Abstract
The eukaryotic ascomycete genus Fusarium comprises many species capable of producing secondary metabolites important for agriculture, health, and biotechnology. Filamentous fungi share common physiological features, but even within Fusarium, there are significant differences that affect the success of biotechnological methods used to unravel biosynthetic pathways. The aim of this review is to describe the different methods that have successfully been used throughout the genus Fusarium to identify the products of novel biosynthetic pathways. The results are presented in tables to give the reader an overview and thereby enable the selection of the most appropriate method to the problem, regarding both species and target products. Significant work has gone into characterization of the underlying molecular genetics of secondary metabolites, but still, the products of only 25–30% of predicted gene clusters have been identified. In this review, we highlight existing knowledge and encourage the development of new techniques and strategies to provide access to the many unknown polyketide and non-ribosomal peptide products that await discovery in Fusarium.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soil-borne ascomycete fungi belonging to the genus Fusarium have high impact on health and agriculture (Nucci and Anaissie 2007; Dean et al. 2012). Like other filamentous fungi, Fusarium has the ability to produce small-specialized compounds, secondary metabolites (SMs), not associated directly with growth or reproduction, although hypothesized to contribute to the fitness of the fungal producer (Brakhage 2013; Ding et al. 2018). Major research achievements have provided an understanding of the biochemical and molecular machinery which controls the formation of these chemical compounds (Keller 2019). Specialized SMs may act as plant hormones or virulence factors, as mycotoxins dangerous to humans and animals, or even as weapons against other microbes (Desjardins and Proctor 2007; Brown et al. 2014). This makes characterization of the compounds and the underlying genetics an important priority. Here, we aim to highlight the technologies that have been used to unravel the secondary metabolism in Fusarium and encourage further initiatives to link biosynthetic gene clusters to their products.
Fusarium comprises more than 100–500 (Leslie and Summerell 2006) species capable of causing infection in plant, humans, and domesticated animals (Summerell et al. 2010). Members of this genus are found in warm and temperate ecosystems throughout the globe, often as plant pathogens contributing to major economic losses due to infected crops (Mcmullen et al. 1997; Windels 2000; Michielse and Rep 2009). Many species are harmless, but species such as F. graminearum and F. oxysporum infect cereals and produce high amounts of mycotoxins rendering entire harvests unfit for consumption (Windels 2000). The speciation of Fusarium has always posed a challenge to researchers due to the lack of distinguishing morphological features. Historically, the number of recognized species has varied between 9 and > 1000, depending on the implemented identification scheme. Currently, many species concepts represent polyphyletic clades comprising several individual species (Leslie and Summerell 2006; Watanabe et al. 2011). Therefore, careful naming of strains is important for this genus. In some cases, the term forma specialis (f. sp.) has been introduced, e.g., categorizing strains of F. oxysporum after the plant they infect (Gordon and Martyn 1997).
Fusarium SMs exhibit an extreme diversity in function and chemical structures. They are usually formed by multi-domain core synthases often cooperating with several decorating enzymes in a pathway to generate the final product. The genes encoding these enzymes are co-regulated and commonly found as neighbors to the core-synthase-encoding gene, and together, they form a biosynthetic gene cluster (BGC) (Keller et al. 1997; Yu and Keller 2005). In addition, gene-encoding transcriptional regulators, transport proteins, and even product detoxification proteins can be encoded by BGCs. Unfortunately, many of these potential BGCs shows little to no expression when grown under the standard laboratory conditions (Gaffoor et al. 2005). Therefore, the potential new SMs may not be produced at all or are present at levels too low to be detected by the standard chemical methods (Wiemann and Keller 2014). Although many molecules have been isolated and described, the full biochemical potential of the collected Fusarium secondary metabolome is yet to be explored. In this review, we cover techniques that have been successful in Fusarium to link gene clusters to biosynthetic compounds and pathways, and suggest new possibilities to be explored in future endeavors.
Classes of secondary metabolites
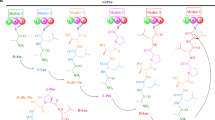
Secondary metabolites are synthesized from small metabolites such as short-chain carboxylic acids and amino acids from the primary metabolism. These precursors are polymerized by large synthase/synthetase enzymes such as iterative polyketide synthases (PKS, types I and III), non-ribosomal peptide synthetases (NRPS), or terpene cyclases (TC). Fusarium species are capable of producing many terpenes (Brock et al. 2013; Burkhardt et al. 2016), some of which are important virulence factors, for example, nivalenol and deoxynivalenol (Marasas et al. 1979; Yoshida and Nakajima 2010; Yang et al. 2018) or plant hormones such as gibberellins (Bömke and Tudzynski 2009; Troncoso et al. 2010). However, the majority of characterized SMs belong to the chemical groups of polyketides (reduced and non-reduced), non-ribosomal peptides, or hybrid compounds (Sieber et al. 2014; Hoogendoorn et al. 2018) (Fig. 1). PKSs are large multi-domain enzymes that as a minimum contain beta-ketosynthase (KS), acyl-transferase (AT), and acyl-carrier protein (ACP) domains which work together in an iterative manner to elongate a polyketide chain with one ketide unit at a time (McDaniel et al. 1994; Bentley and Bennett 1999). Fungal PKSs work in different ways to create structural diversity. Usually, the biosynthesis will start from an acetyl-CoA unit, which is then elongated with malonyl-CoA units through Claisen condensation performed by the KS domain. However, in some cases, the starter unit can stem from another PKS or a fatty acid synthase (Brown et al. 1996). In addition to the KS-AT-ACP domains, PKSs may contain tailoring domains which add to the chemical diversity, e.g., reductase, dehydrogenase, or methyltransferase domains (Meier and Burkart 2009). The PKS type I, which is most predominant in Fusarium (Brown and Proctor 2016), can be further subdivided into reducing or non-reducing PKSs, yielding either fatty acid like or aromatic products. Finally, the tailoring domains can skip an iteration as seen for zearalenone, where only four out of five ketones are fully reduced (Gaffoor and Trail 2006). In addition, similar PKSs may produce different products. It is not surprising that the prediction of the final product based on amino-acid sequence alone has proven impossible.
NRPSs are multi-modular assembly lines catalyzing the formation of small peptides from amino-acid monomers. A full NRPS module contains an adenylation (A), a peptide acyl-carrier (T), and a condensation (C) domain. An NRPS is thus composed of one or more elongation modules (A–T–C) which catalyze the formation of a polypeptide chain. In addition, each module may contain tailoring domains, e.g., epimerization or N-methylation domains, that contribute to the chemical diversity of non-ribosomal peptides (Finking and Marahiel 2004). The compound is released from the synthetase by cyclization, reduction, or hydrolysis, and can be further modified by additional tailoring enzymes encoded by the gene cluster such as cytochrome P450 monooxygenases and dehydrogenases.
At least 500 different NRPS substrates have been reported in filamentous fungi, which comprise non-proteinogenic amino acids, d and l forms, and even hydroxyl acids (Strieker et al. 2010). The A domain contains a binding pocket that recognizes a specific amino-acid substrate (Conti et al. 1997), and substrate prediction algorithms have been developed, first for bacterial NRPSs (Stachelhaus et al. 1999; Challis et al. 2000) and further modified to include eukaryotic NRPSs (Rausch 2005; Bachmann and Ravel 2009; Röttig et al. 2011; Khayatt et al. 2013; Knudsen et al. 2016). The feasibility of using these tools to predict Fusarium NRPS substrate accurately remains to be demonstrated (Wollenberg et al. 2017).
In the case for both NRPS and PKS BGCs, the linking of biosynthetic metabolites to their respective genes relies on experimental evidence. In the following sections, we will cover the current pre-requisites for linking Fusarium metabolite–BGC pairs.
Genomic resources
Knowledge of the genetic basis for SM biosynthesis is essential for genetic manipulation and genome-mining strategies. Genome sequencing has been carried out for several species representative of the genus Fusarium (Cuomo et al. 2007; Ma et al. 2010, 2014; Al-Reedy et al. 2012; Gardiner et al. 2012, 2014; Wiemann et al. 2013; Moolhuijzen et al. 2013; Lysøe et al. 2014; King et al. 2015; Vanheule et al. 2016; Brown and Proctor 2016; Ponts et al. 2018; Walkowiak et al. 2016), revealing a potential for SM production that exceeds the expected (Kroken et al. 2003; Sieber et al. 2014). Comparative analyses of biosynthetic genes show their distribution across the Fusarium metagenome and provide insight into the evolution of BGCs (Ma et al. 2010). Different species of Fusarium are able to produce structurally similar compounds, e.g., fusarubins (PKS3), gibepyrone A (PKS8), and ferricrocin (NRPS2) (Wiemann et al. 2013; Hansen et al. 2015) and, therefore, carry alleles with high levels of homology. To detect BGCs in Fusarium genomes, different bioinformatic complementary tools are used. Protein prediction tools such as SMURF (Khaldi et al. 2010) can predict SM-related genes (Ma et al. 2010; Wiemann et al. 2013) and InterPRO (Apweiler et al. 2000) can determine the protein domain functions (Frandsen et al. 2006). The BLASTP alignment-based algorithm (Altschul et al. 1990) has become a cornerstone in every genetics study and was used in the first Fusarium genetic analyses (Proctor et al. 1999; Linnemannstöns et al. 2002; Kim et al. 2005a; Malz et al. 2005; Varga et al. 2005). CASSIS (Wolf et al. 2016) was developed based on the hypothesis that BGC genes that are co-localized and co-expressed within the same cluster will contain common regulatory patterns in the cluster promoters. SMIPS/CASSIS has been used to identify cluster-specific promoter motifs (Sieber et al. 2014). Finally, the AntiSMASH (Blin et al. 2017) platform combines some of the above methods and includes co-localization comparison analysis (cluster orthology) data to identify BGCs (Wiemann and Keller 2014). These bioinformatic analyses can be combined and aligned with experimental data to form strong evidence in scientific studies.
Three recent studies have analyzed 31 available Fusarium genomes for the presence and distribution of BGCs (Hansen et al. 2012b, 2015; Brown and Proctor 2016; Hoogendoorn et al. 2018) (Fig. 2). Prediction of SM gene clusters (and pseudo-genes) has been carried out and a numbering nomenclature was introduced (Hansen et al. 2012b, 2015). This has been extended to provide a simple system to identify all the PKS and NRPS genes by a number (Brown and Proctor 2016). So far, 67 PKS and 52 NRPS gene clusters have been identified distributed across the Fusarium metagenome. Only 16 out of 67 PKS and 13 out of 52 NRPS Fusarium genes have been linked to their respective biosynthetic product (Table 1).
Some biosynthetic gene clusters are found in the majority of species of Fusarium (PKS3, 7, 8), while others are restricted to a single phylogenetic clade, e.g., PKS29, 30, 31, 32, 33, and 35 from the F. solani complex. The distribution of Fusarium BGCs does not always follow a strict phylogenetic pattern and evidence for horizontal gene transfer events has been reported (Oide et al. 2006; Ma et al. 2010; Gardiner et al. 2012; Sieber et al. 2014).
Activation through cultivation
Although Fusarium does produce many metabolites under cultivation, the majority of BGCs remain silent and their products cryptic. The nature and function of Fusarium SMs probably require specific physiochemical conditions to trigger their activity (Brakhage 2013). To maximize chance of observing fungal metabolites, a popular strategy is to use different cultivation parameters (Bode et al. 2002; Hemphill et al. 2017). However, the activation of cryptic biosynthesis pathways is never guaranteed (Gaffoor et al. 2005). Properties such as pH and nitrogen source are important parameters to control for some metabolite pathways (Linnemannstöns et al. 2002; Kim et al. 2005b; Gomez-Gil et al. 2018). Substituting the nitrogen source glutamine with sodium nitrate in ICI medium leads to the formation of fusarubins instead of bikaverin pigmentation in F. fujikuroi (Studt et al. 2012), which emphasizes the importance of standardized growing medium recipes to strengthen reproducibility (Wiemann et al. 2009; Sørensen and Sondergaard 2014). Biological challenges in the form of co-cultivation with other microorganisms may activate silent biosynthetic pathways leading to an increased metabolite and mycotoxin production as well as changes in growth rate (Müller et al. 2012; Netzker et al. 2015). For instance, F. demicellulare shows enhanced production of fusaristatin A which inhibits the growth of its competitor (Li et al. 2016). The co-cultivation of F. tricinctum and B. subtilis enhances the formation of enniatins and fusaristatin A drastically, and induces the formation of three novel compounds: macrocarpon C, 2-(carboxymethylamino)benzoic acid and (−)-citreoisocoumarinol (Ola et al. 2013). This demonstrates the utility of this approach and confirms the role of SMs as competitive agents.
Transformation methods
The majority of studies mentioned in Table 1 have used genetic manipulation to create a link between genes and the formation of a specific biosynthetic metabolite. A prerequisite for the use of this approach in Fusarium metabolomics was to develop transformation protocols and tools. Protoplast-mediated transformation (PMT) is the most commonly used transformation system in filamentous fungi. Freshly germinated hyphae (Fig. 3a) are treated with commercially available enzymes to remove complex cell-wall components, resulting in the formation of protoplasts (Rodriguez-Iglesias and Schmoll 2015). The protoplasts are usually mixed in an osmotic stabilizing solution such as 1.2 M KCl or 1.2 M sorbitol containing CaCl2 (Fig. 3b). Protoplasts can be mixed with both circular plasmid or linearized double-stranded DNA. Calcium ions are added to open channels in the cytomembrane and thus promote uptake of nucleotides (Olmedo-Monfil et al. 2004). Polyethylene glycol (PEG) is added to promote fusion between exogenous nucleotides and protoplasts (Fig. 3c) (Becker and Lundblad 2001). Transformed protoplasts often require regeneration in osmotically stabilized medium before they are selected. PMT has been applied to transform several Fusarium species with high levels of success (Table 2). PMT has been observed to result in multicopy and ectopic integration events for some species of filamentous fungi (de Groot et al. 1998; Meyer 2008). Indeed, ectopic integration events have been reported in transformants from F. verticillioides (Proctor et al. 1999) and F. fujikuroi (Tudzynski et al. 1996) protoplasts. However, the frequency of homologous recombination-guided integration events depends not only on the transformation host strain in question, but also on the transformation protocol applied (Fernández-Martín et al. 2000).
Overview of protoplast-mediated transformation (a–c) and Agrobacterium tumefaciens-mediated transformation of Fusarium spp. (d). a Mycelial tissue comprising a thick cell wall. b Enzymatic treatment of mycelium releases protoplasts encapsulated by cytomembrane and no cell wall. c Protoplast transformation. Polyethylene glycol (PEG) can form a molecular bridge between cell and nucleotide. DNA uptake is possible through a porous membrane. d Overview of transfer-DNA delivery to the nucleus of Fusarium spp. conidia
The Gram-negative bacterium Agrobacterium tumefaciens is known for its ability to infect plants and during this process transfer the transfer-DNA (T-DNA) region of the Ti plasmid to the genome of the colonized host. The T-DNA regions are bordered by two imperfect inverted repeats (left and right border), and it is possible to introduce exogenous DNA by inserting it between the two border sites (Citovsky et al. 2007). A. tumefaciens is also capable of infecting filamentous fungi when induced by acetosyringone (Fig. 3d) (Idnurm et al. 2017), and a vast arsenal of binary vectors has been developed for this purpose (Frandsen 2011). The T-DNA is usually integrated in the fungal genome as a single copy by homologous recombination (Mullins et al. 2001; Michielse et al. 2005), and has been proven to be effective for targeted gene deletion mediated by homologous recombination (Kistler and Benny 1988; Idnurm et al. 2017). The major bottlenecks in this technique include the preparation of binary vectors and testing of various technical parameters, as an optimized protocol has to be developed for every species (de Groot et al. 1998; Utermark and Karlovsky 2008; Sørensen et al. 2014b). As for PMT, ATMT protocols have been developed for several representatives of Fusarium (Table 3).
Other less used transformation strategies for species of Fusarium include glass bead transformation (Singh and Rajam 2013), electroporation (Liang et al. 2014), and restriction enzyme-mediated integration (Linnemannstöns et al. 1999; Shim and Woloshuk 2001; Inoue et al. 2001; Han et al. 2004). For manipulation of novel strains, we recommend using or expanding the already developed vector systems and protocols reported in the literature.
Low hanging fruits: known secondary metabolites produced under laboratory conditions, identified through knock-out/gene disruption
In this section, we will show how metabolites identified in laboratory cultures can be linked to their underlying genetic machinery. To identify biosynthetic genes, a simple but effective strategy has been to identify putative gene candidates, deleting or disrupting the genes, and then determining the SM complement. The concept of gene replacement, sometimes referred to as knock-out, is based on genomic insertion of a DNA cassette guided by one or two border sequences homologous to the respective target gene or region. The amount of homologous nucleotides required for homologous recombination varies from species to species, but most Fusarium spp. can perform homologous recombination between insert and genome if the segments are 800–1500 bp long (Gaffoor et al. 2005; Oide et al. 2006; Frandsen et al. 2012; Wollenberg et al. 2017; Bahadoor et al. 2018). However, heterologous or ectopic recombination is a common phenomenon (Proctor et al. 1999; Malz et al. 2005) and genetic validation is required to confirm the correct integration of the deletion cassette, e.g., via Southern blot or diagnostic PCR analyses.
Disruption vectors containing a single segment homologous to the target gene are rapidly prepared and introduced into the fungal genome by a single cross-over recombination event guided by either PMT (Gaffoor et al. 2005; Gaffoor and Trail 2006; Brown et al. 2012) or ATMT (Malz et al. 2005) (Fig. 4a). Gaffoor et al. 2005 identified 15 PKS genes in F. graminearum and used RT-PCR to determine under what growth conditions 14 of the genes were expressed. The authors developed a single cross-over gene disruption vector system targeting 500–1750 bp downstream of the start codon of each PKS gene. The mutants provided evidence for PKS12 being responsible for the formation of the red mycelial pigment aurofusarin, PKS10 was responsible for the formation of fusarin C, and PKS3 provided the basis for perithecial pigments, later identified as the fusarubins (Studt et al. 2012) and bostrycoidin (Frandsen et al. 2016). PKS4 and PKS13 were shown to collectively synthesize the mycotoxin zearalenone (Gaffoor and Trail 2006; Lysøe et al. 2006). However, the remaining 11 PKS genes could not be correlated with a phenotype or metabolite (Gaffoor et al. 2005). Meanwhile, a similar approach was used for targeted gene disruption with a vector carrying a 833 bp fragment homologous to the KS domain of PKS12 in F. graminearum, which was introduced via ATMT with A. tumefaciens AGL1 (Malz et al. 2005). The resulting FgΔPKS12 strain showed no PKS12 expression and had an albino phenotype. In addition, F. graminearum was disrupted in the promoter region of aurR1, which turned out to act as a transcriptional activator of several genes in the PKS12 cluster. Later, a similar disruption vector was prepared, linearized, and transformed into protoplasts targeting the KS domain of PKS10 in F. verticillioides (Brown et al. 2012). Disruption mutant FvΔPKS10 provided evidence that this PKS was responsible for formation of fusarin C.
Targeted disruption and gene replacement strategies. a Gene disruption by integration with targeting plasmid-containing segment homologous to part of biosynthetic gene. b Gene replacement by two homologous recombination events replacing the entire open-reading frame with an insertion cassette. c Split-marker gene replacement in protoplasts transformed with two nucleotide fragments
A more popular disruption strategy is based on vectors containing two homologous segments to the target gene separated by a selection marker gene (Fig. 4b). This enables replacing a large portion or the entire biosynthetic gene with the selection marker. Gene replacement has been carried out in most Fusarium spp. guided by either PMT of F. fujikuroi and F. venenatum (Proctor et al. 1999; Song et al. 2004; Wiemann et al. 2009; Niehaus et al. 2014a; Janevska et al. 2016; Studt et al. 2016a) or ATMT of F. graminearum, F. pseudograminearum, F. solani, F. avenaceum, and F. semitectum (Frandsen et al. 2006, 2016; Tobiasen et al. 2007; Ma et al. 2010; Sørensen et al. 2014a, b; Romans-Fuertes et al. 2016; Wollenberg et al. 2017; Bahadoor et al. 2018).
In a study of linking a biosynthetic gene to the formation of fusarin C, Song et al. 2004 produced a knock-out vector able to replace PKS10 from F. venenatum through a double cross-over recombination event. However, Southern blot analysis of the transformed mutants showed that four different types of recombination events between circular plasmid and genome had occurred, all resulting in gene replacement or disruption and the inability to produce fusarin C (Song et al. 2004). The four recombination possibilities between genome and a vector carrying two segments homologous to target genes are: double cross-over leading to gene replacement, integration of the plasmid in the 5′ end of the gene by a single cross-over, integration in the 3′ end of the gene by a single cross-over event, or plasmid integration in both ends of the gene. This agrees with studies in F. verticillioides, where different recombination events have been observed. In one study, only one out of 16 mutants displayed correct gene replacement through double recombination, while 15 mutants had experienced integration of the entire vector in one or more copies, disrupting FvPKS24 (Proctor et al. 1999). Overall, 14% of screened F. pseudograminearum protoplast mutants carried a deletion of the virulence-related FpAH1 gene resulting from double cross-over events (Gardiner et al. 2012). Similarly, successful knock-outs of NRPS32 led to the identification of the novel lipopeptide W493 A and B (Sørensen et al. 2014a).
In contrast to transforming fungi with circular or linearized copies of knock-out vectors, some researchers choose to use the PCR-amplified knock-out cassette directly to minimize the risk of ectopic integration of plasmid backbone elements. This PCR-based PMT approach was used to study the zearalenone gene cluster in F. graminearum (Kim et al. 2005b) and the fusarubins BGC in F. fujikuroi (Studt et al. 2012). Catlett et al. (2003) introduced the split-marker system, where two PCR products each comprised a 3′ or 5′ homologous target region with each two-thirds of the selection marker, together capable of forming an intact deletion cassette when combined through homologous recombination (Fig. 4c) (Catlett et al. 2003; Chung and Lee 2015). Brown et al. (2012) demonstrated targeted gene replacement of FvPKS21 by split marker and PMT. This approach was utilized in the investigation of F. graminearum siderophore biosynthesis pathways (Oide et al. 2006, 2007, 2014) to create knock-outs of NRPS1, 2, and 6, and, furthermore, used for investigation of the aurofusarin pigmentation pathway in F. graminearum (Kim et al. 2005a). Extensive work has been performed describing biosynthetic pathways in F. fujikuroi, and gene replacement experiments have been the key to establishing links between BGCs and the pigment bikaverin (Wiemann et al. 2009), the plant mycotoxin fusaric acid (Studt et al. 2016a), and the peptide drug lead apicidin F (Niehaus et al. 2014a). In one study, knock-out mutants combined with isotopically labeled precursors led to the identification of FfPKS8 as the progenitor of gibepyrone biosynthesis (Janevska et al. 2016).
ATMT with A. tumefaciens strain LBA4404 is used for gene replacement experiments in Fusarium spp. with great success (Idnurm et al. 2017). Based on the ATMT protocol developed by Malz et al. (2005), a vector system for targeted gene deletion was established for F. graminearum (Frandsen et al. 2006, 2008, 2012), allowing characterization of the aurofusarin (PKS12) and fusarubins (PKS3) pigment biosynthesis. Furthermore, ATMT and gene replacement in F. graminearum enabled characterization of ferricrocin (NRPS2), fusaristatin (PKS6-NRPS7), chrysogine (NRPS14), and recently gramillins biosynthesis (NRPS8) (Tobiasen et al. 2007; Sørensen et al. 2014a; Wollenberg et al. 2017; Bahadoor et al. 2018). The F. graminearum ATMT protocol was adapted for F. avenaceum by testing different transformation parameters to identify optimal concentration of spores, co-inoculation time, and incubation temperature. Both F. graminearum and F. avenaceum displayed a high gene-targeting efficiency guided by homologous recombination between T-DNA and the gene of interest. In rare cases (1–10%), the T-DNA integrated ectopically in the recipient genome through non-homologous end-joining (Frandsen et al. 2012; Sørensen et al. 2014b). For gene disruption in F. solani and F. semitectum, ATMT protocols are available relying on the A. tumefaciens AGL-1 strain (Jin et al. 2010; Romans-Fuertes et al. 2016).
To cement the function of a gene, complementation can be carried out by reintroduction of the target gene into the disrupted or replaced gene mutant strains (Proctor et al. 2008; Studt et al. 2016a). Complementation of fumonisin production was carried out by transforming a deficient F. verticillioides strain with a gDNA cosmid library clone carrying PKS24, yielding mutants producing wild-type titers of fumonisins (Proctor et al. 1999). A feasible complementation technique for F. graminearum and F. semitectum has been to PCR amplify a wild-type gene allele including native promoter and terminator sequence in a single fragment which can be co-transformed into protoplasts together with a plasmid containing a selection marker different to what was used to disrupt the target gene originally (Kim et al. 2005a, b; Jin et al. 2010).
Gene replacement is a powerful tool to link genes to function and entire pathways can be resolved in this way. Not only the core synthase can be identified, but the contribution of the other genes in the same cluster to the final product can be determined (Frandsen et al. 2006, 2016; Wiemann et al. 2009; Studt et al. 2012, 2016a; Kakule et al. 2013). However, it is important to bear in mind that for a successful outcome of this strategy, the fungus must produce the target compound under the cultivation conditions used.
Targeted activation
With the introduction of sequencing, the identification of gene clusters has become trivial—but their silence is still a challenge. In consequence, targeted gene activation is used to discover new biosynthetic pathways in Fusarium spp. Core-synthase genes such as PKS and NRPS genes make ideal targets for targeted gene activation. A vector is prepared containing a constitutive promoter and a selection marker between two segments for targeted integration upstream of the biosynthetic gene in question (Fig. 5a). USER cloning has been demonstrated to enable quick assembly of such vectors for targeted promoter replacement in F. graminearum (Frandsen et al. 2008). The pRF-HU2E vector can be easily equipped with suitable homologous sequences upstream from the target gene, enabling promoter swapping to the constitutive A. nidulans PgdpA in front of PKSs and NRPSs. This activated production of gibepyrones A, B, D, and G and polypyrone B (PKS8) (Westphal et al. 2018a), chrysogine (NRPS14) (Wollenberg et al. 2017), orsellinic acid and orcinol (PKS14) (Jørgensen et al. 2014), and three novel bostrycoidin anthrones (PKS3) (Frandsen et al. 2016). Overexpression of FgNRPS4 leads to an increase in surface hydrophobicity, but no specific SM responsible for this phenotype could be identified by chemical analyses (Hansen et al. 2012a). Comparison of knock-out mutants to the wild type in the F. heterosporum PKS69 pathway failed to identify differences in the SM profile on different growing media. However, fusing a copy of the fsdS (PKS69) gene with the constitutive equisetin synthase (PKS18) promoter in a mutant construct resulted in formation of fusaridione A, which is likely the first intermediate in the biosynthetic pathway (Kakule et al. 2013).
Targeted activation can also aim to activate transcriptional regulator genes. Biosynthesis gene clusters often contain a Zn(II)2Cys6-domain gene that acts as a cluster-specific transcription factor (Brown et al. 2007; Brakhage 2013). Examples are the Gip2, Bik5, and Fsr6 transcription factors controlling pigment biosynthesis in F. graminearum and F. fujikuroi (Kim et al. 2006; Studt et al. 2012; Wiemann et al. 2013). Exchanging the native promoter of putative transcription factor APS2 for the β-tubulin promoter in F. semitectum resulted in upregulation of NRPS31 cluster genes and increased formation of apicidin (Jin et al. 2010). Likewise, overexpression of the fusaric acid cluster intrinsic Zn(II)2Cys6 transcription factor Fub10 upregulated gene expression of all cluster proteins including PKS21 (Fub1) and the NRPS-like enzyme Fub8, resulting in product formation. Deletion of the second cluster-specific transcription factor gene FUB12 abolished product derivatization (Studt et al. 2016a). Analysis of BGC promoter regions with the Regulatory Sequence Analysis Tool (RSAT) can reveal conserved transcription factor binding motifs, suggesting that expression is regulated by a single Zn(II)2Cys6 binuclear transcription factor (van Helden et al. 2000; Sørensen et al. 2012a; Sieber et al. 2014; Frandsen et al. 2016).
To ease the process of identifying overexpression mutants, one method has been to amplify the transcription factor genes including the native terminator and fusing it to the pRF-HUEA expression cassette with homologous targeting segments in the F. graminearum PKS12 locus, resulting in albino mutants (Fig. 5b) (Frandsen et al. 2008, 2016). This system was used to overexpress the putative transcription factor Fsr7, resulting in increased formation of three novel toxins: fusarielins F, G, and H (Sørensen et al. 2012a). To ensure high expression, targeted integration into a non-coding locus adjacent to the β-tubulin gene in F. graminearum (Josefsen et al. 2012) has been used for AurR1 overexpression, enabling overproduction of aurofusarin biosynthesis metabolites including novel putative shunt products (Westphal et al. 2018b). Combined overexpression of PKS39 and the cluster-specific transcription factor gave a tenfold increase in metabolite production and enabled characterization of a novel group of metabolites: fujikurins B, C, and D (Wiemann et al. 2013; Von Bargen et al. 2015). Fungal metabolites and their intermediates can be toxic to the producer and it may be necessary to use an inducible expression system. In F. fujikuroi, controlled overproduction of the silent trichosetin gene cluster was obtained by placing the cluster-specific transcription factor gene TF22 under regulation of the inducible, tetracycline-dependent tet-on promoter (Janevska et al. 2017). A novel strategy for production of silent SM genes was developed with elements of the highly producing equisetin polyketide BGC in F. heterosporum. The equisetin synthase eqxS is under regulation of the cluster-specific transcription factor eqxR. eqxR was fused with the inducible/leaky alcA promoter, and the bidirectional promoter peqxS was fused to lovB and lovC from the A. terrus lovastatin nonaketide biosynthesis cluster. Transformation of F. heterosporum with this multi-gene vector construct succeeded in production of the expected lovastatin precursor metabolites. This approach was similarly used to express an uncharacterized biosynthetic pathway in the endophytic fungus NRRL 50135, resulting in the isolation of the anti-tuberculosis agents pyrrolocins A and C (Kakule et al. 2015).
Activation through global regulators and histone modification
The secondary metabolism of filamentous fungi is controlled by complex regulatory network of proteins responding to environmental conditions such as substrate, pH, light and temperature, excellently reviewed by Axel A. Brakhage (2013). In contrast to cluster-specific transcriptional regulator proteins activating a small number of genes, global regulatory proteins control expression of secondary metabolism on higher level (Wiemann and Keller 2014). In approximately 40% of fungal gene clusters, there are no identifiable TF present (Brakhage 2013), and therefore, targeted activation strategies cannot be performed. In such cases, alternative strategies can be used such as manipulation of histone-modifying enzymes or global transcriptional regulator genes (Bok and Keller 2004; Butchko et al. 2012; Giese et al. 2013).
Deletion of the COMPASS protein Ccl1 in F. graminearum and F. fujikuroi significantly altered secondary metabolism (Studt et al. 2017). In both species, SMs produced by genes localized near telomeres were upregulated. These chromosomal regions often have low gene expression mediated by trimethylation of the H3K4 (H3K4me3) and proteins from the COMPASS complex (Palmer and Keller 2010; Zhao et al. 2014). Disruption of the heterochromatin methyltransferase Kmt6 led to transcriptional activation of four novel putative BGCs in F. fujikuroi, leading to isolation of a novel sesquiterpene (Studt et al. 2016b). Likewise, F. graminearum Δkmt6 displayed a drastic change in secondary metabolism profile (Connolly et al. 2013). In F. fujikuroi, the global regulator protein Sge1 is responsible for activation of a number of SM pathways including gibberellic acids, bikaverin, fumonisins, apicidin F, fusarins, and fusaric acid (Michielse et al. 2014, 2015; Studt et al. 2016a). Bikaverin biosynthesis is repressed in nitrogen-rich and ambient pH conditions. Expression of bik genes was repressed by pH-related transcription factor PacC, and a knock-out mutant ΔpacC displayed significant increase in expression of bikaverin cluster genes in comparison with wild-type F. fujikuroi (Wiemann et al. 2009). In addition, the overexpression of the global nitrogen regulator AreA in F. fujikuroi resulted in higher titers of bikaverin, even under repressing conditions (Linnemannstöns et al. 2002). However, global regulators do not result in activation of all biosynthetic genes. In the hunt for the FgNRPS5-NRPS9 product, the overexpression of the known regulator of secondary metabolism FgLaeA did not results in activation of the BGC. Instead, Jia et al. 2019 showed that the overexpression of the cluster-specific transcription factor fgm4 ectopically was required for the formation of the novel virulence factor fusaoctaxin A (Jia et al. 2019). Global regulators may suppress activated gene clusters, but probably cannot in themselves activate clusters. To ensure effectiveness, both types of activation are probably required for a successful outcome.
Chemical analysis
Identification of changes to the secondary metabolome by the aforementioned approaches relies on robust and accurate chemical analyses. High-performance liquid chromatography (HPLC) is often a chosen chemical separation of polyketides and non-ribosomal peptides due to their chemical properties, although gas chromatography was recently used in linking PKS8 to gibepyrones in F. fujikuroi (Janevska et al. 2016). Through the advances in high-resolution mass spectrometry (HRMS), several automated methods have been developed, which have become very helpful in deciphering which fungal SMs are produced (Nielsen et al. 2010). The identification of novel compounds from a complex fungal extract can be accelerated by fast and accurate identification of already known and characterized compounds. This can be achieved through dereplication, where chromatographic and spectroscopic methods are coupled with searches in existing databases (Nielsen et al. 2011). The usefulness of this approach was demonstrated by Klitgaard et al. (2014), who developed a method for automated identification of up to 3000 fungal secondary metabolites (Klitgaard et al. 2014).
Although dereplication holds great potential for working with SM discovery, it has not been widely adapted to studies of Fusarium. MS-based dereplication was used to determine the SM profiles among selected strains from F. solani species complex isolated from human infections (Short et al. 2013). This study led to the identification of several SMs, including citreoisocoumarin, YCM1008A, and lucilactaene, which had not been reported from members of the species complex before. NMR-based dereplication has been used on Fusarium strains isolated from the rhizosphere of Senna spectabilis in which fusaric acid and beauvericin were discovered in F. oxysporum and the depsipeptide HA23 in F. solani (Selegato et al. 2016).
The advances in chemical analyses have also enabled increased use of stable isotopes for elucidation of biosynthetic pathways. A feeding experiment in A. niger with fully labeled 13C8 6-methylsalicylic acid (6-MSA) was used to propose a biosynthetic route ending with yanuthone D as the end product (Holm et al. 2014). A similar approach was used in F. avenaceum, which was fed with 13C14-YWA1 (Klitgaard et al. 2015), the first stable intermediate formed during biosynthesis of aurofusarin. The feeding experiment in F. avenaceum showed that aurofusarin was derived from YWA1, but more interestingly, antibiotic Y (avenacein Y) was also identified as being derived from YWA1. This compound is well known from F. avenaceum, but had not been linked to a biosynthetic pathway. Subsequent genome analyses showed that F. avenaceum contains a gene (aurE, FAVG1_08663) located centrally in the aurofusarin gene cluster, which is not present in the aurofusarin gene cluster in F. graminearum. This gene is predicted to encode an epoxide hydrolase, which could be involved in antibiotic Y biosynthesis.
Outlook
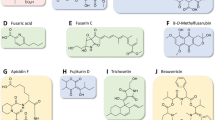
This review has summarized the recent advances in deciphering biosynthetic pathways in Fusarium. The quest is far from finished, as the products are currently unknown for the majority of gene clusters. Ongoing global gene-expression analyses will offer further insight into the biosynthesis of SMs through identification of co-regulated genes (Brown et al. 2012; Jørgensen et al. 2014). Proteomics may supplement this data, as shown in an analysis of an overexpression mutant in F. graminearum (Westphal et al. 2018b). The majority of metabolites are now identified through gene orthology rather than chemical isolation and analysis (Brown and Proctor 2016; Hoogendoorn et al. 2018). Gene homology analyses are now an inherent part of modern metabolomics research due to the strength and swift application of tools such as BLASTP (Altschul et al. 1990) and AntiSMASH (Blin et al. 2017), and resource collections, e.g., the Minimum Information about a Biosynthetic Gene cluster (MIBiG) repository (Epstein et al. 2018). Bioinformatic approaches have helped researchers to predict the product of a Fusarium BGC (Varga et al. 2005) before it was experimentally deduced (Tobiasen et al. 2007; Oide et al. 2014). Currently, a handful of putative metabolites has been assigned to species of Fusarium based on homology (Gaffoor et al. 2005; Hansen et al. 2012b, 2015; Wiemann et al. 2013; Brown and Proctor 2016; Hoogendoorn et al. 2018; Janevska and Tudzynski 2018). Metabolites assigned to Fusarium spp. through homology comprise: depudecin (PKS17), solanapyrone (PKS44), tenellin/fumosorinone (PKS45), alternapyrone (PKS52), 3-methylorsellinic acid (PKS54), oxononal benzaldehyde (PKS55-PKS64), mellein (PKS56), hexadehydro-astechrome (NRPS42), and fumarylalanine (NRPS43) (Hansen et al. 2015; Brown and Proctor 2016; Hoogendoorn et al. 2018). Although prediction tools have proven reliable, many of the predicted metabolites remain to be detected in the fungal organism by chemical analyses. Gene comparisons may be useful in risk assessment as exemplified by the observation of a putative mycotoxin producing synthase in the genome of the biological control strain F. oxysporum Fo47 (Hoogendoorn et al. 2018).
Looking ahead, the majority of detected Fusarium PKS and NRPS BGCs remain to be characterized (Hansen et al. 2015) and their product pathways elucidated. Deletion of biosynthetic gene-producing unwanted metabolites may increase the flux of precursors to a target pathway, resulting in higher yields of the desired compounds (Chiang et al. 2013). Elimination of endogenous biosynthesis pathways may also decrease metabolic background noise in LC–MS, and ease purification of novel products. To control the regulation of endogenous metabolism, two recent studies report novel Cre-loxP-based systems for F. graminearum enabling rapid and effective gene-targeting strategies and selection marker recycling (Connolly et al. 2018; Twaruschek et al. 2018). Major efforts have until now focused on key species such as F. graminearum and F. fujikuroi. The genomes of F. solani and F. avenaceum comprise several untapped PKS and NRPS BGCs not found elsewhere in the metagenome. With the introduction of optimized ATMT protocols for both (Sørensen et al. 2014b; Romans-Fuertes et al. 2016), we expect to contribute new information on SM products to the collected Fusarium metabolome. In addition, we expect that expression in a non-natural host (Munawar et al. 2013; Rugbjerg et al. 2013) will contribute to assessment of enzymatic function.
We hope that this review has shown that data and molecular tools are now available to get insight into the vast number of SMs in Fusarium and evaluate their potential as leads in the biomedical sciences and their impact in the environment.
References
Al-Reedy RM, Malireddy R, Dillman CB, Kennell JC (2012) Comparative analysis of Fusarium mitochondrial genomes reveals a highly variable region that encodes an exceptionally large open reading frame. Fungal Genet Biol 49:2–14. https://doi.org/10.1016/j.fgb.2011.11.008
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Apweiler R, Attwood TK, Bairoch A et al (2000) InterPro–an integrated documentation resource for protein families, domains and functional sites. Bioinformatics 16:1145–1150
Bachmann BO, Ravel J (2009) Methods for in silico prediction of microbial polyketide and nonribosomal peptide biosynthetic pathways from DNA sequence data. Methods Enzymol 458:181–217. https://doi.org/10.1016/S0076-6879(09)04808-3
Bahadoor A, Brauer EK, Bosnich W et al (2018) Gramillin A and B: cyclic lipopeptides identified as the nonribosomal biosynthetic products of Fusarium graminearum. J Am Chem Soc. https://doi.org/10.1021/jacs.8b10017
Becker DM, Lundblad V (2001) Introduction of DNA into yeast cells. Curr Protoc Mol Biol. https://doi.org/10.1002/0471142727.mb1307s27
Bentley R, Bennett JW (1999) Constructing polyketides: from collie to combinatorial biosynthesis. Annu Rev Microbiol 53:411–446. https://doi.org/10.1146/annurev.micro.53.1.411
Blin K, Wolf T, Chevrette MG et al (2017) AntiSMASH 4.0—improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res 45:W36–W41. https://doi.org/10.1093/nar/gkx319
Bode HB, Bethe B, Höfs R, Zeeck A (2002) Big effects from small changes: possible ways to explore nature’s chemical diversity. ChemBioChem 3:619. https://doi.org/10.1002/1439-7633(20020703)3:7%3c619:AID-CBIC619%3e3.0.CO;2-9
Bok J, Keller N (2004) LaeA, a regulator of secondary metabolism in Aspergillus spp. Eukaryot Cell 3:527–535. https://doi.org/10.1128/EC.3.2.527-535.2004
Bömke C, Tudzynski B (2009) Diversity, regulation, and evolution of the gibberellin biosynthetic pathway in fungi compared to plants and bacteria. Phytochemistry 70:1876–1893. https://doi.org/10.1016/j.phytochem.2009.05.020
Brakhage AA (2013) Regulation of fungal secondary metabolism. Nat Rev Microbiol 11:21–32. https://doi.org/10.1038/nrmicro2916
Brock NL, Huss K, Tudzynski B, Dickschat JS (2013) Genetic dissection of sesquiterpene biosynthesis by Fusarium fujikuroi. ChemBioChem 14:311–315. https://doi.org/10.1002/cbic.201200695
Brown DW, Proctor RH (2016) Insights into natural products biosynthesis from analysis of 490 polyketide synthases from Fusarium. Fungal Genet Biol 89:37–51. https://doi.org/10.1016/j.fgb.2016.01.008
Brown DW, Yu JH, Kelkar HS et al (1996) Twenty-five coregulated transcripts define a sterigmatocystin gene cluster in Aspergillus nidulans. Proc Natl Acad Sci USA 93:1418–1422
Brown DW, Butchko RAE, Busman M, Proctor RH (2007) The Fusarium verticillioides FUM gene cluster encodes a Zn(II)2Cys6 protein that affects FUM gene expression and fumonisin production. Eukaryot Cell 6:1210–1218. https://doi.org/10.1128/EC.00400-06
Brown DW, Butchko RAE, Busman M, Proctor RH (2012) Identification of gene clusters associated with fusaric acid, fusarin, and perithecial pigment production in Fusarium verticillioides. Fungal Genet Biol 49:521–532. https://doi.org/10.1016/j.fgb.2012.05.010
Brown DW, Busman M, Proctor RH (2014) Fusarium verticillioides SGE1 is required for full virulence and regulates expression of protein effector and secondary metabolite biosynthetic genes. Mol Plant Microbe Interact 27:809–823. https://doi.org/10.1094/MPMI-09-13-0281-R
Burkhardt I, Siemon T, Henrot M et al (2016) Mechanistic characterisation of two sesquiterpene cyclases from the plant pathogenic fungus Fusarium fujikuroi. Angew Chem Int Ed 55:8748–8751. https://doi.org/10.1002/anie.201603782
Butchko RAE, Brown DW, Busman M et al (2012) Lae1 regulates expression of multiple secondary metabolite gene clusters in Fusarium verticillioides. Fungal Genet Biol 49:602–612. https://doi.org/10.1016/j.fgb.2012.06.003
Catlett NL, Lee B-N, Yoder OC, Turgeon BG (2003) Split-marker recombination for efficient targeted deletion of fungal genes. Fungal Genet Rep 50:9–11. https://doi.org/10.4148/1941-4765.1150
Challis GL, Ravel J, Townsend CA (2000) Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem Biol 7:211–224
Chiang Y-M, Oakley CE, Ahuja M et al (2013) An efficient system for heterologous expression of secondary metabolite genes in Aspergillus nidulans. J Am Chem Soc 135:7720–7731. https://doi.org/10.1021/ja401945a
Chung KR, Lee MH (2015) Split-marker-mediated transformation and targeted gene disruption in filamentous fungi. In: van den Berg M, Maruthachalam K (eds) Genetic transformation systems in fungi, vol 2. Fungal Biology. Springer, Cham. https://doi.org/10.1007/978-3-319-10503-1_15
Citovsky V, Kozlovsky SV, Lacroix B et al (2007) Biological systems of the host cell involved in Agrobacterium infection. Cell Microbiol 9:9–20. https://doi.org/10.1111/j.1462-5822.2006.00830.x
Connolly LR, Smith KM, Freitag M, Madhani HD (2013) The Fusarium graminearum histone H3 K27 methyltransferase KMT6 regulates development and expression of secondary metabolite gene clusters. PLoS Genet 9 10). https://doi.org/10.1371/journal.pgen.1003916
Connolly LR, Erlendson AA, Fargo CM et al (2018) Application of the Cre/lox system to construct auxotrophic markers for quantitative genetic analyses in Fusarium graminearum. Methods Mol Biol 1848:235–263. https://doi.org/10.1007/978-1-4939-8724-5_16
Conti E, Stachelhaus T, Marahiel MA, Brick P (1997) Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. EMBO J 16:4174–4183
Covert SF, Kapoor P, Lee M et al (2001) Agrobacterium tumefaciens-mediated transformation of Fusarium circinatum. Mycol Res 105:259–264. https://doi.org/10.1017/S0953756201003872
Crowhurst RN, Rees-George J, Rikkerink EH, Templeton MD (1992) High efficiency transformation of Fusarium solani f. sp. cucurbitae race 2 (mating population V). Curr Genet 21:463–469
Cuomo CA, Güldener U, Xu J-R et al (2007) The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 317:1400–1402. https://doi.org/10.1126/science.1143708
de Groot MJA, Bundock P, Hooykaas PJ, Beijersbergen AGM (1998) Agrobacterium tumefaciens-mediated transformation of filamentous fungi. Nat Biotechnol 16:839–842. https://doi.org/10.1038/nbt0998-839
Dean R, Van Kan JAL, Pretorius ZA et al (2012) The Top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathol 13:414–430. https://doi.org/10.1111/j.1364-3703.2011.00783.x
Desjardins AE, Proctor RH (2007) Molecular biology of Fusarium mycotoxins. Int J Food Microbiol 119:47–50. https://doi.org/10.1016/j.ijfoodmicro.2007.07.024
Ding M, Li J, Fan X et al (2018) Aquaporin1 regulates development, secondary metabolism and stress responses in Fusarium graminearum. Curr Genet 64:1057–1069. https://doi.org/10.1007/s00294-018-0818-8
Epstein SC, Charkoudian LK, Medema MH (2018) A standardized workflow for submitting data to the Minimum Information about a Biosynthetic Gene cluster (MIBiG) repository: prospects for research-based educational experiences. Stand Genomic Sci 13:1–13. https://doi.org/10.1186/s40793-018-0318-y
Fernández-Martín R, Cerdá-Olmedo E, Avalos J (2000) Homologous recombination and allele replacement in transformants of Fusarium fujikuroi. Mol Gen Genet 263:838–845
Finking R, Marahiel MA (2004) Biosynthesis of nonribosomal peptides. Annu Rev Microbiol 58:453–488. https://doi.org/10.1146/annurev.micro.58.030603.123615
Frandsen RJN (2011) A guide to binary vectors and strategies for targeted genome modification in fungi using Agrobacterium tumefaciens-mediated transformation. J Microbiol Methods 87:247–262. https://doi.org/10.1016/j.mimet.2011.09.004
Frandsen RJN, Nielsen NJ, Maolanon N et al (2006) The biosynthetic pathway for aurofusarin in Fusarium graminearum reveals a close link between the naphthoquinones and naphthopyrones. Mol Microbiol 61:1069–1080. https://doi.org/10.1111/j.1365-2958.2006.05295.x
Frandsen RJN, Andersson JA, Kristensen MB, Giese H (2008) Efficient four fragment cloning for the construction of vectors for targeted gene replacement in filamentous fungi. BMC Mol Biol 9:70. https://doi.org/10.1186/1471-2199-9-70
Frandsen RJN, Frandsen M, Giese H (2012) Targeted gene replacement in fungal pathogens via Agrobacterium tumefaciens-mediated transformation. Methods Mol Biol 835:17–45. https://doi.org/10.1007/978-1-61779-501-5_2
Frandsen RJN, Rasmussen SA, Knudsen PB et al (2016) Black perithecial pigmentation in Fusarium species is due to the accumulation of 5-deoxybostrycoidin-based melanin. Sci Rep 6:26206. https://doi.org/10.1038/srep26206
Gaffoor I, Trail F (2006) Characterization of two polyketide synthase genes involved in zearalenone biosynthesis in Gibberella zeae. Appl Environ Microbiol 72:1793–1799. https://doi.org/10.1128/AEM.72.3.1793-1799.2006
Gaffoor I, Brown DW, Plattner R, Proctor RH (2005) Functional analysis of the polyketide synthase genes in the filamentous fungus. Eukaryot Cell 4:1926–1933. https://doi.org/10.1128/ec.4.11.1926
Gardiner DM, McDonald MC, Covarelli L et al (2012) Comparative pathogenomics reveals horizontally acquired novel virulence genes in fungi infecting cereal hosts. PLoS Pathog 8:e1002952. https://doi.org/10.1371/journal.ppat.1002952
Gardiner DM, Stiller J, Kazan K (2014) Genome sequence of Fusarium graminearum isolate CS3005. Genome Announc 2:e00227–14. https://doi.org/10.1128/genomeA.00227-14
Giese H, Sondergaard TE, Sørensen JL (2013) The AreA transcription factor in Fusarium graminearum regulates the use of some nonpreferred nitrogen sources and secondary metabolite production. Fungal Biol 117:814–821. https://doi.org/10.1016/j.funbio.2013.10.006
Gomez-Gil L, Camara Almiron J, Rodriguez Carrillo PL et al (2018) Nitrate assimilation pathway (NAP): role of structural (nit) and transporter (ntr1) genes in Fusarium oxysporum f. sp. lycopersici growth and pathogenicity. Curr Genet 64:493–507. https://doi.org/10.1007/s00294-017-0766-8
Gordon TR, Martyn RD (1997) The evolutionary biology of Fusarium oxysporum. Annu Rev Phytopathol 35:111–128. https://doi.org/10.1146/annurev.phyto.35.1.111
Graziani S, Vasnier C, Daboussi MJ (2004) Novel polyketide synthase from Nectria haematococca. Appl Environ Microbiol 70:2984–2988. https://doi.org/10.1128/AEM.70.5.2984-2988.2004
Haese A, Schubert M, Herrmann M, Zocher R (1993) Molecular characterization of the enniatin synthetase gene encoding a multifunctional enzyme catalysing N-methyldepsipeptide formation in Fusarium scirpi. Mol Microbiol 7:905–914. https://doi.org/10.1111/j.1365-2958.1993.tb01181.x
Han Y-K, Lee T, Han K-H et al (2004) Functional analysis of the homoserine O-acetyltransferase gene and its identification as a selectable marker in Gibberella zeae. Curr Genet 46:205–212. https://doi.org/10.1007/s00294-004-0528-2
Hansen FT, Droce A, Sørensen JL et al (2012a) Overexpression of NRPS4 leads to increased surface hydrophobicity in Fusarium graminearum. Fungal Biol 116:855–862. https://doi.org/10.1016/j.funbio.2012.04.014
Hansen FT, Sørensen JL, Giese H et al (2012b) Quick guide to polyketide synthase and nonribosomal synthetase genes in Fusarium. Int J Food Microbiol 155:128–136. https://doi.org/10.1016/j.ijfoodmicro.2012.01.018
Hansen FT, Gardiner DM, Lysøe E et al (2015) An update to polyketide synthase and non-ribosomal synthetase genes and nomenclature in Fusarium. Fungal Genet Biol 75:20–29. https://doi.org/10.1016/j.fgb.2014.12.004
Hemphill CFP, Sureechatchaiyan P, Kassack MU et al (2017) OSMAC approach leads to new fusarielin metabolites from Fusarium tricinctum. J Antibiot (Tokyo) 70:726–732. https://doi.org/10.1038/ja.2017.21
Holm DK, Petersen LM, Klitgaard A et al (2014) Molecular and chemical characterization of the biosynthesis of the 6-MSA-derived meroterpenoid yanuthone D in Aspergillus niger. Chem Biol 21:519–529. https://doi.org/10.1016/j.chembiol.2014.01.013
Hoogendoorn K, Barra L, Waalwijk C et al (2018) Evolution and diversity of biosynthetic gene clusters in Fusarium. Front Microbiol 9:1–12. https://doi.org/10.3389/fmicb.2018.01158
Idnurm A, Bailey AM, Cairns TC et al (2017) A silver bullet in a golden age of functional genomics: the impact of Agrobacterium-mediated transformation of fungi. Fungal Biol Biotechnol 4:6. https://doi.org/10.1186/s40694-017-0035-0
Inoue L, Ohara T, Nakami F, Tsuge T (2001) Isolation of pathogenicity mutants of Fusarium oxysporum f. sp. melonis by insertional mutagenesis. J Gen Plant Pathol 67:191–199. https://doi.org/10.1007/PL00013010
Janevska S, Tudzynski B (2018) Secondary metabolism in Fusarium fujikuroi: strategies to unravel the function of biosynthetic pathways. Appl Microbiol Biotechnol 102:615–630. https://doi.org/10.1007/s00253-017-8679-5
Janevska S, Arndt B, Niehaus E-M et al (2016) Gibepyrone biosynthesis in the rice pathogen Fusarium fujikuroi is facilitated by a small polyketide synthase gene cluster. J Biol Chem 291:27403–27420. https://doi.org/10.1074/jbc.M116.753053
Janevska S, Arndt B, Baumann L et al (2017) Establishment of the inducible Tet-on system for the activation of the silent trichosetin gene cluster in Fusarium fujikuroi. Toxins (Basel) 9:E126. https://doi.org/10.3390/toxins9040126
Jia L-J, Tang H-Y, Wang W-Q et al (2019) A linear nonribosomal octapeptide from Fusarium graminearum facilitates cell-to-cell invasion of wheat. Nat Commun 10:922. https://doi.org/10.1038/s41467-019-08726-9
Jin J-M, Lee S, Lee J et al (2010) Functional characterization and manipulation of the apicidin biosynthetic pathway in Fusarium semitectum. Mol Microbiol 76:456–466. https://doi.org/10.1111/j.1365-2958.2010.07109.x
Jørgensen SH, Frandsen RJN, Nielsen KF et al (2014) Fusarium graminearum PKS14 is involved in orsellinic acid and orcinol synthesis. Fungal Genet Biol 70:24–31. https://doi.org/10.1016/j.fgb.2014.06.008
Josefsen L, Droce A, Sondergaard TE et al (2012) Autophagy provides nutrients for nonassimilating fungal structures and is necessary for plant colonization but not for infection in the necrotrophic plant pathogen Fusarium graminearum. Autophagy 8:326–337. https://doi.org/10.4161/auto.8.3.18705
Kakule TB, Sardar D, Lin Z, Schmidt EW (2013) Two related pyrrolidinedione synthetase loci in Fusarium heterosporum ATCC 74349 produce divergent metabolites. ACS Chem Biol 8:1549–1557. https://doi.org/10.1021/cb400159f
Kakule TB, Jadulco RC, Koch M et al (2015) Native promoter strategy for high-yielding synthesis and engineering of fungal secondary metabolites. ACS Synth Biol 4:625–633. https://doi.org/10.1021/sb500296p
Keller NP (2019) Fungal secondary metabolism: regulation, function and drug discovery. Nat Rev Microbiol 17:167–180. https://doi.org/10.1038/s41579-018-0121-1
Keller NP, Hohn TM, Keller Hohn (1997) Metabolic pathway gene clusters in filamentous fungi. Fungal Genet Biol 21:17–29. https://doi.org/10.1006/fgbi.1997.0970
Khaldi N, Seifuddin FT, Turner G et al (2010) SMURF: genomic mapping of fungal secondary metabolite clusters. Fungal Genet Biol 47:736–741. https://doi.org/10.1016/j.fgb.2010.06.003
Khayatt BI, Overmars L, Siezen RJ, Francke C (2013) Classification of the adenylation and acyl-transferase activity of NRPS and PKS systems using ensembles of substrate specific hidden Markov models. PLoS One 8:e62136. https://doi.org/10.1371/journal.pone.0062136
Kim JE, Han KH, Jin J et al (2005a) Putative polyketide synthase and laccase genes for biosynthesis of aurofusarin in Gibberella zeae. Appl Environ Microbiol 71:1701–1708. https://doi.org/10.1128/AEM.71.4.1701-1708.2005
Kim YT, Lee YR, Jin J et al (2005b) Two different polyketide synthase genes are required for synthesis of zearalenone in Gibberella zeae. Mol Microbiol 58:1102–1113. https://doi.org/10.1111/j.1365-2958.2005.04884.x
Kim J-E, Jin J, Kim H et al (2006) GIP2, a putative transcription factor that regulates the aurofusarin biosynthetic gene cluster in Gibberella zeae. Appl Environ Microbiol 72:1645–1652. https://doi.org/10.1128/AEM.72.2.1645-1652.2006
King R, Urban M, Hammond-Kosack MCU et al (2015) The completed genome sequence of the pathogenic ascomycete fungus Fusarium graminearum. BMC Genomics 16:544. https://doi.org/10.1186/s12864-015-1756-1
Kistler HC, Benny UK (1988) Genetic transformation of the fungal plant wilt pathogen, Fusarium oxysporum. Curr Genet 13:145–149. https://doi.org/10.1007/BF00365649
Klitgaard A, Iversen A, Andersen MR et al (2014) Aggressive dereplication using UHPLC–DAD–QTOF: screening extracts for up to 3000 fungal secondary metabolites. Anal Bioanal Chem 406:1933–1943. https://doi.org/10.1007/s00216-013-7582-x
Klitgaard A, Frandsen RJN, Holm DK et al (2015) Combining UHPLC-high resolution MS and feeding of stable isotope labeled polyketide intermediates for linking precursors to end products. J Nat Prod 78:1518–1525. https://doi.org/10.1021/np500979d
Knudsen M, Søndergaard D, Tofting-Olesen C et al (2016) Computational discovery of specificity-conferring sites in non-ribosomal peptide synthetases. Bioinformatics 32:325–329. https://doi.org/10.1093/bioinformatics/btv600
Kroken S, Glass NL, Taylor JW et al (2003) Phylogenomic analysis of type I polyketide synthase genes in pathogenic and saprobic ascomycetes. Proc Natl Acad Sci 100:15670–15675. https://doi.org/10.1073/pnas.2532165100
Leslie JF, Summerell BA (eds) (2006) The Fusarium Laboratory Manual. Blackwell Publishing, Ames
Li G, Kusari S, Golz C et al (2016) Three cyclic pentapeptides and a cyclic lipopeptide produced by endophytic: Fusarium decemcellulare LG53. RSC Adv 6:54092–54098. https://doi.org/10.1039/c6ra10905e
Liang L, Li J, Cheng L et al (2014) A high efficiency gene disruption strategy using a positive–negative split selection marker and electroporation for Fusarium oxysporum. Microbiol Res 169:835–843. https://doi.org/10.1016/j.micres.2014.03.004
Linnemannstöns P, Voss T, Hedden P et al (1999) Deletions in the gibberellin biosynthesis gene cluster of Gibberella fujikuroi by restriction enzyme-mediated integration and conventional transformation-mediated mutagenesis. Appl Environ Microbiol 65:2558–2564
Linnemannstöns P, Schulte J, Del Mar Prado M et al (2002) The polyketide synthase gene pks4 from Gibberella fujikuroi encodes a key enzyme in the biosynthesis of the red pigment bikaverin. Fungal Genet Biol 37:134–148. https://doi.org/10.1016/S1087-1845(02)00501-7
Liuzzi V, Mirabelli V, Cimmarusti M et al (2017) Enniatin and beauvericin biosynthesis in Fusarium species: production profiles and structural determinant prediction. Toxins (Basel) 9:45. https://doi.org/10.3390/toxins9020045
Lysøe E, Klemsdal SS, Bone KR et al (2006) The PKS4 gene of Fusarium graminearum is essential for zearalenone production. Appl Environ Microbiol 72:3924–3932. https://doi.org/10.1128/AEM.00963-05
Lysøe E, Harris LJ, Walkowiak S et al (2014) The genome of the generalist plant pathogen Fusarium avenaceum is enriched with genes involved in redox, signaling and secondary metabolism. PLoS One. https://doi.org/10.1371/journal.pone.0112703
Ma LJ, Van Der Does HC, Borkovich KA et al (2010) Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464:367–373. https://doi.org/10.1038/nature08850
Ma L-J, Shea T, Young S et al (2014) Genome sequence of Fusarium oxysporum f. sp. melonis strain NRRL 26406, a fungus causing wilt disease on melon. Genome Announc 2:2013–2014. https://doi.org/10.1128/genomeA.00730-14
Malz S, Grell MN, Thrane C et al (2005) Identification of a gene cluster responsible for the biosynthesis of aurofusarin in the Fusarium graminearum species complex. Fungal Genet Biol 42:420–433. https://doi.org/10.1016/j.fgb.2005.01.010
Marasas WFO, van Rensburg SJ, Mirocha CJ (1979) Incidence of Fusarium species and the mycotoxins, deoxynivalenol and zearalenone, in corn produced in esophageal cancer areas in Transkei. J Agric Food Chem 27:1108–1112. https://doi.org/10.1021/jf60225a013
Marek ET, Schardl CL, Smith DA (1989) Molecular transformation of Fusarium solani with an antibiotic resistance marker having no fungal DNA homology. Curr Genet 15:421–428
McDaniel R, Ebert-Khosla S, Fu H et al (1994) Engineered biosynthesis of novel polyketides: influence of a downstream enzyme on the catalytic specificity of a minimal aromatic polyketide synthase. Proc Natl Acad Sci USA 91:11542–11546
Mcmullen M, Jones R, Gallenberg D (1997) Scab of wheat and barley: a re-emerging disease of devastating impact. Plant Dis 81:1340–1348
Meier JL, Burkart MD (2009) The chemical biology of modular biosynthetic enzymes. Chem Soc Rev 38:2012. https://doi.org/10.1039/b805115c
Meyer V (2008) Genetic engineering of filamentous fungi—progress, obstacles and future trends. Biotechnol Adv 26:177–185. https://doi.org/10.1016/j.biotechadv.2007.12.001
Michielse CB, Rep M (2009) Pathogen profile update: Fusarium oxysporum. Mol Plant Pathol 10:311–324. https://doi.org/10.1111/j.1364-3703.2009.00538.x
Michielse CB, Hooykaas PJJ, van den Hondel CAMJJ, Ram AFJ (2005) Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Curr Genet 48:1–17. https://doi.org/10.1007/s00294-005-0578-0
Michielse CB, Pfannmüller A, Macios M et al (2014) The interplay between the GATA transcription factors AreA, the global nitrogen regulator and AreB in Fusarium fujikuroi. Mol Microbiol 91:472–493. https://doi.org/10.1111/mmi.12472
Michielse CB, Studt L, Janevska S et al (2015) The global regulator FfSge1 is required for expression of secondary metabolite gene clusters but not for pathogenicity in Fusarium fujikuroi. Environ Microbiol 17:2690–2708. https://doi.org/10.1111/1462-2920.12592
Moolhuijzen PM, Manners JM, Wilcox SA et al (2013) Genome sequences of six wheat-infecting Fusarium species isolates. Genome Announc 1:1. https://doi.org/10.1128/genomea.00670-13
Müller MEH, Steier I, Köppen R et al (2012) Cocultivation of phytopathogenic Fusarium and Alternaria strains affects fungal growth and mycotoxin production. J Appl Microbiol 113:874–887. https://doi.org/10.1111/j.1365-2672.2012.05388.x
Mullins ED, Chen X, Romaine P et al (2001) Agrobacterium-mediated transformation of Fusarium oxysporum: an efficient tool for insertional mutagenesis and gene transfer. Phytopathology 91:173–180. https://doi.org/10.1094/PHYTO.2001.91.2.173
Munawar A, Marshall JW, Cox RJ et al (2013) Isolation and characterisation of a ferrirhodin synthetase gene from the sugarcane pathogen Fusarium sacchari. ChemBioChem 14:388–394. https://doi.org/10.1002/cbic.201200587
Naseema A, Dhanya B, Anjanadevi IP et al (2008) Isolation and regeneration of protoplasts from the mycelium of Fusarium pallidoroseum—a potential biocontrol agent of water hyacinth [Eichhornia crassipes (Mart.) Solms]. J Trop Agric 46:55–57
Netzker T, Fischer J, Weber J et al (2015) Microbial communication leading to the activation of silent fungal secondary metabolite gene clusters. Front Microbiol 6:1–13. https://doi.org/10.3389/fmicb.2015.00299
Niehaus E-M, Janevska S, Von Bargen KW et al (2014a) Apicidin F: characterization and genetic manipulation of a new secondary metabolite gene cluster in the rice pathogen Fusarium fujikuroi. PLoS One 9:1. https://doi.org/10.1371/journal.pone.0103336
Niehaus E-M, Von Bargen KW, Espino JJ et al (2014b) Characterization of the fusaric acid gene cluster in Fusarium fujikuroi. Appl Microbiol Biotechnol 98:1749–1762. https://doi.org/10.1007/s00253-013-5453-1
Niehaus E-M, Kim HK, Münsterkötter M et al (2017) Comparative genomics of geographically distant Fusarium fujikuroi isolates revealed two distinct pathotypes correlating with secondary metabolite profiles. PLoS Pathog 13:e1006670
Nielsen NJ, Tomasi G, Frandsen RJN et al (2010) A pre-processing strategy for liquid chromatography time-of-flight mass spectrometry metabolic fingerprinting data. Metabolomics 6:341–352. https://doi.org/10.1007/s11306-010-0211-1
Nielsen KF, Månsson M, Rank C et al (2011) Dereplication of microbial natural products by LC-DAD-TOFMS. J Nat Prod 74:2338–2348. https://doi.org/10.1021/np200254t
Nihei K, Itoh H, Hashimato K et al (1998) Antifungal cyclodepsipeptides, W493 A and B, from Fusarium sp.: isolation and structural determination. Biosci Biotechnol Biochem 62:858–863. https://doi.org/10.1271/bbb.62.858
Nucci M, Anaissie E (2007) Fusarium infections in immunocompromised patients. Clin Microbiol Rev 20:695–704. https://doi.org/10.1128/CMR.00014-07
Oide S, Moeder W, Krasnoff S et al (2006) NPS6, encoding a nonribosomal peptide synthetase involved in siderophore-mediated iron metabolism, is a conserved virulence determinant of plant pathogenic ascomycetes. Plant Cell Online 18:2836–2853. https://doi.org/10.1105/tpc.106.045633
Oide S, Krasnoff SB, Gibson DM, Turgeon BG (2007) Intracellular siderophores are essential for ascomycete sexual development in heterothallic Cochliobolus heterostrophus and homothallic Gibberella zeae. Eukaryot Cell 6:1339–1353. https://doi.org/10.1128/EC.00111-07
Oide S, Berthiller F, Wiesenberger G et al (2014) Individual and combined roles of malonichrome, ferricrocin, and TAFC siderophores in Fusarium graminearum pathogenic and sexual development. Front Microbiol 5:1–15. https://doi.org/10.3389/fmicb.2014.00759
Ola ARB, Thomy D, Lai D et al (2013) Inducing secondary metabolite production by the endophytic fungus Fusarium tricinctum through coculture with Bacillus subtilis. J Nat Prod 76:2094–2099. https://doi.org/10.1021/np400589h
Olmedo-Monfil V, Cortés-Penagos C, Herrera-Estrella A (2004) Three decades of fungal transformation: key concepts and applications. Recombinant gene expression. Humana Press, New Jersey, pp 297–314
Palmer JM, Keller NP (2010) Secondary metabolism in fungi: does chromosomal location matter? Curr Opin Microbiol 13:431–436. https://doi.org/10.1016/j.mib.2010.04.008
Ponts N, Richard-Forget F, Zhang H et al (2018) Genome sequence of the emerging mycotoxin-producing filamentous fungus Fusarium tricinctum strain INRA104. Genome Announc. https://doi.org/10.1128/genomea.00509-18
Proctor RH, Desjardins AE, Plattner RD, Hohn TM (1999) A polyketide synthase gene required for biosynthesis of fumonisin mycotoxins in Gibberella fujikuroi mating population A. Fungal Genet Biol 27:100–112. https://doi.org/10.1006/fgbi.1999.1141
Proctor RH, Busman M, Seo JA et al (2008) A fumonisin biosynthetic gene cluster in Fusarium oxysporum strain O-1890 and the genetic basis for B versus C fumonisin production. Fungal Genet Biol 45:1016–1026. https://doi.org/10.1016/j.fgb.2008.02.004
Rausch C (2005) Specificity prediction of adenylation domains in nonribosomal peptide synthetases (NRPS) using transductive support vector machines (TSVMs). Nucleic Acids Res 33:5799–5808. https://doi.org/10.1093/nar/gki885
Rodriguez-Iglesias A, Schmoll M (2015) Protoplast transformation for genome manipulation in fungi. Fungal Biol 1:21–40. https://doi.org/10.1007/978-3-319-10142-2_2
Romans-Fuertes P, Sondergaard TE, Sandmann MIH et al (2016) Identification of the non-ribosomal peptide synthetase responsible for biosynthesis of the potential anti-cancer drug sansalvamide in Fusarium solani. Curr Genet 62:799–807. https://doi.org/10.1007/s00294-016-0584-4
Röttig M, Medema MH, Blin K et al (2011) NRPSpredictor2—a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res 39:W362–W367. https://doi.org/10.1093/nar/gkr323
Rugbjerg P, Naesby M, Mortensen UH, Frandsen RJ (2013) Reconstruction of the biosynthetic pathway for the core fungal polyketide scaffold rubrofusarin in Saccharomyces cerevisiae. Microb Cell Fact 12:31. https://doi.org/10.1186/1475-2859-12-31
Salch YP, Beremand MN (1993) Gibberella pulicaris transformants: state of transforming DNA during asexual and sexual growth. Curr Genet 23:343–350. https://doi.org/10.1007/BF00310897
Selegato DM, Freire RT, Tannús A, Castro-Gamboa I (2016) New dereplication method applied to NMR-based metabolomics on different Fusarium species isolated from rhizosphere of Senna spectabilis. J Braz Chem Soc. https://doi.org/10.5935/0103-5053.20160139
Shim W-B, Woloshuk CP (2001) Regulation of fumonisin B1 biosynthesis and conidiation in Fusarium verticillioides by a cyclin-like (C-type) gene, FCC1. Appl Environ Microbiol 67:1607–1612. https://doi.org/10.1128/AEM.67.4.1607-1612.2001
Shiono Y, Tsuchinari M, Shimanuki K et al (2007) Fusaristatins A and B, two new cyclic lipopeptides from an endophytic Fusarium sp. J Antibiot (Tokyo) 60:309–316. https://doi.org/10.1038/ja.2007.39
Short DPG, O’Donnell K, Thrane U et al (2013) Phylogenetic relationships among members of the Fusarium solani species complex in human infections and the descriptions of F. keratoplasticum sp. nov. and F. petroliphilum stat. nov. Fungal Genet Biol 53:59–70. https://doi.org/10.1016/j.fgb.2013.01.004
Sieber CMK, Lee W, Wong P et al (2014) The Fusarium graminearum genome reveals more secondary metabolite gene clusters and hints of horizontal gene transfer. PLoS One. https://doi.org/10.1371/journal.pone.0110311
Singh N, Rajam MV (2013) A simple and rapid glass bead transformation method for a filamentous fungus Fusarium oxysporum. Cell Dev Biol 2(2). https://doi.org/10.4172/2168-9296.1000115
Soliday CL, Dickman MB, Kolattukudy PE (1989) Structure of the cutinase gene and detection of promoter activity in the 5′-flanking region by fungal transformation. J Bacteriol 171:1942–1951
Song Z, Cox RJ, Lazarus CM, Simpson TJ (2004) Fusarin C biosynthesis in Fusarium moniliforme and Fusarium venenatum. ChemBioChem 5:1196–1203. https://doi.org/10.1002/cbic.200400138
Sørensen JL, Sondergaard TE (2014) The effects of different yeast extracts on secondary metabolite production in Fusarium. Int J Food Microbiol 170:55–60. https://doi.org/10.1016/j.ijfoodmicro.2013.10.024
Sørensen JL, Hansen FT, Sondergaard TE et al (2012a) Production of novel fusarielins by ectopic activation of the polyketide synthase 9 cluster in Fusarium graminearum. Environ Microbiol 14:1159–1170. https://doi.org/10.1111/j.1462-2920.2011.02696.x
Sørensen JL, Nielsen KF, Sondergaard TE (2012b) Redirection of pigment biosynthesis to isocoumarins in Fusarium. Fungal Genet Biol 49:613–618. https://doi.org/10.1016/j.fgb.2012.06.004
Sørensen JL, Sondergaard TE, Covarelli L et al (2014a) Identification of the biosynthetic gene clusters for the lipopeptides fusaristatin A and W493 B in Fusarium graminearum and F. pseudograminearum. J Nat Prod 77:2619–2625. https://doi.org/10.1021/np500436r
Sørensen LQ, Larsen JE, Khorsand-Jamal P et al (2014b) Genetic transformation of Fusarium avenaceum by Agrobacterium tumefaciens mediated transformation and the development of a USER-Brick vector construction system. BMC Mol Biol 15:15. https://doi.org/10.1186/1471-2199-15-15
Stachelhaus T, Mootz HD, Marahiel MA (1999) The specificity-conferring code of adenylation domains in nonribosomal peptide synthetases. Chem Biol 6:493–505. https://doi.org/10.1016/S1074-5521(99)80082-9
Strieker M, Tanović A, Marahiel MA (2010) Nonribosomal peptide synthetases: structures and dynamics. Curr Opin Struct Biol 20:234–240. https://doi.org/10.1016/j.sbi.2010.01.009
Studt L, Wiemann P, Kleigrewe K et al (2012) Biosynthesis of fusarubins accounts for pigmentation of Fusarium Fujikuroi perithecia. Appl Environ Microbiol 78:4468–4480. https://doi.org/10.1128/AEM.00823-12
Studt L, Janevska S, Niehaus E-M et al (2016a) Two separate key enzymes and two pathway-specific transcription factors are involved in fusaric acid biosynthesis in Fusarium fujikuroi. Environ Microbiol 18:936–956. https://doi.org/10.1111/1462-2920.13150
Studt L, Rösler SM, Burkhardt I et al (2016b) Knock-down of the methyltransferase Kmt6 relieves H3K27me3 and results in induction of cryptic and otherwise silent secondary metabolite gene clusters in Fusarium fujikuroi. Environ Microbiol 18:4037–4054. https://doi.org/10.1111/1462-2920.13427
Studt L, Janevska S, Arndt B et al (2017) Lack of the COMPASS component Ccl1 reduces H3K4 trimethylation levels and affects transcription of secondary metabolite genes in two plant-pathogenic Fusarium species. Front Microbiol 7:1–17. https://doi.org/10.3389/fmicb.2016.02144
Summerell BA, Laurence MH, Liew ECY, Leslie JF (2010) Biogeography and phylogeography of Fusarium: a review. Fungal Divers 44:3–13. https://doi.org/10.1007/s13225-010-0060-2
Takken FLW, van Wijk R, Michielse CB et al (2004) A one-step method to convert vectors into binary vectors suited for Agrobacterium-mediated transformation. Curr Genet 45:242–248. https://doi.org/10.1007/s00294-003-0481-5
Tobiasen C, Aahman J, Ravnholt KS et al (2007) Nonribosomal peptide synthetase (NPS) genes in Fusarium graminearum, F. culmorum and F. pseudograminearum and identification of NPS2 as the producer of ferricrocin. Curr Genet 51:43–58. https://doi.org/10.1007/s00294-006-0103-0
Troncoso C, González X, Bömke C et al (2010) Gibberellin biosynthesis and gibberellin oxidase activities in Fusarium sacchari, Fusarium konzum and Fusarium subglutinans strains. Phytochemistry 71:1322–1331. https://doi.org/10.1016/j.phytochem.2010.05.006
Tudzynski B, Mende K, Weltring K-M et al (1996) The Gibberella fujikuroi niaD gene encoding nitrate reductase: isolation, sequence, homologous transformation and electrophoretic karyotype location. Microbiology 142:533–539. https://doi.org/10.1099/13500872-142-3-533
Twaruschek K, Spörhase P, Michlmayr H et al (2018) New plasmids for Fusarium transformation allowing positive-negative selection and efficient Cre-loxP mediated marker recycling. Front Microbiol 9:1–14. https://doi.org/10.3389/fmicb.2018.01954
Utermark J, Karlovsky P (2008) Genetic transformation of filamentous fungi by Agrobacterium tumefaciens. Protoc Exch. https://doi.org/10.1038/nprot.2008.83
van Helden J, del Olmo M, Pérez-Ortín JE (2000) Statistical analysis of yeast genomic downstream sequences reveals putative polyadenylation signals. Nucleic Acids Res 28:1000–1010
Vanheule A, Audenaert K, Warris S et al (2016) Living apart together: crosstalk between the core and supernumerary genomes in a fungal plant pathogen. BMC Genomics 17:1–18. https://doi.org/10.1186/s12864-016-2941-6
Varga J, Kocsubé S, Tóth B, Mesterházy Á (2005) Nonribosomal peptide synthetase genes in the genome of Fusarium graminearum, causative agent of wheat head blight. Acta Biol Hung 56:375–388. https://doi.org/10.1556/ABiol.56.2005.3-4.19
Von Bargen KW, Niehaus EM, Krug I et al (2015) Isolation and structure elucidation of fujikurins A–D: products of the PKS19 gene cluster in Fusarium fujikuroi. J Nat Prod 78:1809–1815. https://doi.org/10.1021/np5008137
Walkowiak S, Rowland O, Rodrigue N, Subramaniam R (2016) Whole genome sequencing and comparative genomics of closely related Fusarium Head Blight fungi: Fusarium graminearum, F. meridionale and F. asiaticum. BMC Genomics 17:1014. https://doi.org/10.1186/s12864-016-3371-1
Watanabe M, Yonezawa T, Lee KI et al (2011) Molecular phylogeny of the higher and lower taxonomy of the Fusarium genus and differences in the evolutionary histories of multiple genes. BMC Evol Biol. https://doi.org/10.1186/1471-2148-11-322
Westphal KR, Muurmann AT, Paulsen IE et al (2018a) Who needs neighbors? PKS8 is a stand-alone gene in Fusarium graminearum responsible for production of gibepyrones and prolipyrone B. Molecules. https://doi.org/10.3390/molecules23092232
Westphal KR, Wollenberg R, Herbst F-A et al (2018b) Enhancing the production of the fungal pigment aurofusarin in Fusarium graminearum. Toxins (Basel) 10:485. https://doi.org/10.3390/toxins10110485
Westphal KR, Nielsen KAH, Wollenberg RD, Møllehøj MB, Bachleitner S, Studt L, Lysøe E, Giese H, Wimmer R, Sørensen JL, Sondergaard TE (2019) Fusaoctaxin A, an example of a two-step mechanism for non-ribosomal peptide assembly and maturation in fungi. Toxins 11(5):277
Wiebe MG, Nováková M, Miller L et al (1997) Protoplast production and transformation of morphological mutants of the Quorn® myco-protein fungus, Fusarium graminearum A3/5, using the hygromycin B resistance plasmid pAN7–1. Mycol Res 101:871–877. https://doi.org/10.1017/S0953756296003425
Wiemann P, Keller NP (2014) Strategies for mining fungal natural products. J Ind Microbiol Biotechnol 41:301–313. https://doi.org/10.1007/s10295-013-1366-3
Wiemann P, Willmann A, Straeten M et al (2009) Biosynthesis of the red pigment bikaverin in Fusarium fujikuroi: genes, their function and regulation. Mol Microbiol 72:931–946. https://doi.org/10.1111/j.1365-2958.2009.06695.x
Wiemann P, Sieber CMK, von Bargen KW et al (2013) Deciphering the cryptic genome: genome-wide analyses of the rice pathogen Fusarium fujikuroi reveal complex regulation of secondary metabolism and novel metabolites. PLoS Pathog 9:1. https://doi.org/10.1371/journal.ppat.1003475
Windels CE (2000) Economic and social impacts of Fusarium head blight: changing farms and rural communities in the Northern Great Plains. Phytopathology 90:17–21. https://doi.org/10.1094/PHYTO.2000.90.1.17
Wolf T, Shelest V, Nath N, Shelest E (2016) CASSIS and SMIPS: promoter-based prediction of secondary metabolite gene clusters in eukaryotic genomes. Bioinformatics 32:1138–1143. https://doi.org/10.1093/bioinformatics/btv713
Wollenberg RD, Saei W, Westphal KR et al (2017) Chrysogine biosynthesis is mediated by a two-module nonribosomal peptide synthetase. J Nat Prod 80:2131–2135. https://doi.org/10.1021/acs.jnatprod.6b00822
Yang P, Chen Y, Wu H et al (2018) The 5-oxoprolinase is required for conidiation, sexual reproduction, virulence and deoxynivalenol production of Fusarium graminearum. Curr Genet 64:285–301. https://doi.org/10.1007/s00294-017-0747-y
Yoshida M, Nakajima T (2010) Deoxynivalenol and nivalenol accumulation in wheat infected with Fusarium graminearum during grain development. Phytopathology 100:763–773. https://doi.org/10.1094/PHYTO-100-8-0763
Yu J-H, Keller N (2005) Regulation of secondary metabolism in filamentous fungi. Annu Rev Phytopathol 43:437–458. https://doi.org/10.1146/annurev.phyto.43.040204.140214
Zhao C, Waalwijk C, de Wit PJGM et al (2014) Relocation of genes generates non-conserved chromosomal segments in Fusarium graminearum that show distinct and co-regulated gene expression patterns. BMC Genomics 15:191. https://doi.org/10.1186/1471-2164-15-191
Acknowledgements
This study was funded by Novo Nordisk Foundation (Grant number NNF15OC0016028) to MRN.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nielsen, M.R., Sondergaard, T.E., Giese, H. et al. Advances in linking polyketides and non-ribosomal peptides to their biosynthetic gene clusters in Fusarium. Curr Genet 65, 1263–1280 (2019). https://doi.org/10.1007/s00294-019-00998-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-019-00998-4