Abstract
A growing tumor undergoes processes of heterogenisation and selection resulting in a complex system that facilitates further growth and progression. Immunosuppressive conditions belong to the hallmarks of this microenvironment, as they prevent efficient anti-cancer immunity. Eligible danger signal seems to be the missing link in the chain of events that would lead to successful elimination of cancer. The concept of bacterial cancer therapy is based on the ability of some microbial species to target tumor site and activate the cancer-specific response via pathogen-associated immunostimulatory signals. To date, a number of bacterial species have been shown to colonize tumor tissue, but strains of Salmonella are particularly interesting, since they meet all the requirements for an ideal tumor-targeting agent: a motile, facultatively anaerobic, intracellular microorganism that is prone to genetic manipulations. The most promising, S. Typhimurium, has multiple adaptations that are therapeutically relevant, including broad host specificity, specialized secretion systems and virulence factors with proapoptotic and immunomodulatory properties. Attenuation of wild-type strains has rendered them safe for preclinical and clinical use, while additional genetic modifications can add to their capacity to kill tumor cells and stimulate anti-cancer immunity. Given the recent developments in the field and the spectrum of possibilities offered by S. Typhimurium and its derivatives, it has a good chance of becoming a novel tool in the anticancer toolbox.
Dedication
In memory of Michał Bereta, who ‘infected’ us with the idea of therapeutic Salmonella
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
- Bacterial cancer therapy
- Immunotherapy
- Danger signal
- Cancer vaccine
- Tumor targeting
- Salmonella
- Typhimurium
- Attenuation
How Cancer Avoids Immunity
Cancer Cells May Elicit Immune Response – But Too Little and Too Late
When discussing the relationship between cancer and the immune system, we should bear in mind that the immune system has not evolved particularly to recognize and eliminate cancer cells. The roles of the immune system are to provide defense against pathogens, remove dying cells of the organism and mediate tissue healing. However, due to certain common elements of pathogen invasion and cancer growth, immune system is able to mount a specific, although usually ineffective, anti-tumor response (Goldszmid et al. 2014) and this fact is the basis for the belief that the development of active immunotherapies is possible.
Despite the nature of the trigger, carcinogenesis is considered a slow process, the first stages of which remain frequently ignored by the immune system because the initially mutated cells do not usually express immunogenic tumor-associated antigens (TAAs) and more importantly no danger signal accompanies the early stage of tumor development (Whiteside 2010; Chow et al. 2012). Dendritic cells (DCs) are necessary for the initiation of the adaptive immune response. Unlike immature DCs which phagocytose tissue antigens but are indolent in their processing and presentation, mature DCs are the most potent antigen presenting cells and have the ability to efficiently promote differentiation of effector T cell s . In order to mature fully, DCs must encounter a danger signal: pathogen- or damage-associated molecules that interact with and stimulate Toll-like receptors and/or NOD-like receptors (Chow et al. 2012; Whiteside 2010).
The mechanism of early recognition of transformed cells is not known. It seems reasonable to expect that at the moment of recognition cancer cells already produce mutated proteins or proteins with modified epitopes or express carcinoembrional proteins that may play a role of tumor-specific antigens. It is hypothesized that the danger signal might be provided by dying tumor cells. According to the immunosurveillance hypothesis, the early recognition of transformed cells may lead to eradicating of developing tumors (Chow et al. 2012). However, the extent of this phenomenon is difficult to estimate.
Tumor Growth and Selection Result in a Highly Immunosuppressive Microenvironment
It is believed that frequently, due to weak immunogenicity of TAAs and delayed appearance of a danger signal, the anti-tumor adaptive immune response arises too late and is of too low magnitude to efficiently eradicate tumor cells. The induction of the specific but inefficient anti-tumor immune response results in the so-called cancer immunoediting , the evolutionary process that leads to the selection of the genetically fittest tumor cells. These may further progress into a highly aggressive, malignant population (Dunn et al. 2002; Chow et al. 2012).
The genetic and epigenetic changes that enable tumor escape from the immune system are twofold – they make tumor cells less sensitive to the effects of the immune system (e.g. through increased expression of anti-apoptotic proteins, decreased expression of MHC class I molecules) but they also disarm the immune system by inducing and supporting immunoinhibitory tumor microenvironment . Numerous tumors gain the ability to secrete molecules that limit their infiltration by the immune cells as well as disturb the immune cells maturation and activation.
It is worth noting that the tumor growth itself continuously modulates tumor microenvironment . Hypoxia, a drop in pH due to increased levels of lactate (the glycolytic end-product) (Choi et al. 2013), as well as the accumulation of extracellular adenosine are all hallmarks of tumor growth and all exert immunosuppressive effects by influencing the activities of the immune cells present in tumor millieu (Gabrilovich et al. 2012).
Hypoxia in the tumor tissue leads to necrosis, which induces the recruitment of monocytes and DCs to the site and triggers inflammatory response. Inflammation has paradoxical roles in tumor development (Balkwill and Mantovani 2012). Almost all proinflammatory cytokines and factors (TNF, IL-1, IL-6 , nitric oxide) have been shown by numerous studies to have antitumor as well as tumor promoting activities. The net outcome of tumor-associated inflammation depends on the stage of the mutual relationship between tumor cells and the immune cells present in tumor microenvironment .
Cancer -Associated Immune Cells Facilitate Tumor Progression and Afflict Immunity
Similarly to the differences in phenotypic subsets of the immune cells which participate in the triggering and resolution of inflammation or during the conversion of acute to chronic inflammation , also diverse immune cell populations, often with opposite activities, are seen in tumor and tumor inflammatory microenvironment at different stages of its development.
However, in established tumors, most of the tumor-infiltrating immune cells will demonstrate immunosuppressive and tumor-promoting phenotype. Among them tumor-associated macrophages (TAMs), myeloid-derived suppressor cells (MDSCs) and T regulatory cell s (Tregs) seem to play a crucial role in tumor progression.
Several lines of evidence indicate that in general TAMs have M2-like phenotype, characterized by a high expression of IL-10 and low expression of IL-12. Immunosuppressive activities of TAMs are also attained by the synthesis of TGF-β, iNOS-derived nitric oxide, indoleamine 2,3-dioxygenase (IDO) – a tryptophan-degrading enzyme, and B7-H1, B7-H3 and B7-H4 molecules – negative regulators of T cell immunity. TAMs may aid tumor cell proliferation by expression of growth factors such as EGF, bFGF, VEGFs and PDGF. The growth factors, including VEGF-A, C and D, together with TAM-derived matrix metalloproteases (MMP2, MMP9), and IL-8 (CXCL8) are also responsible for tumor angiogenesis and lymphangiogenesis, which enable tumor growth and spreading. MMPs and other proteases produced by TAMs such as uPA, plasmin and cathepsin B play a double role; they activate and/or release growth factors entrapped in extracellular matrix (ECM) and are responsible for the degradation of the basement membrane and ECM remodeling, thus facilitating tumor invasion and metastasis (Gabrilovich et al. 2012; Whiteside 2010; Galdiero et al. 2013). TAMs also produce PGE2, a prostaglandin with a broad immunoregulatory activity (Kalinski 2012). PGE2 suppresses Th1 cell-mediated immunity and modulates chemokine production, inhibiting recruitment of CD8+, NK and Th1 cells while enhancing attraction of Tregs and MDSCs (Kalinski 2012).
MDSCs are a highly heterogeneous population of cells that can be subdivided into two major groups: monocytic (M-MDSCs) and granulocytic (G-MDSCs) (Youn and Gabrilovich 2010). MDSCs are defined as cells of myeloid origin characterized by the lack or reduced expression of markers of mature myeloid cell s with a strong capacity to suppress cytotoxic T-cell responses (Gabrilovich et al. 2007). Mouse MDSCs are CD11b+Gr1+ and human MDSCs are CD11b+CD33+ and some subsets are CD15+, CD34+ or CD31+. MDSCs, recruited from the bone marrow by tumor-derived attractants, migrate to tumor tissue and to lymph nodes, where they impede activation of T cell s by DCs. MDSCs share certain immunosuppressive instruments with other cells; e.g. they produce TGF-β and IL-10. What is unique, M-MDSCs express simultaneously high levels of iNOS and arginase 1 (ARG1), which degrade L-arginine, the iNOS substrate (Gabrilovich et al. 2012). Upon L-arginine depletion iNOS produce both NO and superoxide radicals that interact to form peroxynitrite (Xia and Zweier 1997). This potent oxidant affects proliferation, migration and effector functions of T cells. For G-MDSCs, prolonged synthesis of reactive oxygen species (ROS), due to high expression of the NOX2 NADPH oxidase, is the major mechanism of immunosuppression . MDSCs also exert their effects through supporting the development of Tregs populations. Mouse MDSCs have been shown to promote clonal expansion of antigen-specific Tregs and conversion of CD4+ lymphocytes into induced Tregs (Gabrilovich et al. 2012).
Tregs (CD4+CD25+FoxP3+) represent a population of immune cells whose physiological role is to prevent autoimmunity and to maintain peripheral tolerance. Tregs support tumor growth by inhibiting activity of tumor-infiltrating effector T cells, DCs, and NK cells. They exert their effects both by contact-dependent suppression through FasL – Fas and PD1 – B7-H1 interactions and by secretion of soluble factors such as TGF-β and IL-10 (Biragyn and Longo 2012).
Also the development and activation of DCs are frequently impaired by tumor microenvironment . DCs in tumors and in tumor-draining lymph nodes show decreased ability to process antigens and to stimulate tumor-specific T-cell responses and what is more, they gain immunosuppressive phenotype (Ma et al. 2013).
It seems that in established tumors, tumor cells and various tumor-infiltrating immune cells have an intense cross-talk, instruct each other and fully cooperate to promote tumor growth. However, the existence of TAAs and the fact that at some stage of tumor development the immune system mounts both humoral and cellular anti-tumor response makes researchers believe that immunotherapies, which destroy the unfavorable balance between a high suppressive and a low activatory response may lead to tumor eradication and prevent its recurrence. This belief may be emphasized by the results of studies involving over 600 patients suffering from colorectal cancer that demonstrated an overwhelming correlation between long term survival of patients after curative surgery and the so called immunoscore reflecting the number of CD8+ and CD45RO activated T cell s in the tumor sections. In 5 years after surgery >86 % of patients with immunoscore 4 (the highest number of analyzed T cells), and only 27.5 % of patients with immunoscore 0 or 1 were still alive (Pages et al. 2009). Thus, it seems that the removal of tumor enabled existing effector cells to control residual disease. There are more examples of a correlation between strong T cell infiltration of different cancers of epithelial origin and good clinical outcomes (Fridman et al. 2012).
The design and desired activities of a growing number of immunotherapeutic drugs and procedures already approved for cancer treatments and those under development include: (i) inhibition of tumor cell growth and induction of tumor cell death by antibody-dependent cell-mediated cytotoxicity (ADCC), complement-dependent cytotoxicity (CDC) and antibody-dependent cellular phagocytosis (ADCP) using TAA-specific monoclonal antibodies (mAb); (ii) expansion and activation of tumor-specific T cells through an adoptive transfer of autologous tumor-infiltrating lymphocytes (TILs) or genetically-engineered T cells as well as by application of anti-CTLA-4 or anti-PD1 mAb; (iii) improvement of TAA presentation and a strength of costimulatory signals utilizing DC-based vaccines ; and (iv) affecting immunosuppressive tumor environment by small-molecule inhibitors of iNOS, ARG1 and IDO.
Another approach to tumor immunotherapy employs viral or bacterial infection . This more holistic strategy aims to simultaneously act on diverse components of the immune response and may have a number of therapeutic advantages, which are highlighted below.
Rationale for Bacterial Cancer Therapy
Bacteria Can Deliver the Danger Signal Needed for Efficient Cancer -Specific Immune Response
Since the discoveries of alpha-fetoprotein and carcinoembryonic antigen in the mid-twentieth century, a still-increasing number of TAAs has been identified as potential targets for vaccine strategies. However, due to poor antigenicity and self-tolerance, the antigen-based approach suffered major setbacks in the clinic. A growing body of evidence shows that the immune response against tumor antigens requires additional stimulation which would be able to break the tolerance towards the tumor and activate a robust, clinically meaningful antitumor immunity. Hence, the danger signal seems to be the missing link in the chain of events that would lead to successful cancer immunotherapy (Matzinger 2012). Efficient immunostimulation, including antigen-presenting cells (APCs) activation that is a starting point for primary and secondary immune responses, requires the presence of a danger signal – a molecular trigger that can be of either endo- or exogenous nature. Cells and tissues undergoing stress, damage or necrotic death release a number of factors that can alarm the immune system, including heat-shock proteins, nucleotides, mitochondria-derived molecules, cytokines and products of extracellular matrix degradation; collectively, these factors are referred to as damage-associated molecular patterns (DAMPs). However, the same pattern-recognition receptors that detect DAMP-mediated danger signals can be also triggered by exogenous molecules derived from pathogens, known as pathogen-associated molecular patterns (PAMPs). This term covers a diverse group of compounds, including microbial and viral DNA with non-methylated CpG sites, as well as mannans, flagellin and bacterial cell wall components like lipopolysaccharide (LPS) and peptidoglycan. The pathogen-derived mediators have a potent immunostimulatory effect – upon interaction with pattern-recognition receptors including Toll-like receptors (TLRs), they strongly activate innate- and initiate adaptive immune responses. Not surprisingly, TLR ligands have been identified as potential cancer therapeutics with a few agents from this group already approved for human use. Up to date, those include: bacillus Calmette-Guérin (BCG), an attenuated strain of Mycobacterium bovis; picibanil (OK-432), a lyophilized preparation of Streptococcus pyogenes; monophosphoryl lipid A (MPL), a detoxified LPS derivative of Salmonella minnesota; and imiquimod (R-837), a synthetic imidazoquinoline. The drugs are followed by a growing number of novel candidates (Vacchelli et al. 2013), however, the efficacy of TLR agonists as single agents for cancer therapy has so far been limited. Moreover, the activation of TLRs seems to be a double-edged sword, as – depending on the tissue context and tumor type – it can promote tumor growth rather than the antitumor response (Lu 2014). For example, prolonged activation of TLR4 via signaling by both PAMPs and DAMPs creates tumor-promoting microenvironment that includes immunosuppressive cytokines, as well as cell populations like Tregs and MDSCs (Mai et al. 2013). However, TLR ligands delivered with an infection have been proved to be clinically effective – intravesical treatment with BCG, which operates as a mixed TLR2/TLR4 agonist, is currently the gold standard of care in non-muscle invasive bladder cancer (Sylvester 2011). Bacterial infection within the tumor tissue can evoke both the damage- and the pathogen-related danger signals; it has been shown that microorganisms within the tumor tissue can indeed promote an inflammatory reaction and potentiate the anti-tumor host response (Avogadri et al. 2005). Notably, bacterial cancer therapies offer many more mechanisms of action than solely the immunostimulatory effect of TLR activation – the most important being the ability to directly influence the suppressive microenvironment.
Many Features of Bacteria Provide Benefits for Cancer Treatment
Successful delivery of the danger signal into the tumor tissue is possible due to a number of bacterial mechanisms that can be therapeutically relevant. Bacteria-based treatments can benefit from microbial metabolism, motility and sensitivity to address the key issues in cancer therapy: low specificity towards cancer tissue and insufficient penetration of the tumor, both of which are limiting to currently used treatment modalities. Cancer cells form a complex and heterogeneous system that is poorly accessible to chemotherapeutic drugs (Saunders et al. 2012), while motile microbial organisms are able to cross biological barriers, act against hemodynamic gradients and preferentially accumulate in the tumor tissue. In contrast to passively-diffusing therapeutics, which produce relatively large drug concentrations in the bloodstream and relatively low drug concentrations in the tumor, bacteria offer unique mechanisms that can facilitate site-specific treatment, highly focused on the tumor and safe to other tissues. The natural ability of bacteria to receive signals via chemoreceptors can be used as a tool to effectively target the unique microenvironment formed by the tumor tissue. Anaerobic bacterial species are able to thrive in the hypoxic areas of tumors, while auxotrophic strains can recognize the tumor microenvironment as a source of nutrients. Moreover, live microorganisms are metabolically active and able to perform specific actions at the tumor site, e.g. produce a prodrug-converting enzyme or a cytotoxic molecule. Strains derived from intracellular pathogens can infect tumor cells and deliver specific proteins or genes into the tumor cells (St Jean et al. 2008). Importantly, bacteria are susceptible to antibiotic treatment and therefore fully manageable in the clinical setting – therapy can be stopped at the onset of adverse effects or when is no longer needed. Taken altogether, the use of bacteria as anticancer agents might have multiple advantages over other therapeutic approaches. Strains of Salmonella are particularly apt to this task, as they can readily address all requirements for an ideal tumor-targeting agent: a motile, facultatively anaerobic, intracellular microorganism that is prone to genetic manipulations (Forbes 2010).
Salmonella – Portrait of a Conqueror
Salmonella Species Are Broad-Host, Perfectly Adapted Intracellular Pathogens
Bacteria from genus Salmonella are enteric pathogens which exploit the common basic strategy to colonize vertebrates, and are able to survive in the host digestive tract as well as in the intracellular niche. Enterobacteriacae family are commensal or pathogenic bacteria, with Escherichia being the closest known genus to Salmonella, since the two branched 120–160 million years ago from the common ancestor (Desai et al. 2013; McClelland et al. 2001; Ochman and Wilson 1987). Two subgroups within the genus are recognized as species: Salmonella bongori and Salmonella enterica. S. bongori strains are typically although not exclusively isolated from reptiles. Isolates more significant for human pathogenesis belong to S. enterica species, divided into six subspecies. S. enterica subspecies enterica are isolated from warm-blooded vertebrates including humans. Gene inactivation and lateral gene transfer are the main driving forces shaping the genome through Salmonella evolution. If we define the core genome as genes which perform the household functions and compare these genes in S. enterica and Escherichia coli, we find a mere 10 % difference, while there is only a 1 % difference among S. enterica serovars (Baker and Dougan 2007). Lateral gene transfer is a source of variability within S. enterica species and 11 % of serovar S. Typhimurium strain LT2 genes are missing in S. Typhi strain CT18 (McClelland et al. 2001). The genetic differences reflect adaptation driven by the interaction with invaded host organisms. Not only physical barriers are defeated by the pathogen, but the resident immune cells of the host, which are utilized in favor of the invader and for its successful propagation.
S. enterica serovars differ in their host preferences and pathogenesis. Most of the strains which are of interest for preventive and therapeutic vaccine development belong to serovar Typhimurium or Typhi. S. enterica subspecies enterica serovar Typhimurium, for the sake of brevity further referred to as S. Typhimurium, is a broad-host range serovar infecting both humans and other animals, and giving rise to enterocolitis or asymptomatic infection . In humans S. Typhimurium is associated with gastroenteritis manifested as short-term, acute inflammation limited to the intestine. On the contrary, host-restricted S. enterica subspecies enterica serovar Typhi (S. Typhi) cause systemic infection known as typhoid fever and humans are the only known host.
Systemic S. Typhimurium Disease in Humans Is Associated with Immune Deficiency
Some aspects of Salmonella interaction with the host immune system were recognized as the source of the host range diversity. Host immune status and the responses to bacteria are crucial for the outcome of infection . Systemic Salmonella infection is usually host-dependent (Ruby et al. 2012) and S. enterica serovars that are not host specific, such as S. Typhimurium , are associated primarily with disease in young animals, suggesting their non-optimal adaptation to a mature immune system, while host-specific serovars (S. Typhi) tend to be more virulent (Baumler et al. 1998).
S. Typhimurium infection in immune competent humans leads to gastroenteritis since the infection does not spread outside the intestine lamina propria (Ruby et al. 2012). However, the infection can lead to bacteremia and severe invasive disease in immune compromised individuals (Gordon 2008). In some laboratory mouse strains S. Typhimurium infection results in systemic inflammation congenial to S. Typhi typhoid fever in humans. Therefore S. Typhimurium infection in mice has become a widely accepted model of typhoid fever and immunity to acute Salmonella infection. In both mice and humans the susceptibility to intracellular pathogens is associated with polymorphism of Nramp1 gene (natural resistance associated macrophage protein 1, or Slc11a1). Nramp1 protein is present in the phagosome membranes in macrophages and neutrophils . It impairs the growth of intracellular pathogens such as Salmonella, which rely on survival and replication inside the phagosome (Forbes and Gros 2003). In mice resistant to lethal Salmonella infection, wild-type Nramp1 removes divalent metal cations essential for bacterial growth from bacteria-containing phagosome. Oral infection with wild-type S. Typhimurium in mice with wild type Nramp1 results in the development of a long-lasting chronic carrier state. The mice showed no apparent signs of illness during the acute phase of infection and oral lethal dose is a few thousand times higher than for mice with mutated Nramp1 (Monack et al. 2004). On the contrary, mice which are susceptible to acute systemic S. Typhimurium infection carry mutated allele of Nramp1. Commonly used laboratory mice such as Balb/c and C57Bl/6 have mutated Nramp1, and wild-type S. Typhimurium cause lethal disease soon after infection, while attenuated strains produce a chronic, persistent infection (Ruby et al. 2012; Voedisch et al. 2009).
Intracellular Lifestyle Predisposes Salmonella to Therapeutic Vaccine Development
The intracellular pathogen invasion scenario comprises of seemingly contradictory events decisive for the balance between survival, replication, spread of the pathogen and the host welfare. Regulated expression of virulence factors allows Salmonella to reach and conquer an intracellular niche and use it for replication and propagation of invasive phenotype. The molecular interactions of bacterial and host factors along with the aforementioned host susceptibility traits indicate the vast plasticity of interactions which can be exploited for the benefit of therapeutic applications. The superb ability of some attenuated S. Typhimurium strains for preferential colonization of solid tumors and the intrinsic immune stimulatory properties prompted the development of anticancer biotherapeutics, with the hope to boost the suppressed immune responses. We will briefly present the various strategies of virulence attenuation, their consequences on the immunity and the results of pre-clinical efficacy of Salmonella-based cancer therapeutics.
Salmonella Virulence Factors Stimulate Immunityand Mediate Intracellular Propagation
The shortest possible description of Salmonella – a Gram-negative, facultatively intracellular, motile enteropathogen – well emphasizes the major factors, which contribute to its virulence.
The molecules common to Salmonella and other bacterial pathogen s , that fall into the category of PAMPs, are: LPS (endotoxin), flagellin, which constructs flagellae, outer-membrane proteins belonging to Omp family, fimbrial- and non-fimbrial adhesins and non-methylated CpG sequences characteristic for bacterial DNA (Fig. 16.1).
The virulence determinants play a bimodal role, as they are important both for a successful bacterial invasion and the initiation of host immunity. Lipopolysaccharide confers bacterial resistance to the serum components and antimicrobial peptides Additionally, LPS on bacterial surface as well as released from bacteria during cell division and upon death binds to TLR4 and activates the expression of pro-inflammatory cytokines, chemokines and co-stimulatory molecules. Fimbriae and non-fimbrial adhesins facilitate colonization of intestinal mucosa and adhesion to epithelial cells (van der Velden et al. 1998; Wagner et al. 2011). Flagellum-mediated motility improves the invasion of epithelial cells and the flagellar structural protein, flagellin, binds to TLR5 and activates the expression of pro-inflammatory mediators in epithelial cells (Steiner 2007). Furthermore, the adaptive immune responses are stimulated by S. Typhimurium outer membrane proteins, which activate DCs maturation (Lee et al. 2010). Binding of non-methylated bacterial DNA to intracellular TLR9 stimulates antigen presentation by DCs, increases the expression of surface TLR9 on epithelial cells and stimulates them to secrete chemokines (Lahiri et al. 2010; Ewaschuk et al. 2007). Invasion of epithelial and phagocytic cells is mediated primarily by type three secretion system 1 (TTSS-1) effector proteins which harness the host cell cytoskeleton to trigger the uptake and formation of Salmonella containing vacuole (SCV). Following the internalization the maturation and trafficking of SCV is guided by TTSS-2 effector proteins to set up a niche permissive for bacterial replication and to delay killing. TTSS-1 and -2 effector proteins regulate the onset of immune response owing to both activatory and inhibitory actions that stimulate cytokine secretion, induce or manipulate the migration of infected cells (Schleker et al. 2012). Plasmid-encoded factors contribute to virulence by interfering with cytoskeletal proteins, mediating resistance to complement-mediated killing or encoding fimbrial proteins.
Elements in the bacterial cell diagram were scaled up relative to the bacterial cell length, e.g. TTSS-1 and flagellar motor, approximately threefold, the widths of flagellar filament and fimbriae, approximately twofold. Schematic representations of TTSS-1 and flagellar motor are based on cryo-electron microscopy data and models presented in (Kawamoto et al. 2013; Radics et al. 2014; Schraidt and Marlovits 2011; Thomas et al. 2006). Similar reconstruction of TTSS-2 is not yet available. Structure of LPS from S. enterica serovar Typhimurium was described in (Olsthoorn et al. 2000).
These molecules constitute a danger signal to leukocytes when bound to pattern recognition receptors (PRR) either in their plasma membrane, as Toll-like receptors (TLRs), or to intracellular receptors (NOD-like receptors and intracellular TLRs). Moreover, Salmonella has extraordinary genes clustered in pathogenicity islands. These genes encode genus-specific virulence factors, which are strictly related to invasion, intracellular survival and replication. Two crucial for pathogenesis and best known Salmonella pathogenicity islands are island 1 and 2 (SPI1 and SPI2), encoding proteins of two distinct type three secretion systems, TTSS-1 and TTSS-2 respectively (Fig. 16.1). Each TTSS consists of proteins that constitute the secretion machinery (structural proteins), chaperones and effector proteins. Type three secretion system structural proteins form a needle-like molecular syringe which delivers the effector proteins into the cytoplasm of eukaryotic cell. SPI effectors harness the host cell signaling pathways and manipulate vesicular trafficking to establish the intracellular niche permissive for bacterial replication.
Type Three Secretion Systems Mediate Invasion and Intra-Phagocyte Survival
Salmonella pathogenicity island 1-encoded proteins facilitate invasion of phagocytes and force bacterial uptake by non-phagocytic epithelial cells of intestinal lining by so called “trigger”-mediated invasion. SPI1 effectors injected by TTSS-1 into the cytoplasm of a eukaryotic cell interfere with actin remodeling and manipulate the cytoskeleton leading to membrane ruffling and formation of a vacuole. After being engulfed by a phagocyte or an epithelial cell, Salmonella resides inside the phagosome, which is modified due to the activities of Salmonella proteins and is therefore termed Salmonella containing vacuole (SCV). Maturation and maintenance of SCV is driven by TTSS-2 effector proteins that manipulate SCV trafficking to endocytic compartment. Salmonella-induced filaments (SIFs) are formed, which are the membrane tubules expanding from the surface of SCV and enriched in late endosomal markers (Garcia-del Portillo et al. 1993; Mota et al. 2009). Further steps of SCV development involve dynein-mediated transport along microtubules triggered by SPI1 and SPI2 TTSS effectors resulting in the juxtanuclear positioning of the SCV. Salmonella-induced tubules interact with the post-Golgi trafficking in epithelial cells, thus the role for acquiring nutrients and membranes or to control host immune response was suggested (Mota et al. 2009). Intra-phagocyte survival of S. Typhimurium may serve to modulate the onset of adaptive immune response, due to the activity of TTSS-2-delivered effector protein SseI, which inhibits the adhesion of infected phagocyte, affecting its migration efficiency and communication with other immune cells (McLaughlin et al. 2009). In some infected epithelial cells not all Salmonella cells are enclosed within SCV, but some replicate in the cytoplasm. These cytoplasmic bacteria are primed for infection of adjacent cells and ready to be shed to the environment, due to the induction of TTSS-1 and flagellar motility (Knodler et al. 2010).
The spatial and temporal regulation of virulence genes is governed by a few two-component systems and regulatory feedbacks. For example, the two-component regulatory system of the membrane sensor PhoQ and the response cytoplasmic kinase PhoP senses the transition from an extra- to intracellular environment and converts it into a transcriptional activation signal for many genes including SPI2. The LPS biosynthesis pathway is modulated as well and less negatively charged LPS molecules are produced, with lower affinity to cationic antimicrobial peptides (Shafer et al. 1984).
S. Typhimurium Infection Resolves in the Intestine or Disseminates into Systemic Disease
Over the natural course of acute typhoid-like infection in humans or mice, the initial step of invasion starts in the small intestine where non-phagocytic epithelial cells and phagocytic M cells of Peyer’s patches, resident phagocytes within lamina propria and tissue-associated DCs, are infected. M cells are the main route of entry for invasive, i.e. SPI1-proficient, Salmonella (Wick 2011). Once Salmonella enters the subepithelial dome of Peyer’s patches, it faces resident cells of intestinal lymphoid tissue, mainly the DCs, which transport bacteria to mesenteric lymph nodes (MLN). The outcome of the infection is significantly affected by the ability of MLN to control the bacterial load (Voedisch et al. 2009; Wick 2011). At this stage the infection can spread to extraintestinal tissues leading to systemic disease, if the bacterial growth is not sufficiently restrained. CD18+ macrophages and DCs play a major role in transporting S. Typhimurium through lymphatics and blood stream to the spleen, liver and bone marrow (Ruby et al. 2012). After reaching the spleen and liver Salmonella establishes the infection foci, where it multiplies mainly inside the phagocytes. Cell death of infected phagocytes enables bacteria release and the formation of new infection foci in the liver and spleen (Mastroeni et al. 2009; Salcedo et al. 2001).
Salmonella Attenuation Improves Safetyand Tumor Colonization
S. Typhimurium is a relatively mild pathogen in immune competent organisms, but systemic infection in immune compromised individuals comprises a serious health risk. Therefore the virulence of therapeutic strains is attenuated in order to achieve the proper safety profile, restrain the natural ability for spread between hosts and to enable the termination of the treatment. Moreover, attenuation provides an opportunity to redirecting the bacterial colonization, as attenuated S. Typhimurium strains colonize solid tumors preferentially over internal organs.
The balance between safety and immunogenicity is critical to achieve eligible immune responses. Over-attenuating may lead to compromised invasion, colonization or insufficient immune stimulation. For instance, in the experimental protective S. Typhi vaccine, simultaneous disruption of aromatic amino acid synthesis pathway and either purine synthesis pathway or virulence regulatory factors, by genetic disruption of aroA, purA or phoP/phoQloci, respectively, resulted in the restriction of side effects, but at the same time decreased immunogenicity preventing the induction of vaccine-specific immunity (Galen and Curtiss 2013).
Attenuation is achieved either by chemical mutagenesis or by genetic engineering techniques. Regardless of the methodology, attenuation results from metabolic and/or virulence deficiencies. Disruption of genes coding for the proteins essential for bacterial fitness, as the enzymes for biosynthesis of amino acids, e.g. aroA mutant strains, leucine and arginine-dependent mutant strain A1-R (described in detail later in this chapter); or purines – purI, purD mutant strains, results in decreased virulence. Likewise, pathogenicity is directly impaired by the inactivation of virulence genes, such as msbB in mutant strains which produce modified LPS, or regulatory genes (catabolite repression regulators crp and cya or two component regulatory system for virulence genes, phoP/phoQ).
Frequently the candidate therapeutic S. Typhimurium strains are equipped with heterologous effector therapeutic protein produced by the bacterial translation apparatus. The expression of heterologous proteins inevitably raises a metabolic burden and forced production of foreign protein often results in restrained growth or impaired invasiveness, further aggravating the attenuation. In order to balance the safety and immunogenicity and avoid the risk of over-attenuation, precisely regulated or delayed expression systems were engineered.
S. Typhimurium lipopolysaccharide synthesis pathway involves more than 30 enzymes and since it is the important structural and virulence factor, altering its structure results in attenuation. LPS structural diversity among microorganisms and its differential recognition by the receptor complex has been denoted as the factor which determines the outcome of the infection (Miller et al. 2005). The conservative core of LPS molecule is built of lipid A and oligosaccharides (core sugars), while Salmonella serotype-specific portion lies within O-antigen polysaccharide. Antimicrobial host responses are initiated through lipid A binding to membrane receptor complex TLR4-MD2-CD14 present on many cell types, among them DCs and macrophages , which secrete proinflammatory cytokines including TNF.
Modification of LPS structure in order to decrease the endotoxic shock induction has already been applied with satisfactory results, as S. Typhimurium VNP20009 strain was safely administered to cancer patients (Toso et al. 2002). The strain is attenuated and auxotrophic due to the disruption of lipid A myristoylation enzyme (inactivation of msbB gene) and purine synthesis pathway gene deletion (purI), respectively. Intravenous administration of VN20009 to tumor bearing mice results in tumor colonization, with the ratio of intratumoral to intrasplenic accumulation exceeding 1000. However, in humans the clinical benefit was not observed, presumably due to insufficient tumor colonization (Toso et al. 2002). The clinical trial will be discussed in detail later in this chapter.
In some studies, intratumoral hemorrhage induced by TNF was attributed to intratumoral Salmonella accumulation (Leschner et al. 2009). The ability of mouse and human to recognize the same lipid A molecule differs. Furthermore, the length and number of acyl side chains is important for TLR4 signaling in humans. Additional diversity in TLR4 responsiveness to LPS comes from the natural gene polymorphism which alters TLR4 signaling outcome in humans (Miller et al. 2005). Recently it has been proposed that the administration of live VNP20009 mixed with molecular wild-type lipid A could overcome the poor clinical tumor targeting (Zhang et al. 2013a). Salmonella accumulation in subcutaneous 4 T1 tumors was lipid A-dose-dependent and bacteria distribution was more homogenic after co-administration. However, the study was completed 2 days after bacteria inoculation, therefore a long-term therapeutic effect was not revealed.
Salmonella Is Able to Induce Several Different Death Modalities in Infected Cells
During natural infection Salmonella enters two types of cells: intestine epithelium including specialized M cells, and immune cells: DCs and macrophages . Salmonella, as an intracellular pathogen, does not immediately kill the cell it has invaded, but rather delays an onset of cell death to earn time to replicate, escape and infect new cells. However, the interplay between infected host cells and bacteria eventually leads to the death of the eukaryotic cells including macrophages. The ability of engulfed Salmonella not only to resist the antimicrobial activity of macrophages, but also to proliferate inside and induce the death of these professional phagocytes is the key strategy of the pathogen to circumvent innate immune defense (Lindgren et al. 1996).
Different types of cell death are induced in Salmonella -containing epithelial cells and macrophages (Ramos-Morales 2012). In cultured human epithelial cells Salmonella triggers an apoptotic cell death program with all the hallmarks of this process including exposure of phosphatidylserine on the cell surface, activation of effector caspase-3, fragmentation of DNA, depolarization of mitochondrial membrane and degradation of cytokeratin 18. However, apoptosis is turned on after a lag period of between 12 and 24 h following bacterial entry (Kim et al. 1998; Paesold et al. 2002). A number of virulence factors stimulate apoptosis by affecting general protein synthesis, inducing actin depolimerization, and tipping the balance between expression of pro- and antiapoptotic proteins. Importantly, some bacterial proteins counteract a rapid induction of apoptosis: SopB activates pro-survival kinase Akt and AvrA inhibits c-Jun-N terminal kinase, which, when activated by stress stimuli, is involved in apoptosis (Ramos-Morales 2012).
In macrophages Salmonella Typhimurium induces a proinflammatory form of programmed cell death – so-called pyroptosis or caspase-1-dependent cell death. This process is induced by TTSS-1-expressing bacteria and occurs in 1–2 h postinfection. Activation of caspase-1 is responsible for maturation of two cytokines: IL-1β and IL-18 and results in cell lysis accompanied by the release of the active proinflammatory mediators (Cookson and Brennan 2001). Pyroptosis requires activation of NLRC4 inflammasome by flagellin and a rod component PrgJ (Zhao et al. 2011). However, the expression of these genes is repressed during systemic infection and in such situation the death of macrophages occurs not earlier than 12–16 h postinfection (Wynosky-Dolfi et al. 2014). This delay is advantageous for Salmonella and is in fact caused by bacteria. It has recently been demonstrated that bacterial tricarboxylic acid (TCA) cycle enzymes impair, by a yet unknown mechanism, an expected rapid canonical activation of NLRP3 inflammasome (Wynosky-Dolfi et al. 2014). Instead, delayed caspase-11- and type I IFN (IFN-I)-dependent noncanonical activation of NLRP3 inflammasome takes place. In the proposed scenario Salmonella through TLR4 stimulates the expression of both IFN-I and caspase-11. IFN-I induces the synthesis of a group of GTPases known as guanylate binding proteins (GBP), which mediate the lysis of Salmonella-containing vacuole. Cytosolic LPS of released bacteria indirectly activates caspase-11/NLRP3 inflammasome followed by activation of caspase-1 and macrophage death (Meunier et al. 2014). It has also been proposed that IFN-I mediates another type of Salmonella- induced cell death in macrophages – so-called necroptosis or programmed necrosis. This type of cell death is triggered by the formation of a necrosis-inducing complex comprised of receptor-interacting protein kinase 1 (RIPK1) and RIPK3 (Lu and Walsh 2012), which is followed by mitochondrial fragmentation (Wang et al. 2012). In contrast to apoptosis, both pyroptosis and necroptosis trigger inflammatory responses (Hu and Zhao 2013).
Autophagy Is an Important Cellular Process Upon Salmonella Infection
Until recently, autophagy was viewed as a process whose primary function in the cell is to degrade faulty organelles and cytosolic macromolecular aggregates as well as to prevent cell death by providing amino acids and other basic compounds during nutritional deprivation (Deretic 2011). On the other hand, due to the fact that autophagosomes and biochemical autophagy markers have often been observed in dying cells, autophagy has been regarded as, distinct from apoptosis, programmed cell death, termed “type II programmed cell death” or “autophagic cell death” (Ryter et al. 2014). However, a growing body of research indicates that autophagy – although often accompanies cell death – usually is not its cause. Therefore, the term “death with autophagy” seems to be more proper in the vast majority of cases than “death by autophagy” (Kroemer and Levine 2008).
Apart from its importance for the cell maintenance, autophagy is presently recognized as a process which plays important roles in innate and adaptive immunity. It has been proposed that autophagy acts as an evolutionary ancient system specialized to eliminate intracellular bacteria and viruses. It also contributes to the presentation of endogenously expressed antigens via major histocompatibility complex class II (MHC-II) (Deretic 2011; Espert et al. 2007). Thus, it is not surprising that autophagy has been observed in macrophages as well as in epithelial cells, fibroblasts and melanoma cells subjected to Salmonella infection (Birmingham et al. 2006; Hernandez et al. 2003; Lee et al. 2014; Thurston et al. 2009).
Autophagy involves three stages: (i) formation of an initiator crescent membrane (phagophore), (ii) growth of the phagophore to enclose cargo in the completed autophagosome and (iii) conversion to the autolysosome by fusion with late endosomes or lysosomes. A phagophore membrane contains phosphatidylethanolamine-anchored LC3-II protein, which interacts with autophagic adapter proteins that recognize targets marked for autophagy by ubiquitination (Deretic 2011).
In epithelial cells a subset of invading Salmonella is released to the cytosol from TTSS-1-damaged SCV shortly after infection (Birmingham et al. 2006). It has been shown that in HeLa cells bacteria that escaped SCV have become ubiquitin-coated and targeted to the autophagy pathway by p62 and nuclear domain 10 protein 52 (NDP52) adapters (Zheng et al. 2009; Thurston et al. 2009). Alternatively, diacylglycerol present in SCV membrane may constitute a signal for antibacterial autophagy (Shahnazari et al. 2010). A number of reports indicate that although autophagy is a transient process peaking in 1–2 h after Salmonella infection, it restricts intracellular growth of bacteria (Birmingham et al. 2006; Thurston et al. 2009; Zheng et al. 2009). Intriguingly and in contrast to the previous reports, Yu et al. (2014) have recently demonstrated, using a live-cell imaging method, that cytosolic Salmonella associated with autophagy components p62 and LC3 replicated efficiently in HeLa cells and, what is more, the replication has been diminished when p62 or LC3 availability has been decreased (Yu et al. 2014). This discrepancy may result from the different time schedules of the experimental design and may be explained, at least partially, by the fact that autophagy regulation is associated with the metabolic status of the cell where amino acid starvation seems to be the key trigger of the program. The transient character of cytosolic amino acid depletion induced by S. Typhimurium infection may explain the transient execution of autophagy. How the bacteria take advantage of certain autophagy elements at later times postinfection remains to be established.
Although autophagy has often been indicated as a possible cause of death of Salmonella -infected cells, in light of current knowledge it should rather be regarded as a natural element of the cell response to the infection that may help in pathogen eradication.
Cell Death Induced by Salmonella Can Promote Cross-Presentation of Tumor Antigens
The studies on the types of death caused by Salmonella were performed mostly using wild type bacteria or bacteria with knockouts in specific genes or gene clusters whose role in Salmonella’s deadly potential was under investigation. Therefore, caution is required if the results are extrapolated to the potential effects of attenuated strains, which are candidates for clinical applications. However, the understanding of the biochemical consequences of particular modes of attenuation allows presuming that at least some of the attenuated strains will execute the death programs in a similar way to the wild type strains.
The primary asset of Salmonella -based therapies could be an enhancement of tumor antigen cross-presentation by DCs that have taken up cancer cells dying due to infection . Much effort has been made to understand the mechanisms of optimal cross presentation. Some researchers indicate specialized subset of DCs, namely CD8+ DCs in mice and thrombomodulin-expressing DC in humans, as possessing superior capability to cross-present antigens (Villadangos and Shortman 2010; Joffre et al. 2012), whereas others postulate that all DCs may be efficient cross-presenters when appropriately activated by CD40L, inflammatory cytokines, and TLR ligands (Dresch et al. 2012). DCs can ingest dying cancer cells and cross-present a panel of tumor antigens; since necrotic cells are a rich source of danger signals, necrosis would be expected as the type of tumor cell death with the highest potential to activate DCs to efficient cross-presentation. Surprisingly, numerous reports from in vitro and in vivo studies showed that tumor apoptotic cells are better at facilitating cross-presentation of tumor antigens to CD8+ T cell s than their necrotic counterparts (Spel et al. 2013). The higher immunogenicity of apoptotic cells may result from their CLEC9A-mediated targeting to storing phagosomes, in which mild pH and reduced rate of proteolysis constitute proper conditions for optimal cross-presentation of cell-associated antigens (Spel et al. 2013). Interestingly, the exposure of DCs to IFN-I further enhances this process. IFN-I seems to prolong the presence of apoptotic cells inside DCs, supporting localization of MHC class I molecules to antigen-storage compartment as well as promoting DCs survival (Lorenzi et al. 2011). Salmonella may strongly promote cross-presentation of tumor antigens since it: (i) induces apoptosis in cells of epithelial origin, (ii) stimulates expression of IFN-I both in macrophages and in non-phagocytic cells, (iii) provides ligands for TLRs, and (iv) stimulates release of proinflammatory cytokines from infected macrophages. Additionally, DAMPs released from necrotic tumor cells, which appear independently of Salmonella infection or originate from non-phagocytosed bacteria-infected apoptotic cells may further increase maturation and activation of DCs. It is noteworthy that also autophagy occurring in antigen-donor cells (e.g. tumor cells or virus-containing cells) may support antigen cross-presentation by DCs immunized with these cells. Uhl et al. demonstrated that virally infected fibroblasts undergoing autophagy facilitated antigen-specific CD8+ T cell cross-priming more efficiently than their apoptotic counterparts (Uhl et al. 2009).
It is also tempting to speculate that Salmonella -induced death of TAMs resulting in the reduction of the number of these immunosuppressive cells in tumors may help to tip the balance of signals in favor of anti-tumor responses. The observation that the killing of TAMs by natural killer T cell s in primary human neuroblastoma suppressed tumor growth may support this concept (Song et al. 2009). Currently, the idea of simultaneous targeting of TAMs and tumor cells is gaining attention as a promising approach for tumor therapy (Germano et al. 2013; Xin et al. 2009).
Salmonella -Mediated Tumor Growth InhibitionInvolves Immune Regulatory Mechanisms
Immune therapy rationale relies on the pre-existence of tumor specific immunity which is not effective in the absence of therapeutic intervention due to a tumor immunosuppressive environment. Therefore the primary value of bacteriotherapy is its potential to stimulate the immune responses to combat suppression in tumor microenvironment and break immune tolerance to tumor. While the elimination of major tumor burden is in the scope of classical treatment, the long-term benefit of immunotherapy would be the induction of tumor-specific memory responses to protect patients from tumor recurrence and metastatic disease. Various approaches have been proposed in the pre-clinical studies to eradicate tumor with Salmonella . Attenuated and auxotrophic strains are either used solely to boost the immunity or as vectors for the delivery of tumor-associated antigens to elicit specific T cell response, or other heterologous therapeutic molecules.
Immunoregulatory cells impede tumor immune surveillance, trigger chronic inflammation and tumor-promoting environment. Modification of phenotype and activity of immune regulatory cell s , such as M2 macrophages and MDSCs was shown to account for Salmonella -induced tumor growth inhibition. Moreover, DC maturation, effective tumor antigen presentation and cytotoxic T cell s responses after Salmonella treatment prove the concept of danger signal-mediated shift form immunosuppressive, tumor fostering environment to antitumor immune responses (Hong et al. 2013; Jarosz et al. 2013; Kaimala et al. 2014).
Auxotrophic S. Typhimurium BRD509E administered intraperitoneally to C57Bl/6 mice bearing subcutaneous B16.F1 melanoma tumors, accumulated in tumors and significantly retarded tumor growth (Kaimala et al. 2014). Tumor colonization was correlated with increased accumulation of CD11b+Gr1+ myeloid cell s in the tumors. These cells had increased surface expression of maturation and activation markers – MHC class II, co-stimulatory molecule CD80 and Sca-1 (Ly-6A, Stem cell antigen-1) in Salmonella -treated, compared to non-treated mice. Moreover, expression of IFNγ was up-regulated in spleens and tumors after Salmonella treatment.
Importantly, intratumoral myeloid cell s from bacteria-treated mice had lower expression of ARG1, suggesting a reversion of suppressive phenotype. Indeed, intratumoral myeloid cells from Salmonella -treated mice were less suppressive towards CD4+ T cell s than those from control mice (Kaimala et al. 2014).
Natural course of Salmonella oral infection results predominantly in mounting the local protective immunity in the mesenteric lymph nodes (Voedisch et al. 2009). Since the exact mechanism of solid tumor colonization is yet to be clarified and bacterial tumor colonization is critical for therapeutic benefit, the optimal route of administration is crucial for the outcome of treatment. Various mutant strains defective in metabolic or virulence factors show diverse extent of tissue colonization in mice. Both orogastric and intravenous administration of different S. Typhimurium strains were shown to inhibit transplantable tumor growth in mouse models. The oral route would be technically easier to conduct in clinical settings and would be of reduced health risk, but some strains do not target distant solid tumors after oral administration.
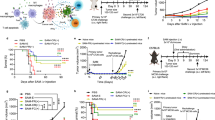
Recently the group led by Siegfried Weiss compared the efficacy of systemic and oral delivery of different S. Typhimurium strains. S. Typhimurium SL1344ΔaroA strain was administered by either of the three routes: intravenous (i.v.), intraperitoneal (i.p.) or oral, to Balb/c mice with subcutaneous CT26 solid tumors. Viable bacteria were recovered from tumor, spleen and liver at all tested time points (1–17 days) after i.v. or i.p. injection, but only on 11th day (not earlier and not later) after oral infection . However, all tested routes resulted in preferential tumor colonization, with about 100 times more bacteria in tumors than in the spleen and liver. Intravenous and intraperitoneal infection led to complete clearance of tumors, while oral infection transiently inhibited tumor growth when bacterial tumor colonization was detectable, but finally the tumors grew as in non-treated control mice (Crull et al. 2011). The same authors provided a comparison of organ colonization by wild type S. Typhimurium strain SL1344 and its two derivatives – deletion mutants of aroA or purA, auxotrophic for aromatic amino acids or purines, respectively. All three strains efficiently colonized tumor and spleen after intravenous injection, but wild-type bacteria caused severe side effects. Colonization with purA-deleted strain was delayed and less efficient than that with aroA-deleted strain (Crull et al. 2011). Finally, the effects of two strains with the same attenuation type but on different genetic background were compared. SL1344ΔaroA colonized tumors more efficiently, leading to their clearance in all mice, while treatment with SL7207, which is also aroA-deficient, eliminated tumors in three out of five mice. These results point out the dependence of attenuation outcome on genetic background (Crull et al. 2011). In line with the aforementioned results, the importance of bacterial tumor colonization, as well as colocalization of tumor antigens and bacterial immune mediators at tumor site was highlighted by the recent study. Attenuated S. Typhimurium Χ9241 strain was injected intratumorally or given orally to Balb/c mice bearing subcutaneous CT26 tumors expressing heterologous tumor antigen, human Her-2/Neu. Intratumoral injection significantly inhibited tumor growth while the oral route was not effective (Hong et al. 2013). Tumor growth inhibition was partially dependent on cytotoxic lymphocyte response, since CD8+ cells depletion reduced the effect. The treatment increased the frequency of splenic M- and G-MDSCs, and intratumoral G-MDSCs. But yet the phenotype of myeloid cell s was distinct from suppressive MDSCs, as a significantly higher percentage produced TNF. The therapeutic effect relied also on the decreased percentage of splenic and tumor-infiltrating CD4+CD25+Foxp3+ regulatory T cell s (Hong et al. 2013). These experimental data support the concept that tumor targeting with attenuated Salmonella not only inhibits its growth, but induces the break of the immune suppressive tumor microenvironment .
Transforming the Perfect Pathogen into a Perfect Drug Candidate
Attenuated S. Typhimurium Inhibits Tumor Growth in Animal Models
While designing S. Typhimurium -based anti-tumor therapies there is a prevalent tendency to improve or enhance therapeutic potential of different attenuated strains by various genetic modifications. These will be described later. Yet, there are also approaches to utilize natural cytotoxic and immunomodulatory capabilities of attenuated bacteria strains without any further modifications. A few examples of the strains that showed promising results when used in animal model s are worth mentioning.
The group led by Robert Hoffman used a sophisticated way of attenuation to develop the A1-R strain. At first, S. Typhimurium (ATCC 14028) was subjected to chemical mutagenesis with nitrosoguanidine (NTG) that resulted in obtaining an auxotrophic A1 strain dependent on an external source of leucine and arginine (Zhao et al. 2005). In the next step, A1 bacteria were passaged through HT-29 tumor-bearing nude mice and recovered from the excised tumor tissue. As a consequence of this selection step, the strain isolated from the tumor, called A1-R, demonstrated improved targeting towards cancer cells in vitro and in vivo in comparison to A1 (Zhao et al. 2006). A1-R without any additional modifications was successfully used in a model therapy of numerous human tumor xenografts in nude mice, significantly suppressing tumor growth in all tested models and causing complete tumor eradication in several of them (Hayashi et al. 2009a; Hayashi et al. 2009b; Kimura et al. 2010; Nagakura et al. 2009; Zhang et al. 2012; Zhao et al. 2006, 2007).
The A1-R strain has also been tested in metastatic tumor models, e.g. in the spontaneous popliteal lymph node metastasis model of human HT-1080 fibrosarcoma in nude mice. In five out of six mice the lymph node metastases were completely eradicated within 7–21 days after the administration of A1-R in the footpad, which was the site of the tumor injection. In contrast, metastases in lymph nodes of all mice from the control group, injected with the solvent instead of bacteria, continued to grow (Hayashi et al. 2009a). In two other nude mice models, the lung metastasis model of human osteosarcoma and human HT-1080 fibrosarcoma, intravenous administration of A1-R bacteria resulted in a strong decrease of the number and size of metastases (Hayashi et al. 2009a, b). Recently, the A1-R strain was demonstrated to have increased efficacy compared to standard chemotherapy in treating patient-derived orthotopic xenograft (PDOX) of pancreatic cancer in nude mice (Hiroshima et al. 2014). The results suggest the clinical potential of A1-R bacterial therapy against this highly lethal type of cancer. All the above examples come from the studies performed on immunodeficient animals and thus the results indicate that the A1-R strain may exert strong cytotoxic effects toward tumors independently of the adaptive immune responses. Presently, the effects of various experimental settings for the evaluation of A1-R applicability in immunocompetent mice are under investigation (Tome et al. 2013; Zhang et al. 2013b).
Another interesting approach was proposed by Yu et al. (2012), who developed a novel strain of S. Typhimurium , YB1, unable to survive in normal tissue. They placed the asd gene, crucial for the synthesis of DAP, an important bacterial cell wall component, under the control of a hypoxia-induced promoter. Limiting the expression of the asd gene to hypoxic conditions makes the bacterium an obligate anaerobe. Such a modification renders YB1 strain non-toxic for normal tissues while maintaining its tumor targeting. The safety and anti-tumor efficacy of three Salmonella strains: SL7207, YB1 and another attenuated Salmonella strain VNP20009 were compared in the MDA-MB-231 breast tumor-bearing nude mice model. Bacteria of all the strains were able to infiltrate and destruct tumors. However, in contrast to deadly SL7207 parental strain, YB1 bacteria were effectively cleared from normal tissues and were barely detectable in the liver 3 days postinfection. They also showed better therapeutic parameters than VNP20009 in MDA-MB-231 tumor model (Yu et al. 2012).
VNP20009 Is Safe But Does Not Colonize Tumors in Humans
VNP20009, msbB, purI-attenuated strain of S. Typhimurium , has been shown to accumulate preferentially at the tumor sites after intravenous administration to the mice bearing spontaneous, syngeneic or human xenograft tumors. The tumor-to-normal tissue ratio ranged from 300:1 to 1000:1 (Clairmont et al. 2000). Also in dogs suffering from different spontaneous tumors and subjected to S. Typhimurium intravenous injections, an accumulation of bacteria was demonstrated in more than 40 % of cases (Thamm et al. 2005). The maximum tolerated dose (MTD) of VNP20009 for mice was estimated to be as high as 0.5 × 108 colony forming units (CFU) per kg of body weight (Lee et al. 2000). Similar values of MTD were established for other mammals: dogs – 3.0 × 107, pigs – 1.9 × 108 and monkeys – 2.5 × 108 CFU/kg (Lee et al. 2000; Thamm et al. 2005). Based on the preclinical data, in November 1999, VNP20009 entered Phase I human clinical trial conducted by Vion Pharmaceuticals Inc. Twenty five patients – 24 with metastatic melanoma and one with metastatic renal cancer – were initially enrolled for the trial. VNP20009 have been administered intravenously for 30 min in doses ranging from 106 to 109 CFU/m2. MTD has been established at 3 × 108 CFU/m2, as higher doses were accompanied by substantial adverse effects including fever, hypotension, anemia and thrombocytopenia. In the majority of cases the bacteria were rapidly cleared from the bloodstream. The serum levels of cytokines: TNF, IL-1β , IL-6 , IL-12 were transiently increased and correlated with Salmonella dose. Unfortunately, only 3 out of 18 tumor biopsies contained viable bacteria and even in those cases tumor growth regression was not observed (Toso et al. 2002). Increasing the infusion time to 4 h, which was applied for the next four patients, did not improve the outcome (Heimann and Rosenberg 2003).
Another pilot clinical trial was performed on three patients in order to evaluate the possible effects of intratumoral application of genetically modified VNP20009 strain called TAPET-CD (Tumor Amplified Protein Expression Therapy – Cytosine Deaminase), expressing E. coli cytosine deaminase (Nemunaitis et al. 2003). In principle, intratumoral localization of the enzyme-expressing bacteria would result in the accumulation of cytotoxic 5-fluorouracil (5-FU) at the tumor site following systemic application of non-toxic 5′-fluorocytosine (5-FC). Indeed, some colonization by TAPET-CD bacteria in tumors of two patients was accompanied by the increased conversion of 5-FC to 5-FU. However, intratumoral route of Salmonella administration did not significantly improve bacterial load in tumor tissue (Nemunaitis et al. 2003).
The clinical trials revealed that VNP20009 do not colonize tumors in patients to the same degree as observed in murine models, which may be a major reason for the lack of its therapeutic effects. Hence attempts were made to increase the tumor targeting capability of Salmonella . Bereta et al. (2007) constructed a VNP20009 strain expressing a fusion protein of OmpA and a single chain antibody fragment (scFv) recognizing carcinoembryonic antigen (CEA), a widespread TAA of human carcinomas. The modified VNP20009 strain showed increased localization in CEA-expressing MC38 tumors compared to wild type-MC38 and inhibited CEA-MC38 tumor growth more efficiently than non-modified, parental VNP20009 (Bereta et al. 2007). A similar approach was also proposed by Massa et al. (2013) who genetically engineered aroA-deficient SL3261 strain to express a fusion protein consisting of OmpA and a camelid single-domain (VHH) antibody against human CD20. They observed that the presence of the antibody at the surface of bacteria improves Salmonella targeting to the mouse and human tumors expressing CD20 and limits bacterial invasion of the liver and spleen (Massa et al. 2013).
Also Salmonella Typhi is taken into account in the development of anti-cancer therapy. Up to date various attenuated S. Typhi strains have been tested mainly as live attenuated oral paratyphoid or typhoid fever vaccines , reviewed in (Roland and Brenneman 2013). When used as a vector in humans S. Typhi turned out to be inferior to S. Typhimurium in eliciting the immune response to a delivered heterologous antigen (H. pylori urease), however this may result from the oral route of bacteria administration and longer persistence of S. Typhimurium in the intestine (Angelakopoulos and Hohmann 2000). S. Typhi Ty21a strain was generated as an anti-angiogenic cancer therapeutic vaccine . It carries a plasmid encoding full length vascular endothelial growth factor receptor-2 (VEGFR-2). S. Typhi Ty21a entered Phase I clinical trial in 2012 (Niethammer et al. 2012), but the results of the study are not yet available.
S. Typhimurium Can Be Optimized Using Genetic Engineering
The failure of the first clinical trials highlights the need for improvement of Salmonella targeting to tumor tissues in humans. This is certainly the biggest challenge facing researchers involved in the development of Salmonella-based therapies. However, even in mouse models attenuated Salmonella strains, although they strongly inhibited the growth of various tumors, rarely led to their complete eradication. Therefore, numerous attempts are being made to increase efficacy of Salmonella-based therapies by introducing genetic modifications into attenuated strains. The modified bacteria, thanks to their intracellular lifestyle, serve as a vector for the delivery of therapeutic molecules into the cells. Salmonella usually gets equipped in a plasmid coding for effector molecules whose expression is controlled by eukaryotic or bacterial promoters. The potential therapeutic advantage of various effector molecules including TAAs, cytokines, apoptosis-inducing factors, prodrug-converting enzymes or short hairpin RNAs (shRNAs) able to silence the expression of a protein of choice have already been tested. The intended effects of these molecules can be divided into three groups: (i) increasing cytotoxic effects of Salmonella infection ; (ii) enhancement and navigation of the immunomodulatory properties of Salmonella; (iii) delivery of TAAs. The therapeutic approaches are summarized in Table 16.1 and several examples of interesting and promising ideas are presented below.
Apoptosis-Inducing Factors Can Increase Cytotoxic Potential of Salmonella
In order to enhance the capability of S. Typhimurium to kill infected cancer cells, several research groups took advantage of apoptosis-inducing factors and equipped bacteria with appropriate expression vectors.
Apoptin (VP3) is a chicken anemia virus (CAV) protein which exhibits p53-independent, tumor cell-specific proapoptotic effects (Zhuang et al. 1995). It does not affect non-malignant cells and this selectivity of action makes apoptin an interesting potential anti-tumor agent. S. Typhimurium LH430 carrying apoptin gene under the control of eukaryotic cytomegalovirus (CMV) early promoter (ST-rC-Apoptin) increased the delay of the growth of human Hep-2 xenografts in nude mice as compared to the effects of ST-rC-EGFP control. The lack of side effects may be explained by an almost thousand times higher accumulation of ST-rC-Apoptin in tumor tissue than in the liver, equally after one or two i.v. injections of Salmonella . The expression of apoptin in tumor tissue was followed by an increased activity of caspases (Guan et al. 2013).
Another Salmonella strain has been equipped in a double proapoptotic weapon: apoptin and TNF-related apoptosis-inducing ligand (TRAIL). Similarly to apoptin, TRAIL induces apoptosis in a wide variety of cancer cells, but hardly in normal cells (Walczak et al. 1999; Yagita et al. 2004). S. Typhimurium SL7207 carrying apoptin gene and TRAIL cDNA under the control of CMV promoter injected intratumorally to human gastric tumor xenografts in nude mice induced higher apoptosis rate than unmodified SL7207 and strongly suppressed tumor growth with its complete eradication in some animals (Cao et al. 2010).
The idea of combining TRAIL expression with Salmonella -tumor targeting was also explored by Ganai et al. (2009). They placed TRAIL cDNA under the bacterial RecA promoter activated during SOS response to DNA damage (Anderson and Kowalczykowski 1998) and thus created a radiation-inducible system for temporal and spatial control of TRAIL expression. Intravenous administration of TRAIL-expressing VNP20009 into mice bearing 4T1 mammary carcinoma followed by 2Gy whole body γ-irradiation 2 days later led to a significant inhibition of tumor growth resulting from the combined effects of Salmonella infection , irradiation and TRAIL expression (Ganai et al. 2009).
It has been shown that second mitochondria-derived activator of caspases (Smac) sensitizes various tumor cells for TRAIL-induced apoptosis (Deng et al. 2002; Zhang et al. 2001). Therefore, Fu et al. (2008) engineered a modified S. Typhimurium SL3261 strain carrying a vector coding for Smac and TRAIL (S.L./SNhTS). Expression of both cDNAs was controlled by the promoter of human telomerase reverse transcriptase (hTERT), highly active in many human cancers and inactive in non-proliferating normal cells (Kim et al. 1994). Indeed, in contrast to normal cells, S.L./SNhTS-infected tumor cells of different origin (LL/2 Lewis lung carcinoma, B16F10 melanoma and 4T1 mammary carcinoma) expressed high levels of exogenous Smac and TRAIL, resulting in an elevated apoptosis rate. In vivo studies demonstrated that orally delivered S.L./SNhTS markedly suppressed tumor growth in all tested mice models, without any detectable side-effects (Fu et al. 2008).
Another approach aiming at the enhancement of proapoptotic properties of Salmonella was proposed by Joeng et al. (2014), who thought not only of the synthesis of a proapoptotic factor, mitochondrial-targeting domain of Noxa (MTD), at the tumor site, but also carefully designed a system of its transport from bacteria to tumor cells (Jeong et al. 2014). The researchers used ΔppGpp Salmonella strain unable to produce a key regulator of vital bacterial processes – guanosine-3′, 5′-bisdiphospate which guarantees very high accumulation of bacteria in the tumor tissue over the liver or spleen. Noxa is a transcriptional target of p53 which contributes to the induction of intrinsic apoptosis pathway via the activation of mitochondrial damage (Seo et al. 2009; Zhang et al. 2011). MTD, its prodeath domain causes extensive necrosis of cells in vitro through an increase of the cytosolic calcium levels (Seo et al. 2009). In the designed system, MTD was expressed as a fusion protein with DS4.3, a cell-penetrating peptide facilitating eukaryotic cell entry after bacteria lysis. To release DS4.3-MTD from bacteria phage lysis genes of a newly characterized Salmonella phage were employed. Both DS4.3-MTD cDNA and phage lysis genes were placed under the control of pBAD, a promoter activated by L-arabinose. CT26 colon carcinoma-bearing mice were intravenously injected with Salmonella carrying pLYSPBAD::DS4.3-MTD. Three days later, when the bacteria accumulated in the tumors over the livers at a ratio of 48,000:1 and over the spleens at a ratio of 65,000:1, daily administration of L-arabinose was started. Massive necrosis of tumor tissue and suppression of tumor growth was observed (Jeong et al. 2014).
Immunomodulatory Properties of Salmonella Can Be Strengthened
Numerous modifications have been introduced to Salmonella to influence immune cells and reinforce anti-tumor immune response by tipping the balance between pro- and anti-tumor activities. The strains of Salmonella producing various cytokines and chemokines such as IL-2 (al-Ramadi et al. 2009; Ha et al. 2012), IL-21 (Wang et al. 2013), TNF (Yoon et al. 2011), IL-18 (Loeffler et al. 2008), CCL21 (Loeffler et al. 2009) or LIGHT (Loeffler et al. 2007) were generated and tested in mouse models. Interestingly, Salmonella expressing specific shRNA may switch off the synthesis of a host immunosuppressing protein (Blache et al. 2012). Two examples of promising approaches are described below.
John C. Reed’s group demonstrated a superiority of CCL21 chemokine-expressing Salmonella strain over a parental strain in inhibiting the growth of CT26 colon, D2F2 breast, and B16 melanoma tumors as well as in limiting CT26-lung colonization in immunocompetent mice (Loeffler et al. 2009). The idea of equipping Salmonella in CCL21 came from the known activities of this chemokine and was supported by the results of experiments in which CCL21 was directly injected into tumors. As CCL21 is a chemoattractant for T lymphocytes and DCs it may therefore stimulate the colocalization of naive T cells and tumor antigen-presenting DCs which may help to mount effective immune response and lead to subsequent tumor eradication. Indeed, observed inhibition of tumor growth by CCL21-expressing Salmonella was accompanied by the increased intratumoral levels of IFNγ, CXCL9, and CXCL10, cytokines known to be induced by CCL21, as well as by enhanced infiltration of tumors by immune cells including CD4+ and CD8+ T lymphocytes. Immunodepletion of different cell populations revealed that both CD4+ and CD8+ T cell s were indispensable for significant inhibition of tumor development by CCL21-expressing Salmonella.
An unconventional therapeutic approach was proposed by Blache et al. (2012), who decided to use Salmonella vector to modulate immunosuppressive tumor microenvironment via silencing of IDO expression. VNP20009 strain of S. Typhimurium carried a plasmid coding for shRNA able to silence IDO expression (shIDO-ST). Bacteria injected intravenously significantly inhibited the growth of subcutaneous B16F10 melanoma as well as considerably diminished a number of lung melanoma foci after intravenous application of the tumor cells and their effects were more pronounced than those of VNP20009 expressing scrambled shRNA. The treatment of mice with shIDO-ST resulted in a significant intratumoral influx of CD11b+Gr1+ which were mostly Ly6G+, accompanied by markedly elevated ROS levels and massive intratumoral cell death. Unexpectedly, the levels of CD4+ and CD8+ T cell s remained unaffected and what is more, shIDO-ST was equally active in normal mice as in mice depleted of CD4+, CD8+, or NK subsets. In contrast, depletion of Gr1+ cells resulted in abrogation of shIDO-ST inhibitory effects. The results indicate that in this tumor model polymorphonuclear leukocytes were obligatory for the shIDO-ST therapeutic efficacy. Interestingly, the silencing of IDO potentiated the colonization of tumor by shIDO-ST (Blache et al. 2012).
Salmonella May Play the Role of a TAA-Expressing Vector
The rationale for using Salmonella as a vector delivering TAAs is that the presentation of antigens to immune cells will be enhanced by the strong danger signals. Up to date, several natural (mAFP, survivin, endoglin) or artificial (β-galactosidase) tumor antigens have been placed under the control of CMV promoter in the plasmids introduced to attenuated Salmonella, which were then used as prophylactic or therapeutic vaccines . This approach assumes that the plasmid carried by the bacteria becomes available to eukaryotic transcriptional machinery which recognizes CMV promoter. Although the mechanism by which the plasmid enclosed in Salmonella is delivered to the cell nucleus is not clear, it is well documented that the TAAs whose expression was controlled by CMV promoter were produced in the cytoplasm of infected cells or in the cytoplasm of phagocytes that engulfed bacteria or infected, apoptotic cells (Paglia et al. 1998; Yrlid and Wick 2000). TAAs produced in this way elicited efficient cell-mediated or both cell-mediated and humoral immune responses (Chou et al. 2006; Fest et al. 2009; Jarosz et al. 2013; Paglia et al. 1998). However, the intracellular location of Salmonella within the SCV may limit the CMV-driven TAAs expression. Another approach has been developed to circumvent this limitation. It utilizes Salmonella’s TTSS to deliver TAAs produced by bacteria inside the SCV to the cytoplasm of the host cell. The transgene that is placed in a plasmid under the control of a bacterial promoter encodes a fusion protein consisting of TAA preceded by a short sequence derived from the N-terminus of a bacterial protein, e.g. SopE, SseF, YopE, SipB exported via TTSS. This sequence contains the so-called secretion and translocation signal which enables an efficient delivery of a fusion protein to the host cell cytoplasm and therefore makes it available for MHC-I presentation. The usefulness of this approach in both prophylactic as well as therapeutic vaccine settings was demonstrated by Russmann et al. and followers (Roider et al. 2011; Russmann et al. 1998). The activation of specific CD8+ T cell s , epitope spreading, increased intratumoral proliferation of CD4+ and CD8+ T lymphocytes, elevated intratumoral levels of granzyme B, increased serum levels of TNF and IFNγ are among the effects that were demonstrated as resulting from oral or intravenous administration of Salmonella-based tumor vaccines. The natural TAAs delivered by Salmonella’s TTSS include NY-ESO-1 (Nishikawa et al. 2006), survivin (Manuel et al. 2011), tyrosinase-related protein 2 (Trp2) (Zhu et al. 2010) and human papilloma virus HPV E7 (Yoon et al. 2014). The chimeric transgenes are usually placed under bacterial promoters activated after the invasion of the host cell. Some researchers introduced additional modifications to the therapeutic settings to further improve TAA-directed immune responses. For example Zhu et al. (2010) designed the genetic construct coding for a fusion protein consisting of a melanoma-specific TRP2 antigen and Hsp70 immunochaperone, which should facilitate the proper presentation of antigenic peptides to cytotoxic T cells (Zhu et al. 2010). Manuel et al. (2011) proposed a combined Salmonella-based therapy in which subsequent intravenous injections of two different Salmonella strains: one – coding for STAT3 shRNA and the second coding for TAA – survivin were applied to mice bearing melanoma tumors (Manuel et al. 2011). The silencing of the tolerogenic transcription factor STAT3 before TAA delivery enabled mounting the strong anti-tumor immune response. More examples of prophylactic and therapeutic approaches utilizing Salmonella can be found in the extensive review by Chorobik et al. (Chorobik et al. 2013).
Other Microbial Species Might Also Be Usefulin Cancer Treatment
Species of Salmonella posses many features of a perfect bacterial cancer therapeutic. However, a number of other microorganisms proved to be advantageous anti-cancer agents. The most promising so far have been strains of Clostridium, Escherichia and Listeria (St Jean et al. 2008), as well as some lactic acid bacteria from Lactococcus, Lactobacillus and Bifidobacterium genera (Tangney 2010).
Clostridium is a genus of Gram-positive bacteria, obligatory anaerobic and capable of producing endospores. Since Clostridium spores can only germinate in oxygen-free conditions, they are particularly suited to target hypoxic or anoxic tumor regions characterized by poor or no perfusion resulting in quiescence and necrosis. This notion sparked interest as early as the 1970s (Heppner and Mose 1978), but the spore treatment alone had limited efficacy against better-vascularized tumor regions, small tumors or metastases. A novel treatment approach, known as combined bacteriolytic therapy, was developed to utilize hypoxia-specific accumulation of bacteria; one of the most effective examples was an attenuated C. novyi-based treatment resulting in lysis of experimental tumors formed by human colon carcinoma and murine melanoma (Dang et al. 2001). Various other Clostridium- based strategies are currently being investigated, including prodrug-, cytokine- and antibody-combined approaches (Umer et al. 2012).
Bacteria from the well-known genus of Escherichia also have a therapeutic potential against cancer . Probiotic E. coli Nissle 1917 was shown to accumulate in tumors and replicate at the border of live and necrotic tumor tissue, while the colonization levels in the spleen and liver versus tumor tissue were very low (Stritzker et al. 2007). The administration of K-12, another strain of E. coli, to mice bearing murine breast carcinomas effectively stimulated an anti-tumor immune response and resulted in major reduction of lung metastases (Weibel et al. 2008). Due to its facultatively anaerobic metabolism, the mechanisms of E. coli tumor targeting are probably similar to those of Salmonella . However, since Escherichia does not invade cells, its possible applications are limited to site-specific expression of proteins within the tumor tissue.
A number of tumor therapies using Listeria have been under development in recent years. L. monocytogenes is a Gram-positive, intracellular pathogen that causes foodborne infections. Most Listeria-based treatments use a strategy different than other bacterial cancer therapies – instead of tumor targeting to perform an intratumoral action, the bacteria are used as live vaccine vectors that can deliver tumor-related antigens to non-tumor cells and thereby stimulate systemic anticancer immune responses. An example of this approach is L. monocytogenes which is able to express E7 antigen of human papilloma virus (HPV )-16 directly within APCs; human trials with patients bearing HPV-induced tumors provided promising clinical data (Le et al. 2012). Another interesting concept utilizing Listeria is targeting tumors via L. monocytogenes-infected MDSCs; the bacteria, labeled with 188Rhenium, successfully delivered the radioactive cargo into the tumor tissue (Chandra and Gravekamp 2013).
Another group of bacteria used for tumor targeting are endosymbiotic strains of Lactococcus, Lactobacillus and Bifidobacterium that share ability to produce lactic acid and are commonly utilized in food and dairy fermentations. Having a long track record of safety in humans, or even health-promoting or probiotic benefits, these bacteria are also able to colonize tumor tissue and can be potentially useful for gene-based treatment and/or detection of cancer . For example, oral administration of obligatory anaerobic B. breve can result in its translocation from the gastrointestinal tract via bloodstream into the tumor, where it was shown to express a reporter gene (Cronin et al. 2010).
Future Perspectives
In order to be successful, any bacterial cancer therapy will need to address a number of issues that are limiting to current treatment options. One of the most important ones, relevant to virtually all chemotherapeutic and biological agents, is the limited accessibility of the tumor tissue to passively-distributed therapeutics. Bacterial motility and environmental sensing can be particularly useful for tumor localization; however, the effective targeting is required not solely in the animal model s , but also in the human context, which to date remains challenging.
Therapeutic bacteria are expected to be safe, but also be able to completely switch the tumor microenvironment from immunosuppression into immunoactivation. While many studies proved the feasibility of this approach, it is important to note that preventing toxic effects by attenuation seems to be a double-edged sword, as it may compromise other therapeutically-relevant features such as invasion, colonization or immunostimulation. The balance between safety and immunogenicity is vital for a clinically-meaningful anticancer effect.
The concept of using bacteria against cancer is not necessary related to a single-agent therapy. A more feasible regimen would include initial treatment with therapeutic strains (administered orally, intravenously or intratumorally, depending on the disease), followed by surgical removal of the tumor mass. Additional follow-up with bacteria to treat the minimal residual disease (MRD) is likely to improve clinical outcomes. Combining bacterial therapy with other treatment modalities may result in stronger impact on the tumor microenvironment , which would improve cancer-specific responses.
Another question is the importance of recurring contact with bacteria to the therapeutic benefits. The role of pre-existing immunity, e.g. against pathogenic Salmonella , is unclear – it may increase clearance of the bacteria on one hand, but also potentiate anti-cancer effects on the other. This issue might be particularly relevant to multiple dosing regimens, in which the bacteria-based therapeutic is applied repeatedly to the same patient – for a prolonged, perhaps even life-long therapy.
Abbreviations
- APC:
-
Antigen-Presenting Cell
- CEA:
-
CarcinoEmbryonic Antigen
- DAMP:
-
Damage-Associated Molecular Pattern
- DC:
-
Dendritic Cell
- IDO:
-
Indoleamine 2,3-DiOxygenase
- IFN-I:
-
type I InterFeroN
- IFNγ:
-
InterFeroN gamma
- IL:
-
InterLeukin
- LPS:
-
LipoPolySaccharide
- MDSC:
-
Myeloid-Derived Suppressor Cells
- MHC:
-
Major Histocompatibility Complex
- MMP:
-
Matrix MetalloProtease
- MTD:
-
Maximum Tolerated Dose
- PAMP:
-
Pathogen-Associated Molecular Pattern
- scFv:
-
single chain variable fragment antibody
- SCV:
-
Salmonella-Containing Vacuole
- shRNA:
-
short hairpin RNA
- SPI:
-
Salmonella Pathogenicity Island
- STAT3:
-
Signal Transducer and Activator of Transcription 3
- TTSS:
-
Type III Secretion System
- TAA:
-
Tumor-Associated Antigen
- TAM:
-
Tumor-Associated Macrophage
- TLR:
-
Toll-Like Receptor
- TNF:
-
Tumor Necrosis Factor
- TRAIL:
-
TNF-Related Apoptosis-Inducing Ligand
- Treg:
-
regulatory T cell
- VEGF:
-
Vascular Endothelial Growth Factor
References
al-Ramadi BK, Fernandez-Cabezudo MJ, El-Hasasna H, Al-Salam S, Bashir G, Chouaib S (2009) Potent anti-tumor activity of systemically-administered IL2-expressing Salmonella correlates with decreased angiogenesis and enhanced tumor apoptosis. Clin Immunol 130(1):89–97. doi:10.1016/j.clim.2008.08.021
Anderson DG, Kowalczykowski SC (1998) Reconstitution of an SOS response pathway: derepression of transcription in response to DNA breaks. Cell 95(7):975–979
Angelakopoulos H, Hohmann EL (2000) Pilot study of phoP/phoQ-deleted Salmonella enterica serovar typhimurium expressing Helicobacter pylori urease in adult volunteers. Infect Immun 68(4):2135–2141
Avogadri F, Martinoli C, Petrovska L, Chiodoni C, Transidico P, Bronte V, Longhi R, Colombo MP, Dougan G, Rescigno M (2005) Cancer immunotherapy based on killing of Salmonella-infected tumor cells. Cancer Res 65(9):3920–3927. doi:10.1158/0008-5472.CAN-04-3002
Baker S, Dougan G (2007) The genome of Salmonella enterica serovar Typhi. Clin Infect Dis 45(Suppl 1):S29–S33. doi:10.1086/518143
Balkwill FR, Mantovani A (2012) Cancer-related inflammation: common themes and therapeutic opportunities. Semin Cancer Biol 22(1):33–40. doi:10.1016/j.semcancer.2011.12.005
Baumler AJ, Tsolis RM, Ficht TA, Adams LG (1998) Evolution of host adaptation in Salmonella enterica. Infect Immun 66(10):4579–4587
Bereta M, Hayhurst A, Gajda M, Chorobik P, Targosz M, Marcinkiewicz J, Kaufman HL (2007) Improving tumor targeting and therapeutic potential of Salmonella VNP20009 by displaying cell surface CEA-specific antibodies. Vaccine 25(21):4183–4192. doi:10.1016/j.vaccine.2007.03.008
Biragyn A, Longo DL (2012) Neoplastic “Black Ops”: cancer’s subversive tactics in overcoming host defenses. Semin Cancer Biol 22(1):50–59. doi:10.1016/j.semcancer.2012.01.005
Birmingham CL, Smith AC, Bakowski MA, Yoshimori T, Brumell JH (2006) Autophagy controls Salmonella infection in response to damage to the Salmonella-containing vacuole. J Biol Chem 281(16):11374–11383. doi:10.1074/jbc.M509157200
Blache CA, Manuel ER, Kaltcheva TI, Wong AN, Ellenhorn JD, Blazar BR, Diamond DJ (2012) Systemic delivery of Salmonella typhimurium transformed with IDO shRNA enhances intratumoral vector colonization and suppresses tumor growth. Cancer Res 72(24):6447–6456. doi:10.1158/0008-5472.CAN-12-0193
Cao HD, Yang YX, Lu L, Liu SN, Wang PL, Tao XH, Wang LJ, Xiang TX (2010) Attenuated Salmonella typhimurium carrying TRAIL and VP3 genes inhibits the growth of gastric cancer cells in vitro and in vivo. Tumori 96(2):296–303
Chandra D, Gravekamp C (2013) Myeloid-derived suppressor cells: cellular missiles to target tumors. Oncoimmunology 2(11):e26967. doi:10.4161/onci.26967
Chen J, Yang B, Cheng X, Qiao Y, Tang B, Chen G, Wei J, Liu X, Cheng W, Du P, Huang X, Jiang W, Hu Q, Hu Y, Li J, Hua ZC (2012) Salmonella-mediated tumor-targeting TRAIL gene therapy significantly suppresses melanoma growth in mouse model. Cancer Sci 103(2):325–333. doi:10.1111/j.1349-7006.2011.02147.x
Choi SY, Collins CC, Gout PW, Wang Y (2013) Cancer-generated lactic acid: a regulatory, immunosuppressive metabolite? J Pathol 230(4):350–355. doi:10.1002/path.4218
Chorobik P, Czaplicki D, Ossysek K, Bereta J (2013) Salmonella and cancer: from pathogens to therapeutics. Acta Biochim Pol 60(3):285–297
Chou CK, Hung JY, Liu JC, Chen CT, Hung MC (2006) An attenuated Salmonella oral DNA vaccine prevents the growth of hepatocellular carcinoma and colon cancer that express alpha-fetoprotein. Cancer Gene Ther 13(8):746–752. doi:10.1038/sj.cgt.7700927
Chow MT, Moller A, Smyth MJ (2012) Inflammation and immune surveillance in cancer. Semin Cancer Biol 22(1):23–32. doi:10.1016/j.semcancer.2011.12.004
Clairmont C, Lee KC, Pike J, Ittensohn M, Low KB, Pawelek J, Bermudes D, Brecher SM, Margitich D, Turnier J, Li Z, Luo X, King I, Zheng LM (2000) Biodistribution and genetic stability of the novel antitumor agent VNP20009, a genetically modified strain of Salmonella typhimurium. J Infect Dis 181(6):1996–2002. doi:10.1086/315497
Cookson BT, Brennan MA (2001) Pro-inflammatory programmed cell death. Trends Microbiol 9(3):113–114
Cronin M, Morrissey D, Rajendran S, El Mashad SM, van Sinderen D, O’Sullivan GC, Tangney M (2010) Orally administered bifidobacteria as vehicles for delivery of agents to systemic tumors. Mol Ther 18(7):1397–1407. doi:10.1038/mt.2010.59
Crull K, Bumann D, Weiss S (2011) Influence of infection route and virulence factors on colonization of solid tumors by Salmonella enterica serovar Typhimurium. FEMS Immunol Med Microbiol 62(1):75–83. doi:10.1111/j.1574-695X.2011.00790.x
Dang LH, Bettegowda C, Huso DL, Kinzler KW, Vogelstein B (2001) Combination bacteriolytic therapy for the treatment of experimental tumors. Proc Natl Acad Sci U S A 98(26):15155–15160. doi:10.1073/pnas.251543698
Deng Y, Lin Y, Wu X (2002) TRAIL-induced apoptosis requires Bax-dependent mitochondrial release of Smac/DIABLO. Genes Dev 16(1):33–45. doi:10.1101/gad.949602
Deretic V (2011) Autophagy in immunity and cell-autonomous defense against intracellular microbes. Immunol Rev 240(1):92–104. doi:10.1111/j.1600-065X.2010.00995.x
Desai PT, Porwollik S, Long F, Cheng P, Wollam A, Clifton S, Weinstock GM, McClelland M (2013) Evolutionary genomics of the Salmonella enterica subspecies. mBio 4(2):e00579–12. doi:10.1128/mBio.00579-12
Dresch C, Leverrier Y, Marvel J, Shortman K (2012) Development of antigen cross-presentation capacity in dendritic cells. Trends Immunol 33(8):381–388. doi:10.1016/j.it.2012.04.009
Dunn GP, Bruce AT, Ikeda H, Old LJ, Schreiber RD (2002) Cancer immunoediting: from immunosurveillance to tumor escape. Nat Immunol 3(11):991–998. doi:10.1038/ni1102-991
Espert L, Codogno P, Biard-Piechaczyk M (2007) Involvement of autophagy in viral infections: antiviral function and subversion by viruses. J Mol Med (Berl) 85(8):811–823. doi:10.1007/s00109-007-0173-6
Ewaschuk JB, Backer JL, Churchill TA, Obermeier F, Krause DO, Madsen KL (2007) Surface expression of Toll-like receptor 9 is upregulated on intestinal epithelial cells in response to pathogenic bacterial DNA. Infect Immun 75(5):2572–2579. doi:10.1128/IAI.01662-06
Fest S, Huebener N, Bleeke M, Durmus T, Stermann A, Woehler A, Baykan B, Zenclussen AC, Michalsky E, Jaeger IS, Preissner R, Hohn O, Weixler S, Gaedicke G, Lode HN (2009) Survivin minigene DNA vaccination is effective against neuroblastoma. Int J Cancer 125(1):104–114. doi:10.1002/ijc.24291
Forbes NS (2010) Engineering the perfect (bacterial) cancer therapy. Nat Rev Cancer 10(11):785–794. doi:10.1038/nrc2934
Forbes JR, Gros P (2003) Iron, manganese, and cobalt transport by Nramp1 (Slc11a1) and Nramp2 (Slc11a2) expressed at the plasma membrane. Blood 102(5):1884–1892. doi:10.1182/blood-2003-02-0425
Fridman WH, Pages F, Sautes-Fridman C, Galon J (2012) The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 12(4):298–306. doi:10.1038/nrc3245
Fu W, Chu L, Han X, Liu X, Ren D (2008) Synergistic antitumoral effects of human telomerase reverse transcriptase-mediated dual-apoptosis-related gene vector delivered by orally attenuated Salmonella enterica Serovar Typhimurium in murine tumor models. J Gene Med 10(6):690–701. doi:10.1002/jgm.1191
Gabrilovich DI, Bronte V, Chen SH, Colombo MP, Ochoa A, Ostrand-Rosenberg S, Schreiber H (2007) The terminology issue for myeloid-derived suppressor cells. Cancer Res 67(1):425. doi:10.1158/0008-5472.CAN-06-3037; author reply 426
Gabrilovich DI, Ostrand-Rosenberg S, Bronte V (2012) Coordinated regulation of myeloid cells by tumours. Nat Rev Immunol 12(4):253–268. doi:10.1038/nri3175
Galdiero MR, Bonavita E, Barajon I, Garlanda C, Mantovani A, Jaillon S (2013) Tumor associated macrophages and neutrophils in cancer. Immunobiology 218(11):1402–1410. doi:10.1016/j.imbio.2013.06.003
Galen JE, Curtiss R 3rd (2013) The delicate balance in genetically engineering live vaccines. Vaccine 32(35):4376-4385. doi:10.1016/j.vaccine.2013.12.026
Ganai S, Arenas RB, Forbes NS (2009) Tumour-targeted delivery of TRAIL using Salmonella typhimurium enhances breast cancer survival in mice. Br J Cancer 101(10):1683–1691. doi:10.1038/sj.bjc.6605403
Garcia-del Portillo F, Zwick MB, Leung KY, Finlay BB (1993) Salmonella induces the formation of filamentous structures containing lysosomal membrane glycoproteins in epithelial cells. Proc Natl Acad Sci U S A 90(22):10544–10548
Germano G, Frapolli R, Belgiovine C, Anselmo A, Pesce S, Liguori M, Erba E, Uboldi S, Zucchetti M, Pasqualini F, Nebuloni M, van Rooijen N, Mortarini R, Beltrame L, Marchini S, Fuso Nerini I, Sanfilippo R, Casali PG, Pilotti S, Galmarini CM, Anichini A, Mantovani A, D’Incalci M, Allavena P (2013) Role of macrophage targeting in the antitumor activity of trabectedin. Cancer Cell 23(2):249–262. doi:10.1016/j.ccr.2013.01.008
Goldszmid RS, Dzutsev A, Trinchieri G (2014) Host immune response to infection and cancer: unexpected commonalities. Cell Host Microbe 15(3):295–305. doi:10.1016/j.chom.2014.02.003
Gordon MA (2008) Salmonella infections in immunocompromised adults. J Infect 56(6):413–422. doi:10.1016/j.jinf.2008.03.012
Guan GF, Zhao M, Liu LM, Jin CS, Sun K, Zhang DJ, Yu DJ, Cao HW, Lu YQ, Wen LJ (2013) Salmonella typhimurium mediated delivery of Apoptin in human laryngeal cancer. Int J Med Sci 10(12):1639–1648. doi:10.7150/ijms.6960
Ha XQ, Yin Q, Zhao HB, Hui L, Wang ML, Peng JH, Dong JZ, Deng ZY, Zhao Y, Zhang YY (2012) Inhibitory effects of the attenuated Salmonella typhimurium containing the IL-2 gene on hepatic tumors in mice. J Biomed Biotechnol 2012:946139. doi:10.1155/2012/946139
Hayashi K, Zhao M, Yamauchi K, Yamamoto N, Tsuchiya H, Tomita K, Hoffman RM (2009a) Cancer metastasis directly eradicated by targeted therapy with a modified Salmonella typhimurium. J Cell Biochem 106(6):992–998. doi:10.1002/jcb.22078
Hayashi K, Zhao M, Yamauchi K, Yamamoto N, Tsuchiya H, Tomita K, Kishimoto H, Bouvet M, Hoffman RM (2009b) Systemic targeting of primary bone tumor and lung metastasis of high-grade osteosarcoma in nude mice with a tumor-selective strain of Salmonella typhimurium. Cell Cycle 8(6):870–875
Heimann DM, Rosenberg SA (2003) Continuous intravenous administration of live genetically modified salmonella typhimurium in patients with metastatic melanoma. J Immunother 26(2):179–180
Heppner F, Mose JR (1978) The liquefaction (oncolysis) of malignant gliomas by a non pathogenic Clostridium. Acta Neurochir (Wien) 42(1–2):123–125
Hernandez LD, Pypaert M, Flavell RA, Galan JE (2003) A Salmonella protein causes macrophage cell death by inducing autophagy. J Cell Biol 163(5):1123–1131. doi:10.1083/jcb.200309161
Hiroshima Y, Zhao M, Maawy A, Zhang Y, Katz MH, Fleming JB, Uehara F, Miwa S, Yano S, Momiyama M, Suetsugu A, Chishima T, Tanaka K, Bouvet M, Endo I, Hoffman RM (2014) Efficacy of Salmonella typhimurium A1-R Versus chemotherapy on a pancreatic cancer Patient-derived orthotopic xenograft (PDOX). J Cell Biochem 115(7):1254–1261. doi:10.1002/jcb.24769
Hong EH, Chang SY, Lee BR, Pyun AR, Kim JW, Kweon MN, Ko HJ (2013) Intratumoral injection of attenuated Salmonella vaccine can induce tumor microenvironmental shift from immune suppressive to immunogenic. Vaccine 31(10):1377–1384. doi:10.1016/j.vaccine.2013.01.006
Hu ZQ, Zhao WH (2013) Type 1 interferon-associated necroptosis: a novel mechanism for Salmonella enterica Typhimurium to induce macrophage death. Cell Mol Immunol 10(1):10–12. doi:10.1038/cmi.2012.54
Jarosz M, Jazowiecka-Rakus J, Cichon T, Glowala-Kosinska M, Smolarczyk R, Smagur A, Malina S, Sochanik A, Szala S (2013) Therapeutic antitumor potential of endoglin-based DNA vaccine combined with immunomodulatory agents. Gene Ther 20(3):262–273. doi:10.1038/gt.2012.28
Jeong JH, Kim K, Lim D, Jeong K, Hong Y, Nguyen VH, Kim TH, Ryu S, Lim JA, Kim JI, Kim GJ, Kim SC, Min JJ, Choy HE (2014) Anti-tumoral effect of the mitochondrial target domain of Noxa delivered by an engineered Salmonella typhimurium. PLoS One 9(1):e80050. doi:10.1371/journal.pone.0080050
Joffre OP, Segura E, Savina A, Amigorena S (2012) Cross-presentation by dendritic cells. Nat Rev Immunol 12(8):557–569. doi:10.1038/nri3254
Kaimala S, Mohamed YA, Nader N, Issac J, Elkord E, Chouaib S, Fernandez-Cabezudo MJ, Al-Ramadi BK (2014) Salmonella-mediated tumor regression involves targeting of tumor myeloid suppressor cells causing a shift to M1-like phenotype and reduction in suppressive capacity. Cancer Immunol Immunother 63(6):587–599. doi:10.1007/s00262-014-1543-x
Kalinski P (2012) Regulation of immune responses by prostaglandin E2. J Immunol 188(1):21–28. doi:10.4049/jimmunol.1101029
Kawamoto A, Morimoto YV, Miyata T, Minamino T, Hughes KT, Kato T, Namba K (2013) Common and distinct structural features of Salmonella injectisome and flagellar basal body. Sci Rep 3:3369. doi:10.1038/srep03369
Kim NW, Piatyszek MA, Prowse KR, Harley CB, West MD, Ho PL, Coviello GM, Wright WE, Weinrich SL, Shay JW (1994) Specific association of human telomerase activity with immortal cells and cancer. Science 266(5193):2011–2015
Kim JM, Eckmann L, Savidge TC, Lowe DC, Witthoft T, Kagnoff MF (1998) Apoptosis of human intestinal epithelial cells after bacterial invasion. J Clin Invest 102(10):1815–1823. doi:10.1172/JCI2466
Kimura H, Zhang L, Zhao M, Hayashi K, Tsuchiya H, Tomita K, Bouvet M, Wessels J, Hoffman RM (2010) Targeted therapy of spinal cord glioma with a genetically modified Salmonella typhimurium. Cell Prolif 43(1):41–48. doi:10.1111/j.1365-2184.2009.00652.x
Knodler LA, Vallance BA, Celli J, Winfree S, Hansen B, Montero M, Steele-Mortimer O (2010) Dissemination of invasive Salmonella via bacterial-induced extrusion of mucosal epithelia. Proc Natl Acad Sci U S A 107(41):17733–17738. doi:10.1073/pnas.1006098107
Kroemer G, Levine B (2008) Autophagic cell death: the story of a misnomer. Nat Rev Mol Cell Biol 9(12):1004–1010. doi:10.1038/nrm2529
Lahiri A, Das P, Vani J, Shaila MS, Chakravortty D (2010) TLR 9 activation in dendritic cells enhances salmonella killing and antigen presentation via involvement of the reactive oxygen species. PLoS One 5(10):e13772. doi:10.1371/journal.pone.0013772
Le DT, Dubenksy TW Jr, Brockstedt DG (2012) Clinical development of Listeria monocytogenes-based immunotherapies. Semin Oncol 39(3):311–322. doi:10.1053/j.seminoncol.2012.02.008
Lee KC, Zheng L-M, Luo X, Clairmont C, Fischer J, Margitich D, Turnier J, Almassian B, Bermudes D, King I (2000) Comparative evaluation of the acute toxic effects in monkeys, pigs and mice of a genetically engineered salmonella strain (VNP20009) being developed as an antitumor agent. Int J Toxicol 19(1):19–25
Lee JS, Jung ID, Lee CM, Park JW, Chun SH, Jeong SK, Ha TK, Shin YK, Kim DJ, Park YM (2010) Outer membrane protein a of Salmonella enterica serovar Typhimurium activates dendritic cells and enhances Th1 polarization. BMC Microbiol 10:263. doi:10.1186/1471-2180-10-263
Lee CH, Lin ST, Liu JJ, Chang WW, Hsieh JL, Wang WK (2014) Salmonella induce autophagy in melanoma by the downregulation of AKT/mTOR pathway. Gene Ther 21(3):309–316. doi:10.1038/gt.2013.86
Leschner S, Westphal K, Dietrich N, Viegas N, Jablonska J, Lyszkiewicz M, Lienenklaus S, Falk W, Gekara N, Loessner H, Weiss S (2009) Tumor invasion of Salmonella enterica serovar Typhimurium is accompanied by strong hemorrhage promoted by TNF-alpha. PLoS One 4(8):e6692. doi:10.1371/journal.pone.0006692
Lindgren SW, Stojiljkovic I, Heffron F (1996) Macrophage killing is an essential virulence mechanism of Salmonella typhimurium. Proc Natl Acad Sci U S A 93(9):4197–4201
Loeffler M, Le’Negrate G, Krajewska M, Reed JC (2007) Attenuated Salmonella engineered to produce human cytokine LIGHT inhibit tumor growth. Proc Natl Acad Sci U S A 104(31):12879–12883. doi:10.1073/pnas.0701959104
Loeffler M, Le’Negrate G, Krajewska M, Reed JC (2008) IL-18-producing Salmonella inhibit tumor growth. Cancer Gene Ther 15(12):787–794. doi:10.1038/cgt.2008.48
Loeffler M, Le’Negrate G, Krajewska M, Reed JC (2009) Salmonella typhimurium engineered to produce CCL21 inhibit tumor growth. Cancer Immunol Immunother 58(5):769–775. doi:10.1007/s00262-008-0555-9
Lorenzi S, Mattei F, Sistigu A, Bracci L, Spadaro F, Sanchez M, Spada M, Belardelli F, Gabriele L, Schiavoni G (2011) Type I IFNs control antigen retention and survival of CD8alpha(+) dendritic cells after uptake of tumor apoptotic cells leading to cross-priming. J Immunol 186(9):5142–5150. doi:10.4049/jimmunol.1004163
Lu H (2014) TLR agonists for cancer immunotherapy: tipping the balance between the immune stimulatory and inhibitory effects. Front Immunol 5:83. doi:10.3389/fimmu.2014.00083
Lu JV, Walsh CM (2012) Programmed necrosis and autophagy in immune function. Immunol Rev 249(1):205–217. doi:10.1111/j.1600-065X.2012.01147.x
Ma Y, Shurin GV, Peiyuan Z, Shurin MR (2013) Dendritic cells in the cancer microenvironment. J Cancer 4(1):36–44. doi:10.7150/jca.5046
Mai CW, Kang YB, Pichika MR (2013) Should a Toll-like receptor 4 (TLR-4) agonist or antagonist be designed to treat cancer? TLR-4: its expression and effects in the ten most common cancers. Onco Targets Ther 6:1573–1587. doi:10.2147/OTT.S50838
Manuel ER, Blache CA, Paquette R, Kaltcheva TI, Ishizaki H, Ellenhorn JD, Hensel M, Metelitsa L, Diamond DJ (2011) Enhancement of cancer vaccine therapy by systemic delivery of a tumor-targeting Salmonella-based STAT3 shRNA suppresses the growth of established melanoma tumors. Cancer Res 71(12):4183–4191. doi:10.1158/0008-5472.CAN-10-4676
Massa PE, Paniccia A, Monegal A, de Marco A, Rescigno M (2013) Salmonella engineered to express CD20-targeting antibodies and a drug-converting enzyme can eradicate human lymphomas. Blood 122(5):705–714. doi:10.1182/blood-2012-12-474098
Mastroeni P, Grant A, Restif O, Maskell D (2009) A dynamic view of the spread and intracellular distribution of Salmonella enterica. Nat Rev Microbiol 7(1):73–80. doi:10.1038/nrmicro2034
Matzinger P (2012) The evolution of the danger theory. Interview by Lauren Constable, Commissioning Editor. Expert Rev Clin Immunol 8(4):311–317. doi:10.1586/eci.12.21
McClelland M, Sanderson KE, Spieth J, Clifton SW, Latreille P, Courtney L, Porwollik S, Ali J, Dante M, Du F, Hou S, Layman D, Leonard S, Nguyen C, Scott K, Holmes A, Grewal N, Mulvaney E, Ryan E, Sun H, Florea L, Miller W, Stoneking T, Nhan M, Waterston R, Wilson RK (2001) Complete genome sequence of Salmonella enterica serovar Typhimurium LT2. Nature 413(6858):852–856. doi:10.1038/35101614
McLaughlin LM, Govoni GR, Gerke C, Gopinath S, Peng K, Laidlaw G, Chien YH, Jeong HW, Li Z, Brown MD, Sacks DB, Monack D (2009) The Salmonella SPI2 effector SseI mediates long-term systemic infection by modulating host cell migration. PLoS Pathog 5(11):e1000671. doi:10.1371/journal.ppat.1000671
Meunier E, Dick MS, Dreier RF, Schurmann N, Broz DK, Warming S, Roose-Girma M, Bumann D, Kayagaki N, Takeda K, Yamamoto M, Broz P (2014) Caspase-11 activation requires lysis of pathogen-containing vacuoles by IFN-induced GTPases. Nature 509(7500):366–370. doi:10.1038/nature13157
Miller SI, Ernst RK, Bader MW (2005) LPS, TLR4 and infectious disease diversity. Nat Rev Microbiol 3(1):36–46. doi:10.1038/nrmicro1068
Monack DM, Bouley DM, Falkow S (2004) Salmonella typhimurium persists within macrophages in the mesenteric lymph nodes of chronically infected Nramp1+/+ mice and can be reactivated by IFNgamma neutralization. J Exp Med 199(2):231–241. doi:10.1084/jem.20031319
Mota LJ, Ramsden AE, Liu M, Castle JD, Holden DW (2009) SCAMP3 is a component of the Salmonella-induced tubular network and reveals an interaction between bacterial effectors and post-Golgi trafficking. Cell Microbiol 11(8):1236–1253. doi:10.1111/j.1462-5822.2009.01329.x
Nagakura C, Hayashi K, Zhao M, Yamauchi K, Yamamoto N, Tsuchiya H, Tomita K, Bouvet M, Hoffman RM (2009) Efficacy of a genetically-modified Salmonella typhimurium in an orthotopic human pancreatic cancer in nude mice. Anticancer Res 29(6):1873–1878
Nemunaitis J, Cunningham C, Senzer N, Kuhn J, Cramm J, Litz C, Cavagnolo R, Cahill A, Clairmont C, Sznol M (2003) Pilot trial of genetically modified, attenuated Salmonella expressing the E. coli cytosine deaminase gene in refractory cancer patients. Cancer Gene Ther 10(10):737–744. doi:10.1038/sj.cgt.7700634
Niethammer AG, Lubenau H, Mikus G, Knebel P, Hohmann N, Leowardi C, Beckhove P, Akhisaroglu M, Ge Y, Springer M, Grenacher L, Buchler MW, Koch M, Weitz J, Haefeli WE, Schmitz-Winnenthal FH (2012) Double-blind, placebo-controlled first in human study to investigate an oral vaccine aimed to elicit an immune reaction against the VEGF-Receptor 2 in patients with stage IV and locally advanced pancreatic cancer. BMC Cancer 12:361. doi:10.1186/1471-2407-12-361
Nishikawa H, Sato E, Briones G, Chen LM, Matsuo M, Nagata Y, Ritter G, Jager E, Nomura H, Kondo S, Tawara I, Kato T, Shiku H, Old LJ, Galan JE, Gnjatic S (2006) In vivo antigen delivery by a Salmonella typhimurium type III secretion system for therapeutic cancer vaccines. J Clin Invest 116(7):1946–1954. doi:10.1172/JCI28045
Ochman H, Wilson AC (1987) Evolution in bacteria: evidence for a universal substitution rate in cellular genomes. J Mol Evol 26(1–2):74–86
Olsthoorn MM, Petersen BO, Duus J, Haverkamp J, Thomas-Oates JE, Bock K, Holst O (2000) The structure of the linkage between the O-specific polysaccharide and the core region of the lipopolysaccharide from Salmonella enterica serovar Typhimurium revisited. Eur J Biochem 267(7):2014–2027
Paesold G, Guiney DG, Eckmann L, Kagnoff MF (2002) Genes in the Salmonella pathogenicity island 2 and the Salmonella virulence plasmid are essential for Salmonella-induced apoptosis in intestinal epithelial cells. Cell Microbiol 4(11):771–781
Pages F, Kirilovsky A, Mlecnik B, Asslaber M, Tosolini M, Bindea G, Lagorce C, Wind P, Marliot F, Bruneval P, Zatloukal K, Trajanoski Z, Berger A, Fridman WH, Galon J (2009) In situ cytotoxic and memory T cells predict outcome in patients with early-stage colorectal cancer. J Clin Oncol 27(35):5944–5951. doi:10.1200/JCO.2008.19.6147
Paglia P, Medina E, Arioli I, Guzman CA, Colombo MP (1998) Gene transfer in dendritic cells, induced by oral DNA vaccination with Salmonella typhimurium, results in protective immunity against a murine fibrosarcoma. Blood 92(9):3172–3176
Radics J, Konigsmaier L, Marlovits TC (2014) Structure of a pathogenic type 3 secretion system in action. Nat Struct Mol Biol 21(1):82–87. doi:10.1038/nsmb.2722
Ramos-Morales F (2012) Impact of Salmonella enterica type III secretion system effectors on the eukaryotic host cell. ISRN Cell Biol 2012:36. doi:10.5402/2012/787934
Roider E, Jellbauer S, Kohn B, Berchtold C, Partilla M, Busch DH, Russmann H, Panthel K (2011) Invasion and destruction of a murine fibrosarcoma by Salmonella-induced effector CD8 T cells as a therapeutic intervention against cancer. Cancer Immunol Immunother 60(3):371–380. doi:10.1007/s00262-010-0950-x
Roland KL, Brenneman KE (2013) Salmonella as a vaccine delivery vehicle. Expert Rev Vaccines 12(9):1033–1045. doi:10.1586/14760584.2013.825454
Ruby T, McLaughlin L, Gopinath S, Monack D (2012) Salmonella’s long-term relationship with its host. FEMS Microbiol Rev 36(3):600–615. doi:10.1111/j.1574-6976.2012.00332.x
Russmann H, Shams H, Poblete F, Fu Y, Galan JE, Donis RO (1998) Delivery of epitopes by the Salmonella type III secretion system for vaccine development. Science 281(5376):565–568
Ryter SW, Mizumura K, Choi AM (2014) The impact of autophagy on cell death modalities. Int J Cell Biol 2014:502676. doi:10.1155/2014/502676
Salcedo SP, Noursadeghi M, Cohen J, Holden DW (2001) Intracellular replication of Salmonella typhimurium strains in specific subsets of splenic macrophages in vivo. Cell Microbiol 3(9):587–597
Saunders NA, Simpson F, Thompson EW, Hill MM, Endo-Munoz L, Leggatt G, Minchin RF, Guminski A (2012) Role of intratumoural heterogeneity in cancer drug resistance: molecular and clinical perspectives. EMBO Mol Med 4(8):675–684. doi:10.1002/emmm.201101131
Schleker S, Sun J, Raghavan B, Srnec M, Muller N, Koepfinger M, Murthy L, Zhao Z, Klein-Seetharaman J (2012) The current Salmonella-host interactome. Proteomics Clin Appl 6(1–2):117–133. doi:10.1002/prca.201100083
Schraidt O, Marlovits TC (2011) Three-dimensional model of Salmonella’s needle complex at subnanometer resolution. Science 331(6021):1192–1195. doi:10.1126/science.1199358
Seo YW, Woo HN, Piya S, Moon AR, Oh JW, Yun CW, Kim KK, Min JY, Jeong SY, Chung S, Song PI, Choi EK, Seol DW, Kim TH (2009) The cell death-inducing activity of the peptide containing Noxa mitochondrial-targeting domain is associated with calcium release. Cancer Res 69(21):8356–8365. doi:10.1158/0008-5472.CAN-09-0349
Shafer WM, Casey SG, Spitznagel JK (1984) Lipid A and resistance of Salmonella typhimurium to antimicrobial granule proteins of human neutrophil granulocytes. Infect Immun 43(3):834–838
Shahnazari S, Yen WL, Birmingham CL, Shiu J, Namolovan A, Zheng YT, Nakayama K, Klionsky DJ, Brumell JH (2010) A diacylglycerol-dependent signaling pathway contributes to regulation of antibacterial autophagy. Cell Host Microbe 8(2):137–146. doi:10.1016/j.chom.2010.07.002
Song L, Asgharzadeh S, Salo J, Engell K, Wu HW, Sposto R, Ara T, Silverman AM, DeClerck YA, Seeger RC, Metelitsa LS (2009) Valpha24-invariant NKT cells mediate antitumor activity via killing of tumor-associated macrophages. J Clin Invest 119(6):1524–1536. doi:10.1172/JCI37869
Spel L, Boelens JJ, Nierkens S, Boes M (2013) Antitumor immune responses mediated by dendritic cells: how signals derived from dying cancer cells drive antigen cross-presentation. Oncoimmunology 2(11):e26403. doi:10.4161/onci.26403
St Jean AT, Zhang M, Forbes NS (2008) Bacterial therapies: completing the cancer treatment toolbox. Curr Opin Biotechnol 19(5):511–517. doi:10.1016/j.copbio.2008.08.004
Steiner TS (2007) How flagellin and toll-like receptor 5 contribute to enteric infection. Infect Immun 75(2):545–552. doi:10.1128/IAI.01506-06
Stritzker J, Weibel S, Hill PJ, Oelschlaeger TA, Goebel W, Szalay AA (2007) Tumor-specific colonization, tissue distribution, and gene induction by probiotic Escherichia coli Nissle 1917 in live mice. Int J Med Microbiol 297(3):151–162. doi:10.1016/j.ijmm.2007.01.008
Sylvester RJ (2011) Bacillus Calmette-Guerin treatment of non-muscle invasive bladder cancer. Int J Urol 18(2):113–120. doi:10.1111/j.1442-2042.2010.02678.x
Tangney M (2010) Gene therapy for cancer: dairy bacteria as delivery vectors. Discov Med 10(52):195–200
Thamm DH, Kurzman ID, King I, Li Z, Sznol M, Dubielzig RR, Vail DM, MacEwen EG (2005) Systemic administration of an attenuated, tumor-targeting Salmonella typhimurium to dogs with spontaneous neoplasia: phase I evaluation. Clinical Cancer Res 11(13):4827–4834. doi:10.1158/1078-0432.CCR-04-2510
Thomas DR, Francis NR, Xu C, DeRosier DJ (2006) The three-dimensional structure of the flagellar rotor from a clockwise-locked mutant of Salmonella enterica serovar Typhimurium. J Bacteriol 188(20):7039–7048. doi:10.1128/JB.00552-06
Thurston TL, Ryzhakov G, Bloor S, von Muhlinen N, Randow F (2009) The TBK1 adaptor and autophagy receptor NDP52 restricts the proliferation of ubiquitin-coated bacteria. Nat Immunol 10(11):1215–1221. doi:10.1038/ni.1800
Tome Y, Zhang Y, Momiyama M, Maehara H, Kanaya F, Tomita K, Tsuchiya H, Bouvet M, Hoffman RM, Zhao M (2013) Primer dosing of S. typhimurium A1-R potentiates tumor-targeting and efficacy in immunocompetent mice. Anticancer Res 33(1):97–102
Toso JF, Gill VJ, Hwu P, Marincola FM, Restifo NP, Schwartzentruber DJ, Sherry RM, Topalian SL, Yang JC, Stock F, Freezer LJ, Morton KE, Seipp C, Haworth L, Mavroukakis S, White D, MacDonald S, Mao J, Sznol M, Rosenberg SA (2002) Phase I study of the intravenous administration of attenuated Salmonella typhimurium to patients with metastatic melanoma. J Clin Oncol 20(1):142–152
Uhl M, Kepp O, Jusforgues-Saklani H, Vicencio JM, Kroemer G, Albert ML (2009) Autophagy within the antigen donor cell facilitates efficient antigen cross-priming of virus-specific CD8+ T cells. Cell Death Differ 16(7):991–1005. doi:10.1038/cdd.2009.8
Umer B, Good D, Anne J, Duan W, Wei MQ (2012) Clostridial spores for cancer therapy: targeting solid tumour microenvironment. J Toxicol 2012:862764. doi:10.1155/2012/862764
Vacchelli E, Eggermont A, Sautes-Fridman C, Galon J, Zitvogel L, Kroemer G, Galluzzi L (2013) Trial watch: toll-like receptor agonists for cancer therapy. Oncoimmunology 2(8):e25238. doi:10.4161/onci.25238
van der Velden AW, Baumler AJ, Tsolis RM, Heffron F (1998) Multiple fimbrial adhesins are required for full virulence of Salmonella typhimurium in mice. Infect Immun 66(6):2803–2808
Villadangos JA, Shortman K (2010) Found in translation: the human equivalent of mouse CD8+ dendritic cells. J Exp Med 207(6):1131–1134. doi:10.1084/jem.20100985
Voedisch S, Koenecke C, David S, Herbrand H, Forster R, Rhen M, Pabst O (2009) Mesenteric lymph nodes confine dendritic cell-mediated dissemination of Salmonella enterica serovar Typhimurium and limit systemic disease in mice. Infect Immun 77(8):3170–3180. doi:10.1128/IAI.00272-09
Wagner C, Polke M, Gerlach RG, Linke D, Stierhof YD, Schwarz H, Hensel M (2011) Functional dissection of SiiE, a giant non-fimbrial adhesin of Salmonella enterica. Cell Microbiol 13(8):1286–1301. doi:10.1111/j.1462-5822.2011.01621.x
Walczak H, Miller RE, Ariail K, Gliniak B, Griffith TS, Kubin M, Chin W, Jones J, Woodward A, Le T, Smith C, Smolak P, Goodwin RG, Rauch CT, Schuh JC, Lynch DH (1999) Tumoricidal activity of tumor necrosis factor-related apoptosis-inducing ligand in vivo. Nat Med 5(2):157–163. doi:10.1038/5517
Wang Z, Jiang H, Chen S, Du F, Wang X (2012) The mitochondrial phosphatase PGAM5 functions at the convergence point of multiple necrotic death pathways. Cell 148(1–2):228–243. doi:10.1016/j.cell.2011.11.030
Wang Y, Chen J, Tang B, Zhang X, Hua ZC (2013) Systemic administration of attenuated in combination with interleukin-21 for cancer therapy. Mol Clin Oncol 1(3):461–465. doi:10.3892/mco.2013.90
Weibel S, Stritzker J, Eck M, Goebel W, Szalay AA (2008) Colonization of experimental murine breast tumours by Escherichia coli K-12 significantly alters the tumour microenvironment. Cell Microbiol 10(6):1235–1248. doi:10.1111/j.1462-5822.2008.01122.x
Whiteside TL (2010) Immune responses to malignancies. J Allergy Clin Immunol 125(2 Suppl 2):S272–S283. doi:10.1016/j.jaci.2009.09.045
Wick MJ (2011) Innate immune control of Salmonella enterica serovar Typhimurium: mechanisms contributing to combating systemic Salmonella infection. J Innate Immun 3(6):543–549. doi:10.1159/000330771
Wynosky-Dolfi MA, Snyder AG, Philip NH, Doonan PJ, Poffenberger MC, Avizonis D, Zwack EE, Riblett AM, Hu B, Strowig T, Flavell RA, Jones RG, Freedman BD, Brodsky IE (2014) Oxidative metabolism enables Salmonella evasion of the NLRP3 inflammasome. J Exp Med 211(4):653–668. doi:10.1084/jem.20130627
Xia Y, Zweier JL (1997) Superoxide and peroxynitrite generation from inducible nitric oxide synthase in macrophages. Proc Natl Acad Sci U S A 94(13):6954–6958
Xin H, Zhang C, Herrmann A, Du Y, Figlin R, Yu H (2009) Sunitinib inhibition of Stat3 induces renal cell carcinoma tumor cell apoptosis and reduces immunosuppressive cells. Cancer Res 69(6):2506–2513. doi:10.1158/0008-5472.CAN-08-4323
Yagita H, Takeda K, Hayakawa Y, Smyth MJ, Okumura K (2004) TRAIL and its receptors as targets for cancer therapy. Cancer Sci 95(10):777–783
Yoon WS, Chae YS, Hong J, Park YK (2011) Antitumor therapeutic effects of a genetically engineered Salmonella typhimurium harboring TNF-alpha in mice. Appl Microbiol Biotechnol 89(6):1807–1819. doi:10.1007/s00253-010-3006-4
Yoon W, Choi JH, Kim S, Park YK (2014) Engineered Salmonella typhimurium expressing E7 fusion protein, derived from human papillomavirus, inhibits tumor growth in cervical tumor-bearing mice. Biotechnol Lett 36(2):349–356. doi:10.1007/s10529-013-1370-8
Youn JI, Gabrilovich DI (2010) The biology of myeloid-derived suppressor cells: the blessing and the curse of morphological and functional heterogeneity. Eur J Immunol 40(11):2969–2975. doi:10.1002/eji.201040895
Yrlid U, Wick MJ (2000) Salmonella-induced apoptosis of infected macrophages results in presentation of a bacteria-encoded antigen after uptake by bystander dendritic cells. J Exp Med 191(4):613–624
Yu B, Yang M, Shi L, Yao Y, Jiang Q, Li X, Tang LH, Zheng BJ, Yuen KY, Smith DK, Song E, Huang JD (2012) Explicit hypoxia targeting with tumor suppression by creating an “obligate” anaerobic Salmonella Typhimurium strain. Sci Rep 2:436. doi:10.1038/srep00436
Yu HB, Croxen MA, Marchiando AM, Ferreira RB, Cadwell K, Foster LJ, Finlay BB (2014) Autophagy facilitates Salmonella replication in HeLa cells. mBio 5(2):e00865–e00914. doi:10.1128/mBio.00865-14
Zhang XD, Zhang XY, Gray CP, Nguyen T, Hersey P (2001) Tumor necrosis factor-related apoptosis-inducing ligand-induced apoptosis of human melanoma is regulated by smac/DIABLO release from mitochondria. Cancer Res 61(19):7339–7348
Zhang L, Lopez H, George NM, Liu X, Pang X, Luo X (2011) Selective involvement of BH3-only proteins and differential targets of Noxa in diverse apoptotic pathways. Cell Death Differ 18(5):864–873. doi:10.1038/cdd.2010.152
Zhang Y, Tome Y, Suetsugu A, Zhang L, Zhang N, Hoffman RM, Zhao M (2012) Determination of the optimal route of administration of Salmonella typhimurium A1-R to target breast cancer in nude mice. Anticancer Res 32(7):2501–2508
Zhang M, Swofford CA, Forbes NS (2013a) Lipid A controls the robustness of intratumoral accumulation of attenuated Salmonella in mice. Int J Cancer 135(3):647–657. doi:10.1002/ijc.28700
Zhang Y, Zhang N, Su S, Hoffman RM, Zhao M (2013b) Salmonella typhimurium A1-R tumor targeting in immunocompetent mice is enhanced by a traditional Chinese medicine herbal mixture. Anticancer Res 33(5):1837–1843
Zhao M, Yang M, Li XM, Jiang P, Baranov E, Li S, Xu M, Penman S, Hoffman RM (2005) Tumor-targeting bacterial therapy with amino acid auxotrophs of GFP-expressing Salmonella typhimurium. Proc Natl Acad Sci U S A 102(3):755–760. doi:10.1073/pnas.0408422102
Zhao M, Yang M, Ma H, Li X, Tan X, Li S, Yang Z, Hoffman RM (2006) Targeted therapy with a Salmonella typhimurium leucine-arginine auxotroph cures orthotopic human breast tumors in nude mice. Cancer Res 66(15):7647–7652. doi:10.1158/0008-5472.CAN-06-0716
Zhao M, Geller J, Ma H, Yang M, Penman S, Hoffman RM (2007) Monotherapy with a tumor-targeting mutant of Salmonella typhimurium cures orthotopic metastatic mouse models of human prostate cancer. Proc Natl Acad Sci U S A 104(24):10170–10174. doi:10.1073/pnas.0703867104
Zhao Y, Yang J, Shi J, Gong YN, Lu Q, Xu H, Liu L, Shao F (2011) The NLRC4 inflammasome receptors for bacterial flagellin and type III secretion apparatus. Nature 477(7366):596–600. doi:10.1038/nature10510
Zheng YT, Shahnazari S, Brech A, Lamark T, Johansen T, Brumell JH (2009) The adaptor protein p62/SQSTM1 targets invading bacteria to the autophagy pathway. J Immunol 183(9):5909–5916. doi:10.4049/jimmunol.0900441
Zhu X, Zhou P, Cai J, Yang G, Liang S, Ren D (2010) Tumor antigen delivered by Salmonella III secretion protein fused with heat shock protein 70 induces protection and eradication against murine melanoma. Cancer Sci 101(12):2621–2628. doi:10.1111/j.1349-7006.2010.01722.x
Zhuang SM, Shvarts A, van Ormondt H, Jochemsen AG, van der Eb AJ, Noteborn MH (1995) Apoptin, a protein derived from chicken anemia virus, induces p53-independent apoptosis in human osteosarcoma cells. Cancer Res 55(3):486–489
Acknowledgements
The authors would like to thank Dr. Grzegorz Bereta for his help with manuscript preparation, especially for the scientific illustration, as well as fruitful discussions. The authors gratefully acknowledge financial support from the Polish National Centre for Research and Development through INNOTECH grant no. 152553 and the funding from the Jagiellonian University within the SET project co-financed by the European Union. Faculty of Biochemistry, Biophysics and Biotechnology of the Jagiellonian University in Kraków is a partner of the Leading National Research Center (KNOW) supported by the Ministry of Science and Higher Education.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this chapter
Cite this chapter
Chorobik, P., Czaplicki, D., Ossysek, K., Bereta, J. (2015). Bacterial Cancer Therapy: How Patients Might Benefit from Salmonella Infections. In: Shurin, M., Thanavala, Y., Ismail, N. (eds) Infection and Cancer: Bi-Directorial Interactions. Springer, Cham. https://doi.org/10.1007/978-3-319-20669-1_16
Download citation
DOI: https://doi.org/10.1007/978-3-319-20669-1_16
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-20668-4
Online ISBN: 978-3-319-20669-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)