Abstract
Honeybees are the most important crop pollinators that contribute significantly to agricultural productivity and profitability worldwide. Microbiota accounts for up to 1–10% of the insect’s biomass. The intestine of European Honeybees, Apis mellifera, have diverse microbiota and are known to be occupied by approximately 70% Gram-negative bacteria, 27% Gram-positive bacteria, and 1% yeast. The native microbiota of the honeybees is known to contribute to their nutrition, growth, digestion, pathogens defense, and insecticide resistance. As with other humans and animals, intestinal dysbiosis might greatly influence these insects’ health status posing a threat to their safe existence. Lactic acid bacteria (LAB) have been discovered in abundance in the honeybee gut and are believed to be of great importance to the honeybee health. Among several symbiotic LAB species isolated from the digestive tract of honeybees, it is found that some of them have the potential to be developed as probiotics. One of the most important health benefits of probiotic LAB in honeybees is their ability to protect against several bee pathogens and contribute to honey’s antimicrobial properties. Hence, the use of probiotics in beekeeping could prevent diseases, enhance bee health, and consequently increase honey production. Although probiotic bacteria isolated from different sources could be used for honeybees, using the host bacteria, i.e., the bacteria from the honeybees’ gut microbiome community would be more desirable for their own health. In this review study, we discuss the important aspects related to Apis mellifera gut microbiome such as composition, perturbation, fermentation, and most important of all, the probiotic bacterial community, mainly LAB species residing in the gut of these insects.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
9.1 Introduction
Apis mellifera (A. mellifera) or European honeybee could colonize virtually all habitable biomes on Earth and adapt to diverse bioclimatic conditions. A. mellifera has been classified as a member of the order Hymenoptera and the superfamily Apoidea (Alatawy et al. 2020). They have been divided into six evolutionary lineages including A lineage (Africa), M lineage (western and northern Europe), C lineage (southern and eastern Europe), O lineage (Caucasus, Turkey, Middle East, Cyprus, Crete), Y lineage (Ethiopia), and S lineage (Syria and Lebanon) (Tihelka et al. 2020). As A. mellifera is an essential pollinator species for natural ecosystems and agricultural production, its health status and, consequently, continuous existence is of great importance.
As obvious from numerous research reports, Apis mellifera depends on its gut microbiome to perform its basic functions and survival. Bacterial communities living in symbiosis with their hosts, also known as probiotics, are essential factors in maintaining host health (Zeinali et al. 2020). Additionally, a close association between Honeybee colony productivity and increased bacterial diversity was discovered recently. Hence, an improved understanding of the honeybees’ gut microbiome can help manage modern challenges to these insects’ health and production.
The gut of the honeybee is a continuous tube starting from mouth to anus and demarcated into foregut (stomodeum), midgut (mesenteron), and the hindgut (proctodeum). In many insects, the hindgut is the gut region bearing the largest microbial populations. In particular, the ileum (the region between the proximal pylorus and distal rectum) is a relatively benign environment, in that it lacks the digestive enzymes of the midgut and, for many terrestrial insects, the desiccation stress of the distal hindgut, where water is actively resorbed from the lumen into insect tissues. Microbial function and growth may also be favored by the ions and metabolites delivered to the hindgut in the filtrate from the Malpighian tubules (Huang et al. 2010). On the contrary, due to midgut epithelium actively secreting immunologically active enzymes as well as several antimicrobial peptides, the midgut shows a hostile environment for microorganisms. Besides, the midgut also contains a region of pH < 3 that mediate many microbial cells degradation (Engel and Moran 2013; Shanbhag and Tripathi 2009).
9.2 Gut Microbiome Composition
In the last couple of years, marked deterioration in honeybee hives’ colony health has been reported that has raised worldwide concerns (Meixner 2010). One of the major reasons for such depurations of honeybee colonies is due to the effect of several honeybee diseases (Genersch et al. 2010). In this context, the honeybee gut’s microbial ecosystem has known to play an essential role in maintaining their health and survival. Thus, understanding the microbial community residing in the gut of different honeybee species could lead us to better health management of the bees that would consequently result in enhanced agriculture productivity and human well-being.
The bee gut microbial community is far simpler than the mammalian microbiota and contains a distinctive community of bacterial species. The composition of the gut bacterial communities of these social insect insects has been shaped by coevolution. These insects’ social behavior provides favorable conditions for the exchange of the symbiont microbes, and a number of these microorganisms are efficiently transmitted between bee colony members and their different generations (Engel and Moran 2013).
The composition of microbial communities in the honeybee gut varies enormously within and between species. Honeybees acquire gut microorganisms from the natural environment via foods, such as nectar, pollen, and water. Hence, honeybees’ gut flora varies according to seasonal or geographical differences in food sources, even among individual honeybees from the same colony (Mohr and Tebbe 2006; Moran et al. 2012).
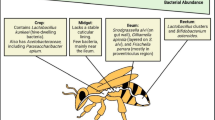
A.mellifera gut microbiota is dominated by only nine bacterial species clusters that are specific to bees and are transmitted through social interactions between individuals (Fig. 9.1). According to available information, there are five main bacterial groups in the honeybees gut including (i) Gram-negative bacteria group (Snodgrassella alvi and Gilliamella apicola), (ii) phylum Proteobacteria to Gram-positive bacteria, Firmicutes (Lactobacillus Firm-4 and Lactobacillus Firm 5 groups), (iii) phylum Actinobacteria (Bifidobacterium asteroids), (iv) a small number of Proteobacteria species (Frischella perrara, Bartonella apis, Parasaccharibacter apium), and (v) Gluconobacter-related species group designated Alpha2.1 (Bottacini et al. 2012; Kwong and Moran 2016).
As seen in Table 9.1, honeybee microbiota occupies distinct metabolic niches in the A. mellifera gut. According to reports, the abundance of Snodgrassella alvi, Frischella perrara, Gilliamella apicola, and Bartonella apis is seen in the ileum while Lactobacillus Firm-4, Lactobacillus Firm-5, as well as Bifidobacterium predominantly reside in the rectum (Kwong and Moran 2016).
Many factors are known to affect the gut microbiota composition and profile of the honeybees, including the age and physiological condition of honeybees. Martinson and Moran in 2012 reported that newly emerged honeybee workers have no or very few gut bacteria, while they uptake bacteria later via contact with the collected honey and bee bread and through trophallactic exchange with nestmates (Martinson et al. 2012). In other research findings, it was stated that larvae’s gut microbial composition differs vastly due to the differences in the bee bread microbial communities (Martinson et al. 2012; Mohr and Tebbe 2006). The observation that certain gut bacteria are maintained in all of the developmental stages of an adult bee, irrespective of differences between species, colonies, and individuals, suggests that distinctive gut bacteria are transferred between generations by eusocial behaviors, such as food exchange between the honeybee populations in a hive (Martinson et al. 2011; Martinson et al. 2012; Vásquez et al. 2012).
Zhi-Xiang Dong and his colleagues, through the use of 16 s rRNA gene sequencing analysis, found that 0-day postemergence (dpe) did not harbor core gut flora in the gut, and the critical points for colonization of the core gut flora were around 1–3 dpe. For example, colonization of Frischella, Gilliamella, and Snodgrassella occurred at 1dpe, while Bifidobacterium, Commensalibacter, and Lactobacillus colonization were significantly detected at 3 dpe (Dong et al. 2020). It is worth mentioning that type of overwintering sugar also influences honeybee gut microbiota. Wang et al. in 2020 via 16 s rRNA sequencing determined bacterial communities in honeybee midguts and hindguts before winter and after bees were fed honey, sucrose, and high-fructose syrup as winter-food. In the midgut, the sucrose group’s microbial diversity was higher than that of the honey and high-fructose syrup groups, but in the hindgut, the microbial diversity of the honey and high-fructose groups was higher than that in the sucrose group. Sucrose increased the relative abundance of Actinobacteria (Bifidobacteriales) and Alphaproteobacteria of honeybee midgut, and honey enriched the Bacteroidetes and Gammaproteobacteria in honeybee hindgut. High-fructose syrup increased the relative abundance of Betaproteobacteria of the midgut. Furthermore, they introduce sucrose as an appropriate overwintering food for honeybee. In this manner, the relative abundance of the dominant microbiota significantly altered with the different sugar types and seasons (Wang et al. 2020).
In another research report, Christina Geldert et al. investigated the effect of phytochemical supplementations on the microbiome diversity and abundance of A. mellifera. They disclosed that phytochemical supplementations are able to enhance gut microbial diversity and significantly increase the abundance of the most represented bacterial genera such as Snodgrassella spp. and Lactobacillus spp. (Geldert et al. 2020).
9.3 Gut Microbiome Perturbation
Exposure to an environmental stressor, including antibiotics as well as the herbicide, is one of the major sources of perturbation to the microbiome that has a detrimental effect on A. mellifera health. Antibiotic treatment of bee colonies has been widely used for over 50 years to prevent bee larvae’s bacterial disease. Some of the most frequently used antibiotics by beekeepers include tetracycline, fumagillin, and tylosin (Genersch et al. 2010). However, these chemical drugs are known to have many side effects, including the emergence of antibiotic resistance. Hence, more attention has been paid to evaluate the effect of antibiotics on the survival and growth of honeybees to identify and pinpoint the main disadvantages of these drugs on honeybee and their environment’s health. Raymann et al., in 2017, assessed the relationship between tetracycline exposure and the size and composition of honeybee gut communities. Their results showed that treatment with tetracycline greatly influenced both the honeybee gut microbiome’s composition and size. According to their observations, tetracycline induced dysbiosis in these insects, which resulted in increased susceptibility to opportunistic pathogens and subsequently led to a significant reduction in bee survival rate (Raymann et al. 2017).
Apart from antibiotics, some agrochemicals such as herbicides can perturb the honeybee gut microbiota and therefore compromise bee health. Shikimate pathway that is found in the bacterial community residing in the bee gut, such as in Snodgrassella alvi, Gilliamella spp., and Bifidobacterium spp., is known to play a key role in the production of essential aromatic compounds such as the amino acids phenylalanine, tryptophan, and tyrosine. In this context, glyphosate, the primary herbicide, inhibits 5-enolpyruvyl-shikimate-3-phosphate synthase (EPSPS) in the shikimate pathway (Motta et al. 2018). Thus, glyphosate via shikimate pathway inhibition and subsequently essential nutrients depletion play a crucial role in bacterial death and reducing beneficial bacteria in bee gut.
Propolis is another critical factor involved in honeybee gut microbiome consistency. In this context, Saelao et al. investigate the association between propolis and microbial community consistency in the honeybee microbiome. They disclosed that propolis insufficiency contributes to significant perturbation in the abundance of several key gut microbiota members. These authors proposed that propolis, via restricting alterations in the microbial community, play a key role in honeybee colony microbial health (Saelao et al. 2020).
9.4 Probiotic Potential of Honeybee Gut-Associated Bacteria
Inappropriate and misuse of antibiotics has led to a rise in antibacterial resistance and diminished the efficacy of these once considered miracle drugs. Since the alarming rise of antibiotic resistance, many strategies and investigations have been carried out to explore other safer ways to treat human ailments without harming the natural immunity of the host, and replacing or augmenting these antibiotics.
In the late nineteenth century, microbiologists identified microflora in healthy individuals’ gastrointestinal tracts that differed from those found in diseased individuals. The beneficial microflora found in the gastrointestinal tract was termed probiotics. FAO and WHO experts defined the term probiotics as “Live microorganisms which when administered in adequate amounts confer a health benefit on the host” (Joint 2002). In other words, probiotics are living microorganisms used to restore gut health by maintaining the intestinal microbiota (Manzanares et al. 2016). Similar to humans and animals, the gut-associated bacterial flora in honeybees has been reported to have the ability to provide health benefits, most important of all, which is the capability to protect them from several honeybee diseases (Li et al. 2017; Schwarz et al. 2016).
Below we discuss some of the beneficial functions carried out by the gut microbiome of honeybees.
9.4.1 Antimicrobial Effects of Honeybee Gut-Associated Bacteria
As stated earlier, the gut microbial community in honeybee A. mellifera protects the host from infection. Schwarz et al. provide experimental support linking parasite susceptibility of honeybee to dysbiosis of their core microbiota. They disclosed that honeybee, in a dysbiosis state, lose their ability to control encounter protozoan Lotmaria passim and lead to L. passim infection in these insects (Schwarz et al. 2016). Nosema ceranae is a gut intracellular parasite of honeybees that destroys epithelial cells and gut tissue integrity. In this context, Jiang Hong Li in 2017 revealed that disruption of bacteria in the honeybee by antibiotic treatment mediates honeybee’s susceptibility to Nosema infection (Li et al. 2017).
Moreover, Huang and Evans in 2020 investigated the effect of Nosema on the gut microbiome via suppression of N. ceranae with specific siRNAs. They found that suppressing N. ceranae led to significant positive effects on gut microbial abundance. These researchers concluded that N. ceranae is negatively correlated with the abundance of 15 identified bacteria (Huang and Evans 2020). In a study conducted by a group of researchers, it was found that the members of the gut microbiome, by lowering the local intestinal pH with the production of lactic acid, antimicrobial metabolites, as well as induction of innate immunity, interfere with the growth of Nosema infection (El Khoury et al. 2018). Furthermore, Streptomycin is an aminoglycoside antibiotic function in protein synthesis inhibition in Gram-negative bacteria. In 2015, through a metagenomics approach, Saraiva identified genes involved in streptomycin biosynthesis in A. mellifera microbiome. The presence of such genes raises the hypothesis about the possible role of normal microbiota in protecting Apis mellifera against pathogenic bacteria and in maintaining the healthy status of the hive (Saraiva et al. 2015).
Paenibacillus larvae, a Gram-positive sporulated bacterium that causes the American foulbrood disease, is an extremely contagious and dangerous pathogen of honeybees. In 2009, Sabate et al. aimed to explore the biological control capability of Bacillus strains associated with the bee intestine and evaluate their influence against P. larvae. They found that Bacillus strains through surfactin synthesis inhibit the growth of P. larvae (Sabaté et al. 2009). These novel findings collectively emphasize the importance of A. mellifera gut bacteria in modulating honeybees’ susceptibility to various infections.
9.4.2 Gut Microbiome Role in Immune Function
Various predators, including parasites, parasitoids, and pathogens, threaten insect health during their life cycle. A complex immune system has evolved in insects for protection against these threats. Several studies have illustrated that gut bacteria are key mediators in immune modulation and are essential for a healthy immune system (Kaltenpoth and Engl 2014). Hemocyte (immune cells), as a crucial element in the innate immune system through phagocytosis, plays a key role in hemolymph pathogen clearance. Vitellogenin (Vg) is a protein engaged in honeybee worker’s stress tolerance, and behavior. Vg is the main zinc carrier in honeybee workers, and zinc deficiency is associated with hemocyte pycnosis (cell death). Thereby, Vg is considered a critical mediator in honeybee immunity and lead to a longer life span. In an experimental investigation by Zheng et al. in 2017, it was found that normal microbiota compared to germ-free bees increase vitellogenin expression almost fivefold (Kaltenpoth and Engl 2014; Zheng et al. 2017). Overall, based on these findings, we can consider the gut microbiome as a major contributing factor for honeybee immune activation.
Furthermore, the Scab phenotype as a prominent immune response factor is triggered by reminiscent of melanization and develops 5–7 days after adult worker bees have emerged. Scab phenotype is characterized by a dark brown to black deposit forming a localized thin band in the pylorus at the midgut-hindgut boundary, in close proximity to the Malpighian tubules of the honeybees. Emery et al. identified significant host gene expression alteration in the pylorus region following Frischella perrara colonization compared to non-colonized bee. Using gene ontology (GO) enrichment analysis, they disclosed that immune-related genes, including irp30, cdc2c, abaecin, apid73, b-guc2, and def-1, were increased in the pylorus region of the screened honeybees. In this manner, Frischella perrara via colonization in a restricted region in the pylorus, as well as immune-related genes activation, play a key role in scab phenotype induction (Emery et al. 2017). Additionally, Horak and his research team investigate the beneficial effect of symbiont Snodgrassella alvi on honeybee immune gene expression. They illustrate that Snodgrassella alvi via expression of host antimicrobial peptides as well as Toll pathway upregulation aid in the clearance of opportunistic pathogen Serratia marcescens from the honeybees gut (Horak et al. 2020).
9.4.3 Gut Microbiome Role in Food Fermentation
Fermentation products such as short-chain fatty acids (SCFAs) are highly beneficial for host energy metabolism. In the fermentation process, A. mellifera gut microbiota members play an important role in breaking saccharides into an array of alcohols, SCFAs, gases, and other organic acids such as acetate and lactate. Acetate kinase (ackA) and L-lactate dehydrogenase (ldh) are the main enzymes responsible for acetate and lactate production, respectively. In turn, acetate production and lactate through increased sucrose sensitivity play a crucial role in honeybee weight gaining (Lee et al. 2018; Zheng et al. 2017). Therefore, A. mellifera gut microbiota through organic acid production plays an important role in honeybee weight gain.
9.4.4 Gut Microbiome Role in Detoxification
Gut microbiota strongly promotes the expression of key enzymes of the honeybee xenobiotic detoxification pathway. Three important enzymes responsible for insect detoxification, including carboxylesterases (COEs), Cytochrome P450 monooxygenases (CYPs, also called P450s), and glutathione S-transferases (GSTs), have been identified recently. Thereby, honeybee gut microbiota enhance host detoxification capability and manipulate host metabolism (Wu et al. 2020). Furthermore, some monosaccharide sugars, including xylose, mannose, rhamnose, and arabinose, have been reported to endorse toxic effects on A. mellifera and decrease their life span. Recently, genes responsible for mannose metabolism, including phosphotransferase systems (PTSs) and mannose-6-phosphate isomerase (MPI), were identified in the Gilliamella apicola genome. Additionally, several genes associated with catabolism of rhamnose, xylose, and arabinose have also been detected in the genome of Gilliamella apicola. Hence, it is concluded that Gilliamella apicola is able to metabolize xylose, mannose, rhamnose, and arabinose and subsequently boost A. mellifera life span (Zheng et al. 2016).
9.4.5 Probiotic Properties of Honeybee-Specific Lactic Acid Bacteria
Lactic acid bacteria (LAB) are a group of Gram-positive lactic acid-producing bacteria present in diverse habitats. LAB belongs to phylum Firmicutes with low G + C in the genome. These bacteria are well known for their role in food fermentation, and a wide variety of strains are routinely employed as starter cultures in the manufacture of dairy, meat, vegetable, and bakery products. Additionally, they have a significant role as starter cultures for cheese and yogurts. One of the factors that make LAB of high importance, especially for human and animal use, is their “generally recognized as safe” (GRAS) status that make these food-grade microorganisms to be employed as probiotics (Åvall-Jääskeläinen and Palva 2005; Choi et al. 2005). While the European Food Safety Authority (EFSA) proclaimed the LAB strain to have QPS (Qualified presumption of safety) status (EFSA 2008). In several findings, the presence of LAB in the gut of honeybees has been reported, where they are shown to provide beneficial effects to their host, and thereby they are potential probiotic candidates.
Majority of honeybee-specific LAB has found significant importance owing to their probiotic potentials. Up to date, thirteen genetically distinct lactic acid-producing bacteria have been identified from the honeybee crop, of which nine are Lactobacilli and four are Bifidobacteria (Olofsson et al. 2014; Olofsson and Vásquez 2008).
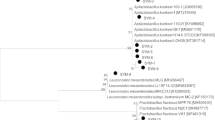
Lactic acid bacteria has been isolated from the gut of several honeybee species, including Apis mellifera, A. dorsata, A. florea, A. nigrocincta, Apis nuluenis, Apis laboriosa, A. cerana indica, Melipona beecheii, Meliponula bacandei, and Trigona sp. (Mathialagan et al. 2018; Niode et al. 2020; Vásquez et al. 2012). Lactobacillus is one of the most important genera within the LAB, which at present includes 175 listed species (Euzéby 1997). Among this group of bacteria, genus Lactobacillus is the most frequent Gram-positive bacteria isolated from different honeybee species’ gut. While Lactobacillus kunkeei has been reported to be one of the most dominant species of this genus residing in their gut (Niode et al. 2020). Table 9.2 depicts a variety of LAB species isolated from different honeybee species around the world.
9.4.6 Antimicrobial Effect of Honeybee Gut-Associated LAB Against Honeybee Diseases
LAB comprises a group of Gram-positive, catalase-negative, non-motile, non-spore-forming facultative anaerobic bacteria that are commonly found as both exogenous and endogenous microbes in healthy individuals. Similar to the LAB found within humans and animals, the honeybee-specific LAB defends their hosts from invasion and colonization of several pathogenic bacteria via the production of a variety of antimicrobial metabolites and modulation of the host immune response (Huang and Evans 2020; Mathialagan et al. 2018; Niode et al. 2020). Hence, honeybee-specific LAB can also provide protection against several honeybee diseases by production of these metabolites.
The antimicrobial effects exerted by these bacteria are owing to their metabolite-producing abilities. These metabolites, also referred to as postbiotic metabolites, includes organic acids (lactic acid, acetate acid, and formic acid) (Olofsson et al. 2016), extracellular proteins, benzoate, bacteriocins, hydrogen peroxide (H2O2), lipopolysaccharides, and lipoteichoic acid volatile compounds (Butler et al. 2013; Olofsson et al. 2016; Olofsson and Vásquez 2008) etc.
Hence, due to the stated health-promoting functions of LAB in honeybees, they are considered safe alternative therapeutic strategy for the control of a number of honeybee diseases, including Paenibacillus larvae (infective bacterial agent of American foulbrood disease), Melissococcus pluton (infective bacterial agent of European foulbrood), Nosemosis, and varroosis (Audisio 2017; Forsgren et al. 2010). In a study conducted by a group of researchers, an organic acid-producing L. johnsonii was shown to inhibit the growth of Nosema ceranae and harbored fumigillin activity (60).
Paratransgenesis has come to mean a Trojan horse strategy, where endogenous microorganisms via effector molecules production inhibit pathogen development. Candidate microorganisms to being practical in honeybee, they should possess several criteria including (1) candidate microorganism should be genetically modifiable for effector molecules expression; (2) ideally the candidate microorganism must be ecologically and functionally fit with other nonpathogenic bee-associated microorganisms, and (3) following reintroduction the modified organism should have no negative impact on honeybee health. In this context, Rangberg et al. investigated L. kunkeei potency in honeybee paratransgenesis. They concluded that L. kunkeei complies with the three criteria required for being a suitable paratransgenic candidate (Rangberg et al. 2015). Similar to these findings, Maddaloni and his co-investigators demonstrated that Fructobacillus fructosus can be used as a powerful tool for honeybee paratransgenesis to control diseases and expand nutrition repertoire (Maddaloni et al. 2014).
9.5 Commonly Used Methods for Microbiome Analysis
Researcher frequently utilizes full-length 16S rRNA gene sequences with nine hypervariable regions (V1–V9) to infer phylogenetic relationship among the microbiome. Therefore, a full-length 16S rRNA amplicon sequencing approach with high accuracy and efficiency can be used for microbial diversity detection in various biological samples. Nanopore DNA sequencer (MinION) containing several significant advantages including rapid library construction, low cost, real-time detection and small size that made it a suitable tool for identifying microbiome composition at the species levels (Shin et al. 2016). However, it was disclosed that bacteria with almost identical 16S rRNA sequences could exhibit high sequence divergence levels at other loci and very different gene repertoires. Thereby, it is difficult to gain insight into intraspecific diversification of bacterial lineages in the gut with 16S rRNA sequencing. Single-cell genomics and transcriptomics can provide reliable context for assembled genome fragments and gene expression activity on the level of individual prokaryotic genomes. In this manner single cell genomics, through allowing direct access to information from individual microorganisms, has the potential to elucidate processes of bacterial diversification (Engel et al. 2014). However, low DNA and mRNA content restrict the yield of reasonable amounts of genetic material for sequencing analysis from a single cell.
Furthermore, the lack of polyadenylation of bacterial mRNA limits its separation from rRNA. Additionally, cell walls and membranes diversity induce a challenge to consistent lysis or permeabilization required for single-cell RNA sequencing (scRNA-seq). These problems impede the characterization of microbes by traditional single-cell sequencing methods (Sharma and Thaiss 2020).
9.6 Microbiome Engineering as a Future Perspective
Engineering of microbiomes is used to modify structures of the microbiota and restore ecological balance. Synthetic biology and engineering principles are frequently applied in microbiome engineering to improve microbiome function. Thereby, microbiome engineering could lead to a breakthrough in agriculture and medicine. In medicine, microbiome engineering enables exploring individual microbes’ contribution and generating potential therapies against metabolic (e.g., phenylketonuria and chronic kidney disease), inflammatory, and immunological diseases, among others. In the case of honeybee, due to their agricultural importance as well as the simple gut microbiome, they are a promising testbed for the nascent field of microbiome engineering (Foo et al. 2017; Leonard 2020). There are several approaches to honeybee microbiome engineering. A plasmid toolkit by combining a broad-host-range (BHR) replicon with a set of modular genetic parts can be applied to bacteria from the A. mellifera gut microbiome. It was disclosed that plasmids constructed using bee microbiome toolkit (BTK) act faithfully in various species of Proteobacteria detected in the A. mellifera gut microbiome. The BTK can be used to express heterologous genes or to repress or disrupt genes in the bacterial chromosome (Leonard 2020). Consequently, microbiome engineering could be employed as a powerful tool for improving A. mellifera health and subsequently agricultural productivity.
9.7 Conclusions
The economic value of commercial honeybee pollination is estimated at over US $220 billion worldwide. Any damage to these insects leads to detrimental consequences not only to our agriculture and production values that ultimately would result in economic losses but might also threaten and endanger our lives on the planet. Hence, intensive research has been done and is still ongoing to find solutions to prevent colony losses and find ways to increase their survival and control the pathogens from harming their viability.
A.mellifera digestive tract is a reservoir of a diverse variety of bacterial communities that play a significant role in these insects’ growth and survival. Recent studies with gut microbiome disclosed the honeybee gut-associated microbial in immune system activation, carbohydrate fermentation, and inhibition of disease in the host. This suggests that the gut bacterial community structure may be considered as an indicator of honeybee health. Since related microbiotas are found across bee species, it strongly suggests a close evolutionary relationship between bacteria and hosts, as well as underscoring the importance of LAB symbionts for bees. Not only are LAB symbionts involved in honeybee food production and preservation, but they are also of importance in host defense against pathogen and transient microbes intercepted during foraging. Hence preserving the balance of these gut bacteria is crucial for maintaining honeybee health and vigor. Tools to engineer a microbial member of these honeybees might play a significant role in beekeeping management issues such as increased colony survival.
References
Alatawy M, Al-Attas SG, Assagaf AI, Al-shehri A, Alghamdi KM, Bahieldin A (2020) Gut microbial communities of adult honey bee workers (Apis Mellifera). Biosci Biotechnol Res Asia 17
Audisio MC (2017) Gram-positive bacteria with probiotic potential for the Apis mellifera L. honey bee: the experience in the northwest of Argentina. Probiotics Antimicrob Proteins 9:22–31
Audisio MC, Torres MJ, Sabaté DC, Ibarguren C, Apella MC (2011) Properties of different lactic acid bacteria isolated from Apis mellifera L. bee-gut. Microbiol Res 166:1–13
Åvall-Jääskeläinen S, Palva A (2005) Lactobacillus surface layers and their applications. FEMS Microbiol Rev 3:511–529
Bonilla-Rosso G, Paredes JC, Das S, Ellegaard KM, Emery O, Garcia-Garcera M, Glover N, Hadadi N, van der Meer JR, Tagini F (2019) Acetobacteraceae in the honey bee gut comprise two distant clades with diverging metabolism and ecological niches. bioRxiv:861260
Bottacini F, Milani C, Turroni F, Sánchez B, Foroni E, Duranti S, Serafini F, Viappiani A, Strati F, Ferrarini A (2012) Bifidobacterium asteroides PRL2011 genome analysis reveals clues for colonization of the insect gut. PLoS One 7:e44229
Butler È, Alsterfjord M, Olofsson TC, Karlsson C, Malmström J, Vásquez A (2013) Proteins of novel lactic acid bacteria from Apis mellifera mellifera: an insight into the production of known extra-cellular proteins during microbial stress. BMC Microbiol 13:235
Choi SS, Kang BY, Chung MJ, Kim SD, Park SH, Kim JS, Kang CY, Ha NJ (2005) Safety assessment of potential lactic acid bacteria Bifidobacterium longum SPM1205 isolated from healthy Koreans. J Microbiol 43:493–498
Cornman RS, Tarpy DR, Chen Y, Jeffreys L, Lopez D, Pettis JS, Evans JD (2012) Pathogen webs in collapsing honey bee colonies. PLoS One 7:e43562
Dong Z-X, Li H-Y, Chen Y-F, Wang F, Deng X-Y, Lin L-B, Zhang Q-L, Li J-L, Guo J (2020) Colonization of the gut microbiota of honey bee (Apis mellifera) workers at different developmental stages. Microbiol Res 231:126370
Duong BTT, Lien NTK, Thu HT, Hoa NT, Lanh PT, Yun B-R, Yoo M-S, Cho YS, Van Quyen D (2020) Investigation of the gut microbiome of Apis cerana honeybees from Vietnam. Biotechnol Lett 42:2309–2317
EFSA (2008) Opinion of the scientific panel on biological hazards on the maintenance of the list of QPS microorganisms intentionally added to food or feed. EFSA J 923:1–48
El Khoury S, Rousseau A, Lecoeur A, Cheaib B, Bouslama S, Mercier P-L, Demey V, Castex M, Giovenazzo P, Derome N (2018) Deleterious interaction between honeybees (Apis mellifera) and its microsporidian intracellular parasite Nosema ceranae was mitigated by administrating either endogenous or allochthonous gut microbiota strains. Front Ecol Evol 6:58
Elzeini HM, Ali A-RA-A, Nasr NF, Elenany YE, Hassan AAM (2020) Isolation and identification of lactic acid bacteria from the intestinal tracts of honey bees, Apis mellifera L., in Egypt. J Apicult Res:1–9
Emery O, Schmidt K, Engel P (2017) Immune system stimulation by the gut symbiont Frischella perrara in the honey bee (Apis mellifera). Mol Ecol 26:2576–2590
Engel P, Moran NA (2013) The gut microbiota of insects–diversity in structure and function. FEMS Microbiol Rev 37:699–735
Engel P, Stepanauskas R, Moran NA (2014) Hidden diversity in honey bee gut symbionts detected by single-cell genomics. PLoS Genet 10:e1004596
Euzéby JP (1997) List of Bacterial Names with Standing in nomenclature: a folder available on the Internet. Int J Syst Evol Microbiol 47:590–592
Foo JL, Ling H, Lee YS, Chang MW (2017) Microbiome engineering: current applications and its future. Biotechnol J 12:1600099
Forsgren E, Olofsson TC, Váasquez A, Fries I (2010) Novel lactic acid bacteria inhibiting Paenibacillus larvae in honey bee larvae. Apidologie 41:99–108
Geldert C, Abdo Z, Stewart JE, HS A (2020) Dietary supplementation with phytochemicals improves diversity and abundance of honey bee gut microbiota. J Appl Microbiol
Genersch E, Von Der Ohe W, Kaatz H, Schroeder A, Otten C, Büchler R, Berg S, Ritter W, Mühlen W, Gisder S (2010) The German bee monitoring project: a long term study to understand periodically high winter losses of honey bee colonies. Apidologie 41:332–352
Horak RD, Leonard SP, Moran NA (2020) Symbionts shape host innate immunity in honeybees. Proc R Soc B 287:20201184
Huang Q, Evans JD (2020) Targeting the honey bee gut parasite Nosema ceranae with siRNA positively affects gut bacteria. BMC Microbiol 20:1–6
Huang SW, Zhang HY, Marshall S, Jackson TA (2010) The scarab gut: a potential bioreactor for bio-fuel production. Insect Sci 17:175–183
Joint F (2002) WHO working group report on drafting guidelines for the evaluation of probiotics in food. London, Ontario, Canada 30
Kaltenpoth M, Engl T (2014) Defensive microbial symbionts in H ymenoptera. Funct Ecol 28:315–327
Khan KA, Ansari MJ, Al-Ghamdi A, Nuru A, Harakeh S, Iqbal J (2017) Investigation of gut microbial communities associated with indigenous honey bee (Apis mellifera jemenitica) from two different eco-regions of Saudi Arabia. Saudi J Biol Sci 24:1061–1068
Kwong WK, Moran NA (2016) Gut microbial communities of social bees. Nat Rev Microbiol 14:374–384
Lee FJ, Miller KI, McKinlay JB, Newton IL (2018) Differential carbohydrate utilization and organic acid production by honey bee symbionts. FEMS Microbiol Ecol 94:fiy113
Leonard SP (2020) Engineering the gut microbiome of honey bees
Li JH, Evans JD, Li WF, Zhao YZ, DeGrandi-Hoffman G, Huang SK, Li ZG, Hamilton M, Chen YP (2017) New evidence showing that the destruction of gut bacteria by antibiotic treatment could increase the honey bee’s vulnerability to Nosema infection. PLoS One 12:e0187505
Maddaloni M, Hoffman C, Pascual D (2014) Paratransgenesis feasibility in the honeybee (A pis mellifera) using F ructobacillus fructosus commensal. J Appl Microbiol 117:1572–1584
Manzanares W, Lemieux M, Langlois PL, Wischmeyer PE (2016) Probiotic and synbiotic therapy in critical illness: a systematic review and meta-analysis. Crit Care 20:262
Martinson VG, Danforth BN, Minckley RL, Rueppell O, Tingek S, Moran NA (2011) A simple and distinctive microbiota associated with honey bees and bumble bees. Mol Ecol 20:619–628
Martinson VG, Moy J, Moran NA (2012) Establishment of characteristic gut bacteria during development of the honeybee worker. Appl Environ Microbiol 78:2830–2840
Mathialagan M, Johnson Y, Thangaraj E (2018) Isolation, characterization and identification of probiotic lactic acid bacteria (LAB) from honey bees. Int J Curr Microbiol App Sci 7:849–906
Meixner MD (2010) A historical review of managed honey bee populations in Europe and the United States and the factors that may affect them. J Invertebr Pathol 103:S80–S95
Mohr KI, Tebbe CC (2006) Diversity and phylotype consistency of bacteria in the guts of three bee species (Apoidea) at an oilseed rape field. Environ Microbiol 8:258–272
Moran NA, Hansen AK, Powell JE, Sabree ZL (2012) Distinctive gut microbiota of honey bees assessed using deep sampling from individual worker bees. PLoS One 7:e36393
Motta EV, Raymann K, Moran NA (2018) Glyphosate perturbs the gut microbiota of honey bees. Proc Natl Acad Sci 115:10305–10310
Niode NJ, Salaki CL, Rumokoy LJ, Tallei TE (2020) Lactic acid bacteria from honey bees digestive tract and their potential as probiotics, International Conference and the 10th Congress of the Entomological Society of Indonesia (ICCESI 2019). Atlantis Press, pp. 236–241
Olofsson TC, Vásquez A (2008) Detection and identification of a novel lactic acid bacterial flora within the honey stomach of the honeybee Apis mellifera. Curr Microbiol 57:356–363
Olofsson TC, Alsterfjord M, Nilson B, Butler È, Vásquez A (2014) Lactobacillus apinorum sp. nov., Lactobacillus mellifer sp. nov., Lactobacillus mellis sp. nov., Lactobacillus melliventris sp. nov., Lactobacillus kimbladii sp. nov., Lactobacillus helsingborgensis sp. nov. and Lactobacillus kullabergensis sp. nov., isolated from the honey stomach of the honeybee Apis mellifera. Int J Syst Evol Microbiol 64:3109
Olofsson TC, Butler È, Markowicz P, Lindholm C, Larsson L, Vásquez A (2016) Lactic acid bacterial symbionts in honeybees–an unknown key to honey’s antimicrobial and therapeutic activities. Int Wound J 13:668–679
Parichehreh S, Tahmasbi G, Sarafrazi A, Imani S, Tajabadi N (2018) Isolation and identification of Lactobacillus bacteria found in the gastrointestinal tract of the dwarf honey bee, Apis florea Fabricius, 1973 (Hymenoptera: Apidae). Apidologie 49:430–438
Rangberg A, Mathiesen G, Amdam G, Diep D (2015) The paratransgenic potential of Lactobacillus kunkeei in the honey bee Apis mellifera. Benefic Microbes 6:513–523
Raymann K, Shaffer Z, Moran NA (2017) Antibiotic exposure perturbs the gut microbiota and elevates mortality in honeybees. PLoS Biol 15:e2001861
Sabaté DC, Carrillo L, Audisio MC (2009) Inhibition of Paenibacillus larvae and Ascosphaera apis by Bacillus subtilis isolated from honeybee gut and honey samples. Res Microbiol 160:193–199
Saelao P, Borba RS, Ricigliano V, Spivak M, Simone-Finstrom M (2020) Honeybee microbiome is stabilized in the presence of propolis. Biol Lett 16:20200003
Saraiva MA, Zemolin APP, Franco JL, Boldo JT, Stefenon VM, Triplett EW, de Oliveira Camargo FA, Roesch LFW (2015) Relationship between honeybee nutrition and their microbial communities. Antonie Van Leeuwenhoek 107:921–933
Schwarz RS, Moran NA, Evans JD (2016) Early gut colonizers shape parasite susceptibility and microbiota composition in honey bee workers. Proc Natl Acad Sci 113:9345–9350
Shanbhag S, Tripathi S (2009) Epithelial ultrastructure and cellular mechanisms of acid and base transport in the Drosophila midgut. J Exp Biol 212:1731–1744
Sharifpour MF, Mardani K, Ownagh A (2016) Molecular identification and phylogenetic analysis of Lactobacillus and Bifidobacterium spp. isolated from gut of honeybees (Apis mellifera) from West Azerbaijan, Iran, Veterinary Research Forum. Faculty of Veterinary Medicine, Urmia University, Urmia, Iran, p 287
Sharma PV, Thaiss CA (2020) Host-microbiome interactions in the era of single-cell biology. Front Cell Infect Microbiol 10
Shin J, Lee S, Go M-J, Lee SY, Kim SC, Lee C-H, Cho B-K (2016) Analysis of the mouse gut microbiome using full-length 16S rRNA amplicon sequencing. Sci Rep 6:29681
Tihelka E, Cai C, Pisani D, Donoghue PC (2020) Mitochondrial genomes illuminate the evolutionary history of the Western honey bee (Apis mellifera). Sci Rep 10:1–10
Vásquez A, Forsgren E, Fries I, Paxton RJ, Flaberg E, Szekely L, Olofsson TC (2012) Symbionts as major modulators of insect health: lactic acid bacteria and honeybees. PLoS One 7:e33188
Wang H, Liu C, Liu Z, Wang Y, Ma L, Xu B (2020) The different dietary sugars modulate the composition of the gut microbiota in honeybee during overwintering. BMC Microbiol 20:1–14
Wu Y, Zheng Y, Chen Y, Wang S, Chen Y, Hu F, Zheng H (2020) Honey bee (Apis mellifera) gut microbiota promotes host endogenous detoxification capability via regulation of P450 gene expression in the digestive tract. J Microbial Biotechnol 13:1201–1212
Zeinali F, Zarch SMA, Mehrjardi MYV, Kalantar SM, Jahan-Mihan A, Karimi-Nazari E, Fallahzadeh H, Hosseinzadeh-Shamsi-Anar M, Rahmanian M, Fazeli MR (2020) Effects of synbiotic supplementation on gut microbiome, serum level of TNF-α, and expression of microRNA-126 and microRNA-146a in patients with type 2 diabetes mellitus: study protocol for a double-blind controlled randomized clinical trial. Trials 21:1–9
Zheng H, Nishida A, Kwong WK, Koch H, Engel P, Steele MI, Moran NA (2016) Metabolism of toxic sugars by strains of the bee gut symbiont Gilliamella apicola. MBio 7
Zheng H, Powell JE, Steele MI, Dietrich C, Moran NA (2017) Honeybee gut microbiota promotes host weight gain via bacterial metabolism and hormonal signaling. Proc Natl Acad Sci 114:4775–4780
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Tootiaie, S., Moharrami, M., Mojgani, N. (2021). Honeybee Gut: Reservoir of Probiotic Bacteria. In: Mojgani, N., Dadar, M. (eds) Probiotic Bacteria and Postbiotic Metabolites: Role in Animal and Human Health. Microorganisms for Sustainability, vol 2. Springer, Singapore. https://doi.org/10.1007/978-981-16-0223-8_9
Download citation
DOI: https://doi.org/10.1007/978-981-16-0223-8_9
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-16-0222-1
Online ISBN: 978-981-16-0223-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)