Abstract
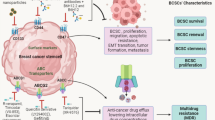
Breast cancers contain small population of tumor-initiating cells called breast cancer stem cells (BCSCs), which are spared even after chemotherapy. Recently, BCSCs are implicated to be a cause of metastasis, tumor relapse, and therapy resistance in breast cancer. BCSCs have unique molecular mechanisms, which can be targeted to eliminate them. These include surface biomarkers, proteins involved in self-renewal pathways, drug efflux transporters, apoptotic/antiapoptotic proteins, autophagy, metabolism, and microenvironment regulation. The complex molecular mechanisms behind the survival of BCSCs and pharmacological targets for elimination of BCSCs are described in this review.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the most common type of cancer in women and is the second leading cause of cancer-related deaths worldwide. According to the World Health Organization (WHO) report, 17.5 million breast cancer-related deaths can be expected per year, by 2050 (Ferlay et al. 2010). Resistance to conventional therapies, metastasis, and relapse of tumors are emerging as major causes of breast cancer-related deaths (Singh and Settleman 2010). Recently, it was identified that breast cancer stem cells (BCSCs) are one of the major responsible factors for therapy resistance, tumor relapse, and metastasis (Al-Hajj et al. 2003; Chen et al. 2013). It was reported that BCSCs express high levels of drug efflux transporters, which can be determined by treatment with Hoechst 33342 dye. The cells containing high levels of the drug efflux transporters expulse Hoechst and are designated as side population (SP) cells (Patrawala et al. 2005). Accumulating evidence has shown that numerous cell lines and tumors contain SP cells and that this cell population possesses a greater capacity for chemoresistance and tumorigenesis than non-SP cells (Britton et al. 2012). The quiescence of BCSCs (i.e., they spend more time in G0 cell cycle phase) and their high DNA repair capacity also makes them to resist apoptosis caused by chemotherapeutic agents (Reya et al. 2001). The pool of BCSCs which are spared by conventional therapies will convert into tumor cells in future, thus leading to tumor relapse (Li et al. 2011).
Unlike bulk tumor cells (non-BCSCs), BCSCs grow in a nonattached (suspension) form when moving from their source to other locations in the body. Due to the nonattached growth nature, BCSCs proliferate in blood stream during cancer metastasis and give rise to spread of tumor. BCSCs promote cancer metastasis by a process called epithelial-to-mesenchymal transition (EMT), in which epithelial cells lose their intercellular adhesion, accompanied by gain of invasive and migratory properties, which is a prerequisite for metastasis (Karnoub et al. 2007; Sabe 2011; Liu et al. 2012; Hu et al. 2015). In this review, we discuss potential avenues for the pharmacological targeting of BCSCs based on their molecular features, including surface biomarkers (CD44, CD133, EpCAM, and ALDH1), proteins involved in self-renewal pathways (Wnt/β signaling proteins, Smo, γ-secretase, STAT-3, etc.), drug efflux transporters (ABCG2 and ABCB1), apoptotic/antiapoptotic proteins (Bcl2, survivin etc.), proteins involved in autophagy, metabolism, epigenetic regulation, and microenvironment regulation (Ablett et al. 2012; Britton et al. 2012; Vinogradov and Wei 2012; Chapellier and Maguer-Satta 2016).
Breast epithelial hierarchy and origin of BCSCs
Understanding the cell of origin of breast cancer is of great importance to unravel the cause of tumor heterogeneity. The mammary epithelium is composed of two types of cell lineages, luminal cells and myoepithelial (basal cells), which are organized into a series of branching ducts that terminates into secretary alveoli and aids in lactation. Luminal cells surround the central lumen and basal cells are located in basal position adjacent to basement membrane of mammary epithelium. It was reported that luminal cells and basal cells originate from multipotent mammary stem cells (MaSCs) during the development of mammary epithelium. Breast epithelial hierarchy suggests that BCSCs can be derived from normal MaSCs, transformed by the deregulation of normal self-renewal (Dontu et al. 2003; Wicha et al. 2003). Compelling body of evidence suggest that although MaSCs are required for the long-term maintenance of mammary gland homeostasis, postnatal glands, luminal, and basal unipotent progenitor cells can independently sustain luminal and basal lineages, respectively, for a long period of time (Van Keymeulen et al. 2011; Rios et al. 2014). Multiple mammary cell types, therefore, can have long-term self-renewal abilities and BCSCs may originate from these precursor cells due to mutations. In addition, it was reported that different breast cancer subtypes may originate from different mammary cell lineages (Lim et al. 2009; Visvader 2011; Visvader et al. 2014). For example, basal-like breast cancer is likely to originate from luminal progenitor cells, whereas multipotent MaSCs are likely the precursor of the claudin-low subtype (Lim et al. 2009; Molyneux et al. 2010).
According to “misplacement somatic stem cell” theory, BCSCs may originate from misplacement of somatic stem cells de novo (Wang et al. 2013a). According to this theory, somatic cells of the normal tissue would undergo successive DNA mutations that allow the cell to evolve and acquire the malignant phenotypes of BCSCs. According to this model, long-lived nature of normal stem cells (NSCs) allows them more time to acquire mutations to become BCSCs. It was also reported that BCSCs can originate from tumor cells via induction of epithelial to mesenchymal transition (EMT) as a part of disease progression or in response to chemotherapeutic agents and environmental stress (Owens and Naylor 2013).
Pharmacological targets of BCSCs
BCSC surface markers
BCSCs express specific biological markers or antigens on their surface that can be used to identify or label them. Fluorescence-activated cell sorting (FACS) uses specific cell surface biomarkers to sort BCSCs (Chen et al. 2013). The expression of these unique surface biomarkers on BCSCs is reported to be associated with chemo/radioresistance in breast cancer (Ablett et al. 2012). A number of therapeutic antibodies, various small molecules, have been proposed for targeting these biomarkers for identifying and elimination of BCSCs. CD44, CD133, EpCAM, and ALDH1 are the important biomarkers of BCSCs. It was reported that triple negative breast cancer (TNBC) cells have higher percentage of these markers compared to other breast cancer subtypes (Croker et al. 2009). It was also reported that BCSC surface markers are enriched in normal tissues adjacent to TNBC cells (Atkinson et al. 2013).
Cluster of differentiation44
Cluster of differentiation44 (CD44) is a surface glycoprotein that is known to participate in a wide variety of cellular functions including regulation of cell adhesion, proliferation, migration, growth, survival, angiogenesis, differentiation, and matrix cell-signaling processes in collaboration with other cellular proteins (Phillips et al. 2006; Honeth et al. 2008; Goodarzi et al. 2014; Yan et al. 2015; Muntimadugu et al. 2016). It was reported that CD44 activates the Rho family of GTPases and initiate recruitment of signaling molecules like T lymphoma invasion and metastasis-inducing protein (Tiam1), p115, Ras-related C3 botulinum toxin substrate-1(Rac1), Rho-associated protein kinase, and proto-oncogene c-Src. These signaling molecules activates the phosphoionositide kinase (PI3K) pathway that is necessary for survival and migration of cancer cells (Bourguignon et al. 2000, 2008, 2009). More recently, it was also reported that CD44 expression is associated with chemoresistance by upregulation of the multidrug resistance receptor through activation of STAT3 (Bourguignon et al. 2008; Louderbough and Schroeder 2011). Clinical studies have shown a positive correlation between expression of CD44-positive BCSCs and tumor aggressiveness in patients with breast cancer (Balic et al. 2006; Yang et al. 2016).
Cluster of differentiation133
Cluster of differentiation133 (CD133) or prominin-1 is another important surface biomarker of BCSCs mainly associated with chemoresistance. It was reported to upregulate antiapoptotic genes like survivin and c-FLIP, promote autophagy, and associate with the Wnt/β-catenin self-renewal signaling pathway and vasculogenic mimicry (VM). The over expression of CD133 has been reported to have negative correlation with breast cancer patient survival (Wu and Wu 2009; Li 2013; Liu et al. 2013; Leon et al. 2016).
Epithelial cellular adhesion molecule
Epithelial cellular adhesion molecule (EpCAM) is a type I transmembrane glycoprotein, belonging to the family of adhesion molecules, which is overexpressed in BCSCs and is associated with poor survival rate in breast cancer (Munz et al. 2009; Königsberg et al. 2011). EpCAM comprise of an extracellular domain (EpEx) and an intracellular domain (EpICD). Cleavage of EpICD (upon EpCAM) activation results in signal transduction and activation of EpICD target genes like Nanog, Oct4, Klf4, and Sox2. These genes are involved in cell cycle regulation and apoptosis (Munz et al. 2009). In addition, it was reported that EpCAM inhibits E-cadherin-mediated adhesion and also activates Wnt/β-catenin pathway to promote survival of BCSCs (Fig. 1).
Cross talk between EpCAM signaling and the Wnt/β-catenin pathway. Activation of the frizzled receptor by members of the Wnt family of ligands induces the inhibition of GSK3β and the subsequent stabilization of β-catenin. Upon nuclear translocation, β-catenin controls Lef-1-dependent transcription. EpICD interacts with the very same components to form a nuclear complex comprised of β-catenin, FHL2, and Lef-1. This nuclear complex binds with promoters of genes like Nanog, Oct4, Klf4, and Sox2 which are involved in cell cycle regulation and stemness of BCSCs. (Source Imrich et al. 2012)
Aldehyde dehydrogenase1 (ALDH1)
Aldehyde dehydrogenases (ALDHs) are superfamily of enzymes which are involved in the oxidation of intracellular aldehydes to carboxylic acids, retinoic acid, and γ-amino butyric acid (GABA) (Ginestier et al. 2007). These enzymes were reported to be overexpressed and associated with chemo/radioresistance in BCSCs (Resetkova et al. 2010; Croker and Allan 2012). It was reported that ALDHhigh subtype in BCSCs was a predictor of poor survival in patients (Ginestier et al. 2007). High ALDH activity also reported to prevent apoptosis due to anticancer agents by metabolizing them into inactive metabolites. It was reported that ALDHs influence various pathways like Wnt, notch, transforming growth factor-β (TGF-β), extracellular signal-regulated kinases (ERK) in ALDHhigh subpopulation of cancer cells that influence proliferation and cell fate, epithelial-to-mesenchymal transition (EMT), retinoic acid synthesis, hypoxia, DNA damage response, and cell migration (Rodriguez-Torres and Allan 2016).
Components of self-renewal pathways
Self-renewal is one of the key features of NSCs, responsible for proliferation and maintenance. The signaling pathways such as Wnt/β-catenin, Notch, Hedgehog (Hh), TGF-β, signal transducer and activator of transcription factor 3 (STAT3), and B cell-specific Moloney murine leukemia virus integration site 1 (Bmi-1) implicated in the self-renewal of NSCs (Clarke et al. 2006). In BCSCs, these self-renewal signaling pathways are deregulated and result in extensive cell proliferation and also considered as an early event in the process of carcinogenesis (Fillmore and Kuperwasser 2008). Inhibition of self-renewal pathways, therefore, can be an attractive strategy for elimination of BCSCs (Liu et al. 2006; Kai et al. 2010) (Table 1).
Wnt/β-catenin signaling pathway
The Wnt/β-catenin signaling pathway is an evolutionarily well-conserved pathway that regulates growth, regeneration, and self-renewal (Branda and Wands 2006). The activation of Wnt/β-catenin pathway occurs when a Wnt ligand binds to the transmembrane receptor that in turn results in binding of the low-density lipoprotein-related receptor (LRP). This leads to the suppression of glycogen synthase kinase-3β (GSK-3β) protein, thereby improving the stability of β-catenin. Consequently, β-catenin forms a complex with the transcription factor/lymphocyte enhancer factor and activates the expression of Wnt target gene such as c-terminus of myc protein (c-myc) and cyclin D1 (Fleming et al. 2008; Choi et al. 2010). Altered activation of Wnt/β-catenin signaling is a key feature of breast cancer where it is considered to be critical for self-renewal of BCSCs and also reported to enhance tumor metastasis by promoting EMT (Zhao et al. 2007; MacDonald et al. 2009) (Fig. 2).
Wnt/β-catenin (a), Notch (b), Hedgehog (c) and STAT3 (d) and Bmi (e) mediated signaling for self-renewal of BCSCs. STAT3 signal transducer and activator of transcription factor 3, TGF-β transforming growth factor-β, Hh Hedgehog, Smo Smoothened, SHh Sonic Hedgehog, IHh Indian hedgehog, DHh desert hedgehog, NICD Notch intracellular domain, NECD Notch extracellular domain, GSK-3β glycogen synthase kinase 3 β, DHh desert Hedgehog, DLL4 delta-like 4 ligand, Bmi-1 B cell-specific Moloney murine leukemia virus integration site 1, JAK Janus kinase, Gli glioma-associated oncogene
Notch signaling pathway
The Notch signaling pathway is essential for differentiation of BCSCs (Yin et al. 2010). It was reported that the aberrant activation of notch signaling in BCSCs is associated with activation of notch target genes which are involved in the maintenance of self-renewal in BCSCs (Artavanis-Tsakonas et al. 1999; Reya et al. 2001; Androutsellis-Theotokis et al. 2006; Hori et al. 2013). This signaling pathway is activated through four Notch receptors (Notch 1–4); among those, Notch 4 and Notch 1 are implicated in self-renewal of BCSCs (Bray 2006; Cerdan and Bhatia 2010; Harrison et al. 2010b; Zhong et al. 2016). Ligand protein binding to Notch receptors leads to their cleavage by γ-secretase to release the Notch intracellular domain (NICD), and following the nuclear translocation, it induces transcriptional activation of Notch target genes to promote survival of BCSCs (Schweisguth 2004). Inhibitors of γ-secretase, therefore, prevent the proteolysis of Notch receptors and suppress the Notch activity in BCSCs. Mastermind-like (MAML) and Delta-like 4 ligand (DLL4) proteins are the other important molecular targets from Notch signaling pathway to inhibit self-renewal of BCSCs (Fig. 2).
Transforming growth factor-β pathway
Transforming growth factor-β (TGF-β)-mediated signaling is essential during the initial phase of development and regeneration of cells (Pryce et al. 2009; Greenow and Clarke 2012). TGF-β ligands binding activates TGF-β type I receptor. The type I receptor triggers the phosphorylation of SMADs (transcription factors) and results in ligand-induced transcription of self-renewal genes in BCSCs (Fig. 2) (Massagué 2000; Liu et al. 2012). TGF-β signaling exerts tumor suppressor effects in normal cells and early carcinomas. However, the mutations in TGF-β results in tumor genesis. As tumors develop and progress, the protective and cytostatic effects of TGF-β will be lost. TGF-β signaling then promotes cancer progression, invasion, and tumor metastasis. TGF-β, therefore, have dual role in both tumor suppression and tumor progression (Moses and Barcellos-Hoff 2011). Higher TGF-β levels in the serum and urine was correlated with poor survival rate and advanced disease state in cancer patients (Tsai et al. 1997). Designing novel therapeutic agents targeting TGF-β is, however, challenging due to its dual role in carcinogenesis. It is necessary to develop drugs that specifically aimed at blocking the prometastatic effects of the TGF-β signaling pathway without affecting its tumor suppressive effects.
Hedgehog signaling pathway
Hedgehog (Hh) signaling pathway is an important pathway that is responsible for the maintenance and self-renewal capacity of the BCSCs. The Sonic Hh (SHh), Desert Hh (DHh), and Indian Hh (IHh) are the three gene homologs of Hh (Ingham and McMahon 2001; Micchelli et al. 2002; Takebe et al. 2011). In Hh pathway, the activation of Smoothened (Smo), a seven-pass transmembrane receptor, is necessary for signaling process. In the presence of Hh ligand, Smo activates the glioma-associated oncogene (Gli) family of transcription factors (Gli1/2/3) to carry out the further downstream signaling required for self-renewal of BCSCs (Svärd et al. 2006). Smo receptor and Gli family of proteins, therefore, can be druggable molecular targets to inhibit self-renewal of BCSCs (Fig. 2).
B cell-specific Moloney murine leukemia virus integration site-1
The B cell-specific Moloney murine leukemia virus integration site-1 (Bmi-1) is one of the polycombcomplex proteins, reported to be involved in the differentiation and self-renewal mechanisms of BCSCs (Alkema et al. 1993; Jacobs et al. 1999; Gil et al. 2005). Bmi-1 affects morphogenesis during embryonic development and in hematopoiesis with a pervasive expression in almost all tissues (Van der Lugt et al. 1994). It is noted that BCSCs are dependent on Bmi-1 for their maintenance and self-renewal (Fig. 2) (Sawa et al. 2005; Borah et al. 2015). In addition, it was reported that Hh signaling act along with Bmi-1 to regulate the self-renewal of BCSCs (Kubo et al. 2004; Liu et al. 2006).
JAK/STAT3 pathway
Signal transducers and activators of transcription (STATs) are a family of transcription factors required for regulation of growth, survival, and differentiation of cells (Darnell Jr et al. 1994; Ihle 2001). So far, seven STAT proteins have been recognized in mammalian cells. Among all, STAT3 plays a key role in carcinogenesis by regulating the transcription of genes involved in cell proliferation, differentiation, apoptosis, angiogenesis, and metastasis (Akira et al. 1994; Yu and Jove 2004). IL-6/JAK/STAT3 is the canonical STAT3 activation signaling pathway, reported to be deregulated in cancer. Since the receptor of IL-6 does not contain a kinase catalytic domain, it induces STAT3 phosphorylation by activating members of the JAK family (Fig. 2) (Ihle et al. 1994; Heim et al. 1995; Ihle and Kerr 1995; Stahl et al. 1995; Ihle 2001). The IL-6/JAK2/STAT3 pathway was found to be active in CD44+/CD24− BCSCs. It was demonstrated that inhibition of STAT3 pathway suppressed growth of xenograft tumors. In addition, it has been reported that cancer cells can be converted into a cancer stem cells via the IL-6/JAK1/STAT3 signaling pathway (Marotta et al. 2011; Wang et al. 2012; Kim et al. 2013; Xiong et al. 2014; Chung and Vadgama 2015). STAT3 is an important molecular drug target for inhibition of this pathway. Recently, it was identified that niclosamide, an anti-helmenthic drug, is an inhibitor of STAT3 phosphorylation (Li et al. 2013; Wang et al. 2013b; Li et al. 2014). In addition, niclosamide is also reported to prevent conversion of non-BCSCs to BCSCs (Kim et al. 2013) and reduced resistance to chemotherapy (Liu et al. 2016a; Liu et al. 2016b) (Table 2). Recently, Wang et al. have shown that leptin-JAK/STAT3 regulate lipid metabolism through fatty acid β-oxidation (FAO) to promote breast cancer stemness and chemoresistance. Blocking FAO and/or depleting leptin sensitized cancer cells to chemotherapy while reducing BCSCs in vivo (Wang et al. 2018).
Apoptotic/antiapoptotic proteins
Deregulation of apoptosis and antiapoptotic (survival) signaling pathways is a characteristic of cancer and a critical determinant of efficacy of chemotherapy (Fig. 3) (Brown and Attardi 2005). In this context, a compelling body of evidence suggests that BCSCs use several mechanisms to deregulate apoptotic/antiapoptotic pathways and promote resistance to treatment (Wicha et al. 2006; Karnoub et al. 2007). The B cell lymphoma2 (Bcl2), FLICE like inhibitory protein (c-FLIP), nuclearfactor-κ-B (NF-κB), phosphatase and tensin homolog (PTEN), mammalian target of rapamycin(m-TOR), and death receptors (DR)-4/5 proteins are the well-characterized regulators of apoptosis and molecular targets for elimination of BCSCs (Martinou and Youle 2011). Bcl2 is an antiapoptotic protein that is reported to be overexpressed in 75% of breast cancer cells (Domen et al. 1998; Honma et al. 2015; Merino et al. 2016). It was reported that breast tumor-targeted gene therapy with pro-apoptotic gene Bcl2 interacting killer (BIK) improved the efficacy of the chemotherapeutic agents against breast cancer (Lang et al. 2011).
Apoptotic and antiapoptotic signaling in cancer stem cells. Bcl2 B cell lymphoma 2, BIK Bcl2 interacting killer, DRs death receptors, IL interleukin, PTEN phosphatase and tensin homolog, PI3-K phosphoinositide 3-kinase, NF-κB nuclear factor-κB, Bad Bcl-2-associated death promoter, PUMA p53 upregulated modulator of apoptosis, IAP inhibitor of apoptosis protein, XIAP X-linked inhibitor of apoptosis protein, IKK I-kappa kinase, c-FLIP FLICE like inhibitory protein (Signore et al. 2013)
Drug efflux transporters
Drug efflux transporters or ATP-binding cassette transporters (or ABC transporters) like P-glycoprotein (P-gp) or ABCB1 and breast cancer resistance protein (BCRP) or ABCG2 are implicated in chemoresistance (Shervington and Lu 2008; Yin et al. 2008). ABCB1 is reported to be expressed and responsible for chemoresistance in breast cancer. Studies have shown that higher expression of CD133 is also accompanied with an elevated ABCB1 efflux activity (Moitra 2015). Hoechst 33342 assay which is used to isolate BCSCs is based on the principle that BCSCs are Hoechst dim due to overexpression of the ABCG2 drug efflux transporter that pumps the dye out of the cells (Kim et al. 2008; Britton et al. 2012). Several inhibitors of ABCG2, like Fumitremorgin C (FTC), Tryprostatin-A, and Tariquidar, have been proposed to kill BCSCs in order to achieve radical cure in breast cancer. However, the clinical application of these compounds is limited due to their low inhibition capacity and off-target effects on the healthy cells (Rabindran et al. 1998; Rabindran et al. 2000; Zhao et al. 2002; Woehlecke et al. 2003; Robey et al. 2004; Peired et al. 2016).
DNA repair capacity
Radiation therapy and chemotherapeutic agents cause DNA damage for induction of apoptosis (Cheung-Ong et al. 2013). BCSCs possess DNA repair ability by activation of various checkpoint mechanisms (Al-Assar et al. 2011; Kim et al. 2012; Peitzsch et al. 2013). DNA damage can be repaired by homology-directed recombination (HDR) or through nonhomologous end joining (NHEJ) (Brandsma and van Gent 2012). The HDR involves resegmentation of the two ends of DNA from 3′ to 5′, formation of single-strand DNA at the 3′ end, assembly of RAD51 filaments (a protein family contributing in the repair of DNA double-stand breaks), and finally repair by annealing at the end of the double-strand break. The HDR repair occurs during the S and G2 phases of the cell cycle. NHEJ utilizes the Lupus Ku autoantigen protein p70/80 (KU70/80) to join the DNA strands. In addition, nucleases, polymerases, DNA-dependent protein kinases, and ligases participate in the NHEJ repair process (Jackson 2002; Jasin and Rothstein 2013). DNA damage checkpoint proteins, like checkpoint kinases (ChK 1/2) are the important molecular targets for prevention of DNA repair and inhibition of BCSCs (Niida and Nakanishi 2006; Yin and Glass 2011).
Oxidative stress
Many anticancer agents and radiation therapy lead to reactive oxygen species (ROS) production to induce apoptosis in cancer cells by either intrinsic or extrinsic pathways (Cook et al. 2004; Sena and Chandel 2012; Sinha et al. 2013). However, BCSCs maintain low ROS levels in addition to high endogenous antioxidant levels (Trachootham et al. 2009). Upregulation of genes encoding the antioxidant enzymes like superoxide dismutase (SOD), catalase (CAT), and glutathione peroxidase (GPO) can be found in BCSCs (Diehn et al. 2009). In addition, BCSCs are regularly localized in the hypoxic regions of the tumors to avoid ROS-mediated apoptosis. This makes BCSCs to avoid oxidative DNA damage and maintain their quiescent state for their survival (Phillips et al. 2006; Gilbertson and Rich 2007). Induction of oxidative stress and reduction of antioxidant defense, however, is not considered as an effective strategy for elimination of BCSCs, due to deleterious effects on healthy cells.
BCSCs metabolism
Differentiated bulk cancer cells rely primarily on glycolysis for production of ATP and to manage high rate of proliferation (Ward and Thompson 2012). In contrast, BCSCs can be highly glycolytic or oxidative phosphorylation (OXPHOS) dependent. In both cases, mitochondrial function is important (Sancho et al. 2016). Inhibition of the mitochondrial metabolic function, therefore, became a potential strategy in recent years for elimination of BCSCs and prevention of tumor relapse. It was reported that BCSCs show distinct glucose and mevalonate metabolism (Ginestier et al. 2012; Dong et al. 2013). It has been recently demonstrated that adenine nucleotide translocator-2 (ANT2), which is involved in glycolytic metabolism (Raaijmakers et al. 2005), and Hexokinase 2 (HK2), which catalyzes the first committed step of glucose metabolism, can be targeted for elimination of BCSCs. Using HK2 conditional knockout mice, it was demonstrated that HK2 is required for tumor initiation and maintenance in breast cancer (Patra et al. 2013). Metformin has been reported to eliminate BCSCs by inhibiting HK2 and thereby enhanced the effects of chemotherapy (Salani et al. 2014) (Fig. 4).
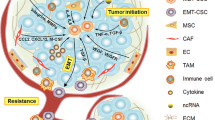
BCSCs microenvironment/niche
BCSCs require a specialized microenvironment or niche which is regulated by various factors for their survival. The factor that regulate BCSCs microenvironment include fibroblast stimuli, immune cells, autocrine signals, and extracellular matrix (ECM) components, as well as physical/chemical factors such as oxygen pressure, nutrients levels, and low pH (Bozorgi et al. 2015; He et al. 2016). Growth factors and cytokines released by tumor cells and cancer-associated fibroblasts and immune cells have strong effects on the survival and metastasis of BCSCs (Culig 2011; Korkaya et al. 2012). It was reported that combination therapy with an IL-6 receptor antibody is required to suppress acquired trastuzumab resistance in breast cancer in vivo. In addition, it was reported that the IL-8 receptor, CXCR1, is highly expressed in BCSCs. Interestingly, chemotherapy may increase the CSC pool by stimulating the release of IL-8, whereas a CXCR1 small molecule inhibitor helped to eliminate the residual BCSC population following docetaxel therapy (Ginestier et al. 2010).
Autophagy proteins
Autophagy is a survival promoting physiological process in BCSCs against various environmental stress, radiation, chemotherapeutic drugs, and hypoxia (Choi et al. 2013). Autophagy plays an important role in breast cancer initiation or transformation of mammary epithelial cells, chemoresistance, and metastasis. Excessive self-eating can promote death, and low levels of autophagy activated in response to cellular stress is believed to promote resistance of breast cancer cell to chemotherapy, radiation, and targeted therapy in most settings (Jain et al. 2013). In BCSCs, it was reported that autophagy is also responsible for chemoresistance, tumor relapse, and metastasis (Sui et al. 2013; Ojha et al. 2015). Mechanisms by which autophagy promotes cancer include induction of the p53 and altering metabolic function of mitochondria (White 2015). It was reported that treatment with autophagy inhibitors or silencing of autophagy-associated genes affects stem cell renewal, differentiation, and stress-resistant abilities that results in elimination of BCSC population and enhanced the sensitivity to chemotherapeutic agents (Mai et al. 2012; Singh et al. 2012). Recently, it was reported that autophagy inhibition with chloroquine (CQ) resulted in elimination of BCSCs in triple negative breast cancer (TNBC) and potentiated the cytotoxic effects of carboplatin (Liang et al. 2016).
Epigenetic regulation of BCSCs
Epigenetic regulation of the genome is one of the primary mechanisms by which genetic code is altered to control cellular developmental hierarchies without change in DNA sequences. Epigenetic mechanisms include histone modifications, DNA methylation, chromatin remodeling, and changes in noncoding RNAs including miRNAs. Emerging evidences suggest that deregulation of various epigenetic mechanisms can contribute to tumor initiation and progression, particularly with respect to maintenance and BCSCs. Histone methylation is a critical factor in epigenetic regulations and is mediated by methyltransferases which catalyze the mono-, di-, or trimethylation of specific lysine residues (Wei et al. 2008). Histone methylation occurs predominantly on lysine (K) and arginine (R) residues (Stallcup 2001). The histone lysine methylation occurs at three different levels: mono-, di-, and trimethylation and commonly associated with gene activation or repression. Histone H3 lysine 4 (H3K4), histone H3 lysine 36 (H3K36), and histone H3 lysine 79 (H3K79) are associated with gene activation and histone H3 lysine 9 (H3K9), histone H3 lysine 27 (H3K27), and histone H4 lysine 20 (H4K20) are associated with gene repression. Aberrations in histone modifications can lead to deregulated gene expression as seen in various human disease and malignancies. It was reported that epigenetic enzymes will be recruited to the E-cadherin promoter by Snail and cause transcriptional silencing of E-cadherin and lead to EMT. Dong et al. have investigated the interaction of Suv39H1 (Snail binding protein) with Snail and identified Suv39H1 is critical for the enrichment of H3K9me3 on the E-cadherin promoter in breast cancer cells and in the induction of EMT (Dong et al. 2012, 2013). It was also reported that the stemness of BCSCs is maintained by the epigenetic marker H3K27me3. In a recent study, Ningning Yan et al. have proposed H3K27me3 as a target for elimination of BCSCs. It was reported that inhibition of H3K27me3 demethylation specifically target BCSCs by inactivation of JMJD3 and UTX, which facilitate target gene activation by catalyzing the conversion of H3K27me3 and H3K27me2 to H3K27me1 and maintain the balance between methylation and demethylation (Yan et al. 2017). Recent studies in epigenomics have, therefore, led to understand the key mechanisms by which epigenetic regulations contribute to tumor progression. Further understanding of the mechanisms involved in epigenetic regulations and testing the epigenetic modulating drugs, offer new avenues for targeting BCSCs.
Future prospects
One major challenge for targeting BCSCs is the molecular cross talk between the self-renewal signaling pathways in BCSCs and NSCs. Multiple developmental signaling pathways implicated in regulating BCSCs, like TGF-β, Wnt, and Notch, have been shown to regulate normal stem and progenitor cells. Selective targeting of BCSCs, therefore, will be challenging. TGF-β is a potent EMT inducer that is reported to be secreted by multiple cell types in tumors (Padua and Massagué 2009). TGF-β is reported to activate EMT programs in both mammary epithelial cells and also BCSCs. In BCSCs, TGF-β activation leads to expression of surface markers CD44highCD24low, and the increase the ability to form mammospheres (Mani et al. 2008; Scheel et al. 2011; Bruna et al. 2012). In normal human mammary cells, efficient activation of EMT requires co-operation of both TGF-β and Wnt signaling pathways. However, such co-operation is reported to be essential only in early developmental stage (Nishita et al. 2000). In adult mammary glands, MaSCs exhibit elevated Wnt signaling (van Amerongen et al. 2012) and the overexpression of Wnt proteins or activation of canonical Wnt by Axin2 mutation or MMP3 overexpression promotes the expansion of MaSCs (Shackleton et al. 2006; Zeng and Nusse 2010; Kessenbrock et al. 2013). In contrast to Wnt, Notch is reported to induce the commitment of MaSCs to luminal-specific progenitors (Bouras et al. 2008). However, basal-like breast cancer is likely to originate from luminal progenitor cells (Molyneux et al. 2010). Notch signaling, therefore, is particularly important for this breast cancer subtype (Harrison et al. 2010a). It was also reported that although TGF-β increases BCSC numbers in claudin-low subtype, it suppresses BCSC in certain basal-like and luminal breast cancer subtypes (Bruna et al. 2012). Similarly, Wnt-overexpressing fibroblasts promoted the growth of one patient-derived xenograft (PDX) model but inhibited another PDX (Green et al. 2013). Future therapeutic strategies can, therefore, be tailored based on the molecular signature of specific tumor subtypes. In addition, understanding the complex differences in the biology of NSCs and BCSCs is necessary for selective targeting of BCSCs. For instance, designing therapeutic strategies to target mutation present only in BCSCs and selective targeting mechanisms of tumor propagation that are distinct from NSC regulation are possible strategies. For example, the cyclopamine (inhibitor of Hh pathway) is inactive in normal cells due to expression of patched (Ptc) gene. Ptc gene products are reported to prevent binding of cyclopamine to its target. However, tumor cells respond well to treatment with cyclopamine due to mutations in Ptc gene. Thus, cyclopamine was expected to selectively kill tumor cells (Goodrich and Scott 1998; Borah et al. 2015). In recent years, researchers have also focused on designing suitable nano-drug delivery systems to specifically target BCSCs. The application of nanocarriers for BCSC-targeting, however, is in its infancy, and many issues need to be well studied in clinical settings (Pindiprolu et al. 2017). Due to complex signaling network and their high dynamic plasticity according to the need of the environment, there are greater chances for development of drug resistance in BCSCs. Multi-targeted anti-BCSC agents need to be designed, therefore, to overcome drug resistance. Various pathways of BCSCs like Hh, notch, and Wnt have multiple points which can be targeted simultaneously. The existence of cross talk among these signaling pathways needs to be understood for designing novel therapeutic agents for targeting BCSCs (Bashyal Insan and Jaitak 2014). Accumulating body of evidences also suggests that although BCSCs are eliminated, non-BCSCs which are left behind will revert back to acquire characteristics of BCSCs. Combination therapy with chemotherapeutic agents and anti-BCSC agent is, therefore, needed to achieve a radical cure.
Conclusion
Compelling body of evidence suggest that the presence of BCSCs is the major cause of tumor relapse, metastasis, and chemoresistance in breast cancer. As discussed in the review, many molecular targets have been identified in BCSCs. They include surface markers, self-renewal pathways, apoptotic pathways, autophagy, metabolism, and microenvironment. The current research is focused on developing anticancer agents against these targets to eliminate BCSCs and to achieve radical cure in breast cancer therapy. The identification of strategies that take advantage of these targets of BCSCs needs to be well studied.
Abbreviations
- ABC:
-
ATP-binding cassette
- ATM:
-
Ataxia telangiectasia-mutated serine/threonine kinase
- Bcl2:
-
B cell lymphoma 2
- BCRP:
-
Breast cancer resistance protein
- BCSCs:
-
Breast cancer stem cells
- BIK:
-
Bcl2 interacting killer
- Bmi-1:
-
B cell-specific Moloney murine leukemia virus integration site 1
- BMP2:
-
Bone morphogenetic protein 2
- CAIX:
-
Carbonic anhydrase-IX
- CAT:
-
Catalase
- CSL:
-
CBF-1/RBPJ-κ in Homo sapiens/Mus musculus, respectively, Suppressor of hairless in Drosophila melanogaster, Lag-1 in Caenorhabditis elegans
- CDK:
-
Cyclin-dependent kinases
- ChKs:
-
Checkpoint kinases
- c-myc:
-
C-terminus of myc protein
- DHh:
-
Desert Hedgehog
- DLL4:
-
Delta-like 4 ligand
- DOX:
-
Doxorubicin
- DRs:
-
Death receptors
- EMT:
-
Epithelial-to-mesenchymal transition
- EpICD:
-
EpCAM intracellular domain
- FAS:
-
Fatty acid synthase
- FTC:
-
Fumitremorgin C
- Gl:
-
Glioma-associated oncogene
- GLUT:
-
Glucose transporter
- GM-CSF:
-
Granulocyte-macrophage colony-stimulating factor
- GPO:
-
Glutathione peroxidase
- GSK-3β:
-
Glycogen synthase kinase 3 β
- HDR:
-
Homology-directed recombination
- Hh:
-
Hedgehog
- HK:
-
Hexose kinase
- IHh:
-
Indian Hedgehog
- IL:
-
Interleukin
- JAK:
-
Janus kinase
- LRP:
-
Low-density lipoprotein-related receptor
- MAML:
-
Mastermind like
- m-TOR:
-
Mammalian target of rapamycin
- Nanog:
-
Gene named after the Tír na nÓg legend
- NHEJ:
-
Nonhomologous end joining
- NICD:
-
Notch intracellular domain
- non-BCSCs:
-
Non-breast cancer stem cells or bulk tumor cells
- Oct-4:
-
Octamer-binding transcription factor
- PDZ:
-
Disheveled PDZ domain
- PI3-k:
-
Phosphoinositide 3-kinase
- PTEN:
-
Phosphatase and tensin homolog
- SCs:
-
Normal stem cells
- SHh:
-
Sonic Hedgehog
- SMADs:
-
Homologs of Sma and MAD proteins
- Smo:
-
Smoothened
- SOD:
-
Superoxide dismutase
- SOX:
-
Sry-related HMG box
- STAT:
-
Signal transducers and activators of transcription
- STAT3:
-
Signal transducer and activator of transcription factor 3
- TGF-β:
-
Transforming growth factor-β
- TR:
-
Thio-redoxin
References
Ablett MP, Singh JK, Clarke RB (2012) Stem cells in breast tumours: are they ready for the clinic? Eur J Cancer 48:2104–2116
Akira S, Nishio Y, Inoue M, Wang X-J, We S, Matsusaka T, Yoshida K, Sudo T, Naruto M, Kishimoto T (1994) Molecular cloning of APRF, a novel IFN-stimulated gene factor 3 p91-related transcription factor involved in the gp130-mediated signaling pathway. Cell 77:63–71
Al-Assar O, Mantoni T, Lunardi S, Kingham G, Helleday T, Brunner TB (2011) Breast cancer stem-like cells show dominant homologous recombination due to a larger S-G2 fraction. Cancer Biol Ther 11:1028–1035
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci 100:3983–3988
Alkema M, Wiegant J, Raap AK, Bems A, van Lohuizen M (1993) Characterization and chromosomal localization of the human proto-oncogene BMI-1. Hum Mol Genet 2:1597–1603
Androutsellis-Theotokis A, Leker RR, Soldner F, Hoeppner DJ, Ravin R, Poser SW, Rueger MA, Bae S-K, Kittappa R, McKay RD (2006) Notch signalling regulates stem cell numbers in vitro and in vivo. Nature 442:823–826
Artavanis-Tsakonas S, Rand MD, Lake RJ (1999) Notch signaling: cell fate control and signal integration in development. Science 284:770–776
Atkinson RL, Yang WT, Rosen DG, Landis MD, Wong H, Lewis MT, Creighton CJ, Sexton KR, Hilsenbeck SG, Sahin AA (2013) Cancer stem cell markers are enriched in normal tissue adjacent to triple negative breast cancer and inversely correlated with DNA repair deficiency. Breast Cancer Res 15:R77
Balic M, Lin H, Young L, Hawes D, Giuliano A, McNamara G, Datar RH, Cote RJ (2006) Most early disseminated cancer cells detected in bone marrow of breast cancer patients have a putative breast cancer stem cell phenotype. Clin Cancer Res 12:5615–5621
Bashyal Insan M, Jaitak V (2014) New approaches to target cancer stem cells: current scenario. Mini Rev Med Chem 14:20–34
Benvenuto M, Masuelli L, De Smaele E, Fantini M, Mattera R, Cucchi D, Bonanno E, Di Stefano E, Frajese GV, Orlandi A (2016) In vitro and in vivo inhibition of breast cancer cell growth by targeting the Hedgehog/GLI pathway with SMO (GDC-0449) or GLI (GANT-61) inhibitors. Oncotarget 7:9250
Borah A, Raveendran S, Rochani A, Maekawa T, Kumar D (2015) Targeting self-renewal pathways in cancer stem cells: clinical implications for cancer therapy. Oncogene 4:e177
Bostad M, Olsen CE, Peng Q, Berg K, Høgset A, Selbo PK (2015) Light-controlled endosomal escape of the novel CD133-targeting immunotoxin AC133–saporin by photochemical internalization—a minimally invasive cancer stem cell-targeting strategy. J Control Release 206:37–48
Bouras T, Pal B, Vaillant F, Harburg G, Asselin-Labat M-L, Oakes SR, Lindeman GJ, Visvader JE (2008) Notch signaling regulates mammary stem cell function and luminal cell-fate commitment. Cell Stem Cell 3:429–441
Bourguignon LY, Zhu H, Shao L, Chen YW (2000) CD44 interaction with tiam1 promotes Rac1 signaling and hyaluronic acid-mediated breast tumor cell migration. J Biol Chem 275:1829–1838
Bourguignon LY, Peyrollier K, Xia W, Gilad E (2008) Hyaluronan-CD44 interaction activates stem cell marker Nanog, Stat-3-mediated MDR1 gene expression, and ankyrin-regulated multidrug efflux in breast and ovarian tumor cells. J Biol Chem 283:17635–17651
Bourguignon LY, Spevak CC, Wong G, Xia W, Gilad E (2009) Hyaluronan-CD44 interaction with protein kinase Cϵ promotes oncogenic signaling by the stem cell marker Nanog and the production of microRNA-21, leading to down-regulation of the tumor suppressor protein PDCD4, anti-apoptosis, and chemotherapy resistance in breast tumor cells. J Biol Chem 284:26533–26546
Bozorgi A, Khazaei M, Khazaei MR (2015) New findings on breast cancer stem cells: a review. J Breast Cancer 18:303–312
Branda M, Wands JR (2006) Signal transduction cascades and hepatitis B and C related hepatocellular carcinoma. Hepatology 43:891–902
Brandsma I, van Gent DC (2012) Pathway choice in DNA double strand break repair: observations of a balancing act. Genome Integr 3:9
Bray SJ (2006) Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 7:678–689
Britton K, Eyre R, Harvey I, Stemke-Hale K, Browell D, Lennard T, Meeson A (2012) Breast cancer, side population cells and ABCG2 expression. Cancer Lett 323:97–105
Brown JM, Attardi LD (2005) The role of apoptosis in cancer development and treatment response. Nat Rev Cancer 5:231–237
Bruna A, Greenwood W, Le Quesne J, Teschendorff A, Miranda-Saavedra D, Rueda OM, Sandoval JL, Vidakovic AT, Saadi A, Pharoah P (2012) TGFβ induces the formation of tumour-initiating cells in claudin low breast cancer. Nat Commun 3:1055
Cerdan C, Bhatia M (2010) Novel roles for Notch, Wnt and Hedgehog in hematopoesis derived from human pluripotent stem cells. Int J Dev Biol 54:955–964
Chapellier M, Maguer-Satta V (2016) BMP2, a key to uncover luminal breast cancer origin linked to pollutant effects on epithelial stem cells niche. Mol Cell Oncol 3:e1026527
Chen B, Dodge ME, Tang W, Lu J, Ma Z, Fan C-W, Wei S, Hao W, Kilgore J, Williams NS (2009) Small molecule–mediated disruption of Wnt-dependent signaling in tissue regeneration and cancer. Nat Chem Biol 5:100–107
Chen D, Pamu S, Cui Q, Chan TH, Dou QP (2012) Novel epigallocatechin gallate (EGCG) analogs activate AMP-activated protein kinase pathway and target cancer stem cells. Bioorg Med Chem 20:3031–3037
Chen K, Huang Y-h, J-l C (2013) Understanding and targeting cancer stem cells: therapeutic implications and challenges. Acta Pharmacol Sin 34:732–740
Cheung-Ong K, Giaever G, Nislow C (2013) DNA-damaging agents in cancer chemotherapy: serendipity and chemical biology. Chem Biol 20:648–659
Choi H, Chun Y-S, Kim T-Y, Park J-W (2010) HIF-2α enhances β-catenin/TCF-driven transcription by interacting with β-catenin. Cancer Res 70:10101–10111
Choi AM, Ryter SW, Levine B (2013) Autophagy in human health and disease. N Engl J Med 368:651–662
Chung SS, Vadgama JV (2015) Curcumin and epigallocatechin gallate inhibit the cancer stem cell phenotype via down-regulation of STAT3–NFκB signaling. Anticancer Res 35:39–46
Clarke MF, Dick JE, Dirks PB, Eaves CJ, Jamieson CH, Jones DL, Visvader J, Weissman IL, Wahl GM (2006) Cancer stem cells—perspectives on current status and future directions: AACR workshop on cancer stem cells. Cancer Res 66:9339–9344
Cook JA, Gius D, Wink DA, Krishna MC, Russo A, Mitchell JB (2004) Oxidative stress, redox, and the tumor microenvironment. Semin Radiat Oncol 14(3):259–266
Croker AK, Allan AL (2012) Inhibition of aldehyde dehydrogenase (ALDH) activity reduces chemotherapy and radiation resistance of stem-like ALDHhiCD44+ human breast cancer cells. Breast Cancer Res Treat 133:75–87
Croker AK, Goodale D, Chu J, Postenka C, Hedley BD, Hess DA, Allan AL (2009) High aldehyde dehydrogenase and expression of cancer stem cell markers selects for breast cancer cells with enhanced malignant and metastatic ability. J Cell Mol Med 13:2236–2252
Culig Z (2011) Cytokine disbalance in common human cancers. Biochim Biophys Acta (BBA)-Mol Cell Res 1813:308–314
Darnell JE Jr, Kerr IM, Stark GR (1994) Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 264:1415–1420
Deng X, Wu X, Weng H, Song F (2016) The siRNA-mediated silencing of Bmi-1 promotes apoptosis and inhibits invasion of MCF-7 breast cancer cells. Xi Bao Yu Fen Zi Mian Yi Xue Za Zhi= Chin J Cell Mol Immunol 32:1036
Diehn M, Cho RW, Lobo NA, Kalisky T, Dorie MJ, Kulp AN, Qian D, Lam JS, Ailles LE, Wong M (2009) Association of reactive oxygen species levels and radioresistance in cancer stem cells. Nature 458:780–783
Domen J, Gandy KL, Weissman IL (1998) Systemic overexpression of BCL-2 in the hematopoietic system protects transgenic mice from the consequences of lethal irradiation. Blood 91:2272–2282
Dong C, Wu Y, Yao J, Wang Y, Yu Y, Rychahou PG, Evers BM, Zhou BP (2012) G9a interacts with Snail and is critical for Snail-mediated E-cadherin repression in human breast cancer. J Clin Invest 122:1469–1486
Dong C, Yuan T, Wu Y, Wang Y, Fan TW, Miriyala S, Lin Y, Yao J, Shi J, Kang T (2013) Loss of FBP1 by Snail-mediated repression provides metabolic advantages in basal-like breast cancer. Cancer Cell 23:316–331
Dontu G, Al-Hajj M, Abdallah WM, Clarke MF, Wicha MS (2003) Stem cells in normal breast development and breast cancer. Cell Prolif 36:59–72
Epenetos A, Kousparou C, Stylianou S (2009) Inhibition of Notch signaling for the treatment of human carcinomas. Cancer Research. Amer Assoc Cancer Research 615 Chestnut St, 17th floor, Philadelphia, PA 19106-4404 USA
Ferlay J, Héry C, Autier P, Sankaranarayanan R (2010) Global burden of breast cancer. In: Li C (ed) Breast cancer epidemiology. Springer, New York
Fillmore CM, Kuperwasser C (2008) Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res 10:1
Fleming HE, Janzen V, Celso CL, Guo J, Leahy KM, Kronenberg HM, Scadden DT (2008) Wnt signaling in the niche enforces hematopoietic stem cell quiescence and is necessary to preserve self-renewal in vivo. Cell Stem Cell 2:274–283
Gil J, Bernard D, Peters G (2005) Role of polycomb group proteins in stem cell self-renewal and cancer. DNA Cell Biol 24:117–125
Gilbertson RJ, Rich JN (2007) Making a tumour's bed: glioblastoma stem cells and the vascular niche. Nat Rev Cancer 7:733–736
Gilboa-Geffen A, Hamar P, Le MT, Wheeler LA, Trifonova R, Petrocca F, Wittrup A, Lieberman J (2015) Gene knockdown by EpCAM aptamer–siRNA chimeras suppresses epithelial breast cancers and their tumor-initiating cells. Mol Cancer Ther 14:2279–2291
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG, Liu S (2007) ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 1:555–567
Ginestier C, Liu S, Diebel ME, Korkaya H, Luo M, Brown M, Wicinski J, Cabaud O, Charafe-Jauffret E, Birnbaum D (2010) CXCR1 blockade selectively targets human breast cancer stem cells in vitro and in xenografts. J Clin Invest 120:485–497
Ginestier C, Monville F, Wicinski J, Cabaud O, Cervera N, Josselin E, Finetti P, Guille A, Larderet G, Viens P (2012) Mevalonate metabolism regulates basal breast cancer stem cells and is a potential therapeutic target. Stem Cells 30:1327–1337
Gong C, Bauvy C, Tonelli G, Yue W, Delomenie C, Nicolas V, Zhu Y, Domergue V, Marin-Esteban V, Tharinger H (2013) Beclin 1 and autophagy are required for the tumorigenicity of breast cancer stem-like/progenitor cells. Oncogene 32:2261–2272
Goodarzi N, Ghahremani MH, Amini M, Atyabi F, Ostad SN, Shabani Ravari N, Nateghian N, Dinarvand R (2014) CD44-targeted docetaxel conjugate for cancer cells and cancer stem-like cells: a novel hyaluronic acid-based drug delivery system. Chem Biol Drug Des 83:741–752
Goodrich LV, Scott MP (1998) Hedgehog and patched in neural development and disease. Neuron 21:1243–1257
Green JL, La J, Yum KW, Desai P, Rodewald L-W, Zhang X, Leblanc M, Nusse R, Lewis MT, Wahl GM (2013) Paracrine Wnt signaling both promotes and inhibits human breast tumor growth. Proc Natl Acad Sci 110:6991–6996
Greenow K, Clarke AR (2012) Controlling the stem cell compartment and regeneration in vivo: the role of pluripotency pathways. Physiol Rev 92:75–99
Grudzien P, Lo S, Albain KS, Robinson P, Rajan P, Strack PR, Golde TE, Miele L, Foreman KE (2010) Inhibition of Notch signaling reduces the stem-like population of breast cancer cells and prevents mammosphere formation. Anticancer Res 30:3853–3867
Gurney A, Axelrod F, Bond CJ, Cain J, Chartier C, Donigan L, Fischer M, Chaudhari A, Ji M, Kapoun AM (2012) Wnt pathway inhibition via the targeting of frizzled receptors results in decreased growth and tumorigenicity of human tumors. Proc Natl Acad Sci 109:11717–11722
Harrison H, Farnie G, Brennan KR, Clarke RB (2010a) Breast cancer stem cells: something out of notching? Cancer Res 70:8973–8976
Harrison H, Farnie G, Howell SJ, Rock RE, Stylianou S, Brennan KR, Bundred NJ, Clarke RB (2010b) Regulation of breast cancer stem cell activity by signaling through the Notch4 receptor. Cancer Res 70:709–718
He L, Gu J, Lim LY, Yuan ZX, Mo J (2016) Nanomedicine-mediated therapies to target breast cancer stem cells. Front Pharmacol 7:313
Heim MH, Kerr IM, Stark GR, Darnell Jr JE (1995) Contribution of STAT SH2 groups to specific interferon signaling by the Jak-STAT pathway. Science 267:1347–1349
Hirsch HA, Iliopoulos D, Tsichlis PN, Struhl K (2009) Metformin selectively targets cancer stem cells, and acts together with chemotherapy to block tumor growth and prolong remission. Cancer Res 69:7507–7511
Hoey T, Yen W-C, Axelrod F, Basi J, Donigian L, Dylla S, Fitch-Bruhns M, Lazetic S, Park I-K, Sato A (2009) DLL4 blockade inhibits tumor growth and reduces tumor-initiating cell frequency. Cell Stem Cell 5:168–177
Honeth G, Bendahl P-O, Ringnér M, Saal LH, Gruvberger-Saal SK, Lövgren K, Grabau D, Fernö M, Borg Å, Hegardt C (2008) The CD44+/CD24-phenotype is enriched in basal-like breast tumors. Breast Cancer Res 10:1
Honma N, Horii R, Ito Y, Saji S, Younes M, Iwase T, Akiyama F (2015) Differences in clinical importance of Bcl-2 in breast cancer according to hormone receptors status or adjuvant endocrine therapy. BMC Cancer 15:1
Hori K, Sen A, Artavanis-Tsakonas S (2013) Notch signaling at a glance. J Cell Sci 126:2135–2140
Hou Z-J, Luo X, Zhang W, Peng F, Cui B, Wu S-J, Zheng F-M, Xu J, Xu L-Z, Long Z-J (2015) Flubendazole, FDA-approved anthelmintic, targets breast cancer stem-like cells. Oncotarget 6:6326
Hu C, Niestroj M, Yuan D, Chang S, Chen J (2015) Treating cancer stem cells and cancer metastasis using glucose-coated gold nanoparticles. Int J Nanomedicine 10:2065
Ihle JN (2001) The Stat family in cytokine signaling. Curr Opin Cell Biol 13:211–217
Ihle JN, Kerr IM (1995) Jaks and Stats in signaling by the cytokine receptor superfamily. Trends Genet 11:69–74
Ihle JN, Witthuhn BA, Quelle FW, Yamamoto K, Thierfelder WE, Kreider B, Silvennoinen O (1994) Signaling by the cytokine receptor superfamily: JAKs and STATs. Trends Biochem Sci 19:222–227
Imrich S, Hachmeister M, Gires O (2012) EpCAM and its potential role in tumor-initiating cells. Cell Adhes Migr 6:30–38
Ingham PW, McMahon AP (2001) Hedgehog signaling in animal development: paradigms and principles. Genes Dev 15:3059–3087
Jackson SP (2002) Sensing and repairing DNA double-strand breaks. Carcinogenesis 23:687–696
Jacobs JJ, Kieboom K, Marino S, DePinho RA, van Lohuizen M (1999) The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature 397:164–168
Jain K, Paranandi KS, Sridharan S, Basu A (2013) Autophagy in breast cancer and its implications for therapy. Am J Cancer Res 3:251–265
Jang G-B, Hong I-S, Kim R-J, Lee S-Y, Park S-J, Lee E-S, Park JH, Yun C-H, Chung J-U, Lee K-J (2015) WNT/β-catenin small-molecule inhibitor CWP232228 preferentially inhibits the growth of breast cancer stem-like cells. Cancer Res 75:1691–1702
Jasin M, Rothstein R (2013) Repair of strand breaks by homologous recombination. Cold Spring Harb Perspect Biol 5:a012740
Kai K, Arima Y, Kamiya T, Saya H (2010) Breast cancer stem cells. Breast Cancer 17:80–85
Karnoub AE, Dash AB, Vo AP, Sullivan A, Brooks MW, Bell GW, Richardson AL, Polyak K, Tubo R, Weinberg RA (2007) Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature 449:557–563
Ke X-Y, Ng VWL, Gao S-J, Tong YW, Hedrick JL, Yang YY (2014) Co-delivery of thioridazine and doxorubicin using polymeric micelles for targeting both cancer cells and cancer stem cells. Biomaterials 35:1096–1108
Kessenbrock K, Dijkgraaf GJ, Lawson DA, Littlepage LE, Shahi P, Pieper U, Werb Z (2013) A role for matrix metalloproteinases in regulating mammary stem cell function via the Wnt signaling pathway. Cell Stem Cell 13:300–313
Kim JB, Ko E, Han W, Shin I, Park SY, Noh D-Y (2008) Berberine diminishes the side population and ABCG2 transporter expression in MCF-7 breast cancer cells. Planta Med 74:1693–1700
Kim S-Y, Rhee JG, Song X, Prochownik EV, Spitz DR, Lee YJ (2012) Breast cancer stem cell-like cells are more sensitive to ionizing radiation than non-stem cells: role of ATM. PLoS One 7:e50423
Kim S-Y, Kang JW, Song X, Kim BK, Yoo YD, Kwon YT, Lee YJ (2013) Role of the IL-6-JAK1-STAT3-Oct-4 pathway in the conversion of non-stem cancer cells into cancer stem-like cells. Cell Signal 25:961–969
Klonisch T, Wiechec E, Hombach-Klonisch S, Ande SR, Wesselborg S, Schulze-Osthoff K, Los M (2008) Cancer stem cell markers in common cancers–therapeutic implications. Trends Mol Med 14:450–460
Königsberg R, Obermayr E, Bises G, Pfeiler G, Gneist M, Wrba F, De Santis M, Zeillinger R, Hudec M, Dittrich C (2011) Detection of EpCAM positive and negative circulating tumor cells in metastatic breast cancer patients. Acta Oncol 50:700–710
Korkaya H, Paulson A, Charafe-Jauffret E, Ginestier C, Brown M, Dutcher J, Clouthier SG, Wicha MS (2009) Regulation of mammary stem/progenitor cells by PTEN/Akt/β-catenin signaling. PLoS Biol 7:e1000121
Korkaya H, Kim G-I, Davis A, Malik F, Henry NL, Ithimakin S, Quraishi AA, Tawakkol N, D'Angelo R, Paulson AK (2012) Activation of an IL6 inflammatory loop mediates trastuzumab resistance in HER2+ breast cancer by expanding the cancer stem cell population. Mol Cell 47:570–584
Kubo M, Nakamura M, Tasaki A, Yamanaka N, Nakashima H, Nomura M, Kuroki S, Katano M (2004) Hedgehog signaling pathway is a new therapeutic target for patients with breast cancer. Cancer Res 64:6071–6074
Lai Y, Yu X, Lin X, He S (2016) Inhibition of mTOR sensitizes breast cancer stem cells to radiation-induced repression of self-renewal through the regulation of MnSOD and Akt. Int J Mol Med 37:369–377
Lang J-Y, Hsu JL, Meric-Bernstam F, Chang C-J, Wang Q, Bao Y, Yamaguchi H, Xie X, Woodward WA, Yu D (2011) BikDD eliminates breast cancer initiating cells and synergizes with lapatinib for breast cancer treatment. Cancer Cell 20:341–356
Leon G, MacDonagh L, Finn SP, Cuffe S, Barr MP (2016) Cancer stem cells in drug resistant lung cancer: targeting cell surface markers and signaling pathways. Pharmacol Ther 158:71–90
Li Z (2013) CD133: a stem cell biomarker and beyond. Exp Hematol Oncol 2:1
Li Y, Zhang T, Korkaya H, Liu S, Lee H-F, Newman B, Yu Y, Clouthier SG, Schwartz SJ, Wicha MS (2010) Sulforaphane, a dietary component of broccoli/broccoli sprouts, inhibits breast cancer stem cells. Clin Cancer Res 16:2580–2590
Li R-J, Ying X, Zhang Y, Ju R-J, Wang X-X, Yao H-J, Men Y, Tian W, Yu Y, Zhang L (2011) All-trans retinoic acid stealth liposomes prevent the relapse of breast cancer arising from the cancer stem cells. J Control Release 149:281–291
Li R, You S, Hu Z, Chen ZG, Sica GL, Khuri FR, Curran WJ, Shin DM, Deng X (2013) Inhibition of STAT3 by niclosamide synergizes with erlotinib against head and neck cancer. PLoS One 8:e74670
Li Y, Li P-K, Roberts MJ, Arend RC, Samant RS, Buchsbaum DJ (2014) Multi-targeted therapy of cancer by niclosamide: a new application for an old drug. Cancer Lett 349:8–14
Liang DH, Choi DS, Ensor JE, Kaipparettu BA, Bass BL, Chang JC (2016) The autophagy inhibitor chloroquine targets cancer stem cells in triple negative breast cancer by inducing mitochondrial damage and impairing DNA break repair. Cancer Lett 376:249–258
Lim E, Vaillant F, Wu D, Forrest NC, Pal B, Hart AH, Asselin-Labat M-L, Gyorki DE, Ward T, Partanen A (2009) Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat Med 15:907–913
Lima RT, Martins LM, Guimaraes JE, Sambade C, Vasconcelos MH (2004) Specific downregulation of bcl-2 and xIAP by RNAi enhances the effects of chemotherapeutic agents in MCF-7 human breast cancer cells. Cancer Gene Ther 11:309–316
Liu S, Dontu G, Mantle ID, Patel S, Ahn N-S, Jackson KW, Suri P, Wicha MS (2006) Hedgehog signaling and Bmi-1 regulate self-renewal of normal and malignant human mammary stem cells. Cancer Res 66:6063–6071
Liu Z, Bandyopadhyay A, Nichols RW, Wang L, Hinck AP, Wang S, Sun L-Z (2012) Blockade of autocrine TGF-β signaling inhibits stem cell phenotype, survival, and metastasis of murine breast cancer cells. J Stem Cell Res Ther 2:1
Liu T, Sun B, Zhao X, Zhao X, Sun T, Gu Q, Yao Z, Dong X, Zhao N, Liu N (2013) CD133+ cells with cancer stem cell characteristics associates with vasculogenic mimicry in triple-negative breast cancer. Oncogene 32:544–553
Liu J, Chen X, Ward T, Mao Y, Bockhorn J, Liu X, Wang G, Pegram M, Shen K (2016a) Niclosamide inhibits epithelial-mesenchymal transition and tumor growth in lapatinib-resistant human epidermal growth factor receptor 2-positive breast cancer. Int J Biochem Cell Biol 71:12–23
Liu J, Chen X, Ward T, Pegram M, Shen K (2016b) Combined niclosamide with cisplatin inhibits epithelial-mesenchymal transition and tumor growth in cisplatin-resistant triple-negative breast cancer. Tumor Biol 37(7):9825–9835
Liu M, Zhang W, Tang W, Wang Y, Zhao X, Wang X, Qi X, Li J (2016c) Isocyclopamine, a novel synthetic derivative of cyclopamine, reverts doxorubicin resistance in MCF-7/ADR cells by increasing intracellular doxorubicin accumulation and downregulating breast cancer stem-like cells. Tumor Biol 37:1919–1931
Lock F, McDonald P, Lou Y, Serrano I, Chafe S, Ostlund C, Aparicio S, Winum J-Y, Supuran C, Dedhar S (2013) Targeting carbonic anhydrase IX depletes breast cancer stem cells within the hypoxic niche. Oncogene 32:5210–5219
Londoño-Joshi AI, Oliver PG, Li Y, Lee CH, Forero-Torres A, LoBuglio AF, Buchsbaum DJ (2012) Basal-like breast cancer stem cells are sensitive to anti-DR5 mediated cytotoxicity. Breast Cancer Res Treat 133:437–445
Louderbough JM, Schroeder JA (2011) Understanding the dual nature of CD44 in breast cancer progression. Mol Cancer Res 9:1573–1586
MacDonald BT, Tamai K, He X (2009) Wnt/β-catenin signaling: components, mechanisms, and diseases. Dev Cell 17:9–26
Mai TT, Moon J, Song Y, Viet PQ, Van Phuc P, Lee JM, Yi T-H, Cho M, Cho SK (2012) Ginsenoside F2 induces apoptosis accompanied by protective autophagy in breast cancer stem cells. Cancer Lett 321:144–153
Mani SA, Guo W, Liao M-J, Eaton EN, Ayyanan A, Zhou AY, Brooks M, Reinhard F, Zhang CC, Shipitsin M (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133:704–715
Marini C, Salani B, Massollo M, Amaro A, Esposito AI, Maria Orengo A, Capitanio S, Emionite L, Riondato M, Bottoni G (2013) Direct inhibition of hexokinase activity by metformin at least partially impairs glucose metabolism and tumor growth in experimental breast cancer. Cell Cycle 12:3490–3499
Marotta LL, Almendro V, Marusyk A, Shipitsin M, Schemme J, Walker SR, Bloushtain-Qimron N, Kim JJ, Choudhury SA, Maruyama R (2011) The JAK2/STAT3 signaling pathway is required for growth of CD44+ CD24–stem cell–like breast cancer cells in human tumors. J Clin Invest 121:2723–2735
Martinou J-C, Youle RJ (2011) Mitochondria in apoptosis: Bcl-2 family members and mitochondrial dynamics. Dev Cell 21:92–101
Massagué J (2000) How cells read TGF-β signals. Nat Rev Mol Cell Biol 1:169–178
Merino D, Lok S, Visvader J, Lindeman G (2016) Targeting BCL-2 to enhance vulnerability to therapy in estrogen receptor-positive breast cancer. Oncogene 35:1877–1887
Micchelli CA, Selva E, Mogila V, Perrimon N (2002) Rasp, a putative transmembrane acyltransferase, is required for Hedgehog signaling. Development 129:843–851
Moitra K (2015) Overcoming multidrug resistance in cancer stem cells. BioMed Res Int 8:635745. https://doi.org/10.1155/2015/635745
Molyneux G, Geyer FC, Magnay F-A, McCarthy A, Kendrick H, Natrajan R, MacKay A, Grigoriadis A, Tutt A, Ashworth A (2010) BRCA1 basal-like breast cancers originate from luminal epithelial progenitors and not from basal stem cells. Cell Stem Cell 7:403–417
Moses H, Barcellos-Hoff MH (2011) TGF-β biology in mammary development and breast cancer. Cold Spring Harb Perspect Biol 3:a003277
Muntimadugu E, Kumar R, Saladi S, Rafeeqi TA, Khan W (2016) CD44 targeted chemotherapy for co-eradication of breast cancer stem cells and cancer cells using polymeric nanoparticles of salinomycin and paclitaxel. Colloids Surf B: Biointerfaces 143:532–546
Munz M, Baeuerle PA, Gires O (2009) The emerging role of EpCAM in cancer and stem cell signaling. Cancer Res 69:5627–5629
Niida H, Nakanishi M (2006) DNA damage checkpoints in mammals. Mutagenesis 21:3–9
Nishita M, Hashimoto MK, Ogata S, Laurent MN, Ueno N, Shibuya H, Cho KW (2000) Interaction between Wnt and TGF-β signalling pathways during formation of Spemann's organizer. Nature 403:781–785
Noguera-Troise I, Daly C, Papadopoulos NJ, Coetzee S, Boland P, Gale NW, Lin HC, Yancopoulos GD, Thurston G (2006) Blockade of Dll4 inhibits tumour growth by promoting non-productive angiogenesis. Nature 444:1032–1037
Ojha R, Bhattacharyya S, Singh SK (2015) Autophagy in cancer stem cells: a potential link between chemoresistance, recurrence, and metastasis. Biores Open Access 4:97–108
Osta WA, Chen Y, Mikhitarian K, Mitas M, Salem M, Hannun YA, Cole DJ, Gillanders WE (2004) EpCAM is overexpressed in breast cancer and is a potential target for breast cancer gene therapy. Cancer Res 64:5818–5824
Owens TW, Naylor MJ (2013) Breast cancer stem cells. Front Physiol 4:225
Padua D, Massagué J (2009) Roles of TGFβ in metastasis. Cell Res 19:89–102
Pan J-X, Ding K, Wang C-Y (2012) Niclosamide, an old antihelminthic agent, demonstrates antitumor activity by blocking multiple signaling pathways of cancer stem cells. Chin J Cancer 31:178–184
Pandey PR, Okuda H, Watabe M, Pai SK, Liu W, Kobayashi A, Xing F, Fukuda K, Hirota S, Sugai T (2011) Resveratrol suppresses growth of cancer stem-like cells by inhibiting fatty acid synthase. Breast Cancer Res Treat 130:387–398
Patra KC, Wang Q, Bhaskar PT, Miller L, Wang Z, Wheaton W, Chandel N, Laakso M, Muller WJ, Allen EL (2013) Hexokinase 2 is required for tumor initiation and maintenance and its systemic deletion is therapeutic in mouse models of cancer. Cancer Cell 24:213–228
Patrawala L, Calhoun T, Schneider-Broussard R, Zhou J, Claypool K, Tang DG (2005) Side population is enriched in tumorigenic, stem-like cancer cells, whereas ABCG2+ and ABCG2− cancer cells are similarly tumorigenic. Cancer Res 65:6207–6219
Peired AJ, Sisti A, Romagnani P (2016) Renal cancer stem cells: characterization and targeted therapies. Stem Cells Int 2016:1–12
Peitzsch C, Kurth I, Kunz-Schughart L, Baumann M, Dubrovska A (2013) Discovery of the cancer stem cell related determinants of radioresistance. Radiother Oncol 108:378–387
Phillips TM, McBride WH, Pajonk F (2006) The response of CD24−/low/CD44+ breast cancer–initiating cells to radiation. J Natl Cancer Inst 98:1777–1785
Pindiprolu S, Krishnamurthy P, Chintamaneni PK, Karri VVSR (2017) Nanocarrier based approaches for targeting breast cancer stem cells. Artif Cells Nanomeds Biotechnol. https://doi.org/10.1080/21691401.2017.1366337
Prud'homme GJ, Glinka Y, Toulina A, Ace O, Subramaniam V, Jothy S (2010) Breast cancer stem-like cells are inhibited by a non-toxic aryl hydrocarbon receptor agonist. PLoS One 5:e13831
Pryce BA, Watson SS, Murchison ND, Staverosky JA, Dünker N, Schweitzer R (2009) Recruitment and maintenance of tendon progenitors by TGFβ signaling are essential for tendon formation. Development 136:1351–1361
Raaijmakers MH, de Grouw EP, Heuver LH, van der Reijden BA, Jansen JH, Scheper RJ, Scheffer GL, de Witte TJ, Raymakers RA (2005) Breast cancer resistance protein in drug resistance of primitive CD34+ 38− cells in acute myeloid leukemia. Clin Cancer Res 11:2436–2444
Rabindran SK, He H, Singh M, Brown E, Collins KI, Annable T, Greenberger LM (1998) Reversal of a novel multidrug resistance mechanism in human colon carcinoma cells by fumitremorgin C. Cancer Res 58:5850–5858
Rabindran SK, Ross DD, Doyle LA, Yang W, Greenberger LM (2000) Fumitremorgin C reverses multidrug resistance in cells transfected with the breast cancer resistance protein. Cancer Res 60:47–50
Resetkova E, Reis-Filho JS, Jain RK, Mehta R, Thorat MA, Nakshatri H, Badve S (2010) Prognostic impact of ALDH1 in breast cancer: a story of stem cells and tumor microenvironment. Breast Cancer Res Treat 123:97–108
Reya T, Morrison SJ, Clarke MF, Weissman IL (2001) Stem cells, cancer, and cancer stem cells. Nature 414:105–111
Ridgway J, Zhang G, Wu Y, Stawicki S, Liang W-C, Chanthery Y, Kowalski J, Watts RJ, Callahan C, Kasman I (2006) Inhibition of Dll4 signalling inhibits tumour growth by deregulating angiogenesis. Nature 444:1083–1087
Rios AC, Fu NY, Lindeman GJ, Visvader JE (2014) In situ identification of bipotent stem cells in the mammary gland. Nature 506:322–327
Robey RW, Steadman K, Polgar O, Morisaki K, Blayney M, Mistry P, Bates SE (2004) Pheophorbide a is a specific probe for ABCG2 function and inhibition. Cancer Res 64:1242–1246
Rodriguez-Torres M, Allan AL (2016) Aldehyde dehydrogenase as a marker and functional mediator of metastasis in solid tumors. Clin Exp Metastasis 33:97–113
Sabe H (2011) Cancer early dissemination: cancerous epithelial–mesenchymal transdifferentiation and transforming growth factor β signalling. J Biochem 149:633–639
Salani B, Del Rio A, Marini C, Sambuceti G, Cordera R, Maggi D (2014) Metformin, cancer and glucose metabolism. Endocr Relat Cancer 21:R461–R471
Samanta D, Gilkes DM, Chaturvedi P, Xiang L, Semenza GL (2014) Hypoxia-inducible factors are required for chemotherapy resistance of breast cancer stem cells. Proc Natl Acad Sci 111:E5429–E5438
Sancho P, Barneda D, Heeschen C (2016) Hallmarks of cancer stem cell metabolism. Br J Cancer 114:1305–1312
Sawa M, Yamamoto K, Yokozawa T, Kiyoi H, Hishida A, Kajiguchi T, Seto M, Kohno A, Kitamura K, Itoh Y (2005) BMI-1 is highly expressed in M0-subtype acute myeloid leukemia. Int J Hematol 82:42–47
Scehnet JS, Jiang W, Kumar SR, Krasnoperov V, Trindade A, Benedito R, Djokovic D, Borges C, Ley EJ, Duarte A (2007) Inhibition of Dll4-mediated signaling induces proliferation of immature vessels and results in poor tissue perfusion. Blood 109:4753–4760
Scheel C, Eaton EN, Li SH-J, Chaffer CL, Reinhardt F, Kah K-J, Bell G, Guo W, Rubin J, Richardson AL (2011) Paracrine and autocrine signals induce and maintain mesenchymal and stem cell states in the breast. Cell 145:926–940
Schweisguth F (2004) Regulation of notch signaling activity. Curr Biol 14:R129–R138
Sena LA, Chandel NS (2012) Physiological roles of mitochondrial reactive oxygen species. Mol Cell 48:158–167
Shackleton M, Vaillant F, Simpson KJ, Stingl J, Smyth GK, Asselin-Labat M-L, Wu L, Lindeman GJ, Visvader JE (2006) Generation of a functional mammary gland from a single stem cell. Nature 439:84–88
Shervington A, Lu C (2008) Expression of multidrug resistance genes in normal and cancer stem cells. Cancer Investig 26:535–542
Signore M, Ricci-Vitiani L, De Maria R (2013) Targeting apoptosis pathways in cancer stem cells. Cancer Lett 332:374–382
Singh A, Settleman J (2010) EMT, cancer stem cells and drug resistance: an emerging axis of evil in the war on cancer. Oncogene 29:4741–4751
Singh BN, Kumar D, Shankar S, Srivastava RK (2012) Rottlerin induces autophagy which leads to apoptotic cell death through inhibition of PI3K/Akt/mTOR pathway in human pancreatic cancer stem cells. Biochem Pharmacol 84:1154–1163
Sinha K, Das J, Pal PB, Sil PC (2013) Oxidative stress: the mitochondria-dependent and mitochondria-independent pathways of apoptosis. Arch Toxicol 87:1157–1180
Stahl N, Farruggella TJ, Boulton TG, Zhong Z (1995) Choice of STATs and other substrates specified by modular tyrosine-based motifs in cytokine receptors. Science 267:1349–1353
Stallcup MR (2001) Role of protein methylation in chromatin remodeling and transcriptional regulation. Oncogene 20:3014–3020
Sui X, Chen R, Wang Z, Huang Z, Kong N, Zhang M, Han W, Lou F, Yang J, Zhang Q (2013) Autophagy and chemotherapy resistance: a promising therapeutic target for cancer treatment. Cell Death Dis 4:e838
Svärd J, Henricson KH, Persson-Lek M, Rozell B, Lauth M, Bergström Å, Ericson J, Toftgård R, Teglund S (2006) Genetic elimination of suppressor of fused reveals an essential repressor function in the mammalian Hedgehog signaling pathway. Dev Cell 10:187–197
Swaminathan SK, Roger E, Toti U, Niu L, Ohlfest JR, Panyam J (2013) CD133-targeted paclitaxel delivery inhibits local tumor recurrence in a mouse model of breast cancer. J Control Release 171:280–287
Takebe N, Harris PJ, Warren RQ, Ivy SP (2011) Targeting cancer stem cells by inhibiting Wnt, Notch, and Hedgehog pathways. Nat Rev Clin Oncol 8:97–106
Thakur R, Trivedi R, Rastogi N, Singh M, Mishra DP (2015) Inhibition of STAT3, FAK and Src mediated signaling reduces cancer stem cell load, tumorigenic potential and metastasis in breast cancer. Sci Rep 5:10194
Trachootham D, Alexandre J, Huang P (2009) Targeting cancer cells by ROS-mediated mechanisms: a radical therapeutic approach? Nat Rev Drug Discov 8:579–591
Tsai J, Jeng J, Chuang L, Yang M, Ho M, Chang W, Hsieh M, Lin Z, Tsai J (1997) Elevated urinary transforming growth factor-beta1 level as a tumour marker and predictor of poor survival in cirrhotic hepatocellular carcinoma. Br J Cancer 76:244–250
van Amerongen R, Bowman AN, Nusse R (2012) Developmental stage and time dictate the fate of Wnt/β-catenin-responsive stem cells in the mammary gland. Cell Stem Cell 11:387–400
Van der Lugt N, Domen J, Linders K, Van Roon M, Robanus-Maandag E, Te Riele H, Van der Valk M, Deschamps J, Sofroniew M, Van Lohuizen M (1994) Posterior transformation, neurological abnormalities, and severe hematopoietic defects in mice with a targeted deletion of the bmi-1 proto-oncogene. Genes Dev 8:757–769
Van Keymeulen A, Rocha AS, Ousset M, Beck B, Bouvencourt G, Rock J, Sharma N, Dekoninck S, Blanpain C (2011) Distinct stem cells contribute to mammary gland development and maintenance. Nature 479:189–193
Vazquez-Martin A, Oliveras-Ferraros C, Del Barco S, Martin-Castillo B, Menendez JA (2011) The anti-diabetic drug metformin suppresses self-renewal and proliferation of trastuzumab-resistant tumor-initiating breast cancer stem cells. Breast Cancer Res Treat 126:355–364
Vinogradov S, Wei X (2012) Cancer stem cells and drug resistance: the potential of nanomedicine. Nanomedicine 7:597–615
Visvader JE (2011) Cells of origin in cancer. Nature 469:314–322
Visvader JE, Rios A, Naiyang F et al (2014) The breast epithelial hierarchy and its implications for tumor heterogeneity. In Proceedings of the 105th annual meeting of the American Association for Cancer Research; 2014 Apr 5-9. AACR, San Diego
Wang X, Wang G, Zhao Y, Liu X, Ding Q, Shi J, Ding Y, Wang S (2012) STAT3 mediates resistance of CD44+ CD24−/low breast cancer stem cells to tamoxifen in vitro. J Biomed Res 26:325–335
Wang RA, Li ZS, Zhang HZ, Zheng PJ, Li QL, Shi JG, Yan QG, Ye J, Wang JB, Guo Y (2013a) Invasive cancers are not necessarily from preformed in situ tumours—an alternative way of carcinogenesis from misplaced stem cells. J Cell Mol Med 17:921–926
Wang Y-C, Chao T-K, Chang C-C, Yo Y-T, Yu M-H, Lai H-C (2013b) Drug screening identifies niclosamide as an inhibitor of breast cancer stem-like cells. PLoS One 8:e74538
Wang T, Fahrmann JF, Lee H, Li Y-J, Tripathi SC, Yue C, Zhang C, Lifshitz V, Song J, Yuan Y (2018) JAK/STAT3-regulated fatty acid β-oxidation is critical for breast cancer stem cell self-renewal and Chemoresistance. Cell Metabolism 27:136–150. e135
Ward PS, Thompson CB (2012) Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 21:297–308
Wei Y, Xia W, Zhang Z, Liu J, Wang H, Adsay NV, Albarracin C, Yu D, Abbruzzese JL, Mills GB (2008) Loss of trimethylation at lysine 27 of histone H3 is a predictor of poor outcome in breast, ovarian, and pancreatic cancers. Mol Carcinog 47:701–706
White E (2015) The role for autophagy in cancer. J Clin Invest 125:42–46
Wicha M, Dontu G, Al-Hajj M, Clarke M (2003) Stem cells in normal breast development and breast cancer. Breast Cancer Res 5:50
Wicha MS, Liu S, Dontu G (2006) Cancer stem cells: an old idea—a paradigm shift. Cancer Res 66:1883–1890
Woehlecke H, Osada H, Herrmann A, Lage H (2003) Reversal of breast cancer resistance protein–mediated drug resistance by tryprostatin A. Int J Cancer 107:721–728
Wright MH, Calcagno AM, Salcido CD, Carlson MD, Ambudkar SV, Varticovski L (2008) Brca1 breast tumors contain distinct CD44+/CD24-and CD133+ cells with cancer stem cell characteristics. Breast Cancer Res 10:R10
Wu Y, Wu PY (2009) CD133 as a marker for cancer stem cells: progresses and concerns. Stem Cells Dev 18:1127–1134
Xiong A, Yang Z, Shen Y, Zhou J, Shen Q (2014) Transcription factor STAT3 as a novel molecular target for cancer prevention. Cancers 6:926–957
Yan Y, Zuo X, Wei D (2015) Concise review: emerging role of CD44 in cancer stem cells: a promising biomarker and therapeutic target. Stem Cells Transl Med 4:1033–1043
Yan N, Xu L, Wu X, Zhang L, Fei X, Cao Y, Zhang F (2017) GSKJ4, an H3K27me3 demethylase inhibitor, effectively suppresses the breast cancer stem cells. Exp Cell Res 359:405–414
Yang F, Cao L, Sun Z, Jin J, Fang H, Zhang W, Guan X (2016) Evaluation of breast cancer stem cells and intratumor stemness heterogeneity in triple-negative breast cancer as prognostic factors. Int J Biol Sci 12:1568–1577
Yin H, Glass J (2011) The phenotypic radiation resistance of CD44+/CD24− or low breast cancer cells is mediated through the enhanced activation of ATM signaling. PLoS One 6:e24080
Yin L, Castagnino P, Assoian RK (2008) ABCG2 expression and side population abundance regulated by a transforming growth factor β–directed epithelial-mesenchymal transition. Cancer Res 68:800–807
Yin L, Velazquez OC, Liu Z-J (2010) Notch signaling: emerging molecular targets for cancer therapy. Biochem Pharmacol 80:690–701
Yip N, Fombon I, Liu P, Brown S, Kannappan V, Armesilla A, Xu B, Cassidy J, Darling J, Wang W (2011) Disulfiram modulated ROS–MAPK and NFκB pathways and targeted breast cancer cells with cancer stem cell-like properties. Br J Cancer 104:1564–1574
You J, Zhao J, Wen X, Wu C, Huang Q, Guan F, Wu R, Liang D, Li C (2015) Chemoradiation therapy using cyclopamine-loaded liquid–lipid nanoparticles and lutetium-177-labeled core-crosslinked polymeric micelles. J Control Release 202:40–48
Yu H, Jove R (2004) The STATs of cancer—new molecular targets come of age. Nat Rev Cancer 4:97–105
Zeng YA, Nusse R (2010) Wnt proteins are self-renewal factors for mammary stem cells and promote their long-term expansion in culture. Cell Stem Cell 6:568–577
Zhang Y, Appleton BA, Wiesmann C, Lau T, Costa M, Hannoush RN, Sidhu SS (2009) Inhibition of Wnt signaling by dishevelled PDZ peptides. Nat Chem Biol 5:217–219
Zhang Y, Zhang H, Wang X, Wang J, Zhang X, Zhang Q (2012) The eradication of breast cancer and cancer stem cells using octreotide modified paclitaxel active targeting micelles and salinomycin passive targeting micelles. Biomaterials 33:679–691
Zhang H, Lu H, Xiang L, Bullen JW, Zhang C, Samanta D, Gilkes DM, He J, Semenza GL (2015) HIF-1 regulates CD47 expression in breast cancer cells to promote evasion of phagocytosis and maintenance of cancer stem cells. Proc Natl Acad Sci 112:E6215–E6223
Zhao S, Smith KS, Deveau AM, Dieckhaus CM, Johnson MA, Macdonald TL, Cook JM (2002) Biological activity of the tryprostatins and their diastereomers on human carcinoma cell lines. J Med Chem 45:1559–1562
Zhao C, Blum J, Chen A, Kwon HY, Jung SH, Cook JM, Lagoo A, Reya T (2007) Loss of β-catenin impairs the renewal of normal and CML stem cells in vivo. Cancer Cell 12:528–541
Zhao F, Ming J, Zhou Y, Fan L (2016) Inhibition of Glut1 by WZB117 sensitizes radioresistant breast cancer cells to irradiation. Cancer Chemother Pharmacol 77:963–972
Zhong Y, Shen S, Zhou Y, Mao F, Lin Y, Guan J, Xu Y, Zhang S, Liu X, Sun Q (2016) nOTch1 is a poor prognostic factor for breast cancer and is associated with breast cancer stem cells. Onco Targets Ther 9:6865–6871
Zhu Y, Zhang X, Liu Y, Zhang S, Liu J, Ma Y, Zhang J (2012) Antitumor effect of the mTOR inhibitor everolimus in combination with trastuzumab on human breast cancer stem cells in vitro and in vivo. Tumor Biol 33:1349–1362
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Pindiprolu, S.K.S.S., Krishnamurthy, P.T. & Chintamaneni, P.K. Pharmacological targets of breast cancer stem cells: a review. Naunyn-Schmiedeberg's Arch Pharmacol 391, 463–479 (2018). https://doi.org/10.1007/s00210-018-1479-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00210-018-1479-3