Abstract
Sulfur is an important key nutrient required for the growth and development of cyanobacteria. Several reports showed the effect of sulfate limitation in unicellular and filamentous cyanobacteria, but such studies have not yet been reported in heterocytous cyanobacteria to ascribe the mechanisms of nitrogen and thiol metabolisms. Thus, the present work was carried out to appraise the impacts of sulfate limitation on nitrogen and thiol metabolisms in Anabaena sp. PCC 7120 by analyzing the contents as well as enzymes of nitrogen and thiol metabolisms. Cells of Anabaena sp. PCC 7120 were exposed to different regimes of sulfate, i.e., 300, 30, 3, and 0 µM. Application of reduced concentration of sulfate showed negative impact on the cyanobacterium. Sulfate-limiting conditions reduces nitrogen-containing compounds in the cells of Anabaena. Additionally, reduced activities of nitrogen metabolic enzymes represented the role of sulfate in nitrogen metabolism. However, decreased activities of thiol metabolic enzymes indicated that sulfate-limited cyanobacterial cells have lower amount of glutathione and total thiol contents. Reduced accumulation of thiol components in the stressed cells indicated that sulfate-limited cells have lower ability to withstand stressful condition. Hence, Anabaena displays differential response to different concentrations of sulfate, and thus, stipulated that sulfur plays an important role in nitrogen and thiol metabolisms. To the best of our knowledge, this is the first report demonstrating the impact of sulfate stress on nitrogen and redox metabolisms in heterocytous cyanobacteria. This preliminary study provides a baseline idea that may help improve the production of paddy.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
According to the FAO (Food and Agriculture Organization) 2020 report, more than 50% of people worldwide eat rice as their primary food [1]. The area of paddy fields accounts for the major cultivated area of kharif grain crops in India and ranks second in the area of rice fields planted in the world (https://worldpopulationreview.com/country-rankings/rice-production-by-country). Planting rice is a common and effective strategy to meet the demands of the growing population. Sulfur is one of the essential nutrients for the growth of the plants, it enters the paddy ecosystem through the application of sulfur fertilizers and gaseous sulfur dioxide.
In the current scenario, decrease in the emission of sulfur dioxide gas escalated the sulfur-limiting environment and posed adverse effects on agriculture as well as agriculturally important microbes [2]. Since sulfur is also an essential nutrient required for the growth and development of cyanobacteria [2]. Sulfur is needed for the proper functioning of several metabolic processes such as photosynthesis, respiration, nitrogen metabolism, redox metabolism, and many more [3, 4]. The decrease in sulfur in the paddy fields disturbs the physiology and metabolic processes of the plant, resulting in less productivity. In order to obtain a good and productive yield, the application of sulfur in the paddy fields is quite significant.
Cyanobacteria are used as bio-inoculants in the paddy fields. They have the potential to fix atmospheric nitrogen into ammonia and other related nitrogenous compounds [5]. Most of the studies focused on the fact that sulfur limitation severely affects the morphology, physiology, and metabolic attributes of rice and other plants, which results in reduced yields [6, 7]. In the paddy field, cyanobacterial growth could also face sulfate stress condition, which might affect nitrogen-fixing capability, thereby reducing agricultural productivity [3, 5,6,7]. Therefore, an understanding of the response of agriculturally important nitrogen-fixing cyanobacteria to sulfate stress condition is particularly relevant for improving soil fertility and paddy field crop production.
The growth of cyanobacteria is hampered by sulfate limitation and produces reactive oxygen species (ROS) that lead to electrolyte leakage and membrane damage resulting programmed cell death [3]. Previous studies have reported the responses of cyanobacteria under sulfate stress condition which are restricted to very few unicellular and filamentous strains [3,4,5,6, 8,9,10]. Sulfate stress causes pigment reduction and lowers photosynthetic processes in Anabaena sp. PCC 7120 (hereafter Anabaena 7120) [3]. Recently, Kharwar and Mishra reported carbon allocation and reduced nitrogen content in the cyanobacterial cells of Anabaena under sulfate stress conditions [3].
Previously, it was reported that cells of Nostoc experiences sulfate stress showed decreased activity of nitrogen metabolic enzymes such as nitrogenase [11, 12]. On the other hand, the effect of sulfate stress on nitrate and nitrite metabolism has been relatively scant. Besides, only meagre information is available on the effect of sulfur deficiency on the thiol metabolism. So far, there have been very few studies in connection with the relationship of sulfate to thiol redox metabolism in cyanobacteria. Hence, the present study was performed to assess the impacts of sulfate limitation on nitrogen and redox metabolisms in the heterocytous cyanobacterium Anabaena 7120. For the first time, we have studied the effect of sulfate stress on the nitrogen and redox metabolisms of the heterocytous cyanobacterium Anabaena 7120. The data will explain how heterocytous cyanobacteria can respond to changing concentrations of sulfate under the stressed conditions. The study also provides information that the sulfur amendment in the rice field should be taken into consideration while increasing productivity, flourishing the growth of cyanobacteria, and further suggesting ways to help in sustain the paddy yield and their productivity.
Materials and Methods
Cyanobacterium Culture Conditions and Experimental Design
Cyanobacterium Anabaena 7120 was a generous gift from Prof. C.P. Wolk (Michigan State University, USA). The cyanobacterium was grown and maintained in the 250 mL Erlenmeyer flask containing 100 mL of BG-11 (pH 7.4) liquid medium at 28°C ± 2°C under continuous illumination of 50–55 μM photons m−2 s−1 with a 14/10h light/dark cycle. The cyanobacterium was manually shaken twice a day.
Magnesium sulfate (MgSO4) salt as a source of sulfate was used in the present study. Various concentrations of MgSO4 were prepared from the stock solution (1 M MgSO4) by dissolving it into the BG-11 medium. The stock solution was prepared in deionized Milli-Q water. On the basis of cell density experiment performed in our previous study [3], the effective concentrations of 300, 30, 3, and 0 µM MgSO4 were selected for the present study. For each experiment, the stock solution of MgSO4 was freshly prepared and high-pressure steam sterilization was done by autoclave. In an attempt to study sulfate stress responses in the cyanobacterium, exponentially grown cultures of Anabaena 7120 were harvested by centrifugation at 10,000 × g for 10 min. Pellets were washed with sterile sulfate-free medium to remove adherent salts. Further, the obtained pellets were dissolved in Milli-Q water and inoculated in the medium containing different sulfate concentrations such as 300, 30, 3, and 0 µM. The experiment was conducted in Erlenmeyer flasks of 250 mL filled with 100 mL of BG-11 culture medium. All the experiments were performed in triplicate. Table 1 describes all the assays performed in the present study and their importance. A comparative account was made between the stressed cyanobacterial cells and control (300 µM sulfate, i.e., normal BG-11 medium) cells with respect to nitrogen-containing compounds and nitrogen metabolic enzymes, components of thiol metabolism (total thiol, oxidized and reduced glutathione), and thiol metabolic enzymes.
Assay of Nitrogen-Containing Compounds and Nitrogen Metabolic Enzymes
Cyanobacterial cells exposed to different concentrations of sulfate were homogenized in the phosphate buffer (pH 7) and centrifuged at 10,000 × g at 25°C for 15 min. The collected supernatants were further used for quantification of nitrate (NO3−) and nitrite (NO2−) as per the protocols of Patterson et al. [13] and Ogawa et al. [14], respectively, and expressed in nmoL mg protein−1. Additionally, intracellular ammonium content was estimated using protocol described by Molins-Legua et al. [15] with the addition of Nessler’s reagent. The absorbance was recorded at 420 nm and expressed in nmoL mg protein−1.
Nitrate and nitrite reductase activities were assayed in cyanobacterial cells treated with different levels of sulfate using modified protocols of Herrero et al. [16] and Herrero and Guerrero [17], respectively. Enzyme activities were expressed in the activity units (U) corresponding to µmoL NO3− removed and/or NO3− produced min−1. While glutamine synthetase (transferase) activity (GS) was assayed as per the protocol of Shapiro and Stadtman [18] and expressed in terms of μmoL γ-glutamyl hydroxamate mg−1 protein min−1.
Assay of Components of Thiol Metabolism
Quantification of total thiol contents has been done according to the protocol of Sedlak and Lindsay [19]. Whereas oxidized and reduced glutathione contents were estimated using the method of Anderson et al. [20]. Oxidized glutathione was calculated by subtracting total glutathione from the reduced glutathione and expressed as µmoL protein−1.
Assay of Thiol Metabolic Enzymes
Cyanobacterial cells were centrifuged (10,000 × g) for 30 min at 4°C, and the obtained pellets were homogenized in 1 mL of phosphate buffer (pH 7.5). Further, homogenates were sonicated for 1 min and recentrifuged at 10,000 × g for 30 min at 4°C. The collected supernatants were further used for enzymatic assays.
SAT activity was estimated by the calorimetric method as given by Nakamura et al. [21]. Whereas OAS-TL activity was determined by measuring the production of cysteine at 560 nm as per the protocol of Kharwar and Mishra [3], and activity was represented in nmoL min−1 mg protein−1 using the standard curve of cysteine. In addition, γ-GCS activity was assayed using the protocol of Orlowki and Meister [22] and expressed in U mg protein−1. Moreover, glutathione reductase (GR) was performed using the modified method of Schaedle and Bassaham [23]. An assay of glutathione peroxidase (GPX) was carried out according to Lawrence and Burke [24] and expressed in terms of U mg protein−1.
Statistical Analysis
At first, all the parameters were checked for analysis of variance, i.e., the normal distribution of residuals, by the Shapiro–Wilk test and the Kolmogorov–Smirnov test, and were found to be satisfactory. Further, one-way ANOVA (analysis of variance) was performed to assess the significant differences between each treatment. The degree of correlation among all the parameters was also analyzed. A matrix dendrogram utilizing the unweighted pair group method with arithmetic mean (UPGMA) algorithm along with Euclidian similarity indices was constructed using PAST 3.0 software for cladistic representation of parameters. Moreover, principle component analysis (PCA) was performed to analyze the correlation between all the studied parameters. All the data analyses were carried out by Sigma 14 and SPSS software (SPSS Inc. Version 21.0, IBM Crop, Armonk, NY) and are represented as mean ± standard error.
Results
Nitrogen-Containing Compounds and Nitrogen Metabolic Enzymes
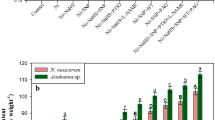
In order to understand the effect of sulfate stress in the heterocytous cyanobacteria on the nitrogen metabolic pathway, we have measured nitrogen-containing compounds and nitrogen metabolic enzymes. The influence of selected different concentrations of sulfate on nitrogen-containing compounds and nitrogen metabolic enzymes is shown in Fig. 1. Supplementation of 30, 3, and 0 µM sulfate substantially affect nitrogen metabolism by decreasing NO3−, NO2−, and NH4+ contents with respect to their control (300 µM sulfate) cells (Fig. 1a). Similarly, limited supplies of sulfate causes significant decrease in the enzyme activities of NR (4.8, 65.2, and 76.8%), NiR (16.2, 47.8, and 57.5%), and GS (7.4, 26.2, and 52%) at 30, 3, and 0 µM sulfate treated cells, respectively, compared to the control, i.e., 300 µM sulfate (Fig. 1b). Reduction in the nitrogen-containing compounds and nitrogen metabolic enzymes suggested that cyanobacterium was difficult to perform nitrogen assimilation efficiently in the limiting condition of sulfate.
a Nitrogen containing components; b Activities of nitrate reductase, nitrite reductase, and glutamine synthetase of the cyanobacterium Anabaena 7120 cells exposed to different concentrations of sulfate. 300 µM sulfate concentration indicated control condition, whereas 30, 3, and 0 µM sulfate concentrations represented as different treatments of sulfate in the present study. Each value is mean of three independent replicates, with bars indicating SE. Bars with different letters indicate that differences were statistically significant at P < 0.05
Thiols and Thiol Metabolic Enzymes Confer Redox Status
Cyanobacteria withstand different types of stressors, including light, nutrients, and oxidative stress. Their regulation of thiol metabolic process is a characteristic of the ability to tolerate such stressful conditions. However, there is few experimental reports of the thiol metabolic components conferring redox status identity in cyanobacteria, and none of the studies were performed in heterocytous cyanobacteria under sulfate stress conditions.
In order to fill the lacuna, we study the thiol metabolic components and enzymes in the heterocytous cyanobacterium. Figure 2a shows that high thiol content was achieved by the cyanobacterial cells at the 300 µM sulfate concentration. The results pertaining to total thiol contents depicted a significant reduction at 30, 3, and 0 µM sulfate concentrations. The lowest (15.8%) and highest (35%) thiol contents were noticed at 0 and 30 µM sulfate, respectively, compared to control (300 µM sulfate) cells (Fig. 2a). Additionally, reduction in the reduced glutathione (GSH) content, i.e., 1.8, 1.7, and 1.5 µM mg protein−1 in the 30, 3, and 0 µM of sulfate treated cells compared to the control (300 µM sulfate) cells, i.e., 2.9 µM mg protein−1 were detected, while increased oxidized glutathione (GSSG) content was noticed upon treatment of cyanobacterial cells with 30, 3, and 0 µM of sulfate as compared to the control (300 µM sulfate) cyanobacterial cells of Anabaena 7120 (Fig. 2b). Thus, the inferred percent increment in the degree of oxidation was 8.7, 9.04, and 10% at 30, 3, and 0 µM of sulfate treatments, respectively, as compared to control (300 µM sulfate) cyanobacterial cells of Anabaena 7120 (Fig. 2c). Changing the concentrations of sulfate from 300 to 0 µM affects the accumulation of total thiol contents and glutathione content. Increasing the GSSG of the cyanobacterial cells treated with 30, 3, and 0 µM sulfate led to the oxidation of glutathione.
a Thiol content; b Oxidized and reduced content of glutathione; c Degree of oxidation of glutathione of the cyanobacterium Anabaena 7120 cells exposed to the different concentrations of sulfate. 300 µM sulfate concentration indicated control condition, whereas 30, 3, and 0 µM sulfate concentrations represented as different treatments of sulfate in the present study. Each value is mean of three independent replicates, with bars indicating SE. Bars with different letters indicate that differences were statistically significant at P < 0.05
We first analyzed the enzymes involved in cysteine biosynthesis, such as SAT and OAS-TL. The negative effect of sulfate limitation was evident on the enzymes involved in cysteine biosynthesis. Percentage reduction in the activity of SAT was 7, 49, and 53.4%, while OAS-TL activity was reduced by 7.5, 11.3, and 14.5% at 30, 3, and 0 µM sulfate supplemented cells, respectively, as compared to the control, i.e., 300 µM sulfate (Fig. 3a and b). Hence, our results indicated that sulfate is important for maintaining cysteine biosynthesis in the test cyanobacterium.
Activities of thiol metabolic enzymes of the cyanobacterium Anabaena 7120 cells exposed to different concentrations of sulfate: a serine acetyl transferase; b O- acetyl serine thiol lyase; c γ-glutamyl cysteine synthetase; d glutathione reductase is represented with the bars and glutathione peroxidase is represented with the line in the diagram. 300 µM sulfate concentration indicated control condition, whereas 30, 3, and 0 µM sulfate concentrations represented as different treatments of sulfate in the present study. Each value is mean of three independent replicates, with bars indicating SE. Different letters indicate that differences were statistically significant at P < 0.05
Then, we analyzed the response of Anabaena 7120 on the thiol metabolic enzymes to different concentrations of sulfate. γ-GCS activity was detected in all the cells of cyanobacteria treated with different sulfate concentrations, and it was decreased during the stress condition. The activity of γ-GCS decreased by 8.5, 39.4, and 35.4% at 30, 3, and 0 µM sulfate treated cells, respectively, compared to control (300 µM sulfate) cells (Fig. 3c).
Next, the activity of GR in the cyanobacterium Anabaena 7120 under stress conditions was determined by harvesting the cyanobacterial cells from different concentrations of sulfate. As shown in Fig. 3d, there was a significant difference (P < 0.05) in the activity of GR for the tested concentrations of sulfate. Sulfate deficiency significantly reduced the activity of GR by 26.8, 91.5, and 87.1% in cells treated with 30, 3, and 0 µM sulfate, respectively, compared to control (300 µM sulfate) cyanobacterial cells (Fig. 3d).
Furthermore, the influence of sulfate stress on the activity of GPX in the test cyanobacterium was investigated. A reversed trend was observed for GPX activity, which was increased from 0.04 U mg protein−1 in control (300 µM sulfate) cells to 0.13, 0.25, and 0.37 U mg protein−1 in 30, 3, and 0 µM sulfate treatments, respectively (Fig. 3d). Therefore, we suggested that the different levels of GPX activity detected in each concentration of sulfate could be due to a differential expression of the gene encoding GPX in the cyanobacteria under sulfate stress conditions.
In summary, changing the concentration of sulfate led to changes in the activities of thiol metabolic enzymes, conferring redox status in the cells of the cyanobacterium (Fig. 3).
Statistical Analysis
A principal component analysis (PCA) was generated by SPSS software to interpret the correlation between the studied parameters under sulfate stress conditions. Results of PCA analysis for all the parameters used in the present study showed that PC1 and PC2 explained 86.258% and 9.033% of the variance, respectively, and thus cumulatively represent 95.291% of the total variance. For maximum variation in PC1, the parameters responsible were ammonia, NR, SAT, GS, nitrate, NiR, thiol, GCS, GR, nitrite, GSH, degree of oxidation (DO), and GPX (Fig. 4a, c), whereas PC2 identified GSH, OAS-TL, nitrite, GR, thiol, GCS, DO, and GSSG in the cyanobacterial cells as dominant variables (Fig. 4a, c). However, these components of the PCA plot are grouped into two clusters, i.e., minor cluster I and major cluster II, sharing an inverse relationship (Fig. 4a, c). Cluster I contained OAS-TL, GSH, and nitrite. Whereas thiol, GR, GCS, NiR, nitrate, SAT, ammonia, GS, and NR were subsumed into cluster II (the major cluster) and positioned closer to cluster I. However, GPX, DO, and GSSG form clusters, i.e., III, IV, and V, respectively, are distinct from both the cluster I and cluster II. PCA showed that all the studied parameters were differentially affected by sulfate, hence, they formed combined as well as independent clusters. Parameters that were similarly affected by sulfate were thiol, GR, GCS, NiR, nitrate, SAT, ammonia, GS, and NiR (cluster I). In the PCA, the studied parameters demonstrated segregation of the different sulfate concentrations with significant variability (Fig. 4a). A clear separation was observed in the parameters of clusters I and II from those clusters III, IV, and V. Cluster I and cluster II were presented in close proximity, showing their similar effects in response to different concentrations of sulfate. Additionally, cluster III, IV, and V were demarcated in close proximity, revealing that they respond to various sulfate concentrations with similar trends.
Statistical relationship among various parameters in control and sulfate limited treated cells of Anabaena 7120. a Biplot of principal component analysis of the parameters in control and sulfate limited cells. 300 µM sulfate concentration indicated control condition, whereas 30, 3, and 0 µM sulfate concentrations represented as different treatments of sulfate in the present study. The contribution to PC 1 is shown on the x-axis, while the contribution to PC 2 is on the y-axis; b Matrix dendrogram showing clustering of studied parameters into clades; c Contributions of the studied parameters to PCs. Parameters are shown on the y-axis, while the degrees of contribution to PCs are on the x-axis. (DO degree of oxidation; GPX glutathione peroxidase; GR glutathione reductase; GSH reduced glutathione; GSSG oxidized glutathione; NiR nitrite reductase; NR nitrate reductase; OAS-TL O-acetylserine (thiol) lyase; SAT serine acetyltransferase; γ-GCS γ-glutamyl cysteine synthetase)
Further, a correlation matrix for all the parameters defining the degree of correlation among various parameters is shown in Table 2. However, cladistic representation of all the data from treatments and control (300 µM sulfate), by UPGMA paired matrix dendrogram using Euclidian similarity indices was analyzed. All the parameters were clustered into two major clades, i.e., I and II. Clade I comprises of two minor subclades, i.e., subclade I and subclade II. The subclade I consisted of ammonia, NiR, SAT, GS, thiol, GCS, GSH, and DO, while subclade II consisted of OAS-TL, GPX, and GSSG. Moreover, parameters like NR, nitrate, nitrite, and GR were subsumed into clade II (Fig. 4b).
Discussion
Sulfate stress is very much expected, as it is actually occurring in the nature because of the decrease in sulfur dioxide gas emissions in the atmosphere. Although high concentrations of sulfate (approximately 28 mM) have been reported from marine ecosystems due to the presence of sulfate salts, while freshwater and agricultural soils contained lower amounts of sulfate [2]. In freshwater ecosystems and agricultural soils, sulfate concentrations range from 10 to 50 µM ensuring the occurrence of sulfur deficiency in common cyanobacterial habitats [2]. Since the cyanobacteria are used as biofertilizer in the paddy field [5,6,7]. It has been reported that the growth and productivity of cyanobacteria were adversely affected due to stress conditions. Thus, the cyanobacteria thriving in such niche might also face sulfate stress. Hence, it is interesting to understand the impact of sulfate stress on the paddy field cyanobacterium and how the cyanobacteria respond in case of sulfate stress. The components and enzyme activities of nitrogen and redox metabolisms of the cyanobacterium Anabaena 7120, were monitored in the current study under selected different concentrations of sulfate, namely 300, 30, 3, and 0 µM, in order to understand how the cyanobacterium responds to each concentration of sulfate and how the sulfate stress affects the paddy field production.
Regulation of the Nitrogen Metabolic Pathway Upon Sulfate Limitation in the cyanobacterium Anabaena 7120
Sulfate limitation alters the nitrate assimilation process of the cyanobacterium Anabaena 7120. Significant decrease in the nitrate and nitrite contents were noticed in the cyanobacterial cells supplemented with 30, 3, and 0 µM as compared to the control (300 µM sulfate) cells. Furthermore, NH4+ content was reduced by limiting sulfate supplies. The gradual decrease in the NH4+, NO2−, and NO3− contents in response to sulfate limitations suggested that reduced sulfate availability alters nitrate metabolism. Flores and Herrero demonstrated that the ABC-type transporter proteins, which likely use ATP as a source of energy, facilitate NO3− and NO2− uptake into the cyanobacterial cells [25, 26]. Reduction in the NO3− and NO2− uptake rates as the result of the disrupted photosynthetic electron transport chain perhaps associated with lower pool of ATP [27]. There was significant decrease in the NO3− and NO2− contents in sulfate-limiting cyanobacterial cells, possibly due to damage in the membrane bound transporter proteins [28], whereas cells supplemented with 300 µM sulfate showed higher enzyme activity, which was supported by increased photosynthetic activity and lower production of ROS. With decreased transportation of nutrients under different treatments of sulfate, i.e., 30, 3, and 0 µM, the MDA content was found to be increased in the cyanobacterial cells along with the high ROS production, which clearly indicated stressful condition in the treated (30, 3, and 0 µM sulfate) cyanobacterial cells of Anabaena 7120 that likely had not occurred in the control cells, i.e., 300 µM sulfate [3]. Moreover, significant decrease in the activities of NR and NiR were observed in the sulfate-limiting cells. Additionally, GS activity was also significantly reduced at 30, 3, and 0 µM sulfate treatments with respect to the control (300 µM sulfate) cells. Decrease in the NiR activity was associated with reduced availability of NO2− ions (as observed in our study). Furthermore, higher NR, NiR, and GS activities at 300 µM sulfate concentrations were connected with higher growth and better photosynthetic activity in terms of chlorophyll a, carotenoids, and phycobiliproteins. Since sulfate stress could strongly inhibit chlorophyll biosynthesis [3], one of the first visible consequences of stress conditions in cyanobacteria is a decrease in the photopigments. The chlorophyll content in the control cyanobacterial cells (300 µM sulfate) was relatively higher than that of cyanobacterial cells treated with 30, 3, and 0 µM sulfate concentrations. Since cyanobacteria require significant amount of sulfur, which plays various roles in the photosynthetic complex, it is present in the Fe-S cluster of cyanobacteria. Enhanced photosynthetic processes that generate ATP in Nostoc muscorum and Phormidium foveolarum have been reported by researchers [28, 29]. At lower concentrations of sulfate, i.e., 3 and 0 µM, an inhibition of growth and lower photosynthetic activities [3] might be the reason for the lesser activity of these enzymes. This decreased activity was probably caused by repression of the enzyme. Sulfate stress modulates NR and NiR activities through nitrate and nitrite uptake, respectively, since the enzyme activity is perhaps determined by nitrate and nitrite flux into the metabolic pool. Different authors reported that excess copper disturbs photosynthesis and redox equilibrium of the cells, disrupting the cells ultrastructure, and finally leading to cell death [30,31,32]. However, under high cadmium and zinc stress, activation of metallothionein (small cysteine-rich proteins) was noticed, which chelates these heavy metals and helps in the survival of cyanobacteria [33]. In accordance with our results, Singh et al. also showed that heavy metals, pesticides, and UV-B stress hampers photosynthetic process of the cyanobacteria, resulting decreased activities of NR and NiR [28, 29]. The GS enzyme plays a key role in nitrogen metabolism. One of the 20 amino acids that make up the standard genetic code, glutamine, is involved in number of biochemical processes, such as synthesis of protein, production of ammonium to control the acid–base balance, providing energy to cells in addition to glucose, donating nitrogen for various anabolic processes, and replenishing citric acid cycle. GS catalyzes the ATP-dependent condensation of NH4+ with glutamate to produce glutamine, making it a crucial enzyme in ammonia assimilation [28, 29]. Hence, reduction in the activities of GS in the cyanobacterial cells treated with 30, 3, and 0 µM sulfate concentrations was presumed with lesser heterocyte frequency as compared to control cells (300 µM sulfate), since GS is located in the heterocyte of the cyanobacterial cells. Another possible explanation for decrease in GS activity is that the C/N balance disrupting cyanobacterial growth. Thus, changes in the status of the above-mentioned parameters reduce the nitrate assimilation process as well as the nitrogen content of the cell. This decrease in nitrogen content indicated that sulfate limitation also inhibits nitrogen uptake by Anabaena 7120 [3]. In the previous study, authors reported a significant impact of salt stress on the cyanobacterial nitrogen metabolism [34].
Sulfate Limitation Leads to Changes in the Sulfur-Containing Metabolites and Enzymes of the Thiol Metabolic Pathway, Indicating an Alteration in the Thiol-Based Redox Buffer of the Cyanobacterium Anabaena 7120
Glutathione is a key regulator of the cellular thiol redox buffer which regulates redox homeostasis and the cell cycle [35, 36]. Therefore, the present study is designed to understand the connection between sulfur, ROS, glutathione formation, and redox equilibrium under different sulfate concentrations in Anabaena 7120. The results showed that the pool of reduced GSH was lower in the sulfate treatment cells as compared to the control, which coincides with the lower activity of γ-GCS, whereas GSSG showed varied response. This results in a smaller GSH/GSSG redox couple in the sulfate stressed cells which favors an oxidize environment. The precise control of intracellular redox status, i.e., maintenance of physiological levels of ROS for mediating normal cellular function (oxidative eustress) while evading excess ROS (distress), is central player for the concept of redox system [36]. Intracellular redox changes affect cell signaling, gene transcription, translation, and cell death [37, 38]. A smaller value of the redox couple, i.e., GSH/GSSG, is reflected by rapid intracellular glutathione oxidation. The higher degree of oxidation in the stressed cells was determined by the increasing oxidized glutathione production. This indicated lower turnover of GSH/GSSG in the stressed cells relative to the control (300 µM sulfate) cells and changes in the glutathione redox potential. Thus, cyanobacterial cells at 30, 3, and 0 µM sulfate are experiences oxidative stress, reflecting that glutathione and sulfate are important for acclimating under the adverse conditions in cyanobacteria. Cameron and Pakrasi demonstrated an increase in GSSG level from the glutathione pool during sulfate starvation in Synechocystis sp. PCC 6803 [35]. Li et al. showed that glutathione is involved in the regulation of several processes through glutathionylation in cyanobacteria [39]. It has been demonstrated that glutathione and cysteine contents are reduced in Synechocystis 6803 during sulfate deficiency [40]. Another probable reason for reduced glutathione content in the stressed cyanobacterial cells was the reduction in NADPH content at 30, 3, and 0 µM sulfate concentrations, as observed in our previous study [3]. Adams et al. reported that the ratio of the reduced and total pyridine nucleotide pools is an index of cellular redox status [41]. NADPH is essential for recycling of glutathione and is related to its antioxidant functions [41]. Hence, there was disturbance in the redox homeostasis of the cyanobacterial cells treated with 30, 3, and 0 µM sulfate concentrations as compared to the control cells (300 µM sulfate). In addition, dramatic decline in the thiol contents was found in the cyanobacterial cells at 30, 3, and 0 µM sulfate as compared to the control (300 µM sulfate). This rapid decrement in the thiol contents also confers oxidative constraints in the cells.
In addition, the gradual reduction in SAT activities was noticed in the cyanobacterial cells at 30, 3, and 0 µM sulfate concentrations, whereas only meagerly significant alteration in OAS-TL enzyme activities was evident at 30, 3, and 0 µM sulfate supplementation as compared to the control (300 µM sulfate) condition, conferring modulations in OAS-TL activity. Sulfate limitations affect cysteine biosynthesis which results in reduced methionine and glutathione contents [40]. Concentration of these sulfur-containing metabolites decreases when sulfate limitations occur in the cells. This might have two reasons: first, upon sulfate limitations, sulfide rather than OAS becomes limiting for cysteine biosynthesis. Secondly, the cell might try to keep the thiol concentration at a certain level. Lower activities of SAT and OAS-TL were related to oxidative stress in sulfate-limiting conditions. OAS-TL and SAT scavenge ROS under stress condition, conferring oxidative stress tolerance [42]. A reduction in intracellular sulfate content is a typical consequence of sulfate deficiency [3]. Sulfate uptake and assimilation are functions of organisms’ demand for sulfur metabolites, viz. sulfide, cysteine, and glutathione, which are involved in arrays of signaling pathways and redox sensors [4].
GR serves as a marker for oxidative stress because it converts glutathione disulfide (GSSG) back to glutathione. Prior research revealed that cyanobacteria that can withstand any kind of stress typically exhibit higher GR activity [35]. Less significant reductions of γ-GCS and GR were observed in the sulfate stressed cells of Anabaena 7120. This reduction in the activities of γ-GCS and GR in the cyanobacterial cells treated with 30, 3, and 0 µM sulfate concentrations was thought to be due to decreased glutathione synthesis. In view of our results in Anabaena 7120, it could be suggested that the gene coding for GR (gor) is differentially transcribed as a function of the sulfur concentration, and the GR activity also depends on the sulfur concentration. γ-GCS is a rate-limiting enzyme in glutathione biosynthesis and plays a central role in glutathione homeostasis. The rapid changes in the glutathione and γ-GCS levels at the 30, 3, and 0 µM sulfate concentrations indicated that glutathione and γ-GCS were perhaps catabolized as a source of sulfur, which is limited during the sulfate-limiting condition in cyanobacteria. These decrease in γ-GCS levels in the stressed cyanobacterial cells represent decrease in the efficiency of cells to maintain the redox poise. Researchers have reported that reduction in glutathione and γ-GCS activity in yeast under nitrogen and sulfur stress conditions, which could be catabolized for amino acids [43, 44]. However, changes in the cellular metabolism were evident in plants during sulfur-limiting conditions [45]. GR was found to regenerate glutathione; thus, reduced levels of GR in the cyanobacterial cells at 30, 3, and 0 µM sulfate concentrations indicated decrease in the glutathione amount. It is one of the most abundant reducing thiol content which catalyzes reduction of glutathione disulfide (GSSG) to the reduced form of glutathione (GSH) by an electron donor, NADPH [46]. However, the lowered activities of these above-mentioned enzymes in the stressed cells of Anabaena 7120 might not necessarily scavenge ROS below their thresholds since the adverse effects of oxidative stress were exhibited by rapid lipid peroxidation in our previous study [3].
By contrast to GR, significant increment of GPX activities was evident in stressed cells of Anabaena 7120 at 30, 3, and 0 µM sulfate concentrations. It is an important enzyme that catalyzes the detoxification of hydrogen peroxide to water and GSSG by using GSH as a reductant. This antioxidative enzyme provides the most vital defense against peroxidative damage to membrane, which are reported to play an important and fundamental role in cellular homeostasis [47]. Thus, cyanobacterial cells at 30, 3, and 0 µM sulfate concentrations experience more oxidative damage. Overall, these results showed that the balance of the GSH/GSSG ratio and ROS homeostasis are regulated by GPX, which represents modulation in the oxidation–reduction system and culminates in altered growth and other processes. In our study, it was observed that the sulfate stressed cyanobacterial cells of Anabaena 7120 showed lesser growth as compared to the control (300 µM sulfate) cells. In general, light and nutrients stress exert comparable effects on cyanobacterial cells [35, 40, 48]. These stresses alter structural characteristics, cellular growth, and metabolism of cyanobacteria and activates the antioxidant system to detoxify ROS as well as regulate redox buffer of the cells [49], but in comparison to sulfate stress, cyanobacteria are fails to activate the antioxidant system due to the lower availability of sulfur for the synthesis of proteins involved in this mechanism. The breakdown of the antioxidant system is one of the critical aspects for the cyanobacteria to maintain their homeostasis under sulfate stress condition. The inability of cyanobacteria to endure disturbed homeostasis exemplifies their poor tolerance mechanisms under such condition. Since heterocytous nitrogen-fixing cyanobacteria are integral components of paddy fields and thus play a crucial role in improving the productivity of rice plants, it is very important that the paddy field was amended with an optimal amount of sulfur, which favors the growth of cyanobacteria and thereby enhances the yield and productivity of rice. The flourishing growth of cyanobacteria in the paddy field helps in bringing about changes in the fertility of the soil, its mineral composition, and many more factors that facilitates the productivity of rice plant [5]. Thus, our findings suggested that sulfate stress alters the nitrogen and redox processes of Anabaena. The present study also demonstrated that the optimal amount of sulfate should be taken into consideration when improving of rice production.
Conclusion
In conclusion, our results showed that sulfur plays a pivotal role in cyanobacterial metabolism (Fig. 5). Alterations in the activities of nitrogen metabolic enzymes as well as nitrogen-containing compounds were noticed, indicating that sulfur is an important nutrient regulating nitrogen metabolism in Anabaena 7120. Additionally, components of thiol and the activities of thiol metabolic enzymes in the cyanobacterium were also reduced, suggested the crucial role of sulfur in maintaining redox status of the cells. Overall, sulfate limitation alters cyanobacterial metabolism and therefore provides evidence for the role of sulfate in redox regulation. It also highlights the cross-talk among the thiol-based redox buffer and nitrogen metabolism of the cyanobacterium Anabaena 7120. This study provides a preliminary idea and expands our knowledge about how the cyanobacterium alters its metabolic pathways during sulfate stress, which subsequently affects the productivity of rice fields. This type of analysis gives an indirect assessment and a first-step study on the impacts of sulfate stress condition in cyanobacteria, which accelerate the growth of agronomically important plants. The task remains to determine the impacts of sulfate stress on the cyanobacterial cells grown in the paddy field in a natural environment.
Data Availability
The datasets of the present study are available from the authors.
Code Availability
Not applicable.
Abbreviations
- APR:
-
APS-reductase
- APS:
-
Adenosine-5’-phopsphosulfate
- ATPS:
-
ATP sulfurylase
- DO:
-
Degree of oxidation
- FAO:
-
Food and agriculture organization
- GPX:
-
Glutathione peroxidase
- GR:
-
Glutathione reductase
- GSH:
-
Reduced glutathione
- GSSG:
-
Oxidized glutathione
- NiR:
-
Nitrite reductase
- NR:
-
Nitrate reductase
- OAS-TL:
-
O-acetyl serine (thiol) lyase
- SAT:
-
Serine acetyltransferase
- γ-GCS:
-
γ-Glutamyl cysteine synthetase
References
FAO (2020) Statistics database. Accessed 10 Feb 2020 http://faostat.fao.org.
Kharwar S, Bhattacharjee S, Chakraborty S, Mishra AK (2021) Regulation of sulfur metabolism, homeostasis and adaptive responses to sulfur limitation in cyanobacteria. Biologia 76(10):2811–2835. https://doi.org/10.1007/s11756-021-00819-5
Kharwar S, Mishra AK (2020) Unravelling the complexities underlying sulfur deficiency and starvation in the cyanobacterium Anabaena sp. PCC 7120. Environ Exp Bot 172:103966. https://doi.org/10.1016/j.envexpbot.2019.103966
Kharwar S, Bhattacharjee S, Mishra AK (2021) Bioinformatics analysis of enzymes involved in cysteine biosynthesis: first evidence for the formation of cysteine synthase complex in cyanobacteria. 3 Biotech. https://doi.org/10.1007/s13205-021-02899-1
Prasad RC, Prasad BN (2001) Cyanobacteria as a source biofertilizer for sustainable agriculture in Nepal. J Plant Sci Bot Orient 1:127–133. https://doi.org/10.1016/j.bbrep.2020.100737
Lunde C, Zygadlo A, Simonsen HT, Nielsen PL, Blennow A, Haldrup A (2008) Sulfur starvation in rice: the effect on photosynthesis, carbohydrate metabolism, and oxidative stress protective pathways. Physiol Plant 134(3):508–521. https://doi.org/10.1111/j.1399-3054.2008.01159.x
Resurreccion AP, Makino A, Bennett J, Mae T (2002) Effect of light intensity on the growth and photosynthesis of rice under different sulfur concentrations. Soil Sci Plant Nutr 48:71–77. https://doi.org/10.1080/00380768.2002.10409173
Kolesinski P, Rydzy M, Szczepaniak A (2017) Is RAF1 protein from Synechocystis sp. PCC 6803 really needed in the cyanobacterial Rubisco assembly process. Photosynth Res 132(2):135–148. https://doi.org/10.1007/s11120-017-0336-4
Kumaresan V, Nizam F, Ravichandran G, Viswanathan K, Palanisamy R, Bhatt P, Arasu MV, Al-Dhabi NA, Mala K, Arockiaraj J (2017) Transcriptome changes of blue-green algae, Arthrospira sp. in response to sulfate stress. Algal Res 23:96–103. https://doi.org/10.1016/j.algal.2017.01.012
Green LS, Grossman AR (1988) Changes in sulfate transport characteristics and protein composition of Anacystis nidulans R2 during sulfur deprivation. J Bacteriol 170(2):583–587. https://doi.org/10.1128/jb.170.2.583-587
Hifney AF, Abdel-Basset R (2014) Induction of hydrogen evolution in Nostoc spp. by nitrogen and/or sulphur deficiency. Int J Curr Microbiol App Sci 3(9):1028–1044
Ortega-Calvo J, Stal LJ (1994) Sulphate-limited growth in the N2-fixing unicellular cyanobacterium Gloeothece (Nägeli) sp. PCC 6909. New Phytol 128(2):273–281. https://doi.org/10.1111/j.1469-8137.1994.tb04011.x
Patterson K, Cakmak T, Cooper A, Lager IDA, Rasmusson AG, Escobar MA (2010) Distinct signalling pathways and transcriptome response signatures differentiate ammonium-and nitrate-supplied plants. Plant Cell Environ 33(9):1486–1501. https://doi.org/10.1111/j.1365-3040.2010.02158.x
Ogawa K, Shiraishi N, Ida S, Nakagawa H, Komamine A (1999) Effects of glutamine on the induction of nitrate reductase and nitrite reductase in cultured spinach cells. J Plant Physiol 154(1):46–50. https://doi.org/10.1016/S0176-1617(99)80316-2
Molins-Legua C, Meseguer-Lloret S, Moliner-Martinez Y, Campíns-Falcó P (2006) A guide for selecting the most appropriate method for ammonium determination in water analysis. Trends Anal Chem 25(3):282–290. https://doi.org/10.1016/j.trac.2005.12.002
Herrero AN, Flores EN, Guerrero MG (1981) Regulation of nitrate reductase levels in the cyanobacteria Anacystis nidulans, Anabaena sp. strain 7119, and Nostoc sp. strain 6719. Bacteriol 145(1):175–180. https://doi.org/10.1128/jb.145.1.175-180.1981
Herrero A, Guerrero MG (1986) Regulation of nitrite reductase in the cyanobacterium Anacystis nidulans. Microbiol 132(9):2463–2468. https://doi.org/10.1128/jb.145.1.175-180.1981
Shapiro BM, Stadtman ER (1970) [130] Glutamine synthetase (Escherichia coli). In Methods in enzymology, Academic Press, 910–922 https://doi.org/10.1016/0076-6879(71)17305-3
Sedlak J, Lindsay RH (1968) Estimation of total, protein-bound, and nonprotein sulfhydryl groups in tissue with Ellman’s reagent. Anal Biochem 25:192–205. https://doi.org/10.1016/0003-2697(68)90092-4
Anderson ME (1985) Determination of glutathione and glutathione disulfide in biological samples. In Methods in enzymology. Academic Press, 113:548-555 https://doi.org/10.1016/s0076-6879(85)13073-9
Nakamura K, Hayama A, Masada M, Fukushima K, Tamura G (1987) Measurement of serine acetyltransferase activity in crude plant extracts by a coupled assay system using cysteine synthase. Plant Cell Physio 28(5):885–891. https://doi.org/10.1093/oxfordjournals.pcp.a077370
Orlowski M, Meister A (1971) Partial reactions catalyzed by γ-glutamylcysteine synthetase and evidence for an activated glutamate intermediate. J Biol Chem 246(23):7095–7105. https://doi.org/10.1016/S0021-9258(19)45858-4
Schaedle M, Bassham JA (1977) Chloroplast glutathione reductase. Plant Physio 59(5):1011–1012. https://doi.org/10.1104/pp.59.5.1011
Lawrence RA, Burk RF (1976) Glutathione peroxidase activity in selenium-deficient rat liver. Biochem Biophys Res Commun 71(4):952–958. https://doi.org/10.1016/0006-291x(76)90747-6
Flores E, Herrero A (1994) Assimilatory nitrogen metabolism and its regulation. In The molecular biology of cyanobacteria Springer, Dordrecht, 487–517 https://doi.org/10.1007/978-94-011-0227-8_16
Rai LC, Tyagi B, Mallick N, Rai PK (1995) Interactive effects of UV-B and copper on photosynthetic activity of the cyanobacterium Anabaena doliolum. Environ Exp Bot 35(2):177–185. https://doi.org/10.1016/0098-8472(94)00046-8
Ohashi Y, Shi W, Takatani N, Aichi M, Maeda SI, Watanabe S, Yoshikawa H, Omata T (2011) Regulation of nitrate assimilation in cyanobacteria. J Exp Bot 62(4):1411–1424. https://doi.org/10.1093/jxb/erq427
Singh VP, Srivastava PK, Prasad SM (2012) UV-B induced differential effect on growth and nitrogen metabolism in two cyanobacteria under copper toxicity. Cell Mol Biol 58(1):85–95. https://doi.org/10.1170/T925
Sheeba RK, Prasad SM (2020) Nostoc muscorum and Phormidium foveolarum differentially respond to butachlor and UV-B stress. Physio Mol Biol Plants 26(4):841–856. https://doi.org/10.1007/s12298-019-00754-5
Nagalakshmi N, Prasad MNV (2001) Responses of glutathione cycle enzymes and glutathione metabolism to copper stress in Scenedesmus bijugatus. Plant Sci 160:291–299. https://doi.org/10.1016/s0168-9452(00)00392-7
Yadav P, Singh RP, Gupta RK (2022) Role of cyanobacteria in germination and growth of paddy seedlings. Int J Phytol Res 2:11–18
Wang Z, Li D, Li G, Liu Y (2010) Mechanism of photosynthetic response in Microcystis aeruginosa PCC7806 to low inorganic phosphorus. Harmful Algae 9:613–619. https://doi.org/10.1016/j.hal.2010.04.012
Blindauer CA (2011) Bacterial metallothioneins: past, present, and questions for the future. J Biol Inorg Chem 16:1011–1024. https://doi.org/10.1007/s00775-011-0790-y
Apte SK, Reddy BR, Thomas J (1987) Relationship between sodium influx and salt tolerance of nitrogen-fixing cyanobacteria. Appl Environ Microbiol 53:1934–1939. https://doi.org/10.1128/aem.53.8.1934-1939.1987
Cameron JC, Pakrasi HB (2010) Essential role of glutathione in acclimation to environmental and redox perturbations in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol 154(4):1672–1685. https://doi.org/10.1104/pp.110.162990
Schafer FQ, Buettner GR (2001) Redox environment of the cell as viewed through the redox state of the glutathione disulfide/glutathione couple. Free Radic Biol Med 30(11):1191–1212. https://doi.org/10.1016/s0891-5849(01)00480-4
Arrigo AP (1999) Gene expression and the thiol redox state. Free Radic Biol Med 27(9–10):936–944. https://doi.org/10.1016/s0891-5849(99)00175-6
Voehringer DW (1999) BCL-2 and glutathione: alterations in cellular redox state that regulate apoptosis sensitivity. Free Radic Biol Med 27(9–10):945–950. https://doi.org/10.1016/s0891-5849(99)00174-4
Li M, Yang Q, Zhang L, Li H, Cui Y, Wu Q (2007) Identification of novel targets of cyanobacterial glutaredoxin. Arch Biochem Biophys 458:220–228. https://doi.org/10.1016/j.abb.2006.12.010
Zechmann B, Tomašić A, Horvat L, Fulgosi H (2010) Subcellular distribution of glutathione and cysteine in cyanobacteria. Protoplasma 246(1):65–72. https://doi.org/10.1007/s00709-010-0126-8
Adams JD Jr, Klaidman LK, Chang ML, Yang J (2001) Brain oxidative stress—analytical chemistry and thermodynamics of glutathione and NADPH. Curr Top Med Chem 1:473–482. https://doi.org/10.2174/1568026013394778
Noji M, Saito K (2002) Molecular and biochemical analysis of serine acetyltransferase and cysteine synthase towards sulfur metabolic engineering in plants. Amino Acids 22(3):231–243. https://doi.org/10.1007/s007260200011
Elskens MT, Jaspers CJ, Penninckx MJ (1991) Glutathione as an endogenous sulphur source in the yeast Saccharomyces cerevisiae. J Gen Microbiol 137:637–644. https://doi.org/10.1099/00221287-137-3-637
Mehdi K, Penninckx MJ (1997) An important role for glutathione and gamma-glutamyltranspeptidase in the supply of growth requirements during nitrogen starvation of the yeast Saccharomyces cerevisiae. Microbiol 143:1885–1889. https://doi.org/10.1099/00221287-143-6-1885
Nikiforova VJ, Kopka J, Tolstikov V, Fiehn O, Hopkins L, Hawkesford MJ, Hesse H, Hoefgen R (2005) Systems rebalancing of metabolism in response to sulfur deprivation, as revealed by metabolome analysis of Arabidopsis plants. Plant Physiol 138:304–318. https://doi.org/10.1104/pp.104.053793
Couto N, Wood J, Barber J (2016) The role of glutathione reductase and related enzymes on cellular redox homoeostasis network. Free Radic Biol Med 95:27–42. https://doi.org/10.1016/j.freeradbiomed.2016.02.028
Zhang J, Hao H, Wu X, Wang Q, Chen M, Feng Z, Chen H (2020) The functions of glutathione peroxidase in ROS homeostasis and fruiting body development in Hypsizygus marmoreus. Appl Microbiol Biotech 104(24):10555–10570. https://doi.org/10.1007/s00253-020-10981-6
Donker VA, Hader DP (1997) Ultraviolet radiation effects on pigmentation in the cyanobacterium Phormidium uncinatum. Acta Protozool 36:49–55
Yadav P, Singh RP, Rana S, Joshi D, Kumar D, Bhardwaj N, Gupta RK, Kumar A (2022) Mechanisms of stress tolerance in cyanobacteria under extreme conditions. Stresses 2(4):531–549. https://doi.org/10.3390/stresses2040036
Acknowledgements
Authors are thankful to the Head, Department of Botany, Banaras Hindu University for the necessary lab and instrument facilities. Surbhi Kharwar is thankful to University Grant Commission, New Delhi, for financial support in the form of Junior and Senior research fellowships.
Funding
This work was supported by the University Grant Commission, New Delhi, and Institute of Eminence (IoE-6031), Banaras Hindu University, Varanasi.
Author information
Authors and Affiliations
Contributions
SK: performed the experiments, statistical analysis, and wrote the manuscript. AKM: reviewed the manuscript. The authors read, reviewed, and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Authors declared they have no conflict of interest.
Ethical Approval
Not applicable.
Consent to Participations
Not applicable.
Consent for Publications
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Kharwar, S., Mishra, A.K. Nitrogen and Redox Metabolism in Cyanobacterium Anabaena sp. PCC 7120 Exposed to Different Sulfate Regimes. Curr Microbiol 80, 265 (2023). https://doi.org/10.1007/s00284-023-03374-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-023-03374-1