Abstract
To gain further insight into the evolution of mitochondrial genomes (mtDNAs) in Phaeophyceae, the first recorded characterization of an Ishigeophycidae mtDNA from Ishige okamurae (Yendo), and only the second recorded characterization of a Dictyotophycidae mtDNA from Dictyopteris divaricata (Okamura) Okamura are presented in this study. The 35,485 bp I. okamurae mtDNA contained 36 protein-coding genes (PCGs), 22 tRNAs, three rRNAs, and four open reading frames (orfs), and the 32,021 bp D. divaricata mtDNA harbored 35 PCGs, 25 tRNAs, three rRNAs, and three orfs. The A + T content in D. divaricata (61.69%) was the lowest recorded in sequenced brown algal mtDNAs. The I. okamurae mtDNA displayed unique genome features including an elevated start-codon usage bias for GTG, while the organization of D. divaricata mtDNA was identical to that of Dictyota dichotoma. Phylogenetic analysis based on the amino acid sequence dataset of 35 PCGs indicated that I. okamurae (Ishigeophycidae) diverged early from the Fucophycidae–Dictyotophycidae complex, which was confirmed by the comparative analysis of the mitogenome structure. The novel mitogenome data made available by this study have improved our understanding of the evolution, phylogenetics, and genomics of brown algae.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Phaeophyceae, or brown algae, are a group of multicellular photosynthetic organisms in the Heterokontophyta (Stramenopiles) (Yang et al. 2012; Charrier et al. 2012). Most brown algae species are found predominantly in marine environments, with only a few freshwater species identified thus far (e.g., Pleurocladia lacustris) (Wehr et al. 2013). Phylogenetic studies have shown that Discosporangiales and Ishigeales were the monophyletic early-diverging lineages, while the other brown algae were clustered into two super-clades representing two subclasses, Dictyotophycidae and Fucophycidae (Guiry and Guiry 2018; Liu et al. 2017a). Brown algae are important primary producers in our oceans, forming essential ecological structures known as marine forests and seaweed beds that provide food and shelter to a diverse range of invertebrates, fishes, sea turtles, birds, and mammals (Laffoley et al. 2011, Witherington et al. 2012; Wikipedia 2018). Large-scale cultivation of commercial kelps (e.g., Saccharina and Undaria) using artificial floating raft systems is commonplace in several East Asian countries (Tseng 2001). On the other hand, some Sargassum species (e.g., Sargassum horneri and Sargassum natans) can maintain high growth rates while in a floating state within the pelagic zone, forming harmful macroalgal blooms known as “golden tides,” which can cause considerable damage to the local environment and economy (Smetacek and Zingone 2013; Liu et al. 2017b).
The launch of several genome sequencing projects to understand the evolutionary history of brown algae led to the complete genome sequences of two species, Ectocarpus siliculosus and Saccharina japonica, to date (Cock et al. 2010; Ye et al. 2015). Considering that more than 2000 brown algal species have been identified worldwide (Guiry and Guiry 2018), our understanding of the evolutionary history of brown algae still remains very limited. Organelle genomes (mtDNAs and cpDNAs) carry important genetic information widely utilized in phylogenetics and comparative genomics implemented to broaden our understanding of Phaeophyceae evolutionary history. The advent of molecular systematics has brought additional insights, reshaping evolutionary concepts associated with brown algae (Silberfeld et al. 2014).
Previous work has unveiled the organelle genome sequences in several brown algal species (e.g., Oudot-Le Secq et al. 2001, 2002, 2006; Le Corguillé et al. 2009; Yotsukura et al. 2010; Zhang et al. 2013; Wang et al. 2013; Liu et al. 2015, 2017a, b; Liu and Pang 2016b). So far, the complete mitochondrial genomes (mtDNAs) of 41 brown algal species have been sequenced (Table 1), which has provided considerable insight into evolutionary and phylogenetic concepts of brown algae. The sequenced mtDNAs are from several orders belonging to two subclasses including the order Dictyotales in subclass Dictyotophycidae (1 species), and four orders in Fucophycidae including Ectocarpales (7 species), Laminariales (15 species), Desmarestiales (1 species), and Fucales (17 species) (i.e., the ELDF complex). There are some general observations of the complete mtDNAs reported thus far including that they are single circular molecules of 31.6–58.5 kb in size; harbor 35–36 protein-coding genes (PCGs), three ribosomal RNA genes (rRNAs), 24–26 transfer RNA genes (tRNAs), and 2–16 open reading frames (orfs); and display highly conserved genome architecture (Liu and Pang 2015a).
Ishige okamurae Yendo grows in the upper intertidal zones of the coastlines of South China, Japan, and Korea (Lee et al. 2009), and is the holotype of the order Ishigeales, subclass Ishigeophycidae (Guiry and Guiry 2018). Previous studies have shown Ishigeales to be a monophyletic early-diverging lineage in Phaeophyceae, yet very little is known about the genome information within this order (Liu et al. 2017a). Dictyopteris divaricata (Okamura) Okamura inhabits littoral and sublittoral rock zones and belongs to the order Dictyotales, subclass Dictyotophycidae (Guiry and Guiry 2018). To date, only one species, Dictyota dichotoma, from Dictyotales has had its mitogenome completely sequenced (Oudot-Le Secq et al. 2006).
In this study, we present the first recorded characterization of an Ishigeophycidae mtDNA from Ishige okamurae (Yendo), and only the second recorded characterization of a Dictyotophycidae mtDNA from Dictyopteris divaricata (Okamura) Okamura. The two novel mtDNA sequences are then compared to the mtDNA sequences reported in Dictyotales, Ectocarpales, Laminariales, Fucales, and Desmarestiales, providing further insights into the evolution of mitogenomes in the class Phaeophyceae.
Materials and Methods
Algal Sampling and DNA Extraction
Adult plants of Ishige okamurae Yendo were collected from the rocky shore of Houjiang, Zhangpu, Fujian Province, China (23°54′N, 117°46′E) on January 2, 2018. Algal thalli of Dictyopteris divaricata (Okamura) Okamura were sampled from the rocky shore of No. 3 bathing beach of Qingdao, Shandong Province, China (36°03′N, 120°22′E) on July 16, 2016 (Liu et al. 2017a). Seaweed samples were transported to the laboratory in coolers (5–8 °C) after collection. Clean thalli without epiphytic brown algae were examined by microscopy and selected for storage in − 80 °C freezers. Frozen tissue from the original algal samples was used for DNA extraction. The algal tissue was ground to fine powder in liquid nitrogen using a Scientz-48 Tissue Grinder (Scientz Biotech, Ningbo, China). Total DNA was extracted using a Plant Genomic DNA Kit (Tiangen Biotech, Beijing, China) according to the manufacturer’s instructions. The concentration and quality of isolated DNA was evaluated by electrophoresis on 1.0% agarose gel.
PCR Amplification, Sequencing, and Assembly
Both species have previously been identified by morphological and molecular methods (data not given; Liu et al. 2017a). The mitochondrial genome sequences of I. okamurae and D. divaricata were amplified using the long PCR and primer walking techniques (Cheng et al. 1994). The novel primer sets were designed and used to amplify the entire I. okamurae and D. divaricata mitochondrial genomes into four and five long fragments, respectively (Supplementary data: Table S1). PCR reactions were carried out in 50 L reaction mixtures containing 32 L of sterile distilled H2O, 10 L of 5 × PrimeSTAR GXL buffer (5 mM Mg2+ plus, Takara, Japan), 4 L of dNTP mixture (2.5 mM each), 1 L of each primer (10 µM), 1 L of PrimeSTAR GXL DNA polymerase (1.25 units/L, Takara, Japan), and 1 L of DNA template (approximately 50 ng). PCR amplification was performed on a TC1000-G Thermal Cycler (Scilogex, USA) with an initial denaturation at 94 °C for 3 min, followed by 30–35 cycles of denaturation at 94 °C for 20 s, annealing at 50–52 °C for 50 s, extension at 68 °C for 1 min/kb, and a final extension at 68 °C for 10 min (Liu et al. 2015). Long PCR products were purified using a QIAquick Gel Extraction Kit (Qiagen, Germany) and Sanger sequencing from both forward and reverse reactions was performed for all fragments, causing the whole mitogenome sequences to be covered twice. In addition, some specific gene sequences for the I. okamurae mtDNA including cox2, nad6, nad11, were amplified and sequenced to confirm their unique characteristics. Sequencing reactions were performed using ABI 3730 XL automated sequencers (Applied Biosystems, USA). The DNA sequences were assembled using Geneious 7.1 software (Biomatters, http://www.geneious.com). This resulted in one scaffold of 35,485 bp for I. okamurae and 32,021 bp for D. divaricata.
Genome Annotation and Analysis
Protein-coding genes (PCGs) and open reading frames (orfs) were annotated using NCBI ORF Finder, Dual Organellar Genome Annotator (DOGMA) (Wyman et al. 2004), and BLAST similarity searches of the non-redundant databases at NCBI (Altschul et al. 1997). Ribosomal RNA genes were delimited by direct comparison to sequenced brown algal orthologues using MEGA v.7.0 (Kumar et al. 2016). Transfer RNA genes were identified by reconstructing their cloverleaf structures using the tRNAscan-SE 1.21 software with default parameters (Schattner et al. 2005; Lowe and Chan 2016). The physical maps of the circular mitochondrial genomes were constructed with Organellar Genome DRAW (OGDraw) (Lohse et al. 2013). Base composition was determined by MEGA v.7.0. The mitochondrial genome sequences of I. okamurae and D. divaricata have been deposited in GenBank with the Accession Numbers MG940857 and MG940856, respectively.
Phylogenetic Analysis
Phylogenetic relationships within Phaeophyceae were analyzed based on the amino acid (aa) sequence dataset of 35 PCGs including 17 ribosomal proteins (rps2–4, 7, 8, 10–14, and 19; rpl2, 5, 6,14, 16, and 31), 10 NADH dehydrogenase complex subunits (nad1–7, 4L, 9, and 11), three cytochrome oxidase subunits (cox1–3), three ATPase subunits (atp6, 8, and 9), apocytochrome b (cob), and a secY-independent transporter protein (tatC). Heterosigma akashiwo (GQ222228) was selected as an out-group taxon for analysis of the aa dataset. We re-annotated the mitochondrial genome of H. akashiwo and found that it shared the same 35 PCGs as Phaeophyceae. The aa sequences were subjected to concatenated alignments using ClustalX 1.83 with the default settings (Thompson et al. 1997). Bayesian Inference (BI) analysis was performed based on the best scoring alternative model of MtREV + G + I using MrBayes v.3.2 (Huelsenbeck and Ronquist 2001). One million generations were run and posterior probabilities determined using the Markov chain Monte Carlo (MCMC) method for tree reconstructions. Every 1000th generation was saved, and the first 100 generations discarded as burn-in. Posterior probability values for the majority-rule consensus trees constructed were calculated. Maximum likelihood (ML) analysis of the aa dataset was run based on the JTT matrix-based model (Jones et al. 1992) with 1000 bootstrap replicates using MEGA v.7.0 (Kumar et al. 2016). Multiple sequence alignment of the brown algal mitochondrial genome sequences was performed using the Mauve Genome Alignment v.2.3.1 (Darling et al. 2004) using the progressive Mauve algorithm (Darling et al. 2010).
Results and Discussion
Genome Features and Codon Usage
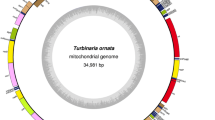
The mitochondrial genomes of I. okamurae and D. divaricata were 35,485 bp and 32,021 bp, respectively (Fig. 1a, b). Comparative analysis of the mtDNA size of 19 brown algal genera indicated that the Dictyotales mtDNAs tend to be smaller (Fig. 2a). The A + T content of I. okamurae and D. divaricata were 64.66% and 61.69%, respectively. To date, the D. divaricata mtDNA displayed the lowest A + T content of all sequenced brown algal mtDNAs (Fig. 2b). The spacer contents were 4.09% and 4.06% in the I. okamurae and D. divaricata mtDNAs, respectively, both of which were lower than all sequenced mtDNAs in Fucophycidae (4.16–6.81%) but higher than D. dichotoma (3.21%) (Fig. 2c). The sequenced mtDNAs in Ishigeophycidae and Dictyotophycidae had an elevated number of reduced spacer regions, suggesting that their mitogenomes evolved to be more compact than Fucophycidae. The I. okamurae genes overlapped by a total of 104 bp in 15 different locations ranging from 1 to 23 bp, while the D. divaricata genes overlapped by 160 bp in 12 locations from 1 to 60 bp. Two overlapping regions, rps8-rpl6 (4 bp) and rpl6-rps2 (1 bp), previously observed to be highly conserved in Fucophycidae and Dictyotophycidae mtDNAs (Liu and Pang 2015a), were also conserved in I. okamurae and D. divaricata.
The mitochondrial genome maps of aIshige okamurae and bDictyopteris divaricata. Annotated genes are colored according to the functional categories. Genes on the inside are transcribed in the clockwise direction, whereas genes on the outside are transcribed in the counterclockwise direction. The ring of bar graphs on the inner circle shows the GC content in dark gray
Most of the PCGs in I. okamurae and D. divaricata mtDNAs had a methionine (ATG) start codon and a TAA stop codon (Fig. 3a, b), which was similar to that in other brown algal mtDNAs (Graf et al. 2017). However, compared with the mtDNAs in Fucophycidae, some novel features in codon usage were observed in mtDNAs of I. okamurae and D. divaricata. An increase in a start-codon usage bias for GTG was observed in the I. okamurae mtDNA where five genes (nad5, nad9, rps2, rps8, and orf136) started with the GTG codon (Fig. 3a). The frequency of stop-codon usage bias for TGA was higher than TAG in I. okamurae and D. divaricata, which was similar to what was reported for D. dichotoma (Fig. 3b, c). However, the usage frequency of TGA was lower than TAG in nearly all mtDNAs of Fucophycidae. The variance of codon usage in brown algal mtDNAs suggests a diverse evolutionary pattern in different lineages.
Novel Features in Genes
The mtDNA of I. okamurae contained a total of 65 genes including 36 protein-coding genes (PCGs), 22 tRNA genes, three rRNA genes, and four orfs (Fig. 1a). The D. divaricata mtDNA harbored 66 genes including 35 protein-coding genes (PCGs), 25 tRNA genes, three rRNA genes, and three orfs (Fig. 1b). The gene content of D. divaricata was identical to that previously reported in D. dichotoma. Unlike the other brown algal mitogenomes, the nad6 gene in the I. okamurae mtDNA was split into two genes, nad6a and nad6b, which were 7.5 kb apart from each other (Fig. 1a). The two nad6 genes can be independently transcribed into two mRNAs and translated into separate polypeptides (Edqvist et al. 2000). The observed nad6 gene in the I. okamurae mtDNA might be associated with the posttranslational fusion of the two polypeptides (Odintsova and Yurina 2005).
The nad11 in I. okamurae was approximately 180 aa longer than its homologues in Fucophycidae and Dictyotophycidae. Based on protein BLAST, the extra 180 aa was predicted to be an Actinobacteria sulfotransferase. The size of atp8 in D. divaricata was identical to that in D. dichotoma but was approximately 30 aa longer than those found in I. okamurae and Fucophycidae. Previous studies showed that the cox2 gene in Fucophycidae mtDNAs contained a large in-frame insertion which introduced an extra region about 761–1010 aa long in the middle of this gene (Oudot-Le Secqet al. 2006, Liu et al. 2015; Graf et al. 2017). However, this insertion was absent in the cox2 of D. divaricata and D. dichotoma. In I. okamurae, however, we observed that a much smaller insertion of approximately 330 aa was present in the cox2 gene.
Two conserved orfs, located in trnW(cca)-trnM3(cau) and atp9-rpl16 spacers, were shared by nearly all known brown algal mtDNAs, with the exception of I. okamurae and Pylaiella littoralis where the conserved orfs located in trnW(cca)-trnM3(cau) are lost. Unique species-specific orfs identified include one orf (orf51) located between nad5 and nad6 in two Dictyotales mtDNA sequences, and three orfs (orf166, orf273, and orf212) with unknown functions only detected in I. okamurae.
Four tRNA genes, trnG(gcc), trnL3(caa), trnD(guc), and trnE(uuc), common in the mtDNAs of Fucophycidae and Dictyotophycidae, were lost from I. okamurae. An extra tRNA gene, trnM(cau), located between rns and trnM1(cau) that transcribed onto different strands was found in I. okamurae only (Fig. 3).
Genome Rearrangement
The structures of brown algal mtDNAs were analyzed and compared by combining the brown algal basal lineages of I. okamurae and D. divaricate with that of Fucophycidae and Dictyotophycidae to understand the evolution of mtDNA genome architecture. The genome organization of I. okamurae mtDNA was different from that of Fucophycidae and Dictyotophycidae, while the architecture of D. divaricata mtDNA was identical to that of D. dichotoma. The brown algal mtDNA demonstrated a highly conserved genome organization at the order level (Liu and Pang 2015a), akin to cpDNAs in Phaeophyceae (Liu et al. 2017a).
Two ribosomal gene blocks, rps8-rpl6-rps2-rps4 and rpl16-rps3-rps9-rpl2-rps13-rps11, and two tRNA gene blocks, MLHCNF and MQL, were observed in all the sequenced mtDNAs of the three subclasses revealing that brown algal mitogenomes are highly conserved (Liu and Pang 2015a). The rps8-rpl6-rps2-rps4 ribosomal gene block was also conserved in the mtDNA of H. akashiwo (Masuda et al. 2011), which belonged to the class Raphidophyceae and has a sister relationship with Phaeophyceae (Guiry and Guiry 2018). Furthermore, the second ribosomal gene block, rpl16-rps3-rps9-rpl2-rps13-rps11, was reduced to the rpl16-rps3-rps9-rpl2 block in H. akashiwo.
Despite generally maintaining a highly conserved mitogenomic architecture, the study observed that two regions, rps14-rns and rrn5-trnM2(cau), experienced greater variability in brown algal mtDNAs, exhibiting structural diversity not only between the three subclasses but also at the inter and intra-order level in Fucophycidae. The SPAV tRNA gene block was present in I. okamurae mtDNA, but absent in Fucophycidae and Dictyotophycidae. The trnM1(cau)-rns-rrn5-rnl rRNA gene block was observed in I. okamurae and H. akashiwo mtDNAs, but was interrupted in the mtDNAs of Fucophycidae and Dictyotophycidae as a result of genome rearrangement events (Fig. 4).
Basal Lineage of Brown Algae
Phylogenetic relationship analyses based on the aa sequence dataset of 35 PCGs showed that the 19 species representing 19 genera from six orders were clustered into three clades, Fucophycidae, Dictyotophycidae, and Ishigeophycidae, with high support values (BI/ML = 100%) (Fig. 5). Fucophycidae and Dictyotophycidae were sister taxa that evolved from a common ancestor forming the Fucophycidae–Dictyotophycidae (FD) complex. The early divergence of I. okamurae (Ishigeophycidae) from the FD complex, which includes Ectocarpales, Laminariales, Desmarestiales, Fucales, and Dictyotales, suggests its basal lineage in brown algae. The polytomy of Fucophycidae, also known as the brown algal crown radiation (BACR) (Rousseau et al. 2001), likely represents a gradual diversification spanning most of the Lower Cretaceous rather than a sudden radiation (Silberfeld et al. 2010).
Phylogenetic tree constructed from analysis of amino acid (aa) sequences of 35 PCGs in brown algal mtDNAs. The tree was rooted with Heterosigma akashiwo (Raphidophyceae). The numbers at internal nodes (BI/ML) indicated Bayesian Inference (BI) posterior probability values and maximum likelihood (ML) bootstrap values, respectively. Branch lengths are proportional to the amount of amino acid substitutions per site, which are indicated by the scale bar below the tree
The phylogenetic relationships uncovered in our study were consistent with the result of phylogeny inferred from the combined dataset of seven genes (Silberfeld et al. 2010, 2014), and was further confirmed by comparative analysis of the mitogenome structure of brown algae. The genome organization of Dictyotophycidae showed greater similarity to that of Fucophycidae when compared to I. okamurae (Supplementary data: Fig. S1). The I. okamurae mtDNA exhibited a novel genome architecture, and more genes appeared in new positions (Fig. 4), suggesting a distant relationship with the FD complex.
The mitogenomic data uncovered in this study emphasize that the Phaeophyceae basal lineage mtDNAs exhibits novel features, suggested an even greater diversity of mitogenome evolution in brown algae than previously thought, providing further insights into the evolutionary history, phylogenetics, and genomics of brown algae.
References
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Amaral-Zettler LA, Dragone NB, Schell J, Slikas B, Murphy LG, Morrall CE, Zettler ER (2017) Comparative mitochondrial and chloroplast genomics of a genetically distinct form of Sargassum contributing to recent “golden tides” in the Western Atlantic. Ecol Evol 7:516–525
Bi YH, Zhou ZG (2016) Complete mitochondrial genome of the brown alga Sargassum vachellianum (Sargassaceae, Phaeophyceae). Mitochondrial DNA Part A 27(4):2796–2797
Charrier B, Bail AL, de Reviers B (2012) Plant proteus: brown algal morphological plasticity and underlying developmental mechanisms. Trends Plant Sci 17(8):468–477
Cheng S, Chang SY, Gravitt P, Respess R (1994) Long PCR. Nature 369:684–685
Cock JM, Sterck L, Rouzé P, Scornet D, Allen AE, Amoutzias G, Anthouard V, Artiguenave F, Aury J-M, Badger JH, Bank Beszteri B, Billiau K, Bonnet E, John H, Bothwell JH, Bowler C, Boyen C, Brownlee C, Carrano CJ, Charrier B, Cho GY, Coelho SM, Collén J, Corre E, Silva CD, Delage L, Delaroque N, Dittami SM, Doulbeau S, Elias M, Farnham G, Gachon CMM, Gschloessl B, Svenja Heesch S, Kamel Jabbari K, Claire Jubin C, Kawai H, Kimura K, Kloareg B, Küpper FC, Lang D, Bail AL, Leblanc C, Lerouge P, Lohr M, Lopez PJ, Martens C, Maumus F, Michel G, Miranda-Saavedra D, Morales J, Moreau H, Motomura T, Nagasato C, Napoli CA, Nelson DR, Nyvall-Collén P, Peters AF, Pommier C, Potin P, Poulain J, Quesneville H, Read B, Rensing SA, Ritter A, Rousvoal S, Samanta M, Samson G, Schroeder DC, Ségurens B, Strittmatter M, Tonon T, Tregear JW, Valentin K, von Dassow P, Yamagishi T, de Peer YV, Wincker P (2010) The Ectocarpus genome and the independent evolution of multicellularity in the brown algae. Nature 465:617–621
Darling AE, Mau B, Blatter FR, Perna NT (2004) Mauve: multiple alignment of conserved genomic sequence with rearrangements. Genome Res 14:1394–1403
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss, and rearrangement. PLoS ONE 5:e11147
Edqvist J, Burger G, Gray MW (2000) Expression of mitochondrial protein-coding genes in Tetrahymena pyriformis. J Mol Biol 297:381–393
Graf L, Kim YJ, Cho GY, Miller KA, Yoon HS (2017) Plastid and mitochondrial genomes of Coccophora langsdorfii (Fucales, Phaeophyceae) and the utility of molecular markers. PLoS ONE 12(11):e0187104
Guiry MD, Guiry GM (2018) AlgaeBase. World-wide electronic publication. National University of Ireland, Galway. http://www.algaebase.org. Accessed 26 Jan 2018
Huelsenbeck JP, Ronquist F (2001) MrBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Hughey JR, Gabrielson PW (2017) The complete mitogenome of the rockweed Fucus distichus (Fucaceae, Phaeophyceae). Mitochondrial DNA Part B Resour 2(1):203–204
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8:275–282
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Laffoley Dd’A, Roe HSJ, Angel MV, Ardron J, Bates NR, Boyd LL, Brooke S, Buck KN, Carlson CA, Causey B, Conte MH, Christiansen S, Cleary J, Donnelly J, Earle SA, Edwards R, Gjerde KM, Giovannoni SJ, Gulick S, Gollock M, Hallet J, Halpin P, Hanel R, Hemphill A, Johnson RJ, Knap AH, Lomas MW, McKenna SA, Miller MJ, Miller PI, Ming FW, Moffitt R, Nelson NB, Parson L, Peters AJ, Pitt J, Rouja P, Roberts J, Roberts J, Seigel DA, Siuda A, Steinberg DK, Stevenson A, Sumaila VR, Swartz W, Trott TM, Vats V (2011) The protection and management of the Sargasso Sea: the golden floating rainforest of the Atlantic Ocean: summary science and supporting evidence case. Sargasso Sea Alliance, Bermuda, pp 44
Le Corguillé G, Pearson G, Valente M, Viegas C, Gschloessl B, Corre E, Bailly X, Peters AF, Jubin C, Vaccherie B, Cock JM, Leblanc C (2009) Plastid genomes of two brown algae, Ectocarpus siliculosus and Fucus vesiculosus: further insights on the evolution of red-algal derived plastids. BMC Evol Biol 9:253
Lee KM, Boo GH, Risomena-Rodriquez R, Shin JA, Boo SM (2009) Classification of the genus Ishige (Ishigeales, Phaeophyceae) in the north Pacific Ocean with recognition of I. foliacea based on plastid rbcL and mitochondrial cox3 gene sequences. J Phycol 45:906–913
Li TY, Qu JQ, Feng YJ, Liu C, Chi S, Liu T (2015) Complete mitochondrial genome of Undaria pinnatifida (Alariaceae, Laminariales, Phaeophyceae). Mitochondrial DNA 26(6):953–954
Liu F, Pang SJ (2015a) Mitochondrial genome of Turbinaria ornata (Sargassaceae, Phaeophyceae): comparative mitogenomics of brown algae. Curr Genet 61:621–631
Liu F, Pang SJ (2015b) Mitochondrial phylogenomics reveals a close relationship between Petalonia fascia (Scytosiphonaceae, Phaeophyceae) and Ectocarpus siliculosus. J Appl Phycol 27:1021–1028
Liu F, Pang SJ (2016a) Complete mitochondrial genome of the brown alga Scytosiphon lomentaria (Scytosiphonaceae, Phaeophyceae). Mitochondrial DNA Part A 27(2):1494–1496
Liu F, Pang SJ (2016b) Chloroplast genome of Sargassum horneri (Sargassaceae, Phaeophyceae): comparative chloroplast genomics of brown algae. J Appl Phycol 28:1419–1426
Liu F, Pang SJ (2016c) Complete mitochondrial genome of the invasive brown alga Sargassum muticum (Sargassaceae, Phaeophyceae). Mitochondrial DNA Part A 27(2):1129–1130
Liu F, Pang SJ (2016d) Mitochondrial genome of Sargassum thunbergii: conservation and variability of mitogenomes within the subgenus Bactrophycus. Mitochondrial DNA Part A 27(5):3186–3188
Liu F, Pang SJ. Li X, Li J (2015) Complete mitochondrial genome of the brown alga Sargassum horneri (Sargassaceae, Phaeophyceae): genome organization and phylogenetic analyses. J Appl Phycol 27:469–478
Liu F, Pang SJ, Chen WZ (2016a) Complete mitochondrial genome of the brown alga Sargassum hemiphyllum (Sargassaceae, Phaeophyceae): comparative analyses. Mitochondrial DNA Part A 27(2):1468–1470
Liu F, Pang SJ, Li J, Li X (2016b) Complete mitochondrial genome of the brown alga Colpomenia peregrina (Scytosiphonaceae, Phaeophyceae): genome characterization and comparative analyses. Mitochondrial DNA Part A 27(2):1601–1603
Liu F, Pang SJ, Luo MB (2016c) Complete mitochondrial genome of the brown alga Sargassum fusiforme (Sargassaceae, Phaeophyceae): genome architecture and taxonomic consideration. Mitochondrial DNA Part A 27(2):1158–1160
Liu F, Jin Z, Wang Y, Bi YP, Melton JT (2017a) Plastid genome of Dictyopteris divaricata (Dictyotales, Phaeophyceae): understanding the evolution of plastid genomes in brown algae. Mar Biotechnol 19:627–637
Liu F, Li XD, Che ZW (2017b) Mitochondrial genome sequences uncover evolutionary relationships of two Sargassum subgenera, Bactrophycus and Sargassum. J Appl Phycol 29:3261–3270
Liu F, Pan J, Zhang ZS, Moejes FW (2018) Organelle genomes of Sargassum confusum (Fucales, Phaeophyceae): mtDNA vs cpDNA. J Appl Phycol 30:2715–2722
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucl Acids Res. https://doi.org/10.1093/nar/gkt289
Lowe TM, Chan PP (2016) tRNAscan-SE On-line: search and contextual analysis of transfer RNA genes. Nucleic Acids Res 44:54–57
Masuda I, Kamikawa R, Ueda M, Oyama K, Yoshimatsu S, Inagaki Y, Sako Y (2011) Mitochondrial genomes from two red tide forming raphidophycean algae Heterosigma akashiwo and Chattonella marina var. marina. Harmful Algae 10:130–137
Odintsova MS, Yurina NP (2005) Genomics and evolution of cellular organelles. Russ J Genet 41(9):957–967
Oudot-Le Secq MP, Fontaine JM, Rousvoal S, Kloareg B, Loiseaux-De Goër S (2001) The complete sequence of a brown algal mitochondrial genome, the Ectocarpale Pylaiella littoralis (L.) Kjellm. J Mol Evol 53:80–88
Oudot-Le Secq MP, Kloareg B, Loiseaux-De Goër S (2002) The mitochondrial genome of the brown alga Laminaria digitata: a comparative analysis. Eur J Phycol 37:163–172
Oudot-Le Secq MP, Loiseaux-De Goër S, Stam WT, Olsen JL (2006) Complete mitochondrial genome of the three brown algae (Heterokonta: Phaeophyceae) Dictyota dichotoma, Fucus vesiculosus and Desmarestia viridis. Curr Genet 49:47–58
Qu JQ, Liu C, Wang XM, Zhang ZB, Chi S, Liu T (2015) Complete mitochondrial genome of Costaria costata shows conservative evolution in Laminariales. Mitochondrial DNA 26(6):919–920
Rousseau F, Burrowes R, Peters AF, Kuhlenkamp R, de Reviers B (2001) A comprehensive phylogeny of the Phaeophyceae based on nrDNA sequences resolves the earliest divergences. Comptes Rendus de l’Acad des Sci Serie III Sci de la Vie (Paris) 324(4):305–319
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:686–689
Silberfeld T, Leigh JW, Verbruggen H, Cruaud C, de Reviers B, Rousseau F (2010) A multi-locus time-calibrated phylogeny of the brown algae (Heterokonta, Ochrophyta, Phaeophyceae): investigating the evolutionary nature of the “brown algal crown radiation”. Mol Phylogenet Evol 56:659–674
Silberfeld T, Rousseau F, de Reviers B (2014) An updated classification of brown algae (Ochrophyta, Phaeophyceae). Cryptogam Algol 35(2):117–156
Smetacek V, Zingone A (2013) Green and golden seaweed tides on the rise. Nature 504:84–88
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Tseng CK (2001) Algal biotechnology industries and research activities in China. J Appl Phycol 13(4):375–380
Wang X (2014) Organellar genome evolution and phylogeographic relationships among populations of freshwater brown algae: Heribaudiella fluviatilis, Bodanella lauterborni and Pleurocladia lacustris. Fordham University, New York
Wang X, Shao Z, Fu W, Yao J, Hu Q, Duan D (2013) Chloroplast genome of one brown seaweed, Saccharina japonica (Laminariales, Phaeophyta): its structural features and phylogenetic analyses with other photosynthetic plastids. Mar Genom 10:1–9
Wehr JD, Stancheva R, Truhn K, Sheath RG (2013) Discovery of the rare freshwater brown alga Pleurocladia lacustris (Ectocarpales, Phaeophyceae) in California Streams. West N Am Nat 73(2):148–157
Wikipedia (2018) Brown algae. Wikipedia, The Free Encyclopedia. https://en.wikipedia.org/wiki/Brown_algae. Accessed 29 Jan 2018
Witherington B, Hirama S, Hardy R (2012) Young sea turtles of the pelagic Sargassum dominated drift community: habitat use, population density, and threats. Mar Ecol Prog Ser 463:1–22
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20(17):3252–3255
Yang EC, Boo GH, Kim HJ, Cho SM, Boo SM, Andersen RA, Yoon HS (2012) Supermatrix data highlight the phylogenetic relationships of photosynthetic stramenopiles. Protist 163:217–231
Ye NH, Zhang XW, Miao M, Fan X, Zheng Y, Xu D, Wang JF, Zhou L, Wang DS, Gao Y, Wang YT, Shi WY, Ji PF, Li DM, Guan Z, Shao CW, Zhuang ZM, Gao ZQ, Qi J, Zhao FQ (2015) Saccharina genomes provide novel insight into kelp biology. Nat Commun 6:6986
Yotsukura N, Shimizu T, Katayama T, Druehl LD (2010) Mitochondrial DNA sequence variation of four Saccharina species (Laminariales, Phaeophyceae) growing in Japan. J Appl Phycol 22:243–251
Zhang J, Wang XM, Liu C, Jin YM, Liu T (2013) The complete mitochondrial genomes of two brown algae (Laminariales, Phaeophyceae) and phylogenetic analysis within Laminaria. J Appl Phycol 25:1247–1253
Zhang L, Wang XM, Liu T, Wang GL, Chi S, Liu C, Wang HY (2016) Complete mitochondrial genome of Kjellmaniella crassifolia (Laminariaceae, Phaeophyceae): Laminaria and Saccharina are distinct genus. Mitochondrial DNA Part A 27(6):4592–4594
Acknowledgements
The authors wish to thank Xingfeng Liu, Wei Luan, Guiyang Li, Zhe Jin, and Jun Qi for their assistance in algal collection and data analysis. This work was financially supported by the Key Research Program of Frontier Sciences, Chinese Academy of Sciences (No. QYZDB-SSW-DQC023), the Marine S&T Fund of Shandong Province for Pilot National Laboratory for Marine Science and Technology (Qingdao) (No. 2018SDKJ0504), the National Natural Science Foundation of China (No. 41876165), the S&T Innovation Project of Pilot National Laboratory for Marine Science and Technology (Qingdao) (No. 2016ASKJ02), the Youth Innovation Promotion Association, Chinese Academy of Sciences (No. 2015164), the Open Research Fund of Key Laboratory of Integrated Marine Monitoring and Applied Technologies for Harmful Algal Blooms, S.O.A. (No. MATHAB201701), and the Earmarked Fund for Modern Agro-industry Technology Research System in Shandong Province of China (No. SDAIT-26-09).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declare that they have no conflict of interest
Ethics Approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
239_2018_9881_MOESM1_ESM.doc
Supplementary material 1 Table S1 The specific primers used to amply the long fragments which were assembled into the complete mitochondrial genomes of Ishige okamurae and Dictyopteris divaricata. (DOC 45 KB)
Rights and permissions
About this article
Cite this article
Liu, F., Zhang, Y., Bi, Y. et al. Understanding the Evolution of Mitochondrial Genomes in Phaeophyceae Inferred from Mitogenomes of Ishige okamurae (Ishigeales) and Dictyopteris divaricata (Dictyotales). J Mol Evol 87, 16–26 (2019). https://doi.org/10.1007/s00239-018-9881-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-018-9881-5