Abstract
The biosynthesis of cysteine is crucial and critically regulated by two enzymes. i.e., serine acetyl transferase (SAT) and O-acetyl serine (thiol) lyase (OAS-TL). A descriptive account on the activity and regulatory mechanism of the enzyme is available in bacteria and plants. But no such studies yet performed in cyanobacteria, to understand the evolutionary aspect of cysteine biosynthesis and its regulation. Therefore, in our study a detailed bioinformatic analysis has been performed to understand all the possible features of cyanobacterial SATs and OAS-TLs. The analysis of SAT and OAS-TL sequences from cyanobacteria depicted that the large genome and morphological complexities favoured acquisition of these genes. Besides, conserved function of these enzymes was presumed by their sequence similarity. Further, the phylogenetic tree consisted of distinct clusters for unicellular, filamentous, and heterocytous strains. Nevertheless, the specificity pocket, SVKDR for OAS-TL having K as catalytic residue was also identified. Additionally, in silico protein modelling of SAT (SrpG) and OAS-TL (SrpH) of Synechococcus elongatus PCC 7942 was performed to gain insight into the structural attributes of the proteins. Finally, here we showed the possibility of hetero-oligomeric bi-enzyme cysteine synthase complex formation upon interaction of SAT and OAS-TL through protein–protein docking analysis thus provides a way to understand the regulation of cysteine biosynthesis in cyanobacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
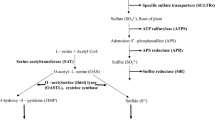
The significance of cysteine as an efficient redox buffer, sulphur assimilator, signalling component, antioxidant, etc. is univocal in all the domains of life (Buchanan and Balmer 2005; Diessner and Schmidt 1980; Foyer and Noctor 2011; Joshi et al. 2019; Kharwar and Mishra 2020; Noctor et al. 2012; Richau et al. 2012; Takahashi et al. 2011). However, cysteine can be toxic above a threshold concentration; thus its homeostasis is maintained precisely inside the cell. The biosynthesis of cysteine commences by enzymes serine acetyl transferase (SAT; EC.2.3.1.30) and cysteine synthase (CS) or O-acetyl serine (thiol) lyase (OAS-TL; EC.4.2.99.8) which catalyses the final step of sulphur assimilation. SAT catalyses rate-limiting step where the acetyl group of acetyl-CoA is transferred to the hydroxyl group of serine to form OAS, followed by β-replacement of acetyl group by OAS-TL (Kopriva and Koprivova 2003), in presence of pyridoxal-5′-phosphate (PLP) as the cofactor:
In plants, SAT and OAS-TL interact to form cysteine synthase complex (CSC), a functional hetero-oligomeric bi-enzyme complex (Bogdanova and Hell 1997; Berkowitz et al. 2002; Droux et al. 1998a, b; Jost et al. 2000; Ruffet et al. 1994; Wirtz et al. 2001). Formation of such hetero-oligomeric bi-enzyme complex strongly depends on the concentrations of two substrates, i.e., O-acetyl-serine (OAS) and sulfide (Buchner et al. 2004; Hopkins et al. 2005; Khan et al. 2010). Studies suggested that sulfide takes part in sulphur homeostasis and serve as a sensor for intracellular sulphur concentration (Yi et al. 2010). Sulphide stabilizes the complex, whereas OAS destabilizes it (Berkowitz et al. 2002; Droux et al. 1998a, b; Kredich 1996; Kredich and Tomkins 1966; Kredich et al. 1969; Wirtz and Hell 2006). Under sulphur limiting condition, OAS accumulates and dissociates bi-enzyme complex which down-regulates the SAT activity, in contrast upon sufficient sulphur supplementation, high sulfide level in the cell enhances binding of OAS-TL to SAT, stabilising hetero-dimeric complex, thus, representing the prime control for the regulation of cysteine biosynthesis (Droux et al. 1998a, b; Kredich and Tomkins 1966; Wirtz and Hell 2006). Moreover, different regulatory mechanisms have been reported in plants such as transcriptional, post-transcriptional, protein–protein interaction, and feed-back control mechanism (Davidian and Kopriva 2010; Yi et al. 2010). Additionally, in bacteria, formation and regulation of CSC by protein–protein interaction and feedback regulation has also been observed (Kredich and Tomkins 1966). Similar to plant and bacteria, in cyanobacteria cysteine synthesis is also catalysed by SAT and OAS-TL, despite the formation of such bi-enzyme complex has not been yet reported. Therefore, in the present study we have tried to elucidate the distribution, abundance, domain configuration, physicochemical properties, and phylogeny of cyanobacterial SAT and OAS-TL proteins using iterative bioinformatics approaches. Besides, the structural attributes, fold, and topology of SAT and OAS-TL were determined by theoretical modelling of SrpG and SrpH from Synechococcus elongatus PCC 7942, respectively. Further, the possible interactions of SrpG and SrpH with other proteins were unravelled by STRING server. Finally, the molecular docking of SrpG and SrpH corroborated the formation of putative heteromeric bi-enzyme cysteine synthase complex in cyanobacteria Thus, this work may provide a comprehensive and updated view of cysteine biosynthesis in cyanobacteria and a broader dimension of its regulation through putative CSC formation.
Methods
Retrieval of serine acetyl transferase (SAT) and O-acetyl serine (thiol) lyase (OAS-TL) sequences
The sequences of cyanobacterial SAT and OAS-TL were retrieved in the FASTA format from non-redundant database of NCBI (http://www.ncbi.nlm.nih.gov/). Sequences with higher identity values were selected and manually curated. Finally, 63 SAT and 80 OAS-TL amino acid sequences from 51 and 67 cyanobacterial strains, respectively, were selected for further studies. The distribution and abundance of the SAT and OAS-TL proteins among different groups of cyanobacteria were analysed.
Multiple sequence alignment and phylogenetic analyses
Multiple sequence alignment (MSA) was performed using MUSCLE in Molecular Evolutionary Genetics Analysis v.10 (MEGA X) software with default parameters, i.e., gap opening penalties − 2.90, gap extension penalties 0.00; hydrophobicity multiplier, i.e., 1.2. Subsequently, the conserved residues among these sequences were visualised as Hidden Markov Model (HMM) logo using Skylign (http://skylign.org/) (Wheeler et al. 2014).
Phylogenetic trees for SAT and OAS-TL were constructed by neighbour-joining (NJ) method (Kumar et al. 2018; Saitou and Nei 1987) using MEGA v10. The align sequences of the protein were further used to construct the phylogenetic trees. JTT matrix-based method was used to compute the evolutionary distances (Jones et al. 1992) and the reliability of each branch was tested with 1000 bootstraps (Felsenstein 1985). Finally, the phylogenetic trees were visualised using iTOL web server (https://itol.embl.de/) (Letunic and Bork 2011).
Domain configuration analysis
All the sequences of SAT and OAS-TL were subjected to CD-search in NCBI against conserve domain database (CDD) with threshold E value 0.01. Only the specific hits were considered among the CD outputs as they are high confidence, top-ranked hits (https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi).
Protein physicochemical characterization
ProtParam (http://web.expasy.org/protparam/) (Gasteiger et al. 2005) were used to analyse the physiochemical properties of SAT and OAS-TL proteins such as amino acid number, molecular weight (MW), number of negatively and positively charged residues, half-life, aliphatic index (AI), instability index (II), isoelectric point (pI), and grand average hydrophobicity (GRAVY) (Ning et al. 2017).
Secondary and tertiary structure prediction
For the structural prediction of SAT and OAS-TL, SrpG and SrpH, of S. elongatus PCC 7942, respectively, were selected based on previous literatures. The folds and features of SrpG and SrpH were analysed by secondary and tertiary structures. The secondary structure of the proteins was generated using PDBsum server (http://www.ebi.ac.uk/pdbsum). Further, the tertiary structures of SrpG and SrpH were generated using RaptorX (http://raptorx.uchicago.edu/StructurePrediction/predict/) (Källberg et al. 2012) and Discovery studio, respectively, and subsequently visualized using UCSF Chimera 1.13.1 software (https://www.cgl.ucsf.edu/chimera/). After that, PROCHEK (http://www.ebi.ac.uk/thornton-srv/software/PROCHECK/) (Laskowski et al. 1996), ProSA (https://prosa.services.came.sbg.ac.at/prosa.php) (Sippl 1993), and VADAR (http://vadar.wishartlab.com/) (Willard et al. 2003) web servers were used to assesses the tertiary protein structures. COFACTOR was used for further analysis of protein (Zhang et al. 2017).
Protein–protein interaction analysis and molecular docking
The functional interactions of the cellular proteins with SrpG and SrpH were predicted using STRING (‘Search Tool for Retrieval of Interacting Genes/Proteins’) version 11.0 database (https://string-db.org), which integrates known and predicted protein–protein interaction (PPIs), were applied to predict functional interactions of the proteins with default parameters (Szklarczyk et al. 2016). Active interaction sources, including text mining, experiments (biochemical/genetic data), databases (previously curated pathway and protein-complex knowledge), neighbourhood, gene fusion, co-occurrence and co-expression as well as species limited to “S. elongatus PCC 7942” and an interaction score > 0.4 were applied to construct the PPI networks. In the networks, the nodes correspond to the proteins and the edges represent the interactions. The network Stats were: number of nodes, i.e., 11; number of edges, i.e., 23; average node degree, i.e., 4.18; and average local clustering coefficient, i.e., 0.871. Additionally, the PPI enrichment p-values are 0.000751 and 0.00063 for SrpG and SrpH, respectively.
Further, the BIOVIA Discovery Studio 2019 was used to perform docking analysis of SrpG and SrpH. ZDOCK program was used for docking, which considered SrpG as the receptor and SrpH as the ligand. ZDOCK protocol provides rigid body docking of two proteins based on the Fast Fourier Correlation Technique. It performs a systematic search on a uniform sample of docked protein poses and uses ZRANK algorithm to predict the optimal interactions. Further, the top scoring docked protein poses were selected to calculate the docking parameters. ZRANK algorithm reranked the initial-stage ZDOCK predictions with detailed electrostatics, van der Waals and desolation energy terms of the pose. For more accurate predictions default angular step size of 6 was selected to perform finer conformational sampling. Water molecules were removed from both the protein structures before running the program. Further, refinement of the docked complex was done using RDOCK (Refined Docked Proteins) by evaluating of electrostatic and desolvation energies. RDOCK is a CHARMm-based protocol to remove clashes and optimize polar and charge interactions. During the RDOCK optimization, each predicted docked complex is subjected to 130 steps of Adopted Basis Newton–Raphson (ABNR) energy minimization. Finally, the docked complex was subsequently visualized using UCSF Chimera 1.13.1 software.
Results and discussions
Distribution and diversity of cyanobacterial serine acetyl transferase (SAT) and O-acetyl serine (thiol) lyase (OAS-TL) proteins
Among 51 cyanobacteria, the distribution of SATs was harboured by 9.8, 11.7, and 78.4% in unicellular, filamentous, and heterocytous strains, respectively (Fig. 1a). Every one of all cyanobacterial organisms possesses a single gene copy for SAT suggesting that these genes are highly conserved throughout the evolutionary history; however, few heterocytous strains, i.e., Cylindrospermum sp. NIES-4074, Nostoc sp. ATCC 43,529, Nostoc sp. PCC 7107, Anabaena sp. PCC 7108, Fischerella sp. PCC 9431, Hapalosiphon sp. MRB220, Trichormus sp. NMC-1, Nostoc sp. NIES-4103, Calothrix sp. NIES-2098, Calothrix sp. NIES-2100, Fischerella sp. NIES-4106, and Tolypothrix sp. PCC 7910 subsumed two copies of the gene. Similarly, 25.3% of OAS-TL was distributed among both unicellular and filamentous cyanobacteria, while 49.2% was harboured by heterocytous ones. Unlike SAT, two copies of OAS-TL were not only observed in the heterocytous strains, i.e., Calothrix sp. PCC 7507, Nostoc sp. PCC 7524, Anabaena sp. PCC 7108, Trichormus sp. NMC-1, Scytonema sp. HK-05, Nodularia sp. NIES-3585, Calothrix sp. NIES-2098, Calothrix sp. NIES-2100, Nostoc sp. ATCC 53789, and Tolypothrix sp. PCC 7910, but also in the filamentous strains such as Microcoleus sp. SU53 and Desertifilum sp. 1PPASB-1220 (Fig. 1a). Besides, two copies of OAS-TL were also harboured by unicellular cyanobacterium, Synechococcus elongatus PCC 11802. The presence of two copies of the SAT and OAS-TL genes, especially, in the heterocytous strains might be a function of larger genome and complex morphology. Despite, genome streaming as a mechanism for adaptation generally attained to reduced energy expenditure in smaller genome might also reduce the copy number of these genes within the picocyanobacteria like Synechococcus (Bhattacharjee and Mishra 2020; Kharwar et al. 2021). The presence of two copies of OAS-TL in unicellular and filamentous cyanobacteria suggested the existence of at least one more other factor apart from genome size and morphology favouring the acquisition of gene. In our study, the acquisition of two copies of OAS-TL in few cyanobacteria such as S. elongatus PCC 11802, Microcoleus sp. SU53 and Desertifilum sp. 1PPASB-1220 perhaps suggested the adaptive advantage in these microorganisms, despite morphology favouring the profusion of gene, further strengthens the previous assumption (Kharwar et al. 2021).
Furthermore, the distribution of SAT and OAS-TL among the habitats of cyanobacteria showed the presence of 79.3% SAT harbouring strains in freshwater and terrestrial habitat, whereas 6.3, 3.1, and 7.9% were subsumed by strains of marine, saline-alkaline lakes, and exclusively freshwater habitats, respectively. Additionally, 3.1% symbionts also consisted of SAT proteins (Fig. 1b). Besides, 52.5% OAS-TL was found in exclusively freshwater ecosystem. 30, 10, 5, 1.25, and 1.25%, consisted of freshwater and terrestrial, followed by marine, saline-alkaline lakes, hotsprings, and symbiotic association, respectively (Fig. 1c). Higher percentage of SAT and OAS-TL in freshwater and terrestrial habitats was correlated with the abundance of heterocytous strains in those ecosystems. Since heterocytous cyanobacteria have larger genome size, they probably subsumed more copies of genes. Besides, the picocyanobacteria like Synechococcus, Synechocystis, and Prochlorococcus are exclusive marine and mostly harbour single gene copies due to genome streamlining (Kharwar et al. 2021). However, the presence of more SAT and OAS-TL in freshwater cyanobacteria compared to marine strains might confer adaptive advantage to the former, since, freshwater, unlike marine, has limited availability of sulphur (Bochenek et al. 2013).
Sequence homology and alignment
Alignment of 60 SAT amino acid sequences revealed considerable identity. The carboxyl-terminal sequence was found to be conserved from the unicellular to heterocytous strains (Fig. S1a). The C-terminal end of SAT plays a crucial role in the formation of cysteine synthase complex since this tail binds OAS-TL in case of E. coli (Mino et al. 1999), Haemophilus influenza (Campanini et al. 2005), Leishmania donovani (Raj et al. 2012), and A. thaliana (Bogdanova and Hell 1997; Feldman-Salit et al. 2012; Francois et al. 2006; Kumaran and Jez 2007), thus conferring the possibility of CSC formation among cyanobacteria. Additionally, the consensus hexapeptide repeat, {V/L/I}-G-XXXX was also observed, signifying the acetyl transferase activity (Dicker and Seetharam 1992; Gorman and Shapiro 2004; Olsen et al. 2007; Vaara 1992; Vuorio et al.1994).
Likewise, sequence alignment of OAS-TL showed significant conserved residues. The highly conserved consensus “SVKDR” motif forming PLP attachment site around the K residue was also evident in HMM logo (Bairoch et al. 1996; Saito et al. 1993). The ɛ-amino group of this K residue forms a salt bridge with the phosphate group of PLP, which is prerequisite for the catalysing the reaction (Saito et al. 1993). The PLP cofactor was covalently bound to K by schiff linkage on alpha helix of the N-terminal region and stays at the large gap made between the N- and C-terminal domains. Furthermore, several conserved G residues were evident which are believed to play a structural and/or functional role in binding the phosphoryl group of the PLP cofactor (Marceau et al. 1988a; b). Moreover, the asparagine loop consists of “TXGNT” motif, a signature sequence of cysteine synthase family, also was conserved in cyanobacterial OAS-TL (Chinthalapudi et al. 2008). This asparagine loop in TSGNT motif is involved in the binding of O-acetyl serine (Fig. S1b).
Phylogenetic analyses
The phylogenetic tree of SAT displayed two distinct clusters i.e., one prominent cluster (cluster I), and a minor cluster (cluster II) (Fig. 2a) as a result of evolutionary changes between amino acid sequences, where GGN, GAKS, and IYQGVTL motifs acquired by simple primitive unicellular life forms (S. elongatus PCC 11,802, S. elongatus PCC 7942, and Synechococcus sp. PCC 6312), the rest of the strains in cluster I subsumed GAG, GGTG, and IYQAVTL motifs which have morphological variabilities (unicellular to heterocytous). These GAG, GGTG, and IYQAVTL motifs showed wide distribution among filamentous and heterocytous strains of cyanobacteria. Despite, the functional significance of these motifs was not evident. Further, three distinct clusters, i.e., clusters I, II, and III were observed in the phylogenetic tree of OAS-TL (Fig. 2b). Clusters I and III subsumed unicellular to heterocytous cyanobacterial strains, whereas cluster II was purely represented by strains of S. elongatus. All the members of cluster I harboured GNS motif except Nostoc sp. TCL240-02 and Nostoc sp. PA-18-2419 consisting GNG. Strains of cluster II were characterised as GPN motif, while GAG motif was harboured by strains in cluster III, except Nostoc sp. ATCC 53789 having GNG motif.
Phylogenetic distribution of SAT (a) and OAS-TL (b) proteins among cyanobacterial strain. Amino acid sequences of SAT and OAS-TL were obtained from the NCBI database and subjected to unrooted phylogenetic tree construction using the neighbour-joining (NJ) method built in the MEGA software. The evolutionary distances were computed using the Poisson correction method and are in the units of the number of amino acid substitutions per site. Bootstrap values are hidden for clarity of the image and all positions containing gaps and missing data were eliminated
As exhibited in phylogenetic tree, the SAT and OAS-TL proteins within same cluster have similar motif compositions but displayed variance among the different clusters. The similar motif arrangements probably indicated conserved protein architecture within the phylogenetic cluster.
Domain configuration of serine acetyl transferase (SAT) and O-acetyl serine (thiol) lyase (OAS-TL)
Comparative analysis of domain architecture of SAT from the cyanobacteria (Scytonema sp., Nostoc sp. PCC 7120, Oscillatoria nigro-viridis, S. elongatus PCC 7942, and Synechocystis sp. CACIAM-05), bacteria (E. coli and Salmonella typhimurium) and plant (Arabidopsis thaliana) showed conserved single functional domain among protein, exhibiting structural uniformity throughout the evolution (Table 1). SATs from various organisms showed catalytic N-terminal serine acetyltransferase (SATase_N), whereas C-terminal possessed well-conserved left-handed parallel β-sheet helices (LβH), due to a hexapeptide repeat sequence, a characteristic of acyl and acetyl transferase family proteins (Fig. S1a). However, OAS-TL belongs to tryptophan synthase beta II super family (Table 1), possesses cystathionine beta-synthase (CBS)-like sequence and PLP dependent enzyme domain (Momany et al. 1995). At the N-terminal, catalytic K (Fig. S1b) was found to be highly conserved in bacteria, cyanobacteria, and A. thaliana indicating conservation of protein functionality (Mozzarelli et al. 2011).
Protein characterization
The molecular weight of SAT ranged among cyanobacteria from 27.31 to 34.48 kDa while in bacteria and A. thaliana, it has a molecular weight of 290 and 42.72 kDa, respectively (Table S1). The pI for cyanobacterial SAT proteins ranged from 5.62 to 9.1, whereas pI of SAT in E. coli, S. typhimurium, and A. thaliana were 6.05, 6.41, and 7.73, respectively. Further, the II values of the studied cyanobacterial SAT except AZB72704.1 and WP_017743387.1 of S. elongatus PCC 11801 and Scytonema hofmannii, respectively, were predicted to be stable having an in vivo half-life of > 16 h. The AI is defined as the relative volume occupied by aliphatic side chains such as alanine, valine, isoleucine, and leucine, which determines thermostability of the protein, ranging from 99.12 to 115.2 for cyanobacterial SATs. GRAVY index indicates the solubility of protein, a higher positive value indicates a greater hydrophobicity and lower solubility in water. The positive GRAVY scores predicted for SAT of E. coli str. K-12 substr. W3110, Salmonella enterica subsp. enterica serovar Typhimurium, Oscillatoria nigro-viridis, and Nostoc sp. PCC 7120 depicted the hydrophobic and non-polar nature of the protein in these organisms, probably suggested their membrane localisation. Contrastingly, the negative GRAVY scores for SAT of Synechocystis sp. PCC 6803, S. elongatus PCC 11801, S. hofmannii, and A. thaliana exhibited hydrophilic nature perhaps indicated the cytosolic localisation of these proteins.
However, the OAS-TL proteins of cyanobacteria ranged from 34.1 to 34.7 kDa, while for bacteria and A. thaliana the molecular weights were noticed to be 34 and 33.8 kDa, respectively. The theoretical pI value of cyanobacterial OAS-TL ranges from 5.21 to 5.82, whereas 5.8 and 5.9 were noticed in bacteria and A. thaliana, respectively, conferred its acidic nature (Table S2). Further, II values were found to be less than 40 depicting stable nature of the studied OAS-TL proteins, while the AI ranging from 89.14 to 110.67 perhaps indicated thermostability of the proteins. The positive GRAVY values of S. elongatus PCC 7942, Scytonema HK-05, and A. thaliana signify their hydrophobic nature, whereas the negative values in case of E. coli, S. typhimurium, Synechocystis, Oscillatoria, and Nostoc sp. 7120 indicated their cytosolic nature (Govardhana and Kumudini 2020).
Structural analyses
The structural features of SAT and OAS-TL proteins were evaluated by analysing protein models of SrpG and SrpH, respectively, from S. elongatus PCC 7942. PROCHECK validates structure of the protein model through the Ramachandran plot and checks the stereochemical nature of the protein structure by analysing residue-by-residue geometry and overall structure geometry. The backbone conformation and overall stereochemical qualities of the modelled SrpG and SrpH were validated by analysing Ramachandran plot (Lovell et al. 2002). In SAT, 91.5% residues appeared in the favoured region, and 8.5% residues in the additionally allowed region, while none of the residues in the generously allowed and disallowed regions, respectively, suggesting that the proposed model is stereo-chemically stable (Fig. 3a, c; Table 2). Besides, 93.5% residues in the favoured region, 5.7 and 0.8% residues in the additionally allowed and generously allowed regions, respectively, and no residues in the disallowed regions were observed (Fig. 4b, d; Table 2) also suggesting the stereo-chemical stability of the SrpH model. Further, the structure quality of the modelled protein structures was also verified by VADAR analysis. VADAR prediction displayed 45, 23, 30, and 25% helix, beta, coil, and turns, respectively, with mean H bond energy of − 2.2 (SD = 0.9) against the expected value of − 2.0 (SD = 0.8) for the SAT model (Fig. S2a), whereas 40, 24, 34, and 24% helix, beta, coil, and turns, respectively, with mean H bond energy of − 1.6 (SD = 1.0) against − 2.0 (SD = 0.8) expected value for OAS-TL model (Fig. S2b). The model was also validated using ProSA-web server and showed that the protein folding energy of the modelled structure, i.e., Z-score which evaluates the energy of each of the amino acids of the protein molecule and verifies the 3D structure by evaluating the local compatibility of the model related to good protein structure. The Z-score values for the 3D models of SrpG and SrpH were − 8.2 and − 9.91, respectively. The Z score of the modelled protein structures was within the acceptable range, i.e., − 10 to 10 and the negative Z score value is considered to be very good quality protein models and, therefore, good and reliable (Wiederstein and Sippl 2007). On the basis of our findings, it is predicted that the quality of modelled proteins is of good quality, and thus, considered for analysing their putative interaction (Fig. 3e, f).
Representation of 3-D models of SAT (a) and OAS-TL (b). Ramachandran plot analysis of SAT (c) and OAS-TL (d) proteins; the plot calculations were computed by PROCHECK server. The red regions in the graph indicate the most allowed regions [A, B, L];, additional allowed regions [a, b, l, p] are indicated as brown, and generously allowed regions[~ a, ~ b, ~ l, ~ p] are indicated as green and yellow shades; results of ProSA analysis of SAT (e) and OAS-TL (f) proteins; topology of SAT (g) and OAS-TL (h) proteins
Protein–protein interaction analysis: STRING analysis of SAT (a) and OAS-TL (b) proteins with their interaction proteins; Ribbon diagram (c); Surface view of the docked model indicating interacting sites (d); COFACTOR analysis depicting binding of OAS-TL with PLP (e). This figure was produced using UCSF Chimera 1.13.1 Visualizer tool
The structure of SrpG composed of 9 N-terminal α-helices and 14 C-terminal β-strands (Fig. 3g). The β-strands in the protein form two sets, i.e., β1-β4-β7-β10-β13 and β2-β5-β8-β11-β3-β6-β9-β12-β14 having antiparallel orientation; however, each of these strands are placed parallel within the set. This topology of the protein is consistent with other β-elimination enzymes (Burkhard et al. 1998). The conserved αβα sandwich fold of SrpH was conferred by 14 α-helices and 10 β-strands (Fig. 3h). The β-strands have both parallel and antiparallel orientations, i.e., β1↑β2↓β3↑β4↑β5↑β6↑β7↓β8↓β9↓β10↓. Among these, β2 and β3; β3 and β4; β4 and β5; β5 and β6; β7 and β8 are flanked by one α-helix, whereas two α-helices were positioned between β6 and β7; β8 and β9. Besides, 3 α-helices were present between β9 and β10 in the structure of SrpH. This topology displayed the typical β-grasp fold as observed in the sulphur carrier proteins, like ThiS (Lehmann et al. 2006) and MoaD (Lake et al. 2001) as well as ubiquitin (Vijay-Kumar et al. 1985).
Molecular docking
STRING analysis exhibited the functional PPI of SrpG and SrpH with the network of various proteins, mostly subsumed in the sulphur metabolism (Fig. 4a, b). The network analysis revealed our target protein interacts with ten different proteins for carrying out its functions. The SrpG and SrpH primarily interacted with each other and CysE (SAT) also. Besides, SrpG showed strong interaction with SIR (sulfite reductase), yet high score of interaction between SrpG and SrpH further strengthened the idea of CSC formation in cyanobacteria.
Further, identification of dimerization interface or active site of both the proteins has been analysed for predicting the possible interaction sites. Notably, among seven and twenty-five residues of SrpG and SrpH, only four and eight residues, respectively, are involved in the interaction (Table 3a). In the docking analysis, the docked pose complex displayed interaction between SrpG and SrpH proteins to form CSC (Fig. 4c, d). Interacting residues of SrpG were H216, G289, W292, and T294, whereas the interacting residues of SrpH were L20, V21, R22, L35, R41, E169, D170, and T299 (Fig. S3; Table 3b). Furthermore, the electrostatics energy was calculated using CHARMm algorithm with a distance-dependent dielectric constant. The electrostatics energy of the docked complex is − 26 kcaL/moL. The negative value of electrostatics energy signifies the stability of the docked complex.
As in bacteria and plants, this potential interaction of SrpG and SrpH showed the possibility of hetero-oligomeric bi-enzyme CSC formation in cyanobacteria, critical for the fine-tuning of cysteine biosynthesis. This is the first study to report CSC formation and their regulation in cyanobacteria. Formation of CSC in cyanobacteria not only facilitates a consensus regulatory loop for cysteine biosynthesis maintained by relative concentration of OAS and sulfide in the cell, but also acts as a sensor for intracellular status of sulphur (Feldman-Salit et al. 2009; Wirtz and Hell 2006). Since the activity of SAT is induced upon CSC formation, whereas OAS-TL activity is reduced upon interaction, an equilibrium between bounded and free SAT and OAS-TL is maintained. Sulfur-sufficient condition accompanied higher sulfide and stabilisation of CSC and activating SAT activity, thereby forming OAS. While upon sulfur-deficient, lower concentration of sulfide and higher OAS triggered dissociation of CSC inducing the inactivation and activation of SAT and OAS-TL, respectively, thus form cysteine (Fig. 5). Therefore, in cyanobacteria, demand-dependent positive regulation of sulfate assimilation by OAS provides homeostasis to cysteine biosynthesis. Additionally, the induction of srpGH by sulphur stress may give cyanobacterial cells of S. elongatus PCC 7942 the ability to convert low level of sulphur into cysteine at a much higher rate than the normal, thus giving this organism a selective advantage for scavenging of sulphur compounds upon sulphur-deprived condition (Nicholson et al. 1995). Similar to bacteria and multicellular organisms such as green algae and plants, CSC in cyanobacteria is a tightly controlled mechanism orchestrated by specialized molecular mechanism.
However, the COFACTOR analysis predicted that 15 amino acid residues, namely K47, I48, N78, G181, V182, G183, T184, G185, G186, T187, G234, I235, S278, P305, and D306 of SrpH bind to PLP (Fig. 4e). Among all these residues, K47 is the conserved amino acid for binding site of PLP in OAS-TL through a schiff base linkage (Chattopadhyay et al. 2007; Rabeh and Cook 2004). A glycine and threonine residues bind the phosphate group and is characteristic of PLP binding proteins (Momany et al. 1992). The SrpH uses PLP as a cofactor for its catalytic activity and follows a ping-pong mechanism which was corroborated by the previous studies (Cook and Wedding 1976; Masada et al. 1975).
Conclusion
The significance of homeostasis of cysteine biosynthesis can be perceived by the involvement of various tiers of regulation underlying this mechanism in bacteria and plants. However, in cyanobacteria the regulatory process of cysteine biosynthesis is lesser understood due to limited knowledge regarding SAT and OAS-TL, the prime enzymes involved in cysteine biosynthesis. Although the presence of these enzymes in cyanobacteria indicated a similar regulatory mechanism as in plant or bacteria, yet the formation of heteromeric bi-enzyme CSC is uncertain. Here, we showed the distribution and abundance of SAT and OAS-TL proteins in cyanobacteria and also elucidated their domain configuration and phylogeny. Our study, for the first time, conferred the formation of putative CSC in cyanobacteria based on theoretical modelling and molecular docking. Though the wet-lab studies are required to affirm the formation of CSC in cyanobacteria, yet these findings will provide useful insights into the functionality and regulation of these genes and thereby extend our knowledge of cysteine biosynthesis in these photoautotrophs.
Data availability
Yes, the authors agreed for the availability of data to ensure data transparency norms.
References
Bairoch A, Bucher P, Hohann K (1996) The PROSITE database, its status in 1995. Nucleic Acids Res 24:189–196. https://doi.org/10.1093/nar/24.1.189
Berkowitz O, Wirtz M, Wolf A (2002) Use of bio molecular interaction analysis to elucidate the regulatory mechanism of the cysteine synthase complex from Arabidopsis thaliana. J Biol Chem 277:30629–30634. https://doi.org/10.1074/jbc.M111632200
Bhattacharjee S, Mishra AK (2020) The tale of caspase homologues and their evolutionary outlook: deciphering programmed cell death in cyanobacteria. J Exp Bot 71:4639–4657. https://doi.org/10.1093/jxb/eraa213
Bochenek M, Etherington GJ, Koprivova A, Mugford ST, Bell TG, Malin G, Kopriva S (2013) Transcriptome analysis of the sulfate deficiency response in the marine microalga Emiliania huxleyi. New Phytol 199:650–662. https://doi.org/10.1111/nph.12303
Bogdanova N, Hell R (1997) Cysteine synthesis in plants: protein-protein interactions of serine acetyltransferase from Arabidopsis thaliana. Plant J 11:251–262. https://doi.org/10.1046/j.1365-313x.1997.11020251.x
Buchanan BB, Balmer Y (2005) Redox regulation: a broadening horizon. Annu Rev Plant Biol 56:187–220. https://doi.org/10.1146/annurev.arplant.56.032604.144246
Buchner P, Takahashi H, Hawkesford MJ (2004) Plant sulphate transporters: co-ordination of uptake, intracellular and long-distance transport. J Exp Bot 55:1765–1773. https://doi.org/10.1093/jxb/erh206
Campanini B, Speroni F, Salsi E, Cook PF, Roderick SL, Huang B, Bettati S, Mozzarelli A (2005) Interaction of serine acetyltransferase with O-acetylserinesulfhydrylase active site: evidence from fluorescence spectroscopy. Protein Sci 14:2115–2124. https://doi.org/10.1110/ps.051492805
Chattopadhyay A, Meier M, Ivaninskii S, Burkhard P, Speroni F, Campanini B, Bettati S, Mozzarelli A, Rabeh WM, Li L, Cook PF (2007) Structure, mechanism, and conformational dynamics of O-acetylserine sulfhydrylase from Salmonella typhimurium: comparison of A and B isozymes. Biochemistry 46:8315–8330. https://doi.org/10.1021/bi602603c
Chinthalapudi K, Kumar M, Kumar S, Jain S, Ala N, Gourinath S (2008) Crystal structure of native O-acetyl-serine sulfhydrylase from Entamoeba histolytica and its complex with cysteine: Structural evidence for cysteine binding and lack of interactions with serine acetyl transferase. Proteins: Struct Funct Bioinform 72:1222–1232. https://doi.org/10.1002/prot.22013
Cook PF, Wedding RT (1976) A reaction mechanism from steady state kinetic studies for O-acetylserine sulfhydrylase from Salmonella typhimurium LT-2. J Biol Chem 251:2023–2029. https://doi.org/10.1016/S0021-9258(17)33649-9
Davidian JC, Kopriva S (2010) Regulation of sulfate uptake and assimilation-the same or not the same? Mol Plant 3:314–325. https://doi.org/10.1093/mp/ssq001
Dicker IB, Seetharam S (1992) What is known about the structure and function of the Escherichia coli protein FirA? Mol Microbiol 6:817–823. https://doi.org/10.1111/j.1365-2958.1992.tb01532.x
Diessner W, Schmidt A (1980) Isoenzymes of cysteine synthase in the cyanobacterium Synechococcus 6301. Z Pjkmsenphysiol 102:57–68. https://doi.org/10.1016/S0044-328X(81)80217-6
Droux M, Ruffet ML, Douce R, Job D (1998a) Interactions between serine acetyltransferase and O-acetylserine (thiol) lyase in higher plants–structural and kinetic properties of the free and bound enzymes. Eur J Biochem 255:235–245. https://doi.org/10.1046/j.1432-1327.1998.2550235.x
Droux M, Ruffet ML, Douce R, Job D (1998b) Interactions between serine kinetic properties of the free and bound enzymes. Eur J Biochem 255:235–245. https://doi.org/10.1046/j.1432-1327.1998.2550235.x
Feldman-Salit A, Wirtz M, Hell R, Wade RC (2009) A mechanistic model of the cysteine synthase complex. J Mole Biol 386:37–59. https://doi.org/10.1016/j.jmb.2008.08.075
Feldman-Salit A, Wirtz M, Lenherr ED, Throm C, Hothorn M, Scheffzek K, Hell R, Wade RC (2012) Allosterically gated enzyme dynamics in the cysteine synthase complex regulate cysteine biosynthesis in Arabidopsis thaliana. Structure 20:292–302. https://doi.org/10.1016/j.str.2011.11.019
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Foyer CH, Noctor G (2011) Ascorbate and glutathione: the heart of the redox hub. Plant Physiol 155:2–18. https://doi.org/10.1104/pp.110.167569
Francois JA, Kumaran S, Jez JM (2006) Structural basis for interaction of O-acetylserine sulfhydrylase and serine acetyltransferase in the Arabidopsis cysteine synthase complex. Plant Cell 18:3647–3655. https://doi.org/10.1105/tpc.106.047316
Gasteiger E, Hoogland C, Gattiker A (2005) Protein identification and analysis tools on the ExPASy server. In: The proteomics protocols handbook. Humana press, pp 571–607
Gorman J, Shapiro L (2004) Structure of serine acetyltransferase from Haemophilus influenzae Rd. Acta Crystallogr Sect D Biol Crystallogr 60:1600–1605. https://doi.org/10.1107/S0907444904015240
Govardhana M, Kumudini BS (2020) In-silico analysis of cucumber (Cucumis sativus L.) Genome for WRKY transcription factors and cis-acting elements. Comp Biol Chem 85:107212. https://doi.org/10.1016/j.compbiolchem.2020.107212
Hopkins L, Parmar S, Blaszczyk A, Hesse H, Hoefgen R, Hawkesford MJ (2005) O-acetylserine and the regulation of expression of genes encoding components for sulfate uptake and assimilation in potato. Plant Physiol 138:433–440. https://doi.org/10.1104/pp.104.057521
Jones DT, Taylor WR, Thornton JM (1992) The rapid generation of mutation data matrices from protein sequences. Comp Appl Biosci 8:275–282. https://doi.org/10.1093/bioinformatics/8.3.275
Joshi P, Gupta A, Gupta V (2019) Insights into multifaceted activities of CysK for therapeutic interventions. 3 Biotech 9:44. https://doi.org/10.1007/s13205-019-1572-4
Jost R, Berkowitz O, Wirtz M, Hopkins L, Hawkesford MJ, Hell R (2000) Genomic and functional characterization of the oas gene family encoding O acetylserine (thiol)lyases, enzymes catalyzing the final step in cysteine biosynthesis in Arabidopsis thaliana. Gene 253:237–247. https://doi.org/10.1016/s0378-1119(00)00261-4
Källberg M, Wang H, Wang S, Peng J, Wang Z, Lu H, Xu J (2012) Template-based protein structure modeling using the RaptorX web server. Nat Protoc 7:1511–1522. https://doi.org/10.1038/nprot.2012.085
Khan MS, Haas FH, Allboje Samami A, Moghaddas Gholami A, Bauer A, Fellenberg K, Reichelt M, Hansch R, Mendel RR, Meyer AJ, Wirtz M, Hell R (2010) Sulfite reductase defines a newly discovered bottleneck for assimilatory sulfate reduction and is essential for growth and development in Arabidopsis thaliana. Plant Cell 22:1216–1231. https://doi.org/10.1105/tpc.110.074088
Kharwar S, Mishra AK (2020) Unraveling the complexities underlying sulfur deficiency and starvation in the cyanobacterium Anabaena sp. PCC 7120. Environ Exp Bot 172:103966. https://doi.org/10.1016/j.envexpbot.2019.103966
Kharwar S, Bhattacharjee S, Mishra AK (2021) Disentangling the impact of sulfur limitation on exopolysaccharide and functionality of Alr2882 by in silico approaches in Anabaena sp. PCC 7120. Appl Biochem Biotech. https://doi.org/10.1007/s12010-021-03501-3
Kopriva S, Koprivova A (2003) Sulphate assimilation: a pathway which likes to surprise. In: Sulphur in plants. Springer, Dordrecht, pp 87–112. https://doi.org/10.1007/978-94-017-0289-8_5
Kredich NM (1996) Biosynthesis of cysteine. In: Neidhardt FC, Curtiss R, Ingraham JL, Lin ECC, Low KB, Magasanik B, Reznikoff WS, Riley M, Schaechter M, Umberger E (eds) Escherichia coli and Salmonella typhimurium. Cellular and molecular biology. ASM Press, Washington D.C., pp 514–527
Kredich NM, Tomkins GM (1966) The enzymic synthesis of L-cysteine in Escherichia coli and Salmonella typhimurium. J Biol Chem 241:4955–4965. https://doi.org/10.1016/S0021-9258(18)99657-2
Kredich NM, Becker MA, Tomkins GM (1969) Purification and characterization of cysteine synthetase, a bifunctional protein complex, from Salmonella typhimurium. J Biol Chem 244:2428–2439. https://doi.org/10.1016/S0021-9258(19)78241-6
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kumaran S, Jez JM (2007) Thermodynamics of the interaction between O-acetylserine sulfhydrylase and the C-terminus of serine acetyltransferase. Biochemistry 46:5586–5594. https://doi.org/10.1021/bi7001168
Lake MW, Wuebbens MM, Rajagopalan KV, Schindelin H (2001) Mechanism of ubiquitin activation revealed by the structure of a bacterial MoeB-MoaD complex. Nature 414:325–329. https://doi.org/10.1038/35104586
Laskowski RA, Rullmannn JA, MacArthur MW, Kaptein R, Thornton JM (1996) AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J Biomol 8:477–486. https://doi.org/10.1007/BF00228148
Lehmann C, Begley TP, Ealick SE (2006) Structure of the Escherichia coli ThiS-ThiF complex, a key component of the sulfur transfer system in thiamin biosynthesis. Biochemistry 45:11–19. https://doi.org/10.1021/bi051502y
Letunic I, Bork P (2011) Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478. https://doi.org/10.1093/nar/gkr201
Lovell SC, Davis IW, Arendall WB III, Lovell SC, Davis IW, Arendall WB III, De Bakker PI, Word JM, Prisant MG, Richardson JS, Richardson DC (2002) Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins: Struct Fun Genet 50:437–450. https://doi.org/10.1002/prot.10286
Marceau M, Lewis SD, Scaefer LA (1988a) The glycine-rich region of Escherichia coli D-serine dehydratase. J Bol Chem 263:16934–16941. https://doi.org/10.1016/S0021-9258(18)37481-7
Marceau M, McFali E, Lewis SD, Scaefer JA (1988b) D-serine dehydratase from Escherichia coli. J Biol Chem 263:16926–16933. https://doi.org/10.1016/j.bbapap.2011.10.017
Masada M, Fukushima K, Tamura G (1975) Cysteine synthase from rape leaves. J Biochem 77:1107–1115. https://doi.org/10.1093/oxfordjournals.jbchem.a130811
Mino K, Yamanoue T, Sakiyama T, Eisaki N, Matsuyama A, Nakanishi K (1999) Purification and characterization of serine acetyltransferase from Escherichia coli partially truncated at the C-terminal region. Biosci Biotech Biochem 63:168–179. https://doi.org/10.1271/bbb.63.168
Momany C, Ernst S, Hackert ML (1992) Ornithine decarboxylase from Lactobacillus embodies a PLP-binding scaffold and a GTP effector site. Am Crysiallogr Assoc Absir 20:44
Momany C, Ghosh R, Hackert ML (1995) Structural motifs for pyridoxal-5′-phosphate binding in decarboxylases: an analysis based on the crystal structure of the Lactobacillus 30a ornithine decarboxylase. Pro Sci 4:849–854. https://doi.org/10.1002/pro.5560040504
Mozzarelli A, Bettati S, Campanini B, Salsi E, Raboni S, Singh R, Spyrakis F, Kumar VP, Cook PF (2011) The multifaceted pyridoxal 5′-phosphate-dependent O-acetylserine sulfhydrylase. Biochim Biophys Acta 1814:1497–1510. https://doi.org/10.1016/j.bbapap.2011.04.011
Nicholson ML, Gaasenbeek M, Laudenbach DE (1995) Two enzymes together capable of cysteine biosynthesis are encoded on a cyanobacterial plasmid. Mol Gen Gene 247:623–632. https://doi.org/10.1007/BF00290354
Ning P, Liu C, Kong J, Lv J (2017) Genome-wide analysis of WRKY transcription factors in wheat (Triticum aestivum L.) and differential expression under water deficit condition. Peer J 5:e3232. https://doi.org/10.7717/peerj.3232
Noctor G, Mhamdi A, Chaouch S, Han YI, Neukermans J, Marquez-Garcia BE, Queval G, Foyer CH (2012) Glutathione in plants: an integrated overview. Plant Cell Environ 35:454–484. https://doi.org/10.1111/j.1365-3040.2011.02400.x
Olsen LR, Vetting MW, Roderick SL (2007) Structure of the E. coli bifunctional GlmU acetyltransferase active site with substrates and products. Protein Sci 16:1230–1235. https://doi.org/10.1110/ps.072779707
Rabeh WM, Cook PF (2004) Structure and mechanism of O-acetylserine sulfhydrylase. J Biol Chem 279:26803–26806. https://doi.org/10.1074/jbc.R400001200
Raj I, Kumar S, Gourinath S (2012) The narrow active-site cleft of O-acetylserine sulfhydrylase from Leishmania donovani allows complex formation with serine acetyltransferases with a range of C-terminal sequences. Acta Crystallogr Sect D Biol Crystallogr 68:909–919. https://doi.org/10.1107/S0907444912016459
Richau KH, Kaschani F, Verdoes M, Pansuriya TC, Niessen S, Stüber K, Colby T, Overkleeft HS, Bogyo M, Van der Hoorn RA (2012) Subclassification and biochemical analysis of plant papain-like cysteine proteases displays subfamily-specific characteristics. Plant Physiol 158:1583–1599. https://doi.org/10.1104/pp.112.194001
Ruffet ML, Droux M, Douce R (1994) Purification and kinetic properties of serine acetyltransferase free of O-acetylserine(thiol)lyase spinach chloroplasts. Plant Physiol 104:597–604. https://doi.org/10.1104/pp.104.2.597
Saito K, Kurosawa M, Murakoshi I (1993) Determination of functional lysine residue of a plant cysteine synthase by site-directed mutagenesis, and the molecular evolutionary implications. FEBS 328:111–114. https://doi.org/10.1016/0014-5793(93)80976-2
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sippl MJ (1993) Recognition of errors in three-dimensional structures of proteins. Proteins: Struct Funct Bioinform 17:355–362. https://doi.org/10.1002/prot.340170404
Szklarczyk D, Morris JH, Cook H, Kuhn M, Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P (2016) The STRING database in 2017: quality controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw937
Takahashi H, Kopriva S, Giordano M, Saito K, Hell R (2011) Sulfur assimilation in photosynthetic organisms: molecular functions and regulations of transporters and assimilatory enzymes. Ann Rev Plant Biol 62:157–184. https://doi.org/10.1146/annurev-arplant-042110-103921
Vaara M (1992) Eight bacterial proteins, including UDP-N-acetylglucosamine acyltransferase (LpxA) and three other transferases of Escherichia coli, consist of a six-residue periodicity theme. FEMS Microbiol Lett 97:249–254. https://doi.org/10.1016/0378-1097(92)90344-n
Vijay-Kumar S, Bugg CE, Wilkinson KD, Cook WJ (1985) Three-dimensional structure of ubiquitin at 2.8 Å resolution. Proc Natl Acad Sci USA 82:3582–3585. https://doi.org/10.1073/pnas.82.11.3582
Vuorio R, Härkönen T, Tolvanen M, Vaara M (1994) The novel hexapeptide motif found in the acyltransferases LpxA and LpxD of lipid A biosynthesis is conserved in various bacteria. FEBS Lett 337:289–292. https://doi.org/10.1016/0014-5793(94)80211-4
Wheeler TJ, Clements J, Finn RD (2014) Skylign: a tool for creating informative, interactive logos representing sequence alignments and profile hidden Markov models. BMC Bioinform 15:7. https://doi.org/10.1186/1471-2105-15-7
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(suppl_2):W407–W410. https://doi.org/10.1093/nar/gkm290
Willard L, Ranjan A, Zhang H, Monzavi H, Boyko RF, Sykes BD, Wishart DS (2003) VADAR: a web server for quantitative evaluation of protein structure quality. Nucleic Acids Res 31:3316–3319. https://doi.org/10.1093/nar/gkg565
Wirtz M, Hell R (2006) Functional analysis of the cysteine synthase protein complex from plants: structural, biochemical and regulatory properties. J Plant Physiol 163:273–286. https://doi.org/10.1016/j.jplph.2005.11.013
Wirtz M, Berkowitz O, Droux M, Hell R (2001) The cysteine synthase complex from plants. Mitochondrial serine acetyltransferase from Arabidopsis thaliana carries a bifunctional domain for catalysis and protein-protein interaction. Eur J Biochem 268:686–693. https://doi.org/10.1046/j.1432-1327.2001.01920.x
Yi H, Galant A, Ravilious GE, Preuss ML, Jez JM (2010) Sensing sulphur conditions: simple to complex protein regulatory mechanisms in plant thiol metabolism. Mol Plant 3:269–279. https://doi.org/10.1093/mp/ssp112
Zhang C, Freddolino PL, Zhang Y (2017) COFACTOR: improved protein function prediction by combining structure, sequence, and protein-protein interaction information. Nucleic Acids Res 45:W291–W299. https://doi.org/10.1093/nar/gkx366
Acknowledgements
Surbhi Kharwar wants to thank University Grants Commission (UGC), New Delhi, India, for financial grant and also grateful to Dr. Vinay Kumar Singh, Information Officer, School of Biotechnology, Banaras Hindu University, Varanasi, India for providing molecular docking facility by Biovia Discovery Studio 2019. Samujjal Bhattacharjee is thankful to Council of Scientific and Industrial Research (CSIR), New Delhi, for awarding Junior Research Fellowship. The authors would also like to take this opportunity to thank Head, Department of Botany, Banaras Hindu University for providing the necessary facilities and encouragement to carry out this work.
Author information
Authors and Affiliations
Contributions
SK and AKM conceived the idea and designed the experiments. SK conducted the experiments and wrote the manuscript. SK, SB, and AKM critically review the manuscript. The manuscript was approved by all the authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
All authors participant agree to publish this work in your esteem Journal: 3 Biotech.
Consent to publish
Yes, all authors have given their consent.
Supplementary Information
Below is the link to the electronic supplementary material.
13205_2021_2899_MOESM3_ESM.pdf
Supplementary file3 Fig. S1. Hidden Markov Model (HMM) logos of cyanobacterial SAT (a) and OAS-TL (b) proteins. Logos were built with Skylign based on the alignment of protein sequences. The height of each amino acid code for each position reflects the weighted probability of occurrence at that position. The first amino acids Val, Leu, or Ile, followed often by Gly in the hexapeptide repeats with a consensus sequence of {V/L/I}-G-X-X-X-X, are marked with red boxes in the classical SAT proteins sequences. The blue box indicates the PLP-binding residues in the protein sequences. The SAT binding sites were indicated by pink represent OAS binding sites. Fig. S2. Structural verification by VADAR analysis of SAT (a) and OAS-TL (b) proteins showing functional accessible surface area, functional residues volume, stero/packing quality index, and 3D profile quality index. Fig. S3. Interacting sites of the SAT and OAS-TL proteins. (PDF 3189 kb)
Rights and permissions
About this article
Cite this article
Kharwar, S., Bhattacharjee, S. & Mishra, A.K. Bioinformatics analysis of enzymes involved in cysteine biosynthesis: first evidence for the formation of cysteine synthase complex in cyanobacteria. 3 Biotech 11, 354 (2021). https://doi.org/10.1007/s13205-021-02899-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-021-02899-1