Abstract
In response to various environmental stimuli and stressors, the budding yeast Saccharomyces cerevisiae can initiate a striking morphological transition from its classic growth mode as isolated single cells to a filamentous form in which elongated cells remain connected post-cytokinesis in multi-cellular pseudohyphae. The formation of pseudohyphal filaments is regulated through an expansive signaling network, encompassing well studied and highly conserved pathways enabling changes in cell polarity, budding, cytoskeletal organization, and cell adhesion; however, changes in metabolite levels underlying the pseudohyphal growth transition are less well understood. We have recently identified a function for second messenger inositol polyphosphates (InsPs) in regulating pseudohyphal growth. InsPs are formed through the cleavage of membrane-bound phosphatidylinositol 4,5-bisphosphate (PIP2), and these soluble compounds are now being appreciated as important regulators of diverse processes, from phosphate homeostasis to cell migration. We find that kinases in the InsP pathway are required for wild-type pseudohyphal growth, and that InsP species exhibit characteristic profiles under conditions promoting filamentation. Ratios of the doubly phosphorylated InsP7 isoforms 5PP-InsP5 to 1PP-InsP5 are elevated in mutants exhibiting exaggerated pseudohyphal growth. Interestingly, S. cerevisiae mutants deleted of the mitogen-activated protein kinases (MAPKs) Kss1p or Fus3p or the AMP-activated kinase (AMPK) family member Snf1p display mutant InsP profiles, suggesting that these signaling pathways may contribute to the regulatory mechanism controlling InsP levels. Consequently, analyses of yeast pseudohyphal growth may be informative in identifying mechanisms regulating InsPs, while indicating a new function for these conserved second messengers in modulating cell stress responses and morphogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
As is true of many fungi, the budding yeast Saccharomyces cerevisiae can exist in more than one morphological type, and the transition between these morphological states is accomplished through a controlled and precise interplay between hundreds of genes enabling dramatic changes in most aspects of cell function (Gimeno et al. 1992; Cullen and Sprague 2012). Under conditions of nitrogen or glucose limitation, certain strains of S. cerevisiae (e.g., Σ1278b) can undergo a transition from the typical budding yeast-like growth form to one in which cells elongate and remain connected following cell division, forming multicellular filaments termed pseudohyphae for their superficial resemblance to multinucleate hyphal tubes (Fig. 1) (Gimeno et al. 1992; Cullen and Sprague 2000). Pseudohyphal filaments can be seen spreading outward from a colony over a solid surface substrate and/or invading the surface below the colony. Pseudohyphal growth is thought to be a scavenging mechanism, enabling non-motile yeast to spread out over a greater surface area in search of regions with more readily available nutrients. Pseudohyphal filamentation has been studied extensively over the years, as the process closely resembles pseudohyphal and hyphal growth transitions that are required for virulence in the related opportunistic human fungal pathogen Candida albicans (Braun and Johnson 1997; Lo et al. 1997; Mitchell 1998). Further, the signaling pathways that enable pseudohyphal growth in S. cerevisiae are conserved, with orthologous signaling systems regulating cell growth and responses to nutrient availability in metazoans (Cook et al. 1996; Erdman et al. 1998; Liu et al. 1993; Madhani and Fink 1997; Mosch et al. 1996; Pan and Heitman 1999).
Morphological changes in S. cerevisiae pseudohyphal growth. Images of yeast cells and colonies grown in media with limited ammonium sulfate as a nitrogen source (low nitrogen media) or in media with normal nutrient availability. Quantification of cell elongation is indicated as the percentage of cells exhibiting a cell length:width ratio of greater than two. Arrowheads indicate elongated cells typical of pseudohyphal growth. Colony images are shown from a culture spread on an agar plate after 3 days growth. Scale bar for cells, 3 µm; scale bars for colony images are each 2 mm
Systematic screens for genes contributing to wild-type pseudohyphal growth have identified over 500 genes either required for pseudohyphal growth or that yield pseudohyphal growth phenotypes upon overexpression (Jin et al. 2008; Ryan et al. 2012; Shively et al. 2013). This collective gene set is broad, and the elucidation of pathways and mechanisms involving these genes warrants significant attention from the biological community, encompassing both ongoing and future studies. Much current and past research into yeast pseudohyphal growth has centered upon key and highly conserved signaling pathways required for wild-type filamentation. Pioneering research into the genetic basis of pseudohyphal growth identified the MAPK cascade of Ste11p, Ste7p, and the MAPK Kss1p (Cook et al. 1997; Liu et al. 1993; Madhani et al. 1997; Roberts and Fink 1994). Among its functions, phosphorylated Kss1p activates pseudohyphal growth through the transcription factors Ste12p and Tec1p, which form a complex that can recognize filamentation-responsive elements (FREs) in the promoters of target genes (Bardwell et al. 1998; Madhani and Fink 1997; Madhani et al. 1997). The mating pathway MAPK Fus3p inhibits pseudohyphal growth by phosphorylating Tec1p in response to pheromone, targeting Tec1p for degradation by the proteasome (Bao et al. 2004). The rat sarcoma (RAS)/protein kinase A pathway regulates pseudohyphal growth through several mechanisms, including phosphorylation of the pseudohyphal growth transcription factor Flo8p by Tpk2p, a catalytic subunit of protein kinase A (Pan and Heitman 1999; Robertson and Fink 1998; Iyer and Bhat 2017). The highly conserved nutrient-sensing target of rapamycin (TOR) pathway regulates pseudohyphal growth through the transcription factor Gcn4p, which in turn regulates expression of the flocculin Flo11p (Boeckstaens et al. 2008; Braus et al. 2003). The AMPK ortholog Snf1p functions in a glucose-sensing and regulatory pathway [reviewed in Simpson-Lavy and Kupiec (2018)], contributing to the control of filamentation by regulating the pseudohyphal growth repressors Nrg1p and Nrg2p at the FLO11 promoter (Kuchin et al. 2002; Lo and Dranginis 1998; Vyas et al. 2003). The FLO11 gene contains an unusually large promoter that is targeted by transcriptional regulators downstream of the MAPK, RAS/protein kinase A, TOR, and Snf1p signaling pathways (Rupp et al. 1999).
The brief overview above highlights a few critical pseudohyphal growth signaling pathways that contribute to broad and striking cellular changes in polarity, budding, cytoskeletal organization, cell cycle progression, and cell–cell adhesion. During pseudohyphal growth, yeast cells exhibit an increase in polarized apical growth occurring at the cell tip. Accordingly, the actin cytoskeleton in cells undergoing pseudohyphal growth is highly polarized, and the polarisome machinery, encompassing the formin Bni1p and the polarity control GTPase Cdc42p, is required for polarized growth during filamentation (Evangelista et al. 1997; Gladfelter et al. 2005). Under nutrient-limiting conditions that induce pseudohyphal growth, both haploid and diploid cells of filamentation-competent strains exhibit distal-unipolar budding, with buds forming predominantly at the distal pole (Gimeno et al. 1992). The distal marker Bud8p is required for unipolar budding during filamentation and is the preferential polar landmark over other positional cues in filamentous cells (Cullen and Sprague 2002; Harkins et al. 2001). Perspectives regarding budding and septin assembly in non-filamentous yeast are presented in Kang and Lew (2017). Cell elongation during pseudohyphal growth is also achieved through a delay in progression through G2/M, extending a period of directed apical growth, relative to uniform isotropic growth spread around the cell cortex (Kron et al. 1994). Filament formation requires enhanced adhesion between cells, and Flo11p is the principal expressed flocculin, with other FLO gene family members located subtelomerically in transcriptionally repressed chromosomal regions (Guo et al. 2000; Lambrechts et al. 1996; Lo and Dranginis 1996). As suggested by its complex transcriptional regulation, alterations in Flo11p levels significantly impact cell adhesion (Fidalgo et al. 2006; Karunanithi et al. 2010). Interestingly, FLO11 expression is regulated epigenetically and can be subject to rapid change, yielding yeast cell populations with heterogeneous adhesion properties (Halme et al. 2004; Verstrepen et al. 2005).
Although much still remains to be understood regarding the changes in cellular properties, signaling pathways, and proteins underlying the pseudohyphal growth transition, even less is known with respect to the changes in metabolites associated with filamentation. Short-chain alcohols, such as 1-butanol, can induce pseudohyphal growth, and these alcohols are now recognized as part of a quorum-sensing mechanism in S. cerevisiae (Chen and Fink 2006; Lorenz et al. 2000). Yeast cells secrete alcohol, such that corresponding alcohol levels roughly gauge cell density and population (Chen and Fink 2006). Tetrahydrofolate (vitamin B9) also induces pseudohyphal growth through uncharacterized mechanisms that impact FLO11 expression levels (Guldener et al. 2004). The phytohormone indole-3-acetic acid is produced in yeasts and is known to induce pseudohyphal filamentation (Rao et al. 2010); its mechanism of action is unclear.
Our studies of pseudohyphal growth signaling pathways inadvertently led us to consider the role of another metabolite, inositol polyphosphate, in the yeast pseudohyphal growth transition. Using quantitative phosphoproteomics to profile changes in protein phosphorylation dependent upon a set of eight kinases required for wild-type pseudohyphal growth (Ste20p, Ste11p, Ste7p, Kss1p, Fus3p, Tpk2p, Snf1p, and Elm1p), we observed differences in the phosphorylation state of several kinases in the InsP biosynthetic pathway (Shively et al. 2015). InsPs are a ubiquitous class of second messengers with an increasingly recognized role in a diverse array of cellular processes.
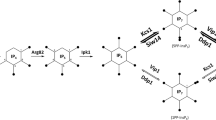
Soluble InsPs are derived from membrane-bound phosphatidylinositol 4,5-bisphosphate (PIP2) through the action of phospholipase C (Flick and Thorner 1993), which cleaves InsP3 from PIP2. The inositol polyphosphate InsP3 is a 6-carbon cyclic alcohol with phosphate groups at the carbon-1, carbon-4, and carbon-5 positions. A variety of InsP species with additional phosphate groups are derived from InsP3 through a sequentially acting set of InsP kinases and phosphatases [reviewed in Monserrate and York (2010)]. Arg82p, the S. cerevisiae ortholog of human IMPK, generates InsP5 from InsP3 through reactions that sequentially add phosphate groups to the carbon-6 and then carbon-3 positions of InsP3 (Hatch and York 2010; Saiardi et al. 1999). The InsP kinase Ipk1p can convert InsP5 to InsP6 (York et al. 1999). Both InsP5 and InsP6 can be pyrophosphorylated, acquiring two phosphate groups at a single carbon position. In S. cerevisiae, the kinases Kcs1p and Vip1p are capable of pyrophosphorylating InsPs (Mulugu et al. 2007). Pyrophosphorylated isoforms of InsP7 and InsP8, as well as the kinases catalyzing the respective reactions, are summarized in Fig. 2. InsP phosphorylation is balanced by dephosphorylation through the phosphatases Siw14p, Ddp1p, and Vip1p, with the latter exhibiting both kinase and phosphatase activity (Pohlmann et al. 2014; Steidle et al. 2016; Wundenberg et al. 2014). The actions of these kinases and phosphatases generate dynamic changes in the cellular abundance and availability of different InsP isoforms, making them strong candidates to act as signal transducers in response to environmental perturbations.
InsPs have indeed come to be recognized as secondary messengers for cellular signal transduction. Perhaps most famously, InsP3 binds to calcium channel receptors, regulating intracellular calcium release; however, this regulatory effect is not observed in S. cerevisiae, being restricted to higher eukaryotes (Michell et al. 1981). InsPs and inositol pyrophosphates have been shown to regulate a broad range of processes, including phosphate sensing, insulin secretion, viral particle release, glycolysis, ribosome synthesis, telomere length, cellular energy dynamics, dynein-driven transport, prion propagation, and amino acid signaling (Azevedo et al. 2009; Chakraborty et al. 2010; Chanduri et al. 2016; Kim et al. 2011; Lee et al. 2007; Saiardi et al. 2005; Szijgyarto et al. 2011; Thota et al. 2015; Wickner et al. 2017, 2018; Wild et al. 2016). Although it has been suggested that InsP3 is the only true second messenger among the InsP species (Shears et al. 2012), there is no doubt that inositol pyrophosphates are important for cellular signaling, particularly since they contain high-energy phosphate bonds (Bennett et al. 2006; Chakraborty et al. 2011). Two mechanisms have been proposed for the actions of pyrophosphates in cell signaling. First, inositol pyrophosphates may bind to proteins allosterically, changing their conformation, localization, and activity (Wu et al. 2016). Secondly, InsPs may regulate signaling through the transfer of a phosphate group to previously phosphorylated serine residues, generating pyrophosphorylated proteins (Bhandari et al. 2007; Saiardi 2016). In the pathogenic fungus Cryptococcus neoformans, InsP7 was found to be crucial for metabolic adaptation to host environment and virulence (Lev et al. 2015). Asp1p, an ortholog of Vip1p in S. pombe, regulates polarized growth and the dimorphic switch (Pohlmann and Fleig 2010). Hence, like membrane constituent phospholipids (Rao et al. 2018), soluble InsPs can regulate physiological responses to various environmental stimuli in yeast.
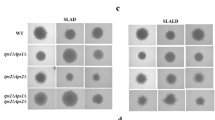
Building on our observation that InsP pathway kinases are differentially phosphorylated in pseudohyphal growth kinase mutants, recent work from our laboratory indicates that InsP signaling regulates pseudohyphal growth (Norman et al. 2018). Genes encoding kinases and phosphatases in the InsP biosynthetic pathway are required for wild-type pseudohyphal growth under conditions of nitrogen limitation. Under these conditions, two isoforms of InsP7, 5PP-InsP5 and 1PP-InsP5, can be distinguished, and elevated ratios of 5PP-InsP5 to 1PP-InsP5 are diagnostic of mutant strains exhibiting exaggerated pseudohyphal filamentation. Overexpression of KCS1, promoting elevated levels of the 5PP-InsP5 isoform of InsP7, is sufficient to drive pseudohyphal filamentation under otherwise non-inducing conditions.
The studies described above indicate a role for inositol polyphosphate signaling in pseudohyphal growth, but the findings also raise an open question as to the signaling pathways and networks that regulate inositol polyphosphate levels. Currently, relatively little is known regarding the regulation of inositol polyphosphate signaling in yeast or other eukaryotes. We find that InsP profiles are perturbed under conditions of nitrogen limitation in S. cerevisiae mutants of the filamentous Σ1278b background singly deleted of the pseudohyphal growth regulatory genes KSS1, FUS3, or SNF1 (Norman et al. 2018). While we lack a regulatory mechanism by which these encoded kinases may modulate InsP kinase phosphorylation, the potential exists for control of the InsP biosynthetic pathway in yeast by the corresponding Kss1p and Fus3p MAPK cascades, as well as by the glucose-responsive AMPK Snf1p pathway. Prior to this work, Arg82p, Ksp1p, and Vip1p were independently identified in proteomic studies as phosphoproteins (Swaney et al. 2013), and VIP1 may be subject to transcriptional regulation under conditions of heat stress by global regulators, such as Xbp1p, as assessed through large-scale chromatin immunoprecipitation analysis (Venters et al. 2011). It is certainly feasible, if not likely, that the InsP pathway is subject to significant regulatory control, both at the level of transcription and translation.
In sum, InsP signaling continues to emerge as a prominent second messenger system. The broad scope of processes impacted by InsP signaling likely reflects the diverse set of proteins bound by various InsP and pyrophosphate species, with the relative levels of particular species affecting substrate protein activity and cellular processes. Inositol pyrophosphates, in particular, may fill important regulatory roles, as their levels have been observed to increase in response to some conditions of stress and nutrient limitation (Dubois et al. 2002; Gibney et al. 2013; Worley et al. 2013). Additional work will be needed to identify and dissect the signaling pathways that act upstream of the InsP biosynthetic pathway. Given the diversity of cell processes affected by InsP signaling, it seems most likely that multiple signaling pathways act in parallel or in a convergent pattern to precisely modulate the respective activities of kinases and phosphatases in the InsP pathway. In that respect, the AMPK and MAPK nutrient/stress-responsive pathways may be part of a larger network affecting InsP levels in response to given cellular states and environmental conditions. On a broader level, analyses of InsP signaling highlight the need to consider the functions and changes associated with other cell metabolites, as ongoing and future metabolite profiling studies across single-cell eukaryotic and metazoan systems hold tremendous promise for the discovery of new and important biology.
References
Azevedo C, Burton A, Ruiz-Mateos E, Marsh M, Saiardi A (2009) Inositol pyrophosphate mediated pyrophosphorylation of AP3B1 regulates HIV-1 Gag release. Proc Natl Acad Sci USA 106:21161–21166. https://doi.org/10.1073/pnas.0909176106
Bao MZ, Schwartz MA, Cantin GT, Yates JR, Madhani H (2004) Pheromone-dependent destruction of the Tec1 transcription factor is required for MAP kinase signaling specificity in yeast. Cell 119:991–1000
Bardwell L, Cook JG, Voora D, Baggott DM, Martinez AR, Thorner J (1998) Repression of yeast Ste12 transcription factor by direct binding of unphosphorylated Kss1 MAPK and its regulation by the Ste7 MEK. Genes Dev 12:2887–2898
Bennett M, Onnebo SM, Azevedo C, Saiardi A (2006) Inositol pyrophosphates: metabolism and signaling. Cell Mol Life Sci 63:552–564. https://doi.org/10.1007/s00018-005-5446-z
Bhandari R, Saiardi A, Ahmadibeni Y, Snowman AM, Resnick AC, Kristiansen TZ, Molina H, Pandey A, Werner JK Jr, Juluri KR, Xu Y, Prestwich GD, Parang K, Snyder SH (2007) Protein pyrophosphorylation by inositol pyrophosphates is a posttranslational event. Proc Natl Acad Sci USA 104:15305–15310. https://doi.org/10.1073/pnas.0707338104
Boeckstaens M, Andre B, Marini AM (2008) Distinct transport mechanisms in yeast ammonium transport/sensor proteins of the Mep/Amt/Rh family and impact on filamentation. J Biol Chem 283:21362–21370. https://doi.org/10.1074/jbc.M801467200
Braun BR, Johnson AD (1997) Control of filament formation in Candida albicans by the transcriptional repressor TUP1. Science 277:105–109
Braus GH, Grundmann O, Bruckner S, Mosch HU (2003) Amino acid starvation and Gcn4p regulate adhesive growth and FLO11 gene expression in Saccharomyces cerevisiae. Mol Biol Cell 14:4272–4284. https://doi.org/10.1091/mbc.e03-01-0042
Chakraborty A, Koldobskiy MA, Bello NT, Maxwell M, Potter JJ, Juluri KR, Maag D, Kim S, Huang AS, Dailey MJ, Saleh M, Snowman AM, Moran TH, Mezey E, Snyder SH (2010) Inositol pyrophosphates inhibit Akt signaling, thereby regulating insulin sensitivity and weight gain. Cell 143:897–910. https://doi.org/10.1016/j.cell.2010.11.032
Chakraborty A, Kim S, Snyder SH (2011) Inositol pyrophosphates as mammalian cell signals. Sci Signal 4:re1. https://doi.org/10.1126/scisignal.2001958
Chanduri M, Rai A, Malla AB, Wu M, Fiedler D, Mallik R, Bhandari R (2016) Inositol hexakisphosphate kinase 1 (IP6K1) activity is required for cytoplasmic dynein-driven transport. Biochem J 473:3031–3047. https://doi.org/10.1042/BCJ20160610
Chen H, Fink GR (2006) Feedback control of morphogenesis in fungi by aromatic alcohols. Genes Dev 20:1150–1161. https://doi.org/10.1101/gad.1411806
Cook JG, Bardwell L, Kron SJ, Thorner J (1996) Two novel targets of the MAP kinase Kss1 are negative regulators of invasive growth in the yeast Saccharomyces cerevisiae. Genes Dev 10:2831–2848
Cook JG, Bardwell L, Thorner J (1997) Inhibitory and activating functions forMAPK Kss1 in the S. cerevisiae filamentous growth signalling pathway. Nature 390:85–88
Cullen PJ, Sprague GF (2000) Glucose depletion causes haploid invasive growth in yeast. Proc Natl Acad Sci USA 97:13461–13463
Cullen PJ, Sprague GF Jr (2002) The roles of bud-site-selection proteins during haploid invasive growth in yeast. Mol Biol Cell 13:2990–3004. https://doi.org/10.1091/mbc.E02-03-0151
Cullen PJ, Sprague GF Jr (2012) The regulation of filamentous growth in yeast. Genetics 190:23–49. https://doi.org/10.1534/genetics.111.127456
Dubois E, Scherens B, Vierendeels F, Ho MM, Messenguy F, Shears SB (2002) In Saccharomyces cerevisiae, the inositol polyphosphate kinase activity of Kcs1p is required for resistance to salt stress, cell wall integrity, and vacuolar morphogenesis. J Biol Chem 277:23755–23763. https://doi.org/10.1074/jbc.M202206200
Erdman S, Lin L, Malczynski M, Snyder M (1998) Pheromone-regulated genes required for yeast mating differentiation. J Cell Biol 140:461–483
Evangelista M, Blundell K, Longtine MS, Chow CJ, Adames N, Pringle JR, Peter M, Boone C (1997) Bni1p, a yeast formin linking cdc42p and the actin cytoskeleton during polarized morphogenesis. Science 276:118–122
Fidalgo M, Barrales RR, Ibeas JI, Jimenez J (2006) Adaptive evolution by mutations in the FLO11 gene. Proc Natl Acad Sci USA 103:11228–11233. https://doi.org/10.1073/pnas.0601713103
Flick JS, Thorner J (1993) Genetic and biochemical characterization of a phosphatidylinositol-specific phospholipase C in Saccharomyces cerevisiae. Mol Cell Biol 13:5861–5876
Gibney PA, Lu C, Caudy AA, Hess DC, Botstein D (2013) Yeast metabolic and signaling genes are required for heat-shock survival and have little overlap with the heat-induced genes. Proc Natl Acad Sci USA 110:E4393–E4402. https://doi.org/10.1073/pnas.1318100110
Gimeno CJ, Ljungdahl PO, Styles CA, Fink GR (1992) Unipolar cell divisions in the yeast S. cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell 68:1077–1090
Gladfelter AS, Kozubowski L, Zyla TR, Lew DJ (2005) Interplay between septin organization, cell cycle and cell shape in yeast. J Cell Sci 118:1617–1628. https://doi.org/10.1242/jcs.02286
Guldener U, Koehler GJ, Haussmann C, Bacher A, Kricke J, Becher D, Hegemann JH (2004) Characterization of the Saccharomyces cerevisiae Fol1 protein: starvation for C1 carrier induces pseudohyphal growth. Mol Biol Cell 15:3811–3828. https://doi.org/10.1091/mbc.e03-09-0680
Guo B, Styles CA, Feng Q, Fink G (2000) A Saccharomyces gene family involved in invasive growth, cell-cell adhesion, and mating. Proc Natl Acad Sci USA 97:12158–12163
Halme A, Bumgarner S, Styles CA, Fink GR (2004) Genetic and epigenetic regulation of the FLO gene family generates cell-surface variation in yeast. Cell 116:405–415
Harkins HA, Page N, Schenkman LR, De Virgilio C, Shaw S, Bussey H, Pringle JR (2001) Bud8p and Bud9p, proteins that may mark the sites for bipolar budding in yeast. Mol Biol Cell 12:2497–2518. https://doi.org/10.1091/mbc.12.8.2497
Hatch AJ, York JD (2010) SnapShot: inositol phosphates. Cell 143:1030–1030.e1. https://doi.org/10.1016/j.cell.2010.11.045
Iyer RS, Bhat PJ (2017) KRH1 and KRH2 are functionally non-redundant in signaling for pseudohyphal differentiation in Saccharomyces cerevisiae. Curr Genet 63:851–859. https://doi.org/10.1007/s00294-017-0684-9
Jin R, Dobry CJ, McCown PJ, Kumar A (2008) Large-scale analysis of yeast filamentous growth by systematic gene disruption and overexpression. Mol Biol Cell 19:284–296
Kang H, Lew DJ (2017) How do cells know what shape they are? Curr Genet 63:75–77. https://doi.org/10.1007/s00294-016-0623-1
Karunanithi S, Vadaie N, Chavel CA, Birkaya B, Joshi J, Grell L, Cullen PJ (2010) Shedding of the mucin-like flocculin Flo11p reveals a new aspect of fungal adhesion regulation. Curr Biol 20:1389–1395
Kim S, Kim SF, Maag D, Maxwell MJ, Resnick AC, Juluri KR, Chakraborty A, Koldobskiy MA, Cha SH, Barrow R, Snowman AM, Snyder SH (2011) Amino acid signaling to mTOR mediated by inositol polyphosphate multikinase. Cell Metab 13:215–221. https://doi.org/10.1016/j.cmet.2011.01.007
Kron SJ, Styles CA, Fink GR (1994) Symmetric cell division in pseudohyphae of the yeast Saccharomyces cerevisiae. Mol Biol Cell 5:1003–1022
Kuchin S, Vyas VK, Carlson M (2002) Snf1 protein kinase and the repressors Nrg1 and Nrg2 regulate FLO11, haploid invasive growth, and diploid pseudohyphal differentiation. Mol Cell Biol 22:3994–4000
Lambrechts MG, Bauer FF, Marmur J, Pretorius IS (1996) Muc1, a mucin-like protein that is regulated by Mss10, is critical for pseudohyphal differentiation in yeast. Proc Natl Acad Sci USA 93:8419–8424
Lee YS, Mulugu S, York JD, O’Shea EK (2007) Regulation of a cyclin-CDK-CDK inhibitor complex by inositol pyrophosphates. Science 316:109–112. https://doi.org/10.1126/science.1139080
Lev S, Li C, Desmarini D, Saiardi A, Fewings NL, Schibeci SD, Sharma R, Sorrell TC, Djordjevic JT (2015) Fungal inositol pyrophosphate IP7 is crucial for metabolic adaptation to the host environment and pathogenicity. MBio 6:e00531–e00515. https://doi.org/10.1128/mBio.00531-15
Liu H, Styles CA, Fink GR (1993) Elements of the yeast pheromone response pathway required for filamentous growth of diploids. Science 262:1741–1744
Lo WS, Dranginis AM (1996) FLO11, a yeast gene related to the STA genes, encodes a novel cell surface flocculin. J Bacteriol 178:7144–7151
Lo WS, Dranginis AM (1998) The cell surface flocculin Flo11 is required for pseudohyphae formation and invasion by Saccharomyces cerevisiae. Mol Biol Cell 9:161–171
Lo HJ, Kohler J, DiDomenico B, Loebenberg D, Cacciapuoti A, Fink GR (1997) Nonfilamentous C. albicans mutants are avirulent. Cell 90:939–949
Lorenz MC, Cutler NS, Heitman J (2000) Characterization of alcohol-induced filamentous growth in Saccharomyces cerevisiae. Mol Biol Cell 11:183–199
Madhani HD, Fink GR (1997) Combinatorial control required for the specificity of yeast MAPK signaling. Science 275:1314–1317
Madhani HD, Styles CA, Fink GR (1997) MAP kinases with distinct inhibitory functions impart signaling specificity during yeast differentiation. Cell 91:673–684
Michell RH, Kirk CJ, Jones LM, Downes CP, Creba JA (1981) The stimulation of inositol lipid metabolism that accompanies calcium mobilization in stimulated cells: defined characteristics and unanswered questions. Philos Trans R Soc Lond B Biol Sci 296:123–138
Mitchell AP (1998) Dimorphism and virulence in Candida albicans. Curr Opin Microbiol 1:687–692
Monserrate JP, York JD (2010) Inositol phosphate synthesis and the nuclear processes they affect. Curr Opin Cell Biol 22:365–373. https://doi.org/10.1016/j.ceb.2010.03.006
Mosch HU, Roberts RL, Fink GR (1996) Ras2 signals via the Cdc42/Ste20/mitogen-activated protein kinase module to induce filamentous growth in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 93:5352–5356
Mulugu S, Bai W, Fridy PC, Bastidas RJ, Otto JC, Dollins DE, Haystead TA, Ribeiro AA, York JD (2007) A conserved family of enzymes that phosphorylate inositol hexakisphosphate. Science 316:106–109. https://doi.org/10.1126/science.1139099
Norman KL, Shively CA, De La Rocha AJ, Mutlu N, Basu S, Cullen PJ, Kumar A (2018) Inositol polyphosphates regulate and predict yeast pseudohyphal growth phenotypes. PLoS Genet 14:e1007493. https://doi.org/10.1371/journal.pgen.1007493
Pan X, Heitman J (1999) Cyclic AMP-dependent protein kinase regulates pseudohyphal differentiation in Saccharomyces cerevisiae. Mol Cell Biol 19:4874–4887
Pohlmann J, Fleig U (2010) Asp1, a conserved 1/3 inositol polyphosphate kinase, regulates the dimorphic switch in Schizosaccharomyces pombe. Mol Cell Biol 30:4535–4547. https://doi.org/10.1128/MCB.00472-10
Pohlmann J, Risse C, Seidel C, Pohlmann T, Jakopec V, Walla E, Ramrath P, Takeshita N, Baumann S, Feldbrugge M, Fischer R, Fleig U (2014) The Vip1 inositol polyphosphate kinase family regulates polarized growth and modulates the microtubule cytoskeleton in fungi. PLoS Genet 10:e1004586. https://doi.org/10.1371/journal.pgen.1004586
Rao RP, Hunter A, Kashpur O, Normanly J (2010) Aberrant synthesis of indole-3-acetic acid in Saccharomyces cerevisiae triggers morphogenic transition, a virulence trait of pathogenic fungi. Genetics 185:211–220. https://doi.org/10.1534/genetics.109.112854
Rao MJ, Srinivasan M, Rajasekharan R (2018) Cell size is regulated by phospholipids and not by storage lipids in Saccharomyces cerevisiae. Curr Genet. https://doi.org/10.1007/s00294-018-0821-0
Roberts RL, Fink GR (1994) Elements of a single MAP kinase cascade in Saccharomyces cerevisiae mediate two developmental programs in the same cell type: mating and invasive growth. Genes Dev 8:2974–2985
Robertson LS, Fink GR (1998) The three yeast A kinases have specific signaling functions in pseudohyphal growth. Proc Natl Acad Sci USA 95:13783–13787
Rupp S, Summers E, Lo HJ, Madhani H, Fink G (1999) MAP kinase and cAMP filamentation signaling pathways converge on the unusually large promoter of the yeast FLO11 gene. EMBO J 18:1257–1269
Ryan O, Shapiro RS, Kurat CF, Mayhew D, Baryshnikova A, Chin B, Lin ZY, Cox MJ, Vizeacoumar F, Cheung D, Bahr S, Tsui K, Tebbji F, Sellam A, Istel F, Schwarzmuller T, Reynolds TB, Kuchler K, Gifford DK, Whiteway M, Giaever G, Nislow C, Costanzo M, Gingras AC, Mitra RD, Andrews B, Fink GR, Cowen LE, Boone C (2012) Global gene deletion analysis exploring yeast filamentous growth. Science 337:1353–1356. https://doi.org/10.1126/science.1224339
Saiardi A (2016) Protein pyrophosphorylation: moving forward. Biochem J 473:3765–3768. https://doi.org/10.1042/BCJ20160710C
Saiardi A, Erdjument-Bromage H, Snowman AM, Tempst P, Snyder SH (1999) Synthesis of diphosphoinositol pentakisphosphate by a newly identified family of higher inositol polyphosphate kinases. Curr Biol 9:1323–1326
Saiardi A, Resnick AC, Snowman AM, Wendland B, Snyder SH (2005) Inositol pyrophosphates regulate cell death and telomere length through phosphoinositide 3-kinase-related protein kinases. Proc Natl Acad Sci USA 102:1911–1914. https://doi.org/10.1073/pnas.0409322102
Shears SB, Ganapathi SB, Gokhale NA, Schenk TM, Wang H, Weaver JD, Zaremba A, Zhou Y (2012) Defining signal transduction by inositol phosphates. Subcell Biochem 59:389–412. https://doi.org/10.1007/978-94-007-3015-1_13
Shively CA, Eckwahl MJ, Dobry CJ, Mellacheruvu D, Nesvizhskii A, Kumar A (2013) Genetic networks inducing invasive growth in Saccharomyces cerevisiae identified through systematic genome-wide overexpression. Genetics 193:1297–1310. https://doi.org/10.1534/genetics.112.147876
Shively CA, Kweon HK, Norman KL, Mellacheruvu D, Xu T, Sheidy DT, Dobry CJ, Sabath I, Cosky EE, Tran EJ, Nesvizhskii A, Andrews PC, Kumar A (2015) Large-scale analysis of kinase signaling in yeast pseudohyphal development identifies regulation of ribonucleoprotein granules. PLoS Genet 11:e1005564. https://doi.org/10.1371/journal.pgen.1005564
Simpson-Lavy K, Kupiec M (2018) A reversible liquid drop aggregation controls glucose response in yeast. Curr Genet 64:785–788. https://doi.org/10.1007/s00294-018-0805-0
Steidle EA, Chong LS, Wu M, Crooke E, Fiedler D, Resnick AC, Rolfes RJ (2016) A novel inositol pyrophosphate phosphatase in Saccharomyces cerevisiae: Siw14 protein selectively cleaves the beta-phosphate from 5-diphosphoinositol pentakisphosphate (5PP-Ip5). J Biol Chem 291:6772–6783. https://doi.org/10.1074/jbc.M116.714907
Swaney DL, Beltrao P, Starita L, Guo A, Rush J, Fields S, Krogan NJ, Villen J (2013) Global analysis of phosphorylation and ubiquitylation cross-talk in protein degradation. Nat Methods 10:676–682. https://doi.org/10.1038/nmeth.2519
Szijgyarto Z, Garedew A, Azevedo C, Saiardi A (2011) Influence of inositol pyrophosphates on cellular energy dynamics. Science 334:802–805. https://doi.org/10.1126/science.1211908
Thota SG, Unnikannan CP, Thampatty SR, Manorama R, Bhandari R (2015) Inositol pyrophosphates regulate RNA polymerase I-mediated rRNA transcription in Saccharomyces cerevisiae. Biochem J 466:105–114. https://doi.org/10.1042/BJ20140798
Venters BJ, Wachi S, Mavrich TN, Andersen BE, Jena P, Sinnamon AJ, Jain P, Rolleri NS, Jiang C, Hemeryck-Walsh C, Pugh BF (2011) A comprehensive genomic binding map of gene and chromatin regulatory proteins in Saccharomyces. Mol Cell 41:480–492. https://doi.org/10.1016/j.molcel.2011.01.015
Verstrepen KJ, Jansen A, Lewitter F, Fink GR (2005) Intragenic tandem repeats generate functional variability. Nat Genet 37:986–990. https://doi.org/10.1038/ng1618
Vyas VK, Kuchin S, Berkey CD, Carlson M (2003) Snf1 kinases with different beta-subunit isoforms play distinct roles in regulating haploid invasive growth. Mol Cell Biol 23:1341–1348
Wickner RB, Kelly AC, Bezsonov EE, Edskes HK (2017) [PSI+] prion propagation is controlled by inositol polyphosphates. Proc Natl Acad Sci USA 114:E8402–E8410. https://doi.org/10.1073/pnas.1714361114
Wickner RB, Edskes HK, Bezsonov EE, Son M, Ducatez M (2018) Prion propagation and inositol polyphosphates. Curr Genet 64:571–574. https://doi.org/10.1007/s00294-017-0788-2
Wild R, Gerasimaite R, Jung JY, Truffault V, Pavlovic I, Schmidt A, Saiardi A, Jessen HJ, Poirier Y, Hothorn M, Mayer A (2016) Control of eukaryotic phosphate homeostasis by inositol polyphosphate sensor domains. Science 352:986–990. https://doi.org/10.1126/science.aad9858
Worley J, Luo X, Capaldi AP (2013) Inositol pyrophosphates regulate cell growth and the environmental stress response by activating the HDAC Rpd3L. Cell Rep 3:1476–1482. https://doi.org/10.1016/j.celrep.2013.03.043
Wu M, Chong LS, Perlman DH, Resnick AC, Fiedler D (2016) Inositol polyphosphates intersect with signaling and metabolic networks via two distinct mechanisms. Proc Natl Acad Sci USA 113:E6757–E6765. https://doi.org/10.1073/pnas.1606853113
Wundenberg T, Grabinski N, Lin H, Mayr GW (2014) Discovery of InsP6-kinases as InsP6-dephosphorylating enzymes provides a new mechanism of cytosolic InsP6 degradation driven by the cellular ATP/ADP ratio. Biochem J 462:173–184. https://doi.org/10.1042/BJ20130992
York JD, Odom AR, Murphy R, Ives EB, Wente SR (1999) A phospholipase C-dependent inositol polyphosphate kinase pathway required for efficient messenger RNA export. Science 285:96–100
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Mutlu, N., Kumar, A. Messengers for morphogenesis: inositol polyphosphate signaling and yeast pseudohyphal growth. Curr Genet 65, 119–125 (2019). https://doi.org/10.1007/s00294-018-0874-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-018-0874-0