Abstract
Many pathogenic bacteria express filamentous appendages, termed pili, on their surface. These organelles function in several important bacterial processes, including mediating bacterial interaction with, and colonization of the host, signalling events, locomotion, DNA uptake, electric conductance, and biofilm formation. In the last decade, it has been established that the tuberculosis-causing bacterium, Mycobacterium tuberculosis, produces two pili types: curli and type IV pili. In this paper, we review studies on M. tuberculosis pili, highlighting their structure and biological significance to M. tuberculosis pathogenesis, and discuss their potential as targets for therapeutic intervention and diagnostic test development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Despite achieving the millennium development goal to decrease incidence rates by 2015, tuberculosis (TB), responsible for 1.5 million deaths in 2013 (WHO 2014), remains a scourge to mankind globally. There is thus an urgent need to identify new drugs, vaccines, and diagnostics for improved TB control. An improved understanding of Mycobacterium tuberculosis genetics, physiology, and its virulence factors would provide knowledge on possible bacterial targets for rationally designed therapeutics and diagnostics.
It is well established that the expression of adherence-mediating molecules, termed adhesins, is crucial to a pathogen’s ability to infect host cells. However, attachment to immune cells can trigger phagocytosis, leading to the destruction of the pathogen (Kline et al. 2009). Expressing adhesins on hydrophobic polymeric structures that extend beyond the bacterial surface limit repulsive forces between the host and pathogen, thereby enabling their interaction from a suitable distance and with less deleterious consequences for the pathogen (Alteri 2005; Kline et al. 2009).
Several saprophytic and pathogenic bacteria express their adhesins on polymeric proteinaceous structures termed pili. Multiple pilus types have been identified in bacteria, each associated with a unique structure and distinct functions. In Gram-negative bacteria, the production of chaperone/usher-assembled, type IV, and curli pili are well documented. Gram-positive bacteria have been reported to produce type IV and sortase-assembled pili (Kline et al. 2010). In general, pili are 1–10 nm wide and 0.07–3 μm long (Telford et al. 2006). They have been implicated in several bacterial processes, including induction of signalling events in host cells, host tissue adhesion, co-aggregation and biofilm formation, immunomodulation, biosensor, motility, DNA uptake, and can act as nanowires that transfer electrons from bacterial cells to extracellular electron acceptors (Källström et al. 1998; Telford et al. 2006; Lovley 2008).
Mycobacteria were generally regarded as a non-piliated genus. However, Alteri (2005) showed, using negative staining and transmission electron microscopy (TEM), that both the fast-growing Mycobacterium smegmatis and Mycobacterium fortuitum and the slow-growing M. tuberculosis produced pili under standard growth conditions. The TB vaccine strain, Mycobacterium bovis BCG, was also found to be piliated (Alteri 2005). Subsequently, two further studies have confirmed piliation by clinical M. tuberculosis isolates, using atomic force microscopy and TEM (Velayati et al. 2012; Hosseini et al. 2014).
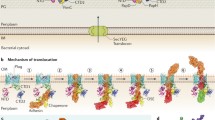
Using TEM and scanning electron microscopy (SEM), Alteri (2005) identified two distinct pili morphotypes produced by M. tuberculosis. The expression of these pili types was found to be influenced by nutritional conditions and/or environmental signals. The two pili types of M. tuberculosis are type IV pili, which are produced in broth-grown cultures (Fig. 1a), and curli pili, which are produced by bacilli cultured on solid media (Fig. 1b). In liquid media, the attenuated M. tuberculosis strain H37Ra expressed significantly less pili compared with virulent strains, alluding to the role of pili as a possible virulence factor of M. tuberculosis (Alteri 2005). In this mini-review, we summarize the current knowledge on the two M. tuberculosis pili types and discuss their potential as targets for the development of anti-TB strategies.
M. tuberculosis pili. a TEM micrograph of broth-grown M. tuberculosis H37Rv showing the expression of rope-like, laterally aggregated type IV pili. b TEM micrograph of agar-grown M. tuberculosis clinical isolate CDC1551 showing the production of coiled, aggregated curli pili. c, d High-resolution SEM images of static-grown M. tuberculosis H37Ra adhering to glass coverslips (c) and adhering to each other in pellicles (d) using pili-like structures. Arrows point to pili fibres (Alteri 2005)
The curli pili of M. tuberculosis (MTP)
Amyloids are a β-sheet-rich fold that many proteins can acquire (Blanco et al. 2012). The production of amyloids by microorganisms is a highly controlled and regulated process and confers advantages to these organisms. These ‘functional amyloids’ play key roles in biofilm formation in organisms such as Pseudomonas spp. (Dueholm et al. 2010), Bacillus subtilis (Romero et al. 2010), Streptococcus mutans (Oli et al. 2012), and Staphylococcus aureus (Schwartz et al. 2012). They also alter the surface properties of Streptomyces coelicolor (Claessen et al. 2003) and Ustilago maydis (Teertstra et al. 2009), thereby enabling spore and aerial hyphae formation in these microbes. Amyloid production is also associated with the virulence and toxicity of Klebsiella pneumoniae (Bieler et al. 2005), Xanthomonas axonopodis (Oh et al. 2007), and Listeria monocytogenes (Bavdek et al. 2012).
Curli pili are the most well studied and characterized bacterial functional amyloid. They are densely tangled and coiled masses of cell surface structures, which are produced by several Enterobacteriaceae. These non-branching proteins are 4–6 nm wide and possess aggregative properties (Epstein and Chapman 2008). They are highly stable structures that are assembled by nucleation-precipitation pathways, involving major and minor curlin subunits (Hammar et al. 1996).
Microscopically, M. tuberculosis curli-like pili (MTP) appear similar in ultrastructure to the curli of Escherichia coli and Salmonella spp. (Olsén et al. 1989; Collinson et al. 1991). MTP are classified as a curli amyloid due to its ability to bind to Congo red and its insolubility in sodium dodecyl sulphate (Alteri et al. 2007). However, MTP subunits display no primary sequence homology to curli (Alteri et al. 2007) and lack the typical β-sheet secondary structure of curlins (Ramsugit et al. 2013).
MTP are 2–3 nm in diameter and comprise subunits that are encoded by the Rv3312A (mtp) ORF, as determined by mass spectroscopy analysis, Western blotting with antibodies against Rv3312A, and immuno-electron microscopy (Alteri et al. 2007). By Western blotting with anti-Rv3312A antibodies, M. smegmatis was found to be unable to produce the MTP pilin subunit protein, which suggested that MTP may be associated with pathogenic mycobacteria (Alteri 2005).
The assembly of curli in E. coli occurs via specific biogenesis pathways, involving seven curli specific genes (csg) that are encoded by the csgDEFG and csgBAC operons (Blanco et al. 2012). The major and minor curlin subunits, CsgA and CsgB, participate in nucleation and polymerization functions, whilst CsgC may be involved in subunit secretion (Gibson et al. 2007; Taylor et al. 2011). Accessory proteins are encoded by the csgDEFG operon (Hammar et al. 1995; Loferer et al. 1997; Chapman et al. 2002; Robinson et al. 2006). The mtp gene is, however, not located in an operon or near other pilus-associated genes (Fig. 2a). The additional proteins that make up the MTP structure, their export and assembly mechanisms, and its association with the complex mycobacterial cell wall are currently unidentified.
Organization of the M. tuberculosis pili-encoding genetic loci. a The curli pili-encoding gene, mtp (Rv3312A), is located between genes involved in intermediary metabolism and respiration and conserved hypotheticals and is not arranged in an operon with other pili-associated genes. b The type IV pili (flp) gene cluster consists of the Rv3654c-Rv3660c ORF’s. The chromosomal coordinates are indicated below each illustration (modified from: http://genolist.pasteur.fr/TubercuList)
The mtp gene is located between genes involved in intermediary metabolism (Fig. 2a), implying that it may be protected from deletion or gene inactivation events (Alteri 2005). Naidoo et al. (2014) showed by amplicon sequencing that 98 % of M. tuberculosis clinical isolates (n = 86) possessed a conserved mtp gene sequence. Hosseini et al. (2014) further reported a 100 % conservation of the mtp (curli) and flp (type IV pili) gene sequences in clinical isolates (n = 36). The mtp gene is present only in M. tuberculosis complex strains and not in non-tuberculous mycobacteria nor other respiratory pathogens (Naidoo et al. 2014).
Using ELISA and immunofluorescent microscopy, Alteri et al. (2007) demonstrated the presence of IgG antibodies against MTP in active TB cases. Thus, MTP are produced during human TB infection (Alteri et al. 2007). TB infection leads to inflammation, tissue damage, and exposure of the extracellular matrix (ECM). M. tuberculosis binds to these areas of tissue damage (Middleton et al. 2002). Using ELISA, MTP were reported to bind to the ECM protein, laminin, in vitro (Alteri et al. 2007). These researchers also showed, using immunofluorescent microscopy, that MTP are produced during the pathogen’s adhesion to A549 epithelial cells. These findings indirectly implicated MTP in functioning as an adherence factor, which may be crucial in mediating a close interaction and colonization of host cells (Alteri et al. 2007).
To assess the function of MTP, a MTP-deficient Δmtp mutant strain and a MTP-overexpressing complemented strain were constructed (Ramsugit et al. 2013). Biofilm formation assays and crystal violet staining identified the role of MTP in in vitro biofilm formation, as the MTP-deficient strain displayed a 68 % reduction in biofilm mass compared with the parental strain (Ramsugit et al. 2013). Pili-like structures were previously observed to mediate the pathogen’s attachment to surfaces (Fig. 1c) and to encase the TB bacilli in pellicle biofilms (Fig. 1d). Based on microscopic observations, their role in in vitro biofilm formation is by mediating cell-to-cell contact (Alteri 2005; Velayati et al. 2012; Ramsugit et al. 2013). Alteri et al. (2007) demonstrated that MTP are expressed during human infection; therefore, formation of M. tuberculosis biofilms in vivo may be possible, although this has yet to be conclusively shown.
MTP also play a fundamental role in the infection of host cells. Adherence and invasion assays revealed that the MTP-deficient mutant displayed a 42 and 69 % decrease in the adhesion to and invasion of THP-1 macrophages, respectively, compared with the parental strain (Ramsugit and Pillay 2014). Adhesion to and invasion of A549 pulmonary epithelial cells by the mutant were also significantly reduced by 69 and 56 %, respectively (Ramsugit S, Pillay B, and Pillay M; submitted). There were no significant differences between cytokine and chemokine levels produced by A549 epithelial cells infected with the wild-type and MTP-deficient strains (Ramsugit S, Pillay B, and Pillay M; submitted). MTP-mediated entry into epithelial cells may therefore be advantageous to the pathogen by suppressing inflammatory responses to invasion into these host cells and possibly innate immune responses.
The type IV pili of M. tuberculosis
Type IV pili are flexible surface-exposed filaments, which tend to form bundles (Berry and Pelicic 2015). They have been well studied and characterized in Gram-negative bacteria. In these organisms, they function in adhesion to host cells, motility (gliding and twitching), microcolony formation, competence, protein secretion, and serve as nanowires that carry electric current (Aas et al. 2002; Mattick 2002; Kirn et al. 2003; Burrows 2005; Reguera et al. 2005; Han et al. 2007; Burrows 2012).
Type IV pili were subsequently found to be produced by several Gram-positive bacteria, including Clostridia (Varga et al. 2006), Streptococcus sanguinis (Xu et al. 2007), and Bacillus spp. (Imam et al. 2011). Using the PilFind algorithm, Imam et al. (2011) showed that Gram-positive bacteria contained a highly diverse set of type IV pili. Type IV pili have been linked to gliding motility of Clostridia (Varga et al. 2006), adherence to the host by organisms such as Clostridium perfringens (Rodgers et al. 2011) and Ruminococcus albus (Rakotoarivonina et al. 2002), and biofilm formation by C. perfringens (Varga et al. 2008).
Since M. tuberculosis is regarded as a non-motile organism, it is tempting to speculate that the M. tuberculosis type IV pili function as an adhesin, which mediates adhesion to host cells and/or biofilm formation. Alteri (2005) provided initial information on the type IV pili of M. tuberculosis and preliminary evidence supporting their possible adhesin function. However, since their discovery, studies on the type IV pili of M. tuberculosis are notably absent in literature and their significance in M. tuberculosis pathogenesis requires further investigation.
M. tuberculosis expresses type IV pili that appear as rope-like bundles, which are encoded by a 5-kb genomic island containing seven genes, including the flp prepilin (Rv3656c) and putative biogenesis genes (Fig. 2b). Rv3654c and Rv3655c encode secreted proteins; Rv3656c codes for a transmembrane protein; and the products of Rv3657c-Rv3660c resemble pili assembly proteins, type II/IV secretion system proteins, and tight adherence (tad) genes (Danelishvili et al. 2010). In addition, M. tuberculosis H37Rv contains two fimbrial low-molecular-weight protein (Flp) pre-pilin peptidases, encoded by Rv0990c and Rv2551c, located away from the flp gene cluster, not shown on the gene map in Fig. 2b.
Flp proteins are small type IV pilins. Their encoding genes are found within the tad loci, together with conserved type IV pili biosynthetic genes and other tad-specific genes (Imam et al. 2011). The M. tuberculosis flp genes are similar to the flp-tad locus of Aggregatibacter actinomycetemcomitans. The M. tuberculosis type IVb pili gene cluster is characterized by a conserved glycine residue, which is located before a signal peptide region, and a conserved glutamate residue, which is five positions from the conserved glycine. These pili belong to the Flp sub-family of type IVb pili, as identified by the presence of a tyrosine residue paired with the conserved glutamate residue (Alteri 2005). The Flp/Tad pili are important virulence factors and mediators of biofilm formation in several pathogenic and environmental bacteria (Tomich et al. 2007) and function in host colonization by Bifidobacterium breve (O’Connell Motherway et al. 2011).
Alteri (2005) showed by gene expression analysis and immunofluorescent microscopy that M. tuberculosis expresses and secretes the Flp protein, thereby confirming that the flp genes are functional. The Flp peptide is capable of self-assembling into polymeric structures at a pH of 4.5–7.4, as evidenced by negative staining and TEM and immuno-electron microscopy. This finding supports the role of the M. tuberculosis Flp homolog as a type IV pilin and may indicate that acidic pH, such as that which is present in the phagosomal vacuole, may trigger type IV pili assembly (Alteri 2005).
Using immunofluorescence microscopy, it was found that Flp pili may have an adhesin function, as evidenced by its expression during the organism’s interaction with U937 human macrophages and A549 epithelial cells (Alteri 2005). Flp pili (or flp genes) are known to function in the adherence to surfaces and the host in pathogens such as A. actinomycetemcomitans (Kachlany et al. 2000, 2001) and Haemophilus ducreyi (Nika et al. 2002; Spinola et al. 2003). M. tuberculosis Flp pili could thus function similarly, although this has yet to be experimentally confirmed.
Alteri (2005) suggested that the M. tuberculosis type IVb pili genes were acquired by horizontal gene transfer. The flp locus has a higher G + C content (70 %) than the M. tuberculosis chromosome (65 %), and Z’ component analysis showed that the increase in G + C content corresponds to the boundary of the type IVb pili genes. This locus is flanked by multiple direct repeats, providing further evidence for the insertion of foreign DNA, as well as confirming that the type IV pili genes are located on a genomic island (Alteri 2005).
Danelishvili et al. (2010) identified that the expression of the flp gene cluster is up-regulated within macrophages, as compared to when the pathogen is extracellularly located. The proteins encoded by Rv3654c and Rv3655c suppress macrophage apoptosis by blocking the extrinsic pathway (Danelishvili et al. 2010). The protein encoded by Rv3660c was also found to be a septum site determining protein which, when overexpressed, induces filamentation and an alternative metabolism in M. tuberculosis (England et al. 2011).
M. tuberculosis pili as potential therapeutic and diagnostic targets
Due to their key functions in microbial pathogenesis, pili (and other adhesins) represent important therapeutic and diagnostic targets (Govender et al. 2014). Although it is unclear whether M. tuberculosis forms biofilms in vivo, several lines of evidence suggest that it could (Ha et al. 2005; Lenaerts et al. 2007; Wang et al. 2013). If M. tuberculosis exists as biofilms in vivo, then drugs (curlicides) that target and block MTP formation could represent a useful anti-biofilming agent to reduce TB persistence, given their essentiality for in vitro biofilm formation (Ramsugit et al. 2013). The cell surface localization and structural role in biofilm formation (including their involvement in the early developmental stages) imply that pili are useful drug targets to prevent biofilm formation or to disrupt existing biofilms (Hett and Hung 2009).
Due to their role in host colonization, M. tuberculosis pili are potential targets for the design of therapeutics to attenuate host infection. Blocking M. tuberculosis entry into host cells may expose the organism for killing by the host immunity and/or drugs. This could involve the use of competitive inhibitors, such as those resembling the host cell receptors or the use of pili analogues, to prevent the initial host–pathogen interaction, thereby limiting disease progression (Ofek et al. 2003; Salminen et al. 2007). Alternatively, the design of peptidomimetic compounds that mimic pili subunit proteins may inhibit pili formation (Evans and Chapman 2014).
Anti-adhesion approaches to prevent infection could be promising to control the spread of M. tuberculosis strains, irrespective of their drug susceptibility or resistance profile (Hansen et al. 1997). In addition, such agents are less likely to lead to the emergence of drug-resistant M. tuberculosis strains, in comparison with antibiotics, which are bactericidal or limit the organism’s growth (Ofek et al. 2003). Carbohydrate analogues of receptors are generally not toxic and immunogenic (Sharon 2006). However, this strategy to TB therapy is disadvantaged by the presence of multiple M. tuberculosis adhesins (Govender et al. 2014). In addition, adhesion to host cells can occur by mechanisms other than adhesin–receptor interactions, such as by hydrophobic and other non-specific interactions (Ofek et al. 2003). Targeting multiple adhesion mechanisms or adhesins may therefore be required to completely disrupt the infection process.
M. tuberculosis pili may be a potential vaccine candidate or used in TB immunotherapy strategies, where anti-pili antibodies may hinder infection by interacting with these extracellular structures. Pili (and other adhesins) are well documented to be excellent immunogens and are therefore prime targets for vaccine development (Klemm and Schembri 2000). However, the diversity of M. tuberculosis adhesins could pose a challenge to their use as vaccine candidates since the influence of other adhesins may still enable infection (Govender et al. 2014). Therefore, complete success would require the targeting of a combination of major adhesins. The mtp gene is highly conserved and unique to the M. tuberculosis complex strains, suggesting that its encoded product, MTP, may be a putative biomarker for a TB diagnostic test (Naidoo et al. 2014). The conserved nature of the M. tuberculosis curlin and type IV pilin genes (Hosseini et al. 2014; Naidoo et al. 2014) implies that antigenic variation may not be a limitation for a pilus-based vaccine.
Concluding remarks and future work
A decade on since the discovery of M. tuberculosis pili, significant insight has been gained on the structure and function of MTP. However, their role during in vivo infection has yet to be determined. The identification of type IV pili in M. tuberculosis was the first reports of a classical virulence factor for the pathogen (Kachlany et al. 2001; Alteri 2005). However, since the pioneering work by Alteri and colleagues, no further studies have explored their role in M. tuberculosis pathogenesis. Gene knockout of the flp gene and in vitro and in vivo assays will clarify the role of this pilus type.
A comparison of what is currently known about the curli and type IV pili of M. tuberculosis to those of other bacteria (Table 1) suggests that significant further characterization of M. tuberculosis pili is needed. A key research area that needs to be explored is the identification of the assembly mechanisms of both pilus types and the host receptors with which they interact. The eventual aim will be to translate this knowledge into useful therapeutics and diagnostics, which can lead to improved TB control and prevention.
References
Aas FE, Wolfgang M, Frye S, Dunham S, Løvold C, Koomey M (2002) Competence for natural transformation in Neisseria gonorrhoeae: components of DNA binding and uptake linked to type IV pilus expression. Mol Microbiol 46:749–760
Alteri CJ (2005) Novel pili of Mycobacterium tuberculosis. Ph.D. Thesis, University of Arizona
Alteri CJ, Xicohténcatl-Cortes J, Hess S, Caballero-Olín G, Girón JA, Friedman RL (2007) Mycobacterium tuberculosis produces pili during human infection. Proc Natl Acad Sci USA 104:5145–5150
Bavdek A, Kostanjšek R, Antonini V, Lakey JH, Dalla Serra M, Gilbert RJ, Anderluh G (2012) pH dependence of listeriolysin O aggregation and pore-forming ability. FEBS J 279:126–141
Berry JL, Pelicic V (2015) Exceptionally widespread nanomachines composed of type IV pilins: the prokaryotic swiss army knives. FEMS Microbiol Rev 39:134–154
Bieler S, Estrada L, Lagos R, Baeza M, Castilla J, Soto C (2005) Amyloid formation modulates the biological activity of a bacterial protein. J Biol Chem 280:26880–26885
Blanco LP, Evans ML, Smith DR, Badtke MP, Chapman MR (2012) Diversity, biogenesis and function of microbial amyloids. Trends Microbiol 20:66–73
Burrows LL (2005) Weapons of mass retraction. Mol Microbiol 57:878–888
Burrows LL (2012) Prime time for minor subunits of the type II secretion and type IV pilus systems. Mol Microbiol 86:765–769
Chapman MR, Robinson LS, Pinkner JS, Roth R, Heuser J, Hammar M, Normark S, Hultgren SJ (2002) Role of Escherichia coli curli operons in directing amyloid fiber formation. Science 295:851–855
Claessen D, Rink R, de Jong W, Siebring J, de Vreugd P, Boersma FGH, Dijkhuizen L, Wösten HAB (2003) A novel class of secreted hydrophobic proteins is involved in aerial hyphae formation in Streptomyces coelicolor by forming amyloid-like fibrils. Genes Dev 17:1714–1726
Collinson SK, Emödy L, Müller KH, Trust TJ, Kay WW (1991) Purification and characterization of thin, aggregative fimbriae from Salmonella enteritidis. J Bacteriol 173:4773–4781
Danelishvili L, Yamazaki Y, Selker J, Bermudez LE (2010) Secreted Mycobacterium tuberculosis Rv3654c and Rv3655c proteins participate in the suppression of macrophage apoptosis. PLoS ONE 5:e10474
Dueholm MS, Petersen SV, Sønderkær M et al (2010) Functional amyloid in Pseudomonas. Mol Microbiol 77:1009–1020
England K, Crew R, Slayden RA (2011) Mycobacterium tuberculosis septum site determining protein, Ssd encoded by rv3660c, promotes filamentation and elicits an alternative metabolic and dormancy stress response. BMC Microbiol 11:79
Epstein EA, Chapman MR (2008) Polymerizing the fibre between bacteria and host cells: the biogenesis of functional amyloid fibres. Cell Microbiol 10:1413–1420
Evans ML, Chapman MR (2014) Curli biogenesis: order out of disorder. Biochim Biophys Acta 1843:1551–1558
Gibson DL, White AP, Rajotte CM, Kay WW (2007) AgfC and AgfE facilitate extracellular thin aggregative fimbriae synthesis in Salmonella enteritidis. Microbiology 153:1131–1140
Govender VS, Ramsugit S, Pillay M (2014) Mycobacterium tuberculosis adhesins: potential biomarkers as anti-tuberculosis therapeutic and diagnostic targets. Microbiology 160:1821–1831
Ha KY, Chung YG, Ryoo SJ (2005) Adherence and biofilm formation of Staphylococcus epidermidis and Mycobacterium tuberculosis on various spinal implants. Spine (Phila Pa 1976) 30:38–43
Hammar M, Arnqvist A, Bian Z, Olsén A, Normark S (1995) Expression of two csg operons is required for production of fibronectin- and congo red-binding curli polymers in Escherichia coli K-12. Mol Microbiol 18:661–670
Hammar M, Bian Z, Normark S (1996) Nucleator-dependent intercellular assembly of adhesive curli organelles in Escherichia coli. Proc Natl Acad Sci USA 93:6562–6566
Han X, Kennan RM, Parker D, Davies JK, Rood JI (2007) Type IV fimbrial biogenesis is required for protease secretion and natural transformation in Dichelobacter nodosus. J Bacteriol 189:5022–5033
Hansen HC, Haataja S, Finne J, Magnusson G (1997) Di-, tri-, and tetravalent dendritic galabiosides that inhibit hemagglutination by Streptococcus suis at nanomolar concentration. J Am Chem Soc 119:6974–6979
Hett EC, Hung DT (2009) Targeting multiple biofilm pathways. Chem Biol 16:1216–1218
Hosseini H, Fooladi AAI, Arjomandzadegan M, Emami N, Bornasi H (2014) Genetics study and transmission electron microscopy of pili in susceptible and resistant clinical isolates of Mycobacterium tuberculosis. Asian Pac J Trop Biomed 4(Suppl 2):S675–S679
Imam S, Chen Z, Roos DS, Pohlschröder M (2011) Identification of surprisingly diverse type IV pili, across a broad range of gram-positive bacteria. PLoS ONE 6:e28919
Kachlany SC, Planet PJ, Bhattacharjee MK, Kollia E, DeSalle R, Fine DH, Figurski DH (2000) Nonspecific adherence by Actinobacillus actinomycetemcomitans requires genes widespread in bacteria and archaea. J Bacteriol 182:6169–6176
Kachlany SC, Planet PJ, DeSalle R, Fine DH, Figurski DH (2001) Genes for tight adherence of Actinobacillus actinomycetemcomitans: from plaque to plague to pond scum. Trends Microbiol 9:429–437
Källström H, Islam MS, Berggren PO, Jonsson AB (1998) Cell signaling by the type IV pili of pathogenic Neisseria. J Biol Chem 273:21777–21782
Kirn TJ, Bose N, Taylor RK (2003) Secretion of a soluble colonization factor by the TCP type 4 pilus biogenesis pathway in Vibrio cholerae. Mol Microbiol 49:81–92
Klemm P, Schembri MA (2000) Bacterial adhesins: function and structure. Int J Med Microbiol 290:27–35
Kline KA, Fälker S, Dahlberg S, Normark S, Henriques-Normark B (2009) Bacterial adhesins in host-microbe interactions. Cell Host Microbe 5:580–592
Kline KA, Dodson KW, Caparon MG, Hultgren SJ (2010) A tale of two pili: assembly and function of pili in bacteria. Trends Microbiol 18:224–232
Lenaerts AJ, Hoff D, Aly S, Ehlers S, Andries K, Cantarero L, Orme IM, Basaraba RJ (2007) Location of persisting mycobacteria in a Guinea pig model of tuberculosis revealed by r207910. Antimicrob Agents Chemother 51:3338–3345
Loferer H, Hammar M, Normark S (1997) Availability of the fibre subunit CsgA and the nucleator protein CsgB during assembly of fibronectin-binding curli is limited by the intracellular concentration of the novel lipoprotein CsgG. Mol Microbiol 26:11–23
Lovley DR (2008) Extracellular electron transfer: wires, capacitors, iron lungs, and more. Geobiology 6:225–231
Mattick JS (2002) Type IV pili and twitching motility. Annu Rev Microbiol 56:289–314
Middleton AM, Chadwick MV, Nicholson AG, Dewar A, Groger RK, Brown EJ, Ratliff TL, Wilson R (2002) Interaction of Mycobacterium tuberculosis with human respiratory mucosa. Tuberculosis 82:69–78
Naidoo N, Ramsugit S, Pillay M (2014) Mycobacterium tuberculosis pili (MTP), a putative biomarker for a tuberculosis diagnostic test. Tuberculosis 94:338–345
Nika JR, Latimer JL, Ward CK, Blick RJ, Wagner NJ, Cope LD, Mahairas GG, Munson RS Jr, Hansen EJ (2002) Haemophilus ducreyi requires the flp gene cluster for microcolony formation in vitro. Infect Immun 70:2965–2975
O’Connell Motherway M, Zomer A, Leahy SC et al (2011) Functional genome analysis of Bifidobacterium breve UCC2003 reveals type IVb tight adherence (Tad) pili as an essential and conserved host-colonization factor. Proc Natl Acad Sci USA 108:11217–11222
Ofek I, Hasty DL, Sharon N (2003) Anti-adhesion therapy of bacterial diseases: prospects and problems. FEMS Immunol Med Microbiol 38:181–191
Oh J, Kim JG, Jeon E, Yoo CH, Moon JS, Rhee S, Hwang I (2007) Amyloidogenesis of type III-dependent harpins from plant pathogenic bacteria. J Biol Chem 282:13601–13609
Oli MW, Otoo HN, Crowley PJ, Heim KP, Nascimento MM, Ramsook CB, Lipke PN, Brady LJ (2012) Functional amyloid formation by Streptococcus mutans. Microbiology 158:2903–2916
Olsén A, Jonsson A, Normark S (1989) Fibronectin binding mediated by a novel class of surface organelles on Escherichia coli. Nature 338:652–655
Rakotoarivonina H, Jubelin G, Hebraud M, Gaillard-Martinie B, Forano E, Mosoni P (2002) Adhesion to cellulose of the Gram-positive bacterium Ruminococcus albus involves type IV pili. Microbiology 148:1871–1880
Ramsugit S, Pillay M (2014) Mycobacterium tuberculosis pili promote adhesion to and invasion of THP-1 macrophages. Jpn J Infect Dis 67:476–478
Ramsugit S, Guma S, Pillay B, Jain P, Larsen MH, Danaviah S, Pillay M (2013) Pili contribute to biofilm formation in vitro in Mycobacterium tuberculosis. Antonie Van Leeuwenhoek 104:725–735
Reguera G, McCarthy KD, Mehta T, Nicoll JS, Tuominen MT, Lovley DR (2005) Extracellular electron transfer via microbial nanowires. Nature 435:1098–1101
Robinson LS, Ashman EM, Hultgren SJ, Chapman MR (2006) Secretion of curli fibre subunits is mediated by the outer membrane-localized CsgG protein. Mol Microbiol 59:870–881
Rodgers K, Arvidson CG, Melville SB (2011) Expression of a Clostridium perfringens type IV pilin by Neisseria gonorrhoeae mediates adherence to muscle cells. Infect Immun 79:3096–3105
Romero D, Aguilar C, Losick R, Kolter R (2010) Amyloid fibers provide structural integrity to Bacillus subtilis biofilms. Proc Natl Acad Sci USA 107:2230–2234
Salminen A, Loimaranta V, Joosten JA, Khan AS, Hacker J, Pieters RJ, Finne J (2007) Inhibition of P-fimbriated Escherichia coli adhesion by multivalent galabiose derivatives studied by a live-bacteria application of surface plasmon resonance. J Antimicrob Chemother 60:495–501
Schwartz K, Syed AK, Stephenson RE, Rickard AH, Boles BR (2012) Functional amyloids composed of phenol soluble modulins stabilize Staphylococcus aureus biofilms. PLoS Pathog 8:e1002744
Sharon N (2006) Carbohydrates as future anti-adhesion drugs for infectious diseases. Biochim Biophys Acta 1760:527–537
Spinola SM, Fortney KR, Katz BP, Latimer JL, Mock JR, Vakevainen M, Hansen EJ (2003) Haemophilus ducreyi requires an intact flp gene cluster for virulence in humans. Infect Immun 71:7178–7182
Taylor JD, Zhou Y, Salgado PS et al (2011) Atomic resolution insights into curli fiber biogenesis. Structure 19:1307–1316
Teertstra WR, van der Velden GJ, de Jong JF, Kruijtzer JA, Liskamp RM, Kroon-Batenburg LM, Müller WH, Gebbink MF, Wösten HA (2009) The filament-specific Rep1-1 repellent of the phytopathogen Ustilago maydis forms functional surface-active amyloid-like fibrils. J Biol Chem 284:9153–9159
Telford JL, Barocchi MA, Margarit I, Rappuoli R, Grandi G (2006) Pili in gram-positive pathogens. Nat Rev Microbiol 4:509–519
Tomich M, Planet PJ, Figurski DH (2007) The tad locus: postcards from the widespread colonization island. Nat Rev Microbiol 5:363–375
Varga JJ, Nguyen V, O’Brien DK, Rodgers K, Walker RA, Melville SB (2006) Type IV pili-dependent gliding motility in the gram-positive pathogen Clostridium perfringens and other clostridia. Mol Microbiol 62:680–694
Varga JJ, Therit B, Melville SB (2008) Type IV pili and the CcpA protein are needed for maximal biofilm formation by the gram-positive anaerobic pathogen Clostridium perfringens. Infect Immun 76:4944–4951
Velayati AA, Farnia P, Masjedi MR (2012) Pili in totally drug resistant Mycobacterium tuberculosis (TDR-TB). Int J MYCO 1:57–58
Wang F, Sambandan D, Halder R et al (2013) Identification of a small molecule with activity against drug-resistant and persistent tuberculosis. Proc Natl Acad Sci USA 110:2510–2517
World Health Organization (2014) Global tuberculosis report 2014. http://apps.who.int/iris/bitstream/10665/137094/1/9789241564809_eng.pdf?ua=1. Accessed 27 Oct 2014
Xu P, Alves JM, Kitten T et al (2007) Genome of the opportunistic pathogen Streptococcus sanguinis. J Bacteriol 189:3166–3175
Acknowledgments
This work was supported by National Research Foundation (NRF) Grants awarded to Dr. M. Pillay (Grant Nos. 77286 and 90508). Dr. S. Ramsugit acknowledges scholarship from the NRF and Canon Collins Trust. We are grateful to Dr. Christopher J. Alteri (University of Michigan Medical School) for providing us with approval to include the micrographs in Fig. 1.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Erko Stackebrandt.
Rights and permissions
About this article
Cite this article
Ramsugit, S., Pillay, M. Pili of Mycobacterium tuberculosis: current knowledge and future prospects. Arch Microbiol 197, 737–744 (2015). https://doi.org/10.1007/s00203-015-1117-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-015-1117-0