Abstract
This study aims to identify lactic acid bacteria (LAB) isolated from honeybees (Apis mellifera workers and larvae) in detail and to determine their functional probiotic properties. A total of 11 strains were classified based on morphological and biochemical characteristics. Preliminary probiotic properties of strains, that were molecularly identified using 16 S rRNA, such as antimicrobial activity, tolerance to digestive conditions, aggregation ability, were investigated. The antimicrobial properties of strains were tested against a wide range of human pathogens. All strains that showed γ-hemolysis and did not contain bacteriophages were considered safe. The strains’ survivability checked for 0.3% bile and 3.0-7.8 pH contents was promising. The highest autoaggregation ranged from 14.7 to 30.76% after 4 h. Tested LAB strains markedly exhibited coaggregation with Listeria monocytogenes and Escherichia coli. According to the results, tested bacteria showed significant antagonistic effects against pathogens, and positive probiotic characteristics compatible with in vitro gastrointestinal tract conditions. The results suggest that Apis mellifera LAB symbionts may have a probiotic potential, and be effective and safe candidates for human use. This study provides an addition to the development of the current knowledge by defining in detail honeybee-associated bacteria and determining their probiotic potential.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Honeybee colonies are exposed to various agroecosystems and many environmental factors that can affect the microbial balance of the hive throughout the year. In this way, they have a rich and unique collection of microorganisms forming normal or transient microflora [1]. A balanced microflora benefits honeybees in many ways, including metabolic and protective functions [2]. The honeybee’s microflora is predominantly composed of lactic acid bacteria, as other animals and humans [3,4,5,6,7]. It has been found that honeybee stomachs, intestines, and products contained more than 45 species of LAB. Among these species, Bifidobacterium asteroides, Bifidobacterium coryneform, Leuconostoc spp., Fructobacillus fructosus, and a number of Lactobacillus species such as Lactobacillus plantarum, Lactobacillus apis, Lactobacillus (Apilactobacillus) kunkeei, Lactobacillus mellifer have been reported to be associated with healthy honeybee colonies. Apilactobacillus kunkeei is frequently isolated from honeybee crop, pollen sac, and larvae [8,9,10,11,12,13,14].
The Food and Agriculture Organization/World Health Organization (FAO/WHO) has redefined probiotics as “Live microorganisms which when consumed in adequate amounts as part of food confer a health benefit on the host” [15, 16]. Probiotics exert dominance by lowering luminal pH, competing for nutrients, adhering to the surface of epithelial cells or mucus, and antagonizing pathogen colonization through aggregation with pathogens [17]. LAB is known as probiotic symbiont in many living things, including honeybees and humans [6]. The most important genera of LAB which can be used as probiotics are Lactobacillus, Streptococcus, Enterococcus, Lactococcus, Pediococcus, and Leuconostoc [17,18,19,20,21].
LAB which is of great importance to human beings is closely related to numerous health and nutritional benefits. Among these are prevention of infectious diseases, maintenance of normal insulin levels in the blood, reducing serum cholesterol, nutrients synthesizing and bioavailability, food preservation and fermentation. Moreover, LAB may play a key role in treating a wide range of conditions such as Helicobacter pylori-associated peptic ulcer, lactose malabsorption, diarrhea, genital and urinary tract infections, allergic reactions, colon cancer [17–18, 22,23,24].
Nowadays, there is an increasing demand for novel LAB strains with probiotic properties [20]. Potentially beneficial bacteria are being explored in numerous alternative sources, including grains, fruits and vegetables, meat and meat products, honey and other hive products. For example, fructophilic lactic acid bacteria (FLAB) that live in symbiosis with insects such as honeybees have been a focus of attention in recent years due to their potential beneficial properties in human health care. The safety of new potential strains, their functional properties such as gastric acid and bile salt resistance, and their effects on the host are among the selection criteria as probiotics [14].
This study focuses on the characterization of LAB strains isolated from worker bees and larval samples (Apis mellifera), as well as on conducting in vitro experiments to detect their functional properties as probiotics.
Materials and methods
Sample collection and bacterial isolation
Honey bee and larvae samples were collected from different apiary regions of Turkey in 2018–2019 by Samsun Veterinary Control Institute Bee Diseases Laboratory. For the samples in which SYM 2-3-7-10 strains were isolated, 3 adults were used from the hive selected for each group, and for the SYM 1-4-5-6-8-9-11 strains, 3 larvae were used from the hive selected for each group. Surface sterilization of the samples was done by dipping in 70% ethanol for 30 s under aseptic conditions, followed by washing 3 times for 5 min with sterile distilled water. Then, the bee samples were mechanically dissected using prefilled 2.0 mL tubes with silica (glass) beads, and 100 µl of each sample were streaked onto de Man, Rogosa, and Sharpe agar (MRSA, Neogen) medium for LAB culture (one sample per plate) [25, 26]. Petri dishes were incubated at 37 °C for 2–3 days using anaerobic jar (Thermo Scientific™ Oxoid™, AnaeroJar™ 2.5 l, Catalog No: AG0025A) and gas pack technique (Thermo Scientific™ Oxoid™, CampyGen™, Catalog No: CN0020C). At the end of the incubation, macroscopic and microscopic morphology characteristics of bacterial colonies grown on MRSA were examined. Colonies with gram positive, fine bacillus and cocci shaped were selected and passaged into MRSA medium by single colony drop cultivation technique. After the cultures were incubated again under the same conditions, a loopful of isolated pure LAB cultures were taken, re-passaged into 3 ml MRS broth (MRSB, Merck), and left to grow for 1 night at 120 rpm in a shaking incubator at 37 °C. Fresh stock cultures were stored in MRSB with 20% glycerol (v v− 1) at -20 °C and − 80 °C until further experiments.
Morphologic and biochemical characterization
All LAB isolates were prepared for use by sub-culturing twice in MRSB for each test. Morphological and biochemical properties of LAB colonies grown at the end of the culture were determined according to Bergey’s Manual of Systematic and Determinative Bacteriology. All strains were examined microscopically by Gram staining and were subjected to a number of biochemical analyses such as various enzyme activities, starch hydrolysis, esculin hydrolysis, and the ability to ferment carbohydrates. Three different incubation temperatures at 18, 37, and 45 °C were tested to determine the optimum growth temperature of the strains [27,28,29,30].
Molecular identification
The genomic DNA of each strain was extracted according to the methodology used by Sambrook et al. [31]. 16S rRNA gene amplification was carried out using universal primers, 27F (5’-AGAGTTTGATCCTGGCTCTCAG-3’) and 1492R (5’-TACCTTGTTACGACTT-3’). The PCR products were examined through 1% (w v− 1) agarose gel electrophoresis and were visualized in a UV transilluminator. Amplicons were sent to Macrogen Inc. (Amsterdam, Netherlands) for purification and sequencing. Obtained raw sequences were edited by BioEdit Software version 7.2.5 and were aligned to closely related species using the Basic Local Alignment Search Tool (BLAST) on the National Center for Biotechnology Information (NCBI) GenBank database. Phylogenetic tree was constructed on the basis of the Neighbor-Joining method in MEGA 11.0.8 Software [32, 33]. The obtained sequences were submitted to Genbank and received accession numbers.
Bacteriophage detection
Bacteriophage contents of LAB strains were determined according to the methods of Kılıç et al. and Trevors et al. [34, 35]. Cultures incubated overnight (18 h) were passaged as 60 µl into 3 ml of MRS-C medium (2.5 ml of 10 mM CaCl2, 250 ml of MRSB). After approximately 2 h of incubation (OD600 nm=0.2), mitomycin C (0.2 µg ml− 1) was added to each culture and incubation process was continued for 5–8 h. Cultures were observed during the incubation period, and those with prophage induction or lysis were centrifuged (12,470 x g, 10 min). The supernatants (lysate) were collected, transferred to new eppendorf tubes, and stored by adding chloroform. 200 µl of the host LAB culture was added into the pre-prepared soft MRS-C medium which was cooled to 50 °C, was quickly mixed by vortexing, and was poured onto the MRSA plates as a second layer. Lysates were dropped onto dried soft agar one by one. Plates were examined at the end of 48 h incubation at 37 °C. After the lysate was dropped on, the areas where the host bacteria did not grow were evaluated as phage plaques, with the expansion of the inhibition zone over time. The formation of this type of inhibition zone was determined as the presence of bacteriophages.
Antimicrobial activity
The agar plug diffusion method and CFS (cell-free culture supernatant) well diffusion method were performed to determine the antimicrobial activity of LAB strains on pathogenic microorganisms [36,37,38]. The indicator culture plates were prepared for inhibition tests and therefore the pathogenic bacteria sourced from frozen stocks were reactivated in the appropriate agar media. 18-h cultures of each pathogenic bacterium (McFarland 0.5) were resuspended into broth media, and then 100 µl of each bacterial suspension was spread on the surface of agar plate homogeneously with a sterile cotton swab.
For the agar well diffusion test of CFS, holes 6 mm in diameter were punched into the indicator culture plates via sterile cork borer. The CFS of LAB strains was prepared from subcultured in MRS broth. The 72-h cultures were centrifuged (10,000 x g, 10 min) and supernatants were filtered through a syringe filter (0.45 µm, Aisimo). The 60 µl of CFS was filled into each well on the agar plates. For the agar plug diffusion test of LAB, the 6 mm diameter agar plugs were aseptically cut from each 24-h LAB culture plated in MRSA, placed on the surface of agar plates (without wells), and previously inoculated with pathogenic microorganisms. All plates were incubated anaerobically for 24 h at 37 °C. The results were evaluated by measuring the inhibition zones around the wells.
Hdyrogene peroxide production
Tetramethylbenzidine (TMB)-Plus medium containing 43 g l− 1 Brucella Agar Base, 10 g soluble starch, 100 ml l− 1 TMB (3,3’,5,5’-tetramethylbenzidine) solution, 1 mg l− 1 horseadish peroxidase, 50 ml bovine serum, 2 mg l− 1 vitamin K (phytomenadione), 5 g l− 1 hemin, 0,12 g l− 1 manganese (II) sulfate monohydrate (MnSO4.H2O) and 0,57 g l− 1 magnesium sulfate (MgSO4) was prepared according to Rabe and Hillier’s and Alpay’s protocols with some modifications [27, 39]. LAB isolates were cultured in TMB-Plus medium at 37 °C anaerobically (using GasPak, CampyGen, AGS) for 72 h. At the end of the incubation, the culture plates were kept in atmospheric air for 15 min. Lactobacillus acidophilus DSM 20,242 was used as positive control. The assay was determined as positive based on observation of dark bluish ring formed around the colonies producing H2O2 in contact with oxygen.
Hemolytic activity
Hemolytic activity was determined by culturing bacterial isolates in Brain Heart Infusion (BHI) agar plates supplemented with 5% (v v− 1) of sheep blood at 37 °C for 24 h. Staphylococcus aureus was used as a positive control. The absence of discoloration zones around the colonies was considered as non-hemolytic activity (γ-hemolysis) [40].
Acid tolerance
Phosphate-buffered saline solutions with pH values of 3.0 and 7.8 were prepared to determine the resistance of potential probiotic microorganisms to the acidic pH digestion process in the stomach and their ability to reach and adapt to the small intestine in vitro. One ml of fresh LAB cultures was centrifuged (10,000 x g, 5 min), pellets were washed twice with phosphate-buffered saline (PBS, pH 7.2) and then resuspended in 1 ml of PBS (pH 3.0 and 7.8) by adjusting the final inoculum size to ≈ 106 CFU ml− 1. The suspensions were incubated in an anaerobic jar using gas pack (Thermo Scientific™ Oxoid™, AnaeroJar™ 2.5 l, Catalog No: AG0025A, Thermo Scientific™ Oxoid™, CampyGen™, Catalog No: CN0020C) for 3 and 4 h at 37 °C incubator (Memmert ULM 600), based on the digestion time of the food in the stomach and intestines. After 3 h, 100 µl of bacterial suspension was serially diluted and spread on standard MRSA plates. All visible colonies were enumerated, and the viability was calculated as quantification of the exact log CFU ml− 1 at the end of 48 h incubation [40,41,42].
Bile salt tolerance
In order to examine the bile salt resistance properties of LAB strains, a Bile-MRS broth medium containing 0.3% (w/v) of bile salts (Sigma Aldrich, B8381-10G) simulating the small intestine system was prepared [43,44,45]. LAB cultures grown overnight in MRSB were centrifuged (10,000 x g, 5 min), cell pellets were washed twice with PBS (pH 7.2) and then resuspended in 1 ml of Bile-MRS broth by adjusting the final inoculum size to ≈ 106 cfu ml− 1. After the samples were incubated in Bile-MRS broth for 4 h, 100 µl of bacterial suspensions were diluted and plated onto standard MRSA plates. By enumerating all visible colonies and quantifying their precise log CFU ml− 1, bacterial viability was assessed compared with negative controls after a 48-h incubation period. Negative controls were consider the strains incubated in MRSB medium for 4 h without exposure to Bile-MRS, then passaged into MRSA medium using the same colony counting method. In addition, LAB cultures were also incubated in Bile-MRS broth directly, and the growth activity was monitored by measuring the turbidity of cultures photometrically (OD600nm) over 20 h via Bioscreen C Automated Microbiology Growth Curve Analysis System (Thermo Scientific, USA). The bacterial growth curve was generated comparing test groups with control groups according to OD data received [40, 42, 43].
Autoaggregation and coaggregation activity
Cultures of 4 ml each of 18-h LAB and indicator pathogenic bacteria (Escherichia coli and Listeria monocytogenes) were centrifuged at 5000 g for 15 min, the pellets were washed twice with PBS (pH 7.2), and then dissolved in 4 ml of PBS by adjusting the bacterial turbidity density to ≈ 106 cfu ml− 1 at 600 nm (OD600 = 0.1–0.2). For the coaggregation test, 2 ml of the pathogen was mixed with 2 ml LAB solution. Prepared all bacterial mixtures were vortexed thoroughly for 10 s and then kept without moving for 4 h at room temperature. Optical densities at 600 nm were measured (Bioscreen C Analysis System) for each mixture at different (1st and 4th h) times. PBS was used as the blank. The percentages of autoaggregation and coaggregation were calculated according to the following Eqs. [46,47,48] :
Results
LAB strains and origins
A total of 11 LAB strains selected for the study were isolated from healthy honeybee samples collected from 10 different apiary locations in Turkey. The strains were named and numerated (Table 1).
Morphologic and biochemical properties of LAB strains
Of the 11 strains indicated to be Gram positive bacteria, all were of the gamma hemolytic type, and all were lactobacilli colonies with off-white or creamy colors, except SYM-9. While strains SYM-8 and SYM-9 showed positive citrate utilization, strains SYM-7, SYM-2, SYM-11 and SYM-3 showed positive cellulase production. Moreover, strains SYM-2, SYM-10 and SYM-11 were determined as lecithinase-producing strains. Eight of the strains grown on TMB-Plus medium were evaluated as H2O2-producing bacteria in view of the fact that dark gray/bluish halos were observed around their colonies. Although all isolates were capable of hydrolyzing starch, none were able to hydrolyze esculin. In general, the strains (other than three strains) have a wide temperature range, especially 5 of strains (SYM-1, SYM-5, SYM-7, SYM-4, and SYM-3) at a much wider temperature (18 - >45 °C) were observed to be able to grow. The optimum growth temperature for all strains was determined as 37 °C, and tolerance to high and low temperature is a strain-specific feature as shown in Table 2.
In the fermentation test, strains that were capable of utilizing a range of carbohydrates (glucose, fructose, galactose, maltose, sucrose, lactose, mannitol, rhamnose, arabinose, trehalose, melibiose, cellobiose, and xylose) as a source in order to produce acidic byproducts were exhibited. It was found that seven strains were capable of utilizing xylose, and three were capable of utilizing lactose. None were able to ferment the trehalose, with the exception of SYM-8. The SYM-7 strain was the only one unable to fermenting melibiose, arabinose and rhamnose (Table 3)
Molecular identification of bacteria
Phylogenetic research pointed out the presence of 11 different phylotypes of LAB. According to the constructed dendrogram, the frequency of the species closely related with the strains studied was Apilactobacillus kunkeei with 72%, Fructobacillus fructosus with 18% and Leuconostoc mesenteroides with 9% (n = 11) (Fig. 1). All strains were honeybee-associated bacteria proven by previous studies [10, 49,50,51,52,53].
The phylogenetic tree of LAB strains isolated from honeybees and their closely-related neighbors from other LAB strains (The evolutionary history was inferred using the Neighbor-Joining method. The percentage of replicate trees in which the associated taxa were clustered together in the bootstrap test (1000 replicates) were shown next to the branches. Evolutionary analyses were constructed in MEGA11. The scale below the figure shows the degree of similarity.)
Antimicrobial activity
Most of the LAB strains exhibited antimicrobial activity on all pathogens tested, according to the results of agar plug diffusion studies; however, the degree of antagonism differed from strain to strain. The highest inhibition activity was observed against Escherichia coli ATCC 25,922, Salmonella enteritis ATCC 13,076, and Bacillus cereus ATCC 43,288. The seven strains were the most effective ones in inhibiting these pathogens. Only one strain displayed an inhibition effect on the growth of Staphylococcus aureus ATCC 29,213 and Pseudomonas aeruginosa ATCC 27,853. The strains with the least antimicrobial activity were determined as SYM-3 and SYM-11. Antagonistic effect of any strain against Candida albicans ATCC 60,193 was not detected. Additionally, CFSs obtained from LAB strains were subjected to agar well diffusion test to ascertain the inhibition efficiency against selected indicator microorganisms in this study. The findings from this experiment indicated that pathogenic microorganisms most affected by LAB-CFSs were Mycobacterium smegmatis RSKK 607, Helicobacter pylori J99, Salmonella enteritis ATCC 13,076, Escherichia coli ATCC 25,922, and Listeria monocytogenes ATCC 43,251, respectively. CFSs of SYM-1 and SYM-5 were the substances with the highest inhibitory efficiency, in particular against Mycobacterium smegmatis RSKK 607 and Helicobacter pylori J99 (Table 4).
Preliminary assessment of probiotic properties
Probiotic selection criterion were based on the ability of the LAB strains to reach the intestine by enduring the acidic pH of the human stomach, as well as to be able to maintain viability in adverse conditions such as pH changes and exposure to bile salts in the intestine [43, 44]. In this context, after each of the strains exposed to PBS at pH 3.0 and 7.8 for 3 h and to the Bile-MRS broth for 4 h was incubated in standard MRS agar medium, their growth activities were measured in terms of log cfu ml− 1 and compared with the negative control. At the end of the incubation period, the bacterial colonies in all petri dishes were counted and the cell densities in 1 mL were calculated. It was determined that the strains that could not resist the pH change developed sensitivity from the 0th hour, while the strains that could not resist the bile were initially able to tolerate the bile when incubated in Bile-MRS, but growth was inhibited after longer exposure. The six strains (SYM-8, SYM-1, SYM-7, SYM-10, SYM-4, and SYM-3) were found to be resistant to pH changes and bile salt. While the strains SYM-2 and SYM-5 were highly able to survive at pH 3.0 and pH 7.8, no viable counts were observed in the inoculated MRSA after exposure to 0.3% bile for 4 h (Fig. 2). The growth curves of LAB strains were created by monitoring the activity in Bile-MRS broth and standard MRSB over a period of time (20-h). LAB strains were able to maintain their proliferation activity to a degree in the Bile-MRS broth, compared to the control groups. The bacterial growth-delaying effect of bile salt was significantly observed from the third hour of incubation. In addition, the growth activity of strain SYM-8, cultivating for 20 h in Bile-MRS broth medium, showed the closest value to the negative control group grown in MRSB. In the strains left to grow in the Bile-MRS medium, delays are observed in the logarithmic growth phase starting from the 3rd hour compared to the control groups. It has been determined that the bacterial growth retarding effect of bile salt can be observed significantly after 2–4 h of incubation, depending on the strains (Figs. 2 and 3).
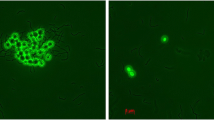
Another criterion addressed to test the probiotic potential of the studied LAB strains was the aggregation ability of the bacteria. Aggregation ability was classified under two groups as autoaggregation and coaggregation. In the analyses performed according to photometrically measured at 600 nm (OD600 = 0.1–0.2), it was determined that the autoaggregation abilities of LAB strains varied between 5.55% and 37.5% for four hours. Results were interpreted according to the cut-off levels proposed by Collado et al. and Suwannaphan [56, 57]. Enterococcus faecium was used as a positive control. The highest increase value was determined for strain SYM-3 (14.7 to 30.76%), and the lowest value was determined for strain SYM-2 (30.3 to 31.31%). The results showed that six strains had remarkable autoaggregation ability and could therefore display good adhesion and competitive activity. The ability to co-aggregate, known as the multicellular colonies formed by the different species by attaching to each other, was examined on Listeria monocytogenes ATCC 43,251 and Escherichia coli ATCC 25,922. The results compared with the values from the positive control group clearly indicated that 45% of the strains were found to coagulate well with E. coli, while 64% were found to have significant coaggregation abilities on L. monocytogenes [56, 57] (Table 5).
One of the main expected targets of the in vitro safety feature of a potential probiotic strain is the absence of bacteriophage infection. Our bacteriophage detection experiment demonstrated that none of our strains did not have phages and was not host (immune) as a result of cross-infection. The occurrence of phage infection between the phages of one module and hosts of another module or the same module is defined as cross-infection [58]. In other words, these strains were not infected by phage when used as hosts, and lysates from same strains were also phage-free. Thus, it was confirmed that all strains were selected from the strains that lack the phage in order to administer the tests of probiotic assessment.
Discussion
Lactic acid bacteria (LAB) represent the group of microorganisms in which the mechanisms of action and utilization efficiency of probiotics are most extensively studied [23]. Many of the lactic acid bacteria that have major nutritional and health benefits are important components of the human and animal microbiota [59, 60]. Many studies have found that LAB has beneficial effects on host health, balances gut flora, can be used as a biotherapeutic or prophylactic agent, and can be studied as new potential strains from a variety of sources [14, 21, 22, 61,62,63,64,65].
This study investigated the probiotic properties of lactic acid bacteria, which are permanent or transient members of the microflora isolated from healthy honey bees as a result of human consumption. Microbial colonization of the honey bee is found in many components with which the bee comes into contact, such as honey, propolis, bee bread and beehive materials [9, 10, 66, 67]. Therefore, there is a possibility of direct or indirect exposure to these bacteria as a result of consuming honey and other bee products. The presence of LAB in honey bees has been extensively studied over the years. By examining the honey bee plant, Vásquez et al. discovered 13 species of bacteria representing the genera Lactobacillus and Bifidobacterium [68]. Among these species, Apilactobacillus kunkeei was found to be dominant.
In a study conducted by Olofsson and Vásquez on honey bees, five strains were found to be related to A. kunkeei, B. asteroides, and Bifidobacterium coryneforme as a result of phylogenetic analysis [9]. Iorizzo et al. identified 24 strains from honeybee A. mellifera ligustica, and Lact. plantarum, A. kunkeei, Lactococcus lactis, F. fructosus were found [7]. Rokop et al. isolated bacteria belonging to genera Lactobacillus and Fructobacillus from bee pollen [69]. In the current study, phylogenetic analysis using culture-based approaches and 16 S rRNA sequencing revealed the presence of strains related to A. kunkeei, F. fructosus and Leuconostoc mesenteroides in healthy honeybees and larvae from different regions of Turkey. The sequences of all isolated ones described in this paper can be accessed via GenBank (Table 1; Fig. 1).
Lactic acid bacteria are defined as Gram-positive, rod, cocci or coccobacillus, nonspore-forming (except Sporolactobacillus), immobile, acid-tolerant, catalase-negative, nitrate reduction-negative, microaerophilic microorganisms producing primarily lactic acid or sole fermentation product [23, 60, 70]. Morphological and biochemical analyzes are important to determine various characteristics of strains and to support species identification (Table 2). Temperature is an important parameter for LAB both in terms of the fermentation process and the metabolic functions of the microorganism. LAB can grow over a wide temperature range (2–53 °C) and generally reproduce optimally at 30–40 °C [71, 72] (Table 2).
Hydrogen peroxide (H2O2), also produced by many LAB species, has an important role in host-microbe interactions [73,74,75]. Besides killing microorganisms, hydrogen peroxide has an important role in host-microbe interactions by acting as incoming signal transduction or second messenger [76, 77]. Hydrogen peroxide production was positive in all but three strains tested. In this context, it can be assumed that the primary antimicrobially active metabolite of these strains, whose antimicrobial activities have been extensively tested, is H2O2 (Table 2). Sugar fermentation tests (Table 3) were found to be important in diagnosing LAB and were tested for this purpose in the study [23, 60, 72, 78]. One of the desired properties of probiotic bacteria selected for use in humans is to increase the use of lactose in the body. It is beneficial for health that bacteria increase the digestion of lactose in the environment and the metabolites formed as a result of lactose fermentation have an antimicrobial effect in the gastrointestinal tract [79]. Accordingly, the results were found to be compatible with the literature and could therefore be used for the preliminary diagnosis of the species. Nectar, water, pollen and varying proportions of monosaccharides from bee foods; glucose, fructose and disaccharide; consists of sucrose [80]. The intin layer, which is composed of polysaccharides in the pollen cytoplasm, contains cellulose, hemicellulose and pectin. Pollen cell walls mainly contain type I rhamnogalacturonan (RG-I) and homogalacturonan, which are the main components of pectin. In addition, pollen can contain many monosaccharides, including arabinogalactan, mannose, xylose, galactose, mannitol and arabinose [81, 82]. Certain carbohydrates are designated as toxic to honeybees because these insects lack suitable digestive enzymes [83, 84]. The main nutrients, simple sugars such as glucose and fructose, are absorbed in the midgut of the bee, and these potential energy sources can be fermented through microbial enzymatic activity [85, 86]. Given their ability to simultaneously participate in the breakdown of complex polysaccharides and metabolize toxic sugars, the role of lactic acid bacteria in improving nutritional tolerance in honeybees as well as maintaining the health of their hosts seems remarkable [87,88,89]. Since these strains were able to partially or fully utilize all carbohydrates tested (except trehalose and xylose) and ferment fructose well, they were accepted as bee microflora-associated bacteria. Some fructophilic lactic acid bacteria (FLAB) have recently been assigned to the genus Fructobacillus (F. fructosus, F. durionis, F. ficulneus, F. pseudoficulneus) and are capable of fermenting fructose found in fermented foods, flowers and fruits [90].
The use and applications of the probiotic LAB have increased tremendously over the past two decades. Extensive screening and characterization is very important in identifying potential probiotic strains. For a probiotic strain to be effective and classified as GRAS (Generally Recognized as Safe), it must have certain properties. The most important selection criteria for probiotics that can be recommended for human use are the ability to survive under difficult conditions of the gastrointestinal system such as gastric acid, bile salts, pancreatic secretions, to adhere to the target area and to have antimicrobial effects against pathogens. A good probiotic candidate should have the aggregation ability to prevent colonization of harmful microorganisms in the host. It should not have pathogenic, toxic, hemolytic or other significant side effects. LAB strains with easily inducible phages cannot be preferred as probiotic strains because the fermentation process can be disrupted by bacteriophage contamination. It is crucial to select probiotic cultures with natural immunity to bacteriophage infection during the probiotic treatment process or milk fermentation to reduce the effects of phage infection [91]. In addition to stating that probiotics intended for humans should be of human origin in view of species-related health effects [37, 92], some probiotic microorganisms widely used today are not of human origin, such as Saccharomyces boulardii [15, 21, 37, 93,94,95]. In this regard, a decrease in cell density was observed as a function of pH change simulating the gastrointestinal system of the strains tested for their probiotic properties (Fig. 2). While the pH of the empty stomach is 1.5-2.0, during the digestion period of about 2 h, the pH of the stomach increases to 3.0 depending on the type and amount of food ingested. Food digestion takes 5–6 h in the small intestine (pH 6.6 ± 0.5) and 12–24 h in the large intestine (pH 7.0 ± 0.7). In other words, the pH of the environment changes between 1.5 and 8.0 during the digestive process in the gastrointestinal tract [96]. On the other hand, one F. fructosus strain showed no viability at pH 7.8, although a significant decrease in cell density (80%) was observed at pH 3.0 (Fig. 2). Another important criterion that is often considered essential for the probiotic strain is the tolerance to the bile content, which ensures the survival of LAB in the small intestine [42, 97]. For the selection of human probiotics, a mean of 0.3% bile concentration in the medium is considered the critical concentration high enough to screen for resistant strains in vitro [45]. With the exception of two strains associated with A. kunkeei, all other LAB strains remained viable after bile salt digestion with an average inhibition of 43.6% (Fig. 2). The pH and bile tolerance of the strains are consistent with results from similar studies [97, 98]. In the analysis carried out to determine the tolerance to bile salts, both the viability of the bacteria when added to the standard medium after 0.3% bile treatment for 4 h and the growth activity in the presence of 0.3% bile for 20 h uninterruptedly, the cell viability of the tested strains was 94% compared to the control group. It was able to protect it with a yield of 50, and it was able to develop in the presence of bile. It has been proven that the strains inoculated directly into Bile-MRS and followed for 20 h in this medium have slowed growth, but that these strains exposed to bile in the previous experiment can preserve their viability. In the light of these data, it can be interpreted that long-term exposure to bile inhibits the growth of the bacterium and perhaps causes it to lose its viability, but when exposed to bile for 4 h, the bacterium tolerates these conditions and maintains its viability. In addition, a decrease in bacterial growth density was observed depending on the time spent in the bile content and the reduced amount of nutrients in the media during the 20-hour incubation, this amount was close to standard growth values (Fig. 3).
Autoaggregation is the ability of the same strains to come together to form bulks. The ability of lactic acid bacteria to autoaggregate is important in terms of adhesion to intestinal epithelial cells and barrier formation [46, 56, 99]. The autoaggregation function of probiotic microorganisms helps to enhance the host’s defense mechanisms in the gut. A potential probiotic strain is expected to have greater than 40% autoaggregation [57]. Juarez Tomas et al. showed in their study that temperature increase, growth medium, and medium pH change can influence the aggregation property [100]. The results prove that the strains have autoaggregation activity (Table 5).
Coaggregation is a process by which probiotic strains compete with pathogens, eliminate them and become dominant in their environment. It is a remarkable feature for treating the gut microbiota suffering from dysbiosis. In particular, microorganisms that have the ability to coaggregate with pathogens have a significant advantage over non-coaggregating microorganisms, which are more easily cleared from the gut environment [47, 56, 101,102,103]. Most of the strains in the study showed promising aggregation abilities against pathogens due to certain combinations. F. fructosus and A. kunkeei showed the highest percentage of coaggregation with E. coli over 4 h. Two strains associated with A. kunkeei exhibited the highest percentage of coaggregation with L. monocytogenes (Table 5). It turns out that the aggregation rates increase with longer incubation times (20–24 h) [48, 56, 100, 102, 104].
The ability to aggregate is not the only thing that makes lactic acid bacteria suitable probiotics for relieving pathogens. They must also exhibit antimicrobial properties. [104]. Various methods are used to determine the antimicrobial effects of microorganisms. Solid media are generally preferred in these methods and the effect of the test culture is to demonstrate inhibition of growth of an indicator strain [105]. The inhibitory effect of probiotics against pathogenic bacteria has been extensively investigated in many studies [14, 37, 42, 97, 102, 106,107,108]. Our honeybee-associated strains showed high antimicrobial activity, particularly against enteropathogenic bacteria such as E. coli and Sal. enteritis, and also moderate antimicrobial activity against Y. pseudotuberculosis, Kl. pneumoniae and L. monocytogenes. Furthermore, effective inhibition was detected on B. cereus, H. pylori and Myco. smegmatis (Table 4). For strains with probiotic potential, it is crucial to be resistant to pathogens such as Salmonella typhimurium, L. monocytogenes, H. pylori, Staph. aureus and to prevent their colonization and infection [109].
Although in vitro studies cannot fully prove a probiotic effect during the selection phase of potential probiotics, they can be used to characterize a possible potential mechanism, to determine the safety of probiotic microorganisms, and to ascertain other information regarding probiotic strains. Therefore, in vitro studies are the first and most important step in the evaluation of probiotics [15, 21, 95]. This should be followed by simulated, randomized, placebo-controlled in vivo studies in humans [15]. Based on the evaluation of the research in general, it was found that the selected strains presented probiotic properties, although these properties varied by genus, species, and strains. We propose these strains, primarily honeybee-associated Apilactobacillus strains, as potential candidates for further in vitro and in vivo research on their potential health benefits and application as novel probiotic additives.
CRediT authorship contribution statement
Şeyma Suyabatmaz devised and planned the project, and analysed the data under the supervision of Şengül Alpay Karaoğlu. Şeyma Suyabatmaz, Şengül Alpay Karaoğlu and Arif Bozdeveci performed the experiments. Rahşan Akpınar provided help with the samples, research tools, and equipment. The manuscript was written by Şeyma Suyabatmaz. Şengül Alpay Karaoğlu revised the manuscript. All authors have read and approved the submitted manuscript.
Data Availability
On reasonable request, all data will be provided.
References
Suyabatmaz Ş, Bozdeveci A, Karaoğlu ŞA (2020) Gastrointestinal Bacterial Flora in Honey Bees. U Bee J 20:19:97–113. https://doi.org/10.31467/uluaricilik.701170
Alberoni D, Gaggìa F, Baffoni L, Di Gioia D (2016) Beneficial Microorganisms for Honey Bees: Problems and Progresses. Appl Microbiol Biotechnol APPL 100:22:9469–9482. https://doi.org/10.1007/s00253-016-7870-4
Rada V, Machova M, Huk J, Marounek M, Duskova D (1997) Microflora in the Honeybee Digestive Tract: Counts, Characteristics and Sensitivity to Veterinary Drugs. Apidologie 28:357–365. https://doi.org/10.1051/apido:19970603
Evans J, Lopez D (2004) Bacterial Probiotics Induce an Immune Response in the Honey Bee (Hymenoptera: Apidae). J Econ Entomol 97:752–756. https://doi.org/10.1603/0022-0493(2004)097[0752:bpiair]2.0.co;2
Killer J, Kopec J, Mrazek J, Rada V, Dubna S, Marounek M (2010) Bifidobacteria in the Digestive Tract of Bumblebees. Anaerobe 16:165–170. https://doi.org/10.1016/j.anaerobe.2009.07.007
Anderson EK, Sheehan T, Eckholm BJ, Mott BM, DeGrandi-Hoffman G (2011) An Emerging Paradigm of Colony Health: Microbial Balance of the Honey Bee and Hive (Apis mellifera). Insect Soc 58:431. https://doi.org/10.1007/s00040-011-0194-6
Iorizzo M, Pannella G, Lombardi SJ, Ganassi S, Testa B, Succi M, Sorrentino E, Petrarca S, De Cristofaro A, Coppola R, Tremonte P (2020) Inter- and Intra-Species Diversity of Lactic Acid Bacteria in Apis mellifera ligustica Colonies. Microorganisms 8:1578. https://doi.org/10.3390/microorganisms8101578
Jones JC, Myerscough MR, Graham S, Oldroyd PB (2004) Honey Bee Nest Thermoregulation: Diversity Promotes Stability. Science 16:402–404. https://doi.org/10.1126/science.1096340
Olofsson TC, Vásquez A (2008) Detection and Identification of a Novel Lactic Acid Bacterial Flora within the Honey Stomach of the Honeybee Apis mellifera. Curr Microbiol 57:356–363. https://doi.org/10.1007/s00284-008-9202-0
Endo A, Salminen S (2013) Honeybees and Beehives are Rich Sources for Fructophilic Lactic Acid Bacteria. Syst Appl Microbiol 36:444–448. https://doi.org/10.1016/j.syapm.2013.06.002
Tamarit D, Ellegaard KM, Wikander J, Olofsson T, Vásquez A, Andersson SGE (2015) Functionally Structured Genomes in Lactobacillus kunkeei Colonizing the Honey Crop and Food Products of Honeybees and Stingless Bees. Genome Biol Evol 7:6:1455–1473. https://doi.org/10.1093/gbe/evv079
Silva MS, Rabadzhiev Y, Eller MR, Iliev I, Ivanova I, Santana WC (2017) Microorganisms in Honey. Honey Anal InTech 11:233–258. https://doi.org/10.5772/67262
Filannino P, Di Cagno R, Addante R, Pontonio E, Gobbetti M (2016) Metabolism of Fructophilic Lactic Acid Bacteria Isolated from the Apis mellifera L. Bee Gut: Phenolic Acids as External Electron Acceptors. Appl Environ Microbiol 82:23. https://doi.org/10.1128/AEM.02194-16
Vergalito F, Testa B, Cozzolino A, Letizia F, Succi M, Lombardi SJ, Tremonte P, Pannella G, Di Marco R, Sorrentino E, Coppola R, Iorizzo M (2020) Potential Application of Apilactobacillus kunkeei for Human Use: Evaluation of Probiotic and Functional Properties. Foods 9:111535. https://doi.org/10.3390/foods9111535
Fijan S (2014) Microorganisms with Claimed Probiotic Properties: An Overview of Recent Literature. Int J Environ Res Public Health 11:5:4745–4767. https://doi.org/10.3390/ijerph110504745
FAO/WHO (2006) Probiotics in Food: Health and Nutritional Properties and Guidelines for Evaluation. Report of a Joint FAO/WHO Expert Consultation on Evaluation of Health and Nutritional Properties of Probiotics in Food Including Powder Milk with Live Lactic Acid Bacteria, Cordoba, Argentina. FAO Food and Nutrition Paper 85, ISSN 0254–4725
Fong W, Li Q, Yu J (2020) Gut Microbiota Modulation: A Novel Strategy for Prevention and Treatment of Colorectal Cancer. Oncogene 39:4925–4943. https://doi.org/10.1038/s41388-020-1341-1
Masood MI, Qadir MI, Shirazi JH, Khan IU (2010) Beneficial Effects of Lactic Acid Bacteria on Human Beings. Crit Rev Microbiol 37:1:91–98. https://doi.org/10.3109/1040841X.2010.536522
Oranusi S, Braide W, Chinakwe E (2013) Probiotic Carrier Potential, Sensory Properties and Microbial Quality of Ugba (Pentaclethra Macrophylla) and Ogiri (Ricinus Communis). Int J Microbiol Res 5:476–481. https://doi.org/10.9735/0975-5276.5.5.476-481
Basavaraju B, Jamil K (2014) Identification and Characterization of Probiotics from New Sources. Int J Sci Res (IJSR) 3:6
George Kerry R, Patra JK, Gouda S, Park Y, Shin H-S, Das G (2018) Benefaction of Probiotics for Human Health: A review. J Food Drug Anal 26:3:927–939. https://doi.org/10.1016/j.jfda.2018.01.002
Savadogo A, Ouattara TAC, Traore SA (2007) Potential of Lactic Acid Bacteria in Human Nutrition. Food 1:1:79–84
Wedajo B (2015) Lactic Acid Bacteria: Benefits, Selection Criteria and Probiotic Potential in Fermented Food. J Prob Health 3:2. https://doi.org/10.4172/2329-8901.1000129
Tarrah A, Castilhos de J, Rossi RC, Duarte V, Ziegler S, Corich DR, Giacomini VA (2018) In vitro Probiotic Potential and Anti-cancer Activity of Newly Isolated Folate-Producing Streptococcus thermophilus Strains. Front Microbiol 9:22–14. https://doi.org/10.3389/fmicb.2018.02214
Kandler O, Weiss N (1986) Genus Lactobacillus. In: Sneath P (ed) Bergey’s Manual of Systematic Bacteriology, vol 2. William&Wilkins, Baltimore, pp 1209–1234
Golman G(1990) Streptococcus and Lactobacillus. In Principles of Bacteriology, Virology and Immunity. Eighth Ed., Parker MT, Collier LH, Duerden BI, (Eds.) London, Melborne, Auckland, 148
Alpay Ş(1999) Investigation of Bacteriophages and Bacteriocins in Vaginal Lactobacilli. Dissertation, Department of Microbiology, Institute of Health Sciences, Karadeniz Technical University
Bergey DH, Whitman BW, Vos P, De, Garrity MG, Jones D, Krieg NR, Ludwig W, Rainey FA, Schleifer KH (2009) Bergey’s Manual of Systematic Bacteriology, Volume 3: The Firmicutes. New York, pp 464–635
Garcia LS, Isenberg HD(2010) Biochemical Tests for the Identification of Aerobic Bacteria. In Clinical Microbiology Procedures Handbook, 3rd Edition, pp 503–642. ASM Press, Washington
Mannan SJ, Rezwan R, Rahman MS, Begum K (2017) Isolation and Biochemical Characterization of Lactobacillus species from Yogurt and Cheese samples in Dhaka Metropolitan Area. Bangla Pharma J 20:1:27–33. https://doi.org/10.3329/bpj.v20i1.32090
Sambrook J, Russell DW (2001) Rapid Isolation of Yeast DNA. In: Sambrook J, Russell DW (eds) Molecular Cloning, a Laboratory Manual. Cold Spring Harbor Laboratory Press, New York, pp 631–632
Saitou N, Nei M (1987) The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Tamura K, Nei M, Kumar S(2004) Prospects for Inferring Very Large Phylogenies by Using the Neighbor-Joining Method. Proceedings of the National Academy of Sciences (USA) 101:11030–11035. https://doi.org/10.1073/pnas.0404206101
Kılıç OA, Pavlova IS, Ma W-G, Tao L (1996) Analysis of Lactobacillus Phages and Bacteriocins in American Dairy Products and Characterization of a Phage Isolated from Yogurt. Appl Environ Microbiol 62:6:2111–2116. https://doi.org/10.1128/aem.62.6.2111-2116.1996
Trevors KE, Holley RA, Kempton AG (1983) Isolation and Characterization of a Lactobacillus plantarum Bacteriophage Isolated from a Meat Starter Culture. J Appl Bacteriol 54:281–288. https://doi.org/10.1111/j.1365-2672.1983.tb02618.x
Balouiri M, Sadiki M, Ibnsouda SK (2016) Methods for in vitro Evaluating Antimicrobial Activity: A Review. J Pharm Anal 6:2:71–79. https://doi.org/10.1016/j.jpha.2015.11.005
Ahmad A, Yap WB, Kofli NT, Ghazali AR (2018) Probiotic potentials of Lactobacillus plantarum Isolated from Fermented Durian (Tempoyak), a Malaysian Traditional Condiment. Food Sci Nutr 1–8. https://doi.org/10.1002/fsn3.672
Pelyuntha W, Chaiyasut C, Kantachote D, Sirilun S (2019) Cell-Free Supernatants from Cultures of Lactic Acid Bacteria Isolated from Fermented Grape as Biocontrol Against Salmonella typhi and Salmonella typhimurium Virulence via Autoinducer-2 and Biofilm Interference. https://doi.org/10.7717/peerj.7555. PeerJ
Rabe KL, Hillier SL (2003) Optimization of Media for Detection of Hydrogen Peroxide Production by Lactobacillus Species. J Clin Microbiol 41:7:3260–3264. https://doi.org/10.1128/JCM.41.7.3260-3264.2003
Maragkoudakis PA, Zoumpopoulou G, Miaris C, Kalantzopoulos G, Pot B, Tsakalidou E (2006) Probiotic Potential of Lactobacillus Strains Isolated from Dairy Products. Int Dairy J 16:3:189–199. https://doi.org/10.1016/j.idairyj.2005.02.009
Jacobsen CN, Rosenfeldt Nielsen V, Hayford AE, Møller PL, Michaelsen KF, Pærregaard A, Sandström B, Tvede M, Jakobsen M(1999) Screening of Probiotic Activities of Forty-Seven Strains of Lactobacillus spp. by in vitro Techniques and Evaluation of the Colonization Ability of Five Selected Strains in Humans. Appl Environ Microbiol 65:11:4949–4956. https://doi.org/10.1128/AEM.65.11.4949-4956.1999
Lee J, Yun HS, Cho KW, Oh S, Kim SH, Chun T, Kim B, Whang KY (2011) Evaluation of Probiotic Characteristics of Newly Isolated Lactobacillus spp.: Immune Modulation and Longevity. Int J Food Microbiol 148:2:80–86. https://doi.org/10.1016/j.ijfoodmicro.2011.05.003
Chung H, Kim Y, Chun S, Ji G (1999) Screening and Selection of Acid and Bile Resistant Bifidobacteria. Int J Food Microbiol 47(1–2):25–32. https://doi.org/10.1016/s0168-1605(98)00180-9
Çomak Göçer ME, Ergin F, Küçükçetin A (2016) Viability of Probiotic Microorganisms in Digestive System Models. Acad Food J 14:2
Chateau N, Deschamps AM, Sassi AH (1994) Heterogeneity of Bile Salts Resistance in the Lactobacillus Isolates of a Probiotic Consortium. Lett Appl Microbiol 18(1):42–44. doi:https://doi.org/10.1111/j.1472-765x.1994.tb00796.x
Kos B, Šušković J, Vuković S, Šimpraga M, Frece J, Matošić S (2003) Adhesion and Aggregation Ability of Probiotic Strain Lactobacillus acidophilus M92. J Appl Microbiol 94:6:981–987. https://doi.org/10.1046/j.1365-2672.2003.01915.x
Rinkinen M, Jalava K, Westermarck E, Salminen S, Ouwehand AC (2003) Interaction Between Probiotic Lactic Acid Bacteria and Canine Enteric Pathogens: A Risk Factor for Intestinal Enterococcus faecium Colonization? Vet Microbiol 92(1–2):111–119. https://doi.org/10.1016/s0378-1135(02)00356-5
Kang CH, Han SH, Kim Y, Paek NS, So JS (2017) In vitro Probiotic Properties of Lactobacillus salivarius MG242 Isolated from Human Vagina. Probiotics and Antimicrobial Proteins 10:2:343–349. https://doi.org/10.1007/s12602-017-9323-5
Moran NA (2015) Genomics of the Honey Bee Microbiome. Curr Opin Insect Sci 10:22–28. https://doi.org/10.1016/j.cois.2015.04.003
Budge GE, Adams I, Thwaites R, Pietravalle S, Drew GC, Hurst GDD, Tomkies V, Boonham N, Brown M (2016) Identifying Bacterial Predictors of Honey Bee Health. J Invertebr Pathol 141:41–44. https://doi.org/10.1016/j.jip.2016.11.003
Parichehreh S, Tahmasbi G, Sarafrazi A, Imani S, Tajabadi N (2018) Isolation and Identification of Lactobacillus Bacteria Found in the Gastrointestinal Tract of the Dwarf Honey Bee, Apis florea Fabricius, 1973 (Hymenoptera: Apidae). Apidologie 49:3:430–438. https://doi.org/10.1007/s13592-018-0569-z
Royan M (2019) Mechanisms of Probiotic Action in the Honeybee. Crit Rev Eukaryot 29:2:95–103. https://doi.org/10.1615/critreveukaryotgeneexpr.2019025358
Behare PV, Ali SA, McAuliff O (2020) Draft Genome Sequences of Fructobacillus fructosus DPC 7238 and Leuconostoc mesenteroides DPC 7261, Mannitol-Producing Organisms Isolated from Fructose-Rich Honeybee-Resident Flowers on an Irish farm. Microbiol Resour Announc 9:50. https://doi.org/10.1128/MRA.01297-20
Wang X, Wang W, Lv H, Zhang H, Liu Y, Zhang M, Wang Y, Tan Z (2021) Probiotics & Antimicro Prot 13:90–101. https://doi.org/10.1007/s12602-020-09658-3
Lazzeri AM, Mangia NP, Mura ME, Floris I, Satta A, Ruiu L (2020) Potential of Novel Food-borne Lactobacillus Isolates against the Honeybee Pathogen Paenibacillus larvae. Biocontrol Sci Technol 1–12. doi:https://doi.org/10.1080/09583157.2020.1769556
Collado MC, Meriluoto J, Salminen S (2008) Adhesion and Aggregation Properties of Probiotic and Pathogen Strains. Eur Food Res Technol 226:1065–1073. https://doi.org/10.1007/s00217-007-0632-x
Suwannaphan S (2021) Isolation, Identification and Potential Probiotic Characterization of Lactic Acid Bacteria from Thai Traditional Fermented Food. AIMS Microbiol 7:4:431–446. https://doi.org/10.3934/microbiol.2021026
Flores C, Valverde S, Weitz J (2013) Multi-scale structure and geographic drivers of cross-infection within marine bacteria and phages. ISME J 7:520–532. doi:https://doi.org/10.1038/ismej.2012.135
Muñoz R, Moreno-Arribas MV, Rivas de las B (2011) Lactic Acid Bacteria. Mol Wine Microbiol 8:191–226. https://doi.org/10.1016/B978-0-12-375021-1.10008-6
Zhang H, Cai Y (eds) (2014) Lactic Acid Bacteria and Animal Health. Lactic Acid Bacteria: Fundamentals and Practice 7:444–459. Springer
Klayraung S, Okonogi S (2009) Antibacterial and Antioxidant Activities of Acid and Bile Resistant Strains of Lactobacillus fermentum Isolated From Miang. Braz J Microbiol 40:757–766. https://doi.org/10.1590/S1517-83822009000400005
Pisano MB, Viale S, Conti S, Fadda ME, Deplano M, Melis MP, Deiana M, Cosentino S (2014) Preliminary Evaluation of Probiotic Properties of Lactobacillus Strains Isolated from Sardinian Dairy Products. BioMed Res Int 1–9. https://doi.org/10.1155/2014/286390
Azat R, Liu Y, Li W, Kayir A, Lin D, Zhou W, Zheng X (2016) Probiotic Properties of Lactic Acid Bacteria Isolated from Traditionally Fermented Xinjiang Cheese. J Zhejiang University-SCIENCE B 17:8. https://doi.org/10.1631/jzus.B1500250
Maldonado Galdeano C, Cazorla SI, Lemme Dumit JM, Vélez E, Perdigón G (2019) Beneficial Effects of Probiotic Consumption on the Immune System. Ann Nutr Metab 74:115–124. https://doi.org/10.1159/000496426
Aziz G, Tariq M, Zaidi AH (2021) Mining Indigenous Honeybee Gut Microbiota for Lactobacillus with Probiotic Potential. Microbiology 167:3:1465–2080. https://doi.org/10.1099/mic.0.001032
Powell JE, Martinson VG, Urban-Mead K, Moran NA (2014) Routes of Acquisition of the Gut Microbiota of the Honey Bee Apis mellifera. Appl Environ Microbiol 80:7378–7387. https://doi.org/10.1128/AEM.01861-14
Kwong WK, Moran NA (2016) Gut Microbial Communities of Social Bees. Nat Rev Microbiol 14:3:74–84
Vásquez A, Forsgren E, Fries I, Paxton RJ, Flaberg E, Szekely L, Olofsson TC (2012) Symbionts as Major Modulators of Insect Health: Lactic Acid Bacteria and Honeybees. PLoS ONE 7:3. https://doi.org/10.1371/journal.pone.0033188
Rokop ZP, Horton MA, Newton ILG (2015) Interactions Between Cooccurring Lactic Acid Bacteria in Honey Bee Hives. Appl Environ Microbiol 81:7261–7270. https://doi.org/10.1128/AEM.01259-15
Gomes AMP, Malcata FX (1999) Bifidobacterium spp. and Lactobacillus acidophilus: Biological, Biochemical, Technological and Therapeutical Properties Relevant for Use as Probiotics. Trends Food Sci Technol 10:4–5. https://doi.org/10.1016/S0924-2244(99)00033-3
Madigan TM and Martinko JM (2005) Brock Biology of Microorganisms (11th edn), Pearson Education, Prentice Hall, ISBN: 0-13-144329-1.
Linjordet SM (2016) A Comparative Analysis of Lactic Acid Bacteria Isolated from Honeybee Gut and Flowers, with Focus on Phylogeny and Plasmid Profile, Master Thesis Dissertation, 18–20. Department of Chemistry, Norwegian University of Life Sciences
Berthier F (1993) On the Screening of Hydrogen Peroxide-Generating Lactic Acid Bacteria. Lett Appl Microbiol 16:150–153. https://doi.org/10.1111/j.1472-765X.1993.tb01381.x
Ocan˜a VS, Pesce de Ruiz Holgado AA, Nader-Macias ME (1999) Selection of Vaginal H2O2-Generating Lactobacillus Species for Probiotic Use. Curr Microbiol 38:279–284. https://doi.org/10.1007/pl00006802
Ito A, Sato Y, Kudo S, Sato S, Nakajima H, Toba T (2003) The Screening of Hydrogen Peroxide-Producing Lactic Acid Bacteria and their Application to Inactivating Psychrotrophic Food-Borne Pathogens. Curr Microbiol 47:3:231–236. https://doi.org/10.1007/s00284-002-3993-1
Marinho HS, Real C, Cyrne L, Soares H, Antunes F (2014) Hydrogen Peroxide Sensing, Signaling and Regulation of Transcription Factors. Redox Biol 2:535–562. https://doi.org/10.1016/j.redox.2014.02.006
Van der Vliet A, Janssen-Heininger YMW (2014) Hydrogen Peroxide as a Damage Signal in Tissue Injury and Inflammation: Murderer, Mediator, or Messenger? J Cell Biochem 115:3:427–435. https://doi.org/10.1002/jcb.24683
Axelsson L, Ahrné S (2000) Lactic Acid Bacteria: Ed. In: Priest FG, Goodfellow M (eds) Applied Microbial Systematics. Springer, Netherlands, pp 367–388
Celikel A, Göncü B, Akın BM, Akın SM (2018) Süt Ürünlerinde Probiyotik Bakterilerin Canlılığını Etkileyen Faktörler. Batman University, Journal of Life Sciences, 8(1/2)
Taylor MA, Robertson AW, Biggs PJ, Richards KK, Jones DF, Parkar SG (2019) The effect of carbohydrate sources: Sucrose, invert sugar and components of mānuka honey, on core bacteria in the digestive tract of adult honey bees (Apis mellifera). PLoS ONE 14(12):e0225845. doi:https://doi.org/10.1371/journal.pone.0225845
Kochansky J, Knox DA, Feldlaufer M, Pettis JS (2001) Screening alternative antibiotics against oxytetracycline-susceptible and –resistant Paenibacillus larvae. Apidologie 32:215–222. https://doi.org/10.1080/00218839.2005.11101142
Li X, Gong H, Yang S, Yang L, Fan Y, Zhou Y (2017) Pectic bee pollen polysaccharide from Rosa rugosa alleviates diet-induced hepatic steatosis and insulin resistance via induction of AMPK/mTOR-mediated autophagy. Molecules 22(5):699. doi: https://doi.org/10.3390/molecules22050699
Barker RJ, Lehner Y (1974) Influence of diet on sugars found by thin-layer chromatography in thoraces of honey bees, Apis mellifera L. J Exp Zool 188:157–164
Johnson RM (2015) Honey bee toxicology. Ann Rev Entomol 60:415–434. doi: https://doi.org/10.1146/annurev-ento-011613-162005
Crailsheim K (1988) Regulation of food passage in the intestine of the honeybee (Apis mellifera L.). J Insect Physiol 34(2):85–90. doi: https://doi.org/10.1016/0022-1910(88)90158-8
Roulston TH, Cane JH (2000) Pollen nutritional content and digestibility for animals. Plant Syst Evol 222:187–209
Lee FJ, Rusch DB, Stewart FJ, Mattila HR, Newton IL (2015) Saccharide breakdown and fermentation by the honey bee gut microbiome. Environ Microbiol 17(3):796–815
Zheng H, Nishida A, Kwong WK, Koch H, Engel P, Steele MI, Moran NA (2016) Metabolism of toxic sugars by strains of the bee gut symbiont Gilliamella apicola. MBio 7(6):e01326–e01316
Ricigliano VA, Fitz W, Copeland DC, Mott BM, Maes P, Floyd AS, Dockstader A, Anderson KE (2017) The impact of pollen consumption on honey bee (Apis mellifera) digestive XVI physiology and carbohydrate metabolism. Archives of Insect Biochemistry and Physiology, 96(2), e21406
Ramos OY, Basualdo M, Libonatti C, Vega MF (2019) Current Status and Application of Lactic Acid Bacteria in Animal Production Systems with a Focus on Bacteria from Honey Bee Colonies. J Appl Microbiol 128:1248–1260. https://doi.org/10.1111/jam.14469
Ventura M, Sozzi T, Turroni F, Matteuzzi D, van Sinderen D (2010) The Impact of Bacteriophages on Probiotic Bacteria and Gut Microbiota Diversity. Genes & Nutrition 6(3):205–207. doi:https://doi.org/10.1007/s12263-010-0188-4
Dunne C (2001) Adaptation of Bacteria to the Intestinal Niche: Probiotics and Gut Disorder. Inflamm Bowel Dis 7(2):136–145. https://doi.org/10.1097/00054725-200105000-00010
Uymaz B (2010) Probiotics and their Use. Pamukkale Univ J Engineering Sci 16:1:95–104
Garneau JE, Moineau S (2011) Bacteriophages of Lactic Acid Bacteria and their Impact on Milk Fermentations. Microb Cell Fact 10. https://doi.org/10.1186/1475-2859-10-S1-S20. :1:S20
Fontana L, Bermudez-Brito M, Plaza-Diaz J, Muñoz-Quezada S, Gil A (2013) Sources, Isolation, Characterisation and Evaluation of Probiotics. Br J Nutr 109:S2:S35–S50. https://doi.org/10.1017/S0007114512004011
Elcioglu O, Kunduhoglu B (2014) Probiotic Characteristics of Natural Lactobacilli Isolated from Traditional Kargi Tulum Cheese. Ital J Food Sci 26:1:31–40
Raghavan KT, Jacob AA, Chandran H (2013) Honey bee Gut flora as a Source of LAB (Lactic Acid Bacteria) with Probiotic Capabilities. J Food Technol 105:126–134
Elzeini HM, Ali ARAA, Nasr NF, Hassan M, Hassan AAM, Elenany YE (2021) Probiotic Capability of Novel Lactic Acid Bacteria Isolated from Worker Honey Bees Gut Microbiota. FEMS Microbiol Lett 368:6. https://doi.org/10.1093/femsle/fnab030
Del Re B, Sgorbati B, Miglioli M, Palenzona D (2000) Adhesion, Autoaggregation and Hydrophobicity of 13 Strains of Bifidobacterium longum. Lett Appl Microbiol 31:6:438–442. https://doi.org/10.1046/j.1365-2672.2000.00845.x
Juarez Tomas MS, Wiese B, Nader-Macias ME (2005) Effects of Culture Conditions on the Growth and Auto-aggregation Ability of Vaginal Lactobacillusjohnsonii CRL 1294. J Appl Microbiol 99:6:1383–1391
Nikolic M, Jovcic B, Kojic M, Topisirovic L (2010) Surface Properties of Lactobacillus and Leuconostoc Isolates from Homemade Cheeses Showing Auto-aggregation Ability. Eur Food Res Technol 231:6:925–931. https://doi.org/10.1007/s00217-010-1344-1
Zhang W, Liu M, Dai X (2013) Biological Characteristics and Probiotic Effect of Leuconostoc lactis Strain Isolated from the Intestine of Black Porgy Fish. Braz J Microbiol 44:3:685–691. https://doi.org/10.1590/S1517-83822013005000053
Bilginer H, Çetin B (2019) Probiotics and in vitro Tests Used for their Determination. Atatürk Univ J Agric Fac 50:3:312–325. https://doi.org/10.17097/ataunizfd.549552
Pachla A, Ptaszyńska AA, Wicha M, Kunat M, Wydrych J, Oleńska E, Małek W (2021) Insight into Probiotic Properties of Lactic Acid Bacterial Endosymbionts of Apis mellifera L. Derived from the Polish Apiary. Saudi J Biol Sci 28:3:1890–1899. https://doi.org/10.1016/j.sjbs.2020.12.040
Coman MM, Verdenelli MC, Cecchini C, Silvi S, Orpianesi C, Boyko N, Cresci A (2014) In vitro Evaluation of Antimicrobial Activity of Lactobacillus rhamnosus IMC 501, Lactobacillus paracasei IMC 502 and SYNBIO against Pathogens. J Appl Microbiol 117:518–527. https://doi.org/10.1111/jam.12544
Sirilun S, Chaiyasut C, Kantachote D, Luxananil P (2010) Characterisation of Non Human Origin Probiotic Lactobacillus plantarum with Cholesterol-Lowering Property. Afr J Microbiol Res 4:10:994–1000
Iorizzo M, Testa B, Lombardi SJ, Ganassi S, Ianiro M, Letizia F, Succi M, Tremonte P, Vergalito F, Cozzolino A, Sorrentino E, Coppola R, Petrarca S, Mancini M, De Cristofaro A (2020) Antimicrobial Activity against Paenibacillus larvae and Functional Properties of Lactiplantibacillus plantarum Strains: Potential Benefits for Honeybee Health. Antibiotics 9:8442. https://doi.org/10.3390/antibiotics9080442
Ebrahimi M, Sadeghi A, Rahimi D, Purabdolah H, Shahryari S (2021) Postbiotic and Anti-aflatoxigenic Capabilities of Lactobacillus kunkeei as the Potential Probiotic LAB Isolated from the Natural Honey. Probiotics Antimicrob Proteins 13:2:343–355. https://doi.org/10.1007/s12602-020-09697-w
Sung-Mee L, Dong-Soon I (2009) Screening and Characterization of Probiotic Lactic Acid Bacteria Isolated from Korean Fermented Foods. J Microbiol Biotechnol 19:2:178–186. https://doi.org/10.4014/jmb.0804.269
Funding
This study was supported and funded by RTEU Scientific Research Projects (FYL-2019-1058).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Declaration of competing interest
No conflict of interest declared.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Suyabatmaz, Ş., Karaoğlu, Ş.A., Bozdeveci, A. et al. Honeybee-associated lactic acid bacteria and their probiotic potential for human use. World J Microbiol Biotechnol 39, 2 (2023). https://doi.org/10.1007/s11274-022-03427-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03427-w