Abstract
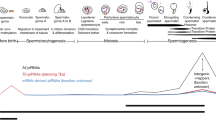
As the most abundant small RNAs, piwi-interacting RNAs (piRNAs) have been identified as a new class of non-coding RNAs with 24–32 nucleotides in length, and they are expressed at high levels in male germ cells. PiRNAs have been implicated in the regulation of several biological processes, including cell differentiation, development, and male reproduction. In this review, we focused on the functions and molecular mechanisms of piRNAs in controlling spermatogenesis, including genome stability, regulation of gene expression, and male germ cell development. The piRNA pathways include two major pathways, namely the pre-pachytene piRNA pathway and the pachytene piRNA pathway. In the pre-pachytene stage, piRNAs are involved in chromosome remodeling and gene expression regulation to maintain genome stability by inhibiting transposon activity. In the pachytene stage, piRNAs mediate the development of male germ cells via regulating gene expression by binding to mRNA and RNA cleavage. We further discussed the correlations between the abnormalities of piRNAs and male infertility and the prospective of piRNAs’ applications in reproductive medicine and future studies. This review provides novel insights into mechanisms underlying mammalian spermatogenesis and offers new targets for diagnosing and treating male infertility.
Graphical Abstract

The piRNA/piRNA pathway functions and applications. The piRNA/piRNA pathway is mainly involved in piRNA biogenesis, regulation of transposons, and binding to mRNAs to control spermatogenesis
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
PiRNAs and piRNA pathways
Non-coding RNAs are classified into short and long non-coding RNAs in terms of nucleotide length, as shown in Table 1. The short non-coding RNAs include microRNAs (miRNAs), small interfering RNAs (siRNAs), small nucleolar RNAs (snoRNAs), and piRNAs [1, 2]. MiRNAs inhibit mRNA translation or cause mRNA degradation by binding to fully or partially complementary sequences in the 3’ untranslated regions (3’ UTRs) of targeting message RNAs (mRNAs) [3, 4], while siRNAs silence gene expression by forming RNA-induced silencing complexes (RISC) to bind perfectly to complementary mRNAs [5]. In addition to binding to mRNAs, snoRNAs direct the cleavage of rRNA. The regulation of mRNA 3’ end processing and selective cleavage by snoRNAs is crucial for regulating gene expression and appropriate protein synthesis [6]. Among all non-coding RNAs, piRNAs are the most abundant small RNAs, and they mediate gene expression by silencing transposable elements to maintain genome stability [7]. PiRNAs are enriched in the testis and ovary, and they play key roles in controlling spermatogenesis and oogenesis [8,9,10,11]. The orderly regulation of piRNA pathways ensures the correct pairing and segregation of chromosomes during meiosis to ensure the transmission and stability of genetic information [12].
The Piwi proteins were first identified in female Drosophila melanogaster, and deletion of Piwi leads to mitotic arrest of female germinal stem cells in Drosophila [26]. Knockdown of Piwi in Caenorhabditis elegans inhibits the self-renewal of germline stem cells [27]. PIWI proteins have been shown to be mainly expressed in male germ cells in mammals, and a diversity of PIWI proteins with temporal and spatial expression exists during germ cell development (Table 2) [28]. There are three types of PIWI proteins in mice, including PIWIL1 (also known as Miwi), PIWIL2 (Mili) and PIWIL4 (Miwi2) [29,30,31], while human PIWI proteins can be divided into four types, namely PIWIL1 (HIWI), PIWIL2 (HILI), PIWIL3 and PIWIL4 (HIWI2) [32, 33]. PiRNAs have been found to interact with PIWI proteins [21,35,36,37,38,39]. In mammals, piRNAs can guide PIWI proteins to silence transposons and regulate gene expression in cells [40]. However, the mechanisms of piRNA generation vary in different species and tissues. Here, we focused on the piRNA generation mechanisms in mammals.
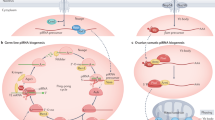
During mammalian spermatogenesis, piRNAs are formed via two stages. At the first wave beginning at the embryonic stage, pre-pachytene piRNAs can be detected in fetal and perinatal germ cells [49, 50]. Pachytene piRNAs are mainly detectable in pachytene spermatocytes [51]. These two waves of piRNA biogenesis play different roles. Pre-pachytene piRNAs are primarily involved in silencing transposons, while pachytene piRNAs mainly participate in gene regulation [52]. There are two main pathways for piRNA generation, including the primary processing pathway and the ping-pong cycle pathway. In the primary processing pathway, precursor RNA transcription of pachytene piRNAs is initiated by MYB proto-oncogene-like 1 (A-myb) and subsequently cleaved to generate piRNA intermediates by phospholipase D family member 6 (MitoPLD) [51]. MitoPLD is located at the mitochondrial outer membrane, and it can process piRNAs from their longer immature forms into shorter intermediate forms, namely pre-piRNAs [53]. PARN-like ribonuclease domain-containing exonuclease 1 (PNLDC1) interacts with Tudor structural domain protein TDRKH, which has been proposed to be a piRNA trimming cofactor [54]. Pre-piRNAs are cleaved to mature length by PNLDC1 and 2’-O-methylation into functional forms by HEN methyltransferase 1 (HENMT1) [55,56,57]. The absence of HENMT1 leads to unmethylation of piRNAs and decreases in their abundance and length [58]. In the ping-pong cycle pathway, primary piRNAs bind to PIWI proteins and cleave complementary mRNAs, which results in the production of secondary piRNAs with a 5’end. These secondary piRNAs are associated with PIWI proteins and cleave complementary target mRNAs again, which generates primary piRNAs with a 3’end. After modification by HENMT1, primary piRNAs are matured into functional molecules. The primary piRNAs and secondary piRNAs are generated in a continuous ping-pong cycle pathway [59,60,61]. The generation process of piRNAs is illustrated in Fig. 1. Any error in the primary processing pathway or the ping-pong pathway can lead to the impaired piRNA biogenesis. In 2020, ribosomes have been identified as key players in pachytene piRNA formation, and they participate in piRNA generation through three distinct mechanisms [62]. First of all, the binding of ribosomes to cleavage products stabilizes the cleavage products for loading PIWI. Ribosomes maintain a stepwise division from the 5’ UTR to the 3’ UTR in piRNA biogenesis. Secondly, the extended ribosome itself acts as a powerful helicase to remove secondary structures and RNA-binding proteins. Finally, ribosomes provide a platform for biological and regulatory proteins to bind to pachytene piRNA precursors [62]. Ribosomes have been shown to direct the fragmentation of mRNAs to produce 3’ UTR piRNAs [63].
The piRNA pathway proteins are involved in piRNA biosynthesis. PiRNA biosynthesis can be divided into two main processes, namely primary piRNA pathway and ping-pong pathway. Primary piRNAs generate the initial piRNAs with the beginning of the ping-pong pathway. These two processes are involved in piRNA generation and functions. Different proteins are required for the primary piRNA pathway and the ping-pong pathway
PiRNAs and their regulation of spermatogenesis
Spermatogenesis is a complex process that comprises three main stages, including self-renewal and differentiation of spermatogonial stem cells (SSCs), two meiotic divisions of spermatocytes, and spermiogenesis with the morphological changes from round spermatids to elongated spermatids [64], and it is derived from primordial germ cells (PGCs) [65]. In the human testis, spermatogonia are classified into type A spermatogonia and type B spermatogonia, while type A spermatogonia are subdivided into the Adark and Apale spermatogonia [66]. Adark spermatogonia are the quiescent SSCs, while Apale spermatogonia are the self-renewing SSCs [67]. Type B spermatogonia further differentiate into spermatocytes that undergo two meiotic divisions to generate spermatids with morphic maturation to form spermatozoa. Unlike human testis, type A spermatogonia in mice are divided into Asingle, Apaired, and Aaligned spermatogonia, which is followed by A1 − 4, intermediate spermatogonia, and type B spermatogonia [68]. This process of spermatogenesis requires the cooperation of various kinds of male germ cells with testicular somatic cells, including Sertoli cells, myoid cells, and Leydig cells [69]. Spermatogenesis is precisely regulated by genetic and epigenetic factors. Notably, numerous studies have highlighted that epigenetic factors, including piRNAs and other ncRNAs, are involved in mediating spermatogenesis.
PiRNAs are essential for the development of male germ cells and somatic cells through multiple mechanisms. Firstly, piRNAs bind to PIWI proteins to form a RISC that effectively inhibits transposons by guiding their methylation and recombination [11, 70]. There are three major types of transposons in mammals, including LINES, SINES, and LTRS, while piRNAs are mainly involved in the regulation of LINES [49, 71, 72]. Secondly, piRNAs can bind to complementary sequences of mRNAs or lncRNAs to promote their degradation, transport, or post-transcriptional regulation. Pachytene piRNAs negatively regulate mRNAs, lncRNAs, and LINE-1 retrotransposons in late spermatocytes, which is largely dependent on retrotransposon sequences and pseudogenes [73]. Thirdly, piRNAs are highly enriched in late spermatocytes and round spermatids [74]. Abnormality of piRNAs/PIWI leads to developmental disorders of male germ cells in Drosophila, zebrafish, nematode, mice, golden hamsters, and humans. Notably, piRNAs are involved in the development and maturation of male germ cells by affecting chromosome remodeling, epigenetic modifications, and apoptosis, and conversely, piRNA dysfunction leads to male infertility [10, 75,76,77,78,79,80]. Finally, piRNA clusters contain partial or complete transposon sequences, and piRNA systems can distinguish between friends and enemies and initiate responses [81]. As such, piRNAs protect the genome of male germline from invasion by new transposable elements [82, 83]. In this review, we elucidated the roles and mechanisms of piRNAs in controlling spermatogenesis, including genome stability, gene expression regulation, and male germ cell development, to illustrate the indispensable functions of piRNAs in maintaining male fertility and reproductive health.
The functions of piRNAs in controlling mammalian spermatogenesis
Abnormality in the expression of one or more genes involved in the piRNA pathway leads to aberrant spermatogenesis, and the phenotypes of mice with piRNA pathway defects include abnormal activation of germ cell transposons, meiotic arrest, and spermiogenesis disorder, which eventually results in male infertility [31, 84,85,86,87]. Several proteins, including Mili, Mov10-like RISC complex RNA helicase 1 (Mov10l1), and Ring finger protein 17 (Rnf17), have been identified as key components involved in the piRNA pathway [67,89,90,91]. PiRNAs play different roles in the pre-pachytene and pachytene stages. During the pre-pachytene stage, piRNAs play a critical role in maintaining genome stability by inhibiting transposon activity [49, 92]. Pachytene piRNAs are mainly involved in regulating mRNA degradation and translation during the post-meiotic stages of spermatogenesis [41, 93, 94]. Notably, piRNAs interact with mRNAs, as evidenced by the findings that piRNAs mediate the stability and translation of mRNAs and that mRNAs in turn participate in the biogenesis and functional regulation of piRNAs [74, 88]. The targets of piRNAs are expressed in the testis, with 74.93% of them being retrotransposons [95]. Therefore, piRNAs are committed to protecting the genome from transposons and participate in the regulation of protein-coding genes. These two processes act synergistically at different stages of testis development [96]. To elucidate the diverse mechanisms by which piRNA function, we summarized the proteins mainly involved in the pre-pachytene piRNA biogenesis and pachytene piRNA biogenesis as well as the deficiency of proteins in the piRNA pathways with spermatogenesis arrest, as we illustrated in Fig. 2.
The expression of piRNAs and PIWI proteins and the relationship between the loss of key proteins in the piRNA/piRNA pathway and spermatogenesis failure. The expression of piRNAs and PIWI proteins is spatio-temporal specific. According to the expression periods, piRNAs are classified into pre-pachytene piRNAs and pachytene piRNAs. Loss of key proteins of the piRNA/piRNA pathway results in spermatogenesis disorder
The pre-pachytene piRNA pathway and spermatogenesis
A series of pre-pachytene piRNA-related proteins have been identified in mammalian spermatogenesis, including Mili [30], Miwi2 [31], Pnldc1 [57], Mov10l1 [97], the homeobox transcription factor Rhox10 [98], Exonuclease domain-containing 1 (Exd1) [99], Glycerol-3-phosphate acyltransferase 2 (Gpat2) [100], Gametocyte-specific factor 1 (Gtsf1) [101], Tdrd12 [102], ATPase activity of mouse Vasa homolog (Mvh) [103], and MORC family CW-type zinc finger 3 (Morc3) [104]. The main function of piRNAs is to inhibit transposon activity during de novo DNA methylation [105, 106]. Mili is expressed in the cytoplasm of prospermatogonia, while Miwi2 is present in the cellular nuclei of these cells. Both Mili and Miwi2 are required for DNA methylation of TE sequences. Mili acts as an upstream factor of Miwi2, and it is involved in the regulation of Miwi2 nuclear localization [107]. As a functional partner of Miwi2, Tdrd9 mutation in fetal testes results in abnormal piRNA profiles in prospermatogonia [108]. In spermatogonia, LINE-1 repression is accomplished primarily through the following three pathways: the Piwi-piRNA pathway, CpG promoter DNA methylation, and G9a-mediated H3K9me2 [109]. Here, we discussed the repression of LINE-1 by the Piwi-piRNA pathway in spermatogonia. PiRNAs complement the LINE-1 sequences and maintain genome stability by directing the Piwi proteins Mili to cleave and disrupt LINE-1 transcripts via the RNA interference (RNAi)-like mechanism [110]. Deletion of Mili severely impacts the self-renewal and differentiation of SSCs by regulating translation [30, 103, 111]. In Mili null mice, spermatogonia are absent and exhibit a Sertoli cell-only (SCO) phenotype [44]. During the transition of PGCs to prospermatogonia, Rhox10 transcriptionally activates Mili, and Mili protein drives the piRNA pathway to mediate the repression of the LINE-1 promoter, thereby inhibiting transposons [112]. Loss of Rhox10 leads to an aberrant number of SSCs and a progressive spermatogenesis disorder [98, 113].
Defects in the processing and maturation of piRNAs impede normal development of male germ cells. Pnldc1 is a trimmer of pre-piRNAs, and it is involved in the cleavage of pre-piRNAs [54, 85]. Abnormal alterations in the length of piRNAs, the number of pachytene piRNAs, and piRNA-processing proteins PIWIL1, PIWIL4, A-MYB and TDRKH, have been observed in non-obstructive azoospermia (NOA) with PNLDC1 mutations [79]. Also, piRNAs 3’end processing is disrupted in Pnldc1 mutant mice, which leads to an accumulation of 3’untrimmed piRNA intermediates, a reduction of pre-pachytene piRNAs and pachytene piRNAs, and defective spermatogenesis [57, 77, 114]. Lack of piRNA trimming and methylation in Pnldc1 and Henmt1 double knockout male mice results in the collapse of the piRNA pathway [115]. A 3’ tail modification of mature piRNAs has been identified in mouse testes, and uridylation initiated by TUT4/7 is the predominant tail form of MIWI-bound piRNAs [116]. In addition, Mov10l1 is an RNA helicase located upstream of PNLDC1 and it is involved in piRNA 5’ end processing. It is noteworthy that two MOV10L1 mutations and one homozygous MOV10L1 mutation are identified in NOA patients [117]. MOV10L1 has been found to be upstream of Piwi proteins during the primary processing of pachytene piRNAs [92]. In mice, Mov10l1 has a similar expression pattern to Mili, and it can bind to pre-piRNAs to initiate the piRNA pathway [97, 118]. Disruption of Mov10l1 helicase activity results in the loss of pre-pachytene piRNAs, the activation of retrotransposons, early meiotic arrest, and male infertility [119, 120]. In testes of cryptorchid boys, expression levels of MOV10L1, PIWIL2, PIWIL4, and TDRD9 are significantly reduced, suggesting that the impaired expression of these genes associated with transposon silencing may lead to genome instability and azoospermia [121]. Defective processing of piRNAs may be an important etiology of NOA patients, as evidenced by the integration of clinical data and in vivo studies.
MVH is crucial for processing intermediates into piRNAs, which ensures the silencing of transposable elements and the maintenance of male fertility [122]. Mvh mutation impairs piRNA binding to Miwi2 and impacts de novo methylation of transposons, resulting in piRNA cycle arrest and spermatocyte arrest at the meiotic stage and male infertility in mice [103]. Gasz (Germ cell protein with Ankyrin repeats, sterile alpha motif, and leucine Zipper) knockout mice exhibit phenotypes similar to Mili knockout mice, and Gasz may participate in the regulation of spermatogenesis by localizing or stabilizing multiple proteins to affect piRNA processing and synthesis [123].
EXD1 acts as an RNA adaptor in the PIWI-EXD1-TDRD12 (PET) complex [99]. The absence of Exd1 causes a reduction in the production of Mili-sliced piRNAs and the biogenesis of Miwi2 piRNAs [99]. In contrast to the phenotype observed in Exd1-deficient mice, Exd2 interacts with Mili and its mutation leads to abnormal regulation of specific piRNA clusters without affecting piRNA biogenesis [124]. Therefore, Exd2 mutations have no obvious effect on male fertility, and the precise function of Exd2 in the piRNA pathway remains to be further elucidated. To elucidate the function of Exd1 and Exd2 in the piRNA pathway is important for understanding the mechanism that maintains genome stability during spermatogenesis [124]. TDRD12, also known as ECAT8, is a unique piRNA biogenesis factor. The complex formed by Tdrd12 with Exd1 and Mili is mainly involved in the biogenesis of secondary piRNAs, while Tdrd12 does not affect the biogenesis of primary piRNAs [125]. Gpat2 is one of the Mili binding proteins, and Gpat2 may act as a scaffold to recruit various factors required for piRNA production, which is essential for primary piRNA biogenesis [126]. Knockdown of Gpat2 results in the impaired piRNA biogenesis in SSCs and more apoptosis in neonatal spermatogonia [100]. Gtsf1 is a component of the Mili/Miwi2 complex, and deficiency of Gtsf1 in mouse prospermatogonia results in abnormal localization of Miwi2. Loss of Miwi2-bound piRNAs leads to defective secondary piRNA biogenesis, reflecting that Gtsf1 is a key factor for piwi-piRNA cleavage of target mRNAs [101]. Morc3 recognizes and binds to H3K4me3 marks on the promoter regions of retrotransposon genes and piRNA clusters, thereby affecting the biogenesis of primary piRNAs. In the embryonic testes, Morc3 is involved in chromosome remodeling and regulates the transcription of piRNA precursors. In addition, Morc3 may be involved in the biogenesis of secondary piRNAs [104].
Collectively, these studies mentioned above implicate that piRNAs assume stage-specific regulation during spermatogenesis. In spermatogonia, a number of transposon-targeting piRNAs are generated, while piRNAs in primary spermatocytes play pivotal roles in the suppression of repetitive sequences and transposons. This stage-specific regulation is essential for maintaining genome stability and ensuring proper spermatogenesis [127].
The pachytene piRNA pathway and spermatogenesis
We summarized the pachytene piRNA pathway proteins affecting mammalian spermatogenesis, including Miwi [128], Tdrd1 [90], Tdrd5 [129], Tdrd9 [108], Ubiquitin-like with PHD and ring finger domains 1 (Uhrf1) [130], Tdrkh [131], MitoPLD [55], Rnf17 [88], Adenosine deaminase domain containing 2 (Adad2) [132], Testis expressed 19 (Tex19) [133], Testis expressed 15 (Tex15) [134], Ubiquitin B (Ubb) [135], BTB domain containing 18 (Btbd18) [136], Maelstrom (Mael) [137], and FKBP prolyl isomerase family member 6 (Fkbp6) [55, 138]. Next, we addressed the specific functions and molecular mechanisms of these proteins involved in piRNA biogenesis.
Miwi, a member of the PIWI protein family, is mainly expressed in male germ cells from pachytene spermatocytes to early spermatids [29]. Miwi deficiency affects centromere assembly in meiosis, resulting in an enhancement in chromosome missegregation at meiosis I, an increase in aneuploidy at meiosis II, and the death of spermatids [139]. Miwi/HIWI functions as piRNAs-guided mRNA degradation [96]. The Miwi/piRNAs mechanism plays a dual role in the regulation of spermatogenesis in mice, and translation activation occurs through the formation of Miwi/piRNAs/eIF3f/HuR supercomplex in round spermatids [128]. At elongating spermatids, the Miwi/piRNAs/CAF1 supercomplex forms and initiates extensive mRNA elimination in spermatids to regulate acrosome formation [128].
Members of the TDRD family (TDRD1-9, 12) are involved in the biogenesis of piRNAs or PIWI protein interaction, with the exception of TDRD3 [140]. However, the specific functions of each protein seem to be distinct [141,142,143], as shown in Table 3. Tdrd1 interacts with Mili, and it is involved in regulating retrotransposons via the Piwi pathway. Tdrd1 deficiency leads to the derepression of LINE-1, and loss of DNA methylation of its regulatory elements causes mislocalization of Miwi2 from nuclei to cytoplasm [42]. Nevertheless, Tdrd1 does not affect piRNA biogenesis in spermatogenesis [43, 144]. Unlike Tdrd1, Tdrd9 is a functional partner of Miwi2 in the Piwi pathway. Tdrd9 mutation in fetal testis affects the silencing of LINE-1 in pre-spermatogonia and piRNAs profiles [108]. Tdrd2, also known as Tdrkh, specifically recruits Miwi to drive the piRNA biogenesis. Miwi is lost in the chromatoid body when Tdrkh is defective in the testis, which causes spermatogenesis arrest at the round spermatid stage [145]. Unlike Tdrkh, MitoPLD deficiency in mice leads to meiotic arrest of spermatocytes due to affecting piRNA generation and distribution [84, 146]. Deletion of Uhrf1 in the testis results in a significant reduction of PIWI proteins and piRNA-associated proteins TDRKH and MVH [130]. Tdrd4, also known as Rnf17, has been shown to participate in the balance of the ping-pong cycle of piRNAs during meiosis, affect piRNA content by inhibiting the generation of secondary piRNAs, and enhance the expression of protein-coding genes crucial for the regulation of spermatogenesis by inhibiting the ping-pong cycle [88]. Interestingly, a testis-specific protein, namely Adad2, interacts with multiple RNA-binding proteins, including Mili, Miwi, Rnf17, and Ythdc2, has been found to be involved in piRNA biogenesis [132]. Adad2 guides Rnf17 to repress ping-pong activity during the biogenesis of pachytene piRNAs. During meiosis, Adad2 knockout can cause mislocalization of Rnf17 followed by the loss of ping-pong suppression, which results in overproduction of secondary piRNAs and ultimately spermatogenesis arrest at the round spermatid stage [147, 148]. Tdrd5 binds directly to the precursors of piRNAs and functions via selectively controlling the processing of the pachytene piRNAs precursors [129]. Tdrd6 interacts with Mili and Miwi, and Tdrd6 knockout mice are arrested at the round spermatid stage and fail to form the elongated spermatids. Abnormal miRNA expression has been observed in Tdrd6 knockout mice, and it remains unclear about whether Tdrd6 affects piRNA biogenesis [144]. In contrast to Tdrd6, LINE-1 expression is abnormal in Tdrd7−/− testis, and Tdrd7 mutant mice have spermatogenesis arrest at the round spermatid stage [149]. Tdrd7 is associated with RNP remodeling at early spermatid stage, and Tdrd6 is related to structural maintenance at a later stage of spermatid development. Chromatoid bodies are important subcellular sites for piRNA biogenesis, containing a large number of proteins required for piRNA biogenesis [150]. Therefore, Tdrd7 may influence not only the expression of LINE-1 in spermatogenesis but also the expression and localization of other piRNA biogenesis proteins, e.g., Mili, Mvh, Mael, and Gasz [149]. Tdrd8, also known as Stk31, has been shown to interact with Miwi in mouse testis by LC/MS assay [151]. However, subsequent experiments found that disruption of Stk31 does not affect male fertility [143], and the function of Stk31 in the testis needs to be further explored. TEX19 is directly associated with piRNAs through its VPTEL domain [133]. Unlike TEX19, TEX15 functions independently of piRNA biogenesis, and it may act as a nuclear effector protein downstream of the piRNA pathway to silence TEs in male germ cells [152]. Meanwhile, TEX15 has been shown to be essential for de novo DNA methylation of TEs regulated by MIWI2-piRNA pathway [134]. Ubb is essential for maintaining piRNA-metabolic proteins, and Ubb knockout in mice results in the reduction of piRNA metabolic process-related proteins and meiotic cell cycle arrest in male germ cells, which leads to azoospermia phenotype [135]. Deletion of Btbd18 reduces the expression level of primary piRNA precursors, which severely impairs piRNA biogenesis. Mice lacking Btbd18 experience a massive loss of spermatocytes due to apoptosis, leading to spermatogenesis arrest, azoospermia, and male infertility. Since no aberrant retrotransposon activation is observed, the phenotype of Btbd18 null mice differs from Miwi knockout mice [136]. Btbd18 enhances the expression level of primary piRNA precursors by promoting transcriptional elongation [136]. During spermatogenesis, both A-MYB and TCFL5 regulate piRNAs. A-Myb is involved in piRNA generation as a transcription factor, and A-Myb null mice generate fewer pachytene piRNAs [51, 153]. Interestingly, half of the promoters of human pachytene piRNAs do not interact with A-Myb [153]. During early meiosis I, A-Myb initiates transcription of Tcfl5 which binds to its own promoter and A-Myb promoter to form a mutually reinforcing positive feedback loop. Tcfl5 regulates the expression of genes required for piRNA maturation and stimulates transcription of evolutionarily young pachytene piRNA genes, while A-Myb is responsible for activating transcription in older pachytene piRNA genes [154]. Spermatogenesis in Tcfl5−/− and Tcfl5+/− mice is arrested at the pachytene/diplotene spermatocyte transition and round/elongating spermatid transition [155, 156], while spermatocytes in A-Myb mutant mice assume abnormal cell cycle progression [157].
The histone H3 family includes two major histone variants CENP-A and H3.3 [164]. H3.3 is encoded by two different genes, namely H3.3a and H3.3b, with the same amino acid sequences [165, 166]. Loss of H3.3b in spermatocytes is associated with the increased expression of RLTR10B and RLTR10B2 retrotransposons as well as downregulation of piRNA clusters. This finding reflects the dual role of H3.3b in controlling spermatogenesis. H3.3 can positively regulate the expression of piRNAs required for meiotic chromatin inactivation and repression of repeat element transcription [167]. Mael is a conserved HMG box structural domain protein that is essential for mouse spermatogenesis [137, 168], and its MAEL domain may have potential nuclease activity or RNA binding capacity to affect piRNA biogenesis [169]. MAEL-related protein complex, including MIWI/PIWIL, TDRD6, TDRD4/RNF17, TDRD, STK31/TDRD8, and TDRD9, has been identified [170]. Mael129 knockout mice exhibit spermatogenesis arrest with acrosome and flagellar malformations, which may be caused by an imbalance between pachytene piRNAs and MIWI [170], and loss of Mael leads to a decrease in the levels of pachytene piRNAs in mouse testes [171, 172]. MAEL is mainly expressed in spermatid mitochondria in the human testis, and loss of Mael can lead to mouse mitochondrial dysfunction and asthenozoospermia [173]. It remains to be further uncovered whether MAEL affects piRNA biogenesis in human spermatogenesis. FKBP6 is required for spermatogenesis, and it is involved in piRNA biogenesis and synaptic complex formation [174]. Recently, Fkbp6-null testicular cells have been found to be arrested at the round spermatids, and Fkbp6 deletion severely affects piRNA levels [175]. Moreover, FKBP6, a molecular chaperone of HSP90, reduces MIWI2-binding piRNAs in Fkbp6-null testes [138, 176]. Hsp90 regulates spermatogenesis by participating in the formation and/or stabilization of MILI-piRNA and MIWI2-piRNA complexes [177]. In addition, piRNAs can directly bind to mRNAs and affect their translation and stability, thereby regulating gene expression. High level of piR-003399 inhibits CDK6 expression and causes cell cycle arrest in mouse spermatogonia at G1 phase and abnormal sperm count, motility, and morphology [178]. The loss of pi6 and pi18 loci on mouse chromosomes 6 and 18 affects sperm motility and acrosome reaction, respectively, thereby preventing spermatids from penetrating the zona pellucida and resulting in male infertility. Pi6 and pi18 piRNAs primarily target mRNA rather than retrotransposons. Pi6 represses gene expression by cleaving mRNA encoding proteins required for spermatogenesis, and it is involved in the piRNA-piRNA precursor interaction network [179]. In summary, the piRNA pathway plays critical roles in spermatogenesis through three primary pathways, including participation in piRNA biogenesis, altering the status of the transposons, and regulating mRNA stability or translation.
Abnormal piRNAs and spermatogenesis failure and male infertility
Mutations or low piRNA levels in blood or semen have recently been linked to male infertility. High-throughput Illumina Hiseq technology has been used to compare the piRNA profiles of testes from unsuccessful sperm retrieval (USR) groups and successful sperm retrieval (SSR) controls. Interestingly, 553 piRNAs have been shown to be specifically expressed in NOA patients with successful sperm retrieval [180], suggesting that these piRNAs may be potential biomarkers for predicting successful sperm retrieval. RNA sequencing of seminal plasma reveals significantly fewer piRNA numbers in infertile patients compared to normal men. In addition, piR-31068, piR-31925, piR-43771, piR-43773, and piR-30198 have been identified as molecular hallmarks of male infertility [181]. In another study, low expression of piR-31704 and piR-39888 has been found in sperm of infertile men, and piRNA levels have been indicated to be correlated with sperm concentration and fertilization rate with intracytoplasmic sperm injection (ICSI) [182]. The levels of piR-1207 and piR-2107 in sperm and seminal plasma of asthenospermia patients are significantly lower than those of normal fertile individuals [183]. RNA sequencing of serum reveals that piR-26399 level exhibits a significant difference between males with reduced fertility and normal controls [184].
The piRNA pathway genes, including HENMT1, PIWIL1, and PIWIL2, are highly expressed in male germ cells during normal spermatogenesis, and their expression levels are decreased in spermatogenesis failure accompanied by germ cell depletion [185]. Significantly, male infertility has been demonstrated to be associated with hypermethylation of the PIWIL2 and TDRD1 promoters, and epigenetic inactivation of PIWI pathway genes can lead to piRNA deficiency and LINE-1 hypomethylation [186]. Aberrant expression of piRNAs is associated with the impaired spermatogenesis, loss of sperm motility, and abnormal sperm morphology in azoospermia (Table 4). Mutations in genes responsible for piRNA generation, e.g., PNLDC1 and TDRD1, contribute to spermatogenesis failure. Notably, mutations in PNLDC1 have been found to cause azoospermia by whole exome sequencing (WES) in three studies [79, 187, 188]. Four PNLDC1 mutations have been identified in 924 NOA patients, reflecting that mutations of PNLDC1 affect meiosis of spermatocytes [79]. A novel PNLDC1 mutation causing oligoasthenozoospermia has been identified by generating male mice with the PNLDC1 R58G mutation [188]. A compound heterozygous missense variant of PNLDC1 has been further found as it causes spermatogenesis arrest [187]. Therefore, PNLDC1 is essential for maintaining normal spermatogenesis by affecting piRNA biogenesis. Mutations in MOV10L1, which is required for piRNA processing, have been identified in two of 414 patients with NOA or severe oligospermia [117]. HIWI and TDRD proteins have been shown to be critical for piRNA biogenesis [189, 190], and notably, HIWI and TDRD polymorphisms are highly related to male infertility [191]. Nine single nucleotide polymorphisms (SNPs) in four human Piwi genes have been identified by the SNP stream ® 12-plex platform and TaqMan methods. Interestingly, the HIWI2 rs508485 has been found to be positively associated with the risk of azoospermia [192], which provides the first epidemiological evidence supporting the involvement of genetic polymorphisms of Piwi in spermatogenesis failure. Variants in piRNA pathway genes have been identified as risk factors for male infertility [191]. In addition, the SNP rs77559927 in piRNA pathway TDRD1 gene has been shown to be associated with a risk of spermatogenesis disorder [193]. Notably, the expression levels of the TDRD gene family are significantly lower in NOA testicular tissues compared to the OA patients with normal spermatogenesis. Abnormal expression of TDRD family genes associated with the piRNA pathway may lead to male infertility [194]. MOV10L1 polymorphisms have been found to be associated with male infertility in 30 infertile men diagnosed with spermatocyte arrest [195]. It is of particular significance to explore the mechanisms by which piRNA-related genes’ polymorphisms control abnormal spermatogenesis. This would help us better understand the pathogenesis of male infertility and develop new treatments for this disease.
In conclusion, aberrant expression of piRNAs and piRNA pathway genes are associated with abnormalities in spermatogenesis and male infertility. Therefore, with the continuous development of single-cell RNA sequencing or RNA deep sequencing, more clues can be uncovered to better understand molecular mechanisms underlying the regulation of piRNAs in male germ cell development and reproductive system diseases, and new approaches would be developed for the diagnosis and treatment of male infertility.
Applications of piRNAs in reproductive medicine
The applications of piRNAs in the male reproductive system
PiRNAs are essential for regulating male germ cell development, normal spermatogenesis and male fertility. In the field of male reproduction, piRNAs might have significant implications as shown in Fig. 3. First of all, piRNAs may be used as biomarkers to assess male reproductive health based upon the quality and quantity of piRNAs in serum or seminal plasma, and piRNAs can be employed for the diagnosis and treatment of reproduction-related diseases. PiRNAs can also be utilized as biomarkers to predict residual spermatogenic conditions in NOA patients. In testicular tissues from NOA patients, a total of 959 piRNAs were significantly differentially expressed between successful and failed sperm extraction groups by RNA-seq, while 553 piRNAs were completely absent in the failed sperm extraction group [180]. Therefore, these piRNAs can be utilized as markers for the development of assisted reproductive technology (ART). Cryptorchidism has been found to be deficient in normal piRNA formation, which affects the development of normal testicular tissues [196, 197]. The expression pattern of piRNAs during spermatogenesis and maturation is closely related to the quality and quantity of sperm, and thus piRNAs can be used as biomarkers of sperm for the diagnosis and treatment of male infertility.
Prospective of piRNAs in the male reproductive system. The applications of piRNAs in the male reproductive system include the following six aspects: as a standard for assessing male reproductive health, selecting high-quality sperm in ART, the diagnosis and treatment of male infertility, the diagnosis and therapy of male reproductive system tumors, biomarkers of reproductive toxicity, and exploring novel mechanisms underlying male germ cell development
PiRNAs also have potential applications in the treatment of reproductive tumors. The majority of piRNAs in testicular germ cell tumors (TGCT) have been found to be lost by RNA sequencing [198]. It has been found that loss of piRNA defense in carcinoma in situ and TGCT cells results in a reduced ability to prevent chromatin instability [198], since neither in germ cell neoplasia in situ (GCNIS) cells nor TGCT cells express PIWI/piRNA pathway genes with no piRNA biogenesis [199, 200]. In particular, piRNAs may have applications in treating tumors because they can inhibit the proliferation and metastasis of cancer cells [201].
PiRNAs can be further used as biomarkers for reproductive toxicity. Fluoride has been shown to change the expression of piRNAs and their lysosomal signaling pathway in the testis, thereby causing testicular damage [202]. The toxicity of nickel nanoparticles (Ni NPs) has been demonstrated to affect spermatogenesis, sperm motility, and fertilization ability. Furthermore, piR-32362,259 enhances Ni NPs-induced GC-1 cell damage by regulating the PI3K/AKT signaling pathway [203]. Together, abnormalities of piRNAs are closely related to the occurrence of male infertility, testicular cancer, and other diseases, and piRNAs can be applied as biomarkers for the diagnosis and treatment of reproductive system diseases.
The applications of piRNAs in drug development
Katalin Kariko and Drew Weissman won the 2023 Nobel Prize in physiology or medicine, for their discoveries on nucleoside base modifications that enable the development of effective mRNA vaccines against COVID-19. At present, the development and application of RNA drugs have aroused great interest. In 2018, two drugs received FDA approval, including Onpattro (patisiran) for the treatment of hereditary amyloidosis and Givlaari (givosiran) for the treatment of acute intermittent porphyria [204, 205]. During COVID-19, Moderna and Pfizer/BioNTech have successfully developed an mRNA vaccine against the Corona Virus [206]. Moreover, non-coding RNAs can be used as targets of drug resistance in cancer cells. By constructing the patient-derived xenograft tumor mouse model, treatment with siRNA targeting carcass has been found to restore sorafenib resistance [207]. The length of piRNAs has been found to be correlated with translation, and short piRNAs result in the impaired MIWI/piRNAs translational activation and inhibit mRNA translation [208]. Future studies may be focused on designing drugs to add exogenous piRNAs to restore normal spermatogenesis. In human spermatogenesis, PNLDC1, MOV10L1, and HIWI mutations have recently been shown to cause azoospermia by affecting piRNA processing [79, 117, 190]. In the near future, small molecule drugs targeting proteins related to these piRNA pathways can be designed to repair the abnormal biosynthesis of piRNAs, rescue spermatogenesis and treat male infertility. It is feasible to achieve this goal by designing small molecule drugs that target piRNAs and their pathway proteins by binding to piRNAs or interfering with piRNA interactions with PIWI or MIWI proteins, thereby affecting piRNA functions. These small molecule compounds can be discovered and optimized by high-throughput screening, bioinformatics analysis, gene expression profiling, and functional experiments. This would offer perspective for the treatment and prevention of male infertility, other reproductive system diseases, and reproductive system tumors.
Conclusions and perspective
In summary, we addressed the functions and mechanisms of piRNAs in regulating spermatogenesis and their abnormality or mutations in male infertility. In general, the piRNA pathway participates in the regulation of spermatogenesis through three main pathways, including the inhibition of transposons, participating in piRNA biogenesis, and binding to mRNAs. We also discussed the correlations between the abnormalities of piRNAs and male infertility. Finally, we pointed out potential applications of piRNAs in reproductive medicine and drug design. Furthermore, piRNAs play essential roles in male reproduction and tumor formation. Future studies on piRNAs might be focused on the following aspects: (i) the roles and regulatory mechanisms of piRNAs in controlling self-renewal and differentiation of SSCs, meiosis of spermatocytes, and spermatogenesis of spermatids; (ii) the functions of piRNAs in mediating the testis environment or niche, particularly somatic cells, including Sertoli cells, myoid cells, and Leydig cells; (iii) the regulatory networks formed by piRNAs, other non-coding RNAs, and genes or proteins to regulate normal spermatogenesis; and (iv) the important applications of piRNAs as biomarkers for the diagnosis of male infertility and cancers as well as novel drug development for the treatment of these diseases.
Data availability
The data is available upon the request from the corresponding author.
References
Alexander RP, Fang G, Rozowsky J, Snyder M, Gerstein MB (2010) Annotating non-coding regions of the genome. Nat Rev Genet 11:559–571. https://doi.org/10.1038/nrg2814
Hombach S, Kretz M (2016) Non-coding RNAs: classification, Biology and Functioning. Adv Exp Med Biol 937:3–17. https://doi.org/10.1007/978-3-319-42059-2_1
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531. https://doi.org/10.1038/nrg1379
Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A et al (2007) A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129:1401–1414. https://doi.org/10.1016/j.cell.2007.04.040
Carthew RW, Sontheimer EJ (2009) Origins and mechanisms of miRNAs and siRNAs. Cell 136:642–655. https://doi.org/10.1016/j.cell.2009.01.035
Liang J, Wen J, Huang Z, Chen XP, Zhang BX, Chu L (2019) Small nucleolar RNAs: insight into their function in Cancer. Front Oncol 9:587. https://doi.org/10.3389/fonc.2019.00587
Weick EM, Miska EA (2014) piRNAs: from biogenesis to function. Development 141:3458–3471. https://doi.org/10.1242/dev.094037
Ozata DM, Gainetdinov I, Zoch A, O’Carroll D, Zamore PD (2019) PIWI-interacting RNAs: small RNAs with big functions. Nat Rev Genet 20:89–108. https://doi.org/10.1038/s41576-018-0073-3
Williams Z, Morozov P, Mihailovic A, Lin C, Puvvula PK, Juranek S et al (2015) Discovery and characterization of piRNAs in the human fetal ovary. Cell Rep 13:854–863. https://doi.org/10.1016/j.celrep.2015.09.030
Dai X, Shu Y, Lou Q, Tian Q, Zhai G, Song J et al (2017) Tdrd12 is essential for germ cell development and maintenance in zebrafish. Int J Mol Sci 18. https://doi.org/10.3390/ijms18061127
Russell SJ, Stalker L, LaMarre J, PIWIs (2017) piRNAs and retrotransposons: complex battles during reprogramming in gametes and early embryos. Reprod Domest Anim 52(Suppl 4):28–38. https://doi.org/10.1111/rda.13053
Gell SL, Reenan RA (2013) Mutations to the piRNA pathway component aubergine enhance meiotic drive of segregation distorter in Drosophila melanogaster. Genetics 193:771–784. https://doi.org/10.1534/genetics.112.147561
Roush S, Slack FJ (2008) The let-7 family of microRNAs. Trends Cell Biol 18:505–516. https://doi.org/10.1016/j.tcb.2008.07.007
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524. https://doi.org/10.1038/nrm3838
Sharp PA (2001) RNA interference–2001. Genes Dev 15:485–490. https://doi.org/10.1101/gad.880001
Semizarov D, Kroeger P, Fesik S (2004) siRNA-mediated gene silencing: a global genome view. Nucleic Acids Res 32:3836–3845. https://doi.org/10.1093/nar/gkh714
Kiss T (2001) Small nucleolar RNA-guided post-transcriptional modification of cellular RNAs. EMBO J 20:3617–3622. https://doi.org/10.1093/emboj/20.14.3617
Li X, Yang L, Chen LL (2018) The Biogenesis, functions, and challenges of Circular RNAs. Mol Cell 71:428–442. https://doi.org/10.1016/j.molcel.2018.06.034
Wang X, Ramat A, Simonelig M, Liu MF (2023) Emerging roles and functional mechanisms of PIWI-interacting RNAs. Nat Rev Mol Cell Biol 24:123–141. https://doi.org/10.1038/s41580-022-00528-0
Iwasaki YW, Siomi MC, Siomi H, PIWI-Interacting RNA (2015) Its Biogenesis and functions. Annu Rev Biochem 84:405–433. https://doi.org/10.1146/annurev-biochem-060614-034258
Wilusz JE, Sunwoo H, Spector DL (2009) Long noncoding RNAs: functional surprises from the RNA world. Genes Dev 23:1494–1504. https://doi.org/10.1101/gad.1800909
Khan MR, Avino M, Wellinger RJ, Laurent B (2023) Distinct regulatory functions and biological roles of lncRNA splice variants. Mol Ther Nucleic Acids 32:127–143. https://doi.org/10.1016/j.omtn.2023.03.004
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK et al (2013) Natural RNA circles function as efficient microRNA sponges. Nature 495:384–388. https://doi.org/10.1038/nature11993
Caldwell AB, Cheng Z, Vargas JD, Birnbaum HA, Hoffmann A (2014) Network dynamics determine the autocrine and paracrine signaling functions of TNF. Genes Dev 28:2120–2133. https://doi.org/10.1101/gad.244749.114
Jeck WR, Sorrentino JA, Wang K, Slevin MK, Burd CE, Liu J et al (2013) Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19:141–157. https://doi.org/10.1261/rna.035667.112
Lin H, Spradling AC (1997) A novel group of pumilio mutations affects the asymmetric division of germline stem cells in the Drosophila ovary. Development 124:2463–2476. https://doi.org/10.1242/dev.124.12.2463
Cox DN, Chao A, Baker J, Chang L, Qiao D, Lin H (1998) A novel class of evolutionarily conserved genes defined by piwi are essential for stem cell self-renewal. Genes Dev 12:3715–3727. https://doi.org/10.1101/gad.12.23.3715
Klattenhoff C, Theurkauf W (2008) Biogenesis and germline functions of piRNAs. Development 135:3–9. https://doi.org/10.1242/dev.006486
Deng W, Lin H (2002) Miwi, a murine homolog of piwi, encodes a cytoplasmic protein essential for spermatogenesis. Dev Cell 2:819–830. https://doi.org/10.1016/s1534-5807(02)00165-x
Kuramochi-Miyagawa S, Kimura T, Ijiri TW, Isobe T, Asada N, Fujita Y et al (2004) Mili, a mammalian member of piwi family gene, is essential for spermatogenesis. Development 131:839–849. https://doi.org/10.1242/dev.00973
Carmell MA, Girard A, van de Kant HJ, Bourc’his D, Bestor TH, de Rooij DG et al (2007) MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell 12:503–514. https://doi.org/10.1016/j.devcel.2007.03.001
Sharma AK, Nelson MC, Brandt JE, Wessman M, Mahmud N, Weller KP et al (2001) Human CD34(+) stem cells express the hiwi gene, a human homologue of the Drosophila gene piwi. Blood 97:426–434. https://doi.org/10.1182/blood.v97.2.426
Sasaki T, Shiohama A, Minoshima S, Shimizu N (2003) Identification of eight members of the Argonaute family in the human genome. Genomics 82:323–330. https://doi.org/10.1016/s0888-7543(03)00129-0
Aravin A, Gaidatzis D, Pfeffer S, Lagos-Quintana M, Landgraf P, Iovino N et al (2006) A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442:203–207. https://doi.org/10.1038/nature04916
Girard A, Sachidanandam R, Hannon GJ, Carmell MA (2006) A germline-specific class of small RNAs binds mammalian piwi proteins. Nature 442:199–202. https://doi.org/10.1038/nature04917
Lau NC, Seto AG, Kim J, Kuramochi-Miyagawa S, Nakano T, Bartel DP et al (2006) Characterization of the piRNA complex from rat testes. Science 313:363–367. https://doi.org/10.1126/science.1130164
Grivna ST, Beyret E, Wang Z, Lin H (2006) A novel class of small RNAs in mouse spermatogenic cells. Genes Dev 20:1709–1714. https://doi.org/10.1101/gad.1434406
Ro S, Park C, Song R, Nguyen D, Jin J, Sanders KM et al (2007) Cloning and expression profiling of testis-expressed piRNA-like RNAs. RNA 13:1693–1702. https://doi.org/10.1261/rna.640307
Thomson T, Lin H (2009) The biogenesis and function of PIWI proteins and piRNAs: progress and prospect. Annu Rev Cell Dev Biol 25:355–376. https://doi.org/10.1146/annurev.cellbio.24.110707.175327
Czech B, Hannon GJ (2016) One Loop to rule them all: the Ping-Pong cycle and piRNA-Guided silencing. Trends Biochem Sci 41:324–337. https://doi.org/10.1016/j.tibs.2015.12.008
Vourekas A, Zheng Q, Alexiou P, Maragkakis M, Kirino Y, Gregory BD et al (2012) Mili and Miwi target RNA repertoire reveals piRNA biogenesis and function of Miwi in spermiogenesis. Nat Struct Mol Biol 19:773–781. https://doi.org/10.1038/nsmb.2347
Reuter M, Chuma S, Tanaka T, Franz T, Stark A, Pillai RS (2009) Loss of the Mili-interacting Tudor domain-containing protein-1 activates transposons and alters the Mili-associated small RNA profile. Nat Struct Mol Biol 16:639–646. https://doi.org/10.1038/nsmb.1615
Wang J, Saxe JP, Tanaka T, Chuma S, Lin H (2009) Mili interacts with tudor domain-containing protein 1 in regulating spermatogenesis. Curr Biol 19:640–644. https://doi.org/10.1016/j.cub.2009.02.061
Unhavaithaya Y, Hao Y, Beyret E, Yin H, Kuramochi-Miyagawa S, Nakano T et al (2009) MILI, a PIWI-interacting RNA-binding protein, is required for germ line stem cell self-renewal and appears to positively regulate translation. J Biol Chem 284:6507–6519. https://doi.org/10.1074/jbc.M809104200
Roovers EF, Rosenkranz D, Mahdipour M, Han CT, He N, Lopes CdeS (2015) Piwi proteins and piRNAs in mammalian oocytes and early embryos. Cell Rep 10:2069–2082. https://doi.org/10.1016/j.celrep.2015.02.062
Tan M, Tol H, Rosenkranz D, Roovers EF, Damen MJ, Stout TAE et al (2020) PIWIL3 Forms a Complex with TDRKH in Mammalian Oocytes. Cells 9. https://doi.org/10.3390/cells9061356
Hasuwa H, Iwasaki YW, Au Yeung WK, Ishino K, Masuda H, Sasaki H et al (2021) Production of functional oocytes requires maternally expressed PIWI genes and piRNAs in golden hamsters. Nat Cell Biol 23:1002–1012. https://doi.org/10.1038/s41556-021-00745-3
Bao J, Zhang Y, Schuster AS, Ortogero N, Nilsson EE, Skinner MK et al (2014) Conditional inactivation of Miwi2 reveals that MIWI2 is only essential for prospermatogonial development in mice. Cell Death Differ 21:783–796. https://doi.org/10.1038/cdd.2014.5
Aravin AA, Sachidanandam R, Girard A, Fejes-Toth K, Hannon GJ (2007) Developmentally regulated piRNA clusters implicate MILI in transposon control. Science 316:744–747. https://doi.org/10.1126/science.1142612
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Totoki Y, Toyoda A, Ikawa M et al (2008) DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev 22:908–917. https://doi.org/10.1101/gad.1640708
Li XZ, Roy CK, Dong X, Bolcun-Filas E, Wang J, Han BW et al (2013) An ancient transcription factor initiates the burst of piRNA production during early meiosis in mouse testes. Mol Cell 50:67–81. https://doi.org/10.1016/j.molcel.2013.02.016
Brennecke J, Aravin AA, Stark A, Dus M, Kellis M, Sachidanandam R et al (2007) Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128:1089–1103. https://doi.org/10.1016/j.cell.2007.01.043
Ipsaro JJ, Haase AD, Knott SR, Joshua-Tor L, Hannon GJ (2012) The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature 491:279–283. https://doi.org/10.1038/nature11502
Izumi N, Shoji K, Sakaguchi Y, Honda S, Kirino Y, Suzuki T et al (2016) Identification and functional analysis of the Pre-piRNA 3’ Trimmer in silkworms. Cell 164:962–973. https://doi.org/10.1016/j.cell.2016.01.008
Watanabe T, Chuma S, Yamamoto Y, Kuramochi-Miyagawa S, Totoki Y, Toyoda A et al (2011) MITOPLD is a mitochondrial protein essential for nuage formation and piRNA biogenesis in the mouse germline. Dev Cell 20:364–375. https://doi.org/10.1016/j.devcel.2011.01.005
Hayashi R, Schnabl J, Handler D, Mohn F, Ameres SL, Brennecke J (2016) Genetic and mechanistic diversity of piRNA 3’-end formation. Nature 539:588–592. https://doi.org/10.1038/nature20162
Ding D, Liu J, Dong K, Midic U, Hess RA, Xie H et al (2017) PNLDC1 is essential for piRNA 3’ end trimming and transposon silencing during spermatogenesis in mice. Nat Commun 8:819. https://doi.org/10.1038/s41467-017-00854-4
Lim SL, Qu ZP, Kortschak RD, Lawrence DM, Geoghegan J, Hempfling AL et al (2015) HENMT1 and piRNA Stability are required for adult male germ cell transposon repression and to define the Spermatogenic Program in the mouse. PLoS Genet 11:e1005620. https://doi.org/10.1371/journal.pgen.1005620
Beyret E, Liu N, Lin H (2012) piRNA biogenesis during adult spermatogenesis in mice is independent of the ping-pong mechanism. Cell Res 22:1429–1439. https://doi.org/10.1038/cr.2012.120
Gunawardane LS, Saito K, Nishida KM, Miyoshi K, Kawamura Y, Nagami T et al (2007) A slicer-mediated mechanism for repeat-associated siRNA 5’ end formation in Drosophila. Science 315:1587–1590. https://doi.org/10.1126/science.1140494
Mohn F, Handler D, Brennecke J, Noncoding RNA (2015) piRNA-guided slicing specifies transcripts for Zucchini-dependent, phased piRNA biogenesis. Science 348:812–817. https://doi.org/10.1126/science.aaa1039
Sun YH, Zhu J, Xie LH, Li Z, Meduri R, Zhu X et al (2020) Ribosomes guide pachytene piRNA formation on long intergenic piRNA precursors. Nat Cell Biol 22:200–212. https://doi.org/10.1038/s41556-019-0457-4
Sun YH, Wang RH, Du K, Zhu J, Zheng J, Xie LH et al (2021) Coupled protein synthesis and ribosome-guided piRNA processing on mRNAs. Nat Commun 12:5970. https://doi.org/10.1038/s41467-021-26233-8
de Kretser DM, Loveland KL, Meinhardt A, Simorangkir D, Wreford N (1998) Spermatogenesis Hum Reprod 13(Suppl 1):1–8. https://doi.org/10.1093/humrep/13.suppl_1.1
Ginsburg M, Snow MH, McLaren A (1990) Primordial germ cells in the mouse embryo during gastrulation. Development 110:521–528. https://doi.org/10.1242/dev.110.2.521
Hermann BP, Sukhwani M, Hansel MC, Orwig KE (2010) Spermatogonial stem cells in higher primates: are there differences from those in rodents? Reproduction 139:479–493. https://doi.org/10.1530/REP-09-0255
Clermont Y (1966) Spermatogenesis in man. A study of the spermatogonial population. Fertil Steril 17:705–721
Ehmcke J, Wistuba J, Schlatt S (2006) Spermatogonial stem cells: questions, models and perspectives. Hum Reprod Update 12:275–282. https://doi.org/10.1093/humupd/dmk001
Law NC, Oatley MJ, Oatley JM (2019) Developmental kinetics and transcriptome dynamics of stem cell specification in the spermatogenic lineage. Nat Commun 10:2787. https://doi.org/10.1038/s41467-019-10596-0
Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF et al (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31:785–799. https://doi.org/10.1016/j.molcel.2008.09.003
Xu M, You Y, Hunsicker P, Hori T, Small C, Griswold MD et al (2008) Mice deficient for a small cluster of Piwi-interacting RNAs implicate Piwi-interacting RNAs in transposon control. Biol Reprod 79:51–57. https://doi.org/10.1095/biolreprod.108.068072
Reuter M, Berninger P, Chuma S, Shah H, Hosokawa M, Funaya C et al (2011) Miwi catalysis is required for piRNA amplification-independent LINE1 transposon silencing. Nature 480:264–267. https://doi.org/10.1038/nature10672
Watanabe T, Cheng EC, Zhong M, Lin H (2015) Retrotransposons and pseudogenes regulate mRNAs and lncRNAs via the piRNA pathway in the germline. Genome Res 25:368–380. https://doi.org/10.1101/gr.180802.114
Gou LT, Dai P, Yang JH, Xue Y, Hu YP, Zhou Y et al (2014) Pachytene piRNAs instruct massive mRNA elimination during late spermiogenesis. Cell Res 24:680–700. https://doi.org/10.1038/cr.2014.41
Theron E, Maupetit-Mehouas S, Pouchin P, Baudet L, Brasset E, Vaury C (2018) The interplay between the Argonaute proteins Piwi and Aub within Drosophila germarium is critical for oogenesis, piRNA biogenesis and TE silencing. Nucleic Acids Res 46:10052–10065. https://doi.org/10.1093/nar/gky695
Loubalova Z, Fulka H, Horvat F, Pasulka J, Malik R, Hirose M et al (2021) Formation of spermatogonia and fertile oocytes in golden hamsters requires piRNAs. Nat Cell Biol 23:992–1001. https://doi.org/10.1038/s41556-021-00746-2
Zhang Y, Guo R, Cui Y, Zhu Z, Zhang Y, Wu H et al (2017) An essential role for PNLDC1 in piRNA 3’ end trimming and male fertility in mice. Cell Res 27:1392–1396. https://doi.org/10.1038/cr.2017.125
Bagijn MP, Goldstein LD, Sapetschnig A, Weick EM, Bouasker S, Lehrbach NJ et al (2012) Function, targets, and evolution of Caenorhabditis elegans piRNAs. Science 337:574–578. https://doi.org/10.1126/science.1220952
Nagirnaja L, Morup N, Nielsen JE, Stakaitis R, Golubickaite I, Oud MS et al (2021) Variant PNLDC1, defective piRNA Processing, and Azoospermia. N Engl J Med 385:707–719. https://doi.org/10.1056/NEJMoa2028973
Megosh HB, Cox DN, Campbell C, Lin H (2006) The role of PIWI and the miRNA machinery in Drosophila germline determination. Curr Biol 16:1884–1894. https://doi.org/10.1016/j.cub.2006.08.051
Leslie M (2013) Cell biology. The immune system’s compact genomic counterpart. Science 339:25–27. https://doi.org/10.1126/science.339.6115.25
Toth KF, Pezic D, Stuwe E, Webster A (2016) The piRNA Pathway guards the germline genome against transposable elements. Adv Exp Med Biol 886:51–77. https://doi.org/10.1007/978-94-017-7417-8_4
Rocha-da-Silva L, Armelin-Correa L, Cantao IH, Flister VJF, Nunes M, Stumpp T (2019) Expression of genome defence protein members in proliferating and quiescent rat male germ cells and the Nuage dynamics. PLoS ONE 14:e0217941. https://doi.org/10.1371/journal.pone.0217941
Huang H, Gao Q, Peng X, Choi SY, Sarma K, Ren H et al (2011) piRNA-associated germline nuage formation and spermatogenesis require MitoPLD profusogenic mitochondrial-surface lipid signaling. Dev Cell 20:376–387. https://doi.org/10.1016/j.devcel.2011.01.004
Nishimura T, Nagamori I, Nakatani T, Izumi N, Tomari Y, Kuramochi-Miyagawa S et al (2018) PNLDC1, mouse pre-piRNA Trimmer, is required for meiotic and post-meiotic male germ cell development. EMBO Rep 19. https://doi.org/10.15252/embr.201744957
Guan Y, Keeney S, Jain D, Wang PJ (2021) Yama, a mutant allele of Mov10l1, disrupts retrotransposon silencing and piRNA biogenesis. PLoS Genet 17:e1009265. https://doi.org/10.1371/journal.pgen.1009265
Kherraf ZE, Cazin C, Bouker A, Fourati Ben Mustapha S, Hennebicq S, Septier A et al (2022) Whole-exome sequencing improves the diagnosis and care of men with non-obstructive azoospermia. Am J Hum Genet 109:508–517. https://doi.org/10.1016/j.ajhg.2022.01.011
Wasik KA, Tam OH, Knott SR, Falciatori I, Hammell M, Vagin VV et al (2015) RNF17 blocks promiscuous activity of PIWI proteins in mouse testes. Genes Dev 29:1403–1415. https://doi.org/10.1101/gad.265215.115
Zheng K, Xiol J, Reuter M, Eckardt S, Leu NA, McLaughlin KJ et al (2010) Mouse MOV10L1 associates with Piwi proteins and is an essential component of the Piwi-interacting RNA (piRNA) pathway. Proc Natl Acad Sci U S A 107:11841–11846. https://doi.org/10.1073/pnas.1003953107
Mathioudakis N, Palencia A, Kadlec J, Round A, Tripsianes K, Sattler M et al (2012) The multiple Tudor domain-containing protein TDRD1 is a molecular scaffold for mouse piwi proteins and piRNA biogenesis factors. RNA 18:2056–2072. https://doi.org/10.1261/rna.034181.112
De Fazio S, Bartonicek N, Di Giacomo M, Abreu-Goodger C, Sankar A, Funaya C et al (2011) The endonuclease activity of Mili fuels piRNA amplification that silences LINE1 elements. Nature 480:259–263. https://doi.org/10.1038/nature10547
Zheng K, Wang PJ (2012) Blockade of pachytene piRNA biogenesis reveals a novel requirement for maintaining post-meiotic germline genome integrity. PLoS Genet 8:e1003038. https://doi.org/10.1371/journal.pgen.1003038
Goh WS, Falciatori I, Tam OH, Burgess R, Meikar O, Kotaja N et al (2015) piRNA-directed cleavage of meiotic transcripts regulates spermatogenesis. Genes Dev 29:1032–1044. https://doi.org/10.1101/gad.260455.115
Saritas G, Main AM, Winge SB, Morup N, Almstrup K (2022) PIWI-interacting RNAs and human testicular function. WIREs Mech Dis 14:e1572. https://doi.org/10.1002/wsbm.1572
Yang Q, Hua J, Wang L, Xu B, Zhang H, Ye N et al (2013) MicroRNA and piRNA profiles in normal human testis detected by next generation sequencing. PLoS ONE 8:e66809. https://doi.org/10.1371/journal.pone.0066809
Zhang P, Kang JY, Gou LT, Wang J, Xue Y, Skogerboe G et al (2015) MIWI and piRNA-mediated cleavage of messenger RNAs in mouse testes. Cell Res 25:193–207. https://doi.org/10.1038/cr.2015.4
Vourekas A, Zheng K, Fu Q, Maragkakis M, Alexiou P, Ma J et al (2015) The RNA helicase MOV10L1 binds piRNA precursors to initiate piRNA processing. Genes Dev 29:617–629. https://doi.org/10.1101/gad.254631.114
Tan K, Song HW, Wilkinson MF (2021) RHOX10 drives mouse spermatogonial stem cell establishment through a transcription factor signaling cascade. Cell Rep 36:109423. https://doi.org/10.1016/j.celrep.2021.109423
Yang Z, Chen KM, Pandey RR, Homolka D, Reuter M, Janeiro BK et al (2016) PIWI Slicing and EXD1 Drive Biogenesis of Nuclear piRNAs from cytosolic targets of the mouse piRNA pathway. Mol Cell 61:138–152. https://doi.org/10.1016/j.molcel.2015.11.009
Shiromoto Y, Kuramochi-Miyagawa S, Nagamori I, Chuma S, Arakawa T, Nishimura T et al (2019) GPAT2 is required for piRNA biogenesis, transposon silencing, and maintenance of spermatogonia in micedagger. Biol Reprod 101:248–256. https://doi.org/10.1093/biolre/ioz056
Yoshimura T, Watanabe T, Kuramochi-Miyagawa S, Takemoto N, Shiromoto Y, Kudo A et al (2018) Mouse GTSF1 is an essential factor for secondary piRNA biogenesis. EMBO Rep 19. https://doi.org/10.15252/embr.201642054
Pandey RR, Homolka D, Olotu O, Sachidanandam R, Kotaja N, Pillai RS (2018) Exonuclease Domain-containing 1 enhances MIWI2 piRNA Biogenesis via its Interaction with TDRD12. Cell Rep 24:3423–3432e3424. https://doi.org/10.1016/j.celrep.2018.08.087
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Takamatsu K, Chuma S, Kojima-Kita K et al (2010) MVH in piRNA processing and gene silencing of retrotransposons. Genes Dev 24:887–892. https://doi.org/10.1101/gad.1902110
Kojima-Kita K, Kuramochi-Miyagawa S, Nakayama M, Miyata H, Jacobsen SE, Ikawa M et al (2021) MORC3, a novel MIWI2 association partner, as an epigenetic regulator of piRNA dependent transposon silencing in male germ cells. Sci Rep 11:20472. https://doi.org/10.1038/s41598-021-98940-7
Molaro A, Falciatori I, Hodges E, Aravin AA, Marran K, Rafii S et al (2014) Two waves of de novo methylation during mouse germ cell development. Genes Dev 28:1544–1549. https://doi.org/10.1101/gad.244350.114
Czech B, Munafo M, Ciabrelli F, Eastwood EL, Fabry MH, Kneuss E et al (2018) piRNA-Guided Genome Defense: from Biogenesis to silencing. Annu Rev Genet 52:131–157. https://doi.org/10.1146/annurev-genet-120417-031441
Manakov SA, Pezic D, Marinov GK, Pastor WA, Sachidanandam R, Aravin AA (2015) MIWI2 and MILI have Differential effects on piRNA Biogenesis and DNA methylation. Cell Rep 12:1234–1243. https://doi.org/10.1016/j.celrep.2015.07.036
Shoji M, Tanaka T, Hosokawa M, Reuter M, Stark A, Kato Y et al (2009) The TDRD9-MIWI2 complex is essential for piRNA-mediated retrotransposon silencing in the mouse male germline. Dev Cell 17:775–787. https://doi.org/10.1016/j.devcel.2009.10.012
Di Giacomo M, Comazzetto S, Sampath SC, Sampath SC, O’Carroll D (2014) G9a co-suppresses LINE1 elements in spermatogonia. Epigenetics Chromatin 7:24. https://doi.org/10.1186/1756-8935-7-24
Di Giacomo M, Comazzetto S, Saini H, De Fazio S, Carrieri C, Morgan M et al (2013) Multiple epigenetic mechanisms and the piRNA pathway enforce LINE1 silencing during adult spermatogenesis. Mol Cell 50:601–608. https://doi.org/10.1016/j.molcel.2013.04.026
Lee JH, Engel W, Nayernia K (2006) Stem cell protein Piwil2 modulates expression of murine spermatogonial stem cell expressed genes. Mol Reprod Dev 73:173–179. https://doi.org/10.1002/mrd.20391
Tan K, Kim ME, Song HW, Skarbrevik D, Babajanian E, Bedrosian TA et al (2021) The rhox gene cluster suppresses germline LINE1 transposition. Proc Natl Acad Sci U S A 118. https://doi.org/10.1073/pnas.2024785118
Song HW, Bettegowda A, Lake BB, Zhao AH, Skarbrevik D, Babajanian E et al (2016) The homeobox transcription factor RHOX10 drives mouse Spermatogonial Stem Cell Establishment. Cell Rep 17:149–164. https://doi.org/10.1016/j.celrep.2016.08.090
Bronkhorst AW, Ketting RF (2018) Trimming it short: PNLDC1 is required for piRNA maturation during mouse spermatogenesis. EMBO Rep 19. https://doi.org/10.15252/embr.201845824
Gainetdinov I, Colpan C, Cecchini K, Arif A, Jouravleva K, Albosta P et al (2021) Terminal modification, sequence, length, and PIWI-protein identity determine piRNA stability. Mol Cell 81:4826–4842e4828. https://doi.org/10.1016/j.molcel.2021.09.012
Zhao MZ, Lin DH, Zuo H, Wei H, Wang X, Gou LT et al (2022) piRNA 3’ uridylation facilitates the assembly of MIWI/piRNA complex for efficient target regulation in mouse male germ cells. Cell Res 32:1030–1033. https://doi.org/10.1038/s41422-022-00659-1
Li L, Tan YQ, Lu LY (2022) Defective piRNA Processing and Azoospermia. N Engl J Med 386:1675–1676. https://doi.org/10.1056/NEJMc2116008
Zhu X, Zhi E, Li Z (2015) MOV10L1 in piRNA processing and gene silencing of retrotransposons during spermatogenesis. Reproduction 149:R229–235. https://doi.org/10.1530/REP-14-0569
Zheng K, Xiol J, Reuter M, Eckardt S, Leu NA, McLaughlin KJ et al (2010) Mouse MOV10L1 associates with Piwi proteins and is an essential component of the Piwi-interacting RNA (piRNA) pathway. P Natl Acad Sci USA 107:11841–11846. https://doi.org/10.1073/pnas.1003953107
Fu Q, Pandey RR, Leu NA, Pillai RS, Wang PJ (2016) Mutations in the MOV10L1 ATP hydrolysis motif cause piRNA Biogenesis failure and male sterility in mice. Biol Reprod 95. ARTN 103. https://doi.org/10.1095/biolreprod.116.142430
Hadziselimovic F, Hadziselimovic NO, Demougin P, Krey G, Oakeley EJ (2011) Deficient expression of genes involved in the endogenous defense system against transposons in cryptorchid boys with impaired mini-puberty. Sex Dev 5:287–293. https://doi.org/10.1159/000335188
Wenda JM, Homolka D, Yang Z, Spinelli P, Sachidanandam R, Pandey RR et al (2017) Distinct roles of RNA helicases MVH and TDRD9 in PIWI slicing-triggered mammalian piRNA Biogenesis and function. Dev Cell 41:623–637e629. https://doi.org/10.1016/j.devcel.2017.05.021
Ma L, Buchold GM, Greenbaum MP, Roy A, Burns KH, Zhu H et al (2009) GASZ is essential for male meiosis and suppression of retrotransposon expression in the male germline. PLoS Genet 5:e1000635. https://doi.org/10.1371/journal.pgen.1000635
Olotu O, Dowling M, Homolka D, Wojtas MN, Tran P, Lehtiniemi T et al (2023) Intermitochondrial cement (IMC) harbors piRNA biogenesis machinery and exonuclease domain-containing proteins EXD1 and EXD2 in mouse spermatocytes. Andrology 11:710–723. https://doi.org/10.1111/andr.13361
Pandey RR, Tokuzawa Y, Yang Z, Hayashi E, Ichisaka T, Kajita S et al (2013) Tudor domain containing 12 (TDRD12) is essential for secondary PIWI interacting RNA biogenesis in mice. Proc Natl Acad Sci U S A 110:16492–16497. https://doi.org/10.1073/pnas.1316316110
Shiromoto Y, Kuramochi-Miyagawa S, Daiba A, Chuma S, Katanaya A, Katsumata A et al (2013) GPAT2, a mitochondrial outer membrane protein, in piRNA biogenesis in germline stem cells. RNA 19:803–810. https://doi.org/10.1261/rna.038521.113
Quenerch’du E, Anand A, Kai T (2016) The piRNA pathway is developmentally regulated during spermatogenesis in Drosophila. RNA 22:1044–1054. https://doi.org/10.1261/rna.055996.116
Dai P, Wang X, Gou LT, Li ZT, Wen Z, Chen ZG et al (2019) A translation-activating function of MIWI/piRNA during Mouse Spermiogenesis. Cell 179:1566–1581e1516. https://doi.org/10.1016/j.cell.2019.11.022
Ding D, Liu J, Midic U, Wu Y, Dong K, Melnick A et al (2018) TDRD5 binds piRNA precursors and selectively enhances pachytene piRNA processing in mice. Nat Commun 9:127. https://doi.org/10.1038/s41467-017-02622-w
Dong J, Wang X, Cao C, Wen Y, Sakashita A, Chen S et al (2019) UHRF1 suppresses retrotransposons and cooperates with PRMT5 and PIWI proteins in male germ cells. Nat Commun 10:4705. https://doi.org/10.1038/s41467-019-12455-4
Saxe JP, Chen M, Zhao H, Lin H (2013) Tdrkh is essential for spermatogenesis and participates in primary piRNA biogenesis in the germline. EMBO J 32:1869–1885. https://doi.org/10.1038/emboj.2013.121
Lu Y, Nagamori I, Kobayashi H, Kojima-Kita K, Shirane K, Chang HY et al (2023) ADAD2 functions in spermiogenesis and piRNA biogenesis in mice. Andrology 11:698–709. https://doi.org/10.1111/andr.13400
Tarabay Y, Achour M, Teletin M, Ye T, Teissandier A, Mark M et al (2017) Tex19 paralogs are new members of the piRNA pathway controlling retrotransposon suppression. J Cell Sci 130:1463–1474. https://doi.org/10.1242/jcs.188763
Schopp T, Zoch A, Berrens RV, Auchynnikava T, Kabayama Y, Vasiliauskaite L et al (2020) TEX15 is an essential executor of MIWI2-directed transposon DNA methylation and silencing. Nat Commun 11:3739. https://doi.org/10.1038/s41467-020-17372-5
Han B, Jung BK, Park SH, Song KJ, Anwar MA, Ryu KY et al (2021) Polyubiquitin gene Ubb is required for upregulation of Piwi protein level during mouse testis development. Cell Death Discov 7:194. https://doi.org/10.1038/s41420-021-00581-2
Zhou L, Canagarajah B, Zhao Y, Baibakov B, Tokuhiro K, Maric D et al (2017) BTBD18 regulates a Subset of piRNA-Generating loci through transcription elongation in mice. Dev Cell 40:453–466e455. https://doi.org/10.1016/j.devcel.2017.02.007
Soper SF, van der Heijden GW, Hardiman TC, Goodheart M, Martin SL, de Boer P et al (2008) Mouse maelstrom, a component of nuage, is essential for spermatogenesis and transposon repression in meiosis. Dev Cell 15:285–297. https://doi.org/10.1016/j.devcel.2008.05.015
Xiol J, Cora E, Koglgruber R, Chuma S, Subramanian S, Hosokawa M et al (2012) A role for Fkbp6 and the chaperone machinery in piRNA amplification and transposon silencing. Mol Cell 47:970–979. https://doi.org/10.1016/j.molcel.2012.07.019
Hsieh CL, Xia J, Lin H (2020) MIWI prevents aneuploidy during meiosis by cleaving excess satellite RNA. EMBO J 39:e103614. https://doi.org/10.15252/embj.2019103614
Siomi MC, Mannen T, Siomi H (2010) How does the royal family of Tudor rule the PIWI-interacting RNA pathway? Genes dev. 24:636–646. https://doi.org/10.1101/gad.1899210
Kojima K, Kuramochi-Miyagawa S, Chuma S, Tanaka T, Nakatsuji N, Kimura T et al (2009) Associations between PIWI proteins and TDRD1/MTR-1 are critical for integrated subcellular localization in murine male germ cells. Genes Cells 14:1155–1165. https://doi.org/10.1111/j.1365-2443.2009.01342.x
Chen C, Jin J, James DA, Adams-Cioaba MA, Park JG, Guo Y et al (2009) Mouse Piwi interactome identifies binding mechanism of Tdrkh Tudor domain to arginine methylated Miwi. Proc Natl Acad Sci U S A 106:20336–20341. https://doi.org/10.1073/pnas.0911640106
Zhou J, Leu NA, Eckardt S, McLaughlin KJ, Wang PJ (2014) STK31/TDRD8, a germ cell-specific factor, is dispensable for reproduction in mice. PLoS ONE 9:e89471. https://doi.org/10.1371/journal.pone.0089471
Vasileva A, Tiedau D, Firooznia A, Muller-Reichert T, Jessberger R (2009) Tdrd6 is required for spermiogenesis, chromatoid body architecture, and regulation of miRNA expression. Curr Biol 19:630–639. https://doi.org/10.1016/j.cub.2009.02.047
Ding D, Liu J, Dong K, Melnick AF, Latham KE, Chen C (2019) Mitochondrial membrane-based initial separation of MIWI and MILI functions during pachytene piRNA biogenesis. Nucleic Acids Res 47:2594–2608. https://doi.org/10.1093/nar/gky1281
Yang R, Zhang B, Zhu W, Zhu C, Chen L, Zhao Y et al (2023) Expression of phospholipase D Family Member 6 in bovine testes and its molecular characteristics. Int J Mol Sci 24. https://doi.org/10.3390/ijms241512172
Xiong M, Yin L, Gui Y, Lv C, Ma X, Guo S et al (2023) ADAD2 interacts with RNF17 in P-bodies to repress the Ping-pong cycle in pachytene piRNA biogenesis. J Cell Biol 222. https://doi.org/10.1083/jcb.202206067
Chukrallah LG, Potgieter S, Chueh L, Snyder EM (2023) Two RNA binding proteins, ADAD2 and RNF17, interact to form a heterogeneous population of novel meiotic germ cell granules with developmentally dependent organelle association. PLoS Genet 19:e1010519. https://doi.org/10.1371/journal.pgen.1010519
Tanaka T, Hosokawa M, Vagin VV, Reuter M, Hayashi E, Mochizuki AL et al (2011) Tudor domain containing 7 (Tdrd7) is essential for dynamic ribonucleoprotein (RNP) remodeling of chromatoid bodies during spermatogenesis. Proc Natl Acad Sci U S A 108:10579–10584. https://doi.org/10.1073/pnas.1015447108
Chuma S, Hosokawa M, Tanaka T, Nakatsuji N (2009) Ultrastructural characterization of spermatogenesis and its evolutionary conservation in the germline: germinal granules in mammals. Mol Cell Endocrinol 306:17–23. https://doi.org/10.1016/j.mce.2008.11.009
Bao J, Wang L, Lei J, Hu Y, Liu Y, Shen H et al (2012) STK31(TDRD8) is dynamically regulated throughout mouse spermatogenesis and interacts with MIWI protein. Histochem Cell Biol 137:377–389. https://doi.org/10.1007/s00418-011-0897-9
Yang F, Lan Y, Pandey RR, Homolka D, Berger SL, Pillai RS et al (2020) TEX15 associates with MILI and silences transposable elements in male germ cells. Genes Dev 34:745–750. https://doi.org/10.1101/gad.335489.119
Ozata DM, Yu T, Mou H, Gainetdinov I, Colpan C, Cecchini K et al (2020) Evolutionarily conserved pachytene piRNA loci are highly divergent among modern humans. Nat Ecol Evol 4:156–168. https://doi.org/10.1038/s41559-019-1065-1
Yu T, Biasini A, Cecchini K, Saflund M, Mou H, Arif A et al (2022) A-MYB/TCFL5 regulatory architecture ensures the production of pachytene piRNAs in placental mammals. RNA 29:30–43. https://doi.org/10.1261/rna.079472.122
Xu W, Zhang Y, Qin D, Gui Y, Wang S, Du G et al (2022) Transcription factor-like 5 is a potential DNA- and RNA-binding protein essential for maintaining male fertility in mice. J Cell Sci 135. https://doi.org/10.1242/jcs.259036
Galan-Martinez J, Berenguer I, Del Carmen Maza M, Stamatakis K, Girones N, Fresno M (2022) TCFL5 deficiency impairs the pachytene to diplotene transition during spermatogenesis in the mouse. Sci Rep 12:10956. https://doi.org/10.1038/s41598-022-15167-w
Bolcun-Filas E, Bannister LA, Barash A, Schimenti KJ, Hartford SA, Eppig JJ et al (2011) A-MYB (MYBL1) transcription factor is a master regulator of male meiosis. Development 138:3319–3330. https://doi.org/10.1242/dev.067645
Chuma S, Hosokawa M, Kitamura K, Kasai S, Fujioka M, Hiyoshi M et al (2006) Tdrd1/Mtr-1, a tudor-related gene, is essential for male germ-cell differentiation and nuage/germinal granule formation in mice. Proc Natl Acad Sci U S A 103:15894–15899. https://doi.org/10.1073/pnas.0601878103
Zhang H, Liu K, Izumi N, Huang H, Ding D, Ni Z et al (2017) Structural basis for arginine methylation-independent recognition of PIWIL1 by TDRD2. Proc Natl Acad Sci U S A 114:12483–12488. https://doi.org/10.1073/pnas.1711486114
Pan J, Goodheart M, Chuma S, Nakatsuji N, Page DC, Wang PJ (2005) RNF17, a component of the mammalian germ cell nuage, is essential for spermiogenesis. Development 132:4029–4039. https://doi.org/10.1242/dev.02003
Sun YH, Jiang F, Li XZ (2018) Disruption of Tdrd5 decouples the stepwise processing of long precursor transcripts during pachytene PIWI-interacting RNA biogenesis. Biol Reprod 99:684–685. https://doi.org/10.1093/biolre/ioy110
Yabuta Y, Ohta H, Abe T, Kurimoto K, Chuma S, Saitou M (2011) TDRD5 is required for retrotransposon silencing, chromatoid body assembly, and spermiogenesis in mice. J Cell Biol 192:781–795. https://doi.org/10.1083/jcb.201009043
Akpinar M, Lesche M, Fanourgakis G, Fu J, Anastassiadis K, Dahl A et al (2017) TDRD6 mediates early steps of spliceosome maturation in primary spermatocytes. PLoS Genet 13:e1006660. https://doi.org/10.1371/journal.pgen.1006660
Filipescu D, Szenker E, Almouzni G (2013) Developmental roles of histone H3 variants and their chaperones. Trends Genet 29:630–640. https://doi.org/10.1016/j.tig.2013.06.002
Bramlage B, Kosciessa U, Doenecke D, Differentiation (1997) 62: 13–20. https://doi.org/10.1046/j.1432-0436.1997.6210013.x
Szenker E, Ray-Gallet D, Almouzni G (2011) The double face of the histone variant H3.3. Cell Res 21:421–434. https://doi.org/10.1038/cr.2011.14
Fontaine E, Papin C, Martinez G, Le Gras S, Nahed RA, Hery P et al (2022) Dual role of histone variant H3.3B in spermatogenesis: positive regulation of piRNA transcription and implication in X-chromosome inactivation. Nucleic Acids Res 50:7350–7366. https://doi.org/10.1093/nar/gkac541
Costa Y, Speed RM, Gautier P, Semple CA, Maratou K, Turner JM et al (2006) Mouse MAELSTROM: the link between meiotic silencing of unsynapsed chromatin and microRNA pathway? Hum Mol Genet 15:2324–2334. https://doi.org/10.1093/hmg/ddl158
Zhang D, Xiong H, Shan J, Xia X, Trudeau VL (2008) Functional insight into Maelstrom in the germline piRNA pathway: a unique domain homologous to the DnaQ-H 3’-5’ exonuclease, its lineage-specific expansion/loss and evolutionarily active site switch. Biol Direct 3:48. https://doi.org/10.1186/1745-6150-3-48
Castaneda J, Genzor P, van der Heijden GW, Sarkeshik A, Yates JR 3rd, Ingolia NT et al (2014) Reduced pachytene piRNAs and translation underlie spermiogenic arrest in Maelstrom mutant mice. EMBO J 33:1999–2019. https://doi.org/10.15252/embj.201386855
Sato K, Siomi MC (2015) Functional and structural insights into the piRNA factor Maelstrom. FEBS Lett 589:1688–1693. https://doi.org/10.1016/j.febslet.2015.03.023
Pandey RR, Pillai RS (2014) Primary piRNA biogenesis: caught up in a Maelstrom. EMBO J 33:1979–1980. https://doi.org/10.15252/embj.201489670
Cheng YS, Chen HY, Lin YC, Lin YS, Yeh YC, Yeh YH et al (2023) The MAEL expression in mitochondria of human spermatozoa and the association with asthenozoospermia. Andrology. https://doi.org/10.1111/andr.13408
Crackower MA, Kolas NK, Noguchi J, Sarao R, Kikuchi K, Kaneko H et al (2003) Essential role of Fkbp6 in male fertility and homologous chromosome pairing in meiosis. Science 300:1291–1295. https://doi.org/10.1126/science.1083022
Wyrwoll MJ, Gaasbeek CM, Golubickaite I, Stakaitis R, Oud MS, Nagirnaja L et al (2022) The piRNA-pathway factor FKBP6 is essential for spermatogenesis but dispensable for control of meiotic LINE-1 expression in humans. Am J Hum Genet 109:1850–1866. https://doi.org/10.1016/j.ajhg.2022.09.002
Taipale M, Tucker G, Peng J, Krykbaeva I, Lin ZY, Larsen B et al (2014) A quantitative chaperone interaction network reveals the architecture of cellular protein homeostasis pathways. Cell 158:434–448. https://doi.org/10.1016/j.cell.2014.05.039
Ichiyanagi T, Ichiyanagi K, Ogawa A, Kuramochi-Miyagawa S, Nakano T, Chuma S et al (2014) HSP90alpha plays an important role in piRNA biogenesis and retrotransposon repression in mouse. Nucleic Acids Res 42:11903–11911. https://doi.org/10.1093/nar/gku881
Zhang L, Meng X, Xiang Z, Li D, Han X (2018) From the cover: roles of mmu_piR_003399 in microcystin-leucine Arginine-Induced Reproductive toxicity in the Spermatogonial cells and Testis. Toxicol Sci 161:159–170. https://doi.org/10.1093/toxsci/kfx209
Wu PH, Fu Y, Cecchini K, Ozata DM, Arif A, Yu T et al (2020) The evolutionarily conserved piRNA-producing locus pi6 is required for male mouse fertility. Nat Genet 52:728–739. https://doi.org/10.1038/s41588-020-0657-7
Cao C, Wen Y, Wang X, Fang N, Yuan S, Huang X (2018) Testicular piRNA profile comparison between successful and unsuccessful micro-TESE retrieval in NOA patients. J Assist Reprod Genet 35:801–808. https://doi.org/10.1007/s10815-018-1134-4
Hong Y, Wang C, Fu Z, Liang H, Zhang S, Lu M et al (2016) Systematic characterization of seminal plasma piRNAs as molecular biomarkers for male infertility. Sci Rep 6:24229. https://doi.org/10.1038/srep24229
Cui L, Fang L, Shi B, Qiu S, Ye Y (2018) Spermatozoa expression of piR-31704, piR-39888, and piR-40349 and their correlation to sperm concentration and fertilization rate after ICSI. Reprod Sci 25:733–739. https://doi.org/10.1177/1933719117725822
Shen L, Yu C, Wu Y, Jiang Y, Sun Y, Yao J et al (2022) Transcriptomic landscape of GC-2spd(ts) cell in response to the downregulation of piR-1207/2107. Andrologia 54:e14522. https://doi.org/10.1111/and.14522
Kumar K, Trzybulska D, Tsatsanis C, Giwercman A, Almstrup K (2019) Identification of circulating small non-coding RNAs in relation to male subfertility and reproductive hormones. Mol Cell Endocrinol 492:110443. https://doi.org/10.1016/j.mce.2019.05.002
Hempfling AL, Lim SL, Adelson DL, Evans J, O’Connor AE, Qu ZP et al (2017) Expression patterns of HENMT1 and PIWIL1 in human testis: implications for transposon expression. Reproduction 154:363–374. https://doi.org/10.1530/REP-16-0586
Heyn H, Ferreira HJ, Bassas L, Bonache S, Sayols S, Sandoval J et al (2012) Epigenetic disruption of the PIWI pathway in human spermatogenic disorders. PLoS ONE 7:e47892. https://doi.org/10.1371/journal.pone.0047892
Sha Y, Li L, Yin C (2022) Defective piRNA Processing and Azoospermia. N Engl J Med 386:1675. https://doi.org/10.1056/NEJMc2116008
Wang X, Tan YQ, Liu MF (2022) Defective piRNA Processing and Azoospermia. N Engl J Med 386:1674–1675. https://doi.org/10.1056/NEJMc2116008
Gou LT, Kang JY, Dai P, Wang X, Li F, Zhao S et al (2017) Ubiquitination-deficient mutations in human piwi cause male infertility by impairing histone-to-Protamine Exchange during Spermiogenesis. Cell 169:1090–1104e1013. https://doi.org/10.1016/j.cell.2017.04.034
Hasuwa H, Ishino K, Siomi H, Human (2018) PIWI (HIWI) is an azoospermia factor. Sci China Life Sci 61:348–350. https://doi.org/10.1007/s11427-017-9149-0
Kamaliyan Z, Pouriamanesh S, Soosanabadi M, Gholami M, Mirfakhraie R (2018) Investigation of piwi-interacting RNA pathway genes role in idiopathic non-obstructive azoospermia. Sci Rep 8:142. https://doi.org/10.1038/s41598-017-17518-4
Gu A, Ji G, Shi X, Long Y, Xia Y, Song L et al (2010) Genetic variants in Piwi-interacting RNA pathway genes confer susceptibility to spermatogenic failure in a Chinese population. Hum Reprod 25:2955–2961. https://doi.org/10.1093/humrep/deq274
Zhu XB, Lu JQ, Zhi EL, Zhu Y, Zou SS, Zhu ZJ et al (2016) Association of a TDRD1 variant with spermatogenic failure susceptibility in the Han Chinese. J Assist Reprod Genet 33:1099–1104. https://doi.org/10.1007/s10815-016-0738-9
Babakhanzadeh E, Khodadadian A, Rostami S, Alipourfard I, Aghaei M, Nazari M et al (2020) Testicular expression of TDRD1, TDRD5, TDRD9 and TDRD12 in azoospermia. BMC Med Genet 21:33. https://doi.org/10.1186/s12881-020-0970-0
Sarkardeh H, Totonchi M, Asadpour O, Sadighi Gilani MA, Zamani Esteki M, Almadani N et al (2014) Association of MOV10L1 gene polymorphisms and male infertility in azoospermic men with complete maturation arrest. J Assist Reprod Genet 31:865–871. https://doi.org/10.1007/s10815-014-0240-1
Ge W, Chen M, Tian W, Chen J, Zhao Y, Xian H et al (2021) Global 3’UTR shortening and down-regulation of repeated element related piRNA play crucial roles in boys with cryptorchidism. Genomics 113:633–645. https://doi.org/10.1016/j.ygeno.2021.01.006
Hadziselimovic F, Hadziselimovic NO, Demougin P, Krey G, Oakeley E (2015) Piwi-pathway alteration induces LINE-1 transposon derepression and infertility development in cryptorchidism. Sex Dev 9:98–104. https://doi.org/10.1159/000375351
Rounge TB, Furu K, Skotheim RI, Haugen TB, Grotmol T, Enerly E (2015) Profiling of the small RNA populations in human testicular germ cell tumors shows global loss of piRNAs. Mol Cancer 14:153. https://doi.org/10.1186/s12943-015-0411-4
Gainetdinov IV, Skvortsova YV, Kondratieva SA, Klimov A, Tryakin AA, Azhikina TL (2018) Assessment of piRNA biogenesis and function in testicular germ cell tumors and their precursor germ cell neoplasia in situ. BMC Cancer 18:20. https://doi.org/10.1186/s12885-017-3945-6
Ferreira HJ, Heyn H, Garcia del Muro X, Vidal A, Larriba S, Munoz C et al (2014) Epigenetic loss of the PIWI/piRNA machinery in human testicular tumorigenesis. Epigenetics 9:113–118. https://doi.org/10.4161/epi.27237
Han H, Fan G, Song S, Jiang Y, Qian C, Zhang W et al (2021) piRNA-30473 contributes to tumorigenesis and poor prognosis by regulating m6A RNA methylation in DLBCL. Blood 137:1603–1614. https://doi.org/10.1182/blood.2019003764
Li Y, Min C, Zhao Y, Wang J, Wang J (2021) Effects of fluoride on PIWI-interacting RNA expression profiling in testis of mice. Chemosphere 269:128727. https://doi.org/10.1016/j.chemosphere.2020.128727
Kong L, Wu Y, Hu W, Liu L, Xue Y, Liang G (2021) Mechanisms underlying reproductive toxicity induced by nickel nanoparticles identified by comprehensive gene expression analysis in GC-1 spg cells. Environ Pollut 275:116556. https://doi.org/10.1016/j.envpol.2021.116556
Setten RL, Rossi JJ, Han SP (2019) The current state and future directions of RNAi-based therapeutics. Nat Rev Drug Discov 18:421–446. https://doi.org/10.1038/s41573-019-0017-4
Hu B, Zhong L, Weng Y, Peng L, Huang Y, Zhao Y et al (2020) Therapeutic siRNA: state of the art. Signal Transduct Target Ther 5:101. https://doi.org/10.1038/s41392-020-0207-x
Fortner A, Schumacher D (2021) First COVID-19 vaccines receiving the US FDA and EMA Emergency Use Authorization. Discoveries (Craiova) 9(e122). https://doi.org/10.15190/d.2021.1
Xu J, Ji L, Liang Y, Wan Z, Zheng W, Song X et al (2020) CircRNA-SORE mediates sorafenib resistance in hepatocellular carcinoma by stabilizing YBX1. Signal Transduct Target Ther 5:298. https://doi.org/10.1038/s41392-020-00375-5
Wang X, Lin DH, Yan Y, Wang AH, Liao J, Meng Q et al (2023) The PIWI-specific insertion module helps load longer piRNAs for translational activation essential for male fertility. Sci China Life Sci 66:1459–1481. https://doi.org/10.1007/s11427-023-2390-5
Acknowledgements
This work was supported by the grants from the National Nature Science Foundation of China (32170862, 31872845), Major Scientific and Technological Projects for Collaborative Prevention and Control of Birth Defect in Hunan Province (2019SK1012), Developmental Biology and Breeding (2022XKQ0205), Research Team for Reproduction Health and Translational Medicine of Hunan Normal University (2023JC101), Natural Science Foundation of Hunan Province (2023JJ30424), Hunan Provincial Innovation Foundation for Postgraduate (CX20230465), and a grant from the Shanghai Key Laboratory of Reproductive Medicine.
Author information
Authors and Affiliations
Contributions
L.D., W.C., D.Z. and Y.C. wrote the manuscript. Z.H. was responsible for study conception, revising the manuscript, and final approval of the manuscript. All authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Du, L., Chen, W., Zhang, D. et al. The functions and mechanisms of piRNAs in mediating mammalian spermatogenesis and their applications in reproductive medicine. Cell. Mol. Life Sci. 81, 379 (2024). https://doi.org/10.1007/s00018-024-05399-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05399-6