Abstract
Turbinaria ornata (Turner) J. Agardh is a perennial brown alga native to coral reef ecosystems of tropical areas of the Pacific and Indian Ocean. Very little is known about its organellar genome structure. In the present work, the complete mitochondrial genome sequence of T. ornata was determined and compared with other reported brown algal mtDNAs. The circular mitogenome of 34,981 bp contains a basic set of 65 mitochondrial genes. The structure and organization of T. ornata mitogenome is very similar to Sargassum species. Turbinaria ornata genes overlap by a total of 164 bp in 12 different locations from 1 to 66 bp, and the non-coding sequences are 1872 bp, constituting approximate 5.35 % of the genome. The total spacer size has positive correlation with the brown algal mitogenome size with the correlation coefficient of 0.7972. Several regions displaying greater inconsistency (rnl-trnK spacer, cox2 gene, cox3-atp6 spacer, rps14-rns middle region and trnP-rnl spacer) have been identified in brown algal mtDNAs. The observed uncertainty regarding the position and support values of some branches might be closely associated with the heterogeneity of evolutionary rate.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brown algae (Phaeophyceae) have successfully occupied almost all marine coasts in cold, temperate and tropical zones, and play important ecological roles in the marine ecosystem. They have developed diverse morphology from the giant kelp of the order Laminariales (Macrocystis and Nereocystis) to the microscopic, filamentous representatives of the Ectocarpales (Laminariocolax and Microsyphar) (Charrier et al. 2012). So far, brown algae comprise a multitude of taxa including 19 orders, 62 families, 473 genera and nearly 2000 species (Guiry and Guiry 2014). Phylogenetic analyses are required to develop a complete inventory of the Phaeophyceae to enable us to understand evolutionary biodiversity over time (Silberfeld et al. 2014). Many attractive features of the mitochondrial genome (e.g., its abundance in the cell, accelerated substitution rates, and genome-level information content) have made it an invaluable tool in phylogenetics, phylogeography, and population genetics at various taxonomic levels (Liu et al. 2015; Liu and Pang 2015).
Complete mtDNA sequences from under-investigated categories would supplement and improve our understanding of mitochondrial genome evolution in the Phaeophyceae. Its remarkable stability and universality of mitogenome content in major groups of stramenopiles have allowed analyses of mitochondrial genome sequences and gene order to reconstruct deep node phylogenies and to reassess relationships among closely related species (Ravin et al. 2010; Liu et al. 2015). However, there is still scarce information on mitogenomes of brown algae, and only 25 complete brown algal mtDNAs have been reported (Oudot-Le Secq et al. 2001; Yotsukura et al. 2010; Zhang et al. 2013; Liu and Pang 2015). The known brown algal mitochondrial genomes are circular, double-stranded DNA molecules, ranging in length from 31.6 to 58.5 kb (Table 1). Their organization and gene content are highly conserved. Brown algal mtDNAs usually harbor three ribosomal RNA genes, 24–26 transfer RNA genes, 35–36 protein-encoding genes, several open reading frames (ORFs), and some non-coding regions containing signals for transcription and replication.
Turbinaria J.V. Lamouroux is a relatively species-poor genus of the order Fucales (Wynne 2002; Le Lann et al. 2008), and to date, there are 22 species accepted taxonomically in Turbinaria (Guiry and Guiry 2014). Turbinaria species display great morphological plasticity in response to different environmental conditions, thus molecular data are used to delineate species boundaries (Prathep et al. 2007; Rohfritsch et al. 2007, 2010; Draisma et al. 2010). Turbinaria ornata (Turner) J. Agardh is a perennial brown alga native to coral reef ecosystems of tropical areas of the Pacific and Indian Ocean. Since the 1980s, this alga has markedly expanded its abundance and geographical distribution in French Polynesia, due to dispersal of detached, reproductively mature floating fronds (Stewart 2004, 2006; Stiger and Payri 2005). The aggregations of this alga could provide the spatial refuge from herbivory to other algal species on coral reefs and increasing local algal species richness (Bittick et al. 2010). Despite its ecological importance, very little is known about its genome sequence, which has restricted our understanding of its genetic diversity, interspecific taxonomic positions and evolutionary relationships.
In this study, the complete mitochondrial genome sequence of T. ornata has been determined, and comparatively analyzed with published brown algal mtDNAs from Fucales, Desmarestiales, Ectocarpales, Laminariales and Dictyotales, to gain more general insights into the evolution of brown algal mitogenomes. To our knowledge, this is the first report of a completely annotated mitogenome sequence for the genus Turbinaria.
Materials and methods
Sample collection and DNA extraction
Adult plants of Turbinaria ornata (Turner) J. Agardh were collected on the intertidal rocks from Dadonghai Bay, Sanya, Hainan Province, China (37°09′N, 122°32′E) on 1st April, 2014. Plants were transported to the laboratory in coolers (5–8 °C) within 48 h after collection. Fresh tissue was ground to a fine powder in liquid nitrogen for DNA extraction. Total genomic DNA was extracted using a Plant Genomic DNA Kit according to the manufacturer’s instructions (Tiangen Biotech, Beijing, China). The concentration and quality of isolated DNA were evaluated via electrophoresis on a 1.0 % agarose gel.
PCR amplification and sequencing
Based on the known mitogenome sequence data in the Fucales, seven pairs of PCR primers were designed to amplify the entire T. ornata mitogenome with lengths of 7.3, 7.6, 4.0, 1.9, 4.9, 5.9 and 6.8 kb. The primer pairs were as follows: T-23S-F (5′-CTTGGAAGCAGCCATTCTT-3′, this study)/T-trnQ-R (5′-ACCAAAAACGGGCGCCTTAC-3′, this study), T-trnW-F (5′-AACTTTGGTTTCCAAAGCCAACG-3′, this study)/T-atp6-R (5′-ATAAGTCAAATAAACCAATACCT-3′, this study), CAF4A (Kogame et al. 2005)/cox1-R (5′-AACCAAAACAAATGCTGATAAA-3′, this study), T-cox1-F (5′-GGCATCACTATGCTTTTAACGG-3′, this study)/T-cob-R (5′-CAATAATATGAGAATCTACCCAAC-3′, this study), T-nad9-F (5′-CATTTGAGGTGTTTGGACATGAG-3′, this study)/T-nad4-R (5′-CCAAAAACACCAATAATTAGAAGC-3′, this study), cox2-F/trnS-R (Liu et al. 2015), and T-trnS-F (5′-GGAGATGTGGCTGAGTGGCT-3′, this study)/T-23S-R (5′-CTCTTCTCGATGTAGCTACCCG-3′, this study).
PCR was performed in 50 μL reactions containing 32 μL of sterile distilled H2O, 10 μL of 5× PrimeSTAR GXL buffer (5 mM Mg2+ plus, Takara, Japan), 4 μL of dNTP mixture (2.5 mM each), 1 μL of each primer (10 μM), 1 μL of PrimeSTAR GXL DNA polymerase (1.25 units μL−1, Takara, Japan), and 1 μL of DNA template (approximate 50 ng). PCR amplification was carried out on a T-Gradient Thermoblock Thermal Cycler (Whatman Biometra, Goettingen, Germany) with an initial denaturation at 94 °C for 3 min, followed by 30 cycles of denaturation at 94 °C for 20 s, annealing at 50–52 °C for 50 s, extension at 68 °C for 1 min/kb, and a final extension at 68 °C for 10 min. PCR products were purified using the Qiaquick Gel Extraction Kit (Qiagen, Germany). Sequencing reactions were performed using ABI 3730 XL automated sequencers (Applied Biosystems, USA).
Genome assembly and annotation
The DNA sequences were manually edited and assembled using the BioEdit program (http://www.mbio.ncsu.edu/BioEdit/bioedit.html). DNA sequences of the complete mitochondrial genome of T. ornata were determined by comparison with published mitochondrial genome sequences of the Fucales (Oudot-Le Secq et al. 2006; Liu et al. 2014a, c, 2015; Liu and Pang 2015). The locations of the protein-coding genes were determined with DOGMA (Wyman et al. 2004), ORF Finder, and Blastx searches (GenBank). Transfer RNA genes were identified by reconstructing their cloverleaf structures using the tRNAscan-SE 1.21 software with default parameters (Schattner et al. 2005). Ribosomal RNA genes were identified by BLAST searches (Altschul et al. 1997) of the nonredundant databases at the National Center for Biotechnology Information. The map of the mitochondrial genome was built using Organellar Genome DRAW (Lohse et al. 2013). The complete mitogenome sequence of T. ornata is deposited in GenBank with the accession number KM501562.
Phylogenetic analyses
Phylogenetic analyses were carried out using 20 reported brown algal mitogenome sequences (Table 1). Sequences of individual genes were separately aligned and manually adjusted using MEGA 5.2 software (Tamura et al. 2011). The sequences of 21 brown algal mitochondrial genomes were compared using Blast Ring Image Generator (BRIG) software (Alikhan et al. 2011). Phylogenetic relationships within Phaeophyceae were analyzed based on four datasets: (1) three ribosomal RNA genes (rnl, rns, and rrn5), (2) 25 conserved transfer RNA genes, (3) 35 functionally known protein-coding genes and (4) their concatenated protein sequences (rps2-4, 7, 8, 10-14, 19; rpl2, 5, 6, 14, 16, 31; nad1-7, 9, 11; cob; cox1-3; atp6, 8, 9; and tatC). Each dataset was subjected to concatenated alignments using ClustalX 1.83 with the default settings (Thompson et al. 1997). Base composition and pairwise comparison were examined by MEGA 5.2. The identity percentages of 21 brown algal mtDNA sequences including 3 rRNA, 25 tRNA and 35 protein-coding genes were evaluated using the BioEdit v7.0.8.0 software (Hall 1999). Neighbor-joining (NJ) and maximum likelihood (ML) trees were conducted with 1000 bootstrap replicates using MEGA5.2. Dictyota dichotoma served as the outgroup according to Oudot-Le Secq et al. (2006). The evolutionary distances were computed based on the Kimura two-parameter model (Kimura 1980) for DNA sequences and on the Whelan And Goldman + Freq. model (Whelan and Goldman 2001) for protein sequences. Final results are presented using gaps as fifth character states for bases in DNA sequences.
Results and discussion
Genome features
The circular T. ornata mitogenome is 34,981 bp in length within the range of known Fucales mitogenomes from 34,609 bp in Sargassum horneri to 36,392 bp in Fucus vesiculosus. The reported Fucales mtDNAs are shorter than that of Desmarestiales, Ectocarpales (not including Colpomenia peregrina), and Laminariales (Table 1). The overall AT content of the T. ornata mitogenome is 64.18 % and falls between Sargassum (62.47–63.84 %) and F. vesiculosus (65.55 %). The T. ornata mitogenome harbors 65 densely packed genes which are the same genes found in species of Sargassum (Liu et al. 2015; Liu and Pang 2015), including 35 protein-coding genes, three rRNAs (rnl, rns and rrns), 25 tRNAs, and two conserved open reading frames (ORFs) (Fig. 1). No introns were found in all the genes. Two genes, trnY situated between trnA and rps10 and orf379 in F. vesiculosus were absent in T. ornata and Sargassum (Oudot-Le Secq et al. 2006), indicating that the deletion event might have occurred in the common ancestor of Turbinaria and Sargassum.
The structure and organization of the T. ornata mitogenome is identical to Sargassum species. Genes are encoded on both strands of the mtDNA, suggesting the presence of at least two main transcriptional units. There is no inverted repeat in the T. ornata mitogenome. Only one tandem repeat with the period size of 22 bp was observed in the nad4 gene, which was also found in the mitogenomes of S. muticum and S. fusiforme, but not in other Fucales.
Genes in the T. ornata mitogenome
Coding sequences within the T. ornata mtDNA were initially identified by a comparison to mitochondrial gene sequences of reported Sargassum species. The 35 protein-coding genes including 17 ribosomal proteins (rps2-4, 7, 8, 10-14, and 19; rpl2, 5, 6,14, 16, and 31), 10 NADH dehydrogenase complex subunits (nad1-7, 4L, 9 and 11), three cytochrome oxidase subunits (cox1-3), three ATPase subunits (atp6, 8 and 9), apocytochrome b (cob), and secY-independent transporter protein (tatC), exist in all sequenced brown algal mitogenomes.
All protein-coding genes start with a typical initiation codon ATG in the T. ornata mitogenome, while unconventional start codons GTG, CTG and TTG are rarely used in other brown algal mtDNAs (Liu et al. 2015). In brown algal mtDNAs, most protein-coding genes terminate with TAA stop codon, which account for 63.16–89.74 %. In the T. ornata mitogenome, 26 protein-coding genes (70.27 %) terminate with TAA stop codon, seven with TAG and four with TGA (Table 2). No incomplete stop codon is found here or in other brown algal mitogenomes.
A total of 25 tRNA genes are detected in the T. ornata mitogenome, which could transport 19 amino acids, but are not sufficient for messenger RNA translation inside the mitochondria. The trnT gene responsible for the transport of threonine was neither detected in the T. ornata mitogenome, nor was it found in other brown algal mtDNAs. However, considering the amino acid Thr is present in many predicted protein products, the trnT gene might have been shifted to the nuclear genome in very early period of brown algal evolution, but such assumption needs to be determined with further genetic evidence.
Unlike the other 24 tRNA genes that could fold into the standard cloverleaf secondary structure, the tRNA gene situated between nad4 and nad5 was variable in structure and function among the different brown algal mitogenomes (Table 3). In the Ectocarpales and Laminariales a 15-bp insertion has been found in the D-loop, which has changed the conventional structure of tRNA. This modified tRNA gene might play a new role instead of old function (Oudot-Le Secq et al. 2006). The insertion was not found in the mitogenomes of Fucales, Desmarestiales, and Dictyotales. In the mtDNAs of T. ornata and Sargassum, this region is predicted as trnI with the anticodon TAT, but the anticodon is TTA for a stop codon in F. vesiculosus, and TTT for the trnK gene in D. dichotoma and Desmarestia viridis.
Overlapping regions
Brown algal mitogenomes usually contain 10–14 overlapping regions, with the exception of P. littoralis which contain 17. The T. ornata genes overlap by a total of 164 bp in 12 different locations from 1 to 66 bp in length (Table 2). All are found in species in the Sargassaceae. Among them, the nad1-tatC overlapping region is the largest one with the size of 66 bp, which is similar to other species in the Fucales: 60 bp in S. thunbergii, 63 bp in S. muticum, 66 bp in S. horneri and F. vesiculosus, and 90 bp in S. hemiphyllum and S. fusiforme.
Ten of the overlapping regions are highly conserved at the family level with respect to length and sequence, including the overlaps of rps8-rpl6 (4 bp), rpl6-rps2 (1), rps2-rps4 (11), rps12-rps7 (7), rps19-rpl2 (14), rps13-rps11 (11), cox2-nad4 (10), nad5-nad6 (16), nad3-rps14 (4), and rrn5-trnM (3). The rpl16-rps3 region is 17 bp in T. ornata, which is identical to that in other species of Sargassaceae except for S. fusiforme, where this region is 32 bp in size with a 15-bp insertion. Comparative analyses of overlapping regions at the order level reveal that seven of the overlapping regions (rps8-rpl6, rpl6-rps2, rps13-rps11, cox2-nad4, nad5-nad6, nad3-rps14, and rrn5-trnM) identified are highly conserved in Fucalean mitogenomes. However, only two of the overlapping regions which are exactly conserved among brown algae of the five orders are rps8-rpl6 and rpl6-rps2 (Oudot-Le Secq et al. 2006).
Non-coding sequences
The non-coding sequences comprise 1872 bp in total, approximately 5.35 % of the T. ornata mitogenome sequence. This is higher than Sargassum (4.29–4.65 %) and lower than F. vesiculosus (5.62 %) (Table 2). On the whole, the spacer content of the Fucales mitogenomes are significantly low compared with that of Desmarestiales (6.06 %), Ectocarpales (6.02–6.81 %. not including C. peregrina: 4.16 %), and Laminariales (5.83–6.49 %), indicating that the Fucales mitogenomes are more compact and economical than that of other orders in the Ectocarpales, Laminariales, Desmarestiales, and Fucales (ELDF) complex (Liu and Pang 2014c).
The total spacer size is positively correlated with brown algal genome size. We compared the spacer size with the mitogenome size of 21 brown algae and estimated their correlation coefficient to be 0.7972 (Fig. 2). Like the vast majority of the mitogenomes, the AT content of intergenic spacer regions is significantly rich in brown algal mtDNAs. In the T. ornata mtDNA, the AT content of non-coding regions is 75.48 %, which is in range of known brown algal mtDNAs from 67.41 % in Pylaiella littoralis to 83.92 % in C. peregrina.
Mitochondria originated from a proteobacterial ancestor through an endosymbiotic event (Liu et al. 2015). A positive correlation has been identified between the increase in AT richness and the DNA loss for these nonfunctional DNA regions in endosymbiotic bacteria (Gómez-Valero et al. 2004). Large intergenic spacer regions normally limited in number and size tend to be eliminated from mitogenomes due to deletional bias and inactivity. The largest spacer region in the T. ornata mtDNA is located between cox3 and atp6 with the size of 251 bp. We previously found that the cox3-atp6 spacer region is the most variable in Sargassum mitogenomes with respect to sequence length and identity, which was directly responsible for the variance in Sargassum genome size (Liu and Pang 2015). The size of this region is 1299 bp in F. vesiculosus containing a conserved orf379, while it has been reduced to 20–113 bp in Sargassum species of subgenus Bactrophycus.
Gene arrangements
The gene order or arrangement of mtDNAs among the species in the orders ELDF has high similarity, varying in ORF number and tRNA gene position. These ELDF arrangement patterns are apparently different from that of Dictyotales representing a more ancestral brown algal lineage (Rousseau et al. 2001; Phillips et al. 2011), in which more genes appear in different positions (Oudot-Le Secq et al. 2006; Liu and Pang 2014c).
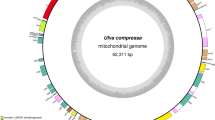
The mitogenome sequences of Desmarestiales, Ectocarpales, Laminariales and Dictyotales are more incongruent with that of T. ornata than five other species in Fucales, considering that more gap regions are detected based on the BRIG map (Fig. 3). The overall mitogenome sequences are fairly conserved in the ELDF, and most regions of the mtDNAs have more than 70 % of sequence identity relative to T. ornata. However, several regions such as the rnl-trnK spacer, cox2 gene, cox3-atp6 spacer, rps14-rns middle region and trnP-rnl spacer, display greater inconsistency. This indicates that mitochondrial variation is more active in these regions during the process of evolution. For example, the cox2 gene in ELDF mtDNAs contains an in-frame insertion with different sizes (2.28–2.99 kb); and the rps14-rns middle region has a high frequency of gene rearrangement.
Comparison of the T. ornata mitogenome against 20 published brown algae. The innermost ring shows GC content (black), and the second innermost one shows the T. ornata mitogenome as reference sequence (purple). The remaining rings show BLAST comparisons of other complete brown algal genomes against T. ornata, which from inside to outside are Fucales (S. muticum, S. hemiphyllum, S. fusiforme, S. horneri and F. vesiculosus. aqua), Desmarestiales (D. viridis. blue), Ectocarpales (C. peregrina, S. lomentaria, P. fascia, E. siliculosus, and P. littoralis. green), Laminariales (U. pinnatifida, C. costata, L. digitata, L. hyperborea, S. japonica, S. longissima, S. angustata, and S. coriacea. yellow), and Dictyotales (D. dichotoma. red). The intensity of the ring color denotes the degree of sequence conservation at that region. Gaps or white spaces indicate highly variable regions (<50 % sequence similarity), six of which are annotated in the outermost words (black). The figure was generated using BRIG software
Sequence identity and phylogenetic analyses
Species in orders ELDF share the identity values from 68.0 to 99.9 % (Table 4), which are significantly higher than that between D. dichotoma and the others (58.7–60.5 %). Previous studies indicated that tRNA genes were most conserved in brown algal mtDNAs, followed by rRNAs and protein-coding genes, with intergenic spacer regions being the most variable (Liu et al. 2014b, c). To examine the phylogenetic relationships within the Phaeophyceae, ML and NJ trees were constructed on the bases of three datasets: 3 rRNA genes, 25 tRNA genes, and 35 protein-coding genes. The current analyses based on different categories of genes produce some incongruent topographies, which are clearly displayed by the variation in the overall topology of phylogenetic trees (Figs. 4, 5).
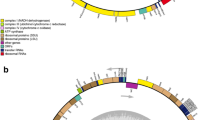
Phylogenetic trees depicting brown algal relationships inferred from a three mitochondrial rRNA genes (rnl, rns and rrn5) and b 25 mitochondrial tRNA genes. Dictyota dichotoma was designated as the outgroup. Numbers on branches indicate maximum likelihood (ML, above) and neighbor-joining (NJ, below) bootstrap values
Phylogenetic trees depicting brown algal relationships inferred from a nucleotide and b amino acid datasets of 35 mitochondrial protein-coding genes (rps2-4, rps7, rps8, rps10-14, rps19; rpl2, rpl5, rpl6, rpl14, rpl16, rpl31; nad1-7, nad9, nad11; cob; cox1-3; atp6, atp8, atp9; and tatC). Dictyota dichotoma was designated as the outgroup. Numbers on branches indicate maximum likelihood (ML, above) and neighbor-joining (NJ, below) bootstrap values
Based on 3 rRNAs or 35 PCG datasets, 5 resolved clades, representing 5 different orders Ectocarpales, Laminariales, Desmarestiales, Fucales, and Dictyotales, are clearly formed with strong support values. Analysis of 25 tRNAs confirmed the ELDF affinity, but failed to cluster F. vesiculosus with five species in the Sargassaceae. The inconsistency of the Desmarestiales position, and the variation of branching patterns in the interior of Ectocarpales, Laminariales, and Fucales clades are observed in the present study. Considering the common origin of brown algal mtDNAs, the observed uncertainty regarding the position and support values of some branches is closely associated with the heterogeneity in evolutionary rate of different categories of genes.
References
Alikhan N, Petty NK, Zakour NLB, Beatson SA (2011) BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genom 12:402
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bittick SJ, Bilotti ND, Peterson HA, Stewart HL (2010) Turbinaria ornata as an herbivory refuge for associate algae. Mar Biol 157:317–323
Charrier B, Bail AL, de Reviers B (2012) Plant Proteus: brown algal morphological plasticity and underlying developmental mechanisms. Trends Plant Sci 17(8):468–477
Draisma SGA, Ballesteros E, Rousseau F, Thibaut T (2010) DNA sequence data demonstrate the polyphyly of the genus Cystoseira and other Sargassaceae genera (Phaeophyceae). J Phycol 46:1329–1345
Gómez-Valero L, Latorre A, Silva FJ (2004) The evolutionary fate of nonfunctional DNA in the bacterial endosymbiont Buchnera aphidicola. Mol Biol Evol 21(11):2172–2181
Guiry MD, Guiry GM (2014) AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org. Accessed on 18 December 2014
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acid Symp Ser 41:95–98
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kogame K, Uwai S, Shimada S, Masuda M (2005) A study of sexual and asexual populations of Scytosiphon lomentaria (Scytosiphonaceae, Phaeophyceae) in Hokkaido, northern Japan, using molecular marker. Eur J Phycol 40:313–322
Le Lann K, Kervarec N, Payri C, Deslandes E, Stiger Pouvreau V (2008) Discrimination of allied species within the genus Turbinaria (Fucales, Phaeophyceae) using HRMAS NMR spectroscopy. Talanta 74:1079–1083
Li TY, Qu JQ, Feng YJ, Liu C, Chi S, Liu T (2014) Complete mitochondrial genome of Undaria pinnatifida (Alariaceae, Laminariales, Phaeophyceae). Mitochondrial DNA. doi:10.3109/19401736.2013.865172
Liu F, Pang SJ (2014a) Complete mitochondrial genome of the brown alga Scytosiphon lomentaria (Scytosiphonaceae, Phaeophyceae). Mitochondrial DNA. doi:10.3109/19401736.2014.953108
Liu F, Pang SJ (2014b) Complete mitochondrial genome of the invasive brown alga Sargassum muticum (Sargassaceae, Phaeophyceae). Mitochondrial DNA. doi:10.3109/19401736.2014.933333
Liu F, Pang SJ (2014c) Mitochondrial phylogenomics reveals a close relationship between Petalonia fascia (Scytosiphonaceae, Phaeophyceae) and Ectocarpus siliculosus. J Appl Phycol. doi:10.1007/s10811-014-0386-3
Liu F, Pang SJ (2015) Mitochondrial genome of Sargassum thunbergii: conservation and variability of mitogenomes within the subgenus Bactrophycus. Mitochondrial DNA. doi:10.3109/19401736.2015.1007328
Liu F, Pang SJ, Chen WZ (2014a) Complete mitochondrial genome of the brown alga Sargassum hemiphyllum (Sargassaceae, Phaeophyceae): comparative analyses. Mitochondrial DNA. doi:10.3109/19401736.2014.953096
Liu F, Pang SJ, Li J, Li X (2014b) Complete mitochondrial genome of the brown alga Colpomenia peregrina (Scytosiphonaceae, Phaeophyceae): genome characterization and comparative analyses. Mitochondrial DNA. doi:10.3109/19401736.2014.958688
Liu F, Pang SJ, Luo MB (2014c) Complete mitochondrial genome of the brown alga Sargassum fusiforme (Sargassaceae, Phaeophyceae): genome architecture and taxonomic consideration. Mitochondrial DNA. doi:10.3109/19401736.2014.936417
Liu F, Pang SJ, Li X, Li J (2015) Complete mitochondrial genome of the brown alga Sargassum horneri (Sargassaceae, Phaeophyceae): genome organization and phylogenetic analyses. J Appl Phycol 27:469–478
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucl Acids Res. doi:10.1093/nar/gkt289
Oudot-Le Secq MP, Fontaine JM, Rousvoal S, Kloareg B, Loiseaux-De Goër S (2001) The complete sequence of a brown algal mitochondrial genome, the Ectocarpale Pylaiella littoralis (L.) Kjellm. J Mol Evol 53:80–88
Oudot-Le Secq MP, Kloareg B, Loiseaux-De Goër S (2002) The mitochondrial genome of the brown alga Laminaria digitata: a comparative analysis. Eur J Phycol 37:163–172
Oudot-Le Secq MP, Loiseaux-De Goër S, Stam WT, Olsen JL (2006) Complete mitochondrial genome of the three brown algae (Heterokonta: Phaeophyceae) Dictyota dichotoma, Fucus vesiculosus and Desmarestia viridis. Curr Genet 49:47–58
Phillips N, Kapraun DF, Garreta AG, Siguan MAR, Rull JL, Soler NS, Lewis R, Kawai H (2011) Estimates of nuclear DNA content in 98 species of brown algae (Phaeophyta). AoB Plants. doi:10.1093/aobpla/plr001
Prathep A, Wichachucherd B, Thongroy P (2007) Spatial and temporal variation in density and thallus morphology of Turbinaria ornata in Thailand. Aquat Bot 86:132–138
Qu JQ, Liu C, Wang XM, Zhang ZB, Chi S, Liu T (2014) Complete mitochondrial genome of Costaria costata shows conservative evolution in Laminariales. Mitochondrial DNA. doi:10.3109/19401736.2013.863290
Ravin NV, Galachyants YP, Mardanov AV, Beletsky AV, Petrova DP, Sherbakova TA, Zakharova YR, Likhoshway YV, Skryabin KG, Grachev MA (2010) Complete sequence of the mitochondrial genome of a diatom alga Synedra acus and comparative analysis of diatom mitochondrial genomes. Curr Genet 56:215–223
Rohfritsch A, Payri C, Stiger V, Bonhomme F (2007) Molecular and morphological relationships between two closely related species, Turbinaria ornata and T. conoides (Sargassaceae, Phaeophyceae). Biochem Syst Ecol 35:91–98
Rohfritsch A, Payri C, Stiger V, Bonhomme F (2010) Habitat-related allelic variation revealed by an anonymous DNA locus in reef-dwelling Turbinaria ornata (Fucales, Phaeophyceae). Bot Mar 53:189–192
Rousseau F, Burrowes R, Peters AF, Kuhlenkamp R, de Reviers B (2001) A comprehensive phylogeny of the Phaeophyceae based on nrDNA sequences resolves the earliest divergences. Comptes Rendus de l’Academie des Sciences. Serie III. Sciences de la Vie (Paris) 324:305–319
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:686–689
Silberfeld T, Rousseau F, de Reviers B (2014) An updated classification of brown algae (Ochrophyta, Phaeophyceae). Cryptogamie Algologie 35(2):117–156
Stewart HL (2004) Hydrodynamic consequences of maintaining an upright posture by different magnitudes of stiffness and buoyancy in the tropical alga Turbinaria ornata. J Mar Syst 49:157–167
Stewart HL (2006) Ontogenetic changes in buoyancy, breaking strength, extensibility, and reproductive investment in a drifting macroalga Turbinaria ornata (Phaeophyta). J Phycol 42:43–50
Stiger V, Payri CE (2005) Natural settlement dynamics of a young population of Turbinaria ornata and phonological comparisons with older populations. Aquat Bot 81:225–243
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20(17):3252–3255
Wynne MJ (2002) Turbinaria foliosa sp. nov. (Fucales, Phaeophyceae) from the Sultanate of Oman, with a census of currently recognized species in the genus Turbinaria. Phycol Res 50:283–293
Yotsukura N, Shimizu T, Katayama T, Druehl LD (2010) Mitochondrial DNA sequence variation of four Saccharina species (Laminariales, Phaeophyceae) growing in Japan. J Appl Phycol 22:243–251
Zhang J, Wang XM, Liu C, Jin YM, Liu T (2013) The complete mitochondrial genomes of two brown algae (Laminariales, Phaeophyceae) and phylogenetic analysis within Laminaria. J Appl Phycol 25:1247–1253
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 41206146, 41176135), the Scientific Research Foundation for Outstanding Young Scientists of Shandong Province (No. BS2013HZ004), the Special Fund of the membership of Youth Innovation Promotion Association of Chinese Academy of Sciences (No. 2015164), the 863 Hi-Tech Research and Development Program of China (No. 2012AA10A413), the Science and Technology Service Network Program of Chinese Academy of Sciences (No. KFJ-EW-STS-060), and the Knowledge Innovation Programs of Chinese Academy of Sciences (No. KSCX2-EW-B-14).
Compliance with Ethical standards
The authors declare that they have no conflict of interest. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with human participants or animals performed by any of the authors. Informed consent was obtained from all individual participants included in the study.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Liu, F., Pang, S. Mitochondrial genome of Turbinaria ornata (Sargassaceae, Phaeophyceae): comparative mitogenomics of brown algae. Curr Genet 61, 621–631 (2015). https://doi.org/10.1007/s00294-015-0488-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-015-0488-8