Abstract
Plants are sessile organisms. This intriguing nature provokes the question of how they survive despite the continual perturbations caused by their constantly changing environment. The large amount of knowledge accumulated to date demonstrates the fascinating dynamic and plastic mechanisms, which underpin the diverse strategies selected in plants in response to the fluctuating environment. This phenotypic plasticity requires an efficient integration of external cues to their growth and developmental programs that can only be achieved through the dynamic and interactive coordination of various signaling networks. Given the versatility of intrinsic structural disorder within proteins, this feature appears as one of the leading characters of such complex functional circuits, critical for plant adaptation and survival in their wild habitats. In this review, we present information of those intrinsically disordered proteins (IDPs) from plants for which their high level of predicted structural disorder has been correlated with a particular function, or where there is experimental evidence linking this structural feature with its protein function. Using examples of plant IDPs involved in the control of cell cycle, metabolism, hormonal signaling and regulation of gene expression, development and responses to stress, we demonstrate the critical importance of IDPs throughout the life of the plant.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Throughout evolution plants have developed an extraordinary ability to overcome fluctuating and drastic environmental changes. Their sessile nature has imposed the selection of particular defense strategies allowing them efficient and effective adjustment or acclimation responses to these conditions, as well as skilled mechanisms to tolerate and survive them. The different endurance strategies selected in these organisms are the result of complex structural and interconnected regulatory networks, which have evolved in an intimate relationship with developmental programs. For instance, in many plant species, the reproductive stage waits for favorable climatic conditions to instrument a crucial set of processes for their perpetuation; root architecture modifies according to the availability of water, phosphorus and other nutrients; and orthodox seeds once desiccated can remain dormant for many years without significant loss in viability until they find sufficient water to germinate [1, 2]. This outstanding resourcefulness suggests mechanisms that make them capable of detecting diverse changes in the plant cell milieu, imposed by the external environment or by developmental programs.
Many molecular response mechanisms are efficiently adapted for rapid detection of subtle environmental fluctuations, as can be observed in mechano-sensitivity, ion channels, proton pumps and post-translational protein modifications. The modification of protein structure also seems to be an efficient and effective transducer of a great diversity of signals. This phenomenon is commonly associated with changes in protein conformation produced by phosphorylation or acetylation, or by interactions between proteins or other partners such as nucleic acids or other small molecules acting as substrates, cofactors, and allosteric regulators [3, 4]. However, an adaptation that has received less attention is that related to the intrinsic plasticity found in those proteins that have the ability to present different transient structures depending on the nature of their surroundings.
During the last decade, we have witnessed significant advances in the identification and characterization of many proteins showing intrinsic structural disorder. This has increased our knowledge of their functional relevance, structural properties and dynamics, as well as mechanisms of action (For review see [5,6,7]). Intrinsically disordered proteins (IDPs) are widely distributed in all domains of life. Although only a few complete proteomes from the different domains are currently available, various bioinformatic studies agree that Eukaryota proteomes show a higher average of disorder, compared to those of Bacteria, which in turn present higher disorder than those of Archaea. Interestingly, the predicted disorder in eukaryote proteomes spans a broad range of score values, with both very low and very high disorder [8, 9]. Overall, current information indicates that the level of disorder is higher in eukaryotic organisms than in prokaryotes. Even more important is the observation that protein superfamilies, which have undergone massive diversification during evolution present more structural disorder than other families. These data also correlate with the expansion of the number of cell types in an organism, revealing a positive relationship between proteome disorder and organism complexity [10, 11].
The accumulated knowledge on IDPs has revealed their functional versatility resulting from their peculiar properties. For example, IDPs can form ensembles with different structural conformations, allowing variability in the exposed surfaces [7, 12,13,14]. This structural plasticity confers to IDPs the ability to differentially exhibit different post-translational modification sites and/or recognition motifs, depending on specific conditions to interact transiently, but specifically with proteins or nucleic acids. With this in mind, it is not surprising the central roles that IDPs play in cellular functions, achieving regulatory and signaling roles as well as acting as scaffold or assembly proteins.
In this review, we present a general panorama of the available knowledge on protein disorder in plants. We have put together this information in the context of fundamental biological processes such as development, metabolism and stress responses, which in spite of the limited number of studies unveil the functional relevance of these proteins in the life of plants. The different IDPs referred to in this work are compiled in Tables 1 and 2.
IDPs distribution in plants
In recent years, the discovery and characterization of proteins with different amounts of structural disorder has revealed their high representation in plants [15,16,17,18]. Large-scale analysis of IDPs and intrinsically disordered regions (IDRs) in Arabidopsis thaliana, a widely used experimental model in plant biology, has shown that approximately 30% of its proteome is mostly disordered [10, 16], whereas Zea mays and Glycine max proteomes contain an even higher proportion of disorder (~50%) [19]. Interestingly, the chloroplast and mitochondrial proteomes show a significantly lower occurrence of disorder (between 2 and 19%) when compared to nuclear proteomes of different plant species. The abundance of disorder in these organellar proteomes is comparable to that of Archaea and bacteria, in accordance with the bacterial origin of the genes encoding their proteins [20]. The IDPs encoded in these organellar genomes are mostly involved in translation, transcription or RNA biosynthesis, and some are structural constituents of ribosomes, having in common the ability to form large complexes or to interact with numerous partners as expected from their intrinsic structural flexibility [5, 20]. It is interesting to note that for those proteins with paralogues of nuclear origin, both copies tend to show similarly low levels of disorder, suggesting again a common extra-nuclear origin or functional constraints [20]. Furthermore, the recent data obtained from the examination of the distribution of genes encoding IDPs in the genomes of A. thaliana and Oryza sativa indicate that they are not randomly arranged and that their organization may result from high recombination rates and chromosomal rearrangements. These observations are in accordance with the location of genes for proteins with highly disordered content within recombination hotspots and possessing high G + C content; this codon usage related to the over-representation of specific amino acid residues in IDPs (e.g., Arg, Gly, Ala and Pro) [19].
In silico analyses of the Arabidopsis proteome and of proteins from other plant species have found that IDPs are highly represented in functions related to cell cycle, nucleic acid metabolism, protein synthesis, hormone signaling and regulation of gene expression, development and responses to stress [16, 17, 19, 21,22,23]. This last functional category seems to be particularly associated with plant IDPs, including proteins involved in detection and signaling of external stimuli, chaperone activities and secondary metabolism; all essential functions for the phenotypic plasticity needed for plant adaptation and survival, as will be further discussed in this review. It should be noted that the examples we present in this review cannot be unequivocally classified and they may belong to several functional classes, which alludes to their functional promiscuity.
IDPs in plant development
The study of plant development and the characterization of the mechanisms involved have identified many proteins playing major control roles in this process. Further detailed analyses have revealed the presence of IDRs in some of these proteins. Germination and early seedling development [24], adventitious shoot formation [25], xylem development [26], photomorphogenesis [27], phytohormone signaling and response [28], flowering [29], and vegetative and reproductive growth [30] are some of the processes where IDR-containing proteins appear as key players. Interestingly, the structural plasticity arising from IDRs of several of these IDPs has been shown to be essential for proper function.
TCP (TB1-CYC-PCF1) transcription factors
The appropriate development and function of vegetative (leaves, shoot and roots) and reproductive (flowers) organs is orchestrated by several proteins, which are subjected to adaptable but precise spatio-temporal control, resulting in a timely fine-tuning of cell proliferation, expansion and differentiation [31]. Many of these proteins are transcription factors, some of which contain IDRs of significant length, that by interacting with other proteins and/or binding to DNA decode a specific signal in the activation or repression of gene expression. The TCP [from TEOSINTE BRANCHED1 (TB1), CYCLOIDEA (CYC) and PROLIFERATING CELL NUCLEAR ANTIGEN FACTOR1 (PCF1)] protein family consists of plant-specific transcription factors involved in plant shape developmental control. Bioinformatic analyses have shown that these transcription factors are IDPs [30, 32]. TCPs are classified as class I or class II according to the characteristics of their conserved and non-canonical basic helix-loop-helix (bHLH) DNA-binding domain [30, 32]. Class I TCP transcription factors participate in organ shape and growth, pollen development, germination, and inflorescence and flower development [33]. Class II TCPs, in addition to their redundant function in the regulation of lateral organ morphogenesis, also participate in endosperm, cotyledon, leaf, petal and stamen development, as well as other aspects of plant development and other processes [33]. Some of the functions assigned to TCP transcription in plant growth and development are a consequence of their involvement in the biosynthesis of some phytohormones, such as brassinosteroids and jasmonic acid, and other metabolites with biological activity such as flavonoids [21]. Analysis of the 24 Arabidopsis TCP protein sequences has shown a differential structural disorder content between the two TCP classes; with class I being more disordered than class II [30]. Biochemical analysis of TCP8, a class I TCP shows three IDRs of more than 50 residues in length containing a cluster of serine residues, at least one of which is phosphorylated [30]. In addition, the IDR located in the TCP8 C-terminal region corresponds to a trans-activation domain (TAD), which is required for the formation of high-order TCP8 homo-oligomers [30]. The identification of molecular recognition features (MoRFs) in the TCP8 TAD [30] and evidence of its requirement to bind TCP15 and PNM1, a pentatricopeptide repeat protein [34], are consistent with TCPs’ function as mediators of different stimuli or signals (Fig. 1a). Furthermore, they demonstrate the importance of IDRs as protein domains able to confer the ability to recognize various different partners, a feature needed for precise and flexible control.
Schematic representation of two examples of plant proteins containing IDRs that participate in developmental and metabolic processes. a TCP8 is a plant-specific transcription factor involved in plant shape developmental control. TCP8 contains three IDRs (represented by curved lines). In these IDRs, there are conserved serine residues, from which at least one is phosphorylated (fill blue small circle in the middle IDR). The IDR at the C-terminal region corresponds to a trans-activation domain (TAD) required for the formation of TCP8 homo-oligomers. This TAD is also required to bind different partners, such as TCP15 or PNM (red irregular oval). The IDR at the amino-terminal region (purple irregular line) is part of the TCP8 DNA-binding domain; this disordered region gains structure when TCP8 binds to DNA. b CP12 plays a key role in the regulation of the Calvin cycle by translating changes in light availability into the modulation of GAPDH and PRK enzyme activities. CP12 is a scaffold protein (represented by curved lines at the top of this panel) that forms a ternary complex with GAPDH (blue and red irregular ovals) and PRK (brown irregular oval) (GAPDH-CP12-PRK) (represented by the association of the three components at the bottom of the panel). During the formation of the GAPDH-CP12-PRK complex, GAPDH associates with CP12 by conformational selection. Upon this interaction, the CP12 N-terminal region remains in a fuzzy state, serving as a linker that facilitates the interaction with PRK. Once the complex is formed, it dimerizes to form a native complex in which there are two dimers of PRK, two tetramers of GAPDH and two monomers of CP12 (figure at the bottom right of this panel). Using this mechanism, it seems that CP12 is able to modulate GAPDH and PRK activities
NAC (NAM-ATA-CUC2) transcription factors
Another fundamental aspect of plant development is the maintenance of the shoot apical meristem (SAM). NAC (NAM/ATAF/CUC2) transcription factors constitute one of the largest families described in plants that, in addition to their involvement in other processes, control key aspects of SAM maintenance [35]. A conserved and folded DNA-binding domain defines these transcription factors; however, an additional feature of some NAC transcription factors is the presence of a variable and disordered TAD [24]. This characteristic has been experimentally confirmed for several NAC TADs [21, 36], such as ANAC019, involved in germination and early seedling development; HvNAC005 and HvNAC013, in senescence; NTL8, ANAC013, NAP, ANAC046 and SOG1 in germination and senescence; CUC1 in adventitious shoot formation and ANAC012 in xylem fiber development [24, 25, 37, 38]. It is known that TADs from HvNAC013 and ANAC046 interact with the RST (RCD1-SRO-TAF4) multi-binding domain of the hub protein RCD1 (RADICAL-INDUCED CELL DEATH1), a regulator of developmental, hormonal and stress responses [37]. Differing from the folding-upon-binding phenomenon, no structural rearrangement of the two disordered TADs occur upon binding to RCD1 [37], indicating that these ensembles might function as fuzzy complexes.
Elongated hypocotyl (HY5), bZIP transcription factor
Light is absolutely required for plant life. The presence or absence of light causes developmental reprogramming. The light-dependent modulation of plant development is known as photomorphogenesis. This developmental program leads to cotyledon expansion, hypocotyl shortening and chloroplast development [27]. HY5 (Elongated hypocotyl) is a bZIP transcription factor that positively regulates photomorphogenesis [39]. Disorder within the N-terminal region of HY5, responsible for the interaction with its negative regulator COP1, a multifunctional E3 ubiquitin ligase, has been demonstrated by various biophysical methods including limited proteolysis, mass spectrometry, circular dichroism (CD) and nuclear magnetic resonance (NMR) [27]. It is proposed that this disordered character might modulate the interaction with its partners, although functional characterization is still needed.
Cryptochromes (CRYs), blue light receptors
Plants are able to sense light quality (or wavelength) using different proteins such as phytochromes, phototropins and cryptochromes (CRY). Cryptochromes are blue light receptors that control developmental processes such as seedling de-etiolation, growth by elongation and initiation of flowering [40, 41]. CRYs consist of two domains: a conserved light-sensing N-photolyase-homologous region (PHR) of about 500 residues, and a C-terminal tail of variable sequence and length (CRY C-terminal Extension, CCE) [42, 43]. The CCE tail interacts with the PHR domain in a globular well-defined structure. Light activation of the Arabidopsis receptors CRY1 and CRY2 releases the CCE tail from the PHR, inducing the unfolding of the tail and allowing the interaction of both the PHR and the CCE with other proteins (e.g., COP1 and SPA1, a suppressor of phytochrome A1) to promote the blue light signal transduction pathway [44,45,46]. The light-induced disordered state of CRY receptors has been characterized by several biophysical methods such as limited proteolysis, CD, NMR and X-ray crystallography [47,48,49]. It is possible that plant CRYs use their disordered CCE region to efficiently recognize diverse binding partners through high-specificity/low-affinity interactions, potentially expanding the repertoire of plant signaling pathways coordinated by light [17].
HDC1 (histone deacetylase complex 1)
Regulation of chromatin accessibility is an important event of gene expression control, fundamental in developmental processes to fulfill the cell requirements within its organismal context. This process depends on the action of multiprotein complexes that control different modifications in DNA and histones [50]. One of these complexes is the histone deacetylase complex (HDAC), which in plants consists of histone deacetylases, co-repressors and histone-binding proteins [51]. HDC1 (HISTONE DEACETYLASE COMPLEX1) is a protein component of Arabidopsis HDAC containing a disordered N-terminal region [52, 53]. Interestingly, an HDC1 knockout mutant shows impaired leaf growth and delayed flowering, demonstrating its participation in plant development [52]. As expected for an IDP, HDC1 interacts with a wide variety of partners (HDA6, HDA19, SNL3, SNL2, SAP18, ING2 and MSI1) [53]. Deletion of the N-terminal disordered region considerably weakens HDC1 interaction with those proteins. This result together with evidence obtained from complementation experiments shows that the HDC N-terminal IDR plays a significant role in the coordination of flowering and petiole development [53].
BRI1 and BKI1, brassinosteroids signaling proteins
Brassinosteroids (BRs) are plant hormones that control a variety of growth and developmental processes, such as vascular differentiation, leaf development, stem elongation, flowering, senescence, stomatal development and male fertility [54,55,56]. BRs are perceived at the cell surface by BRI1 (BRASSINOSTEROID INSENSITIVE 1), a leucine-rich repeat receptor-like kinase (LRR-RLK) and its co-receptor BAK1 (BRI1-ASSOCIATED RECEPTOR KINASE 1) [57]. In the absence of BRs, the cytosolic kinase activity of BRI1 is maintained at low levels by auto-inhibition through its C terminus and by interacting with the repressor protein BKI1 (BRI1 KINASE INHIBITOR 1) [58]. When BRs bind to the extracellular domain of BRI1, the intracellular kinase domain is activated through auto- and trans-phosphorylation. BKI1 is then phosphorylated by BRI1 and released to the cytosol [59]. In contrast to animal LRR toll-like receptors, the extracellular region of the BR receptor contains a superhelix of twenty-five twisted LRRs; moreover, a ~70 amino acid ‘island’ domain has been localized between LRRs 21 and 22, which together constitute a hormone binding region. BR binding causes a conformational change in the BRI1 receptor that leads to its auto-phosphorylation. Remarkably, the ‘island’ domain connects to the LRR core through two long-disordered loops that become fully ordered upon binding to the steroid ligand. This makes the receptor competent to interact with other proteins, a conversion that may be necessary for receptor activation. It has been proposed that the BRI1 IDR may be an LRR receptor adaptation for efficient detection of small ligands [60]. Further participation of protein structural disorder is evident in this BR sensing protein ensemble, as the BKI1 C-terminal region presents high levels of disorder, particularly, at the BRI1 interacting motif (BIM). It is interesting to note that even though angiosperm BKI1 orthologues are highly diverse, the BIM IDR shows a high degree of conservation [61]. This, together with the finding that the absence of the IDR leads to increased BR sensitivity, establishes its relevance in BR signaling in plants [61].
Luminidependens (LD), a plant prion
The most diverse group of plants corresponds to the flowering plants (angiosperms). Flowering needs to be precisely controlled to generate flowers in an optimal time frame, where environmental conditions match with the presence of pollinators to promote fertilization and reproduction processes [62] Flowering often follows vernalization, a process achieved after a prolonged period of cold (winter), which ensures flowering in the spring [63]. Interestingly, Chakrabortee and collaborators found that a high proportion of proteins related to flowering in Arabidopsis are predicted to contain prion-like domains (PrDs) [29]. Some of these proteins are involved in transcription or regulation of RNA stability in the autonomous flowering pathway: Luminidependens (LD), Flowering Locus PA (FPA), Flowering Locus Y (FY) and Flowering Locus CA (FCA) [29]. Prions are proteins that retain the molecular memory of the cell because they are able to adopt different conformations and can be self-perpetuating [64]. PrDs are enriched in glutamine, asparagine, glycine, proline, serine and tyrosine and it has been shown that they are intrinsically disordered [65, 66]. LD is the first protein reported to have prion-like properties in plants, and can fully complement the activity of the Sup35 PrD, a well-characterized yeast prion [29]. As expected for a prion-like protein, LD protein shows a high level of structural disorder (64.6% according to PONDR, this work) [67], indicating that it is an IDP, even though this property has not been experimentally tested. Notably, LD orthologues from different plant species (Z. mays, O. sativa, Phaseolus vulgaris and Physcomitrella patens) also show a high percentage of disorder (51–66%, this work) [67]. As mentioned above, LD, along with a substantial percentage of Arabidopsis PrD-containing proteins, participates in flowering processes. This suggests that these proteins may play adaptive roles in the plant environmental memory required for fast responses to changing conditions, fine-tuning reproductive functions and consequently plant species preservation.
GRAS (GAI-RGA-SCR) transcription factors
The plant-specific GRAS [GIBBERELLIC ACID INSENSITIVE (GAI), REPRESSOR OF GAI (RGA), SCARECROW (SCR)] protein family is essential in diverse developmental processes, acting as integrators of signals from different plant growth regulatory inputs (for an extensive review refer to 68). GRAS proteins modulate gene expression through interaction with different transcription factors, thereby controlling their activities. Along with the conserved and folded GRAS domain, GRAS proteins are characterized by a disordered N-domain enriched in MoRFs [69]. Remarkably, the predicted MoRFs exclusively reside in the N-domain conserved motifs that define each subfamily, suggesting that structural disorder permits interactions with different proteins [17]. As has been established for other unstructured proteins, GRAS IDRs containing MoRFs experience disorder-to-order transitions when interacting with their ligands [17, 68,69,70]. GRAS proteins are classified in ten subfamilies. One of these subfamilies, composed of DELLA (Asp-Glu-Leu–Leu-Ala) proteins, is particularly important for hormonal regulation because DELLA proteins participate as negative regulators of gibberellic acid (GA)-induced plant growth. These are negatively regulated under increasing GA, as GA binds to its receptor (GID), prompting the interaction of the GID-GA complex with the disordered N-domain of DELLAs. This, in turn, promotes the degradation of the DELLA proteins through the ubiquitin–proteasome pathway, resulting in derepression of plant growth [71]. This interaction is mediated by the conserved DELLA and VHYNP motifs localized in an IDR that upon binding to the GID1/GA complex, experiences a disorder-to-order transition [70]. The participation of GRAS IDRs in this signaling pathway highlights their prevalence and function among hub network proteins, operating as integrators of environmental and developmental cues in plants.
MAP65-1, a microtubule associated protein
MAP65-1 is a microtubule (MT)-bundling protein implicated in central spindle formation and cytokinesis in animals, yeast and plants [72]. The Arabidopsis genome has nine genes encoding MAP65 proteins [73]. All these proteins have an N-terminal dimerization domain and an MT-binding domain. The MT-binding domain is localized at the second half of the MAP65-1 protein. The N-terminal region of this part of the MAP65-1 protein contains a conserved sequence responsible for MT binding, whereas its C-terminal region is more variable and predicted to be disordered [74, 75]. It was recently shown that Arabidopsis MAP65-1 is phosphorylated by Aurora α-kinases at two amino acid residues within its C-terminal disordered tail. The phosphorylation of these residues renders its detachment from MTs, leading to cell cycle progression, suggesting that the unfolded structure in MAP65-1 is required to modulate the accessibility of the two phosphorylatable residues to Aurora kinases, hence ensuring appropriate cell proliferation during plant development [75].
NRPE1, the largest subunit of Pol V
The RNA-directed DNA methylation (RdDM) pathway may act to repress the transcription of transposable elements to maintain genome integrity, mostly during critical plant development stages [76]. In A. thaliana, the canonical RdDM pathway is characterized by the participation of heterochromatic 24 nt small RNAs (hc-siRNAs) which are mainly produced by the interplay between RNA POLYMERASE IV (POLIV) and RNA-DEPENDENT RNA POLYMERASE 2 (RDR2). These enzymes generate a double stranded RNA that is subsequently trimmed into a 24 nt duplex by a type III ribonuclease, DICER-LIKE 3 (DCL3) [77, 78]. The generated hc-siRNAs are then methylated by HEN1 at the 3’ end of each strand [79] to be exported to the cytoplasm where one strand associates with the ARGONAUTE 4 (AGO4) complex [80]. The complex is then imported to the nucleus where hc-siRNA pairs may bind by base complementarity to a scaffold long non-coding RNA produced by RNA POLYMERASE V (POLV) [81]. The association of AGO4 in the silencing complex allows a physical interaction between this protein and POLV carboxy-terminal domain (CTD) via AGO hooks (described below) aided by the function of KTF1/SPT5L (Suppressor of Ty insertion 5—such as a homologue of SPT5 Pol II-associated elongation factor) [82]. This triggers the recruitment of a plethora of proteins which remove active chromatin marks and establish repressive ones, such as DNA methylation, DNA and histone modifications and chromatin remodeling features (reviewed extensively in [83]).
A peculiarity of the RdDM pathway in plants is the participation of two plant-exclusive RNA polymerases, POLIV and POLV. The catalytic domain of these polymerases is highly conserved, but their specific activities are conferred by their largest subunits; NRPD1 for POLIV, and NRPE1 for POLV [76, 84]. These subunits possess a characteristic carboxy-terminal domain which, in the case of NRPE1, contains a region rich in GW, WG and GWG amino acid residue arrangements, known as AGO hooks [84, 85]. This region constitutes an AGO-binding platform necessary for the interaction between NRPE1 and AGO4 and the consequent small RNA-directed DNA methylation [86]. Besides NRPE1, AGO hooks are also present in other AGO-binding proteins with up to 45 repeats. Along with their repetitive character, AGO-binding platforms have been predicted to be IDRs [87]. Interestingly, whereas the AGO-binding platform of NRPE1 orthologues is highly divergent in the primary sequence, the intrinsic disorder and the presence of AGO hooks are hallmarks of AGO-binding platforms across NRPE1 s. These characteristics are also extended to other AGO-binding proteins such as SPT5L, suggesting that this repetitive disordered structure is required to interact with a broad repertoire of targets, presumably regardless of sequence conservation [84]. Moreover, the evolutionary analyses reported by Trujillo et al. [84] suggest that this repetitive disordered array has been conserved to allow rapid sequence divergence while maintaining key functions in these proteins.
Protein disorder in plant metabolism
Large-scale computational approaches have found that IDP functions seem to be more common in signaling and regulation processes, whereas structural order is more frequent in proteins involved in catalysis, in binding of small ligands and in membrane proteins (channels or transporters) [88]. However, this dichotomy contrasts with the description of some enzymes containing IDRs in loops or tails, which participate in the modification of protein conformation upon substrate binding, and thus expose catalytic residues and contribute to catalysis [89,90,91]. Furthermore, one must consider the role of some IDRs as sites for post-translational modifications, acting as switches of activation/inactivation or as modulators of their own activity. Many of these IDR-containing proteins are involved in the fundamental housekeeping of the plant.
In this section, we will describe those IDPs known to participate in different aspects of plant metabolism; some of them involved in photosynthesis, in metal binding or in antioxidant mechanisms.
Chloroplast protein 12 (CP12)
Few studies have investigated the role of protein structural disorder in the plant photosynthetic machinery. However, with the advancement of the characterization of proteins implicated in this process, more data are emerging showing the impact of intrinsic disorder in this essential plant function. An example of this is the chloroplast protein 12 (CP12), a well-characterized scaffold protein that forms a ternary complex with glyceraldehyde-3-phosphate dehydrogenase (GAPDH) and phosphoribulokinase (PRK), named the GAPDH-CP12-PRK complex. CP12, present in most photosynthetic organisms, also regulates GAPDH and PRK activities [92, 93]. CP12 is a small protein (8.5 kDa) encoded in the nuclear genome and translocated to chloroplasts; although it contains cysteine residues, it has been shown to have all the properties of an IDP. Because its degree of disorder is higher in vascular plant orthologues than in eukaryotic algae, it has been proposed that CP12 has evolved to become more flexible, which correlates with its increased multifunctionality [94, 95]. In the plant kingdom, CP12 proteins share common features; however, their N termini, in addition to being highly disordered, show high sequence variability [95, 96]. During the formation of the GAPDH-CP12 or PRK-CP12 binary complexes, CP12 structural disorder remains, in particular in its N-terminal region, indicating that these are fuzzy complexes. These observations have suggested that the fuzziness of this association could facilitate the binding of either GAPDH or PRK [97, 98]. The integration of the different lines of evidence suggests a model for the formation of the GAPDH-CP12-PRK complex, where GAPDH associates with CP12 by conformational selection; first recognizing specific conformation(s) in CP12 to establish the binding. Upon this interaction event, the CP12N-terminal remains in a fuzzy state acting as a linker to facilitate the association with PRK. Once the GAPDH-CP12-PRK complex is formed, it dimerizes to form the native complex, composed finally of two dimers of PRKs, two tetramers of GAPDH, and probably, two monomers of CP12 [93, 96, 97] (Fig. 1b).
CP12 plays a key role in the regulation of the Calvin cycle, transducing changes in light availability such as those occurring during the day–night transition. This event leads to the generation of a hyperoxidant state, which is detected by the two cysteine residues in the CP12 C terminus forming a disulfide bridge. This leads to a conformational change in CP12, resulting in its N-terminal region folding into α-helix [96], which subsequently prevents the entrance of the NADPH cofactor in the GAPDH catalytic site. In the night-to-day transition, the conformation is reversed; the disulfide bridge is reduced by thioredoxin permitting NADPH entry and resulting in GAPDH activation. This inhibiting effect exerted by the CP12 also occurs on the PRK enzyme, as part of the complex. Interestingly, accumulating evidence indicates that CP12 assembles in larger supramolecular complexes, as happens in Chlamydomonas reinhardtii, where the GAPDH-CP12-PRK complex associates with aldolase [92], thus suggesting additional roles in other metabolic processes [23]. From the differential lines of evidence, it can be concluded that CP12, as with some other IDPs, has a moonlighting activity, being able to act as a scaffold for GAPDH and PRK [93], as a regulator of these enzyme activities, and as a protective shield against oxidative damage [23, 99, 100].
Glyceraldehyde-3-phosphate dehydrogenase (GAPDH)
GAPDH plays a central role in glycolysis and gluconeogenesis. In vascular plants, GAPDH can exist as heterotetramers of two GapA and two GapB (A2B2) subunits, as homotetramers of four GapA subunits (A4) or as hexadecamer of eight GapA and eight GapB subunits (A8B8). Interestingly, the GapB subunit also contains a C terminus highly similar to the CP12 C-terminal IDR [101]. The presence of two cysteine residues in this region permits photosynthetic NADPH-dependent GAPDH containing the GapB subunits to detect redox changes. Oxidative conditions induce the formation of a disulfide bridge in its CP12-like C terminus, promoting the NAD-dependent arrangement of higher homo-oligomers that result in auto-inhibition of its NADPH-dependent catalytic activity. This conformational change and complex formation is needed for the reduction of 1,3-bisphosphoglycerate to produce glyceraldehyde-3-phosphate [101,102,103]. This intrinsically disordered feature of GapB confers on A2B2 GADPH a CP12-autonomous regulation by the redox status of the cell.
Ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) activase
Rubisco (Ribulose-1,5-bisphosphate carboxylase/oxygenase), the most abundant protein on Earth [104, 105], is an enzyme responsible for fixing atmospheric CO2 into RuBP (ribulose 1,5-bisphosphate) to produce two phosphoglycerate molecules. The activity of this enzyme depends on the binding of Mg2+ ions and the carbamylation of a lysine residue located in its active site; however, the binding of RuBP can reduce the efficiency of carbamylation, and consequently the activation of the enzyme [106]. Nature has solved this limitation through proteins known as Rubisco activases that because of their ATPase and chaperone activity, allow Rubisco carbamylation by removing RuBP from the active site and giving access to CO2 molecules. Photosynthetic organisms present two Rubisco activase isoforms (α and β) [107] containing a C-terminal extension (20–50 amino acid residues) which is predicted as an intrinsically disordered region [23]. As is the case for CP12, this IDR contains two highly conserved cysteine residues in α isoforms [108, 109], responsible for the light regulation of Rubisco activase. This control is achieved by the action of thioredoxin f on the two cysteine residues, such that upon oxidation, inhibition of α isoform activity by light is abolished [110, 111]. The Rubisco activase function can be recovered by the reduction of the C-terminal disulfide bridge by thioredoxin f, depending on the redox status of the chloroplasts [112]. Interestingly, in spite of the functional or structural differences among Rubisco activases in diverse photosynthetic organisms, their C-terminal IDRs have been conserved; for example, in the case of cyanobacteria, they are involved in carboxysome targeting [23, 113]. Overall, intrinsic disorder in Rubisco activase strongly suggests that it is a conserved feature responsible for its functional versatility as an ATPase, a chaperone and as a fine-tuning regulator that has contributed to the broad adaptability of the photosynthetic process.
Manganese stabilizing protein (MSP)
Plants capture sunlight through the light-harvesting complex (LHC) or antenna complex as part of Photosystem II (PSII). This complex of proteins and pigments is embedded in thylakoid membranes and connects the antenna to the chlorophylls in the reaction center. The photons captured by PSII initiate a chain of redox states through electron transfer reactions needed for the oxidation of two water molecules to O2. This photolysis reaction takes place in the oxygen-evolving complex (OEC), one of the PSII subunits. The different polypeptides of PSII are needed for an efficient O2 evolution; in particular, three extrinsic proteins of 17, 23, and 33 kDa, which are located on the luminal side of PSII. This last protein, also termed manganese stabilizing protein (MSP), is required to maintain stability and an efficient cycling of the four oxidizing manganese atoms [114,115,116,117]. MSP lacks a compact structure and is composed of 55% turns and random coils. These properties, together with its amino acid composition and other features, establish its intrinsic structural disorder. In vitro experiments suggest that the structural flexibility of this protein is required for its function, possibly by facilitating effective protein–protein interactions as an integral member of PSII [118]. Moreover, it has been shown that conserved charged amino acid residues in MSP are important for the retention of Cl− ions, to maintain their concentration at the levels needed for the effective redox reactions of the manganese cluster [119]. Again, MSP exemplifies the participation of protein structural disorder as an essential attribute to achieve precise and opportune roles in a complex system able to adjust to the changing environment.
Alb3, a thylakoid membrane protein
The membrane invertase protein Alb3 controls the insertion, folding and assembly of a diverse group of proteins into the thylakoid membrane of chloroplasts. Alb3 interacts with chloroplast signal recognition particles (cpSRP) in the thylakoid membrane through its C-terminal intrinsically disordered region. This IDR has two conserved positively charged motifs needed for the association with cpSRPs that follows a coupled binding and folding mechanism [120]. Once the Alb3-cpSRP complex is formed, it participates in the post-translational insertion of the light-harvesting chlorophyll a/b-binding protein (LHCP), a highly abundant protein in thylakoid membranes. The insertion of LHCP into these membranes strictly requires the involvement of cpSRP and Alb3. Alb3 is also needed for the targeting and insertion of cytochrome b 6 into the thylakoid membrane [121]. Cytochrome b 6 is a largely disordered protein in aqueous solution, but by interaction with lipids from the membrane it folds into an α-helical structure just before its membrane insertion [122]. An additional function assigned to the Alb3 C-terminal IDR is the light-dependent modulation of Alb3 stability [123].
Polyphenol oxidases (PPOs)
Tyrosinases and catecholases from plants and fungi are generally named polyphenol oxidases (PPOs). In plants, PPOs mediate the production of melanin, responsible for the brown color in fruits when they suffer damage. They are nuclear-encoded and are transported to the chloroplast thylakoid lumen, where they can be in a soluble form or in a weak association to the thylakoid membranes. They are activated by the proteolytic cleavage of their C-terminal region. Using bioinformatic approaches to analyze multiple plant PPO sequences, it was found that the region between the N-terminal and C-terminal corresponds to a disordered linker essential to establish those conditions in which the PPOs are processed and may be activated [124]. This prediction suggests that the PPO IDR may acquire certain levels of order depending on the environment. Although experimental data are needed, the presence of a conserved phosphorylation site within this IDR suggests auto-regulation of PPO activities and/or that PPOs have roles as signaling molecules [124].
Phosphoenolpyruvate carboxylase (PEPC)
Carbon assimilation is not only accomplished by the activity of Rubisco, but also by PEPC (Phosphoenolpyruvate carboxylase), a ubiquitous enzyme in plants. PEPC also plays a critical role in plants with C4 photosynthesis and crassulacean acid metabolism (CAM), by producing oxaloacetate from HCO3 [125]. Two types of PEPC enzymes have been described in plants, known as plant-type PEPC (PTPC) and a distantly related bacterial-type PEPC (BTPC). The BTPC enzymes show low sequence identity with PTPCs, they lack the typical serine phosphorylation motif located in the PTPC N-terminal region and they are encoded in all plant genomes sequenced to date. Of particular interest is the fact that BTPCs contain an insertion of approximately 142 amino acid residues predicted as a structurally disordered region. This IDR seems to be highly divergent and a distinctive characteristic of BTPCs. PEPC enzymes are organized as oligomers, from which two classes have been identified: class-1 oligomers consisting of PTPC homotetramers, and class-2 complexes corresponding to heterotetramers composed of three PTPC and one BTPC subunits [125]. Recently, it was demonstrated that the BTPC IDR from the castor oil plant (Ricinus communis) is needed for its association with the PTPC subunit in a class-2 PEPC complex. Furthermore, even though the N-terminal region conserved in PTPCs is not conserved in BTPCs, it was thought that these enzymes were non-phosphorylatable. However, it has been shown that RcBTPC is phosphorylated in vivo at least at two serine residues. One of these modifications occurs at serine-451, a highly conserved target residue located within the IDR of these proteins. This event exerts a regulatory role, causing the inhibition of the catalytic activity of the enzyme within the class-2 PEPC complex [126].
UreG G protein
GTP-binding proteins (G proteins) are GTPases that catalyze the hydrolysis of GTP to yield GDP and inorganic phosphate. The structure of the catalytic domain of this enzyme is usually a β-sheet delimited by flexible regions of α-helices and loops. The binding of GTP or GDP activates (in the case of GTP) or inactivates (in the case of GDP) these GTPases, associations stabilized by the binding of specific protein regulators that promote conformational modifications. UreG is a bacterial-type G protein involved in urease maturation that has been demonstrated to belong to the class of intrinsically disordered enzymes. The structural disorder in UreG is mostly concentrated in a region of ~50 residues localized in the center of its protein sequence, which seems to influence the structure of the GTP-binding pocket [127,128,129]. In plants, one gene shows sequence similarity with bacterial UreG GTPase, which functions as an urease accessory protein, promoting optimal urease activation by allowing nickel or zinc incorporation in its active site and the GTP-dependent CO2 transfer required for lysine carbamylation. This protein has been characterized in soybean (G. max), where it has shown a differential binding affinity to Ni2+ and Zn2+. Furthermore, it has the highest affinity for Zn2+ described to date for any UreG protein. This observation suggests a role for UreG as a Zn2+ accumulator protein that may modulate the available levels of this metal in the cell. Analysis of its quaternary structure indicates that UreG is monomeric in solution and that dimers can be formed and stabilized upon Zn2+ binding, due to conformational rearrangements in the protein. The association with Zn2+ decreases the levels of secondary structure, but perhaps stabilizes the subsequent dimerization by facilitating the folding of the active site domain. However, this binding alone is not enough to yield a high UreG activity, suggesting that additional factors are needed to achieve its optimal GTPase activity [130]. UreG further illustrates the functional versatility conferred by intrinsic disorder to proteins with catalytic and regulatory roles in plant metabolism.
Jaburetox, an intrinsically disordered insecticidal polypeptide
Ureases are nickel-dependent metallo-enzymes that catalyze the hydrolysis of urea into ammonia and CO2 [131]. It was discovered that canatoxin, considered an isoform of a jack bean urease (from seeds of Canavalia ensiformis), corresponds to a 10-kDa peptide (JBU) produced from urease hydrolysis by cathepsin-like enzymes. This JBU peptide is toxic to mammals, fungi and insects. One of the major urease isoforms from jack bean seeds shows toxicity to hemiptera insects independent of its ureolytic activity, and instead its effect is produced by the action of digestive enzymes present in the insect gut [132, 133]. This entomotoxic activity is caused by an internal peptide (pepcanatox) product of this hydrolysis. Jaburetox, a recombinant version of the in vivo generated peptide, is derived from the N-terminal sequence of the C. ensiformis urease isoform and possesses a potent insecticidal effect on crop pests [133, 134]. A motif present on this peptide is also found in pore-forming and neurotoxic peptides which present membrane-disturbing activities [135]. A large hydrodynamic radius, together with light scattering, CD and NMR spectroscopic data, shows that Jaburetox is a monomeric disordered peptide with an α-helix motif by its N terminus and two turn-like structures in the central region and by the C terminus of the peptide. It is suggested that the Jaburetox IDP might act as a membrane protein or as a scaffold protein, but evidence for this is still lacking, therefore a comprehensive view of its insecticidal activity remains elusive [136].
Protein structural disorder in plant abiotic stress responses
Prediction of structural intrinsic disorder from plant proteomes reveals a noteworthy participation of IDPs in plant responses to their environment and to stress conditions. However, not many abiotic stress response proteins have been confirmed as IDPs and there is limited information about their function. Here, we compile those stress-responsive IDPs for which there is evidence of function and structural organization.
Late embryogenesis abundant (LEA) proteins
Late embryogenesis abundant (LEA) proteins belong to an emblematic group of IDPs distinctively involved in plant stress responses, notably, in adverse conditions of low water availability. LEA proteins can be classified into seven groups or families based on amino acid sequence similarity, although nomenclature can vary. In this review, we will follow that proposed by Battaglia et al. [137], who also report the presence of distinctive motifs for each family, some of which correspond to MoRFs [138, 139]. LEA proteins do not show significant sequence similarity with any other proteins of known function, making their characterization a challenging task. LEA proteins are considered ubiquitous in the Viridiplantae kingdom because they have been found in angiosperms, gymnosperms, non-vascular plants and algae [137, 140]. Although for some time they were considered exclusive to plants, interestingly they have also been detected in other organisms including insects, nematodes, crustaceans, rotifers and bacteria [141,142,143,144,145]. In all cases, their abundance is related to water deficit, but some also respond to other stress conditions. In general, LEA proteins are highly hydrophilic with a high content of glycine residues or other small amino acids, and they are usually deficient in tryptophan and cysteine residues; all characteristics of IDPs [137, 146, 147]. These properties are conserved in a wider group of water deficit response proteins, the ‘hydrophilins’, which are conserved across all domains of life [148]. As is documented for other IDPs, LEA proteins possess key qualities that enable them to perform more than one function; this ‘moonlighting’ characteristic will be described below. As in the case of IDPs involved in development and metabolism, the plasticity and molecular flexibility of LEA proteins appear to be central to their function (Fig. 2A).
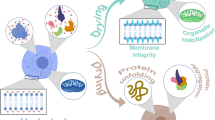
Schematic representation of two examples of plant IDPs that participate in abiotic and biotic stress responses. A LEA proteins (represented as purple curved lines) belong to a representative group of plant IDPs involved in plant abiotic stress responses. LEA proteins are able to prevent the inactivation of reporter enzymes under in vitro partial dehydration and freeze–thaw treatments. One action mechanism supported by different lines of evidence indicates that LEA proteins function as chaperones during water deficit a by interacting with their protein target(s) (green irregular ovals) and avoiding the damage (denaturation represented by green irregular lines emerging from the green ovals) caused by the effects of low water availability in the cell. The possibility that LEA proteins may bind and recognize their targets by conformational selection under water deficit has been suggested by in vitro data. In addition, there is evidence indicating that LEA proteins are able to stabilize membrane (double blue circles) integrity b during water deficit, by interaction through the amphipathic regions present in some LEA proteins. It has been suggested that LEA proteins might achieve more stable conformations upon membrane association. It has been proposed that this interaction induces LEA protein folding. b An additional attribute of at least some LEA proteins is their ability to bind metal ions (Fe3+, Ni2+, Cu2+, Co2+ and Zn2+) (small gray fill circles), which in some cases, by these means scavenge reactive oxygen species (c). For some LEA proteins, metal binding promotes a reduction in the content of structural disorder; however, this is not a common observation. In this panel, continue arrows represent the protective effect of LEA proteins, whereas discontinuous arrows indicate the consequent damage produced by stress in the absence of these proteins. B Biotic stress produced by plant pathogens has led to the selection of refined mechanisms to detect their presence and to mount complex inducible responses to efficiently counteract their attack. The participation of IDPs along the different steps of pathogen invasion, from their perception to the plant defense response has been documented. The RbohD protein (green curved lines), which belongs to the NADPH oxidase family, represents an example of this. This protein, partially integrated in the membrane, is responsible for the early generation of ROS, upstream of calcium and phosphorylation signaling. The RbohD cytoplasmic N terminus possesses an IDR, which contains EF-hand motifs involved in calcium binding. The malleable nature of this region results in extended conformational changes induced by the synergistic effect of calcium binding and its phosphorylation, which in turn modulates the interaction with small GTPase proteins (orange irregular oval); a process needed to set up the plant protection response against pathogens
One of the most general functions across the LEA group is an ability to protect the integrity of other enzymes. This has been demonstrated using several non-plant reporter enzymes with in vitro partial dehydration and freeze–thaw treatments, whereby the presence of LEAs prevents inactivation, denaturation and consequent aggregation of enzymes such as lactate (LDH) and malate dehydrogenases (MDH), citrate synthase (CS), β-glucosidase G (βglG), and glucose oxidase/peroxidase (GOD/POD) [137, 146, 149,150,151,152,153,154,155]. In the case of group 3 LEA proteins from Pisum sativum (PsLEAm), this protective effect has been demonstrated on plant proteins such as mitochondrial rhodanase and fumarase [156]. The protective activities resemble that of small heat shock proteins (sHSPs), which circumvent protein aggregation upon heat shock treatments in the absence of ATP [157]. Hence, it appears that LEA proteins may function as chaperones during water deficit stress. These observations suggest a protective role specifically against protein damage caused when water limitation inhibits cellular functions. Furthermore, it appears that this unique function cannot be provided by other types of chaperones [149, 158] (Cuevas-Velazquez et al. unpublished).
The different lines of evidence from in vitro enzyme assays have led to two main hypotheses to explain the LEA protein protecting activity. Because high concentrations of LEA proteins are able to prevent inactivation and aggregation of other proteins, it has been proposed that they may act as ‘molecular shields’. Given their large hydrodynamic radius in aqueous solution, they may create a protein molecular net, thereby promoting the alignment of their hydrophilic amino acid residues around the surface of a target protein, and in this way, prevent the loss of its bulk water and consequent changes in its native structure [159, 160]. However, there is also evidence showing that small amounts of LEA proteins (down to 1:1–1:5 ratios of LEA:reporter enzyme) are also capable of protecting target proteins to a similar degree [149, 161,162,163,164,165,166]. This indicates that LEA proteins may function in a ‘chaperone-like’ mode, where interaction is required to select and protect their targets, binding as monomers or oligomers [149, 161, 167, 168] (Fig. 2A(a)). This hypothesis is supported by evidence indicating that these disordered proteins can fold in α-helix under high osmolarity or high macromolecular crowding, prevalent conditions under water deficit, which would lead to a decrease in their hydrodynamic radius [138, 139, 147, 169]. Crucially, this property seems to be associated with their chaperone-like activity [139]. With this in mind, and considering the role of conformational plasticity [138, 139, 147, 169,170,171], it is possible that LEA proteins may bind and recognize their targets following a mechanism that resembles conformational selection under water deficit, the natural conditions under which they accumulate in the cell. Although similar lines of evidence have been obtained for LEA proteins from different groups (LEA2, LEA3, LEA4, LEA7), further experimentation is needed to support these alternative mechanisms. In particular, it is imperative to establish strategies to obtain in vivo data that could help provide a more comprehensive view of their action mechanisms.
The intrinsic disorder of LEA proteins may also function in the stabilization of membrane integrity under stress (Fig. 2A(b)). Some LEA proteins, mainly those from groups 2–4, are able to bind in vitro to lipid vesicles [172,173,174,175,176]. In some cases, these vesicles have been produced using phospholipids and galactolipids common to plant chloroplast and mitochondrial envelops [169, 177], or they have been obtained from thylakoid membrane fractions of spinach leaf tissue [178]. Interestingly, α-helix folding upon vesicle binding has also been shown for some LEA proteins [179, 180]. For group 2 LEA proteins (dehydrins), it has been found that the K-segment, a distinctive motif of this family is necessary for liposome binding, which is consistent with its amphipathic nature [178, 180, 181] (Fig. 2A(b)). Distinctive motifs of LEA3 and LEA4 proteins also present amphipathic properties, which help explain their ability to bind to lipid vesicles surfaces [137, 182, 183].
Unfortunately, to date there remains no evidence of any of these activities in vivo. Despite this, in vitro functions correlate with the accumulation of LEA proteins in plant tissues under low water potentials induced by dehydration or by cold or freezing temperatures; conditions in which enzymes can be inactivated and membrane injuries are produced. Interestingly, some Arabidopsis LEA proteins are required for plant optimal adjustment to cold, water deficit and/or salinity, as can be inferred from the phenotypes produced by mutants lacking genes encoding LEA proteins from group 1 [184], group 2 [185], group 3 [169] and group 4 [186]. Additionally, the acquisition of tolerance to water limitation or low temperatures by the overexpression of several LEA proteins (from groups 2–4 and 7) in different plant species strongly supports their role as protector molecules under these stress conditions [187,188,189,190,191,192,193,194,195,196,197].
As can be seen with IDPs involved in plant development and metabolism, dynamic structural order can be attained by interaction with metal ions. LEA proteins have also been shown to bind metal ions (Fe3+, Ni2+, Cu2+, Co2+ and Zn2+) and scavenge reactive oxygen species (reviewed in [198, 199]). LEA proteins showing high affinity to these metals include LEA2 or dehydrins, LEA3 and LEA4 [191, 198,199,200]. Acid dehydrins (RAB17 and VCaB45) are able to bind calcium, possibly to modulate intracellular calcium levels, thereby acting as ionic buffers during water deficit: a hypothesis that still needs to be tested [201, 202]. The metal binding properties in these proteins have been attributed to the abundance of histidine residues or to the presence of metal binding motifs (HX3H or HH) [203]. Importantly, for some LEA proteins, it is known that metal binding can promote the gain of an ordered conformation [204].
Remarkably, a group 2 LEA protein (ITP, iron transport protein) has been shown to carry iron in vivo and bind iron in vitro (Fig. 2A(c)). This protein was found associated with iron in phloem exudate from R. communis L. [205]. ITP also binds Ni2+, Cu2+, Zn2+ and Mn2+ in vitro, preferentially binding to Fe3+ but not to Fe2+. This indicates that this LEA protein may function as a phloem micronutrient transport protein [205], opening up the possibility that this novel function may exist for other LEA proteins or IDPs able to bind iron or other micronutrients.
For some group 2 proteins, it has been shown that their phosphorylation is required for the metal association to occur [202, 206, 207]. Group 2 and group 4 proteins can also circumvent the production of reactive oxygen species (ROS), given their capacity to bind metals able to promote ROS generation (Fig. 2A(c)). Evidence for this activity has been obtained in vitro and in vivo [200, 204, 208]. This mechanism could be advantageous under abiotic stress conditions such as water deficit, when ROS production and sensitivity to secondary stresses are exacerbated.
The multifunctionality of LEA proteins and the role of metal ions are reinforced by data indicating that group 2 and group 7 LEA proteins can also bind nucleic acids. Group 2 LEA proteins (CuCOR15, VvDHN1a and WCI16) have been shown to associate with DNA and RNA. In the case of CuCOR15 and WCI16, this occurs in the presence of physiological concentrations of Zn2+ [209,210,211]. This evidence suggests that nucleic acids need similar protection from the effects of water limitation.
DNA binding has also been demonstrated for a group 7 LEA protein (ASR1, ABA [Abscisic acid] stress ripening 1), a widely occurring plant LEA protein that does not exist in A. thaliana. Strikingly, in addition to its protein protective role, ASR1 can also function as a transcription factor. It has been shown that ASR1 is able to bind to the regulatory regions of genes related to cell wall synthesis and remodeling, as well as genes encoding membrane channels implicated in water and solute trafficking [212]. Grape ASR1, VvMSA, recognizes specific sites in the regulatory region of the hexose transport 1 (Ht1) gene [213], and ASR orthologues are also involved in sugar and amino acid accumulation in species such as maize and potato [214, 215].
Phosphorylation of IDRs in some LEA groups may also play a role in LEA protein function. Members of group 2, 4, 6 contain phosphorylatable motifs and in vivo and in vitro phosphorylation has been verified for group 2 LEA proteins (dehydrins/DHNs) in Arabidopsis, wheat, maize, and other plants [201, 207, 216,217,218,219,220,221,222]. Although the role of this modification is not well understood, for group 2 LEA proteins it may be needed to modulate membrane interaction and lipid phase transition [178, 180], as well as nuclear localization [223, 224]. However, although phosphorylation pattern correlates with tolerance to water limitation, it is unknown whether this modification is required to modulate LEA protein protective activity and/or target selectivity by allowing the display of different binding motifs and/or MoRFs.
The multifunctionality found in vitro for the different LEA proteins is often compatible with their in vivo intracellular localization, suggesting that there may be both subcellular specialization as well as redundancy. LEA proteins from all groups have been localized to cytosol, nucleus, mitochondria and chloroplast [137, 199, 225]. However, at least in the case of group 3 LEA proteins, the most diverse LEA family, not all its members show the same localization. Some are found in cytosol or nucleus, others in the chloroplast and some others only in mitochondria [226]. This implies a requirement and possible functional specificity of LEA proteins during the plant stress response.
Further evidence of their deep functional divergence, as well as their ubiquitousness, can be seen in the high conservation of most LEA families throughout the Plantae kingdom’s evolution. LEA proteins from group 1–4 can be detected in genomes from the most recent angiosperms through to the bryophytes, including the liverwort Marchantia, the most basal plant model described to date. Group 6 and 7 LEA proteins have been found only in seed plants, and in the case of group 7 LEA proteins, do not seem to be present in all phyla [137, 155, 186, 227, 228]. The broad distribution and conservation of these plant IDPs throughout evolution illustrate not only the relevance of these proteins for the organisms in this kingdom, but also the importance of disorder for the various functions they achieve.
The ubiquity of LEA proteins across land plants is testament to their versatility. Even though LEA protein action mechanisms remain elusive, the intrinsically disordered nature of these proteins matches their apparent ‘moonlighting’ character, as exhibited by diverse data where the same LEA protein is able to protect proteins and membranes, and bind metals and/or nucleic acids (see Table 2). These characteristics are compatible with their ability to utilize the same or overlapping regions to exert distinct effects and to switch functions by adopting different conformations upon binding [229].
Small heat shock proteins (sHSPs)
Small heat shock proteins (sHSPs) are ubiquitous molecular chaperones, which play important roles in protein homeostasis and in plant responses to stress. sHSPs are classified in 11 subfamilies, six localized to cytoplasm/nucleus and five to organelles. These chaperones bind diverse partially unfolded polypeptides maintaining their refolding capacity until they can return to their native structure with the help of other chaperone proteins, such as HSP70. In this way, sHSPs protect cells from the loss of essential proteins and from the penalties caused by protein aggregation. Commonly these proteins respond to high temperatures, but also to other stress conditions, and some may also be produced even under optimal growth conditions. In contrast to other molecular chaperones, sHSPs form large and dynamic oligomers with different stoichiometry. All sHSPs contain a core α-crystallin domain bordered by a short C-terminal region and an N-terminal extension of variable length and sequence (for review see [157, 230]). Both regions participate in the recognition of—and binding to—clients and in the formation of their oligomers (for review see [230, 231]). It has been proposed that during heat stress the oligomeric sHSPs undergo conformational rearrangements leading to their dissociation. These structural changes enable the interaction of these chaperones with hydrophobic patches in the partially denatured clients, subsequently forming large soluble complexes, protecting protein clients form further damage. Biochemical and biophysical evidence indicate that the intrinsically disordered N-terminal arm is able to present different interaction sites revealing a mechanism to efficiently protect the integrity of many different substrates in the cell [157, 232]. Although many questions still remain unanswered regarding mechanistic details and in vivo evidence is required, sHSPs offer a view of the need for structural plasticity and promiscuity to maintain cell functions during stress.
Glycine-rich RNA-binding proteins (GR-RBPs)
Although LEA proteins are important to the plant cold stress response, other IDPs are known to play a protective role. Plants exposed to low temperatures experience a slowing down or even a pause of their metabolic processes and this may result, directly or indirectly, in damage to macromolecules and cellular structures [233]. Among the proteins synthesized to overcome the impairment that cold and other abiotic and biotic stresses cause to macromolecules are the so-called glycine-rich RNA-binding proteins (GR-RBPs) [234, 235]. Some of the functions characterized for GR-RBPs are the facilitation of mRNA transport and participation in splicing and translation: roles mediated by their RNA chaperone activity [236, 237]. GR-RBPs contain an RNA recognition motif (RRM) in the N-terminal region and a disordered glycine-rich region (GR) at their C-terminal end, and they can be classified into eight groups, each one with apparently different roles [238, 239]. In Arabidopsis, AtGR-RBP7, in addition to being a circadian regulator and promoter of flowering and mRNA splicing, accumulates in response to cold stress [236, 237, 240,241,242,243,244]. Deletion of AtGR-RBP7 leads to low-temperature sensitive phenotypes, highlighting its role in optimal plant adjustment to cold stress [245]. NMR analysis confirms the structural disorder of the GR domain for NtGR-RBP1, an AtGR-RBP7 orthologue from Nicotiana tabacum [246]. As expected, NtGR-RBP1 is shown to bind RNA and single stranded DNA through the RRM. Furthermore, the NtGR-RBP1 GR interacts transiently with its RRM domain, promoting self-association to effectively increase its local concentration, and hence its affinity for nucleic acids. These findings suggest a mechanism for the unfolding of non-native structures in RNA by NtGR-RBP1, which may be involved in enhancing its RNA chaperone activity [246].
Vesicle-inducing protein in plastid 1 (VIPP1)
The integrity of thylakoid membranes is crucially important for photosynthesis and chloroplast functions. Multiple reports have shown the participation of a protein called VIPP1 (Vesicle inducing protein in plastid 1) in thylakoid membrane biogenesis and thylakoid membrane maintenance during drought, heat and osmotic stress [247, 248], not only in cyanobacteria and green algae, but also in vascular plants [249,250,251]. The evolutionary emergence of this protein seems to be specific to oxygenic photosynthetic organisms [251]. Recent evidence suggests that while VIPP1 may have multiple roles in plastids, it strongly protects the chloroplast envelope [252]. The N-terminal region of VIPP1 presents high sequence similarity to its bacterial orthologue PspA (Phage shock protein A) [253], which plays a central role in the well-characterized bacterial system Psp, involved in the protection of membrane integrity under various stresses [254]. During membrane damage, PspA and VIPP1 bind to membranes forming high-order oligomeric effector complexes able to repair the inner membrane and conserve its integrity [255, 256]. Interestingly, this occurs despite the absence of transmembrane domains in these proteins [252, 253]. CD spectroscopy studies show that PspA and VIPP1 N-terminal peptides are disordered in solution and fold upon membrane association, as occurs in a typical membrane amphipathic helix [257, 258]. The membrane binding of these proteins depends on differences in stored curvature elastic stress, a feature of damaged membranes [259]. These observations suggest that the folding transition associated with PspA and VIPP N-terminal membrane binding might act as a stress-sensing mechanism controlling the effector function of these proteins.
During the evolution of photosynthetic organisms, the PspA orthologue VIPP1 has acquired an additional C-terminal tail (Vc) that also presents the characteristics of an intrinsically disordered region [260]. Using live imaging experiments performed in vivo in Arabidopsis, with GFP (green fluorescent protein) translational fusions of VIPP1 or VIPP1 lacking Vc (VIPP1ΔVc), it was shown that Vc enables VIPP1 to form oligomeric effector complexes along cell envelopes, whereas VIPP1ΔVc leads to the formation of irregular aggregates of VIPP1 particles. The expression of VIPP1ΔVc complemented the vipp1 knock out mutation in Arabidopsis, but exhibited sensitivity to heat shock. Furthermore, transgenic plants over-expressing wild-type VIPP1 showed enhanced tolerance against heat shock. Vipp1 knockout Arabidopsis mutants show reduced content and other structural defects of thylakoid membranes, as well as reduced photosynthetic activity. In addition to its role in membrane biogenesis, it has been proposed that VIPP1 may also function as a lipid transfer protein, delivering structural lipids into thylakoid or envelope membranes [253]. Overall, these data suggest that the involvement of the Vc disordered region in the formation of the oligomeric effector complexes might be relevant for the control of VIPP1 association/dissociation states. Under conditions of membrane stress, this IDR may permit the insertion of their amphipathic helix into the lipid bilayer to relax the curvature elastic stress in membranes [259].
Dehydration-responsive element binding protein 2A (DREB2A)
Dehydration-responsive element binding protein 2A (DREB2A) is a key transcription factor for drought and heat stress tolerance in Arabidopsis. DREB2A induces the expression of dehydration and heat stress responsive genes [261]. This transcription factor contains several IDRs allowing it to interact with multiple proteins, a characteristic consistent with interactome data showing that DREB2A is a hub protein with 26 nodes [21]. DREB2A may interact with its negative regulators DRIP1 and DRIP2 (DREB2A-interacting protein1 and 2), with ribosomal proteins such as RPL15 (ribosomal protein L15), other transcription factors such as RCD1 (Radical cell death 1), and the transcription co-regulator MED25 (Mediator 25), among others [21]. It has been shown that MED25 binds to one of the DREB2A IDRs and that this interaction results in a gain of ordered structure in this region. Similarly, the binding of DREB2A to its canonical DNA sequence also leads to an increase in the secondary structure of the protein. Data also show that DREB2A conformational changes induced by DNA binding reduce its interaction with the MED25 acid domain, which does not exclude the possibility that this modification may promote its association to another Mediator subunit close by [262]. RCD1 controls DREB2A function, and is itself rapidly removed during abiotic stress [263]. O’Shea et al. [264] showed by NMR spectroscopy that DREB2A undergoes coupled folding and binding with α-helix formation upon interaction with RCD1.
bZIP28, a transcription factor in the unfolded protein response
Under adverse conditions such as heat stress, pathogenesis and by inhibition of protein glycosylation [265,266,267], the demand for protein folding can exceed the capacity of protein homoeostasis systems. This results in the increase of misfolded or unfolded proteins in the endoplasmic reticulum (ER) lumen. This series of events leads to ER stress that subsequently induces the unfolded protein response (UPR) to fulfill the requirement of protein folding and degradation [268]. Two branches of the UPR signaling pathway have been described in plants: one involving the membrane-associated basic leucine zipper (bZIP) transcription factors and the other involving a bifunctional protein, with kinase and ribonuclease activities, known as inositol-requiring enzyme 1 (IRE1), which functions as an RNA splicing factor [269]. In Arabidopsis, bZIP28 is an ER membrane-associated transcription factor; its N-terminal region contains a transcriptional activation domain oriented towards the cytoplasm, while its disordered C-terminal tail localizes to the ER lumen [270]. It is proposed that bZIP28 senses ER stress through its interaction with the ER chaperone BiP (binding immunoglobulin protein), a master regulator of the ER stress sensor. Under non-stress conditions, BiP binds to bZIP28 IDRs present in its lumen-facing tail and retains it in the ER. Upon stress, BiP is competed away from bZIP28 by the accumulation of misfolded proteins in the ER, releasing bZIP28 and allowing its exit from the ER, to move towards the Golgi apparatus [271]. Then, bZIP28 is cleaved by proteases, releasing its transcriptional activation domain that will be translocated to the nucleus to up-regulate stress response genes [271]. The bZIP28 IDR represents one additional example of the role of IDRs in controlling signaling in plant stress responses.
Protein structural disorder in plant biotic stress responses
From germination to reproduction, plants confront a large diversity of parasitic organisms that can cause disease. These pathogens include viruses, bacteria, fungi, nematodes and insects that exploit resources and replication systems in plants [272]. Infection by these organisms has driven plants to evolve refined mechanisms to detect their presence and to mount complex inducible responses to efficiently counteract their attack. As in other plant processes, plant defense systems are tightly regulated, many of them through the participation of kinases and phosphatases that modulate the phosphorylation status of key control proteins [272,273,274]. Marín and Ott [22] have reported the prediction and extensive compilation of different IDPs involved in this process. Because this information has been recently published, in the present work we include only a summary of the material for which functional and structural evidence is available.
Plants are able to specifically recognize their aggressors through receptors localized at the cell membrane. These receptors include LRR-RLKs, a common class of receptors in plants, where intrinsic disorder is present. An example of this is the aforementioned BAK1, an RLK that in this process functions as a co-receptor of the two of the best-characterized pathogen LRR-RLK receptors, FLS2 (Flagellin-sensing 2) and EFR (EF-Tu receptor) [275]. The relevance of the BAK1 IDR C-region resides in its ability to discriminate between two signal transduction pathways, even though the same phosphorylation site (Tyr-610) inside this region is involved in both brassinosteroid sensing and in the pathogen defense response [276]. It is well established that plant perception of pathogens is accompanied by an oxidative burst, where RbohD plays a central role (Fig. 2b). This protein belongs to the NADPH oxidase family, responsible for the early generation of ROS, upstream of calcium and protein phosphorylation signaling. Different experimental evidence supports the presence of an IDR in the RbohD cytoplasmic N terminus, a region that contains an EF-hand motif involved in calcium binding. The malleable nature of this region results in extended conformational changes induced by the synergistic effect of calcium binding and its phosphorylation, which in turn modulates the interaction with small GTPase proteins [277, 278]; fundamental events to set up protection responses to pathogenic agents (Fig. 2b). Following perception at the cell envelope, the signaling process continues in the cytosol, where different molecules play relevant roles. One of these protein molecules is the HSP90 molecular chaperone that given its refolding capacity, is an essential participant in many signaling pathways in plants and animals [279]. The structural organization of this chaperone shows an N-terminal region with an ATPase domain and a linker region composed of charged residues that connect its middle domain with the dimerization region localized at the C terminus. Interestingly, the N-terminal domain undergoes consecutive conformational changes upon ATP binding, leading to the formation of a transient dimer with different co-chaperone partners. The association of HSP90 with the RAR1 co-chaperone results in an order-to-disorder transition of this ATP domain, which enables its movement to allow the entrance of the catalytic loop localized at the middle HSP90 region [280]. These interaction events are essential for the competence of RAR1 function, which together with the SGT1 co-chaperone, is needed to activate the majority of R proteins, detectors of pathogen effector molecules, by mediating NLR (nucleotide binding leucine-rich repeat receptor) function [281]. This signal pathway flows towards a MAP kinase cascade, whose activation ends in the phosphorylation of transcription factors (e.g., WRKY33) that induce the expression of defense genes. Two of these MAP kinases, MEK and MEKK1, show long disordered regions in their N termini that, in the case of MEKK1, have been shown to play a regulatory role; their removal results in a constitutively active kinase [22, 282]. The reprogramming of those genes encoding the proteins that will counteract pathogen incursion needs the action of transcription factors (TFs). Various TF families are involved in this process including MYC, MYB, TGA, WRKY and ERF. Among the TFs known to have a role in the plant pathogen response are MYC2, MYB30, TGA3, WRKY1, WRKY4, WRKY52, WRKY53 and ERF. All these proteins contain, in addition to their DNA-binding domains, IDR-containing linker domains with regulatory functions [21]. Some of these linker domain IDRs have been shown to interact with co-transcription factors that might contribute to the modulation of the spatio-temporal expression of target genes and to the selectivity required to distinguish the identity of particular pathogens (for review see [283]).
Computational analyses using available plant genome sequences predict the presence of significant structural disorder in many more proteins implicated in plant pathogen responses. However, as yet there is no experimental support for their structure or function. Hence, new discoveries await our curiosity and creativity.
Conclusions and future directions
Plants provide a clear picture of the importance of intrinsic disorder in eukaryote protein function. The structural flexibility and molecular promiscuity afforded to a wealth of plant proteins with intrinsically disordered domains have ensured pivotal and multifunctional roles in core processes, including development and metabolism as well as biotic and abiotic stress responses. Technical and experimental barriers to the study of IDPs have limited IDP research in planta, and up to now, there has been a strong reliance on interpretation and extrapolation from in vitro analyses; in particular, for those which are highly disordered and function under stress. It is hoped that the recent explosion in molecular genetic technologies will pave the way for further exploration of the in vivo mechanisms and interactions of plant IDPs. We are only beginning to understand their place in the story of plant evolution and their essential functions in life as a whole.
References
Jones R, Ougham H, Thomas H, Waaland S (2013) The molecular life of plants. Wiley-Blackwell, Hoboken
Raven PH, Evert RF, Eichhorn SE (1999) Biology of plants. W.H. Freeman, Worth Publishers, New York
Bögre L (2001) Cell signaling mechanisms in plants. In: eLS. Wiley, New York. doi:10.1002/9780470015902.a0020134
Hetherington AM, Bardwell L (2011) Plant signaling pathways: a comparative evolutionary overview. Curr Biol 21(9):R317–R319. doi:10.1016/j.cub.2011.04.013
Calabretta S, Richard S (2015) Emerging roles of disordered sequences in RNA-binding proteins. Trends Biochem Sci 40(11):662–672. doi:10.1016/j.tibs.2015.08.012
Oldfield CJ, Dunker AK (2014) Intrinsically disordered proteins and intrinsically disordered protein regions. Annu Rev Biochem 83(1):553–584. doi:10.1146/annurev-biochem-072711-164947
Wright PE, Dyson HJ (2015) Intrinsically disordered proteins in cellular signaling and regulation. Nat Rev Mol Cell Biol 16(1):18–29. doi:10.1038/Nrm3920
Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF, Jones DT (2004) Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol 337(3):635–645. doi:10.1016/j.jmb.2004.02.002
Vucetic S, Brown CJ, Dunker AK, Obradovic Z (2003) Flavors of protein disorder. Proteins 52(4):573–584. doi:10.1002/prot.10437
Schad E, Tompa P, Hegyi H (2011) The relationship between proteome size, structural disorder and organism complexity. Genome Biol 12(12):R120. doi:10.1186/gb-2011-12-12-r120
Vogel C, Chothia C (2006) Protein family expansions and biological complexity. PLoS Comput Biol 2(5):e48. doi:10.1371/journal.pcbi.0020048
Dosztanyi Z, Chen J, Dunker AK, Simon I, Tompa P (2006) Disorder and sequence repeats in hub proteins and their implications for network evolution. J Proteome Res 5(11):2985–2995. doi:10.1021/pr060171o
Fuxreiter M, Tompa P, Simon I, Uversky VN, Hansen JC, Asturias FJ (2008) Malleable machines take shape in eukaryotic transcriptional regulation. Nat Chem Biol 4(12):728–737. doi:10.1038/nchembio.127
Mittag T, Forman-Kay JD (2007) Atomic-level characterization of disordered protein ensembles. Curr Opin Struct Biol 17(1):3–14. doi:10.1016/j.sbi.2007.01.009
Peng Z, Yan J, Fan X, Mizianty MJ, Xue B, Wang K, Hu G, Uversky VN, Kurgan L (2015) Exceptionally abundant exceptions: comprehensive characterization of intrinsic disorder in all domains of life. Cell Mol Life Sci 72(1):137–151. doi:10.1007/s00018-014-1661-9
Pietrosemoli N, García-Martín JA, Solano R, Pazos F (2013) Genome-wide analysis of protein disorder in Arabidopsis thaliana: implications for plant environmental adaptation. PLoS One 8(2):e55524. doi:10.1371/journal.pone.0055524
Sun X, Rikkerink EHA, Jones WT, Uversky VN (2013) Multifarious roles of intrinsic disorder in proteins illustrate its broad impact on plant biology. Plant Cell 25:38–55. doi:10.1105/tpc.112.106062
Dunker AK, Bondos SE, Huang F, Oldfield CJ (2015) Intrinsically disordered proteins and multicellular organisms. Semin Cell Dev Biol 37:44–55. doi:10.1016/j.semcdb.2014.09.025
Yruela I, Contreras-Moreira B (2013) Genetic recombination is associated with intrinsic disorder in plant proteomes. BMC Genom 14(1):772. doi:10.1186/1471-2164-14-772
Yruela I, Contreras-Moreira B (2012) Protein disorder in plants: a view from the chloroplast. BMC Plant Biol 12:165. doi:10.1186/1471-2229-12-165
Kragelund BB, Jensen MK, Skriver K (2012) Order by disorder in plant signaling. Trends Plant Sci 17(11):625–632. doi:10.1016/j.tplants.2012.06.010
Marín M, Ott T (2014) Intrinsic disorder in plant proteins and phytopathogenic bacterial effectors. Chem Rev 114:6912–6932. doi:10.1021/cr400488d
Thieulin-Pardo G, Avilan L, Kojadinovic M, Gontero B (2015) Fairy “tails”: flexibility and function of intrinsically disordered extensions in the photosynthetic world. Front Mol Biosci 2:23. doi:10.3389/fmolb.2015.00023
Jensen MK, Kjaersgaard T, Nielsen MM, Galberg P, Petersen K, O’Shea C, Skriver K (2010) The Arabidopsis thaliana NAC transcription factor family: structure-function relationships and determinants of ANAC019 stress signaling. Biochem J 426:183–196. doi:10.1042/BJ20091234
Taoka K, Yanagimoto Y, Daimon Y, Hibara K, Aida M, Tasaka M (2004) The NAC domain mediates functional specificity of CUP-SHAPED COTYLEDON proteins. Plant J 40(4):462–473. doi:10.1111/j.1365-313X.2004.02238.x
Ko JH, Yang SH, Park AH, Lerouxel O, Han KH (2007) ANAC012, a member of the plant-specific NAC transcription factor family, negatively regulates xylary fiber development in Arabidopsis thaliana. Plant J 50(6):1035–1048. doi:10.1111/j.1365-313X.2007.03109.x
Yoon MK, Shin J, Choi G, Choi BS (2006) Intrinsically unstructured N-terminal domain of bZIP transcription factor HY5. Proteins Struct Funct Bioinf 65(4):856–866. doi:10.1002/prot.21089
Wang J, Jiang JJ, Wang J, Chen L, Fan SL, Wu JW, Wang XL, Wang ZX (2014) Structural insights into the negative regulation of BRI1 signaling by BRI1-interacting protein BKI1. Cell Res 24(11):1328–1341. doi:10.1038/cr.2014.132
Chakrabortee S, Kayatekin C, Newby GA, Mendillo ML, Lancaster A, Lindquist S (2016) Luminidependens (LD) is an Arabidopsis protein with prion behavior. Proc Natl Acad Sci USA 113(21):6065–6070. doi:10.1073/pnas.1604478113
Valsecchi I, Guittard-Crilat E, Maldiney R, Habricot Y, Lignon S, Lebrun R, Miginiac E, Ruelland E, Jeannette E, Lebreton S (2013) The intrinsically disordered C-terminal region of Arabidopsis thaliana TCP8 transcription factor acts both as a transactivation and self-assembly domain. Mol BioSyst 9(9):2282–2295. doi:10.1039/c3mb70128j
Ingram GC, Waites R (2006) Keeping it together: co-ordinating plant growth. Curr Opin Plant Biol 9(1):12–20. doi:10.1016/j.pbi.2005.11.007
Viola IL, Guttlein LN, Gonzalez DH (2013) Redox modulation of plant developmental regulators from the class I TCP transcription factor family. Plant Physiol 162(3):1434–1447. doi:10.1104/pp.113.216416
Aguilar-Martinez JA, Sinha N (2013) Analysis of the role of Arabidopsis class I TCP genes AtTCP7, AtTCP8, AtTCP22, and AtTCP23 in leaf development. Front Plant Sci. doi:10.3389/fpls.2013.00406
Hammani K, Gobert A, Hleibieh K, Choulier L, Small I, Giege P (2011) An Arabidopsis dual-localized pentatricopeptide repeat protein interacts with nuclear proteins involved in gene expression regulation. Plant Cell 23(2):730–740. doi:10.1105/tpc.110.081638
Olsen AN, Ernst HA, Lo Leggio L, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10(2):79–87. doi:10.1016/j.tplants.2004.12.010
Sun XL, Rikkerink EHA, Jones WT, Uversky VN (2013) Multifarious roles of intrinsic disorder in proteins illustrate its broad impact on plant biology. Plant Cell 25(1):38–55. doi:10.1105/Tpc.112.106062
Kjaersgaard T, Jensen MK, Christiansen MW, Gregersen P, Kragelund BB, Skriver K (2011) Senescence-associated barley NAC (NAM, ATAF1,2, CUC) transcription factor interacts with radical-induced cell death 1 through a disordered regulatory domain. J Biol Chem 286(41):35418–35429. doi:10.1074/jbc.M111.247221
O’shea C, Kryger M, Stender EGP, Kragelund BB, Willemoes M, Skriver K (2015) Protein intrinsic disorder in Arabidopsis NAC transcription factors: transcriptional activation by ANAC013 and ANAC046 and their interactions with RCD1. Biochem J 465:281–294. doi:10.1042/BJ20141045
Oyama T, Shimura Y, Okada K (1997) The Arabidopsis HY5 gene encodes a bZIP protein that regulates stimulus-induced development of root and hypocotyl. Gene Dev 11(22):2983–2995. doi:10.1101/Gad.11.22.2983
Ahmad M (2016) Photocycle and signaling mechanisms of plant cryptochromes. Curr Opin Plant Biol 33:108–115. doi:10.1016/j.pbi.2016.06.013
Chaves I, Pokorny R, Byrdin M, Hoang N, Ritz T, Brettel K, Essen LO, van der Horst GTJ, Batschauer A, Ahmad M (2011) The cryptochromes: blue light photoreceptors in plants and animals. Annu Rev Plant Biol 62:335–364. doi:10.1146/annurev-arplant-042110-103759
Cashmore AR, Jarillo JA, Wu YJ, Liu D (1999) Cryptochromes: blue light receptors for plants and animals. Science 284(5415):760–765
Lin C, Shalitin D (2003) Cryptochrome structure and signal transduction. Annu Rev Plant Biol 54:469–496. doi:10.1146/annurev.arplant.54.110901.160901
Liu B, Zuo Z, Liu H, Liu X, Lin C (2011) Arabidopsis cryptochrome 1 interacts with SPA1 to suppress COP1 activity in response to blue light. Genes Dev 25(10):1029–1034. doi:10.1101/gad.2025011
Wang H, Ma L-G, Li J-M, Zhao H-Y, Deng XW (2001) Direct interaction of Arabidopsis cryptochromes with COP1 in light control development. Science 294(5540):154–158. doi:10.1126/science.1063630
Wang Q, Barshop WD, Bian M, Vashisht AA, He R, Yu X, Liu B, Nguyen P, Liu X, Zhao X, Wohlschlegel JA, Lin C (2015) The blue light-dependent phosphorylation of the CCE domain determines the photosensitivity of Arabidopsis CRY2. Mol Plant 8(4):631–643. doi:10.1016/j.molp.2015.03.005
Brautigam CA, Smith BS, Ma ZQ, Palnitkar M, Tomchick DR, Machius M, Deisenhofer J (2004) Structure of the photolyase-like domain of cryptochrome 1 from Arabidopsis thaliana. Proc Natl Acad Sci USA 101(33):12142–12147. doi:10.1073/pnas.0404851101
Partch CL, Clarkson MW, Ozgur S, Lee AL, Sancar A (2005) Role of structural plasticity in signal transduction by the cryptochrome blue-light photoreceptor. Biochem 44(10):3795–3805. doi:10.1021/bi047545g
Kondoh M, Shiraishi C, Muller P, Ahmad M, Hitomi K, Getzoff ED, Terazima M (2011) Light-induced conformational changes in full-length Arabidopsis thaliana cryptochrome. J Mol Biol 413(1):128–137. doi:10.1016/j.jmb.2011.08.031
Pfluger J, Wagner D (2007) Histone modifications and dynamic regulation of genome accessibility in plants. Curr Opin Plant Biol 10(6):645–652. doi:10.1016/j.pbi.2007.07.013
Mehdi S, Derkacheva M, Ramstrom M, Kralemann L, Bergquist J, Hennig L (2016) The WD40 domain protein MSI1 functions in a histone deacetylase complex to fine-tune abscisic acid signaling. Plant Cell 28(1):42–54. doi:10.1105/tpc.15.00763
Perrella G, Lopez-Vernaza MA, Carr C, Sani E, Gossele V, Verduyn C, Kellermeier F, Hannah MA, Amtmann A (2013) Histone deacetylase complex1 expression level titrates plant growth and abscisic acid sensitivity in Arabidopsis. Plant Cell 25(9):3491–3505. doi:10.1105/tpc.113.114835
Perrella G, Carr C, Asensi-Fabado MA, Donald NA, Paldi K, Hannah MA, Amtmann A (2016) The histone deacetylase complex 1 protein of Arabidopsis has the capacity to interact with multiple proteins including histone 3-binding proteins and histone 1 variants. Plant Physiol 171(1):62–70. doi:10.1104/pp.15.01760
Clouse SD (2011) Brassinosteroid signal transduction: from receptor kinase activation to transcriptional networks regulating plant development. Plant Cell 23(4):1219–1230. doi:10.1105/tpc.111.084475
Kim TW, Wang ZY (2010) Brassinosteroid signal transduction from receptor kinases to transcription factors. Annu Rev Plant Biol 61:681–704. doi:10.1146/annurev.arplant.043008.092057
Xie HB, Vucetic S, Iakoucheva LM, Oldfield CJ, Dunker AK, Uversky VN, Obradovic Z (2007) Functional anthology of intrinsic disorder. 1. Biological processes and functions of proteins with long disordered regions. J Proteome Res 6(5):1882–1898. doi:10.1021/pr060392u
Nam KH, Li JM (2002) BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 110(2):203–212. doi:10.1016/S0092-8674(02)00814-0
Wang XL, Chory J (2006) Brassinosteroids regulate dissociation of BKI1, a negative regulator of BRI1 signaling, from the plasma membrane. Science 313(5790):1118–1122. doi:10.1126/science.1127593
Wang XF, Kota U, He K, Blackburn K, Li J, Goshe MB, Huber SC, Clouse SD (2008) Sequential transphosphorylation of the BRI1/BAK1 receptor kinase complex impacts early events in brassinosteroid signaling. Dev Cell 15(2):220–235. doi:10.1016/j.devcel.2008.06.011
Hothorn M, Belkhadir Y, Dreux M, Dabi T, Noel JP, Wilson IA, Chory J (2011) Structural basis of steroid hormone perception by the receptor kinase BRI1. Nature 474(7352):467–471. doi:10.1038/nature10153
Jiang JJ, Wang T, Wu ZH, Wang J, Zhang C, Wang HJ, Wang ZX, Wang XL (2015) The intrinsically disordered protein BKI1 is essential for inhibiting BRI1 signaling in plants. Mol Plant 8(11):1675–1678. doi:10.1016/j.molp.2015.07.012
Song YH, Ito S, Imaizumi T (2013) Flowering time regulation: photoperiod- and temperature-sensing in leaves. Trends Plant Sci 18(10):575–583. doi:10.1016/j.tplants.2013.05.003
Schmitz RJ, Amasino RM (2007) Vernalization: a model for investigating epigenetics and eukaryotic gene regulation in plants. BBA Gene Struct Expr 1769(5–6):269–275. doi:10.1016/j.bbaexp.2007.02.003
Shorter J, Lindquist S (2005) Prions as adaptive conduits of memory and inheritance. Nat Rev Genet 6(6):435–450. doi:10.1038/nrg1616
Mukhopadhyay S, Krishnan R, Lemke EA, Lindquist S, Deniz AA (2007) A natively unfolded yeast prion monomer adopts an ensemble of collapsed and rapidly fluctuating structures. Proc Natl Acad Sci USA 104(8):2649–2654. doi:10.1073/pnas.0611503104
Malinovska L, Kroschwald S, Alberti S (2013) Protein disorder, prion propensities, and self-organizing macromolecular collectives. BBA Proteins Proteom 1834(5):918–931. doi:10.1016/j.bbapap.2013.01.003
Romero P, Obradovic Z, Li XH, Garner EC, Brown CJ, Dunker AK (2001) Sequence complexity of disordered protein. Proteins Struct Funct Genet 42(1):38–48. doi:10.1002/1097-0134(20010101)42:1<38:Aid-Prot50>3.0.Co;2-3
Sun XL, Jones WT, Rikkerink EHA (2012) GRAS proteins: the versatile roles of intrinsically disordered proteins in plant signaling. Biochem J 442:1–12. doi:10.1042/BJ20111766
Sun X, Xue B, Jones WT, Rikkerink E, Dunker AK, Uversky VN (2011) A functionally required unfoldome from the plant kingdom: intrinsically disordered N-terminal domains of GRAS proteins are involved in molecular recognition during plant development. Plant Mol Biol 77:205–223. doi:10.1007/s11103-011-9803-z
Sun XL, Jones WT, Harvey D, Edwards PJB, Pascal SM, Kirk C, Considine T, Sheerin DJ, Rakonjac J, Oldfield CJ, Xue B, Dunker AK, Uversky VN (2010) N-terminal domains of DELLA proteins are intrinsically unstructured in the absence of interaction with GID1/gibberellic acid receptors. J Biol Chem 285(15):11557–11571. doi:10.1074/jbc.M109.027011
Daviere JM, Achard P (2016) A pivotal role of DELLAs in regulating multiple hormone signals. Mol Plant 9(1):10–20. doi:10.1016/j.molp.2015.09.011
Smertenko AP, Chang H-Y, Wagner V, Kaloriti D, Fenyk S, Sonobe S, Lloyd C, Hauser M-T, Hussey PJ (2004) The Arabidopsis microtubule-associated protein AtMAP65-1: molecular analysis of its microtubule bundling activity. Plant Cell 16(8):2035–2047. doi:10.1105/tpc.104.023937
Hussey PJ, Hawkins TJ, Igarashi H, Kaloriti D, Smertenko A (2002) The plant cytoskeleton: recent advances in the study of the plant microtubule-associated proteins MAP-65, MAP-190 and the Xenopus MAP215-like protein, MOR1. Plant Mol Biol 50(6):915–924. doi:10.1023/A:1021236307508
Smertenko AP, Kaloriti D, Chang H-Y, Fiserova J, Opatrny Z, Hussey PJ (2008) The C-terminal variable region specifies the dynamic properties of Arabidopsis microtubule-associated protein MAP65 isotypes. Plant Cell 20(12):3346–3358. doi:10.1105/tpc.108.063362
Boruc J, Weimer A, Stoppin-Mellet V, Mylle E, Kosetsu K, Cedeno C, Jaquinod M, Njo M, De Milde L, Tompa P, Gonzalez N, Inze D, Beeckman T, Vantard M, van Damme D (2016) Phosphorylation of MAP65-1 by Arabidopsis Aurora kinases is required for efficient cell cycle progression. Plant Physiol. doi:10.1104/pp.16.01602
Matzke MA, Mosher RA (2014) RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat Rev Genet 15(6):394–408. doi:10.1038/nrg3683
Blevins T, Podicheti R, Mishra V, Marasco M, Wang J, Rusch D, Tang H, Pikaard CS (2015) Identification of Pol IV and RDR2-dependent precursors of 24 nt siRNAs guiding de novo DNA methylation in Arabidopsis. Elife 4:e09591. doi:10.7554/eLife.09591
Henderson IR, Zhang X, Lu C, Johnson L, Meyers BC, Green PJ, Jacobsen SE (2006) Dissecting Arabidopsis thaliana DICER function in small RNA processing, gene silencing and DNA methylation patterning. Nat Genet 38(6):721–725. doi:10.1038/ng1804
Yu B, Bi L, Zhai J, Agarwal M, Li S, Wu Q, Ding SW, Meyers BC, Vaucheret H, Chen X (2010) siRNAs compete with miRNAs for methylation by HEN1 in Arabidopsis. Nucleic Acids Res 38(17):5844–5850. doi:10.1093/nar/gkq348
Ye R, Wang W, Iki T, Liu C, Wu Y, Ishikawa M, Zhou X, Qi Y (2012) Cytoplasmic assembly and selective nuclear import of Arabidopsis Argonaute4/siRNA complexes. Mol Cell 46(6):859–870. doi:10.1016/j.molcel.2012.04.013
Wierzbicki AT, Ream TS, Haag JR, Pikaard CS (2009) RNA polymerase V transcription guides ARGONAUTE4 to chromatin. Nat Genet 41(5):630–634. doi:10.1038/ng.365
Rowley MJ, Avrutsky MI, Sifuentes CJ, Pereira L, Wierzbicki AT (2011) Independent chromatin binding of ARGONAUTE4 and SPT5L/KTF1 mediates transcriptional gene silencing. PLoS Genet 7(6):e1002120. doi:10.1371/journal.pgen.1002120
Matzke MA, Kanno T, Matzke AJ (2015) RNA-directed DNA methylation: the evolution of a complex epigenetic pathway in flowering plants. Annu Rev Plant Biol 66:243–267. doi:10.1146/annurev-arplant-043014-114633
Trujillo JT, Beilstein MA, Mosher RA (2016) The Argonaute-binding platform of NRPE1 evolves through modulation of intrinsically disordered repeats. New Phytol 212(4):1094–1105. doi:10.1111/nph.14089
Till S, Lejeune E, Thermann R, Bortfeld M, Hothorn M, Enderle D, Heinrich C, Hentze MW, Ladurner AG (2007) A conserved motif in Argonaute-interacting proteins mediates functional interactions through the Argonaute PIWI domain. Nat Struct Mol Biol 14(10):897–903. doi:10.1038/nsmb1302
El-Shami M, Pontier D, Lahmy S, Braun L, Picart C, Vega D, Hakimi MA, Jacobsen SE, Cooke R, Lagrange T (2007) Reiterated WG/GW motifs form functionally and evolutionarily conserved ARGONAUTE-binding platforms in RNAi-related components. Genes Dev 21(20):2539–2544. doi:10.1101/gad.451207
Pfaff J, Meister G (2013) Argonaute and GW182 proteins: an effective alliance in gene silencing. Biochem Soc Trans 41(4):855–860. doi:10.1042/bst20130047
Radivojac P, Clark WT, Oron TR, Schnoes AM, Wittkop T, Sokolov A, Graim K, Funk C, Verspoor K, Ben-Hur A, Pandey G, Yunes JM, Talwalkar AS, Repo S, Souza ML, Piovesan D, Casadio R, Wang Z, Cheng J, Fang H, Gough J, Koskinen P, Toronen P, Nokso-Koivisto J, Holm L, Cozzetto D, Buchan DW, Bryson K, Jones DT, Limaye B, Inamdar H, Datta A, Manjari SK, Joshi R, Chitale M, Kihara D, Lisewski AM, Erdin S, Venner E, Lichtarge O, Rentzsch R, Yang H, Romero AE, Bhat P, Paccanaro A, Hamp T, Kassner R, Seemayer S, Vicedo E, Schaefer C, Achten D, Auer F, Boehm A, Braun T, Hecht M, Heron M, Honigschmid P, Hopf TA, Kaufmann S, Kiening M, Krompass D, Landerer C, Mahlich Y, Roos M, Bjorne J, Salakoski T, Wong A, Shatkay H, Gatzmann F, Sommer I, Wass MN, Sternberg MJ, Skunca N, Supek F, Bosnjak M, Panov P, Dzeroski S, Smuc T, Kourmpetis YA, van Dijk AD, ter Braak CJ, Zhou Y, Gong Q, Dong X, Tian W, Falda M, Fontana P, Lavezzo E, Di Camillo B, Toppo S, Lan L, Djuric N, Guo Y, Vucetic S, Bairoch A, Linial M, Babbitt PC, Brenner SE, Orengo C, Rost B, Mooney SD, Friedberg I (2013) A large-scale evaluation of computational protein function prediction. Nat Methods 10(3):221–227. doi:10.1038/nmeth.2340
Fersht AR, Knill-Jones JW, Bedouelle H, Winter G (1988) Reconstruction by site-directed mutagenesis of the transition state for the activation of tyrosine by the tyrosyl-tRNA synthetase: a mobile loop envelopes the transition state in an induced-fit mechanism. Biochemistry 27(5):1581–1587. doi:10.1021/bi00405a028
McElheny D, Schnell JR, Lansing JC, Dyson HJ, Wright PE (2005) Defining the role of active-site loop fluctuations in dihydrofolate reductase catalysis. Proc Natl Acad Sci USA 102(14):5032–5037. doi:10.1073/pnas.0500699102
Schulenburg C, Hilvert D (2013) Protein conformational disorder and enzyme catalysis. Top Curr Chem 337:41–67. doi:10.1007/128_2012_411
Erales J, Lorenzi M, Lebrun R, Fournel A, Etienne E, Courcelle C, Guigliarelli B, Gontero B, Belle V (2009) A new function of GAPDH from Chlamydomonas reinhardtii: a thiol-disulfide exchange reaction with CP12. Biochemistry 48:6034–6040. doi:10.1021/bi900569h
Graciet E, Gans P, Wedel N, Lebreton S, Camadro JM, Gontero B (2003) The small protein CP12: a protein linker for supramolecular complex assembly. Biochemistry 42:8163–8170. doi:10.1021/bi034474x
Groben R, Kaloudas D, Raines CA, Offmann B, Maberly SC, Gontero B (2010) Comparative sequence analysis of CP12, a small protein involved in the formation of a Calvin cycle complex in photosynthetic organisms. Photosynth Res 103:183–194. doi:10.1007/s11120-010-9542-z
Stanley DN, Raines CA, Kerfeld CA (2013) Comparative analysis of 126 cyanobacterial genomes reveals evidence of functional diversity among homologs of the redox-regulated CP12 protein. Plant Physiol 161(2):824–835. doi:10.1104/pp.112.210542
Fermani S, Trivelli X, Sparla F, Thumiger A, Calvaresi M, Marri L, Falini G, Zerbetto F, Trost P (2012) Conformational selection and folding-upon-binding of intrinsically disordered protein CP12 regulate photosynthetic enzymes assembly. J Biol Chem 287:21372–21383. doi:10.1074/jbc.M112.350355
Mileo E, Lorenzi M, Erales J, Lignon S, Puppo C, Le Breton N, Etienne E, Marque SRA, Guigliarelli B, Gontero B, Belle V (2013) Dynamics of the intrinsically disordered protein CP12 in its association with GAPDH in the green alga Chlamydomonas reinhardtii: a fuzzy complex. Mol BioSyst 9:2869–2876. doi:10.1039/c3mb70190e
Moparthi SB, Thieulin-Pardo G, Mansuelle P, Rigneault H, Gontero B, Wenger J (2014) Conformational modulation and hydrodynamic radii of CP12 protein and its complexes probed by fluorescence correlation spectroscopy. FEBS J 281:3206–3217. doi:10.1111/febs.12854
Marri L, Thieulin-Pardo G, Lebrun R, Puppo R, Zaffagnini M, Trost P, Gontero B, Sparla F (2014) CP12-mediated protection of Calvin–Benson cycle enzymes from oxidative stress. Biochimie 97:228–237. doi:10.1016/j.biochi.2013.10.018
Lopez-Calcagno PE, Howard TP, Raines CA (2014) The CP12 protein family: a thioredoxin-mediated metabolic switch? Front Plant Sci 5:9. doi:10.3389/fpls.2014.00009
Pohlmeyer K, Paap BK, Soll J, Wedel N (1996) CP12: a small nuclear-encoded chloroplast protein provides novel insights into higher-plant GAPDH evolution. Plant Mol Biol 32:969–978. doi:10.1007/BF00020493
Matsumura H, Kai A, Maeda T, Tamoi M, Satoh A, Tamura H, Hirose M, Ogawa T, Kizu N, Wadano A, Inoue T, Shigeoka S (2011) Structure basis for the regulation of glyceraldehyde-3-phosphate dehydrogenase activity via the intrinsically disordered protein CP12. Structure 19:1846–1854. doi:10.1016/j.str.2011.08.016
Sparla F, Pupillo P, Trost P (2002) The C-terminal extension of glyceraldehyde-3-phosphate dehydrogenase subunit B acts as an autoinhibitory domain regulated by thioredoxins and nicotinamide adenine dinucleotide. J Biol Chem 277(47):44946–44952. doi:10.1074/jbc.M206873200
Ellis RJ (1979) The most abundant protein in the world. Trends Biochem Sci 4:241–244. doi:10.1016/0968-0004(79)90212-3
Raven JA (2013) RuBisCO: still the most abundant protein of Earth? New Phytol 198:1–3. doi:10.1111/nph.12197
Cleland WW, Andrews TJ, Gutteridge S, Hartman FC, Lorimer GH (1998) Mechanism of RuBisCO: the carbamate as general base. Chem Rev 98:549–562. doi:10.1021/cr970010r
Gontero B, Salvucci ME (2014) Regulation of photosynthetic carbon metabolism in aquatic and terrestrial organisms by RuBisCO activase, redox-modulation and CP12. Aquat Bot 118:14–23. doi:10.1016/j.aquabot.2014.05.011
Zhang N, Portis AR (1999) Mechanism of light regulation of RuBisCO: a specific role for the larger RuBisCO activase isoform involving reductive activation by Thioredoxin-f. Proc Natl Acad Sci USA 96:9438–9443. doi:10.1073/pnas.96.16.9438
Stotz M, Mueller-Cajar O, Ciniawsky S, Wendler P, Hartl FU, Bracher A, Hayer-Hartl M (2011) Structure of green-type RuBisCO activase from tobacco. Nat Struct Mol Biol 18:1366–1370. doi:10.1038/nsmb.2171
Shen JB, Ogren WL (1992) Alteration of spinach ribulose-1,5-bisphosphate carboxylase/oxygenase activase activities by site-directed mutagenesis. Plant Physiol 99(3):1201–1207. doi:10.1104/pp.99.3.1201
Wang D, Portis AR (2006) Increased sensitivity of oxidized large isoform of ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO) activase to ADP inhibition is due to an interaction between its carboxyl extension and nucleotide-binding pocket. J Biol Chem 281:25241–25249. doi:10.1074/jbc.M604756200
Carmo-Silva AE, Salvucci ME (2013) The regulatory properties of RuBisCO activase differ among species and affect photosynthetic induction during light transitions. Plant Physiol 161:1645–1655. doi:10.1104/pp.112.213348
Zarzycki J, Axen SD, Kinney JN, Kerfeld CA (2013) Cyanobacterial-based approaches to improving photosynthesis in plants. J Exp Bot 64(3):787–798. doi:10.1093/jxb/ers294
Seidler A (1996) The extrinsic polypeptides of photosystem II. Biochim Biophys Acta Bioenerg 1277:35–60. doi:10.1016/S0005-2728(96)00102-8
Lydakis-Simantiris N, Betts SD, Yocum CF (1999) Leucine 245 is a critical residue for folding and function of the manganese stabilizing protein of photosystem II. Biochemistry 38:15528–15535. doi:10.1021/bi991599m
Shutova T, Irrgang KD, Shubin V, Klimov VV, Renger G (1997) Analysis of pH-induced structural changes of the isolated extrinsic 33 kilodalton protein of photosystem II. Biochemistry 36:6350–6358. doi:10.1021/bi963115h
Yi X, McChargue M, Laborde S, Frankel LK, Bricker TM (2005) The manganese-stabilizing protein is required for photosystem II assembly/stability and photoautotrophy in higher plants. J Biol Chem 280(16):16170–16174. doi:10.1074/jbc.M501550200
Lydakis-Simantiris N, Hutchison RS, Betts SD, Barry BA, Yocum CF (1999) Manganese stabilizing protein of photosystem II is a thermostable, natively unfolded polypeptide. Biochemistry 38:404–414. doi:10.1021/bi981847z
Popelkova H, Betts SD, Lydakis-Symantiris N, Im MM, Swenson E, Yocum CF (2006) Mutagenesis of basic residues R151 and R161 in manganese-stabilizing protein of photosystem II causes inefficient binding of chloride to the oxygen-evolving complex. Biochemistry 45:3107–3115. doi:10.1021/bi0523759
Falk S, Ravaud S, Koch J, Sinning I (2010) The C terminus of the Alb3 membrane insertase recruits cpSRP43 to the thylakoid membrane. J Biol Chem 285:5954–5962. doi:10.1074/jbc.M109.084996
Kroliczewski J, Piskozub M, Bartoszewski R, Kroliczewska B (2016) ALB3 insertase mediates cytochrome b6 co-translational import into the thylakoid membrane. Sci Rep 6:34557. doi:10.1038/srep34557
Bryson EA, Rankin SE, Carey M, Watts A, Pinheiro TJ (1999) Folding of apocytochrome c in lipid micelles: formation of alpha-helix precedes membrane insertion. Biochemistry 38(30):9758–9767. doi:10.1021/bi990119o
Urbischek M, Nick von Braun S, Brylok T, Gügel IL, Richter A, Koskela M, Grimm B, Mulo P, Bölter B, Soll J, Ankele E, Schwenkert S (2015) The extreme Albino3 (Alb3) C terminus is required for Alb3 stability and function in Arabidopsis thaliana. Planta 242(3):733–746. doi:10.1007/s00425-015-2352-y
Marusek CM, Trobaugh NM, Flurkey WH, Inlow JK (2006) Comparative analysis of polyphenol oxidase from plant and fungal species. J Inorg Biochem 100:108–123. doi:10.1016/j.jinorgbio.2005.10.008
O’Leary B, Park J, Plaxton WC (2011) The remarkable diversity of plant PEPC (phosphoenolpyruvate carboxylase): recent insights into the physiological functions and post-translational controls of non-photosynthetic PEPCs. Biochem J 436:15–34. doi:10.1042/BJ20110078
Dalziel KJ, O’Leary B, Brikis C, Rao SK, She Y-M, Cyr T, Plaxton WC (2012) The bacterial-type phosphoenolpyruvate carboxylase isozyme from developing castor oil seeds is subject to in vivo regulatory phosphorylation at serine-451. FEBS Lett 586:1049–1054. doi:10.1016/j.febslet.2012.02.054
Musiani F, Ippoliti E, Micheletti C, Carloni P, Ciurli S (2013) Conformational fluctuations of UreG, an intrinsically disordered enzyme. Biochemistry 52:2949–2954. doi:10.1021/bi4001744
Neyroz P, Zambelli B, Ciurli S (2006) Intrinsically disordered structure of Bacillus pasteurii UreG as revealed by steady-state and time-resolved fluorescence spectroscopy. Biochemistry 45:8918–8930. doi:10.1021/bi060227s
Zambelli B, Cremades N, Neyroz P, Turano P, Uversky VN, Ciurli S (2012) Insights in the (un)structural organization of Bacillus pasteurii UreG, an intrinsically disordered GTPase enzyme. Mol BioSyst 8:220–228. doi:10.1039/c1mb05227f
Real-Guerra R, Staniscuaski F, Zambelli B, Musiani F, Ciurli S, Carlini CR (2012) Biochemical and structural studies on native and recombinant Glycine max UreG: a detailed characterization of a plant urease accessory protein. Plant Mol Biol 78(4–5):461–475. doi:10.1007/s11103-012-9878-1
Karplus PA, Pearson MA, Hausinger RP (1997) 70 years of crystalline urease: what have we learned? Acc Chem Res 30:330–337. doi:10.1021/ar960022j
Carlini CR, Oliveira AE, Azambuja P, Xavier-Filho J, Wells MA (1997) Biological effects of canatoxin in different insect models: evidence for a proteolytic activation of the toxin by insect cathepsin-like enzymes. J Econ Entomol 90:340–348. doi:10.1093/jee/90.2.340
Staniscuaski F, Ferreira-Dasilva CT, Mulinari F, Pires-Alves M, Carlini CR (2005) Insecticidal effects of canatoxin on the cotton stainer bug Dysdercus peruvianus (Hemiptera: Pyrrhocoridae). Toxicon 45(6):753–760. doi:10.1016/j.toxicon.2005.01.014
Mulinari F, Stanisçuaski F, Bertholdo-Vargas LR, Postal M, Oliveira-Neto OB, Rigden DJ, Grossi-de-Sá MF, Carlini CR (2007) Jaburetox-2Ec: an insecticidal peptide derived from an isoform of urease from the plant Canavalia ensiformis. Peptides 28:2042–2050. doi:10.1016/j.peptides.2007.08.009
Menez A (1998) Functional architectures of animal toxins: a clue to drug design? Toxicon 36(11):1557–1572. doi:10.1016/S0041-0101(98)00148-2
Lopes FC, Dobrovolska O, Real-Guerra R, Broll V, Zambelli B, Musiani F, Uversky VN, Carlini CR, Ciurli S (2015) Pliable natural biocide: Jaburetox is an intrinsically disordered insecticidal and fungicidal polypeptide derived from jack bean urease. FEBS J 282:1043–1064. doi:10.1111/febs.13201
Battaglia M, Olvera-Carrillo Y, Garciarrubio A, Campos F, Covarrubias AA (2008) The enigmatic LEA proteins and other hydrophilins. Plant Physiol 148(1):6–24. doi:10.1104/pp.108.120725
Rivera-Najera LY, Saab-Rincon G, Battaglia M, Amero C, Pulido NO, Garcia-Hernandez E, Solorzano RM, Reyes JL, Covarrubias AA (2014) A group 6 late embryogenesis abundant protein from common bean is a disordered protein with extended helical structure and oligomer-forming properties. J Biol Chem 289(46):31995–32009. doi:10.1074/jbc.M114.583369
Cuevas-Velazquez CL, Saab-Rincon G, Reyes JL, Covarrubias AA (2016) The unstructured N-terminal region of Arabidopsis group 4 late embryogenesis abundant (LEA) proteins is required for folding and for chaperone-like activity under water deficit. J Biol Chem 291(20):10893–10903. doi:10.1074/jbc.M116.720318
Shih M-D, Hoekstra FA, Hsing Y-IC (2008) Late embryogenesis abundant proteins. Adv Bot Res 48:211–255. doi:10.1016/S0065-2296(08)00404-7
Hand SC, Jones D, Menze MA, Witt TL (2007) Life without water: expression of plant LEA genes by an anhydrobiotic arthropod. J Exp Zool A Ecol Genet Physiol 307(1):62–66. doi:10.1002/jez.a.343
Stacy RAP, Aalen RB (1998) Identification of sequence homology between the internal hydrophilic repeated motifs of Group 1 late-embryogenesis-abundant proteins in plants and hydrophilic repeats of the general stress protein GsiB of Bacillus subtilis. Planta 206(3):476–478. doi:10.1007/S004250050424
Tunnacliffe A, Lapinski J, McGee B (2005) A putative LEA protein, but no trehalose, is present in anhydrobiotic bdelloid rotifers. Hydrobiologia 546:315–321. doi:10.1007/s10750-005-4239-6
Browne J, Tunnacliffe A, Burnell A (2002) Anhydrobiosis: plant desiccation gene found in a nematode. Nature 416(6876):38. doi:10.1038/416038a
Hand SC, Menze MA, Toner M, Boswell L, Moore D (2011) LEA proteins during water stress: not just for plants anymore. Annu Rev Physiol 73:115–134. doi:10.1146/annurev-physiol-012110-142203
Hincha DK, Thalhammer A (2012) LEA proteins: IDPs with versatile functions in cellular dehydration tolerance. Biochem Soc Trans 40:1000–1003. doi:10.1042/Bst20120109
Hundertmark M, Popova AV, Rausch S, Seckler R, Hincha DK (2012) Influence of drying on the secondary structure of intrinsically disordered and globular proteins. Biochem Biophys Res Commun 417(1):122–128. doi:10.1016/J.Bbrc.2011.11.067
Garay-Arroyo A, Colmenero-Flores JM, Garciarrubio A, Covarrubias AA (2000) Highly hydrophilic proteins in prokaryotes and eukaryotes are common during conditions of water deficit. J Biol Chem 275(8):5668–5674. doi:10.1074/jbc.275.8.5668
Reyes JL, Rodrigo MJ, Colmenero-Flores JM, Gil JV, Garay-Arroyo A, Campos F, Salamini F, Bartels D, Covarrubias AA (2005) Hydrophilins from distant organisms can protect enzymatic activities from water limitation effects in vitro. Plant Cell Environ 28(6):709–718. doi:10.1111/J.1365-3040.2005.01317.X
Goyal K, Walton LJ, Tunnacliffe A (2005) LEA proteins prevent protein aggregation due to water stress. Biochem J 388:151–157. doi:10.1042/BJ20041931
Sanchez-Ballesta MT, Rodrigo MJ, LaFuente MT, Granell A, Zacarias L (2004) Dehydrin from citrus, which confers in vitro dehydration and freezing protection activity, is constitutive and highly expressed in the flavedo of fruit but responsive to cold and water stress in leaves. J Agr Food Chem 52(7):1950–1957. doi:10.1021/Jf035216
Tantos A, Friedrich P, Tompa P (2009) Cold stability of intrinsically disordered proteins. FEBS Lett 583(2):465–469. doi:10.1016/J.Febslet.2008.12.054
Brini F, Saibi W, Amara I, Gargouri A, Masmoudi K, Hanin M (2010) Wheat dehydrin DHN-5 exerts a heat-protective effect on beta-glucosidase and glucose oxidase activities. Biosci Biotech Biochem 74(5):1050–1054. doi:10.1271/Bbb.90949
Drira M, Saibi W, Brini F, Gargouri A, Masmoudi K, Hanin M (2013) The K-segments of the wheat dehydrin DHN-5 are essential for the protection of lactate dehydrogenase and beta-glucosidase activities in vitro. Mol Biotechnol 54(2):643–650. doi:10.1007/s12033-012-9606-8
Campos F, Cuevas-Velazquez C, Fares MA, Reyes JL, Covarrubias AA (2013) Group 1 LEA proteins, an ancestral plant protein group, are also present in other eukaryotes, and in the archaea and bacteria domains. Mol Genet Genom 288(10):503–517. doi:10.1007/s00438-013-0768-2
Grelet J, Benamar A, Teyssier E, Avelange-Macherel MH, Grunwald D, Macherel D (2005) Identification in pea seed mitochondria of a late-embryogenesis abundant protein able to protect enzymes from drying. Plant Physiol 137(1):157–167. doi:10.1104/pp.104.052480
Santhanagopalan I, Basha E, Ballard KN, Bopp NE, Vierling E (2015) Model chaperones: small heat shock proteins from plants. In: Tanguay RM, Hightower LE (eds) The big book on small heat shock proteins. Springer International Publishing, Cham, pp 119–153. doi:10.1007/978-3-319-16077-1_5
Reyes JL, Campos F, Wei H, Arora R, Yang YI, Karlson DT, Covarrubias AA (2008) Functional dissection of hydrophilins during in vitro freeze protection. Plant Cell Environ 31(12):1781–1790. doi:10.1111/J.1365-3040.2008.01879.X
Tunnacliffe A, Wise MJ (2007) The continuing conundrum of the LEA proteins. Naturwissenschaften 94(10):791–812. doi:10.1007/S00114-007-0254-Y
Hughes SL, Schart V, Malcolmson J, Hogarth KA, Martynowicz DM, Tralman-Baker E, Patel SN, Graether SP (2013) The importance of size and disorder in the cryoprotective effects of dehydrins. Plant Physiol 163(3):1376–1386. doi:10.1104/pp.113.226803
Cuevas-Velazquez CL, Rendon-Luna DF, Covarrubias AA (2014) Dissecting the cryoprotection mechanisms for dehydrins. Front Plant Sci 5:583. doi:10.3389/fpls.2014.00583
Kazuoka T, Oeda K (1994) Purification and characterization of Cor85-oligomeric complex from cold-acclimated spinach. Plant Cell Physiol 35(4):601–611
Houde M, Daniel C, Lachapelle M, Allard F, Laliberte S, Sarhan F (1995) Immunolocalization of freezing-tolerance-associated proteins in the cytoplasm and nucleoplasm of wheat crown tissues. Plant J 8(4):583–593. doi:10.1046/J.1365-313x.1995.8040583.X
Bravo LA, Gallardo J, Navarrete A, Olave N, Martinez J, Alberdi M, Close TJ, Corcuera LJ (2003) Cryoprotective activity of a cold-induced dehydrin purified from barley. Physiol Plant 118(2):262–269. doi:10.1034/J.1399-3054.2003.00060.X
Nakayama K, Okawa K, Kakizaki T, Honma T, Itoh H, Inaba T (2007) Arabidopsis Cor15am is a chloroplast stromal protein that has cryoprotective activity and forms oligomers. Plant Physiol 144(1):513–523. doi:10.1104/pp.106.094581
Kovacs D, Kalmar E, Torok Z, Tompa P (2008) Chaperone activity of ERD10 and ERD14, two disordered stress-related plant proteins. Plant Physiol. doi:10.1104/pp.108.118208
Olvera-Carrillo Y, Luis Reyes J, Covarrubias AA (2011) Late embryogenesis abundant proteins: versatile players in the plant adaptation to water limiting environments. Plant Signal Behav 6(4):586–589. doi:10.4161/psb.6.4.15042
Kovacs D, Agoston B, Tompa P (2008) Disordered plant LEA proteins as molecular chaperones. Plant Signal Behav. doi:10.4161/psb.3.9.6434
Thalhammer A, Bryant G, Sulpice R, Hincha DK (2014) Disordered cold regulated15 proteins protect chloroplast membranes during freezing through binding and folding, but do not stabilize chloroplast enzymes in vivo. Plant Physiol 166(1):190–201. doi:10.1104/Pp.114.245399
Szalaine Agoston B, Kovacs D, Tompa P, Perczel A (2011) Full backbone assignment and dynamics of the intrinsically disordered dehydrin ERD14. Biomol NMR Assign 5(2):189–193. doi:10.1007/s12104-011-9297-2
Shih MD, Hsieh TY, Lin TP, Hsing YIC, Hoekstra FA (2010) Characterization of two soybean (Glycine max L.) LEA IV proteins by circular dichroism and Fourier transform infrared spectrometry. Plant Cell Physiol 51(3):395–407. doi:10.1093/Pcp/Pcq005
Ismail AM, Hall AE, Close TJ (1999) Purification and partial characterization of a dehydrin involved in chilling tolerance during seedling emergence of cowpea. Plant Physiol 120(1):237–244. doi:10.1104/Pp.120.1.237
Hara M, Terashima S, Kuboi T (2001) Characterization and cryoprotective activity of cold-responsive dehydrin from Citrus unshiu. J Plant Physiol 158(10):1333–1339. doi:10.1078/0176-1617-00600
Koag MC, Wilkens S, Fenton RD, Resnik J, Vo E, Close TJ (2009) The K-segment of maize DHN1 mediates binding to anionic phospholipid vesicles and concomitant structural changes. Plant Physiol 150(3):1503–1514. doi:10.1104/pp.109.136697
Koag MC, Fenton RD, Wilkens S, Close TJ (2003) The binding of maize DHN1 to lipid vesicles. Gain of structure and lipid specificity. Plant Physiol 131(1):309–316. doi:10.1104/Pp.011171
Soulages JL, Kim K, Arrese EL, Walters C, Cushman JC (2003) Conformation of a group 2 late embryogenesis abundant protein from soybean. Evidence of poly (l-proline)-type II structure. Plant Physiol 131(3):963–975. doi:10.1104/Pp.015891
Rahman LN, McKay F, Giuliani M, Quirk A, Moffatt BA, Harauz G, Dutcher JR (2013) Interactions of Thellungiella salsuginea dehydrins TsDHN-1 and TsDHN-2 with membranes at cold and ambient temperatures-surface morphology and single-molecule force measurements show phase separation, and reveal tertiary and quaternary associations. Biochim Biophys Acta 1828(3):967–980. doi:10.1016/j.bbamem.2012.11.031
Eriksson SK, Kutzer M, Procek J, Grobner G, Harryson P (2011) Tunable membrane binding of the intrinsically disordered dehydrin Lti30, a cold-induced plant stress protein. Plant Cell 23(6):2391–2404. doi:10.1105/Tpc.111.085183
Clarke MW, Boddington KF, Warnica JM, Atkinson J, McKenna S, Madge J, Barker CH, Graether SP (2015) Structural and functional insights into the cryoprotection of membranes by the intrinsically disordered dehydrins. J Biol Chem 290(45):26900–26913. doi:10.1074/jbc.M115.678219
Eriksson S, Eremina N, Barth A, Danielsson J, Harryson P (2016) Membrane-induced folding of the plant stress dehydrin Lti30. Plant Physiol 171(2):932–943. doi:10.1104/pp.15.01531
Dure L (1993) Structural motifs in lea proteins. Struct Motifs Lea Proteins 10:91–103. doi:10.1104/pp.105.072967
Tolleter D, Jaquinod M, Mangavel C, Passirani C, Saulnier P, Manon S, Teyssier E, Payet N, Avelange-Macherel MH, Macherel D (2007) Structure and function of a mitochondrial late embryogenesis abundant protein are revealed by desiccation. Plant Cell 19(5):1580–1589. doi:10.1105/Tpc.107.050104
Hundertmark M, Dimova R, Lengefeld J, Seckler R, Hincha DK (2011) The intrinsically disordered late embryogenesis abundant protein LEA18 from Arabidopsis thaliana modulates membrane stability through binding and folding. BBA Biomembr 1808(1):446–453. doi:10.1016/J.Bbamem.2010.09.010
Manfre AJ, Lanni LM, Marcotte WR (2006) The Arabidopsis group 1 LATE EMBRYOGENESIS ABUNDANT protein AtEM6 is required for normal seed development. Plant Physiol 140(1):140–149. doi:10.1104/Pp.105.072967
Saavedra L, Svensson J, Carballo V, Izmendi D, Welin B, Vidal S (2006) A dehydrin gene in Physcomitrella patens is required for salt and osmotic stress tolerance. Plant J 45(2):237–249. doi:10.1111/J.1365-313x.2005.02603.X
Olvera-Carrillo Y, Campos F, Reyes JL, Garciarrubio A, Covarrubias AA (2010) Functional analysis of the group 4 late embryogenesis abundant proteins reveals their relevance in the adaptive response during water deficit in Arabidopsis. Plant Physiol 154(1):373–390. doi:10.1104/pp.110.158964
Cheng Z, Targolli J, Huang X, Wu R (2002) Wheat LEA genes, PMA80 and PMA1959, enhance dehydration tolerance of transgenic rice (Oryza sativa L.). Mol Breed 10(1):71–82. doi:10.1023/a:1020329401191
Figueras M, Pujal J, Saleh A, Save R, Pages M, Goday A (2004) Maize Rabl7 overexpression in Arabidopsis plants promotes osmotic stress tolerance. Ann Appl Biol 144(3):251–257. doi:10.1111/j.1744-7348.2004.tb00341.x
Xu D, Duan X, Wang B, Hong B, Ho THD, Wu R (1996) Expression of a late embryogenesis abundant protein gene, HVA1, from barley confers tolerance to water deficit and salt stress in transgenic rice. Plant Physiol 110(1):249–257. doi:10.1104/pp.110.1.249
Park B-J, Liu Z, Kanno A, Kameya T (2005) Genetic improvement of Chinese cabbage for salt and drought tolerance by constitutive expression of a B. napus LEA gene. Plant Sci 169(3):553–558. doi:10.1016/j.plantsci.2005.05.008
Liu Y, Wang L, Xing X, Sun L, Pan J, Kong X, Zhang M, Li D (2013) ZmLEA3, a multifunctional group 3 LEA protein from maize (Zea mays L.), is involved in biotic and abiotic stresses. Plant Cell Physiol 54(6):944–959. doi:10.1093/pcp/pct047
Jha B, Lal S, Tiwari V, Yadav SK, Agarwal PK (2012) The SbASR-1 gene cloned from an extreme halophyte Salicornia brachiata enhances salt tolerance in transgenic tobacco. Mar Biotechnol (N Y) 14(6):782–792. doi:10.1007/s10126-012-9442-7
Hu W, Huang C, Deng X, Zhou S, Chen L, Li Y, Wang C, Ma Z, Yuan Q, Wang Y, Cai R, Liang X, Yang G, He G (2013) TaASR1, a transcription factor gene in wheat, confers drought stress tolerance in transgenic tobacco. Plant Cell Environ 36(8):1449–1464. doi:10.1111/pce.12074
Liu J, Jia C, Dong F, Wang J, Zhang J, Xu Y, Xu B, Jin Z (2013) Isolation of an abscisic acid senescence and ripening inducible gene from litchi and functional characterization under water stress. Planta 237(4):1025–1036. doi:10.1007/s00425-012-1820-x
Jeanneau M, Gerentes D, Foueillassar X, Zivy M, Vidal J, Toppan A, Perez P (2002) Improvement of drought tolerance in maize: towards the functional validation of the Zm-Asr1 gene and increase of water use efficiency by over-expressing C4-PEPC. Biochimie 84(11):1127–1135
Kalifa Y, Gilad A, Konrad Z, Zaccai M, Scolnik PA, Bar-Zvi D (2004) The water- and salt-stress-regulated Asr1 (abscisic acid stress ripening) gene encodes a zinc-dependent DNA-binding protein. Biochem J 381(Pt 2):373–378. doi:10.1042/bj20031800
Yang C-Y, Chen Y-C, Jauh GY, Wang C-S (2005) A lily ASR protein involves abscisic acid signaling and confers drought and salt resistance in Arabidopsis. Plant Physiol 139(2):836–846. doi:10.1104/pp.105.065458
Hara M, Shinoda Y, Kubo M, Kashima D, Takahashi I, Kato T, Horiike T, Kuboi T (2011) Biochemical characterization of the Arabidopsis KS-type dehydrin protein, whose gene expression is constitutively abundant rather than stress dependent. Acta Physiol Plant 33(6):2103–2116. doi:10.1007/s11738-011-0749-1
Graether SP, Boddington KF (2014) Disorder and function: a review of the dehydrin protein family. Front Plant Sci 5:576. doi:10.3389/fpls.2014.00576
Liu G, Xu H, Zhang L, Zheng Y (2011) Fe binding properties of two soybean (Glycine max L.) LEA4 proteins associated with antioxidant activity. Plant Cell Physiol 52(6):994–1002. doi:10.1093/pcp/pcr052
Plana M, Itarte E, Eritja R, Goday A, Pages M, Martinez MC (1991) Phosphorylation of maize RAB-17 protein by casein kinase 2. J Biol Chem 266(33):22510–22514
Heyen BJ, Alsheikh MK, Smith EA, Torvik CF, Seals DF, Randall SK (2002) The calcium-binding activity of a vacuole-associated, dehydrin-like protein is regulated by phosphorylation. Plant Physiol 130(2):675–687. doi:10.1104/pp.002550
Hara M, Fujinaga M, Kuboi T (2005) Metal binding by citrus dehydrin with histidine-rich domains. J Exp Bot 56(420):2695–2703. doi:10.1093/Jxb/Eri262
Hara M, Kondo M, Kato T (2013) A KS-type dehydrin and its related domains reduce Cu-promoted radical generation and the histidine residues contribute to the radical-reducing activities. J Exp Bot 64(6):1615–1624. doi:10.1093/jxb/ert016
Kruger C, Berkowitz O, Stephan UW, Hell R (2002) A metal-binding member of the late embryogenesis abundant protein family transports iron in the phloem of Ricinus communis L. J Biol Chem 277(28):25062–25069. doi:10.1074/Jbc.M201896200
Alsheikh MK, Heyen BJ, Randall SK (2003) Ion binding properties of the dehydrin ERD14 are dependent upon phosphorylation. J Biol Chem 278(42):40882–40889. doi:10.1074/jbc.M307151200
Alsheikh MK, Svensson JT, Randall SK (2005) Phosphorylation regulated ion-binding is a property shared by the acidic subclass dehydrins. Plant Cell Environ 28(9):1114–1122. doi:10.1111/J.1365-3040.2005.01348.X
Xu J, Zhang YX, Wei W, Han L, Guan ZQ, Wang Z, Chai TY (2008) BjDHNs confer heavy-metal tolerance in plants. Mol Biotechnol 38(2):91–98. doi:10.1007/s12033-007-9005-8
Hara M, Shinoda Y, Tanaka Y, Kuboi T (2009) DNA binding of citrus dehydrin promoted by zinc ion. Plant Cell Environ 32(5):532–541. doi:10.1111/J.1365-3040.2009.01947.X
Sasaki K, Christov NK, Tsuda S, Imai R (2014) Identification of a novel LEA protein involved in freezing tolerance in wheat. Plant Cell Physiol 55(1):136–147. doi:10.1093/pcp/pct164
Rosales R, Romero I, Escribano MI, Merodio C, Sanchez-Ballesta MT (2014) The crucial role of Φ- and K-segments in the in vitro functionality of Vitis vinifera dehydrin DHN1a. Phytochemistry 108:17–25. doi:10.1016/j.phytochem.2014.10.006
Ricardi MM, Gonzalez RM, Zhong S, Dominguez PG, Duffy T, Turjanski PG, Salgado Salter JD, Alleva K, Carrari F, Giovannoni JJ, Estevez JM, Iusem ND (2014) Genome-wide data (ChIP-seq) enabled identification of cell wall-related and aquaporin genes as targets of tomato ASR1, a drought stress-responsive transcription factor. BMC Plant Biol 14:29. doi:10.1186/1471-2229-14-29
Hayes MA, Davies C, Dry IB (2007) Isolation, functional characterization, and expression analysis of grapevine (Vitis vinifera L.) hexose transporters: differential roles in sink and sourcetissues. J Exp Bot 58(8):1985–1997. doi:10.1093/jxb/erm061
Virlouvet L, Jacquemot MP, Gerentes D, Corti H, Bouton S, Gilard F, Valot B, Trouverie J, Tcherkez G, Falque M, Damerval C, Rogowsky P, Perez P, Noctor G, Zivy M, Coursol S (2011) The ZmASR1 protein influences branched-chain amino acid biosynthesis and maintains kernel yield in maize under water-limited conditions. Plant Physiol 157(2):917–936. doi:10.1104/pp.111.176818
Frankel N, Nunes-Nesi A, Balbo I, Mazuch J, Centeno D, Iusem ND, Fernie AR, Carrari F (2007) ci21A/Asr1 expression influences glucose accumulation in potato tubers. Plant Mol Biol 63(5):719–730. doi:10.1007/s11103-006-9120-0
Brini F, Hanin M, Lumbreras V, Irar S, Pages M, Masmoudi K (2007) Functional characterization of DHN-5, a dehydrin showing a differential phosphorylation pattern in two Tunisian durum wheat (Triticum durum Desf.) varieties with marked differences in salt and drought tolerance. Plant Sci 172(1):20–28. doi:10.1016/j.plantsci.2006.07.011
Bonhomme L, Valot B, Tardieu F, Zivy M (2012) Phosphoproteome dynamics upon changes in plant water status reveal early events associated with rapid growth adjustment in maize leaves. Mol Cell Proteomics 11(10):957–972. doi:10.1074/mcp.M111.015867
Mayank P, Grossman J, Wuest S, Boisson-Dernier A, Roschitzki B, Nanni P, Nuhse T, Grossniklaus U (2012) Characterization of the phosphoproteome of mature Arabidopsis pollen. Plant J 72(1):89–101. doi:10.1111/j.1365-313X.2012.05061.x
Yao Q, Bollinger C, Gao J, Xu D, Thelen JJ (2012) P(3)DB: an integrated database for plant protein phosphorylation. Front Plant Sci 3:206. doi:10.3389/fpls.2012.00206
Jiang X, Wang Y (2004) Beta-elimination coupled with tandem mass spectrometry for the identification of in vivo and in vitro phosphorylation sites in maize dehydrin DHN1 protein. Biochemistry 43(49):15567–15576. doi:10.1021/bi0483965
Röhrig H, Schmidt J, Colby T, Brautigam A, Hufnagel P, Bartels D (2006) Desiccation of the resurrection plant Craterostigma plantagineum induces dynamic changes in protein phosphorylation. Plant Cell Environ 29(8):1606–1617. doi:10.1111/j.1365-3040.2006.01537.x
Gao J, Xu D (2012) Correlation between posttranslational modification and intrinsic disorder in protein. Pac Symp Biocomput. doi:10.1142/9789814366496_0010
Riera M, Figueras M, López C, Goday A, Pagès M (2004) Protein kinase CK2 modulates developmental functions of the abscisic acid responsive protein Rab17 from maize. Proc Natl Acad Sci USA 101(26):9879–9884. doi:10.1073/pnas.0306154101
Goday A, Jensen AB, Culianez-Macia FA, Mar Alba M, Figueras M, Serratosa J, Torrent M, Pages M (1994) The maize abscisic acid-responsive protein Rab17 is located in the nucleus and interacts with nuclear localization signals. Plant cell 6(3):351–360. doi:10.1105/tpc.6.3.351
Hara M (2010) The multifunctionality of dehydrins: an overview. Plant Signal Behav 5(5):503–508. doi:10.4161/psb.11085
Candat A, Paszkiewicz G, Neveu M, Gautier R, Logan DC, Avelange-Macherel MH, Macherel D (2014) The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 26(7):3148–3166. doi:10.1105/tpc.114.127316
Dominguez PG, Carrari F (2015) ASR1 transcription factor and its role in metabolism. Plant Signal Behav 10(4):e992751. doi:10.4161/15592324.2014.992751
Gao J, Lan T (2016) Functional characterization of the late embryogenesis abundant (LEA) protein gene family from Pinus tabuliformis (Pinaceae) in Escherichia coli. Sci Rep 6:19467. doi:10.1038/srep19467
Tompa P, Szasz C, Buday L (2005) Structural disorder throws new light on moonlighting. Trends Biochem Sci 30(9):484–489. doi:10.1016/j.tibs.2005.07.008
Basha E, O’Neill H, Vierling E (2012) Small heat shock proteins and alpha-crystallins: dynamic proteins with flexible functions. Trends Biochem Sci 37(3):106–117. doi:10.1016/j.tibs.2011.11.005
Waters ER (2013) The evolution, function, structure, and expression of the plant sHSPs. J Exp Bot 64(2):391–403. doi:10.1093/jxb/ers355
Jaya N, Garcia V, Vierling E (2009) Substrate binding site flexibility of the small heat shock protein molecular chaperones. Proc Natl Acad Sci USA 106(37):15604–15609. doi:10.1073/pnas.0902177106
Graham D, Patterson BD (1982) Responses of plants to low, nonfreezing temperatures: proteins, metabolism, and acclimation. Annu Rev Plant Physiol 33(1):347–372. doi:10.1146/annurev.pp.33.060182.002023
Mangeon A, Junqueira RM, Sachetto-Martins G (2010) Functional diversity of the plant glycine-rich proteins superfamily. Plant Signal Behav 5(2):99–104. doi:10.4161/psb.5.2.10336
Khan F, Sultana T, Deeba F, Naqvi S (2013) Dynamics of mRNA of glycine-rich RNA-binding protein during wounding, cold and salt stresses in Nicotiana tabacum. Pak J Bot 45:297–300
Kim JY, Kim WY, Kwak KJ, Oh SH, Han YS, Kang H (2010) Glycine-rich RNA-binding proteins are functionally conserved in Arabidopsis thaliana and Oryza sativa during cold adaptation process. J Exp Bot 61(9):2317–2325. doi:10.1093/jxb/erq058
Kwak KJ, Park SJ, Han JH, Kim MK, Oh SH, Han YS, Kang H (2011) Structural determinants crucial to the RNA chaperone activity of glycine-rich RNA-binding proteins 4 and 7 in Arabidopsis thaliana during the cold adaptation process. J Exp Bot 62(11):4003–4011. doi:10.1093/jxb/err101
Freire MA (2012) The Zea mays glycine-rich RNA-binding protein MA16 is bound to a ribonucleotide(s) by a stable linkage. J Plant Res 125(5):653–660. doi:10.1007/s10265-012-0476-8
Lorkovic ZJ (2009) Role of plant RNA-binding proteins in development, stress response and genome organization. Trends Plant Sci 14(4):229–236. doi:10.1016/j.tplants.2009.01.007
Heintzen C, Nater M, Apel K, Staiger D (1997) AtGRP7, a nuclear RNA-binding protein as a component of a circadian-regulated negative feedback loop in Arabidopsis thaliana. Proc Natl Acad Sci 94(16):8515–8520. doi:10.1073/pnas.94.16.8515
Staiger D, Zecca L, Wieczorek Kirk DA, Apel K, Eckstein L (2003) The circadian clock regulated RNA-binding protein AtGRP7 autoregulates its expression by influencing alternative splicing of its own pre-mRNA. Plant J 33(2):361–371. doi:10.1046/j.1365-313X.2003.01629.x
Streitner C, Danisman S, Wehrle F, Schoning JC, Alfano JR, Staiger D (2008) The small glycine-rich RNA binding protein AtGRP7 promotes floral transition in Arabidopsis thaliana. Plant J 56(2):239–250. doi:10.1111/j.1365-313X.2008.03591.x
Kim JS, Park SJ, Kwak KJ, Kim YO, Kim JY, Song J, Jang B, Jung CH, Kang H (2007) Cold shock domain proteins and glycine-rich RNA-binding proteins from Arabidopsis thaliana can promote the cold adaptation process in Escherichia coli. Nucleic Acids Res 35(2):506–516. doi:10.1093/nar/gkl1076
Streitner C, Koster T, Simpson CG, Shaw P, Danisman S, Brown JW, Staiger D (2012) An hnRNP-like RNA-binding protein affects alternative splicing by in vivo interaction with transcripts in Arabidopsis thaliana. Nucleic Acids Res 40(22):11240–11255. doi:10.1093/nar/gks873
Kim JS, Jung HJ, Lee HJ, Kim KA, Goh CH, Woo Y, Oh SH, Han YS, Kang H (2008) Glycine-rich RNA-binding protein 7 affects abiotic stress responses by regulating stomata opening and closing in Arabidopsis thaliana. Plant J 55(3):455–466. doi:10.1111/j.1365-313X.2008.03518.x
Khan F, Daniels MA, Folkers GE, Boelens R, Saqlan Naqvi SM, van Ingen H (2014) Structural basis of nucleic acid binding by Nicotiana tabacum glycine-rich RNA-binding protein: implications for its RNA chaperone function. Nucleic Acids Res 42(13):8705–8718. doi:10.1093/nar/gku468
McCain DC, Croxdale J, Markley JL (1989) Thermal damage to chloroplast envelope membranes. Plant Physiol 90(2):606–609. doi:10.1104/pp.90.2.606
Dekov I, Tsonev T, Yordanov I (2000) Effects of water stress and high-temperature stress on the structure and activity of photosynthetic apparatus of Zea mays and Helianthus annuus. Photosynthetica 38(3):361–366. doi:10.1023/a:1010961218145
Vothknecht UC, Otters S, Hennig R, Schneider D (2012) Vipp1: a very important protein in plastids? J Exp Bot 63(4):1699–1712. doi:10.1093/jxb/err357
Zhang L, Sakamoto W (2013) Possible function of VIPP1 in thylakoids: protection but not formation? Plant Signal Behav 8(2):e22860. doi:10.4161/psb.22860
Zhang L, Sakamoto W (2015) Possible function of VIPP1 in maintaining chloroplast membranes. Biochim Biophys Acta 1847(9):831–837. doi:10.1016/j.bbabio.2015.02.013
Zhang L, Kato Y, Otters S, Vothknecht UC, Sakamoto W (2012) Essential role of VIPP1 in chloroplast envelope maintenance in Arabidopsis. Plant Cell 24(9):3695–3707. doi:10.1105/tpc.112.103606
Kroll D, Meierhoff K, Bechtold N, Kinoshita M, Westphal S, Vothknecht UC, Soll J, Westhoff P (2001) VIPP1, a nuclear gene of Arabidopsis thaliana essential for thylakoid membrane formation. Proc Natl Acad Sci USA 98(7):4238–4242. doi:10.1073/pnas.061500998
Flores-Kim J, Darwin AJ (2016) The phage shock protein response. Annu Rev Microbiol 70:83–101. doi:10.1146/annurev-micro-102215-095359
Hankamer BD, Elderkin SL, Buck M, Nield J (2004) Organization of the AAA+ adaptor protein PspA is an oligomeric ring. J Biol Chem 279(10):8862–8866. doi:10.1074/jbc.M307889200
Lenn T, Gkekas CN, Bernard L, Engl C, Jovanovic G, Buck M, Ying L (2011) Measuring the stoichiometry of functional PspA complexes in living bacterial cells by single molecule photobleaching. Chem Commun 47(1):400–402. doi:10.1039/C0CC01707H
Drin G, Casella J-F, Gautier R, Boehmer T, Schwartz TU, Antonny B (2007) A general amphipathic [alpha]-helical motif for sensing membrane curvature. Nat Struct Mol Biol 14(2):138–146. doi:10.1038/nsmb1194
Drin G, Antonny B (2010) Amphipathic helices and membrane curvature. FEBS Lett 584(9):1840–1847. doi:10.1016/j.febslet.2009.10.022
McDonald C, Jovanovic G, Wallace BA, Ces O, Buck M (2016) Structure and function of PspA and Vipp1 N-terminal peptides: insights into the membrane stress sensing and mitigation. Biochim Biophys Acta 1859(1):28–39. doi:10.1016/j.bbamem.2016.10.018
Zhang L, Kondo H, Kamikubo H (2016) VIPP1 has a disordered C-terminal tail necessary for protecting photosynthetic membranes against stress. Plant Physiol 171(3):1983–1995. doi:10.1104/pp.16.00532
Sato H, Mizoi J, Tanaka H, Maruyama K, Qin F, Osakabe Y, Morimoto K, Ohori T, Kusakabe K, Nagata M, Shinozaki K, Yamaguchi-Shinozaki K (2014) Arabidopsis DPB3-1, a DREB2A interactor, specifically enhances heat stress-induced gene expression by forming a heat stress-specific transcriptional complex with NF-Y subunits. Plant Cell 26(12):4954–4973. doi:10.1105/tpc.114.132928
Blomberg J, Aguilar X, Brännström K, Rautio L, Olofsson A, Wittung-Stafshede P, Björklund S (2012) Interactions between DNA, transcriptional regulator Dreb2a and the Med25 mediator subunit from Arabidopsis thaliana involve conformational changes. Nucleic Acids Res 40(13):5938–5950. doi:10.1093/nar/gks265
Vainonen JP, Jaspers P, Wrzaczek M, Lamminmaki A, Reddy RA, Vaahtera L, Brosche M, Kangasjarvi J (2012) RCD1-DREB2A interaction in leaf senescence and stress responses in Arabidopsis thaliana. Biochem J 442:573–581. doi:10.1042/BJ20111739
O’Shea C, Staby L, Bendsen SK, Tidemand FG, Redsted A, Willemoës M, Kragelund BB, Skriver K (2016) Structures and short linear motif of disordered transcription factor regions provide clues to the interactome of the cellular hub radical-induced cell death1. J Biol Chem. doi:10.1074/jbc.M116.753426
Che P, Bussell JD, Zhou W, Estavillo GM, Pogson BJ, Smith SM (2010) Signaling from the endoplasmic reticulum activates brassinosteroid signaling and promotes acclimation to stress in Arabidopsis. Sci Signal 3(141):ra69. doi:10.1126/scisignal.2001140
Gao H, Brandizzi F, Benning C, Larkin RM (2008) A membrane-tethered transcription factor defines a branch of the heat stress response in Arabidopsis thaliana. Proc Natl Acad Sci USA 105(42):16398–16403. doi:10.1073/pnas.0808463105
Urade R (2009) The endoplasmic reticulum stress signaling pathways in plants. BioFactors 35(4):326–331. doi:10.1002/biof.45
Howell SH (2013) Endoplasmic reticulum stress responses in plants. Annu Rev Plant Biol 64:477–499. doi:10.1146/annurev-arplant-050312-120053
Deng Y, Humbert S, Liu JX, Srivastava R, Rothstein SJ, Howell SH (2011) Heat induces the splicing by IRE1 of a mRNA encoding a transcription factor involved in the unfolded protein response in Arabidopsis. Proc Natl Acad Sci USA 108(17):7247–7252. doi:10.1073/pnas.1102117108
Srivastava R, Deng Y, Shah S, Rao AG, Howell SH (2013) BINDING PROTEIN is a master regulator of the endoplasmic reticulum stress sensor/transducer bZIP28 in Arabidopsis. Plant Cell 25(4):1416–1429. doi:10.1105/tpc.113.110684
Srivastava R, Deng Y, Howell SH (2014) Stress sensing in plants by an ER stress sensor/transducer, bZIP28. Front Plant Sci 5:59. doi:10.3389/fpls.2014.00059
Dangl JL, Jones JD (2001) Plant pathogens and integrated defence responses to infection. Nature 411(6839):826–833. doi:10.1038/35081161
Delteil A, Gobbato E, Cayrol B, Estevan J, Michel-Romiti C, Dievart A, Kroj T, Morel JB (2016) Several wall-associated kinases participate positively and negatively in basal defense against rice blast fungus. BMC Plant Biol 16:17. doi:10.1186/s12870-016-0711-x
Rasmussen MW, Roux M, Petersen M, Mundy J (2012) MAP kinase cascades in Arabidopsis innate immunity. Front Plant Sci 3:169. doi:10.3389/fpls.2012.00169
Shiu SH, Bleecker AB (2001) Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc Natl Acad Sci USA 98(19):10763–10768. doi:10.1073/Pnas.181141598
Kemmerling B, Schwedt A, Rodriguez P, Mazzotta S, Frank M, Qamar SA, Mengiste T, Betsuyaku S, Parker JE, Mussig C, Thomma BP, Albrecht C, de Vries SC, Hirt H, Nurnberger T (2007) The BRI1-associated kinase 1, BAK1, has a brassinolide-independent role in plant cell-death control. Curr Biol 17(13):1116–1122. doi:10.1016/j.cub.2007.05.046
Ogasawara Y, Kaya H, Hiraoka G, Yumoto F, Kimura S, Kadota Y, Hishinuma H, Senzaki E, Yamagoe S, Nagata K, Nara M, Suzuki K, Tanokura M, Kuchitsu K (2008) Synergistic activation of the Arabidopsis NADPH oxidase AtrbohD by Ca2+ and phosphorylation. J Biol Chem 283(14):8885–8892. doi:10.1074/jbc.M708106200
Oda T, Hashimoto H, Kuwabara N, Akashi S, Hayashi K, Kojima C, Wong HL, Kawasaki T, Shimamoto K, Sato M, Shimizu T (2010) Structure of the N-terminal regulatory domain of a plant NADPH oxidase and its functional implications. J Biol Chem 285(2):1435–1445. doi:10.1074/jbc.M109.058909
Sangster TA, Queitsch C (2005) The HSP90 chaperone complex, an emerging force in plant development and phenotypic plasticity. Curr Opin Plant Biol 8(1):86–92. doi:10.1016/j.pbi.2004.11.012
Shirasu K, Schulze-Lefert P (2003) Complex formation, promiscuity and multi-functionality: protein interactions in disease-resistance pathways. Trends Plant Sci 8(6):252–258. doi:10.1016/S1360-1385(03)00104-3
Zhang MH, Kadota Y, Prodromou C, Shirasu K, Pearl LH (2010) Structural basis for assembly of Hsp90-Sgt1-CHORD protein complexes: implications for chaperoning of NLR innate immunity receptors. Mol Cell 39(2):269–281. doi:10.1016/j.molcel.2010.05.010
Kong Q, Qu N, Gao MH, Zhang ZB, Ding XJ, Yang F, Li YZ, Dong OX, Chen S, Li X, Zhang YL (2012) The MEKK1-MKK1/MKK2-MPK4 kinase cascade negatively regulates immunity mediated by a mitogen-activated protein kinase kinase kinase in Arabidopsis. Plant Cell 24(5):2225–2236. doi:10.1105/tpc.112.097253
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444(7117):323–329. doi:10.1038/nature05286
Segrest JP, De Loof H, Dohlman JG, Brouillette CG, Anantharamaiah GM (1990) Amphipathic helix motif: classes and properties. Protein Struct Funct Bioinform 8(2):103–117. doi:10.1002/prot.340080202
Ellis J (1987) Proteins as molecular chaperones. Nature 328(6129):378–379. doi:10.1038/328378a0
Hendrick JP, Hartl FU (1995) The role of molecular chaperones in protein folding. FASEB J 9(15):1559–1569
Schunemann D (2004) Structure and function of the chloroplast signal recognition particle. Curr Genet 44(6):295–304. doi:10.1007/s00294-003-0450-z
Rosenblad MA, Samuelsson T (2004) Identification of chloroplast signal recognition particle RNA genes. Plant Cell Physiol 45(11):1633–1639. doi:10.1093/pcp/pch185
Weikl TR, Paul F (2014) Conformational selection in protein binding and function. Protein Sci 23(11):1508–1518. doi:10.1002/pro.2539
Csermely P, Palotai R, Nussinov R (2010) Induced fit, conformational selection and independent dynamic segments: an extended view of binding events. Trends Biochem Sci 35(10):539–546. doi:10.1016/j.tibs.2010.04.009
Cushman JC (2001) Crassulacean acid metabolism. A plastic photosynthetic adaptation to arid environments. Plant Physiol 127(4):1439–1448. doi:10.1104/pp.010818
Wright PE, Dyson HJ (1999) Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J Mol Biol 293(2):321–331. doi:10.1006/Jmbi.1999.3110
Wright PE, Dyson HJ (2009) Linking folding and binding. Curr Opin Struct Biol 19(1):31–38. doi:10.1016/j.sbi.2008.12.003
Tompa P, Fuxreiter M (2008) Fuzzy complexes: polymorphism and structural disorder in protein–protein interactions. Trends Biochem Sci 33(1):2–8. doi:10.1016/J.Tibs.2007.10.003
Janson J-C (2011) Protein purification: principles, high resolution methods, and applications, 3rd edn. Wiley, New York. doi:10.1002/9780470939932
Oldfield CJ, Cheng YG, Cortese MS, Romero P, Uversky VN, Dunker AK (2005) Coupled folding and binding with alpha-helix-forming molecular recognition elements. Biochemistry 44(37):12454–12470. doi:10.1021/Bi050736e
Roberts EH (1973) Predicting the storage life of seeds. Seed Sci Technol 1:499–514
McDonald C, Jovanovic G, Ces O, Buck M (2015) Membrane stored curvature elastic stress modulates recruitment of maintenance proteins PspA and Vipp1. MBio 6(5):e01188–e01467. doi:10.1128/mBio.01188-15
Vogt L, Vinyard DJ, Khan S, Brudvig GW (2015) Oxygen-evolving complex of Photosystem II: an analysis of second-shell residues and hydrogen-bonding networks. Curr Opin Chem Biol 25:152–158. doi:10.1016/j.cbpa.2014.12.040
Wang M, Kaufman RJ (2014) The impact of the endoplasmic reticulum protein-folding environment on cancer development. Nat Rev Cancer 14(9):581–597. doi:10.1038/nrc3800
Acknowledgements
We thank Jose L. Reyes for critical reviewing this manuscript and for his kindness in helping us with the design and drawing of the figures in this manuscript. This work was partially supported by a Grant from CONACyT-México (221448). CLC-V and CC are supported by post-doctoral fellowships from CONACyT (221448) and Newton Fund-Mexican Academy of Sciences-CONACyT, respectively, and PSR-P and DFR-L by CONACyT PhD fellowships.
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
Amphipathic helix A type of α-helix in which one face along the long axis of the helix is polar and the opposing face is nonpolar [284].
Chaperone A type of protein that ensures the proper folding of nascent polypeptidic chains by binding to them, stabilizing one structural conformation and releasing from the folded protein [285, 286].
Chloroplast signal recognition particle (CSRP) A protein complex formed by cpSRP43 and cpSRP54 involved in the transport and insertion of proteins of the LHC into the membrane of thylakoids [287, 288].
Conformational selection A protein binding mechanism in which the binding protein structure shifts among different conformations and the ligand “selects” one such conformation, stabilizing the structure of the binding protein [289, 290].
Crassulacean acid metabolism (CAM) A photosynthetic pathway in which CO2 is “stored” in the form of malate in the vacuole during the night and released during the day for RuBisCO-mediated carbon fixation [291].
Folding-upon-binding A mechanism in which IDPs or IDRs fold upon binding to their targets [292, 293].
Fuzzy complex A mechanism in which the parts of the proteins involved in the formation of macromolecular complexes do not fold, but remain in a disordered state [294].
Hydrodynamic radius The radius required for a perfect solid sphere to achieve the same frictional coefficient as the protein analyzed [295].
Hydrophilins Any protein that has a high glycine and/or small amino acid residues content (>6%) and a high hydrophilicity index (>1.0) [148, 167].
Intrinsically disordered protein (IDP) These proteins lack a fixed three-dimensional structure. These proteins fail to form a stable conformation and cannot be adequately described by a single equilibrium 3D structure, yet they exhibit biological activity [7, 296].
Intrinsically disordered region (IDR) Significant disordered sequences contained in a protein with structured globular domains [7].
Light-harvesting complex (LHC) In green plants, these complexes contain multiple proteins and molecules such as chlorophylls a and b and carotenoids, which increase the capacity for photon capture during the photosynthesis [1].
Molecular recognition feature (MORF) Protein structural element or feature that mediates the binding events of disordered regions with other proteins or nucleic acids. This element undergoes coupled binding and folding within a disorder region [296].
Moonlight activity The ability of a protein to fulfill several, apparently unrelated functions [229].
Orthodox seed Seeds that acquire desiccation tolerance during development maintain dormancy and might be stored under dry environments [297].
Store curvature elasticity stress Physical torque stress that occurs within a bilayer when lipids in the constituent monolayers are forced to adopt an unfavorable packing conformation. This stress results in hydrophobic cavities in the membrane, which are known as lipid-packing defects [298].
Oxygen-evolving complex (OEC) A Mn4O5Ca cluster embedded in the Photosystem II (PSII) protein complex. It is considered the active site of water oxidation where O2 is produced and protons are released into the lumen during the light reactions of photosynthesis [299].
Unfolded protein response (UPR) Signaling pathways triggered to maintain a productive endoplasmic reticulum protein-folding environment [300].
Rights and permissions
About this article
Cite this article
Covarrubias, A.A., Cuevas-Velazquez, C.L., Romero-Pérez, P.S. et al. Structural disorder in plant proteins: where plasticity meets sessility. Cell. Mol. Life Sci. 74, 3119–3147 (2017). https://doi.org/10.1007/s00018-017-2557-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-017-2557-2