consistent with FaCHS transcript level results, FaMYB10 in light-treated fruits exhibited higher expression levels compared to dark control fruits throughout the experimental period (0–8 days). However, dark control fruits exhibited a continued increase in FaMYB10 transcript levels from days 2 to 8. Light-treated and dark control fruits showed the lowest increase in FaMYB10 transcript levels at day 2, and significant differences were observed between expression levels (P < 0.05) (Fig. 2c). FaMYB10 transcript levels under light conditions increased to the highest levels detected during the experiment at day 4, which were significantly higher than the dark control (P < 0.001). Fruit transcript levels under light-treatment conditions decreased as dark control levels increased from days 6 to 8, and significant differences were observed in FaMYB10 transcript levels (day 6 and day 8; P < 0.01 and P < 0.05, respectively) (Fig. 2c).
ABA effect on FaMYB10 expression in strawberry fruit
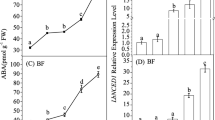
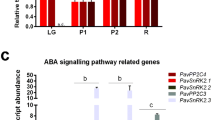
Former studies reported ABA was a key regulator in strawberry fruit ripening (Jia et al. 2011, 2013b; Chai et al. 2011). The relationship between ABA and FaMYB10 in strawberry fruit was characterized by analyzing the down-regulation effects of the ABA biosynthesis gene (FaNCED1), and the ABA receptor (FaCHLH/ABAR) by in planta RNAi. FaNCED1 expression levels in pRI-nced1i-infiltrated (Fanced1 AmRNAi fruits) and pRI-chlhi-infiltrated fruits (Fachlh AmRNAi fruits) were found 1.7-fold lower (P < 0.001), and no change relative to vector control fruits was identified, respectively (Fig. 3a). FaCHLH/ABAR expression levels in Fachlh AmRNAi and Fanced1 AmRNAi fruits were found 1.6-fold lower (P < 0.01) and 1.6-fold higher (P < 0.01) relative to vector control fruits, respectively (Fig. 3b). FaABI3 (ABA-responsive gene), FaCHS, and FaMYB10 transcript levels exhibited a significant decrease in Fachlh AmRNAi (P values ranged from <0.05 to 0.01) and Fanced1 AmRNAi fruits (all genes; P < 0.01) (Fig. 3c–e). Fachlh AmRNAi and Fanced1 AmRNAi fruits showed anthocyanin reductions compared to the vector control. Total anthocyanin concentrations in Fachlh AmRNAi (295.2 ± 23.1 μg/g fresh weight), and Fanced1 AmRNAi fruits (265.4 ± 41.8 μg/g) were 1.2-fold (P < 0.001) or 1.4-fold (P < 0.001) lower relative to control fruits (366.5 ± 33.2 μg/g fresh weight), respectively (Fig. 3f).
FaNCED1 or FaCHLH/ABAR down-regulation effects on anthocyanin content, and FaMYB10 and flavonoid pathway gene expression levels in strawberry fruits. FaNCED1 (a), FaCHLE/ABAR (b), FaABI3 (c), FaCHS (d) and FaMYB10 (e) were detected by qRT-PCR in Fanced1 AmRNAi (black bars), Fachlh AmRNAi (gray bars), and fruits infiltrated with a control vector (white bars). Relative expression levels for each gene were normalized to FaGAPDH levels. In addition, normalized expression of each gene in the control vector-infiltrated fruits was set to 1. Data are expressed as mean ± SD (n = 3–6). f Anthocyanin content was measured for Fanced1 AmRNAi (black bars), Fachlh AmRNAi (gray bars), and control vector-infiltrated fruits (white bars) (n = 6–10). Asterisks indicate significant differences among Fanced1 AmRNAi or Fachlh AmRNAi fruits, and control vector-infiltrated fruits (Student’s t test; *P < 0.05, **P < 0.01, and ***P < 0.001)
ABA effects on strawberry fruit coloration, and the relationship between ABA and FaMYB10 in strawberry fruit was confirmed by injecting ABA (50 μM) or ABA synthesis inhibitor (fluridone, 500 μM) into Wh stage harvested strawberry fruits, which were subsequently treated to the presence or absence of light. We confirmed light conditions induced an increase in anthocyanin content compared to dark conditions (Table 1). A significant increase in anthocyanin accumulation (P < 0.001) was detected for ABA treatments under light conditions compared to the dark control. Fluridone treatments, however, significantly decreased anthocyanin content under both light and dark conditions (P < 0.001). The decreased anthocyanin content resulting from fluridone treatment was recovered by light, and anthocyanin reached levels comparable to dark control conditions.
The relationship between ABA and anthocyanin accumulation was examined by performing qRT-PCR analysis of the following genes: FaABI3, FaCHS, and FaMYB10 in treated strawberry fruits. FaABI3 transcript levels were, respectively, increased and decreased by ABA- and fluridone-treated fruits (Fig. 4a). FaCHS and FaMYB10 transcript levels were positively associated with anthocyanin accumulation, and increased and decreased by ABA- and fluridone-treated fruits, respectively (Fig. 4b, c). We also evaluated FaANS and FaFGT (flavonoid pathway genes) transcript levels, and results showed expression patterns of these genes were similar to FaCHS and FaMYB10, although significant differences compared to control fruits were not detected (Supplemental Fig. S1).
ABA or ABA synthesis inhibitor effects on FaMYB10 and flavonoid pathway gene expression levels in strawberry fruits. FaABI3 (a), FaCHS (b) and FaMYB10 (c) were detected using qRT-PCR for ABA 50 μM or fluridone 500 μM injected fruits. Relative expression levels for each gene were normalized to FaGAPDH levels. In addition, normalized expression of dark control fruits was set to 1. Data are expressed as mean ± SD (n = 3–5). Black and white bars depict, respectively dark control and light-treated samples. Asterisks indicate significant differences between fruits, and dark control vector-infiltrated fruits (Student’s t test; *P < 0.05 and **P < 0.01). Comparisons between light and dark in each treatment group were also performed (Student’s t test; *P < 0.05 and **P < 0.01)
FaMYB10 effects on anthocyanin accumulation in strawberry fruit
Changes in anthocyanin accumulation during strawberry fruit ripening were initially measured by investigating anthocyanin content. Strawberry fruit growth and maturation are divided into six different stages (Fait et al. 2008). FaMYB10 expression patterns during strawberry fruit ripening were analyzed by qRT-PCR. FaMYB10 expression exhibited increased levels commensurate with fruit ripening, and the highest levels were detected in the Re stage (Fig. 5a).
FaMYB10 gene expression level and anthocyanin content in Famyb10 AmRNAi and FaMYB10-OE strawberry fruits. a FaMYB10 gene expression levels were detected by qRT-PCR during the six strawberry fruit developmental stages. FaMYB10 relative expression levels were normalized against FaGAPDH. In each assay, the normalized expression level was set to 1 for the SG stage. Data are expressed as mean ± SD (n = 4). b FaMYB10 relative expression levels in Famyb10 AmRNAi (black bars), FaMYB10-OE (gray bars), and fruit infiltrated with a control vector (white bars). FaMYB10 relative expression levels were normalized against FaGAPDH. In addition, normalized expression of each gene in the control vector-infiltrated fruits was set to 1. Data are expressed as mean ± SD (n = 4–5). c Famyb1 AmRNAi fruit phenotypes. d FaMYB1-OE fruits. e Control vector fruits. f Anthocyanin content was measured for Famyb10 AmRNAi (black bars), FaMYB10-OE (gray bars), and control vector-infiltrated fruits (white bars) (n = 4–6). Asterisks indicate significant differences among Famyb10 AmRNAi or FaMYB10-OE fruits, and control vector-infiltrated fruits (one-way ANOVA with post hoc Dunnet’s test; **P < 0.01 and ***P < 0.001)
Therefore, the relationship between FaMYB10 and strawberry fruit coloration was investigated by performing in planta RNAi-mediated down-regulation, or FaMYB10 over-expression. FaMYB10 expression levels in pRI-myb10i-(Famyb10 AmRNAi) or pRI-MYB10-OE-(FaMYB10-OE) infiltrated fruits were found fivefold lower and 1.7-fold higher relative to vector control fruits, respectively (Fig. 5b). In addition, significant differences between Famyb10 AmRNAi and FaMYB10-OE fruits compared to the control vector-infiltrated fruits were observed (P < 0.001).
Famyb10 AmRNAi and FaMYB10-OE fruits showed coloration reduction and accumulation (Fig. 5c, d) compared to the vector control (Fig. 5e). Total anthocyanin concentration in Famyb10 AmRNAi (171.0 ± 13.5 μg/g fresh weight) and FaMYB10-OE (1,032.7 ± 153.3 μg/g fresh weight) fruits were also 1.8-fold lower (P < 0.01) and 3.2-fold higher (P < 0.001) relative to control fruits (318.8 ± 24.8 μg/g fresh weight), respectively (Fig. 5f).
FaMYB10 effects on anthocyanin accumulation were further investigated by determining cyanidin 3-glucoside and pelargonidin 3-glucoside content, which are main strawberry fruit anthocyanins in Famyb10 AmRNAi and FaMYB10-OE fruits. Cyanidin 3-glucoside in Famyb10 AmRNAi (1.54 ± 0.83 μg/g fresh weight) and FaMYB10-OE fruits (292.39 ± 75.42 μg/g fresh weight) content was 3.4-fold lower (P < 0.01) and 56.4-fold higher (P < 0.001) relative to control fruits (5.18 ± 1.48 μg/g fresh weight), respectively (Table 2). Pelargonidin 3-glucoside in Famyb10 AmRNAi (73.49 ± 17.77 μg/g fresh weight) and FaMYB10-OE (1,114.50 ± 186.62 μg/g fresh weight) fruit content was also 6.4-fold lower (P < 0.001) and 2.4-fold higher (P < 0.001) relative to control fruits (473.94 ± 55.83 μg/g fresh weight), respectively (Table 2).
FaMYB10 effects on flavonoid pathway gene expression in strawberry fruit
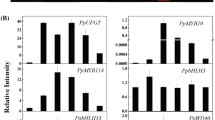
Results showed FaMYB10 expression level was positively correlated with anthocyanin accumulation in strawberry fruit. FaMYB10 effects on anthocyanin biosynthesis were better characterized by performing qRT-PCR analysis of flavonoid pathway genes in Famyb10 AmRNAi and FaMYB10-OE fruits. Expression levels in all flavonoid pathway genes, with the exception of the FaDFR1 gene were significantly reduced in Famyb10 AmRNAi compared to control fruits (P values ranged from <0.05 to 0.001). In FaDFR1, Famyb10 AmRNAi expression levels exhibited a significant increase compared to the control (P < 0.05). FaMYB10-OE fruits exhibited significantly increased expression levels for genes in the first five steps of the flavonoid pathway (FaCHS, FaCHI FaF3H, FaF3′H, and FaDFR2) compared to the control (P values ranged from <0.05 to 0.001), with the exception of the FaDFR1 gene (Fig. 6). Gene expression during the final two steps of the flavonoid pathway (FaANS and FaFGT) also showed increased FaMYB10-OE fruit expression, although significant differences compared to control fruits were not detected.
Flavonoid pathway gene expression profiling in Famyb10 AmRNAi and FaMYB10-OE strawberry fruits. The following flavonoid pathway genes were detected by qRT-PCR: FaCHS (a), FaCHI (b), FaF3H (c), FaF3′H (d), FaDFR1 (e), FaDFR2 (f), FaANS (g) and FaFGT (h). Relative expression levels for each gene were normalized to FaGAPDH levels. In addition, normalized expression of each gene in the control vector-infiltrated fruits was set to 1. Data are expressed as mean ± SD (n = 3–5). Asterisks indicate significant differences among Famyb10 AmRNAi (black bars) or FaMYB10-OE fruits (gray bars), and control vector-infiltrated fruits (white bars) (one-way ANOVA with post hoc Dunnet’s test; *P < 0.05, **P < 0.01, and ***P < 0.001)
Discussion
Strawberry plant transformation with FaMYB10 resulted in elevated anthocyanin content in several plant tissues (Lin-Wang et al. 2010). In the present study, we characterized FaMYB10 during strawberry fruit ripening, and analyzed FaMYB10 regulation and function using in planta transient assays we previously developed (Miyawaki et al. 2012; Kadomura-Ishikawa et al. 2013). FaMYB10 down-regulation and over-expression exhibited respective significant decreased and increased anthocyanin content in strawberry fruits, and were accompanied by consistent flavonoid pathway gene transcript levels. We also determined cyanidin 3-glucoside and pelargonidin 3-glucoside content, and revealed not only pelargonidin 3-glucoside but also cyanidin 3-glucoside content increased and decreased in FaMYB10 over-expressed and down-regulated fruits, respectively. FaMYB10, in particular, strongly affected cyanidin 3-glucoside content in strawberry fruits. In Fragaria ananassa cv. Sachinoka, pelargonidin 3-glucoside was the predominant pigment (approximately 90 % of the total anthocyanin content), and cyanidin 3-glucoside was the minor pigment (approximately 10 % of the total anthocyanin content) in mature fruits (Yoshida et al. 2002). Cyanidin 3-glucoside is a 3′,4′-hydroxylated compound, and the 3′-hydroxylation of phenolic compounds is catalyzed by F3′H. Results showed high FaF3′H levels during early stages, which markedly decreased at stages following Wh (Carbone et al. 2009; present data Suppl. Fig. S2), resulting in low cyanidin 3-glucoside content in mature fruits. We showed FaMYB10 affected FaF3′H levels, which influenced cyanidin 3-glucoside content in strawberry fruit. These results confirmed the relationship between FaMYB10 and anthocyanin accumulation in fruits by different assay systems. In addition, results indicated FaMYB10 functioned as a positive anthocyanin biosynthesis regulator in strawberry fruit. However, FaF3′H exhibited low levels during later fruit stages (Carbone et al. 2009; our data Suppl. Fig. S2), while FaMYB10 showed high levels from strong induction at later fruits stages during normal fruit ripening. A lack of congruency between FaMYB10 and FaF3′H expression patterns in fruit coloration was observed. It is possible an unknown FaF3′H regulator, which competes with FaMYB10 might exist in strawberry fruits. However, further analysis is required to confirm this possibility.
Lin-Wang et al. (2010) reported FaMYB10 activated the Arabidopsis DFR promoter in tobacco leaves. Consistent with heterologous assays, FaMYB10 affected FaDFR2, but not FaDFR1 levels in Famyb10 AmRNAi and FaMYB10-OE fruits. Medina-Puche et al. (2014) showed decreased DFR (FaDFR2) gene levels in FaMYB10 down-regulated fruits, consistent with our result, which suggested FaMYB10 might activate FaDFR2 but not FaDFR1 expression in strawberry fruit. It is possible FaDFR1 and FaDFR2 might serve different roles in strawberry fruit ripening and coloration.
In fruits of many climacteric and non-climacteric fruits, including apple (Malus pumila), peach (Amygdalus persica), pear (Pyrus communis), grape (Vitis labruscana), Vaccinium sp., bilberry (Vaccinium myrtillus), cranberry (Vaccinium microcarpum), and litchi (Litchi chinensis), flavonoid pathway gene expression was light induced, resulting in anthocyanin accumulation (Dussi et al. 1995; Kim et al. 2003; Kataoka and Beppu 2004; Zhou and Singh 2004; Jia et al. 2005; Wei et al. 2011; Azuma et al. 2012; Uleberg et al. 2012). In addition, several studies demonstrated the integral role of light in strawberry fruit coloration (Anttonen et al. 2006; Carbone et al. 2009; Josuttis et al. 2010; Kadomura-Ishikawa et al. 2013), confirmed by results of the present study. Furthermore, our results indicated initial increased FaMYB10 transcript levels after light treatment, followed by increased flavonoid pathway gene (FaCHS) expression, resulting in anthocyanin accumulation. FaMYB10 transcript levels were also significantly increased by light compared to the dark control, indicating FaMYB10 expression was positively regulated by light. However, FaMYB10 transcripts increased under dark conditions, suggesting other factors regulated anthocyanin biosynthesis in strawberry fruit.
Studies recently reported the important role of ABA in strawberry fruit ripening. Down-regulation of the ABA synthesis enzyme (NCED1) and ABA receptor (CHLH/ABAR and PYR1) showed an unpigmented phenotype (Jia et al. 2011, 2013b; Chai et al. 2011). Daminato et al. (2013) showed ABA treatment induced FaMYB10 transcript levels in strawberry fruit. Most recently, Medina-Puche et al. (2014) showed that ABA synthesis inhibitor treatment and NCED1 down-regulation in strawberry fruits decreased FaMYB10 levels. In the present study, we showed ABA and fluridone treatments, respectively, increased and decreased anthocyanin fruit content, with corresponding increased and decreased FaMYB10 and FaCHS gene transcript levels. We also evaluated FaANS and FaFGT transcript levels, which were directly associated with anthocyanin biosynthesis, and expression patterns were similar to FaMYB10 and FaCHS (Suppl. Fig. S1). Furthermore, FaNCED1 and FaCHLH down-regulation resulted in the same fruit phenotype as fluridone-treated fruit. These results suggested FaMYB10 expression was positively regulated by ABA, and influenced strawberry fruit anthocyanin content. This observation is one explanation for increased anthocyanin content in fruit under dark conditions, and endogenous ABA might be affected. Indeed, Symons et al. (2012) revealed ABA levels gradually increased during fruit ripening, and the highest level was detected at the Re stage.
The results of this study provided evidence that anthocyanin biosynthesis was regulated by light and ABA in strawberry fruit (Fig. 7). Additive effects were observed in anthocyanin accumulation under combined light and ABA, suggesting light and ABA signaling pathways are independent. Most recently, we showed FaPHOT2 was associated with anthocyanin biosynthesis induced by light in strawberry fruit (Kadomura-Ishikawa et al. 2013), and former studies reported CRY association in anthocyanin biosynthesis (Ahmad et al. 1995; Giliberto et al. 2005; Chatterjee et al. 2006). Anthocyanin content decreased in the Arabidopsis CRY1 mutant, and CRY2-deficient seedlings relative to the wild type (Ahmad et al. 1995; Li et al. 2013). Li et al. (2013) reported anthocyanins accumulated in Arabidopsis seedlings over-expressed for apple CRY2. These reports indicated CRYs might also be associated with anthocyanin biosynthesis in strawberry. In the signaling pathways, light and ABA promoted FaMYB10 expression, resulting in anthocyanin accumulation via accelerated flavonoid pathway gene expression. Recent studies revealed auxin negatively regulated FaMYB10 in strawberry fruit (Daminato et al. 2013; Medina-Puche et al. 2014). Daminato et al. (2013) also reported two MADS-box transcription factors, SHATTERPROOF-like (FaSHP) and FaMADS9 were involved in the ABA and auxin signaling pathways, although the function of these genes has not been characterized. Therefore, detailed signal transduction in each signaling pathway, and crosstalk among the signaling pathways remains elusive. Efforts are currently under way to determine the signal transductions and crosstalk between these signaling pathways.
Author contribution statement
YK-I, KM, AT and SN designed and YK-I conducted the research. YK-I and KM contributed new reagents and analytic tools. YK-I, KM, AT, TM, and SN analyzed data. YK-I wrote the manuscript. All authors read and approved the manuscript.
Abbreviations
- ABA:
-
Abscisic acid
- CHS:
-
Chalcone synthase
- CHI:
-
Chalcone isomerase
- CHLH/ABAR:
-
Magnesium chelatase H subunit
- F3′H:
-
Flavonoid 3′-hydroxylase
- F3H:
-
Flavanone 3-hydroxylase
- DFR:
-
Dihydroflavonol-4-reductase
- ANS:
-
Anthocyanidin synthase
- FGT:
-
Flavonoid glycosyltransferase
- TF:
-
Transcription factors
- NCED:
-
9-cis epoxycarotenoid dioxygenase
References
Aharoni A, De Vos CH, Wein M, Sun Z, Greco R, Kroon A, Mol JN, O’Connell AP (2001) The strawberry FaMYB1 transcription factor suppresses anthocyanin and flavonol accumulation in transgenic tobacco. Plant J 28:319–332. doi:10.1046/j.1365-313X.2001.01154.x
Ahmad M, Lin C, Cashmore AR (1995) Mutations throughout an Arabidopsis blue-light photoreceptor impair blue-light-responsive anthocyanin accumulation and inhibition of hypocotyl elongation. Plant J 8:653–658. doi:10.1046/j.1365-313X.1995.08050653.x
Alexander L, Grierson D (2002) Ethylene biosynthesis and action in tomato: a model for climacteric fruit ripening. J Exp Bot 53:2039–2055. doi:10.1093/jxb/erf072
Almeida JR, D’Amico E, Preuss A, Carbone F, de Vos CH, Deiml B, Mourgues F, Perrotta G, Fischer TC, Bovy AG, Martens S, Rosati C (2007) Characterization of major enzymes and genes involved in flavonoid and proanthocyanidin biosynthesis during fruit development in strawberry (Fragaria xananassa). Arch Biochem Biophys 465:61–71. doi:10.1016/j.abb.2007.04.040
Anttonen MJ, Hoppula KI, Nestby R, Verheul MJ, Karjalainen RO (2006) Influence of fertilization, mulch color, early forcing, fruit order, planting date, shading, growing environment, and genotype on the contents of selected phenolics in strawberry (Fragaria × ananassa Duch.) fruits. J Agric Food Chem 54:2614–2620. doi:10.1021/jf052947w
Azuma A, Yakushiji H, Koshita Y, Kobayashi S (2012) Flavonoid biosynthesis-related genes in grape skin are differentially regulated by temperature and light conditions. Planta 236:1067–1080. doi:10.1007/s00425-012-1650-x
Baudry A, Heim MA, Dubreucq B, Caboche M, Weisshaar B, Lepiniec L (2004) TT2, TT8, and TTG1 synergistically specify the expression of BANYULS and proanthocyanidin biosynthesis in Arabidopsis thaliana. Plant J 39:366–380. doi:10.1111/j.1365-313X.2004.02138.x
Borevitz JO, Xia Y, Blount J, Dixon RA, Lamb C (2000) Activation tagging identifies a conserved MYB regulator of phenylpropanoid biosynthesis. Plant Cell 12:2383–2394. doi:10.1105/tpc.12.12.2383
Carbone F, Preuss A, De Vos RC, D’Amico E, Perrotta G, Bovy AG, Martens S, Rosati C (2009) Developmental, genetic and environmental factors affect the expression of flavonoid genes, enzymes and metabolites in strawberry fruits. Plant Cell Environ 32:1117–1131. doi:10.1111/j.1365-3040.2009.01994.x
Chai YM, Jia HF, Li CL, Dong QH, Shen YY (2011) FaPYR1 is involved in strawberry fruit ripening. J Exp Bot 62:5079–5089. doi:10.1093/jxb/err207
Chatterjee M, Sharma P, Khurana JP (2006) Cryptochrome 1 from Brassica napus is up-regulated by blue light and controls hypocotyl/stem growth and anthocyanin accumulation. Plant Physiol 141:61–74. doi:10.1104/pp.105.076323
Chernys JT, Zeevaart JAD (2007) Characterization of the 9-cis-epoxycarotenoid dioxygenase gene family and the regulation of abscisic acid biosynthesis in avocado. Plant Physiol 124:343–353. doi:10.1104/pp.124.1.343
Cominelli E, Gusmaroli G, Allegra D, Galbiati M, Wade HK, Jenkins GI, Tonelli C (2008) Expression analysis of anthocyanin regulatory genes in response to different light qualities in Arabidopsis thaliana. J Plant Physiol 165:886–894. doi:10.1016/j.jplph.2007.06.010
Daminato M, Guzzo F, Casadoro G (2013) A SHATTERPROOF-like gene controls ripening in non-climacteric strawberries, and auxin and abscisic acid antagonistically affect its expression. J Exp Bot 64:3775–3786. doi:10.1093/jxb/ert214
Davies C, Boss PK, Robinson SP (1997) Treatment of grape berries, a nonclimacteric fruit with a synthetic auxin, retards ripening and alters the expression of developmentally regulated genes. Plant Physiol 115:1155–1161. doi:10.1104/pp.115.3.1155
Deikman J (1997) Molecular mechanisms of ethylene regulation of gene transcription. Physiol Plant 100:561–566. doi:10.1111/j.1399-3054.1997.tb03061.x
Dubos C, Le Gourrierec J, Baudry A, Huep G, Lanet E, Debeaujon I, Routaboul JM, Alboresi A, Weisshaar B, Lepiniec L (2008) MYBL2 is a new regulator of flavonoid biosynthesis in Arabidopsis thaliana. Plant J 55:940–953. doi:10.1111/j.1365-313X.2008.03564.x
Dussi MC, Sugar D, Wrolstad RE (1995) Characterizing and quantifying anthocyanins in red pears and the effect of light quality on fruit color. J Am Soc Hortic Sci 120:785–789
Fait A, Hanhineva K, Beleggia R, Dai N, Rogachev I, Nikiforova VJ, Fernie AR, Aharoni A (2008) Reconfiguration of the achene and receptacle metabolic networks during strawberry fruit development. Plant Physiol 148:730–750. doi:10.1104/pp.108.120691
Finkelstein RR, Gampala SS, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14:S15–S45. doi:10.1105/tpc.010441
Gapper NE, McQuinn RP, Giovannoni JJ (2013) Molecular and genetic regulation of fruit ripening. Plant Mol Biol 82:575–591. doi:10.1007/s11103-013-0050-3
Giliberto L, Perrotta G, Pallara P, Weller JL, Fraser PD, Bramley PM, Fiore A, Tavazza M, Giuliano G (2005) Manipulation of the blue light photoreceptor cryptochrome 2 in tomato affects vegetative development, flowering time, and fruit antioxidant content. Plant Physiol 137:199–208. doi:10.1104/pp.104.051987
Giovannoni JJ (2004) Genetic regulation of fruit development and ripening. Plant Cell 16:S170–S180. doi:10.1105/tpc.019158
He J, Giusti MM (2010) Anthocyanins: natural colorants with health-promoting properties. Annu Rev Food Sci Technol 1:163–187. doi:10.1146/annurev.food.080708
Hirayama T, Shinozaki K (2007) Perception and transduction of abscisic acid signals: keys to the function of the versatile plant hormone ABA. Trends Plant Sci 12:343–351. doi:10.1016/j.tplants.2007.06.013
Hoffmann T, Kalinowski G, Schwab W (2006) RNAi-induced silencing of gene expression in strawberry fruit (Fragaria × ananassa) by agroinfiltration: a rapid assay for gene function analysis. Plant J 48:818–826. doi:10.1111/j.1365-313X.2006.02913.x
Jia HJ, Araki A, Okamoto G (2005) Influence of fruit bagging on aroma volatiles and skin coloration of “Hakuho” peach (Prunus persica Batsch). Postharvest Biol Technol 35:61–68. doi:10.1016/j.postharvbio.2004.06.004
Jia HF, Chai YM, Li CL, Lu D, Luo JJ, Qin L, Shen YY (2011) Abscisic acid plays an important role in the regulation of strawberry fruit ripening. Plant Physiol 157:188–199. doi:10.1104/pp.111.177311
Jia H, Wang Y, Sun M, Li B, Han Y, Zhao Y, Li X, Ding N, Li C, Ji W, Jia W (2013a) Sucrose functions as a signal involved in the regulation of strawberry fruit development and ripening. New Phytol 198:453–465. doi:10.1111/nph.12176
Jia HF, Lu D, Sun JH, Li CL, Xing Y, Qin L, Shen YY (2013b) Type 2C protein phosphatase ABI1 is a negative regulator of strawberry fruit ripening. J Exp Bot 64:1677–1687. doi:10.1093/jxb/ert028
Jiang Y, Joyce DC (2003) ABA effects on ethylene production, PAL activity, anthocyanin and phenolic contents of strawberry fruit. Plant Growth Regul 39:171–174. doi:10.1023/A:1022539901044
Jin H, Cominelli E, Bailey P, Parr A, Mehrtens F, Jones J, Tonelli C, Weisshaar B, Martin C (2000) Transcriptional repression by AtMYB4 controls production of UV-protecting sunscreens in Arabidopsis. EMBO J 19:6150–6161. doi:10.1093/emboj/19.22.6150
Josuttis M, Dietrich H, Treutter D, Will F, Linnemannstöns L, Krüger E (2010) Solar UVB response of bioactives in strawberry (Fragaria × ananassa Duch. L.): a comparison of protected and open-field cultivation. J Agric Food Chem 58:12692–12702. doi:10.1021/jf102937e
Kadomura-Ishikawa Y, Miyawaki K, Noji S, Takahashi A (2013) Phototropin 2 is involved in blue light-induced anthocyanin accumulation in Fragaria × ananassa fruits. J Plant Res 126:847–857. doi:10.1007/s10265-013-0582-2
Kami C, Lorrain S, Hornitschek P, Fankhauser C (2010) Light-regulated plant growth and development. Curr Top Dev Biol 91:29–66. doi:10.1016/S0070-2153(10)91002-8
Kano Y, Asahira T (1981) Roles of cytokinin and abscisic acid in the maturing of strawberry fruits. J Jpn Soc Hortic Sci 50:31–36
Karppinen K, Hirvelä E, Nevala T, Sipari N, Suokas M, Jaakola L (2013) Changes in the abscisic acid levels and related gene expression during fruit development and ripening in bilberry (Vaccinium myrtillus L.). Phytochemistry 95:127–134. doi:10.1016/j.phytochem.2013.06.023
Kataoka I, Beppu K (2004) UV irradiance increases development of red skin color and anthocyanins in “Hakuho” peach. HortScience 39:1234–1237
Kim SH, Lee JR, Hong ST, Yoo YK, An G, Kim SR (2003) Molecular cloning and analysis of anthocyanin biosynthesis genes preferentially expressed in apple skin. Plant Sci 165:403–413. doi:10.1016/S0168-9452(03)00201-2
Klee HJ, Giovannoni JJ (2011) Genetics and control of tomato fruit ripening and quality attributes. Annu Rev Genet 45:41–59. doi:10.1146/annurev-genet-110410-132507
Kobashi K, Gemma H, Iwahori S (1999) Sugar accumulation in peach fruit as affected by abscisic acid (ABA) treatment in relation to some sugar metabolizing enzymes. J Jpn Soc Hortic Sci 68:465–470
Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10:236–242. doi:10.1016/j.tplants.2005.03.002
Kortstee AJ, Khan SA, Helderman C, Trindade LM, Wu Y, Visser RG, Brendolise C, Allan A, Schouten HJ, Jacobsen E (2011) Anthocyanin production as a potential visual selection marker during plant transformation. Transgenic Res 20:1253–1264. doi:10.1007/s11248-011-9490-1
Li C, Jia H, Chai Y, Shen Y (2011) Abscisic acid perception and signaling transduction in strawberry: a model for non-climacteric fruit ripening. Plant Signal Behav 6:1950–1953. doi:10.4161/psb.6.12.18024
Li YY, Mao K, Zhao C, Zhao XY, Zhang RF, Zhang HL, Shu HR, Hao YJ (2013) Molecular cloning and functional analysis of a blue light receptor gene MdCRY2 from apple (Malus domestica). Plant Cell Rep 32:555–566. doi:10.1007/s00299-013-1387-4
Lin-Wang K, Bolitho K, Grafton K, Kortstee A, Karunairetnam S, McGhie TK, Espley RV, Hellens RP, Allan AC (2010) An R2R3 MYB transcription factor associated with regulation of the anthocyanin biosynthetic pathway in Rosaceae. BMC Plant Biol 21(10):50. doi:10.1186/1471-2229-10-50
Lopes-da-Silva F, Escribano-Bailòn MT, Pérez Alonso JJ, Rivas-Gonzalo JC, Santos-Buelga C (2007) Anthocyanin pigments in strawberry. LWT Food Sci Technol 40:374–382. doi:10.1016/j.lwt.2005.09.018
Matsui K, Umemura Y, Ohme-Takagi M (2008) AtMYBL2, a protein with a single MYB domain, acts as a negative regulator of anthocyanin biosynthesis in Arabidopsis. Plant J 55:954–967. doi:10.1111/j.1365-313X.2008.03565.x
Medina-Puche L, Cumplido-Laso G, Amil-Ruiz F, Hoffmann T, Ring L, Rodríguez-Franco A, Caballero JL, Schwab W, Muñoz-Blanco J, Blanco-Portales R (2014) MYB10 plays a major role in the regulation of flavonoid/phenylpropanoid metabolism during ripening of Fragaria × ananassa fruits. J Exp Bot 65:401–417. doi:10.1093/jxb/ert377
Miyawaki K, Fukuoka S, Kadomura Y, Hamaoka H, Mito T, Ohuchi H, Schwab W, Noji S (2012) Establishment of a novel system to elucidate the mechanisms underlying light-induced ripening of strawberry fruit with an Agrobacterium-mediated RNAi technique. Plant Biotechnol 29:271–277. doi:10.5511/plantbiotechnology.12.0406a
Nakashima K, Yamaguchi-Shinozaki K (2013) ABA signaling in stress-response and seed development. Plant Cell Rep 32:959–970. doi:10.1007/s00299-013-1418-1
Pan QH, Li MJ, Peng CC, Zhang N, Zou X, Zou KQ, Wang XL, Yu XC, Wang XF, Zhang DP (2005) Abscisic acid activates acid invertases in developing grape berry. Physiol Plant 125:157–170. doi:10.1111/j.1399-3054.2005.00552.x
Qin X, Zeevaart JA (1999) The 9-cis-epoxycarotenoid cleavage reaction is the key regulatory step of abscisic acid biosynthesis in water-stressed bean. Proc Natl Acad Sci USA 96:15354–15361. doi:10.1016/j.jplph.2009.01.013
Rodrigo MJ, Marcos JF, Alférez F, Mallent MD, Zacarías L (2003) Characterization of Pinalate, a novel Citrus sinensis mutant with a fruit specific alteration that results in yellow pigmentation and decreased ABA content. J Exp Bot 54:727–738. doi:10.1093/jxb/erg083
Rodrigo MJ, Alquezar B, Zacarı´as L (2006) Cloning and characterization of two 9-cis-epoxycarotenoid dioxygenase genes, differentially regulated during fruit maturation and under stress conditions, from orange (Citrus sinensis L. Osbeck). J Exp Bot 57:633–643. doi:10.1093/jxb/erj048
Rook F, Hadingham SA, Li Y, Bevan MW (2006) Sugar and ABA response pathways and the control of gene expression. Plant Cell Environ 29:426–434. doi:10.1111/j.1365-3040.2005.01477.x
Salvatierra A, Pimentel P, Moya-Leon MA, Caligari PD, Herrera R (2010) Comparison of transcriptional profiles of flavonoid genes and anthocyanin contents during fruit development of two botanical forms of Fragaria chiloensis ssp. chiloensis. Phytochemistry 71:1839–1847. doi:10.1016/j.phytochem.2010.08.005
Symons GM, Chua YJ, Ross JJ, Quittenden LJ, Davies NW, Reid JB (2012) Hormonal changes during non-climacteric ripening in strawberry. J Exp Bot 63:4741–4750. doi:10.1093/jxb/ers147
Tamagnone L, Merida A, Parr A, Mackay S, Culianez-Macia FA, Roberts K, Martin C (1998) The AmMYB308 and AmMYB330 transcription factors from Antirrhinum regulate phenylpropanoid and lignin biosynthesis in transgenic tobacco. Plant Cell 10:135–154. doi:10.1105/tpc.10.2.135
Uleberg E, Rohloff J, Jaakola L, Trôst K, Junttila O, Häggman H, Martinussen I (2012) Effects of temperature and photoperiod on yield and chemical composition of northern and southern clones of bilberry (Vaccinium myrtillus L.). J Agric Food Chem 60:10406–10414. doi:10.1021/jf302924m
Wei YZ, Hu FC, Hu GB, Li XJ, Huang XM, Wang HC (2011) Differential expression of anthocyanin biosynthetic genes in relation to anthocyanin accumulation in the pericarp of Litchi chinensis Sonn. PLoS ONE 6:e19455. doi:10.1371/journal.pone.0019455
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. doi:10.1146/annurev.arplant.57.032905.105444
Yoshida Y, Koyama N, Tamura H (2002) Color and anthocyanin composition of strawberry fruit: changes during fruit development and differences among cultivars, with special reference to the occurrence of pelargonidin 3-malnoylglucoside. J Japan Soc Hort Sci 71:355–361
Zhang M, Leng P, Zhang G, Li X (2009a) Cloning and functional analysis of 9-cis-epoxycarotenoid dioxygenase (NCED) genes encoding a key enzyme during abscisic acid biosynthesis from peach and grape fruits. J Plant Physiol 166:1241–1252. doi:10.1016/j.jplph.2009.01.013
Zhang M, Yuan B, Leng P (2009b) The role of ABA in triggering ethylene biosynthesis and ripening of tomato fruit. J Exp Bot 60:1579–1588. doi:10.1093/jxb/erp026
Zhou Y, Singh BR (2004) Effect of light on anthocyanin levels in submerged, harvested cranberry fruit. J Biomed Biotechnol 2004:259–263. doi:10.1155/S1110724304403027
Acknowledgments
We thank Koichi Hayashi for cultivating and donating strawberry fruit samples. We also thank Toshifumi Miki and Keisuke Hirota (Tokushima Agricultural Research Center, Japan) for invaluable advice, and additional strawberry fruit samples. Ourgenic Co. Ltd (Tokushima, Japan) supported this research. LED-Life project of the University of Tokushima, Japan, also provided financial support for this work.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kadomura-Ishikawa, Y., Miyawaki, K., Takahashi, A. et al. Light and abscisic acid independently regulated FaMYB10 in Fragaria × ananassa fruit. Planta 241, 953–965 (2015). https://doi.org/10.1007/s00425-014-2228-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-014-2228-6