Abstract
Our planet and its inhabitants have been shaped by micro organisms for over 3.5 billion years. Various studies have been carried out on interactions between these microbes and plants but traditionally emphasized on pathogenic interactions. After numerous depictions and attempts to evaluate the incident and their impersonation to boost microbial diversity linked with flora, it is pretended that somewhat a dinky portion of plant-interacting microbes are pathogenic or harmful comparable to the beneficial one. Mostly microbes inhabiting plant-related alcoves have impartial or advantageous roles in plant buildup. Soil microbiome symbolizes the immense reservoir of biological diversity acknowledged in the world so far. In the present chapter, we counsel the perception that for sustainable development of agriculture we have to understand the mechanisms by which plants interact with their microbiome. This interaction may directly affect the plant health and development which ultimately leads to the organization of novel microbiome-driven strategies. In addition, for the benefit of agriculture and food production, we also discussthe potential for root microbiome modification. So, in the above-mentioned context this article will help the researchers to uncover the critical areas of the microbiome that many researchers were not able to explore earlier.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
15.1 Introduction
Microbes are considered a prime requisite for the prolongation of life on Earth, yet we figure out the little bit about the plurality of beneficial microbes present in the environments such as soils, oceans, in the atmosphere and even those living on and in our own bodies (Turner et al. 2013). Nowadays, understanding the plant-associated microbial communities and their impact on plant health and development is gaining much interest for sustainable agriculture (Beirn et al. 2017). Soil is referred to as the greatest reservoir of microbial diversity consisting of both beneficial and harmful microbes and plant roots are in a close association to these microbial communities (Gams 2007; Bue’e et al. 2009; Berendsen et al. 2012). This narrow soil zone around the plant root is known as rhizosphere and known as the most active zone for soil microbial activities. Root microbiome or microbial community associated with the plant root is also mentioned as the plant’s second genome since composite genome size of the communities of microbes is much larger compared to the plant genome (Lareen et al. 2016). The structure of the rhizosphere communities of microbes is influenced by root exudates in a very coordinated way. However, regarding this connection between microbiome assemblage, root exudates, and plant development our knowledge is still limited (Chaparro et al. 2014).
A blooming recognition of biodiversity and its function in combination with advances in data analytics technologies and omics is needed for sustaining the brisk advances in microbiome exploration and research (Srivastava et al. 2015). One propulsive motivation regarding harnessing beneficial microbes and reducing impacts of detrimental microbes in both humans and crop plants is nearly common (Busby et al. 2017). Joshua Lederberg was the first who antecedently used the word “microbiome” and defined it as the “ecological community of commensal microorganisms, symbionts or pathogens, which literally occupy a space in our body” (Lederberg and McCray 2001). Recently, in the presence of specific sets of collective microbes, it was found that most of the features of the host have been linked with the human body which contributes as a great reservoir of microbes proved during the Human Microbiome Project (Turnbaugh et al. 2006; Djikeng et al. 2011; The Human Microbiome Project Consortium 2012). Just a while ago, the utilization of this idiom has been widely tested with the contrasting sets of microbes found in particular hosts or populates of present surroundings (Boon et al. 2014; Ofek et al. 2014). Boon et al. (2014) projected that the most excellent ever description of “microbiome” would relate to the set of genes encountered in partnership of a specific environment with the host, thus showing the less influence of the link among taxonomy and range of capabilities of the microbial community members. The conceptual importance of the microbiome to plants is made doable only by using detected presumptions, which carry out various imperative activities in association with significant microbial faction. This issue has been addressed by some authors, viz. Bulgarelli et al. (2013), Turner et al. (2013), and Rout (2014).
In the present chapter, special emphasis has been put on the characterization of phyto-microbiomes by not only depicting the sub-divisions where the microbes live (phyllosphere, endosphere, and rhizosphere) but also by discussing the need of interactions among plants and microorganisms (Srivastava et al. 2011). The efforts related to microbiome will improve the prognosis of ecosystem response and will assist the progress of the development of new, robust, microbiome-based solutions against significant challenges of our time. So, in the present article, the perception is that for more sustainable development of agriculture we have to understand the mechanisms by which plants interact with the beneficial microbiome.
15.2 Plant-Associated Microbial Communities in Cropping System
Being represented as the most biodiverse ecosystem on Earth, soil characteristics in the form of matrix and texture are crucial for the soil to serve as the reservoir of microbes that interact with flora and fauna of the particular surroundings (Vogel et al. 2009). The soil microbiome is culpable for their signature series of actions to achieve a significant result which are directly related to plant health occurring under this environment (Attwood et al. 2019; Compant et al. 2019). For instance, the soil microbiome has the capability to suppress various phytopathogenic diseases by impressing some restrictions on physiological functions of the related pathogens vital in infecting and colonizing plant tissues (Weller et al. 2002; Mendes et al. 2011). Evenly, the soil microbiomes also pass on a certain degree of resistance to the system against “invaders,” thereby connecting the microbiome diversity to its peculiar ability to limit the situation or prohibit the survival and dissemination of exogenic microorganisms (van Elsas et al. 2012).
Astounding potential has been shown by plant-associated microbiomes during the improvement of plant yields in farming/cropping systems (Lyu et al. 2020). There are many pieces of evidence which prove that the biological technologies using microbes or their metabolites are beneficial in the enhancement of crop yield by nutrient uptake, by managing pests/pathogens and also by mitigating the plant abiotic stress responses. However, for utter realization of the technological potential of microbes, their consistency and efficacy under the wide-ranging extent of the real-world environment need to be upgraded. While the use of biopesticides and biofertilizers is rapidly expanding to cover a wide range of soils, crop varieties, and environments, crop breeding programs have yet to incorporate the selection of beneficial plant-microbe interactions in order to breed “microbe-optimized plants.” Developing attempts to delve into microbiome engineering could lead to microbial consortia that are exceptionally adapted to hold the plants. The combination of all the three approaches should be unified for the enormous benefits and also to boost agricultural production significantly (Trivedi et al. 2017).
Undoubtedly, development of some traditional agricultural practices has not so far been able to meet the future requirements because they are neither economical nor environmentally feasible. So, there is an urgent need for complimentary feasible approaches to meet the global food demands of the booming population. Another way to develop advanced and improved sustainable crop production methods is to embellish the beneficial plant-associated microbiome having the potential to increase crop growth and vigor, biotic/abiotic stress tolerance, nutrient use efficiency, and disease resistance (Fig. 15.1). If this potentiality of microbes could be harnessed under real-world conditions, it will be very effective in the improvement of farm productivity and produce quality sustainably framing healthy environmental, social, and robust economic outcomes (Trivedi et al. 2017). Because microbe-based formulations can raise the crop yields and also prove promising to or replace, reduce, or at least complement the chemical source of pesticides and fertilizers. Nowadays, many agro-chemical companies have initiated attempts to explore individual microorganisms which can be used as biocontrol or biofertilizer products and also to develop carrier-based inoculants of beneficial strains. An increase of 10–20% in the production of economically important crops has resulted in large-scale field trials (Pérez-Jaramillo et al. 2016).
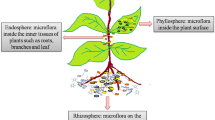
Considering the microbiome an effective constituent of the host and also responsible for biotic and abiotic changes in the environmental conditions, it is important to understand the composition of the most important diverse plant microbiomes in a better way. Recent studies are more focused to understand the major phylogenetic and functional factors that can influence microbial assembly. The plant microbiome can be separated into three major components, i.e., rhizosphere, endosphere, and phyllosphere (Hardoim et al. 2008; Hirsch and Mauchline 2012) (Fig. 15.2). The rhizospheric zone of soil contributes as a most active narrow zone for soil microbial activities beneficial for plant metabolism (Bulgarelli et al. 2013; Philippot et al. 2013). Endosphere is composed of microbial habitats in the plant system the members of which inhabit inner tissues of the plant asymptomatically (Hallmann et al. 1997; Hardoim et al. 2008), and the phyllosphere is inhabited by the microbes capable to colonize plant surfaces (Lindow and Brandl 2003; Lambais et al. 2006).
15.3 Plant Selection for the Rhizosphere Microbial Communities
The soil region under the influence of the roots is known as the rhizosphere zone of a plant (Hiltner 1904; Hartmann et al. 2008; Philippot et al. 2013). Root exudates determine soil microflora by releasing some selective growth substrates thereby selectively influencing the expansion of fungi and bacteria that colonize the rhizosphere zone of a plant by altering the soil chemistry of the environs. In return, microbes also influence the quantity, components, and composition of miscellaneous root exudates by affecting cell metabolism, root cell leakage, and plant nutrition. Microbial communities of the rhizosphere zone can vary on the basis of differences in rhizo-deposition and exudation in structure and species composition of specific root zones at different locations or soil type, plant species, nutritional status, age, and stress, be it biotic or abiotic (De Leij et al. 1994; Mahaffee and Kloepper 1997; Lupwayi et al. 1998; Griffiths et al. 1999). Some root exudates secreted during the growth of new roots in the elongation zone just following the root tips are beneficial for the growth of colonizers of primary root that can efficiently utilize organic acids and degradable sugars. But in the older root zones, primarily carbon is put in place of safe-keeping as sloughed cells and consisting of more recalcitrant materials, including lignified hemicellulose and cellulose so that bacteria and fungi in these older root zones are probably adapted for oligotrophic conditions. Other nutritionally specific zones include the sites of lateral root emergence and non-growing root tips, distinctly known as nutrient-rich environments colonized by mature communities of microbes (Yang and Crowley 2000). In nutshell, taking all these above-mentioned effects together, plants have evolved to grant the rhizosphere zone to attract specific microbes necessary for the growth and development of plants (Mendes et al. 2011; Prashar et al. 2014).
15.4 Endosphere—A Niche for Intimate Friends
Diverse communities of various microbes consisting of bacteria, archaebacteria, fungi, and other protistic taxa spending some parts of their life cycle by living inside the plants are technically known as endophytes (Hardoim et al. 2015; Pavithra et al. 2020). Further, over the coming years the concept of endophyte is likely to change and expectedly will evolve by some researchers realizing that plant tissues could be colonized by bacteria as much as fungi living inside plants (Hallmann et al. 1997). This concept of endophytes proves that plants are firmly associated with the microbes present in their surroundings particularly with those living inside the plants rather than living alone as lone entities. Recently emerged concept of the “plant microbiome,” i.e., the composite genomes of microbes living associated with plants has paved the way for the stimulation of new ideas regarding the evolution of plants where selective forces do not act solely on the plant genome itself but rather on the entire plant, including its microbial community. Hologenome approach for vertical transmission of beneficial traits provided by endophytes to plants is the best example used to explain the Lamarckian concepts of acquired heritable characters (Rosenberg et al. 2009).
While the observation of the microbial cells presents inside the plant tissues, De Bary (1866) was the first person who described the presence of other non-pathogenic organisms inside plants. This observation was explored at the end of the last century until after the emergence of the endophytic concept. Universally, the concept behind endophytes is based on the capacity to identify the microbial cells from formerly surface-sterilized plant tissues (Hallmann et al. 1997); in nutshell this proves that these microorganisms are not epiphytes. A functional definition for endophytes was also provided by Petrini (1991), as “Organisms which colonize plant tissues internally while spending some part of their life cycle without causing any apparent harm.” Endophytic communities have been divided into two sub-groups, viz. “obligate” and “facultative” by some authors (Das et al. 2021) (Ref., if any, please). Hardoim et al. (2008) classify the obligate endophytes as those which depend on plant metabolism for their survival, being spread among plants by the activity of vectors or by vertical transmission. The endophytes living outside the host during a certain stage of their life cycle are known as facultative endophytes, which are recruited by the plant from adjacent communities in the form of bulk soil, mainly through the rhizosphere zone (Andreote et al. 2014; Harman and Uphoff 2019).
In nutshell, the endophytic microbes play crucial roles in plant growth, development, and diversification. The increased awareness and information related to endophytes provide intuitiveness into the complexity of the plant microbiome. The basic nature of plant–endophyte interactions ranges from mutualism to pathogenicity depending on the set of biotic and abiotic factors including environmental conditions, microbes, and plant genotypes along with their dynamic connection of interactions within the plant biome. Accordingly, the latest insights into evolution, plant ecosystem functioning, and multipartite interactions are fulfilled by the concept of endophytism (Hardoim et al. 2015).
15.5 Microbial Groups Living in the Phyllosphere
A third component of the plant microbiome, i.e., the phyllosphere is made up of microbes colonizing the aerial plant tissues although this term can be used for any extrinsic plant surfaces (Vorholt 2012). The phyllosphere contributed as an enormous environment on Earth that is supposed to be an area of 6.4 × 108 km2 fully colonized by microbes (Morris and Kinkel 2002).
Our knowledge regarding the phyllosphere microbiology or the microbiome of aerial parts of plants has historically lagged comparative to the rhizosphere or the below-ground habitat of plants, particularly concerning fundamental investigations such as which microbes are prevalent and what activities do they perform there. However, for the enhancement in this regard recently floated cultivation-independent studies have revealed that a few bacterial phyla predominate in the aerial parts of several distinct plants and the plant factors which are involved in shaping these phyllosphere communities, feature specific adaptations, and exhibit multipartite relationships both with host plants and among community members. Awareness regarding the structural principles of indigenous microbial phyllosphere populations will help us develop an understanding of the phyllosphere microbiota also showing advantageous promotion of plant growth and protection.
Many fungi whether they are filamentous or yeasts-like, bacteria, algae, and at lower densities some protozoa and nematodes are comprised by the phyllosphere community (Lindow and Brandl 2003). Out of all, the bacterial community is the major group presented in the phyllosphere the numbers being between 105 and 107 cells/cm2 (Andrews and Harris 2000). Microflora of phyllosphere is characterized to thrive well in an oligotrophic environment, i.e., live under harsh environmental conditions where there is an unavailability of nutrients and under fluctuating conditions of humidity, presence of ultraviolet radiations along with a wide range of temperature and pH (Andrews and Harris 2000).
The microbial communities of the phyllosphere performing an efficient role in the processes related to plant growth, for example, by fixation of nitrogen, biosynthesizing phytohormones along with the protection of plants facing biotic stresses (Jones 1970; Freiberg 1998; Brandl et al. 2001; Kishore et al. 2005). These above-mentioned activities make them the main contributor of many global processes such as the sequestration of carbon (Bulgarelli et al. 2013), and they can also be potentially used for the sustainable development of agriculture. Thus, microbes present in the phyllospheric region play a promising role to offer the eco-friendly protection of plants (Lindow and Brandl 2003).
15.6 Microbial Interaction Across the Cropping Systems
Farming/cropping systems are broadly grouped into (1) those which use chemical or synthetic means of pest control and nutrient fertilization (a.k.a. conventional), and (2) those which favor an integrated system with the goal of sustainability (a.k.a. organic). Within each system, a number of management techniques may be used which collectively alter the above-ground and below-ground biodiversity, including chemical use, fertilization, irrigation, crop rotation or crop-fallow rotations, co-cropping, and livestock grazing. Several studies broadly comparing organic and conventional systems have shown differences in crop production, competition by weeds, pests, or microbial pathogens (Pollnac et al. 2009). Notably, organic farming, and often the increased soil organic matter associated with organic farms, is selected for a higher overall microbial diversity (Flohre et al. 2011; Chaudhry et al. 2012; Pershina et al. 2015; Hartmann et al. 2015; Ishaq et al. 2016).
15.7 Soil Fertilization
Soil fertilization utilizes organic matter (mulching) or chemical supplementation to add nutrients back into the soil. Long-term use of mineral fertilizers has been shown to increase bacterial and fungal diversity, microbial biomass carbon, as well as dehydrogenase and another enzyme activity (Luo et al. 2015). However, these benefits are variable depending on the type and source of minerals. Using only mineral nitrogen (typically ammonium sulfate) does not increase soil microbial diversity (Ramirez et al. 2010; Luo et al. 2015; Zhalnina et al. 2015) and may even reduce it (Campbell et al. 2010). Phosphorous-only supplementation has a similar lack of effect (Zhalnina et al. 2015) except where it was limiting (Su et al. 2015). This reduction may be driven by a shift toward more acidic soil which tends to reduce total microbial diversity and shift toward acid-tolerant species, such as within the bacterial phylum Acidobacteria (Lauber et al. 2009; Rousk et al. 2010; Fierer et al. 2013; Zhalnina et al. 2015). It may also be a function of the relative type and amount of plant residues (Roesch et al. 2007), or a change in nutrient availability and the C:N ratio in soil (Ramirez et al. 2010; Zhalnina et al. 2015).
Animal manure has been shown to be significantly more effective at increasing microbial biomass than mineral fertilization (Hartmann et al. 2015; Luo et al. 015). Integrated livestock grazing has recently re-emerged as an alternative method of crop-residue removal, specifically in organic systems (McKenzie et al. 2016). Its implementation has been slow, especially in large production systems, as the use of grazing livestock can be time- and labor-intensive. Inputs of feces and urine from livestock grazing increases soil organic carbon and nitrogen (Liu et al. 2015), as well as total microbial biomass (Liu et al. 2012, 2015). However, this may only be reflected in bacterial biomass and not an increase in fungal biomass (Taddese et al. 2007). In systems where grazing pressure is high, this effect can be reversed as soil nutrients are lost to erosion caused by a lack of plant cover material (Mofidi et al. 2012; Chen et al. 2015; Liu et al. 2015).
15.8 Cover Crops
Cover crops are grown as an alternative to fallowing or leaving a field unplanted to rest. They provide additional economic benefit (Adusumilli and Fromme 2016; Duzy et al. 2016), feed for livestock (Sulc and Franzluebbers 2014), reduce erosion, and facilitate weed and insect pest management (Dabney et al. 2001; McKenzie et al. 2016; Duzy et al. 2016). Specifically, cover crops can reduce weed seed production via competitive exclusion (Gallandt et al. 1998), or survivability of weed seeds gets decreased by recruiting a microbial community which contributes to seed decay (Dabney et al. 1996; Liebman and Davis 2000). Mineralization of cover crop residues causes an increment in the organic matter of soil (Reeves 1994; Hartwig and Ammon 2002), which can increase cation exchange capacity, and enhance the cycling of macronutrients (Kamh et al. 1999).
Not only do the additional inputs of organic matter from cover crop residues encourage microbial diversity, but they also allow the above-ground biomass to generate more below-ground biomass (Wild 1993; Reeves 1994; Hu et al. 1999; Hartwig and Ammon 2002; Snapp et al. 2004). Crop rotations can also improve soil quality and microbial diversity (Ghimire et al. 2014). The use of legumes as a cover crop or in rotation, or other crops which encourage rhizobial symbiotic bacteria to biologically fix nitrogen, and the subsequent mineralization of those nitrogen-rich plant residues back into the soil can provide usable available nitrogen for other plant species (Snapp et al. 2004; Biederbeck et al. 2005). For example, bacterial liter increased most in response to clover (Trifolium repens L.) conditioning compared with wheat (Triticum aestivum L.), ryegrass (Lolium perenne L.), bentgrass (Agrostis capillaris L.), or sucrose conditioning (Grayston et al. 1998). Additionally, microbial communities differed strongly among the four cover crop conditioning species (Grayston et al. 1998).
15.9 Tillage
In both organic and conventional systems, tillage is the most common method of incorporating crop residues back into soil, as well as redistributing weed seeds either further into soil to prevent germination or onto the surface where they may be eaten. Due to the disruptive nature of tillage in the first 30–50 cm of topsoil, significant detriment can be done by physically destroying mycorrhizal root colonization (McGonigle et al. 1990). Moreover, soil microbial diversity and density is highly correlative to soil depth and local factors (e.g., oxygen content, UV light, moisture). Thus, intensive soil tillage can drastically decrease soil microbial diversity and density, specifically bacterial and fungal, through erosion and wind dispersion of microorganisms or nutrients, or through selective culling of sub-surface species brought to the surface (Lupwayi et al. 1998; Castillo et al. 2006; De Quadros et al. 2012; Mathew et al. 2012; Fierer et al. 2013; García-Orenes et al. 2013; Ghimire et al. 2014). However, the addition of soil organic matter through mulching may attenuate some of these adverse effects (García-Orenes et al. 2013; Ghimire et al. 2014). No-till systems typically have more soil carbon (Brevik 2013).
15.10 Chemical Control and Bioremediation of Farmland
Chemical control used for managing agricultural systems has been shown to alter the microbial community, notably in decreasing diversity (elFantroussi et al. 1999; Lupwayi et al. 2004; Lo 2010). However, the persistence of pesticides and other chemical contaminants in soil is also of concern for biological systems in natural and agricultural settings, not only because they may accrue and affect other beneficial organisms and soil health indicators, but many contain heavy metals which are toxic (Hussain et al. 2009). Additionally, the local water sources and runoff may add contaminants from exogenous sources. Phyto, microbial, or combined bioremediation of chemical contamination has been sought to degrade or detoxify pesticides (i.e., herbicides, insecticides, fungicides, rodenticides), heavy metals, and antibiotics.
For the above purpose, bacteria belonging to the genera Acinetobacter, Alcaligenes, Arthrobacter, Bacillus, Burkholderia, Corynebacterium, Flavobacterium, Micrococcus, Mycobacterium, Pseudomonas, Sphingomonas, and Rhodococcus, and the fungus Phanerochaetechrysosporium are just a few of the microorganisms shown to degrade different types of hydrocarbons from petroleum spills (Kuhad et al. 2004; Hussain et al. 2007a, b, 2009; Das and Chandran 2011). The degradation of chemicals, the sequestration of heavy metals, or the detoxification of heavy metal compounds by microorganisms is dependent on the nature of the compound, as well as on the ambient conditions of the environment (Kuhad et al. 2004; Singh 2008). Endosulfan degradation depends on soil type and oxygen content (Kumar and Philip 2006a, b), as well as on soil texture, organic matter content, inoculum concentration, pH, and specificity of bacterial strains used (Hussain et al. 2007a). Similarly, dichlorodiphenyltrichloroethane (DDT), metoxychlor, and gamma-hexachlorocyclohexane (gamma-HCH) degradation processes are dependent on temperature (Baczynski et al. 2010). HCH degradation was also shown to be dependent on oxygen content and nitrate concentration (Langenhoff et al. 2002). An additional nutrient source, such as molasses, is often needed to increase the rate of chemical degradation in culture (Lamichhane et al. 2012; Hussain et al. 2014).
Field trials have been focused on removing chemical and metal contamination from soil or water runoff, either using direct application of microorganisms or by the use of a “biobed” as a biological filter or retaining system to remove contaminants from farm wastewater (Antonious 2012). The bacterium Mycobacterium gilvum was successfully used to degrade polycyclic aromatic hydrocarbons, and increase soil bacterial diversity, on a vegetable farm (Ma et al. 2018). A strain of Arthrobacter and another of Bacillus were used to reduce metal contamination in soil, improve rice biomass production, and reduce the amount of metal accumulated in rice (Du et al. 2016). Halophilic bacteria were used to remove the salt left behind after the March 2011 tsunami in Japan, as well as green compost to restore organic matter that had been washed away (Azizul and Omine 2013). Furthermore, bacteria that are able to mitigate salt-stress in plants can promote growth into similarly affected areas (Cao et al. 2008; Nabti et al. 2015).
The concept of remediating soil diversity toward a “more natural” community has been slower to take root. A study of pre-agricultural prairie soil reported a very different bacterial community than that found in the human-associated agricultural soil (Fierer et al. 2013). Notably, prairie soils were dominated by the bacterial phylum Verrucomicrobia, whereas agricultural soil shows a dominance of Proteobacteria, Bacteroidetes, or Firmicutes (Lauber et al. 2009; Ishaq et al. 2016). Verrucomicrobia grow more slowly, but survive better in nutrient-limiting soils. Likewise, Acidobacteria are also known to survive under nutrient-limiting (oligotrophic) conditions (Fierer et al. 2012; Koyama et al. 2014; Greening et al. 2015; Kielak et al. 2016). Moreover, Verrucomicrobia from pre-agricultural soil contained more genes for carbohydrate metabolism than nitrogen metabolism (Fierer et al. 2013), suggesting that their abundance in agricultural soil may be negatively selected for by the use of nitrogen fertilizer. And, as Proteobacteria produce the quorum-sensing molecule AHL which triggers beneficial and pathogenic responses from bacteria, selecting for these species under agricultural conditions may be contributing to plant disease dynamics.
15.11 Understanding and Exploiting Plant Beneficial Microbes
We all know about the link between the trillions of microorganisms that exist inside the body related to our health. Earlier studies have already proved that some sort of depressions and food allergies has been prevented by the activities of some healthy microbiome present inside the body. These types of miniatures also perform a favorable role in plant growth and development similar to what they do inside the human body. Various methods are adopted for the addition of growth-promoting bacteria or fungi by using a variety of ways like an addition inside the seed coats, suspended in water and sprinkled on plant or soil surfaces, or mixed into mulches that are added to the soil or placed around plant stems which will be beneficial for the plant growth and development with a robust yield (Ishaq 2017; Srivastava et al. 2020). Nowadays, this microscopic world attracts more attention of researchers looking for some new cost-effective eco-friendly techniques for the betterment of agriculture production (Srivastava et al. 2019).
15.12 Nitrogen Fixation
The entire range of flora present around the natural ecosystem is closely associated with microbes, including bacteria and fungi (Finkel et al. 2017). The symbiotic relationship between the plants and microbes has been proved by evidence of fossils since the last four million years. To cope with the numerous challenges of the environment, plants rely on microbes for growth since their evolution. For example, various growth-promoting microorganisms such as fungi and bacteria can fix nitrogen from the air and then make it available to plants for growth and development. Most of these microbes inhabit the surface of roots often inside special structures known as root nodules (Gage 2004).
The first formal description of the enhancive role of soil microbes on agricultural plant growth was given by Lorenz Hiltner in 1904, though agricultural treatments anticipated to customize the microbial activities present in the soil have been utilized since Roman times. For example, the Romans used to plant alfalfa and clover that form close associations with nitrogen-fixing bacteria during the cultural practices like crop rotations which were helpful in improving soil fertility. They didn’t exactly know the scientific reason of this practice but now we do know that these plants can elevate the nitrogen content of the soil.
There are also enormous fungal species that are able to build a symbiotic association with the roots of different higher plants by forming some structures known as mycorrhizae (Denison and Kiers 2011). Mycorrhizae spread into the root system of the plant in association and the fungal mycelium intermingles or passes through the plant cells/tissues with the help of some specific structures like Hartig net, vesicles, and arbuscules. by which nutrient uptake or exchanges occur for enhancing plant growth and development. By using these specific structures fungi enable the plant to scavenge water from the surrounding soil and also allow the plant to exchange sugars for nutrients. Microbes which are able to establish a mutually beneficial partnership by forming specialized structures with the plants are scientifically recognized as symbiotes (Skriabin 1923) (Table 15.1).
15.13 Balancing Action of Lodgepole Pine
Lodgepole pine is a perfect plant host example of beneficial non-symbiotic or free-living bacteria and fungi (Beirn et al. 2017) which can live in the soil, plant roots, on plant surfaces, or even within the plant tissues. Beneficial free-living microbes, especially endophytes that live within plant tissues, whether they are acting as symbiotes or not, are able to suppress the phytopathogens along with other harmful organisms thereby affecting the plant metabolism and health. Additionally, endophytes can also facilitate plant growth directly by regulating plant hormones, activating plant immune responses, and also by providing supplementary nutrients to the plant (Timmusk et al. 2017).
Despite the fact that endophytes have shown enormous potential needed for the betterment of the plant health, discovering some non-symbiotic microbes that usually produce significant positive growth responses under extremely variable field conditions still remain a challenging task. Addition of any beneficial microbe that can improve the growth of plants may not constantly be sufficient because some other group of microorganisms present in the surroundings as a part of the plant’s environment will also be affecting the interaction between the beneficial microbe and the plant. For example, a bacterium being necessary for the growth of lodgepole pine can be thwarted by the occurrence of another contending bacterium (Bent et al. 2001). According to the study this effect was shown by both, i.e., by endophytic plant-beneficial bacterium as well as in case of those simply living on the root surface of the plant.
15.14 Microbial Cocktail
The sole meaning of the term is that if we isolate a phyto-beneficial microbe under a protected environment, then it is not certain to be compatible under field conditions unless it is scientifically tested exhaustively. During deciding the specific conditions necessary for the optimum activities of plant-beneficial microorganisms it may also be mandatory to consider the complete composition of the microbiome to which the particular flora is supposed to be exposed either it is soil, root, or plant surface. Exploring the microbes that can be exploited in agriculture for crop production is a prime quest having a long debate (Schloter et al. 2018). During the study about these arduous interactions of plants-microbiomes along with the favorable environmental conditions, we also have to reveal that in what way or manner these age-old traditional know-how and practices can be refined and utilized in the future.
15.15 Advancement Required for Improving Microbiome in Future
Treasured information regarding the genome desperately augments our knowledge related to the diversity of microbial metabolic pathways used to access the novel and innovative traits (Trivedi et al. 2017). These recent discoveries of novel genes evolved some genetically engineered plants for disease resistance, stress, and herbicide tolerance, and last but the ultimate is for crop improvement program (Macdonald and Singh 2014). Nevertheless, most of these breakthroughs were brought with the successful conclusion by inserting some minor or along with the combo of many targeted genes (multiplexing). Forthcoming and imminent researches have to focus on integrating the distinct plan of action, for example, more than one gene of importance simultaneously be incorporated in transgenic plants by using the multigenic approach. Other than this, we have to apply new tools and resources to initiate intricate heterologous pathways into plants (Shih et al. 2016) which possess the key to frame the useful clusters of synthetic genomes from microbiomes, enable the shuffling and stacking of stress-tolerance and disease resistance traits between the crop plants. Further intensification of the rate of novel gene discovery will be fueled up by using the novel efficiency developed in the trait discovery. For example, forward genetic screening based on the CRISPR–Cas9 will be helpful in the future learning of plant–microbiome interactions to surpass particular genes and evolve as a best holistic strategy while explaining the process behind the plant–microbiome interactions along with the uncovering of novel genes needed for biotechnological applications and innovations (Barakate and Stephens 2016).
The integration of microbe-optimized crops for distinct types of soil, microbial biofertilizers, optimized microbiomes, biocontrol microbes, and soil amendments would be the ultimate purpose of action behind the enhancement of plant–microbe interactions. Undoubtedly, being principally untapped, this area should be entitled to considerable research attempts which will prove promising to address the issue of food security by improving crop yields in a sustainable and eco-friendly manner. Globally, the emerging microbiome along with existing microbial technologies and correlated overtures offer advance and most sustainable methods for use to enhance agriculture productivity. Furthermore, if the existing scientific and technological challenges in this area can be planned out along with advanced work strategies (e.g., product registration, safety requirements) emerging microbial-based solutions can potentially reconstruct the field of agriculture sustainably. Above all, it is universally proven that the aforementioned approach can enable us to achieve the multiple sustainable development goals (SDGs) if put into action in a truly systematic way (Table 15.2).
15.16 Conclusions
Integration of the beneficial plant–microbe and microbiome interactions is the need of the hour which may prove as a promising sustainable solution for the enhancement of our agricultural production. Holistic ecological studies and reductionist mechanistic discoveries both form a beneficial tactic for the study of plant microbiomes during their interactions. Both schools of thought are substantiating reflective awareness into the ecological operations that take command over plant–microbe interactions likewise the specific molecular mechanisms work behind them. The induction of enormous microbial isolates and of synthetic microbial communities if blended with genetic resources of plants will significantly grant us to tide over the chasm lying in between and to conduct reductionist hypothesis-driven studies in increasingly complex ecological contexts up to field tests. These significant advances will contribute to the next green revolution by potentially revolutionizing knowledge regarding the interactions of plants and microbes occurring in natural ecosystem being utilized in agriculture. This chapter notifies the role of microbial diversity that can be beneficial for flora and will help the researchers to uncover the critical areas of microbiome that many researchers had not been able to explore earlier.
References
Adusumilli N, Fromme D (2016) Evaluating benefits and costs of cover crops in cotton production systems in Northwestern Louisiana. Southern Agricultural Economics Association, 2016 Annual Meeting
Andreote FD, Gumiere T, Durrer A (2014) Exploring interactions of plant microbiomes. Sci Agric 71(6):528–539
Andrews JH, Harris RF (2000) The ecology and biogeography of microorganisms on plant surfaces. Annu Rev Phytopathol 38:145–180
Antonious GF (2012) On-farm bioremediation of dimethazone and trifluralin residues in runoff water from an agricultural field. J Environ Sci Heal Part B 47:608–621
Attwood GT, Wakelin SA, Leahy SC, Rowe S, Clarke S, Chapman DF, Muirhead R, Jacobs JME (2019) Applications of the soil, plant and rumen microbiomes in pastoral agriculture. Front Nutr 6(107):1–17
Azizul M, Omine K (2013) Bioremediation of agricultural land damaged by tsunami. In: Chamy R (ed) Biodegradation of hazardous special products. IntechOpen Limited, London, pp 39–50
Baczynski TP, Pleissner D, Grotenhuis T (2010) Anaerobic biodegradation of organochlorine pesticides in contaminated soil-significance of temperature and availability. Chemosphere 78:22–28
Barakate A, Stephens J (2016) An overview of CRISPR-based tools and their improvements: new opportunities in understanding plant–pathogen interactions for better crop protection. Front Plant Sci 7:765
Beirn LA, Hempfling JW, Schmid CJ, Murphy JA, Clarke BB, Crouch JA (2017) Differences among soil-inhabiting microbial communities in Poa annua Turf throughout the growing season. Crop Sci 57:S-262–S-273
Bent E, Tuzun S, Chanway CP, Enebak S (2001) Alterations in plant growth and in root hormone levels of lodgepole pines inoculated with rhizobacteria. Can J Microbiol 47(9):793–800
Berendsen RL, Pieterse CMJ, Bakker PAHM (2012) The rhizosphere microbiome and plant health. Trends Plant Sci 17(8):478–486
Biederbeck VO, Zentner RP, Campbell CA (2005) Soil microbial populations and activities as influenced by legume green fallow in a semiarid climate. Soil Biol Biochem 37:1775–1784
Boon E, Meehan CJ, Whidden C, Wong DHJ, Langille MGI, Beiko RG (2014) Interactions in the microbiome: communities of organisms and communities of genes. FEMS Microbiol Rev 38:90–118
Brandl MT, Quinones B, Lindow SE (2001) Heterogeneous transcription of an indoleacetic acid biosynthetic gene in Erwinia herbicola on plant surfaces. Proc Natl Acad Sci U S A 98:3454–3459
Brevik E (2013) The potential impact of climate change on soil properties and processes and corresponding influence on food security. Agriculture 3:398–417
Bue’e M, Reich M, Murat C, Morin E, Nilsson RH, Uroz S, Martin F (2009) Pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184:454–456
Bulgarelli D, Spaepen SS, Themaat EVL, Shulze-Lefert P (2013) Structure and functions of the bacterial microbiota of plants. Ann Rev Plant Biol 64:807–838
Busby PE, Soman C, Wagner MR, Friesen ML, Kremer J, Bennett A, Morsy M, Eisen JA, Leach JE, Dangl JL (2017) Research priorities for harnessing plant microbiomes in sustainable agriculture. PLoS Biol 15(3):e2001793
Campbell BJ, Polson SW, Hanson TE, Mack MC, Schuur EAG (2010) The effect of nutrient deposition on bacterial communities in arctic tundra soil. Environ Microbiol 12:1842–1854
Cao YR, Chen SY, Zhang JS (2008) Ethylene signaling regulates salt stress response: an overview. Plant Signal Behav 3:761–763
Castillo CG, Rubio R, Rouanet JL, Borie F (2006) Early effects of tillage and crop rotation on arbuscular mycorrhizal fungal propagules in an Ultisol. Biol Fertil Soils 43:83–92
Chaparro JM, Badri DV, Vivanco JM (2014) Rhizosphere microbiome assemblage is affected by plant development. ISME J 8(4):790–803
Chaudhry V, Rehman A, Mishra A, Chauhan PS, Nautiyal CS (2012) Changes in bacterial community structure of agricultural land due to long-term organic and chemical amendments. Microb Ecol 64:450–460
Chen W, Huang D, Liu N, Zhang Y, Badgery WB, Wang X, Shen Y (2015) Improved grazing management may increase soil carbon sequestration in temperate steppe. Sci Rep 5:10892
Compant S, Samad A, Faist H, Sessitsch A (2019) A review on the plant microbiome: ecology, functions, and emerging trends in microbial application. J Adv Res 19:29–37
Dabney SM, Schreiber JD, Rothrock CS, Johnson JR (1996) Cover crops affect sorghum seedling growth. Agron J 88:961
Dabney SM, Delgado JA, Reeves DW (2001) Using winter cover crops to improve soil and water quality. Commun Soil Sci Plant Anal 32:1221–1250
Das N, Chandran P (2011) Microbial degradation of petroleum hydrocarbon contaminants: an overview. Biotechnol Res Int 2011:941810
Das AK, Chosdon T, Nawale R, Koder SB, Bhosale RK, Rana M, Srivastava S (2021) Role of rhizospheric microbiome against soil-borne pathogens. Biopestic Int 17(1):1–15
De Bary A (1866) Morphologie und Physiologie der Pilze, Flechten und Myxomyceten Vol. 2. Leipzig: Hofmeister’s Handbook of Physiological Botany
De Leij FAAM, Whipps JM, Lynch JM (1994) The use of colony development for the characterization of bacterial communities in soil and on roots. MicrobEcol 27:81–97
De Quadros PD, Zhalnina K, Davis-Richardson A, Fagen JR, Drew J, Bayer C, Camargo FAO, Triplett EW (2012) The effect of tillage system and crop rotation on soil microbial diversity and composition in a subtropical acrisol. Diversity 4:375–395
Denison RF, Kiers ET (2011) Life histories of symbiotic rhizobia and mycorrhizal fungi. Curr Biol 21(18):R775–R785
Djikeng A, Nelson BJ, Nelson KE (2011) Implications of the human microbiome research for the developing world. In: Nelson KE (ed) Metagenomics of the human body. Springer, New York, NY, pp 317–336
Du RY, Wen D, Zhao PH, Chen Y, Wang FH (2016) Effect of bacterial application on metal availability and plant growth in farmland-contaminated soils. J Bioremed Biodegr 7:341
Duzy LM, Price AJ, Balkcom KS, Aulakh JS (2016) Assessing the economic impact of inversion tillage, cover crops, and herbicide regimes in Palmer Amaranth (Amaranthus palmeri) infested cotton. Int J Agron 2016:1–9
elFantroussi S, Verschuere L, Verstraete W, Top EM (1999) Effect of phenylurea herbicides on soil microbial communities estimated by analysis of 16S rRNA gene fingerprints and community-level physiological profiles. Appl Environ Microbiol 65:982–988
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6:1007–1017
Fierer N, Ladau J, Clemente JC, Leff JW, Owens SM, Pollard KS, Knight R, Gilbert JA, McCulley RL (2013) Reconstructing the microbial diversity and function of pre-agricultural tallgrass prairie soils in the United States. Science 342:621–624
Finkel OM, Castrillo G, Paredes SH, Gonzalez IS, Dangl JL (2017) Understanding and exploiting plant beneficial microbes. Curr Opin Plant Biol 38:155–163
Flohre A, Rudnick M, Traser G, Tscharntke T, Eggers T (2011) Does soil biota benefit from organic farming in complex vs. simple landscapes? Agric Ecosyst Environ 141:210–214
Freiberg E (1998) Microclimatic parameters influencing nitrogen fixation in the phyllosphere in a costa Rican premontane rain forest. Oecologia 117:9–18
Gage DJ (2004) Infection and invasion of roots by symbiotic, nitrogen-fixing rhizobia during nodulation of temperate legumes. Microbiol Mol Biol Rev 68(2):280–300
Gallandt ER, Liebman M, Huggins DR (1998) Improving soil quality: implications for weed management. J Crop Prod 2:95–121
Gams W (2007) Biodiversity of soil-inhabiting fungi. Biodivers Conserv 16:69–72
García-Orenes F, Morugán-Coronado A, Zornoza R, Cerdà A, Scow K (2013) Changes in soil microbial community structure influenced by agricultural management practices in a mediterraneanagro-ecosystem. PLoS One 8:e80522
Ghimire R, Norton JB, Stahl PD, Norton U (2014) Soil microbial substrate properties and microbial community responses under irrigated organic and reduced-tillage crop and forage production systems. PLoS One 9:e103901
Grayston SJ, Wang SQ, Campbell CD, Edwards AD (1998) Selective influence of plant species on microbial diversity in the rhizosphere. Soil Biol Biochem 30:369–378
Greening C, Carere CR, Rushton-Green R, Harold LK, Hards K, Taylor MC, Morales SE, Stott MB, Cook GM (2015) Persistence of the dominant soil phylum Acidobacteria by trace gas scavenging. Proc Natl Acad Sci U S A 112:10497–10502
Griffiths BS, Ritz K, Ebblewhite N, Dobson G (1999) Soil microbial community structure: effects of substrate loading rates. Soil Biol Biochem 31:145–153
Hallmann J, Quadt-Hallmann A, Mahaffee WF, Kloepper JW (1997) Bacterial endophytes in agricultural crops. Can J Microbiol 43:895–914
Hardoim P, van Overbeek L, van Elsas J (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol 16:463–471
Hardoim PR, van Overbeek LS, Berg G, Pirttilä AM, Compant S, Campisano A, Döring M, Sessitsch A (2015) The hidden world within plants: ecological and evolutionary considerations for defining functioning of microbial endophytes. Microbiol Mol Biol Rev 79(3):293–320
Harman GE, Uphoff N (2019) Symbiotic root-endophytic soil microbes improve crop productivity and provide environmental benefits. Scientifica 2019:9106395. https://doi.org/10.1155/2019/9106395
Hartmann A, Rothballer M, Schmid M (2008) Lorenz Hiltner, a pioneer in rhizosphere microbial ecology and soil bacteriology research. Plant Soil 312:7–14
Hartmann M, Frey B, Mayer J, Mäder P, Widmer F (2015) Distinct soil microbial diversity under long-term organic and conventional farming. ISME J 9:1177–1194
Hartwig NL, Ammon HU (2002) Cover crops and living mulches. Weed Sci 50:688–699
Hiltner L (1904) Uber neuereErfahrungen und Probleme auf dem Gebiete der Bodenbakteriologieunterbesonderdenberucksichtigung und Brache. Arb Dtsch Landwirtsch Gesellschaft 98:59–78
Hirsch PR, Mauchline TH (2012) Who’s who in the plant root microbiome? Nat Biotechnol 30:961–962
Hu SJ, van Bruggen AHC, Grünwald NJ (1999) Dynamics of bacterial populations in relation to carbon availability in a residue-amended soil. Appl Soil Ecol 13:21–30
Hussain S, Arshad M, Saleem M, Khalid A (2007a) Biodegradation of α- and β-endosulfan by soil bacteria. Biodegradation 18:731–740
Hussain S, Arshad M, Saleem M, Zahir ZA (2007b) Screening of soil fungi for in vitro degradation of endosulfan. World J Microbiol Biotechnol 23:939–945
Hussain S, Siddique T, Arshad M, Saleem M (2009) Bioremediation and phytoremediation of pesticides: recent advances. Crit Rev Environ Sci Technol 39:843–907
Hussain A, Iqbal Quzi J, Abdullah Shakir H (2014) Implication of molasses as electron donor for biological sulphate reduction. Am J Environ Eng 4:7–10
Ishaq SL (2017) Plant-microbial interactions in agriculture and the use of farming systems to improve diversity and productivity. AIMS Microbiol 3(2):335–353
Ishaq SL, Johnson SP, Miller ZJ, Lehnhoff EA, Olivo S, Yeoman CJ, Menalled FD (2016) Impact of cropping systems, soil inoculum, and plant species identity on soil bacterial community structure. Microb Ecol 73(2):417–434
Jones K (1970) Nitrogen fixation in phyllosphere of Douglas Fir Pseudotsuga-Douglasii. Ann Bot 34:239–244
Kamh M, Horst WJ, Amer F, Mostafa H, Maier P (1999) Mobilization of soil and fertilizer phosphate by cover crops. Plant Soil 211:19–27
Kielak AM, Barreto CC, Kowalchuk GA, van Veen JA, Kuramae EE (2016) The ecology of Acidobacteria: moving beyond genes and genomes. Front Microbiol 7:744
Kishore GK, Pande S, Podile AR (2005) Biological control of late leaf spot of peanut (Arachis hypogaea) with chitinolytic bacteria. Phytopathology 95:1157–1165
Koyama A, Wallenstein MD, Simpson RT, Moore JC (2014) Soil bacterial community composition altered by increased nutrient availability in Arctic tundra soils. Front Microbiol 5:516
Kuhad RC, Johri AK, Singh A, Ward OP (2004) Bioremediation of pesticide-contaminated soils. In: Singh A, Ward OP (eds) Applied bioremediation and phytoremediation. Springer, Berlin Heidelberg, pp 35–54
Kumar M, Philip L (2006a) Enrichment and isolation of a mixed bacterial culture for complete mineralization of endosulfan. J Environ Sci Heal Part B 41:81–96
Kumar M, Philip L (2006b) Bioremediation of endosulfan contaminated soil and water-optimization of operating conditions in laboratory scale reactors. J Hazard Mater 136:354–364
Lambais MR, Crowley DE, Cury JC, Bull RC, Rodrigues RR (2006) Bacterial diversity in tree canopies of the Atlantic forest. Science 312:1917–1917
Lamichhane KM, Babcock RW Jr, Turnbull SJ, Schenek S (2012) Molasses enhanced phyto and bioremediation reatability study of explosives contaminated Hawaiian soils. J Hazard Mater 243:334–339
Langenhoff AAM, Staps JJM, Pijls C, Rijnaarts HH (2002) Intrinsic and stimulated in situ biodegradation of hexachlorocyclohexane (HCH). Water Air Soil Pollut Focus 2:171–181
Lareen A, Burton F, Schafer P (2016) Plant root-microbe communication in shaping root microbiomes. Plant Mol Biol 90:575–587
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120
Lederberg J, McCray AT (2001) ‘Ome sweet Omics’ - a genealogical treasury of words. Scientist 15:8
Liebman M, Davis AS (2000) Integration of soil, crop and weed management in low-external-input farming systems. Weed Res 40:27–47
Lindow SE, Brandl MT (2003) Microbiology of the phyllosphere. Appl Environ Microbiol 64(4):1875–1883
Liu N, Zhang Y, Chang S, Kan H, Lin L (2012) Impact of grazing on soil carbon and microbial biomass in typical steppe and desert steppe of Inner Mongolia. PLoS One 7:e36434
Liu N, Kan HM, Yang GW, Zhang YJ (2015) Changes in plant, soil, and microbes in a typical steppe from simulated grazing: explaining potential change in soil C. Ecol Monogr 85:269–286
Lo CC (2010) Effect of pesticides on soil microbial community. J Environ Sci Health B 45:348–359
Luo P, Han X, Wang Y, Han M, Shi H, Liu N, Bai H (2015) Influence of long-term fertilization on soil microbial biomass, dehydrogenase activity, and bacterial and fungal community structure in a brown soil of Northeast China. Ann Microbiol 65:533–542
Lupwayi NZ, Rice WA, Clayton GW (1998) Soil microbial diversity and community structure under wheat as influenced by tillage and crop rotation. Soil Biol Biochem 30:1733–1741
Lupwayi NZ, Harker KN, Clayton GW, Turkington TK, Rice WA, O’Donovan JT (2004) Soil microbial biomass and diversity after herbicide application. Can J Plant Sci 84:677–685
Lyu D, Backer R, Subramanian S, Smith DL (2020) Phytomicrobiome coordination signals hold potential for climate change-resilient agriculture. Front Plant Sci 11(634):1–7
Ma L, Deng F, Yang C, Guo C, Dang Z (2018) Bioremediation of PAH-contaminated farmland: field experiment. Environ Sci Pollut Res 25(1):64–72
Macdonald C, Singh B (2014) Harnessing plant-microbe interactions for enhancing farm productivity. Bioengineered 5:5–9
Mahaffee WF, Kloepper JW (1997) Temporal changes in the bacterial communities of soil, rhizosphere, and endorhiza associated with field grown cucumber (Cucumis sativus L.). Microb Ecol 34:210–223
Mathew RP, Feng Y, Githinji L, Ankumah R, Balkcom KS (2012) Impact of no-tillage and conventional tillage systems on soil microbial communities. Appl Environ Soil Sci 2012:1–10
McGonigle TP, Evans DG, Miller MH (1990) Effect of degree of soil disturbance on mycorrhizal colonization and phosphorus absorption by maize in growth chamber and field experiments. New Phytol 116:629–636
McKenzie SC, Goosey HB, O’Neill KM, Menalled FD (2016) Integration of sheep grazing for cover crop termination into market gardens: agronomic consequences of an ecologically based management strategy. Renew Agric Food Syst 74:1–14
Mendes R, Kruijt M, Bruijn I, Dekkers E, van der Voort M, Schneider JHM, Piceno YM, Santis TZ, Andersen GL, Bakker PAHM, Raaijmakers JM (2011) Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332:1097–1100
Mofidi M, Rashtbari M, Abbaspour H, Ebadi A, Sheidai E, Motamedi J (2012) Impact of grazing on chemical, physical and biological properties of soils in the mountain rangelands of Sahand, Iran. Rangel J 34(3):297–303
Morris CE, Kinkel LL (2002) Fifty years of phyllosphere microbiology: significant contributions to research in related fields. In: Lindow SE, Hecht-Poinar EI, Elliott V (eds) Phyllosphere microbiology. APS Press, St. Paul, MN, pp 365–375
Nabti E, Schmid M, Hartmann A (2015) Application of halotolerant bacteria to restore plant growth under salt stress. In: Maheshwari D, Saraf M (eds) . Springer, Halophiles, pp 235–259
Ofek M, Voronov-Goldman M, Hadar Y, Minz D (2014) Host signature effect on plant root-associated microbiomes revealed through analyses of resident vs active communities. Environ Microbiol 16:2157–2167
Pavithra G, Bindal S, Rana M, Srivastava S (2020) Role of endophytic microbes against plant pathogens: a review. Asian J Plant Sci 19(1):54–62
Pérez-Jaramillo JE, Mendes R, Raaijmakers JM (2016) Impact of plant domestication on rhizosphere microbiome assembly and functions. Plant Mol Biol 90:635–644
Pershina E, Valkonen J, Kurki P, Ivanova E, Chirak E, Korvigo I, Provorov N, Andronov E (2015) Comparative analysis of prokaryotic communities associated with organic and conventional farming systems. PLoS One 10(12):e0145072
Petrini O (1991) Fungal endophytes of tree leaves. In: Andrews JH, Hirano SS (eds) Microbial ecology of leaves. Springer-Verlag, New York, NY, pp 179–197
Philippot L, Raaijmakers JM, Lemanceau P, van der Putten WH (2013) Going back to the roots: the microbial ecology of the rhizosphere. Nat Rev Microbiol 11:789–799
Pollnac FW, Maxwell BD, Menalled FD (2009) Using species-area curves to examine weed communities in organic and conventional spring wheat systems. Weed Sci 57:241–247
Prashar P, Kapoor N, Sachdeva S (2014) Rhizosphere: its structure, bacterial diversity and significance. Rev Environ Sci Biotechnol 13:63–77
Ramirez KS, Lauber CL, Knight R, Bradford MA, Fierer N (2010) Consistent effects of nitrogen fertilization on soil bacterial communities in contrasting systems. Ecology 91(12):3463–3470
Reeves DW (1994) Cover crops and rotations. In: Hatfield JL, Stewart BA (eds) Advances in soil science: crop residue management. Lewis Publishers, Boca Raton, FL, pp 125–172
Roesch LFW, Fulthorpe RR, Riva A, Casella G, Hadwin AK, Kent AD, Daroub SH, Camargo FA, Farmerie WG, Triplett EW (2007) Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1(4):283–290
Rosenberg E, Sharon G, Zilber-Rosenberg I (2009) The hologenome theory of evolution contains Lamarckian aspects within a Darwinian framework. Environ Microbiol 11:2959–2962
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG, Knight R, Fierer N (2010) Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4(10):1340–1351
Rout ME (2014) The plant microbiome. Genomes Herbaceous Land Plants 69:279–309
Schloter M, Nannipieri P, Sørensen SJ, van Elsas JD (2018) Microbial indicators for soil quality. Biol Fertil Soils 54:1–10
Shih PM, Vuu K, Mansoori N, Ayad L, Louie KB, Bowen BP, Northen TR, Loqué D (2016) A robust gene-stacking method utilizing yeast assembly for plant synthetic biology. Nat Commun 7:13215
Singh DK (2008) Biodegradation and bioremediation of pesticide in soil: concept, method and recent developments. Indian J Microbiol 48:35–40
Skriabin KI (1923) Simbioziparazitizm v prirode. Petrograd 1923
Snapp SS, Swinton SM, Labarta R, Mutch D, Black JR, Leep R, Nyiraneza J, O’Neil K (2004) Evaluating cover crops for benefits, costs and performance within cropping system niches. Agron J 97(1):322–332
Srivastava S, Singh VP, Kumar R, Srivastava M, Sinha A, Simon S (2011) In vitro evaluation of carbendazim 50% WP, antagonists and botanicals against Fusarium oxysporumf. sp. psidii associated with rhizosphere soil of guava. Asian J Plant Pathol 5(1):46–53
Srivastava S, Singh VP, Rastogi J, Shrivastava SK, Sinha A (2015) Efforts to conserve our microbial biodiversity. In: Sinha A, Srivastava S, Kumar R (eds) Microbial biodiversity: a boon for agriculture sustainability. Biotech Books, New Delhi, pp 1–8. ISBN: 978-81-7622-341-6
Srivastava S, Bindal S, Rana M, Kumar R, Yadav S, Singh JP, Sinha A (2019) Biotechnological role of fungal microbes in sustainable agriculture. Plant Arch 19(1):107–110
Srivastava S, Kumar R, Bindal S, Singh VP, Rana M, Singh JP, Sinha A (2020) Ancient, mid-time, and recent history of seed pathology. In: Kumar R, Gupta A (eds) Seed-borne diseases of agricultural crops: detection, diagnosis & management. Springer, Singapore, pp 81–103
Su JQ, Ding LJ, Xue K, Yao HY, Quensen J, Bai SJ, Wei WX, Wu JS, Zhou J, Tiedje JM, Zhu YG (2015) Long-term balanced fertilization increases the soil microbial functional diversity in a phosphorus-limited paddy soil. Mol Ecol 24(1):136–150
Sulc RM, Franzluebbers AJ (2014) Exploring integrated crop-livestock systems in different ecoregions of the United States. Eur J Agron 57:21–30
Taddese G, Peden D, Hailemariam A, Makonnen YJ (2007) Effect of livestock grazing on soil microorganisms of cracking and self-mulching vertisol. Ethopain Vet J 11(1):141–150
The Human Microbiome Project Consortium (2012) Structure, function and diversity of the healthy human microbiome. Nature 486:207–214
Timmusk S, Behers L, Muthoni J, Muraya A, Aronsson AC (2017) Perspectives and challenges of microbial application for crop improvement. Front Plant Sci 8:49
Trivedi P, Schenk PM, Wallenstein MD, Singh BK (2017) Tiny microbes, big yields: enhancing food crop production with biological solutions. Microb Biotechnol 10(5):999–1003
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444:1027–1103
Turner TR, James EK, Poole PS (2013) The plant microbiome. Genome Biol 14(209):1–10
van Elsas JD, Chiurazzi M, Mallon CA, Elhottova D, Krištufek V, Salles JF (2012) Microbial diversity determines the invasion of soil by a bacterial pathogen. Proc Natl Acad Sci U S A 109:1159–1164
Vogel TM, Simonet P, Jansson JK, Hirsch PR, Tiedje JM, van Elsas JD, Bailey MJ, Nalin R, Philippot L (2009) TerraGenome: a consortium for the sequencing of a soil metagenome. Nat Rev Microbiol 7:252
Vorholt JA (2012) Microbial life in the phyllosphere. Nat Rev Microbiol 10:828–840
Weller DM, Raaijmakers JM, Gardener BBM, Thomashow LS (2002) Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu Rev Phytopathol 40:309–348
Wild A (1993) Soils and the environment. Cambridge University Press, Cambridge
Yang CH, Crowley DE (2000) Rhizosphere microbial community structure in relation to root location and plant iron nutritional status. Appl Environ Microbiol 66(1):345–351
Zhalnina K, Dias R, de Quadros PD, Davis-Richardson A, Camargo FA, Clark IM, McGrath SP, Hirsch PR, Triplett EW (2015) Soil pH determines microbial diversity and composition in the park grass experiment. MicrobEcol 69(2):395–406
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Mehta, C.M. et al. (2022). Understanding the Microbiome Interactions Across the Cropping System. In: Singh, U.B., Rai, J.P., Sharma, A.K. (eds) Re-visiting the Rhizosphere Eco-system for Agricultural Sustainability. Rhizosphere Biology. Springer, Singapore. https://doi.org/10.1007/978-981-19-4101-6_15
Download citation
DOI: https://doi.org/10.1007/978-981-19-4101-6_15
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-19-4100-9
Online ISBN: 978-981-19-4101-6
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)