Abstract
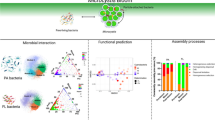
The estuarine system functions as natural filters due to its ability to facilitate material transformation, planktonic bacteria play a crucial role in the cycling of complex nutrients and pollutants within estuaries, and understanding the community composition and assembly therein is crucial for comprehending bacterial ecology within estuaries. Despite extensive investigations into the composition and community assembly of two bacterial fractions (free-living, FLB; particle-attached, PAB), the process by which bacterioplankton communities in these two habitats assemble in the nearshore and offshore zones of estuarine ecosystems remains poorly understood. In this study, we conducted sampling in the Yangtze River Estuary (YRE) to investigate potential variations in the composition and community assembly of FLB and PAB in nearshore and offshore regions. We collected 90 samples of surface, middle, and bottom water from 16 sampling stations and performed 16S rRNA gene amplicon analysis along with environmental factor measurements. The results unveiled that the nearshore communities demonstrated significantly greater species richness and Chao1 indices compared to the offshore communities. In contrast, the nearshore communities had lower values of Shannon and Simpson indices. When compared to the FLB, the PAB exhibit a higher level of biodiversity and abundance. However, no distinct alpha and beta diversity differences were observed between the bottom, middle, and surface water layers. The community assembly analysis indicated that nearshore communities are predominantly shaped by deterministic processes, particularly due to heterogeneous selection of PAB; In contrast, offshore communities are governed more by stochastic processes, largely due to homogenizing dispersal of FLB. Consequently, the findings of this study demonstrate that nearshore and PAB communities exhibit higher levels of species diversity, while stochastic and deterministic processes exert distinct influences on communities among near- and offshore regions. This study further sheds new light on our understanding of the mechanisms governing bacterial communities in estuarine ecosystems.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Estuaries are one of the most important marine ecosystems due to their role as the transitional zone between terrestrial rivers and the saline open sea. The high variability of land salinity and nutrient input make estuaries a terrigenous nutrient enriched and ecological sensitive coastal environment [1, 2]. Planktonic bacteria play a crucial role in the cycling of complex matter within estuaries; they transform particulate organic matter (POM) and dissolved organic matter (DOM), derived from ocean and rivers, into biomass that serves as food for microbial food webs and transfer energy and carbon to higher trophic levels [3, 4]. In shallow and turbid estuarine systems, bacterial communities undergo adaptation and specialization driven by sediment resuspension caused by currents, tides, and wind [5, 6]. The sinking of POM is responsible for transporting carbon and nutrients from the surface layer to the deep ocean [7]. During this sinking process, POM is often colonized and simultaneously decomposed by particle-attached (PA) microbes, leading to the release of DOM into the surrounding seawater, which serves as a fuel source for free-living (FL) microbes [8]. Recently, several studies have focused on investigating PA and FL bacterial (PAB and FLB) communities in marine [9,10,11,12], lacustrine [13, 14], and estuarine environments [15, 16]. Comparing PAB and FLB communities can yield valuable insights into the potential interactions between the two communities and the difference in ecological processes present within them [17, 18].
PAB and FLB communities exhibit differences in taxonomic composition, physiological metabolism, lifestyle, and ecological behavior [10, 19, 20]. For example, PAB are larger and occur in higher local concentrations than FLB in water [19]. Bacterial carbon production measurements have shown significantly higher activity for PAB compared to FLB in both freshwater and estuarine samples [20]; In terms of taxonomy, Proteobacteria were found to be the most abundant microbes in the PAB community, whereas Actinobacteria dominated the FLB community [10]. Due to their smaller size, FLB are more easily dispersed compared to PAB [13]. The ecological and oceanographic processes drive the response of ocean microbiomes to environmental changes [21]; there is a lack of knowledge regarding the community assembly processes of FLB and PAB communities in estuarine ecosystems, specifically between nearshore and offshore regions. In the community assembly theory, deterministic processes refer to environmental selection, including environmental filtering and interactions among species (competition, predation, and facilitation). Stochastic processes refer to dispersal and ecological drift, the dispersal means the movement of species across space, speciation is the generation of new genetic variation, and ecological drift represents random changes in species’ relative abundance over time due to the inherent [22]. The estuary is a transitional zone between land and ocean interactions, with significant changes in salinity and abundant habitats, making it a breeding ground for many marine fish; microbial activity strongly influences these organisms [23]. It is of both theoretical and practical significance to elucidate the community assembly of estuarine microorganisms.
The Yangtze River, also known as the Changjiang River, is the third longest river in the world and receives a substantial amount of nutrients from its basin [24]. Our previous study in Yangtze River Estuary (YRE) have shown that PAB community are less affected by environmental filtration, while homogeneous selection and drift were important processes in the FLB community assembly [16], However, geographical comparisons are lacking. In this study, 16 sampling stations were selected at the Yangtze River Estuary across near- and offshore regions, and water samples from the surface, middle, and bottom layers were collected. The differences in community structure and assembly between FLB and PAB in nearshore and offshore regions were determined using 16S rRNA gene amplicon sequencing and the neutral community model (NCM) analysis [25]. Community assembly is simultaneously influenced by factors that are relatively deterministic and factors that are more stochastic. The deterministic class includes selection imposed by the abiotic environment and both antagonistic and synergistic species interactions. The stochastic class includes unpredictable disturbance, probabilistic dispersal and random birth-death events [26]. This study aims to provide insights into (1) bacterial structures and diversity between the FLB and PAB communities and (2) bacterial community assembly process within the nearshore and offshore regions of the YRE.
Materials and Methods
Sample Collection and Environmental Factor Measurement
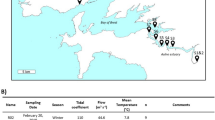
Seawater samples were collected from 16 sampling stations along the YRE in October 2020, using conductivity-temperature-depth sensors (Sea-Bird Electronics SBE 32) and Go-Flo® bottles (Fig. 1). Samples were collected from the surface, middle, and bottom of the water columns, with the middle samples missing from stations N0, N1, and M0 due to limited water depth (<20 m). To obtain particle-attached (PA) and free-living (FL), microbial cells were sequentially filtered using 3-μm and 0.22-μm membrane filters (Millipore Corporation, Billerica, MA, USA) under mild vacuum pressure (<33.3 kPa). All samples were immediately stored at −20 °C on board and subsequently transferred to −80 °C in the laboratory for DNA extraction. In total, 90 samples were collected and categorized into four groups: surface layer (~1-m depth), middle layer (20-m depth), bottom layer (water layer above sediment), nearshore (<50 m)/offshore (>50 m) and FL/PA fractions.
An overview of geographical location in the Yangtze River Estuary (dark circles). A total of 90 samples were collected from 16 sites in October 2020. The map was generated with the Ocean Data View (http://odv.awi.de); water depths are indicated with colored column on the right side. Stations belong to near- or offshore region were shown in light green and light red boxes, respectively
Environmental variables, including temperature, salinity, depth, pH, chlorophyll a (Chla), suspended solids (SS), total alkalinity (ALK), chemical oxygen demand (COD), and dissolved oxygen (DO), were measured on-site using a CTD system. ammonia nitrogen (NH4+), nitrite (NO2−), silicate (SiO44−) and phosphorus (PO43−) were analyzed using the SmartChem automatic nutrient analyzer (Smartchem 200, Alliance, France); nitrate (NO3−) was determined using the copper-cadmium reduction method following the procedure described by Sorte and Basak [27], All physical and chemical parameters adhere to the Marine Monitoring Specification (GB17378, 2007). To gain insights into the influence of water flow on bacterial communities, information on ocean currents at different depths (1 m, 20 m, and 50 m) was also collected using the CTD system.
DNA Extraction, PCR, and Sequencing
DNA from water samples was extracted using the PowerSoil DNA Isolation Kit (MO BIO, San Diego, CA, USA), following the kit instructions. DNA quality was assessed using 1% agarose gel electrophoresis and the Nanodrop 2000 (Thermo, Waltham, MA, USA). For bacterial 16S rRNA gene amplification, 515F and 806R primers were utilized. The reaction mixture consisted of 4 μL FastPfu buffer, 2 μL dNTPs (2.5 mM), 0.80 μL forward primers (5 μM), 0.80 μL reverse primers (5 μM), 0.40 μL FastPfu polymerase, 0.20 μL bovine serum albumin (BSA), 10 ng DNA, and double-distilled H2O, with a total volume of 20 μL. Amplification reactions were conducted using an ABI GeneAmp® 9700 PCR instrument (ABI, Waltham, MA, USA). The PCR products were used to construct paired-end libraries following the manufacturer's instructions. High-throughput sequencing was performed by Majorbio Bio-Pharm Technology Co., Ltd. (Shanghai, China) on the Illumina MiSeq platform.
Amplicon Analysis
Raw reads were processed and analyzed using the QIIME2 platform [28]. Primer excision and quality control were performed using VSEARCH [29]. ASV (amplicon sequence variant) clustering was conducted by calling unoise3 in USEARCH [29], Taxonomy annotations for each ASV were performed using the RDP classifier [30] against the Silva database (v138) [31]. ASVs annotated as “Chloroplast,” “Eukaryote,” “Archaea,” and “Mitochondria” were filtered out. ASVs table was generated by mapping primers-removed reads to the representative sequences of ASVs. To account for sequencing depth, the ASVs table was rarefied based on the lowest reads number (12,259) of all samples using the “rarefy” function of vegan package [32] (Fig. S1).
Statistical Analysis
All statistical analyses were performed using R v4.0 [33]. Bacterial diversity was calculated as the Shannon, Chao1, Simpson, and Richness indices using the picante package [34]. The statistically significant difference in alpha diversity between FLB and PAB was analyzed using the two-sided Wilcoxon signed-rank test from the ggsignif package [35] and plotted with the ggplot2 package [36]. A principal coordinate analysis (PCoA) analysis was conducted using the vegan package, the dissimilarity matrix was calculated by the Bray-Curtis dissimilarity method, R2 and p value was also calculated, the R2 indicates the variance can be explained by geographical distance, the p value indicates the significance of the variance [37]. Full linkage hierarchical clustering based on Bray-Curtis dissimilarity was performed using the vegan package [32]. Non-metric multidimensional scaling (NMDS) analysis was also performed using the vegan and ggplot2 packages [38]. Mantel test analysis was conducted using the vegan package [39]. Heatmaps were generated using the pheatmap package to visualize the Spearman correlations between environmental factors and the top 30 phyla [40]. The vegan and randomForest packages were used to calculate and visualize the contribution of biomarker species among nearshore and offshore communities [41]. Venn diagram were generated using the UpSetR package [42].
Specificity-Occupancy Plots
The term “specificity” refers to the distribution of microbes exclusively within certain communities, while “occupancy” refers to the broad distribution range of microbes. In order to examine the distribution of ASVs across sampling sites within the same habitat and their specificity to that habitat, we calculated the occupancy and specificity of individual ASVs for each habitat and projected them onto maps [43]. The ASVs with a relative abundance greater than 1% were selected from the ASV table for each habitat, and from the table, specificity and occupancy were calculated in Jiang et.al [43]. In our study, specificity is defined by the number of individual ASVs (S) in the samples of a habitat (H) (Eq. 1), and occupancy is defined by the relative frequency of occurrence of S in the samples of H (Eq. 2).
N-individual S,H is the mean number of individual ASVs across all samples in habitat H, while N-individual S is the sum of the mean number of individual S over all habitats; N-sites S,H is the number of samples in H where S is present, while N-sites H is the total number of samples in H [44]. These two metrics were then used as the axes in the specificity-occupancy plots. To find specialist taxa attributable to each habitat type, ASVs with specificity and occupancy greater or equal to 0.7 was selected according to Gweon’s method [45].
Community Assembly Analysis
The Stegen’s null model based analyses [26] were conducted using the iCAMP package [46]. This analysis was based on the β-nearest taxon index (βNTI) and the Bray-Curtis-based Raup-Crick index (RCbray) [46]. Values of |βNTI| > 1.96 indicate determinism, which can be further classified as homogeneous selection (HoS, βNTI < −1.96) or heterogeneous selection (HeS, βNTI > 1.96). Conversely, values of |βNTI| ≤ 1.96 suggest stochasticity. Integrating the RCbray value, the community assembly processes can be categorized as homogenizing dispersal (HD, RCbray < −0.95), dispersal limitation (DL, RCbray > +0.95), or drift and others processes (with |RCbray| < 0.95) representing weak selection, weak dispersal, diversification, and drift processes. The null community was randomized 999 times to obtain average null expectations [47]. The neutral community model (NCM) was also fitted using the Hmisc package to assess the contributions of stochastic processes in shaping FL and PA communities [48].
Results
Overview of Water Properties
A total of 13 environmental factors including salinity, temperature, SS, DO, NO2−, NO3−, PO43−, NH4+, COD, SiO44−, pH, total alkalinity, and Chla were measured (Table S1). The values for these factors are as follows: salinity ranged from 21.96 to 34.50 g/L, SS ranged from 0.60 to 534.70 mg/L, temperature ranged from 13.95 to 20.81 °C, COD ranged from 0.02 to 2.81 mg/L, DO ranged from 6.78 to 8.51 mg/L, Chla ranged from 0.05 to 0.35 μg/L, NO2−ranged from 0.0004 to 0.004 mg/L, NH4+ ranged from 0.001 to 0.012 mg/L, NO3− ranged from 0.01 to 0.48 mg/L, PO43− ranged from 0.002 to 0.02 mg/L, SiO44− ranged from 0.06 to 1.49 mg/L, and pH ranged from 8.06 to 8.24. The alkalinity values were in the range of 21.96 to 34.50 g/L. Additionally, ocean currents were measured at different depths (1 m, 20 m, and 50 m) (Fig. S2). In the YRE, the water forms an eddy, and the velocity increases from the surface to the bottom, ranging from 0.1 to 0.2 m/s.
Community Structure and Diversity of FL and PA Bacteria
A total of 6,117,739 high-quality bacterial sequences were obtained from the 90 samples, revealing the presence of 51 phyla, 320 orders, and 758 genera in the seawater of the YRE. Among these taxa, phyla including Proteobacteria, SAR324, Actinobacteriota, Bacteroidota, Marinimicrobia, Dadabacteria, Chloroflexi, Gemmatimonadota, Nitrospinota, Cyanobacteria, and Verrucomicrobiota were shared across all samples, indicating their widespread presence. Additionally, Margulisbacteria was exclusively detected in offshore samples, while Desulfobacterota was only found in the nearshore region. The phylum PAUC34f exhibited distribution in both nearshore and offshore environments.
The distribution of microbial taxa in the surface, middle, and bottom layers showed similar patterns. Proteobacteria, Actinobacteriota, and Bacteroidota were consistently dominant phyla across all three layers, with Proteobacteria exhibiting particularly high relative abundance (57.33%) at station P. Other abundant taxa included SAR324, Chloroflexi, Marinimicrobia, Planctomycetota, Dadabacteria, Cyanobacteria, and Gemmatimonadota (Fig. S3). However, notable differences were observed between the FLB and PAB communities. For instance, Planctomycetota (7.56%) displaying a significant advantage in the PAB community, but only 0.14% in the FLB community (Fig. 2). On the other hand, Gemmatimonadota (0.03%), Nitrospinota (1.19%), and PAUC34f (0.31%) exhibited greater dominance in the FLB community compared to only 0.01%, 0.38%, and 0.03% in the PAB community (Fig. 2), Cyanobacteria (1.6%) and Verrucomicrobiota (2.1%) showed higher relative abundance in the PAB community, but only 0.15%, 0.61% in the FLB community. Furthermore, PAUC34f (0.31%) and Margulisbacteria (0.30%) were exclusively detected in the FLB communities, while Desulfobacterota (0.36%) was found solely in PAB communities. Proteobacteria and Actinobacteriota were the two most abundant phyla in both the FLB and PAB communities. Subtle variations between the FLB and PAB communities among sample stations were also observed. For instance, in the FLB communities, Actinobacteriota (77.54%) exhibited the highest relative abundance at the N0 site, while Planctomycetota (3.50%) dominated at the P2 site. In the PAB communities, Alphaproteobacteria (29.02%) displayed the highest relative abundance at the N3 site. Random forest analysis was employed to identify microbial biomarkers at the ASV level, ASVs belongs to Desulfobacterota, Myxococcota, Acidobacteriota, and Cyanobacteria are biomarkers for offshore communities, while Planctomycetota Firmicutes SAR324 and Marinimicrobia are characteristic phyla in nearshore regions (Fig. S4).
Dominant bacterioplankton composition and alpha diversity between FL and PA bacterial communities among near- and offshore zones. A ASVs with Relative abundance of sequences ≥ 2% was calculated, the top ten phyla for each sample were used, others represent the phyla not unassigned at the phylum level. B Richness, Chao1, Shannon, and Simpson index. p values of Tukey’s HSD (honestly significant difference) test between groups are indicated on the top of bar plots. ***p < 0.001
Significant differences on alpha diversity were observed between near- and offshore (p < 0.001, Fig. 2B). In comparison to the offshore communities, the nearshore communities exhibited significantly higher species richness and Chao1 indices, while the Shannon and Simpson indices were lower. The four diversity indices of PAB community were higher than FLB community, indicating greater diversity and richness of the PAB community (Fig. 2B). However, no significant differences (p > 0.05) were found between the surface layer, middle layer, and bottom layer (Fig. S5). These community differences were also supported by PCoA analysis. The nearshore communities showed distinct separation between the FAB and PAB communities (R2 = 0.217, p = 0.01) compared to the offshore communities (R2 = 0.14, p = 0.001) (Fig. 3). No significant differences in beta-diversity were found between the three layers (R2 = 0.20–0.065, p > 0.05). The NMDS analysis further confirmed these findings (Fig. S6).
Principal coordinate analysis (PCoA) of bacterioplankton communities based on Bray–Curtis dissimilarity. Communities within ellipses are associated with 95% confidence intervals. The red, green, pink, light green, and dark blue circles represent FLB (free-living bacterial communities), PAB (particle-attached bacterial communities), bottom, middle, and surface water, respectively. Samples within green and red boxes are bacterial communities of near- and offshore, respectively. R2 indicates the variance can be explained by geographical distance, the p value indicates the significance of the variance
Effects of Environmental Factors on Bacterial Communities
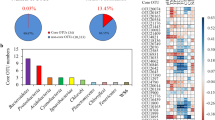
The Mantel test was conducted to analyze the influence of environmental factors on bacterial community composition (Table 1). For the nearshore communities, several environmental factors showed significant effects (p < 0.01), including COD, DO, SS, PO43−, SiO44−, and NO3−. Conversely, in offshore communities, significant effects were observed for DO, Chla, temperature, and salinity. pH, Alk, NO2−, and NH4+ had no discernible influence on both nearshore and offshore communities. In summary, the nearshore community was primarily influenced by COD, SS, PO43−, SiO44−, and NO3−, whereas offshore communities were most influenced by Chla, temperature, and salinity. Among these factors, Salinity (r = 0.21) exerted the strongest influence on the offshore communities, while SiO44− (r = 0.26) had the most pronounced impact on the nearshore communities. The correlation analysis between environmental factors and bacterial taxa at the phylum level was also conducted on different lifestyles (FLB and PAB) (Fig. S7). In addition to SS, other environmental factors have varying degrees of influence on bacterial taxa in FLB; moreover, Chla, temperature, DO, NO2−, and NH4+ present a significantly correlation (p < 0.05) with almost all taxa in FLB. However, there are no environmental factor shows universal and significant (p < 0.05) correlation with these taxa (Fig. S7B).
Occupancy and Specificity of ASVs in the Near- and Offshore Regions
To investigate the distribution of different ASVs across sampling stations within the same regions (nearshore and offshore) and their specificity to the regions, the occupancy and specificity of individual ASVs were calculated. To identify specialist taxa associated with each lifestyle and region, ASVs with a specificity and occupancy of 0.7 or greater (indicated by dotted boxes in Fig. 4A, B) were examined. specificity-occupancy analysis indicates that the occupancy rate of ASVs originating from nearshore exhibits greater variability, whereas the occupancy rate of ASVs originating from offshore remains relatively consistent (Fig. 4A, B). Since ASVs related to Cyanobacteria, Planctomycetota, Actinobacteriota, Verrucomicrobiota, Desulfobacterota and PAUC34f were mainly found in PAB, while ASVs of Marinimicrobia and SAR324 mainly lives in FLB, the higher occupancy of nearshore regions is attributed to the absence of Marinimicrobia, which enhances specificity (Fig. 4A, B). ASVs with a specificity and occupancy of 0.7 or greater of each lifestyle showed varying occupancy but were commonly found in most offshore sites (Fig. 4B). Furthermore, overall ASVs were more frequent and prevalent in offshore habitats (Fig. 4B). These differences in ASVs shared by nearshore and offshore were due to a combination of different occurrence frequencies and habitat selections of the ASVs. Proteobacteria, SAR324 (Marine group B), Bacteroidota, Dadabacteria, Chloroflexi, Bdellovibrionota, Nitrospinota, and Gemmatimonadota were found in all four specialist groups: FLB-nearshore, PAB-nearshore, FLB-offshore, PAB-offshore (Fig. 4D). Notably, Planctomycetota (7.56%) displaying a significant advantage in the PAB community, but only 0.14% in the FLB community (Fig. 2). On the other hand, Gemmatimonadota (0.03%), Nitrospinota (1.19%), and PAUC34f (0.31%) exhibited greater dominance in the FLB community compared to only 0.01%, 0.38%, and 0.03% in the PAB community (Fig. 2). Importantly, the result of the Venn diagram shows no specialists were identified between the nearshore and offshore communities (Fig. 4D).
Specificity-occupancy plots showing ASVs that different between the PA and FL in the nearshore and offshore water from the Yangtze River Estuary. A, B The x-axis represents occupancy, i.e., how well an ASV is distributed in each habitat across all sites; the y-axis represents specificity, i.e., whether ASVs are also found in other habitats. C Venn diagrams showing the numbers of unique and shared ASVs among the four bacterial communities. D Detailed bacterial phyla of unique and shared ASVs. Dotted box on the top right are ASVs having a spec-occ > 0.7
Microbial Community Assembly Mechanisms in Near- and Offshore Regions
The null model based analyses were employed to explore the relative contributions of deterministic and stochastic processes to community assembly in nearshore and offshore bacterial communities. βNTI confirmed the contribution of determinism community assembly processes in the nearshore communities and stochastic processes among offshore bacterial communities (Fig. S8). The large proportion of heterogeneous selection (50.44%) and homogeneous selection (19.59%) suggested that deterministic processes exerted greater influence on nearshore community (Fig. 5A). In turn, the relative high proportion of homogenizing dispersal (82.81%) in the offshore communities indicated that stochastic processes played greater role in governing community assembly (Fig. 5C). FLB communities were mainly influenced by heterogeneous selection, while PAB communities were mainly influenced by homogenizing dispersal and drift (Fig. 5B, D) across all regions. The NCM analysis successfully captured a significant portion of the relationship between the occurrence frequency of ASVs and their mean relative abundances (Fig. 6). For FLB communities, the NCM explained 47.30%, 87.20%, and 53.90% of the community variance in the nearshore, offshore, and both regions, respectively. Similarly, for PAB communities, the NCM explained 28.90%, 72.30%, and 55.20% of the community variance in the nearshore, offshore, and both regions, respectively. Moreover, the NCM analysis revealed a higher explained community variance in the offshore planktonic community (ranging from 72.30 to 87.20%, with a mean value of 79.75%) compared to the nearshore community (ranging from 28.90 to 47.30%, with a mean value of 38.10%). Additionally, the Nm-value, representing species dispersal, was higher for bacterial taxa in the offshore regions (Nm = 29,291–54,229) compared to the nearshore region (Nm = 300–34,306) (Fig. 6), which indicate that the dispersal of planktonic bacteria species is higher in offshore environments than in nearshore regions. Both the null model and NCM analysis suggested that the nearshore communities are shaped by deterministic processes and offshore communities are governed by stochastic processes.
Relative importance of different ecological processes between near- and offshore communities. A, B Nearshore. C, D Offshore. Values on the ring indicate the fraction of ecological processes (deterministic: homogeneous and heterogeneous selection; stochastic: dispersal limitations and homogenizing dispersal; drift and others).
Fit of the neutral community model (NCM) of community assembly. The predicted occurrence frequencies for nearshore, offshore, and all represent bacterial communities from nearshore (green colored), offshore (pink colored), and both region (white colored), respectively. The solid blue lines indicate the best fit to the NCM as in Sloan et al., and the dashed blue lines represent 95% confidence intervals around the model prediction. ASVs that occur more or less frequently than predicted by the NCM are shown in green or red colors. Nm represents the metacommunity size multiplied by immigration, and R2 represents the fit to the model
Discussion
The Yangtze River basin supports a dense population and well-developed industry and agriculture. The YRE, adjacent to the Zhoushan fishing grounds, serves as a traditional fishing area [49], also a representative estuarine ecosystem. Recent studies have extensively investigated the bacterial compositions and community characteristics in this ecosystem [16, 50,51,52]. Our study aimed to announce a comprehensive understanding of community assembly of the free-living and particle-attached bacterioplankton by surveying 16 sampling stations in the YRE and its adjacent waters. The results unveiled significant differences among nearshore, offshore, FLB and PAB communities, but no distinct differences were observed between the bottom, middle, and surface water layers (mainly α diversity and β diversity).
Distinct Microbial Communities Between Near- and Offshore Regions
Despite the alpha diversity being similar across different water depths, distinct microbial communities were observed between nearshore and offshore regions of the YRE. This can be attributed to the enrichment of heterotrophic bacteria resulting from nutrient-rich river inputs in the nearshore region, whereas the relatively oligotrophic offshore environment limits the growth of these microbes [53]. In contrast to previous research [54], no significant differences in diversity were observed among different layers in this study. The frequent mixing of water may be the primary factor contributing to this phenomenon, the in situ current pattern in the YRE region, including the south China Sea surface current, the China coastal current, and the Kuroshio branch current, contributes to water dynamics from the bottom to the surface (Fig. S2) [55], promoting efficient mixing of bacterioplankton in the water column, the high dissolved oxygen (DO) values (>6 mg/L, Table S1) also indicate the efficient mixing in the YRE during sampling [56].
The higher richness and diversity of bacterial species in PAB community has been observed in open ocean environments [57], the number of ASVs in the PAB community is also found significantly higher than that in the FLB community. The particulate organic matter (POM) serves as a nutrient source for PAB, and their higher capacity for rapid decomposition and utilization of organic carbon may explain this phenomenon [58, 59]. We found that Margulisbacteria was exclusively present in offshore samples, while Desulfobacterota was detected solely in the nearshore region (Fig. 2, Fig. 6D), suggesting potential environmental preferences. Margulisbacteria has been reported to specifically attach to spirochetes and thrive in ocean waters [60, 61], while Desulfobacterota prefers nutrient-rich environments [62] and not sensitive to contamination [63]. In consistent with previous studies [64], Planctomycetota shows a closer association with particle-attached (PA) samples (Fig. 2), This may be due to the preference of Planctomycetota microbes for an anaerobic environment, and the organic aggregates developed by PAB provide an anaerobic microenvironment [65]. Previous studies have revealed that Verrucomicrobia was widely distributed across the marine envrionment [66]; our results further demonstrated that Verrucomicrobia was more dominant in the nearshore community than in offshore. The random forest analysis also identified Verrucomicrobia as an important indicator taxon for the nearshore community (Fig. S4). Fluctuations in the abundance of Verrucomicrobia in particle-attached bacterial communities have been reported in several Arctic studies [67]; the preference of Verrucomicrobia for organic matter-rich marine environments [68] and the abundance of organic matter in nearshore waters due to terrigenous input [69] could be the underlying reason.
Environmental Factors Have Different Influences on Near- and Offshore Communities
The nearshore communities were primarily influenced by COD, SS, PO43−, SiO44−and NO3−, while offshore communities were mainly influenced by Chla, temperature, and salinity. Among these factors, salinity (r = 0.208) and Si (r = 0.259) exhibit the strongest influences on offshore and nearshore communities, respectively. Salinity gradient has been identified as a crucial impact factor on bacterioplankton in estuary ecosystems [70]. SS, carried by river inflow, supplies a large amount of nutrients, could be due to the enrichment of microbial communities in nearshore region, which has been frequently reported [71]. Further investigation is needed to determine the impacts of other environmental factors on nearshore and offshore bacterial communities. Numerous studies have shown significant correlations between PAB and FLB and environmental parameters [15, 72]. The distinction between these two communities is primarily attributed to the different microhabitats they inhabit in aquatic ecosystems. Attached bacteria reside on particle surfaces, while FLB float freely in the water column. Previous studies have indicated that FLB bacteria are more sensitive to environmental variables compared to PAB [15, 73]. In our study, the Chla, temperature, DO, NO2−, and NH4+ significantly influences (p < 0.05) the majority of phylum-level taxa in FLB, while only a limited number of taxa are affected by detected environmental factors, which also demonstrate that a higher proportion of free-living variation can be explained by environmental parameters (as shown in the heatmap, Fig. S7) compared to PAB. This can be attributed to the fact that PAB attached to particles act as “buffers” or micro-islands [73, 74], while FL cells are directly exposed to the surrounding water. Therefore, particle size plays a crucial role, as larger particles to which bacteria are attached make them less sensitive to changes in the surrounding water [73]. Additionally, the high bacterial density on particles facilitates efficient signaling and quorum sensing, quorum sensing signals, such as acylated homoserine lactones, which are required mainly when the population reaches high densities [75], and the nutrient ratio within particles may differ significantly from that in the surrounding seawater. These factors collectively explain why PAB are less influenced by changes in environmental parameters compared to FLB.
Within Habitat Specificity and Overlap Was Observed Across the Spatial Scale
Planktonic communities in dynamic water columns are generally considered to be “heterogeneous” in both space and time [76]. Despite our study did not take time variation into consideration, this heterogeneous was also observed across the spatial scale. Compared with the nearshore habitats, both FLB and PAB communities in offshore showed higher specificity (Fig. 4B) across the sampling stations, which indicate that the offshore environments are more selective for a large and stable core community. The constant mixing of water from terrestrial rivers and dynamic fluctuations in salinity and nutrient levels within estuarine ecosystems provide ample opportunities for microbial communities to exchange between diverse habitats. The presence of this phenomenon suggests that microorganisms in these aquatic ecosystems possess the capability to adapt different lifestyle in order to optimize nutrient acquisition from the surrounding environment [77]. As shown in the specificity-occupancy plots (Fig. 4), we found that the majority of the ASVs, both FLB and PAB, have high occupancy in YRE, which particularly evident in offshore communities, meaning that most of them are also found in all other sites across the estuarine ecosystem. This may because of the continuous input of fresh water and nutrient from Yangtze River make up a rapidly changing environment and make residents lives there present the rapid exchange among habitats. The mechanisms underlying the spatial homogeneity observed in both near- and offshore communities remain unclear at present. Our study did not explore the physical and chemical characteristics of the PAB and FLB themselves, thus further investigation is needed to determine whether these factors contribute to stabilizing nutrient levels in the overlying water column, thereby fostering similar community structures. Alternatively, it is possible that other factors associated with an attached lifestyle play a role.
Bacterial Community Assembly Process Influenced by Randomness
Homogeneous selection refers to stable environments where few phylogenetic shifts occur due to consistent abiotic and biotic conditions, whereas heterogeneous selection results from fluctuating environments causing higher phylogenetic diversity [26, 78]. Dispersal is categorized into homogenizing dispersal and dispersal limitation. High dispersal homogenizes communities leading to minor taxonomic variation; while limited dispersal increases taxonomic diversity. If neither dominates, “undominated” or “drift” governs community formation [26, 78]. Analyses based on null model suggested the deterministic and stochastic processes shaped the bacterial communities in nearshore and offshore region, respectively (Fig. 5A, C), which is consistent with the result of NCM analysis (Fig. 6). The results could be due to the relatively unrestricted dispersal is in offshore region [79], where hydrological mixing may increase dispersal-related processes and ecological drift [80, 81]. The NCM analysis explained community variance for PAB and FLB is 53.9% and 55.2%, respectively. This is consistent with the conclusion of our previous study [16], which indicates that both FLB and PAB communities are shaped by stochastic processes in the YRE region. However, a detailed analysis of the near- and offshore regions reveals distinct results. For FLB, 47.30% and 87.20% of the explained community variance is observed in the nearshore and offshore regions, respectively, whereas for PAB, 28.90% and 72.30% of the explained community variance is observed in the nearshore and offshore regions, respectively. This suggests that PAB community in the nearshore are impacted by a stronger effect of deterministic processes, while FLB community in the offshore region are significantly impacted by stochastic processes. Further evidences were provided by the results of null model analysis. Although, PAB in nearshore are predominantly governed by HeS, DR, and DL, compared with HeS (50.44%) and DR (4.17%), the HeS exerts the paramount influence, accounting for 44.14% of the total impact on PAB. Seminally, among the HD (56.93%), HeS (1.65%), and HoS (0.12%) in offshore region, the HeS strongly shapes the assembly of FLB (Fig. 5). The YRE is eutrophic [82], and the availability of nutrients for planktonic microbes makes it compatible with nutrient-rich substrates. PAB abundances are commonly high at the more locations along the nutrient-rich coastal zones [83], high nutrient levels in nearshore regions could be impose a strong selective pressure by selecting for copiotrophs; however, there is more randomness in more oligotrophic waters in offshore region because of a relaxed selective pressure. which may explain why the PAB community is shaped by deterministic processes in the nearshore region and FLB community is predominantly shape by stochastic processes in the offshore region. In this study, FLB in nearshore and offshore have distinct assembly pattern, and the same situation was also observed in PAB; further investigation is needed to determine this phenomenon.
Conclusion
The present investigation offers an extensive characterization of the PAB and FLB communities in the nearshore and offshore regions of the YRE, encompassing their biodiversity, community structure, the influence of environmental factors, and microbial community assembly mechanisms. Distinct from the FLB community, the PAB community exhibits a higher alpha diversity. In contrast to the offshore region, the nearshore communities display a significantly elevated species richness and Chao1 indices. Margulisbacteria was exclusively identified in offshore samples, whereas Desulfobacterota was solely detected in the nearshore area. Planctomycetota demonstrates a notable advantage within the PAB community, yet Nitrospinota exhibits greater predominance in the FLB community. The nearshore communities are significantly influenced by COD, DO, SS, PO43−, SiO44−, and NO3−, while the offshore communities are mainly affected by DO, Chla, temperature, and salinity. In the nearshore region, deterministic processes play a vital role in shaping bacterial community assembly, whereas the dispersal of planktonic bacteria species is more prominent in offshore regions.
Data Availability
All sequence data generated in this study have been submitted to the NCBI Sequence Read Archive under the accession number PRJNA877030.
References
Suzzi AL, Gaston TF, McKenzie L, Mazumder D, Huggett MJ (2022) Tracking the impacts of nutrient inputs on estuary ecosystem function. Sci Total Environ 811:152405. https://doi.org/10.1016/j.scitotenv.2021.152405
Santos M, Amorim A, Brotas V, Cruz JPC, Palma C, Borges C, Favareto LR, Veloso V, Damaso-Rodrigues ML, Chainho P, Felix PM, Brito AC (2022) Spatio-temporal dynamics of phytoplankton community in a well-mixed temperate estuary (Sado Estuary, Portugal). Sci Rep 12:16423. https://doi.org/10.1038/s41598-022-20792-6
Thibault A, Derenne S, Parlanti E, Anquetil C, Sourzac M, Budzinski H, Fuster L, Laverman A, Roose-Amsaleg C, Viollier E (2019) Dynamics of organic matter in the Seine Estuary (France): bulk and structural approaches. Mar Chem 212:108–119
He C, Pan Q, Li P, Xie W, He D, Zhang C, Shi Q (2019) Molecular composition and spatial distribution of dissolved organic matter (DOM) in the Pearl River Estuary, China. Environ Chem 17:240–251
Manning A, Langston W, Jonas P (2010) A review of sediment dynamics in the Severn Estuary: influence of flocculation. Mar Pollut Bull 61:37–51. https://doi.org/10.1016/j.marpolbul.2009.12.012
Briones EE (2004) Current knowledge of benthic communities in the Gulf of Mexico. In: Withers K, Nippers M (eds) Environmental Analysis of the Gulf of Mexico, pp 108–136
Mari X, Passow U, Migon C, Burd AB, Legendre L (2017) Transparent exopolymer particles: effects on carbon cycling in the ocean. Prog Oceanogr 151:13–37. https://doi.org/10.3390/gels7030075
Azam F, Malfatti F (2007) Microbial structuring of marine ecosystems. Nat Rev Microbiol 5:782–791. https://doi.org/10.1038/nrmicro1747
Li J, Gu L, Bai S, Wang J, Su L, Wei B, Zhang L, Fang J (2021) Characterization of particle-associated and free-living bacterial and archaeal communities along the water columns of the South China Sea. Biogeosciences 18:113–133. https://doi.org/10.5194/bg-18-113-2021
Suzuki S, Kaneko R, Kodama T, Hashihama F, Suwa S, Tanita I, Furuya K, Hamasaki K (2017) Comparison of community structures between particle-associated and free-living prokaryotes in tropical and subtropical Pacific Ocean surface waters. J Oceanogr 73:383–395. https://doi.org/10.1007/s10872-016-0410-0
Yuan H, Li T, Li H, Wang C, Li L, Lin X, Lin S (2021) Diversity distribution, driving factors and assembly mechanisms of free-living and particle-associated bacterial communities at a subtropical marginal sea. Microorganisms 9:2445. https://doi.org/10.3390/microorganisms9122445
Catão CPE, Pollet T, Garnier C, Barry-Martinet R, Rehel K, Linossier I, Tunin-Ley A, Turquet J, Briand J (2021) Temperate and tropical coastal waters share relatively similar microbial biofilm communities while free-living or particle-attached communities are distinct. Mol Ecol 30:2891–2904
Liu K, Hou J, Liu Y, Hu A, Wang M, Wang F, Chen Y, Gu Z (2019) Biogeography of the free-living and particle-attached bacteria in Tibetan lakes. FEMS Microbiol Ecol 95:fiz088. https://doi.org/10.1093/femsec/fiz088
Zhao D, Xu H, Zeng J, Cao X, Huang R, Shen F, Yu Z (2017) Community composition and assembly processes of the free-living and particle-attached bacteria in Taihu Lake. FEMS Microbiol Ecol 93:1–5. https://doi.org/10.1093/femsec/fix062
Li J-L, Salam N, Wang P-D, Chen L-X, Jiao J-Y, Li X, Xian W-D, Han M-X, Fang B-Z, Mou X-Z, Li W-J (2018) Discordance between resident and active bacterioplankton in free-living and particle-associated communities in estuary ecosystem. Microb Ecol 76:637–647. https://doi.org/10.1007/s00248-018-1174-4
Shi J, Zuo Y, Qu W, Liu X, Fan Y, Cao P, Wang J (2022) Stochastic processes shape the aggregation of free-living and particle-attached bacterial communities in the Yangtze River Estuary, China. J Basic Microbiol 62:1514–1525. https://doi.org/10.1002/jobm.202100666
Guo X-p, Lu D-p, Niu Z-s, Feng J-n, Chen Y-r, Tou F-y, Liu M, Yang Y (2018) Bacterial community structure in response to environmental impacts in the intertidal sediments along the Yangtze Estuary, China. Mar Pollut Bull 126:141–149. https://doi.org/10.1016/j.marpolbul.2017.11.003
Mohit V, Archambault P, Toupoint N, Lovejoy C (2014) Phylogenetic differences in attached and free-living bacterial communities in a temperate coastal lagoon during summer, revealed via high-throughput 16S rRNA gene sequencing. Appl Environ Microbiol 80:2071–2083. https://doi.org/10.1128/AEM.02916-13
Caron DA, Davis PG, Madin LP, Sieburth JM (1982) Heterotrophic bacteria and bacterivorous protozoa in oceanic macroaggregates. Science 218:795–797. https://doi.org/10.1126/science.218.4574.795
Crump BC, Baross JA, Simenstad CA (1998) Dominance of particle-attached bacteria in the Columbia River estuary, USA. Aquat Microb Ecol 14:7–18. https://doi.org/10.3354/ame014007
James CC, Barton AD, Allen LZ, Lampe RH, Rabines A, Schulberg A, Zheng H, Goericke R, Goodwin KD, Allen AE (2022) Influence of nutrient supply on plankton microbiome biodiversity and distribution in a coastal upwelling region. Nat Commun 13:e2448. https://doi.org/10.1038/s41467-022-30139-4
Skouroliakou DI, Breton E, Irion S, Artigas LF, Christaki U (2022) Stochastic and deterministic processes regulate phytoplankton assemblages in a temperate coastal ecosystem. Microbiol Spectr 10:e0242722. https://doi.org/10.1128/spectrum.02427-22
Courrat A, Lobry J, Nicolas D, Laffargue P, Amara R, Lepage M, Girardin M, Le Pape O (2009) Anthropogenic disturbance on nursery function of estuarine areas for marine species. Estuar Coast Shelf S 81:179–190. https://doi.org/10.1016/j.ecss.2008.10.017
De Vriend HJ, Wang ZB, Ysebaert T, Herman PM, Ding P (2011) Eco-morphological problems in the Yangtze Estuary and the Western Scheldt. Wetlands 31:1033–1042. https://doi.org/10.1007/s13157-011-0239-7
Caswell H (1976) Community structure: a neutral model analysis. Ecoll Monogr 46:327–354. https://doi.org/10.2307/1942257
Stegen JC, Lin X, Konopka AE, Fredrickson JK (2012) Stochastic and deterministic assembly processes in subsurface microbial communities. ISME J 6:1653–1664. https://doi.org/10.1038/ismej.2012.22
Sorte K, Basak A (2010) Development of a modified copper-cadmium reduction method for rapid assay of total nitric oxide. Ana Methods-UK 2:944–947. https://doi.org/10.1039/C0AY00029A
Hall M, Beiko RG (2018) 16S rRNA Gene Analysis with QIIME2. In: Beiko R, Hsiao W, Parkinson J (eds) Microbiome Analysis. Methods in Molecular Biology, vol 1849. Humana Press, New York, pp 113–129. https://doi.org/10.1007/978-1-4939-8728-3_8
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Lan Y, Wang Q, Cole JR, Rosen GL (2012) Using the RDP classifier to predict taxonomic novelty and reduce the search space for finding novel organisms. PLoS One 7:e32491. https://doi.org/10.1371/journal.pone.0032491
Dueholm MS, Andersen KS, McIlroy SJ, Kristensen JM, Yashiro E, Karst SM, Albertsen M, Nielsen PH (2020) Generation of comprehensive ecosystem-specific reference databases with species-level resolution by high-throughput full-length 16S rRNA gene sequencing and automated taxonomy assignment (AutoTax). mbio 11:e01557–e01520. https://doi.org/10.1128/mbio.01557-20
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’hara R, Simpson GL, Solymos P, MHH S, Wagner H (2013) Package ‘vegan’. Community Ecol 2:1–295. https://doi.org/10.1016/B978-0-12-407863-5.00021-6
Van der Loo MP (2012) Learning RStudio for R statistical computing. Packt Publishing Ltd
Jost L (2007) Partitioning diversity into independent alpha and beta components. Ecology 88:2427–2439. https://doi.org/10.1890/06-1736.1
Ahlmann-Eltze C, Patil I (2021) ggsignif: R package for displaying significance brackets for ‘ggplot2’. A Preprint 1:1–3
Wickham H, Chang W, Wickham MH (2016) Package ‘ggplot2’. Create elegant data visualisations using the grammar of graphics Version, vol 2, pp 1–189. https://doi.org/10.1071/EN19051
Huson DH, Weber N (2013) Microbial community analysis using MEGANMethods Enzymol. Elsevier, pp 465–485
Dexter E, Rollwagen-Bollens G, Bollens SM (2018) The trouble with stress: a flexible method for the evaluation of nonmetric multidimensional scaling. Limnol Oceanogr Methods 16:434–443. https://doi.org/10.1002/lom3.10257
Lord FM, Novick MR (2008) Statistical theories of mental test scores. IAP
Myers L, Sirois MJ (2006) Spearman correlation coefficients, differences between. Wiley
Biau G, Scornet E (2016) A random forest guided tour. Test 25:197–227. https://doi.org/10.1007/s11749-016-0481-7
Jake RC, Alexander L, Nils G (2017) UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33(18):2938–2940. https://doi.org/10.1093/bioinformatics/btx364
Jiang C, Chen H, Grossart HP, Zhang Q, Stoks R, Zhao Y, Ju F, Liu W, Yang Y (2022) Frequency of occurrence and habitat selection shape the spatial variation in the antibiotic resistome in riverine ecosystems in eastern China. Environ Microbiome 17:e53. https://doi.org/10.1186/s40793-022-00447-9
Dufrêne M, Legendre P (1997) Species assemblages and indicator species: the need for a flexible asymmetrical approach. Ecol Monogr 67:345–366. https://doi.org/10.1890/0012-9615(1997)067[0345:SAAIST]2.0.CO;2
Jing H, Liu H (2012) Contrasting bacterial dynamics in subtropical estuarine and coastal waters. Estuar Coast 35:976–990. https://doi.org/10.1007/s12237-012-9504-0
Ning D, Yuan M, Wu L, Zhang Y, Guo X, Zhou X, Yang Y, Arkin AP, Firestone MK, Zhou J (2020) A quantitative framework reveals ecological drivers of grassland microbial community assembly in response to warming. Nat Commun 11:4717. https://doi.org/10.1038/s41467-020-18560-z
He Q, Wang S, Hou W, Feng K, Li F, Hai W, Zhang Y, Sun Y, Deng Y (2021) Temperature and microbial interactions drive the deterministic assembly processes in sediments of hot springs. Sci Total Environ 772:145465. https://doi.org/10.1016/j.scitotenv.2021.145465
Venkataraman A, Bassis CM, Beck JM, Young VB, Curtis JL, Huffnagle GB, Schmidt TM (2015) Application of a neutral community model to assess structuring of the human lung microbiome. mbio 6:e02284–e02214. https://doi.org/10.1128/mBio.02284-14
Wei Q, Wang B, Zhang X, Ran X, Fu M, Sun X, Yu Z (2021) Contribution of the offshore detached Changjiang (Yangtze River) diluted water to the formation of hypoxia in summer. Sci Total Environ 764:142838. https://doi.org/10.1016/j.scitotenv.2020.142838
Guo XW, Linnan Huang, Lingfeng (2020) Spatiotemporal patterns in diversity and assembly process of marine protist communities of the Changjiang (Yangtze River) plume and its adjacent waters. Front Microbiol 11: 579290. https://doi.org/10.3389/fmicb.2020.579290
Jiang XJ, Zhu Z, Wu JN, Lian EG, Liu DY, Yang SY, Zhang RF (2022) Bacterial and protistan community variation across the Changjiang estuary to the ocean with multiple environmental gradients. Microorganisms 10:991. https://doi.org/10.3390/microorganisms10050991
Ye FH, Chunchen Wang, Yu Wu, Jiapeng Hong, Yiguo (2021) Biogeographic pattern of methanogenic community in surface water along the Yangtze River. Geomicrobiolo J 38: 588-597. https://doi.org/10.1080/01490451.2021.1905113
Brankovits D, Pohlman JW, Niemann H, Leigh MB, Leewis MC, Becker KW, Iliffe TM, Alvarez F, Lehmann MF, Phillips B (2017) Methane- and dissolved organic carbon-fueled microbial loop supports a tropical subterranean estuary ecosystem. Nat Commun 8:1835. https://doi.org/10.1038/s41467-017-01776-x
Wu DM, Dai QP, Liu XZ, Fan YP, Wang JX (2019) Comparison of bacterial community structure and potential functions in hypoxic and non-hypoxic zones of the Changjiang Estuary. PLoS One 14:e0217431. https://doi.org/10.1371/journal.pone.0217431
Hu J, Kawamura H, Li C, Hong H, Jiang Y (2010) Review on current and seawater volume transport through the Taiwan Strait. J Oceanogr 66:591–610. https://doi.org/10.1007/s10872-010-0049-1
Ulloa O, Canfield DE, DeLong EF, Letelier RM, Stewart FJ (2012) Microbial oceanography of anoxic oxygen minimum zones. Proc Natl Acad Sci U S A 109:15996–16003. https://doi.org/10.1073/pnas.1205009109
Bižić-Ionescu M, Zeder M, Ionescu D, Orlić S, Fuchs BM, Grossart HP, Amann R (2015) Comparison of bacterial communities on limnic versus coastal marine particles reveals profound differences in colonization. Environ Microbiol 17:3500–3514. https://doi.org/10.1111/1462-2920.12466
Zhang R, Liu B, Lau SC, Ki JS, Qian PY (2007) Particle-attached and free-living bacterial communities in a contrasting marine environment: Victoria Harbor, Hong Kong. FEMS Microbiol Ecol 61:496–508. https://doi.org/10.1111/j.1574-6941.2007.00353.x
Rieck A, Herlemann DP, Jürgens K, Grossart H-P (2015) Particle-associated differ from free-living bacteria in surface waters of the Baltic Sea. Front Microbiol 6:1297. https://doi.org/10.3389/fmicb.2015.01297
Utami YD, Kuwahara H, Igai K, Murakami T, Sugaya K, Morikawa T, Nagura Y, Yuki M, Deevong P, Inoue T (2019) Genome analyses of uncultured TG2/ZB3 bacteria in ‘Margulisbacteria’specifically attached to ectosymbiotic spirochetes of protists in the termite gut. ISME J 13:455–467. https://doi.org/10.1038/s41396-018-0297-4
PB MC, Schulz F, Castelle C, Kantor R, Shih P, Sharon I, Santini J, Olm M, Amano Y, Thomas B (2019) Hydrogen-based metabolism as an ancestral trait in lineages sibling to the Cyanobacteria. Nat Commun 10:463–463. https://doi.org/10.1038/s41467-018-08246-y
Jia B, Li Y, Zi X, Gu X, Yuan H, Jeppesen E, Zeng Q (2023) Nutrient enrichment drives the sediment microbial communities in Chinese mitten crab Eriocheir sinensis culture. Environ Res 15:115281. https://doi.org/10.1016/j.envres.2023.115281
Di Cesare A, Pjevac P, Eckert E, Curkov N, Šparica MM, Corno G, Orlić S (2020) The role of metal contamination in shaping microbial communities in heavily polluted marine sediments. Environ Pollut 265:114823. https://doi.org/10.1016/j.envpol.2020.114823
Ganesh S, Parris DJ, DeLong EF, Stewart FJ (2014) Metagenomic analysis of size-fractionated picoplankton in a marine oxygen minimum zone. ISME J 8:187–211. https://doi.org/10.1038/ismej.2013.144
Bianchi D, Weber TS, Kiko R, Deutsch C (2018) Global niche of marine anaerobic metabolisms expanded by particle microenvironments. Nat Geosci 11:263–268. https://doi.org/10.1038/s41561-018-0081-0
Freitas S, Hatosy S, Fuhrman JA, Huse SM, Mark Welch DB, Sogin ML, Martiny AC (2012) Global distribution and diversity of marine Verrucomicrobia. ISME J 6:1499–1505. https://doi.org/10.1038/ismej.2012.3
Cardman Z, Arnosti C, Durbin A, Ziervogel K, Cox C, Steen A, Teske A (2014) Verrucomicrobia are candidates for polysaccharide-degrading bacterioplankton in an arctic fjord of Svalbard. Appl Environ Microbiol 80:3749–3756. https://doi.org/10.1128/AEM.00899-14
Bergen B, Herlemann DPR, Labrenz M, Jürgens K (2014) Distribution of the verrucomicrobial clade Spartobacteria along a salinity gradient in the Baltic Sea. Environmental Microbiology Reports 6:625–630. https://doi.org/10.1111/1758-2229.12178
St. Pierre KA, Oliver AA, Tank SE, BPV H, Giesbrecht I, CTE K, Jackson JM, Lertzman KP, Floyd WC, Korver MC (2020) Terrestrial exports of dissolved and particulate organic carbon affect nearshore ecosystems of the Pacific coastal temperate rainforest. Limnol Oceanogr 65:2657–2675. https://doi.org/10.1002/lno.11538
Ohore OE, Wei Y, Wang Y, Nwankwegu AS, Wang Z (2022) Tracking the influence of antibiotics, antibiotic resistomes, and salinity gradient in modulating microbial community assemblage of surface water and the ecological consequences. Chemosphere 305:135428. https://doi.org/10.1016/j.chemosphere.2022.135428
Boesch DF (2002) Challenges and opportunities for science in reducing nutrient over-enrichment of coastal ecosystems. Estuaries 25:886–900. https://doi.org/10.1007/BF02804914
Hanson CA, Fuhrman JA, Horner-Devine MC, Martiny JB (2012) Beyond biogeographic patterns: processes shaping the microbial landscape. Nat Rev Microbiol 10:497–506. https://doi.org/10.1038/nrmicro2795
Yung C-M, Ward CS, Davis KM, Johnson ZI, Hunt DE (2016) Insensitivity of diverse and temporally variable particle-associated microbial communities to bulk seawater environmental parameters. Appl Environ Microbiol 82:3431–3437. https://doi.org/10.1128/AEM.00395-16
Lyons M, Ward J, Gaff H, Hicks R, Drake J, Dobbs FC (2010) Theory of island biogeography on a microscopic scale: organic aggregates as islands for aquatic pathogens. Aquat Microb Ecol 60:1–13. https://doi.org/10.3354/ame01417
Gram L, Grossart H-P, Schlingloff A, Kiørboe T (2002) Possible quorum sensing in marine snow bacteria: production of acylated homoserine lactones by Roseobacter strains isolated from marine snow. Appl Environ Microbiol 68:4111–4116. https://doi.org/10.1128/AEM.68.8.4111-4116.2002
Ruiz-González C, Niño-García JP, del Giorgio PA (2015) Terrestrial origin of bacterial communities in complex boreal freshwater networks. Ecol Lett 18:1198–1206. https://doi.org/10.1111/ele.12499
Grossart H-P (2010) Ecological consequences of bacterioplankton lifestyles: changes in concepts are needed. Environ Microbiol Rep 2:706–714. https://doi.org/10.1111/j.1758-2229.2010.00179.x
Stegen JC, Lin X, Fredrickson JK, Konopka AE (2015) Estimating and mapping ecological processes influencing microbial community assembly. Front Microbiol 6:370. https://doi.org/10.3389/fmicb.2015.00370
Zhou J, Deng Y, Zhang P, Xue K, Liang Y, Van Nostrand JD, Yang Y, He Z, Wu L, Stahl DA, Hazen TC, Tiedje JM, Arkin AP (2014) Stochasticity, succession, and environmental perturbations in a fluidic ecosystem. Proc Natl Acad Sci U S A 111:E836–E845. https://doi.org/10.1073/pnas.1324044111
Meyerhof MS, Wilson JM, Dawson MN, Michael Beman J (2016) Microbial community diversity, structure and assembly across oxygen gradients in meromictic marine lakes, Palau. Environ Microbiol 18:4907–4919. https://doi.org/10.1111/1462-2920.13416
Bracken LJ, Wainwright J, Ali G, Tetzlaff D, Smith M, Reaney S, Roy A (2013) Concepts of hydrological connectivity: research approaches, pathways and future agendas. Earth-Sci Rev 119:17–34. https://doi.org/10.1016/j.earscirev.2013.02.001
Chen Y, Liu R, Sun C, Zhang P, Feng C, Shen Z (2012) Spatial and temporal variations in nitrogen and phosphorous nutrients in the Yangtze River Estuary. Mar Pollut Bull 64:2083–2089. https://doi.org/10.1016/j.marpolbul.2012.07.020 Epub 2012 Aug 19
Bachmann J, Heimbach T, Hassenrück C, Kopprio GA, Iversen MH, Grossart HP, Gärdes A (2018) Environmental drivers of free-living vs. particle-attached bacterial community composition in the Mauritania upwelling system. Front Microbiol 9:2836. https://doi.org/10.3389/fmicb.2018.02836
Funding
This work was funded by the Key R&D projects in Zhejiang Province (Grant Numbers: 2021C02047, 2022C02040, 2023C03120), the Fundamental Research Fund for the Provincial Universities of Zhejiang (Grant Number: 2021JD003), the National Natural Science Foundation of China (Grant Number: Q20C060004), the Science and Technology Program of Zhoushan (Grant Number: 2019C21011), the Research Project of Ecological Environment Protection and Restoration of Yangtze River in Zhoushan (Grant Number: 2019C21011), and the Natural Science Foundation of Zhejiang Province (Grant Number: SZGXZS2020068).
Author information
Authors and Affiliations
Contributions
JJW, WDX conceived and designed the study. JHC, PLC performed the fieldwork. WQ, CYH performed DNA quantification and sequencing in the laboratory. WDX, JJD reviewed the literature and wrote the manuscript. YYZ, XZL, JLL, PDW, WJL improved and corrected the manuscript. The above authors approved this manuscript.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xian, WD., Ding, J., Chen, J. et al. Distinct Assembly Processes Structure Planktonic Bacterial Communities Among Near- and Offshore Ecosystems in the Yangtze River Estuary. Microb Ecol 87, 42 (2024). https://doi.org/10.1007/s00248-024-02350-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02350-x