Abstract
Purpose
4-hydroxycyclophosphamide (4HCY) is the principal precursor to the cytotoxic metabolite of cyclophosphamide (CY), which is often used as first-line treatment of children with cancer. There is conflicting data regarding the relationship between CY efficacy, toxicity, and pharmacokinetics with the genes encoding proteins involved in 4HCY pharmacokinetics, specifically its formation and elimination.
Methods
We evaluated germline pharmacogenetics in children with various malignancies receiving their first CY dose. Using linear regression, we analyzed the associations between two pharmacokinetic outcomes – how fast a child cleared CY (i.e., CY clearance) and the ratio of the 4HCY/CY exposure, specifically area under the plasma concentration-time curve (AUC), and 372 single nucleotide polymorphisms (SNP) in 14 drug-metabolizing transporters or enzymes involved in 4HCY formation or elimination.
Results
Age was associated with the ratio of 4HCY/CY AUC (P = 0.004); Chemotherapy regimen was associated with CY clearance (P = 0.003). No SNPs were associated with CY clearance or the ratio of 4HCY/CY AUC after controlling for a false discovery rate.
Conclusion
Age and chemotherapy regimen, but not germline pharmacogenomics, were associated with CY clearance or the ratio of 4HCY/CY AUC. Other methods, such as metabolomics or lipidomics, should be explored.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The prodrug cyclophosphamide (CY) is widely used to treat newly diagnosed pediatric cancer patients.[1] There is substantive interpatient variability in the efficacy and toxicity of CY-based combination chemotherapy regimens. This variability in clinical outcomes may be partially attributed to the pharmacokinetic variability of CY and its metabolites (Supplemental Fig. S1). CY is metabolized to 4-hydroxycyclophosphamide (4HCY), which is the precursor to CY’s primary cytotoxic metabolite phosphoramide mustard.[2] An understanding of 4HCY pharmacokinetics is critical because 4HCY, in turn, forms phosphoramide mustard, which covalently cross-links DNA. Because phosphoramide mustard does not cross cell membranes easily, 4HCY in the plasma transporting into the cell is a key step of CY’s cytotoxic activity via covalent cross-linking of DNA.[3].
The efficacy of CY relies upon its conversion to 4HCY by the cytochrome P450 (CYP) isozymes and myeloperoxidase (MPO). This conversion is mediated by numerous CYPs, specifically CYP2B6, CYP2C9, CYP2C19, CYP3A4, and CYP3A5.[3, 1] Exposure to 4HCY is further mediated by 4HCY elimination by glutathione S-transferases (GSTs), aldehyde dehydrogenases (ALDHs), and ATP-binding cassette (ABC) transporters.[3] As such, previous studies have focused on the pharmacokinetic and pharmacogenetic nature of the 4HCY formation and clearance in predominantly adult cancer patient populations, as we previously reviewed [4, 5].
Enthusiasm has been increasing for the potential of germline pharmacogenomics to enhance care and reduce disparities in cancer care. However, both non-genetic and genetic factors can influence the pharmacokinetics of CY and its metabolites (Supplemental Fig. S1). Furthermore, the pharmacogenetic associations between clinical outcomes and genes expected to affect 4HCY pharmacokinetics may differ with age and CY dose.[1] The contribution of the CYP enzymes involved in 4HCY formation varies with CY concentration. When children reach adulthood, levels of enzyme activity vary depending on the enzyme; children between the ages of 1–14 years have greater hepatic CYP2B6, CYP2C9, CYP2C19, CYP3A4/5, and GST activity (normalized for body weight).[6, 7] In this study, we evaluated the ratio of 4HCY/CY AUC (i.e., 4HCY formation clearance) with genetic polymorphisms, specifically those associated with 4HCY formation or 4HCY elimination. We hypothesized that non-genetic factors and genetic polymorphisms in 4HCY-pharmacokinetic enzymes are associated with CY clearance or the ratio of 4HCY/CY AUC in children.
Materials & methods
Patient population
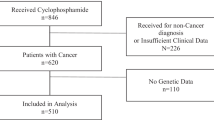
This was an ancillary prospective study in children who received cyclophosphamide from June 2006 to January 2013 under the aegis of a protocol approved by the Seattle Children’s Hospital Institutional Review Board. The inclusion criterion was: < 18 years of age at the time of enrollment; scheduled to receive IV cyclophosphamide (any dose, any number of doses, any dosing frequency) as their first course of therapy. The exclusion criteria were: body weight less than 5 kg (kg); patients who received prior chemotherapy (excluding corticosteroids).
Demographic data were taken from the participants’ medical charts [age, sex, height, body surface area, and clinical information (disease, chemotherapy regimen)]. The regimen administered was not affected by participation in this study. All participants received similar antiemetics, antibiotics, and antifungals per institutional standard practice guidelines; co-administration of CYP3A inhibitors was avoided when feasible. Two children received fluconazole concomitant with CY.
All participants’ parents gave informed consent before participation, as well as written assent for children 14–17 years.
Pharmacogenomic
We analyzed the associations between CY clearance and 4HCY formation clearance and the single nucleotide polymorphisms (SNPs) in 14 drug-metabolizing- enzymes or transporters involved in 4HCY disposition. The genes of interest were: cytochrome P450 (CYP) 2B6, CYP2C9, CYP2C19, CYP3A4, CYP3A5, Myeloperoxidase (MPO), aldehyde dehydrogenase (ALDH) 1A1, ALDH3A1, glutathione S-transferase (GST) A1, GSTM1, GSTP1, GSTT1, ATP binding cassette (ABC) ABCC2, ABCC3, and ABCC4 (Supplemental Table 1). We interrogated the ABC genes based on the discovery of ABCC2 contributing to 4HCY elimination in the rat model[8] and the paucity of data evaluating the SNPs potentially influencing the activity of these transporters (Supplemental Table S1).
From a 10 mL whole blood sample, total genomic DNA was isolated, according to the manufacturer’s instructions, using the Gentra Purgene® DNA Purification Kit (Gentra Systems, Minneapolis, MN). DNA was genotyped using Illumina’s GoldenGate Genotyping assay on the VeraCode Platform (Illumina, San Diego, CA) following manufacturer recommendation (VeraCode Assay Guide 11,312,819 rev A1).[9] 250 ng of genomic DNA was aliquoted into 96-well plates, processed accordingly, and scanned on the BeadXpress reader using GenomeStudio software (v2011.1). To ensure quality control (QC) of the data, 90 samples representing 30 parent-parent-child Centre d’Étude du Polymorphisme Humain (CEPH) trios (Utah residents with ancestry from northern and western Europe)[10] were genotyped to assess the performance of the Illumina oligonucleotide pool assay (OPA) before analysis of study samples. Assay accuracy was verified by comparing genotypes to publicly available genotype data for these samples from HapMap (http://www.hapmap.org/), 1000Genomes (http://www.1000genomes.org), dbSNP (http://www.ncbi.nlm.nih.gov/projects/SNP/), and by assessing inheritance errors. Two external control samples from the HapMap project were included on each plate to confirm the reliability and reproducibility of the genotyping across the study plates. Intra- and inter-plate duplicates comprised > 10% of all samples. Laboratory personnel were blinded to all research information about the samples. Other QC procedures included the use of barcodes on samples and plates, dedicated materials and working space, and a visual review of SNP cluster plots by two laboratory staff members. Samples with weak signals, discordant duplicates, and outliers were repeated at least once.
Any SNPs with less than 90% concordance with public data for CEPH controls were considered failed and excluded from the analysis. Among successfully genotyped SNPs, sample and SNP call rates were > 85%. All non-ancestry SNPs were in Hardy-Weinberg equilibrium (p > 0.001).
Of the 394 genetic polymorphisms evaluated, 372 passed quality control and were included in the analyses. Of the 81 children enrolled in this study, 72 had pharmacogenomic data available.
Pharmacokinetic analysis of CY and metabolites
CY clearance and the ratio of 4HCY/CY AUC (i.e., 4HCY formation clearance) were the pharmacokinetic outcomes of interest. Pharmacokinetic samples were drawn immediately before CY administration, upon completion of the 30-minute infusion, and at 2.5, 4.5, 6.5, 22.5, and 24 h after the start of the first CY dose. Sample processing and quantitation of CY and metabolites (i.e., carboxyethylphosphoramide mustard (CEPM), dechlorocyclophosphamide (DCCY) and ketocyclophosphamide (ketoCY)) were analyzed separately from 4HCY, using previously published methods [2].
A non-compartmental model was used to calculate the AUC from time 0 to infinity after the first CY dose of that chemotherapy cycle using WinNonlin Version 2.0 (Pharsight, Mountain View, CA). Of the 72 children with pharmacogenomic data available, CY clearance could be estimated in 57 children, and the ratio of 4HCY/CY AUC could be estimated for 49 children. CY clearance or the ratio of 4HCY/CY AUC could not be estimated in a small proportion of children because the concentrations were below the limit of quantitation to accurately estimate the CY AUC (which was used to estimate both pharmacokinetic outcomes) or 4HCY AUC (which was used to estimate the ratio of 4HCY/CY AUC).
Statistical analysis
Both pharmacokinetic outcomes, specifically the ratio of 4HCY/CY AUC and CY clearance, were log-transformed prior to analysis to better approximate a normal distribution. Because age and chemotherapy regimen may affect CY pharmacokinetics (Supplemental Figs. S1),[3] we first evaluated the associations between these clinical variables and each pharmacokinetic outcome. We then used linear regression to evaluate the association between these pharmacokinetic outcomes and each SNP separately, adjusted for age and chemotherapy regimen. Our analysis did not assume that specific genotypes would affect the pharmacokinetic outcomes in any particular direction or by any joint effects; instead, we took an unbiased approach of testing across the genotypes (two or three, depending on the number of variants in our population) for each of the 372 SNPs. Variants expressed in less than 5 participants (i.e., < 5 observations) were combined with the other less common variant for SNPs with three genotypes. SNPs with only two genotypes were dropped if the variant had < 5 observations. The Benjamini-Hochberg procedure was used to control the false discovery rate (FDR < 0.05).[11] All analyses were conducted in Stata (v18, StataCorp, College Station, TX).
Results
The participant characteristics by chemotherapy regimen are described in Supplemental Tables 2 and by chemotherapy regimen in Supplemental Table S3. The median age was 5.7 years (range: 0.59–17.5); there were 39 male and 42 female participants. Children enrolled in this study were diagnosed with acute lymphoblastic leukemia, Ewing sarcoma, rhabdomyosarcoma, lymphoma, neuroblastoma, or other malignancies. The majority of children received CY doses of 1000 mg/m2 or higher.
CY clearance was not associated with age (p = 0.49, Fig. 1A), but it did differ by chemotherapy regimen (p < 0.0031, Fig. 1B), with participants treated with the VAC regimen having the greatest CY clearance. The ratio of 4HCY/CY AUC was significantly associated with age such that each 1-year increase in age was associated with a 4% reduction in the ratio of 4HCY/CY AUC (P = 0.0041, Fig. 1C). The ratio of 4HCY/CY AUC was not associated with the chemotherapy regimen (p = 0.80, Fig. 1D).
Relationship of CY clearance (L/m2/h) with age (A) and chemotherapy regimen (B), and Ratio of 4HCY/CY AUC with age (C) and chemotherapy regimen (D). The ratio of HCY/CY AUC was associated with age such that there was a 4% reduction with each one-year increase in age. CY clearance was associated with the chemotherapy regimen
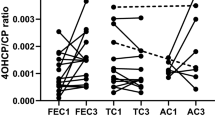
One child had two CY pharmacokinetic assessments, with separate informed consent being obtained at each time. Their first set of CY pharmacokinetic outcomes was used in the pharmacogenomics analysis. Appreciable within-patient variability was observed with their CY pharmacokinetics, which was assessed with their first CY dose of the topotecan/CY regimen at age 9 months and their first CY dose with CY/doxorubicin/vincristine at age 12 months (i.e., 12 weeks later). The concomitant chemotherapy drugs are presumed not to affect CY pharmacokinetics since topotecan, doxorubicin, and vincristine did not affect 4hydroxylation of CY in human liver microsomes.[1] Their CY clearance went from 6.26 to 7.49 L/m2/hour, and their ratio of 4HCY/CY AUC went from 0.126 to 0.199, respectively.
Table 1 shows the SNP associations with CY clearance (L/m2/hr) and the ratio of 4HCY/CY AUC. Of the 372 SNPs evaluated, 336 had at least one polymorphic allele in addition to the wild type in this cohort. Twenty-one SNPs were nominally significant at P < 0.05 for CY clearance, and sixteen were nominally significant at P < 0.05 for the ratio of 4HCY/CY AUC. All SNPs with allele frequencies included in the analysis are provided in Supplemental Table S4. Supplemental Table S5 contains a full list of the SNPs evaluated with their associated p-values and FDR. The majority of these SNPs were variants in the ABC genes. Most of the associations were inverse, with a slower CY clearance and a lower ratio of 4HCY/CY AUC among those with a genetic variant compared to those with the wild-type gene. However, no SNPs remained statistically significant after controlling for multiple testing (FDR < 0.05).
Discussion
In this exploratory analysis, we evaluated non-genetic and genetic factors associated with CY clearance and the ratio of 4HCY/CY AUC. Age and chemotherapy regimen did influence the ratio of 4HCY/CY AUC and CY clearance, respectively. However, among the 372 germline SNPs in 4HCY pharmacokinetic enzymes test, none were significantly associated with CY clearance or the ratio of 4HCY/CY AUC in children. The pro-drug CY has a complex metabolic schema, with a wide range of parent drug AUC and even greater variability in the AUCs of its metabolites, such as acrolein and phosphoramide mustard.[3] After administration, the majority of CY is metabolized to 4HCY (~ 75%) or DCCY (~ 5%); a significant portion is eliminated unchanged in the urine (~ 20%). An abbreviated summary of the drug-metabolizing enzymes or transporters involved in the pharmacokinetics of CY and its metabolites is shown in Supplemental Fig. S1; a more comprehensive summary is located in Supplemental Table S1. The non-genetic factors affecting CY clearance and 4HCY metabolism or elimination in pharmacokinetic studies of children are also shown in Supplemental Fig. 1. Cyclophosphamide induces cytochrome P450[12] enzyme activity, which is the putative mechanism of CY clearance being slower[13,14,15] and lower 4HCY exposure after the first CY dose[13] compared to subsequent days of CY administration. Older children, aged 3–19 years, have a slower CY clearance[15] than children aged < 2 years; however, within children < 2 years of age, body weight and age are not associated with CY clearance.[16] Specifically, 49 older (aged 3–19 years) children receiving CY showed an average CY clearance of 30.5 ± 17.8 mL/min/m2 [15] compared to a faster CY clearance of 46.6 ± 17.8 mL/min/m2 in 25 children aged < 2 years receiving CY 100–1500 mg/m2 (5–75 mg/kg).[16] Younger infants with medulloblastoma exhibited higher exposures to 4HCY than older infants.[17] However, age was not associated with 4HCY formation or elimination in children with neuroblastoma.[13] These suggest that the diagnosis or the appropriate treatments before CY administration may influence CY and its metabolite’s pharmacokinetics. Compared to children diagnosed with cancer, children diagnosed with Fanconi anemia have a slower CY clearance and no apparent metabolite formation.[18] Hepatic[14] and renal function[13] have also been associated with CY clearance and 4HCY disposition, respectively, in children. Thus, a significant number of non-genetic factors affect CY and its metabolites’ pharmacokinetics; these non-genetic factors may have obscured the influence of genetic polymorphisms on CY and its metabolites’ pharmacokinetics.
Early adult data suggested that efficacy, safety, or pharmacokinetics were associated with germline pharmacogenomics. As summarized in Supplemental Table S1, the majority focused on GST polymorphisms (children) or genetic polymorphisms of the CYP enzymes relevant to 4HCY formation (adults). Our analysis suggests that no promising germline SNPs are associated with CY clearance or the ratio of 4HCY/CY AUC. Our data agree with two Children’s Oncology Group studies showing that these genotypes were not associated with induction response to topotecan/CY in children diagnosed with high-risk neuroblastoma, [19] or 3-year event-free survival in children with intermediate-risk rhabdomyosarcoma treated with VAC.[4] These negative genotype-phenotype associations indicate that germline pharmacogenetic testing is likely not viable for personalizing chemotherapy in patients treated with CY-based regimens.
There are limitations to these data. Given the relative rarity of pediatric cancer, participant numbers for this trial were limited, which likely hampered our ability to detect meaningful differences in the pharmacokinetic endpoints – CY clearance and the ratio of 4HCY/CY AUC – between genotypic groups. Further, evaluating the pharmacogenomic association was not feasible because of insufficient variation in some SNPs. Drug-drug interactions between CY and other chemotherapy or supportive care agents may confound 4HCY formation. However, of the concomitant drugs administered to these participants, pharmacologically relevant concentrations of D-actinomycin, cytarabine, doxorubicin, 6-mercaptopurine, topotecan, and vincristine did not reduce 4HCY formation in CYP2B6 Supersomes® for pooled human liver microsomes.[1] Furthermore, the supportive care drugs were prescribed based on institutional standard practice. Thus, the possibility of drug-drug interactions was minimized as much as possible.
Given that numerous non-genetic factors may influence CY pharmacokinetics, evaluating if CY pharmacokinetics are associated with the metabolome or lipidome should be explored.[20, 21] The development of robust metabolomic platforms can facilitate understanding of the in vitro and in vivo actions of drugs.[20] The metabolome’s concentrations represent sensitive downstream markers of genomic changes and cells’ and tissues’ responses to external stimuli.[20] Metabolomics and lipidomics may enable a deeper mechanistic understanding of the pharmacokinetic disposition, drug–target interactions, and systems physiology from the molecular (genomic, proteomic, metabolomic) to cellular to whole–body levels.
In summary, after controlling for multiple testing, we did not find a significant association between CY pharmacokinetics and germline SNPs that encode an enzyme or transporter involved in 4HCY formation or 4HCY elimination However, the child’s age and the chemotherapy regimen were associated with CY pharmacokinetics.
Data availability
No datasets were generated or analysed during the current study.
References
1. Raccor BS, Claessens AJ, Dinh JC, Park JR, Hawkins DS, Thomas SS, Makar KW, McCune JS, Totah RA (2012) Potential contribution of cytochrome P450 2B6 to hepatic 4-hydroxycyclophosphamide formation in vitro and in vivo. Drug Metab Dispos 40 (1):54–63. doi:10.1124/dmd.111.039347
2. Kalhorn TF, Howald WN, Cole S, Phillips B, Wang J, Slattery JT, McCune JS (2006) Rapid quantitation of cyclophosphamide metabolites in plasma by liquid chromatography-mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci 835 (1–2):105–113
3. de Jonge ME, Huitema AD, Rodenhuis S, Beijnen JH (2005) Clinical pharmacokinetics of cyclophosphamide. Clin Pharmacokinet 44 (11):1135–1164
4. Pinto N, Navarro SL, Rimorin C, Wurscher M, Hawkins DS, McCune JS (2021) Pharmacogenomic associations of cyclophosphamide pharmacokinetic candidate genes with event-free survival in intermediate-risk rhabdomyosarcoma: A report from the Children’s Oncology Group. Pediatr Blood Cancer 68 (11):e29203. doi: 29210.21002/pbc.29203. doi:10.1002/pbc.29203
5. Pinto N, Ludeman SM, Dolan ME (2009) Drug focus: Pharmacogenetic studies related to cyclophosphamide-based therapy. Pharmacogenomics 10 (12):1897–1903. doi:10.2217/pgs.09.134
6. Hines RN, McCarver DG (2002) The ontogeny of human drug-metabolizing enzymes: phase I oxidative enzymes. J Pharmacol Exp Ther 300 (2):355–360
7. McCarver DG, Hines RN (2002) The ontogeny of human drug-metabolizing enzymes: phase II conjugation enzymes and regulatory mechanisms. J Pharmacol Exp Ther 300 (2):361–366
8. Qiu R, Kalhorn TF, Slattery JT (2004) ABCC2-mediated biliary transport of 4-glutathionylcyclophosphamide and its contribution to elimination of 4-hydroxycyclophosphamide in rat. J Pharmacol Exp Ther 308 (3):1204–1212
9. Shen R, Fan JB, Campbell D, Chang W, Chen J, Doucet D, Yeakley J, Bibikova M, Wickham Garcia E, McBride C, Steemers F, Garcia F, Kermani BG, Gunderson K, Oliphant A (2005) High-throughput SNP genotyping on universal bead arrays. Mutat Res 573 (1–2):70–82. doi:10.1016/j.mrfmmm.2004.07.022
10. Thorisson GA, Smith AV, Krishnan L, Stein LD (2005) The International HapMap Project Web site. Genome Res 15 (11):1592–1593. doi:10.1101/gr.4413105
11. Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate - a practical and powerful approach to multiple testing. J Royal Statist Soc Serial B 57 (1):289–300
12. Lindley C, Hamilton G, McCune JS, Faucette S, Shord SS, Hawke RL, Wang H, Gilbert D, Jolley S, Yan B, LeCluyse EL (2002) The effect of cyclophosphamide with and without dexamethasone on cytochrome P450 3A4 and 2B6 in human hepatocytes. Drug Metab Dispos 30 (7):814–822
13. McCune JS, Salinger DH, Vicini P, Oglesby C, Blough DK, Park JR (2009) Population pharmacokinetics of cyclophosphamide and metabolites in children with neuroblastoma: a report from the Children’s Oncology Group. J Clin Pharmacol 49 (1):88–102
14. Chinnaswamy G, Errington J, Foot A, Boddy AV, Veal GJ, Cole M (2011) Pharmacokinetics of cyclophosphamide and its metabolites in paediatric patients receiving high-dose myeloablative therapy. Eur J Cancer 47 (10):1556–1563. doi:10.1016/j.ejca.2011.03.008
15. Veal GJ, Cole M, Chinnaswamy G, Sludden J, Jamieson D, Errington J, Malik G, Hill CR, Chamberlain T, Boddy AV (2016) Cyclophosphamide pharmacokinetics and pharmacogenetics in children with B-cell non-Hodgkin’s lymphoma. Eur J Cancer 55:56–64. doi:10.1016/j.ejca.2015.12.007
16. Barnett S, Errington J, Sludden J, Jamieson D, Poinsignon V, Paci A, Veal GJ (2021) Pharmacokinetics and Pharmacogenetics of Cyclophosphamide in a Neonate and Infant Childhood Cancer Patient Population. Pharmaceuticals (Basel) 14 (3). doi:10.3390/ph14030272
17. Campagne O, Zhong B, Nair S, Lin T, Huang J, Onar-Thomas A, Robinson G, Gajjar A, Stewart CF (2020) Exposure-Toxicity Association of Cyclophosphamide and Its Metabolites in Infants and Young Children with Primary Brain Tumors: Implications for Dosing. Clin Cancer Res 26 (7):1563–1573. doi:10.1158/1078-0432.CCR-19-2685
18. Yule SM, Price L, Cole M, Pearson AD, Boddy AV (1999) Cyclophosphamide metabolism in children with Fanconi’s anaemia. Bone Marrow Transplant 24 (2):123–128. doi:10.1038/sj.bmt.1701868
19. Children’s Oncology Group. Comparing Two Different Myeloablation Therapies in Treating Young Patients Who Are Undergoing a Stem Cell Transplant for High-Risk Neuroblastoma. https://classic.clinicaltrials.gov/ct2/show/results/NCT00567567. Accessed February 18 2024
20. McCune JS, Nakamura R, O’Meally D, Randolph TW, Sandmaier BM, Karolak A, Hockenbery D, Navarro SL (2022) Pharmacometabonomic association of cyclophosphamide 4-hydroxylation in hematopoietic cell transplant recipients. Clin Transl Sci 15 (5):1215–1224. doi:10.1111/cts.13239
21. Navarro SL, Zheng Z, Randolph TW, Nakamura R, Sandmaier BM, Hockenbery D, McCune JS (2022) Lipidomics of cyclophosphamide 4-hydroxylation in patients receiving post-transplant cyclophosphamide. Clin Transl Sci 15 (11):2772–2780. doi:10.1111/cts.13404
Acknowledgements
This work is supported in part by grants from the National Institutes of Health (R03CA178104; R21CA162059; R01GM129863, R01HL91744S1, M01-RR00037), Alex’s Lemonade Stand; Children’s Foundation; and Fraternal Order of Eagles - Auburn. We appreciate the children and their families who participated in this study. We also appreciate the contributions of their health care team. We also appreciate the contributions of the research staff involved in data generation and management, especially Meagan J. Bemer, MS, Karen W. Makar, PhD, Brian Phillips, BS, and Linda Risler, BS.
Author information
Authors and Affiliations
Contributions
SLN and JSM wrote the main manuscript text; JSM prepared the figures. All authors reviewed and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Navarro, S.L., Pinto, N., Hawkins, D.S. et al. Pharmacogenomic associations of cyclophosphamide pharmacokinetic candidate genes with 4hydroxycyclophosphamide formation in children with Cancer. Cancer Chemother Pharmacol (2024). https://doi.org/10.1007/s00280-024-04703-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00280-024-04703-2