Abstract
Vegetable crops have been cultivated since they were domesticated. These are the principal source of nutrients required in the growth and development of human beings. Rapid advances in the methods of next-generation sequencing technology and high throughput genotyping protocols have resulted in the collection and publication of reference genomes of major vegetable species. The large-scale genetic resource reorganization strategy has revealed the process of vegetable crops domestication and improvement of the essential traits through breeding procedures. The utilization of genetic mapping strategies and identification of quantitative trait locus has resulted in the exploration of significant molecular markers linked to essential traits in vegetables. Furthermore, the genome-based breeding approach is employed in most important vegetable crops families, such as Solanaceae and Brassicaceae, and allowing molecular selection at the single-base level. As a result, genome-wide molecular markers are extensively used for efficient genotyping in most vegetable crops. Molecular breeding has emerged as a key method for vegetables. Besides this, genome editing technology can dramatically increase vegetable breeding efficiency. This chapter examines the current scenario of genome-based molecular breeding tactics and genome editing approaches employed in significant vegetable crops to give insights into next-generation molecular breeding for a growing global population.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
13.1 Introduction
Abiotic stresses such as extremely low or high temperature, increase in soil salinity status, drought, and floods have an impact on vegetable production. Climate change significantly impacts various developmental stages of vegetables such as vegetative growth, flowering, and fruiting (Spaldon et al. 2015; Khar et al. 2022). The effects of high temperature and unpredictable rainfall can impede average plant growth and development, affecting crop yields. Environmental stress has a significant impact on soil organic matter decomposition, the nutrient cycle, and plant nutrients as well as water availability. The intensity and duration of the extreme environment, on the other hand, determines the crop growth cycle, biomass accumulation, and ultimately the range of economic benefits. Vegetable crops yields in Asia are expected to fall by 2510 per cent between 2020 and 2050, with South and Central Asia experiencing the steepest declines (Cruz et al. 2007; Ashraf et al. 2022). Several initiatives have been launched to reduce the potential impact of climate change on vegetable production. Advanced molecular breeding approaches have been used to improve the yields and quality of important vegetable crops and combat the abiotic stresses of climate change.
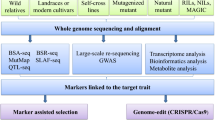
NGS (next-generation sequencing) technology has considerably accelerated vegetable crops genomic sequencing and resequencing, which resulted in the publication of chromosomal sequences of some major vegetable crops. The order is determined by the specific locus of the corresponding gene to a marker. Based on genotyping techniques, these linked markers can be used to perform molecular marker-assisted breeding and identify genetic relationships between different traits of vegetable crops. Various genome sequencing methods also contribute to the development of new molecular markers (Saha et al. 2022). Resequencing techniques rely heavily on the reference genome and bring researchers together to analyse sequence diversity at the genomic level. The primary approach, known as bulked segregant analysis (BSA), was assumed to be whole-genome resequencing employing mass separation analysis. BSA is based on the gene identification and marker development procedures, tested on a variety of vegetable crops. Creating the F2 progeny’s divergent phenotypes acquires the SNP index of the two diversified gene pools and matches them to the vegetable reference genome. Besides this, QTLseq and MutMap can also be utilised. As a result, there is a lot less doubt about other genes and phenotypes (Takagi et al. 2013). MutMap was also used to build a DNA pool from a single individual with the F2 isolation mutant phenotype and compare it to wildtype and parental reference sequences (Abe et al. 2012). It can also be used to locate the markers inside the genome. Large-scale resequencing has evolved into a more generic strategy based on improved approaches for finding markers linked with vegetable crop attributes.
Transcriptome and metabolic analyses help the researchers to domesticate genes and comprehend molecular regulatory networks at the genomic and transcriptional levels by altering gene expression patterns and analysing metabolic pathways. Existing molecular markers and reference genomes for vegetable crops can be employed to alter the vegetable genomes. CRISPR/Cas9 is a standard genome editing technique used to improve a variety of vegetable crops’ undesirable traits (Khan et al. 2019; Satish Chhapekar et al. 2022). Compared to other breeding methods, this form of targeted breeding is more efficient and time-saving (Chen et al. 2019; Zahid et al. 2022). This chapter summarises the progress of research on molecular methods used to create vegetable crops resistant to abiotic stress, including molecular breeding and genome editing approaches.
13.2 Signal Transduction Mechanism Against Abiotic Stresses in Vegetable Crops
Plants respond to stress in various ways, including altered gene expression, cellular metabolism, growth rates, and vegetable crops yields. A sudden change in environmental conditions typically causes plant stress. Abiotic stress imposed on plants by environmental factors can be physical or chemical. Drought, flood, extreme temperatures, salt and mineral toxicity, are the abiotic stresses impacting vegetable crops yields and seed production. The roots are the plant‘s first line of defence against abiotic stress. The disruption of the Na + / K + ratio in the cytoplasm (Sahoo and Mohapatra 2020) of plant cells is one of the major responses to abiotic stress, including excess salt. Signalling is divided into three stages. First, according to the information received, the cell recognises extracellular signalling molecules. When a signalling molecule binds to a receptor, the signalling mechanism modifies the receptor protein and the response when the signal elicits a specific cellular response. Sensors’ perception of the signal is the first step in the signalling pathway, and the generation of secondary signalling molecules follows it. It is usually a non-protein molecule such as Ca2 +, IP (Inositol phosphate), or ROS (reactive oxygen species). Each secondary signalling molecule can initiate a protein phosphorylation cascade that regulates the activity of specific transcription factors. Given the abundance of stress at the cellular level and its individual effects, cells develop a sensor that can detect stress (cold, drought, salt) and selectively facilitate the transmission of specific stress signals to cellular targets. A typical two-component system includes HIK (Histidine kinase), which is located on the membrane and senses the input signal, and Rreg (response regulator), which mediates the output signal (Sahoo and Mohapatra 2020).
13.2.1 ROS and Calcium
ROS are formed as a by-product of respiration and photosynthesis, and they are required for average plant growth. Single-electron transfer converts the O2 molecule to superoxide anions, then reduced to hydroxyl radicals by H2O2. These three molecules are reactive oxygen species (ROS), which cause oxidative damage to proteins, DNA, and lipids. Drought, low and high temperatures, and other stressors activate ROS, leading to oxidative stress (Sahoo and Mohapatra 2020). Ca2 + is another non-protein molecule that influences how vegetable crops respond to abiotic stress. The interaction of inflow and outflow produces and maintains Ca2+ spikes in the cytosol. Plants become hypersensitive to ionic and cold stress when Ca2 + homeostasis is disrupted due to overexpression of the Ca2 + /H+ antiporter calcium exchanger 1 (CAX1), which loads Ca2 + into the vacuole. ABA and mechanical stress signals mobilise Ca2 + from intracellular storage, and extracellular Ca2 + influx is crucial in cold stress signal transmission (Sahoo and Mohapatra 2020).
13.2.2 Phospholipids, CDPKs and MAPKs
These are critical components of the plasma membrane that can generate signalling molecules. Phosphorylated phosphatidylinositol (PI) is a significant and complex class of signalling precursors. PIP (Phosphatidylinositol phosphate) and PIP2 (Phosphatidylinositol bisphosphate) plant PI isomers complicate signal transduction. PLC (Phospholipase C) catalyses phospholipid degradation, resulting in IP3 (Inositol-1, 4, 5-triphosphate) and DAG (Diacyl glycerol). Individually, these serve as second messengers. IP3 levels in the cell have a direct impact on ABA-induced gene expression (Sahoo and Mohapatra 2020). Plants that overexpress the enzyme involved in IP3 catabolism delay ABA induction of KIN1, RD29A, and RD22. Calcium-dependent protein kinases (CDPKs) are among the most numerous plant protein kinase subfamilies. It has a Ca2 + sensor kinase hybrid structure, a calmodulin-like domain, and a ser / thr protein kinase domain at the N-terminus (Sahoo and Mohapatra 2020). It is crucial in the signalling of abiotic stress. MAP kinase is a key player in signalling pathways initiated by various surface receptors. The MAPK target can be found in a variety of intracellular compartments. MAPK acts as a physical link between the cytoplasmic and nuclear signalling pathways (Sahoo and Mohapatra 2020).
13.3 Salinity Tolerance in Vegetable Crops
Excessive salt in agricultural land has been a problem since agriculture’s inception. Irrigation has enabled agriculture to be expanded to semi-arid and arid lands and has contributed to a significant increase in food production over the last 40 years, resulting in the accumulation of large amounts of water and high salt. Soil degradation due to increased salinity now affects about 20% of the world’s irrigated areas, instead of arid areas and deserts, which currently occupy 1/4thof the planet’s total land area (Yeo et al. 1999). SOS3, SOS2, and SOS1 were identified as three significant additives that maintain ion homeostasis across salinity stress by molecular and biochemical mechanisms (Yamaguchi and Blumwald 2005; Flowers 2004). The membrane-bound Ca2 + sensor SOS3, which recruits the ser/thr protein kinase SOS2 to the moving membrane, detects Ca2 + signalling induced by the salt stress. SOS3 and SOS2 activate the plasma membrane Na+/H+ antiporter SOS1 and restores mobile ion balance via Na+ ion outflow (Sahoo and Mohapatra 2020). The SOS3/SOS2 complex regulates the Na+ region of the cytosol further by activating and dispersing these ions via the vacuolar transporter NHX1. Access to Na2+ ions is further complicated by the recreational inhibition of the plasma membrane transporter HKT1. SOS2 also regulates the Ca2+ ion domain in the cytosol by controlling the vacuolar Ca2+ channel, CAX1. The entire mechanism results in vegetable crops salt tolerance.
13.4 Cold Tolerance in Vegetable Crops
Cold stress, is the most significant biological stress, reducing vegetable crops productivity by affecting plant quality and postharvest lifespan. Plants adapt to such lethal cold stresses by acquiring cold and frost resistance through a process known as acclimation. This cold stress is transmitted via multiple signalling pathways, including ROS, protein kinases, protein phosphatases, ABA, and Ca2+. Moreover, ABA has proven to be the best (Sahoo and Mohapatra 2020). Plants have a cold acclimation mechanism that helps them survive below-freezing temperatures, and this mechanism is induced by the COR (Cold response) gene. COR78 (cold induction) or RD29A (dehydration reaction) are the critical COR gene for studying gene regulation in cold and osmotic stress. The RD29A promoter region contains a 9-base pair sequence known as the dehydration response element (DRE), which response to cold stress. The CBF gene is activated by ICE1 (Inducer of CBF expression). ICE1 is a constitutive transcription factor that binds to the CBF3 promoter and induces its expression when activated by cold stress. CBF binds to the CRT / DRE cis-element of the COR gene and induces gene expression to provide freeze resistance.
13.5 Drought Tolerance in Vegetable Crops
The global climate is changing due to rising temperatures and CO2 levels in the atmosphere. Because of the severe drought, the soil moisture available to the plant is constantly increasing, causing the plants to die prematurely. Stunted growth is a plant‘s first reaction after a drought has been imposed on the crop. ABA is an important plant hormone that aids in responding to various stress signals during drought stress. The use of ABA on plants mimics the effects of stressful conditions. In vegetable crops, ABA-mediated gene expression can be explained by two types of signalling pathways, i.e., ABA-dependent and independent signalling pathways. AREB / ABF and MYC / MYB are the ABA-dependent regulons (Sahoo and Mohapatra 2020). CBF / DREB and NAC / ZFHD are ABA-independent regulons.
13.6 Genomics of Major Vegetable Crops and Identification of Genes for Abiotic Stresses
Plant genomic information research has bolstered the field of life sciences. Whole-genome sequencing can be used to investigate a species’ entire genome and determine gene function. It is a valuable tool for figuring out how plants grow, develop, and differentiate at the molecular level. The advancement of high-throughput sequencing and the reduction of sequencing costs have made genome sequencing projects on various plant species more feasible. Due to their critical economic relevance and high nutrient content, vegetable crops have become an attractive subject for genetic research. Genome studies of numerous vegetable crops have been performed, including wild relatives and current cultivars. These sequenced vegetable crops can be grouped principally into Solanaceae, Cucurbitaceae, Brassicaceae, and other families (Dias and Ortiz 2021). Many major vegetable species in the Solanaceae family, including tomato, pepper, potato, and eggplant, have their genomic sequences published (Fig. 13.1).
List of reference genome information in vegetable crops. (Adapted from Hao et al. 2020); Left side: Crop name with sequence accession; Right side: Genome size in Mega Base pair (Mb)
On the other hand, wild tomatoes have a higher stress tolerance than farmed tomatoes. This discovery suggests that some critical wild tomato genes can be adapted to survive in abiotic stress (Hao et al. 2020; Pradhan et al. 2021). The following are the primary genome-based methodologies for finding genes relevant to vegetable crops. BSAseq is a popular method for resequencing the whole genome. This entails generating a new batch of descendants and comparing the resequence results to the compiled reference genome. This technique relies mainly on the reference genome to swiftly identify potential regions. Multiple candidate genes for significant features, such as in tomato for internode length (Schrager-Lavelle et al. 2019), Grey press (Chen et al. 2017a, b) and Carcass count (Gao et al. 2016a, b; Yu et al. 2021) in cucumber and, fruit shape in watermelon (Dou et al. 2018b) etc., by using this method. Some other genes are also identified by using this method in vegetables which can be useful during the phenotypic selection of genotypes in abiotic stresses (Table 13.1).
Other BSAseq-derived approaches, such as the QTLseq quantitative trait and the ethyl methane sulfonate (EMS) mutant MutMap, have been used to develop abiotic stress tolerance vegetable crops. QTLseq has been frequently utilised to find genes that control the quantitative features of vegetable crops since publishing reference genomes for numerous vegetable crops. This method has successfully identified quantitative features that suggest continuous phenotypic changes in different offspring of vegetable crops. It was also used to find the linkage marker Ef2.1 (Shu et al. 2018) linked to broccoli and cabbage blooms for tolerance to abiotic stress.
Significant QTL linked to cucumber early flowering during the abiotic stresses, particularly drought, has also been identified (Lu et al. 2014). This makes it easier to choose early-blooming cucumber types and shortens the cucumber growing cycle. Fruit weight and quantity of tomato loci (Illa-Berenguer et al. 2015), as well as several genes for other qualities such as ornamental cabbage locus leaves (Illa-Berenguer et al. 2015), has also been identified. In addition, QTLseq identified watermelon seed coat colour, cucumber sub-gynecology, and cucumber sub-gynecology (Win et al. 2019). The relevance of the reference genome for rapid molecular breeding is further demonstrated by the identification of candidate genes from carcinogenic EMS vegetable variations. Several pooled sequencing approaches have also been established (Abe et al. 2012), and MutMap has been proposed as a standard gene identification approach based on NGS technology. Individuals with mutant traits in the isolated F2 group can pool their DNA. Two light green shell genes (Lun et al. 2015; Zhou et al. 2015) are found in cucumber by using MutMap technique (Hao et al. 2018), and leaf-altered genes that control cucumber fruit or leaf colour (Cao et al. 2018) has also been identified. MutMap is also used to find genes for Chinese cabbage fruits, golden leaves, and yellow tomato fruits (Fu et al. 2019). MutMap also successfully identifies random genes for plant height and leaf morphological variants.
To construct a high-density linkage map and uncover candidate genes for the Rfd1 gene, researchers used the BSRseq approach. BSAseq and BSRseq were also employed on Bell peppers at the same time to develop chloroplast and immature Bell pepper colours (Garcia et al. 2016). SLAFseq is another method for simplifying the genome and performing whole-genome resequencing. For SLAFseq applications to succeed, it is also necessary to have a known reference genome and bioinformatics foundation. SLAFseq may analyse a species’ genetic diversity by simplifying its genome, and bioinformatics opens up new paths for investigating genomic alterations in evolutionary species (Xu et al. 2014). Cucumbers, peppers, and watermelons can all benefit from SLAFseq to help them figure out their complex features against abiotic stresses. For example, genes associated with cucumber fruit thickness, aphid resistance, and flooding were discovered using SLAFseq (Liang et al. 2016; Xu et al. 2015a, b; Zhu et al. 2016). It was also utilised to create a high-density map with 2634 watermelon SNPs and significantly lower distances between linked markers (Shang et al. 2016). Furthermore, SLAFseq, in conjunction with other genome-based sequencing technologies, can produce accurate and modest linkage mapping. SLAFseq and BSAseq, for example, have been used to define genes linked to pepper’s first floral segment (Xu et al. 2016) and the melon flavour gene (Zhang et al. 2016; Abe et al. 2012). These finding suggests that, a combination of BSAseq and RNAseq is required, including a reference genome to identify candidate genes for abiotic stress tolerance in vegetable crops (Table 13.1).
13.7 Genome-Wide Association Analysis in Major Vegetable Crops for Abiotic Stresses
Genome-wide association analysis (GWAS) is an effective method based on genotype-phenotype correlation in a group of individuals, which looks upon genetic variation of complex traits in vegetable crops in the reference genome sequence (Saidou et al. 2014). This can be done by collecting a variety of cultural accessions, crossing lines, sequencing, and comparing genetic diversity between them, especially for traits influenced by various factors. For example, bitterness in cucumbers is an undesirable feature, hence cultivars that are not bitter after long periods of domestication were chosen. Critical GWAS studies on cucumbers have also aided in understanding the genetic mechanism of bittern production using 115 cucumber varieties and the rapid and precise detection of bitterness using linked molecular markers (Shang et al. 2014), particularly during the employment of abiotic stress.
GWAS can also be used to uncover links between fructose and volatile organic acids, which are prevalent in tomato fruit flavours during drought and salinity stress (Zhang et al. 2015; Zhao et al. 2019). Capsaicinoid content is a crucial component of pepper flavour, and GWAS has been used in several studies to explore capsaicinoids. Many transport factors and ankyrin-like proteins may be involved in capsaicinoid synthesis, and the C. annum accession has been used to find markers connected to capsaicin and dihydrocapsaicin levels (Nimmakayala et al. 2016). Another GWAS sample, including the C. annuum accession, discovered candidate genes or QTLs that modulate capsaicinoid concentration. High-density SNP mapping identified 69 QTL regions and 5 candidate genes (Han et al. 2018) during abiotic stresses. GWAS, on the other hand, has several drawbacks. Some GWAS markers produce false-positive results, and others are not repeatable among cultivars. As a result, GWAS-identified molecular markers in vegetable crops will need to be confirmed through genotyping or polymorphism amplification while studying them in the abiotic stress environment.
13.8 Genome Based Molecular Marker Discovery in Vegetable Breeding
Marker-assisted selection (MAS) has become a popular genotype-based breeding approach for increasing vegetable crops breeding performance. MAS can pre-emptively eliminate undesirable genetic backgrounds or nudge alleles back to the intended genetic region (Fig. 13.2). A sufficient number of molecular markers linked with the trait and their polymorphisms are also essential considerations. Molecular markers integrating RLFPs or SSRs were previously advanced without genomic information. On the other hand, those molecular markers are far less polymorphic and have a narrow range of expression.
With the introduction of the reference genome for maximal vegetable vegetation and the advancement of the NGS era for genome sequencing and resequencing, the number of SSR markers has increased dramatically. For the black spikes of ripe cucumber fruit and the uniform colour of orange and immature fruit, near-range SSR markers were mapped (Li et al. 2013; Yang et al. 2014). In addition, in Brassica rapa L. var. purpurea, a high-density map using SSR markers was employed to identify the markers linked to traits related to abiotic stresses (Wang et al. 2017). In tomatoes, OVATE has been discovered to be a good indicator for fruit shape, as well as CYC B, SlMYB12, and SlMYB75 for fruit colour (Rodríguez et al. 2013; Ballester et al. 2010; Fernandez-Moreno et al. 2016; Hwang et al. 2016; Jian et al. 2019). For cucumbers, a novel targeted SSRseq approach has been devised (Jian et al. 2019). It also has higher coverage in high-throughput sequences and is less expensive than previously advanced SSR markers, allowing genotyping of many SSR markers across different accessions. This procedure can also broaden markers so that distinct cucumber ecotypes can be distinguished. SNPs, and SSR markers are important markers for vegetable plants because of the vast range and frequency of genomic modifications (Hwang et al. 2016).
As a result, SNP markers can be used to accurately genotype vegetables and plants and create accurate genetic maps for abiotic stresses (Nimmakayala et al. 2014).The molecular linkage map of tomato was also constructed in the presence of an excessive density map with more than one molecular marker (SSR, and SNP) from the preliminary 2 cM distance among the markers (Tanksley et al. 1992). As a result of the disclosure of the complete genome collection of several pepper cultivars, the genetic map has advanced dramatically (Zhang et al. 2019b). There are now 5569 SNPs in those saturation maps (Zhang et al. 2019b) available for different abiotic stresses. The essential qualities of Chinese cabbage are also linked to some molecular indications. For example, in Brassica plant breeding, flower colour is a flag function that boosts production by attracting pollinating insects. Zhang et al. (2019a) discovered markers in Chinese cabbage orange flowers and followed them through marker assisted breeding to discern different flower colour styles. These comprehensive genetic maps gave markers for inclinations in vegetable crops and increased MAS breeding for abiotic stresses. Table 13.2 summarises the genes or QTLs associated with abiotic stress tolerance in major vegetable crops.
13.9 Application of Transgenic and Gene-Editing Technology in Vegetable Breeding
Physical mutagenesis, EMS mutagenesis, and TDNA insertion can all produce a large number of mutagens, but the mechanisms behind them are yet understood (Chaudhary et al. 2019). Finding the correct strain takes a long time and many individuals. A genome-based gene editing approach for vegetable crops is currently being used to knock off-target genes in vegetable crops. Because of their stable gene transfer method and rich genomic information, tomatoes are an appealing food crop for proving the efficacy of gene editing. Tomato CRISPR has knocked out many genes, including regulation of the drought-tolerant gene SlMAPK3 (Wang et al. 2017). PROCERA is a gene that responds to gibberellin (Tomlinson et al. 2019) in significant vegetable crops during abiotic stresses (Soyk et al. 2017). Future studies on the domestication of wild tomatoes will also employ genome editing techniques. Wildtype tomatoes with good resistance properties can be genetically modified (Li et al. 2018; Zsögön et al. 2018). The cucumber is a model plant for squash, and its solitary nature makes reproduction difficult. As a result, selecting a female cucumber variety as a breeding target can increase cucumber yields while saving time and money. According to Hu (2017), all-female cucumber materials are conserved by boosting the cucumber genetic transformation mechanism and knocking down CsWIP, which regulates the whole female phenotype during multiple stresses. This method was also used to knock out the cucumber-specific short fruit genes sf1 and sf2 and provide the framework for molecular fruit development regulation (Xin et al. 2019; Zhang et al. 2019a). The CRISPR technique can benefit other Cucurbitaceae vegetable crops (Tian et al. 2018). For example, changing the watermelon genome revealed ClWIP1’s genetic role, and female plants were kept to promote hybrid plant purity (Zhang et al. 2019a). CRISPR technology has also helped salt-tolerant squash varieties (Huang et al. 2019).
13.10 Conclusion
With the completion of the genetic sequencing of major vegetable crops, genomic information has significantly accelerated molecular breeding. By utilizing genetic data, genomic sequencing, and genotyping, several new identification approaches have been developed and applied to the majority of vegetable crops in order to identify major genes tolerance to abiotic stresses. For marker assisted breeding, several molecular markers that are directly related to essential properties of vegetable crops and linked to abiotic stress tolerance genes have also been established for major vegetable crops. Advances in gene-editing technology have also paved the way for developing new vegetables varieties with improved characters. However, in the case of vegetable farming, the breeding strategy must be improved.
Even though the genetic sequences of the majority of vegetable crops have been identified, many approaches, such as MutMap Plus and MutMap Gap, have yet to be successfully applied to vegetables for identifying genes associated with abiotic stress tolerance in order to combat climate change. These methodologies, which are reliable in other model cultures, should be used to discover target genes and related molecular markers in vegetable crops. Researchers and breeders can improve communication and information exchange by using connected data. Molecular identifiers can thus be effectively transformed into widely used markers in this manner. Superior traits in vegetable crops and cultivar genotypes can also been studied to better understand the molecular process of superior trait development and to provide a theoretical foundation for molecular breeding.
References
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30(2):174–178
Anithakumari AM, Nataraja KN, Visser RG, van der Linden CG (2012) Genetic dissection of drought tolerance and recovery potential by quantitative trait locus mapping of a diploid potato population. Mol Breed 30(3):1413–1429
Ashraf U, Mahmood S, Shahid N, Imran M, Siddique M, Abrar M (2022) Multi-omics approaches for strategic improvements of crops under changing climatic conditions. In: Principles and practices of OMICS and genome editing for crop improvement. Springer, Cham, pp 57–92
Ballester AR, Molthoff J, de Vos R, Hekkert BTL, Orzaez D, Fernandez-Moreno JP, Tripodi P, Grandillo S, Martin C, Heldens J, Ykema M (2010) Biochemical and molecular analysis of pink tomatoes: deregulated expression of the gene encoding transcription factor SlMYB12 leads to pink tomato fruit color. Plant Physiol 152(1):71–84
Borovsky Y, Monsonego N, Mohan V, Shabtai S, Kamara I, Faigenboim A, Hill T, Chen S, Stoffel K, Van Deynze A, Paran I (2019) The zinc-finger transcription factor cc LOL 1 controls chloroplast development and immature pepper fruit color in Capsicum chinense and its function is conserved in tomato. The Plant J 99(1):41–55
Cao W, Du Y, Wang C, Xu L, Wu T (2018) Cscs encoding chorismate synthase is a candidate gene for leaf variegation mutation in cucumber. Breed Sci:18023
Chaudhary J, Deshmukh R, Sonah H (2019) Mutagenesis approaches and their role in crop improvement. Plan Theory 8(11):467
Chen F, Fu B, Pan Y, Zhang C, Wen H, Weng Y, Chen P, Li Y (2017b) Fine-mapping identifies CsGCN5 encoding a histone acetyltransferase as putative candidate gene for tendril-less1 mutation (td-1) in cucumber. TheorAppl Genet 130(8):1549–1558
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu Rev Plant Biol 70:667–697
Chen S, Zhao H, Zou C, Li Y, Chen Y, Wang Z, Jiang Y, Liu A, Zhao P, Wang M, Ahammed GJ (2017a) Combined inoculation with multiple arbuscular mycorrhizal fungi improves growth, nutrient uptake and photosynthesis in cucumber seedlings. Front Microbiol 8:2516
Cheng Q, Wang P, Liu J, Wu L, Zhang Z, Li T, Gao W, Yang W, Sun L, Shen H (2018) Identification of candidate genes underlying genic male-sterile msc-1 locus via genome resequencing in Capsicum annuum L. TheorAppl Genet 131(9):1861–1872
Cruz RV, Harasawa H, Lal M, Wu S, Anokhin Y, Punsalmaa B, Honda Y, Jafari M, Li C, Huu N (2007) Asia climate change 2007: impacts, adaptation and vulnerability. In: Parry ML, Canziani OF, Palutikof JP, van der Linden PJ, Hanson CE (eds) Contribution of working group ii to the fourth assessment report of the intergovernmental panel on climate change. Cambridge University Press, Cambridge, pp 469–506
Dias JS, Ortiz R (2021) New strategies and approaches for improving vegetable cultivars. In: The basics of human civilization. CRC Press, pp 349–381
Dou J, Lu X, Ali A, Zhao S, Zhang L, He N, Liu W (2018b) Genetic mapping reveals a marker for yellow skin in watermelon (Citrulluslanatus L.). PLoS One 13(9):e0200617
Dou J, Zhao S, Lu X, He N, Zhang L, Ali A, Kuang H, Liu W (2018a) Genetic mapping reveals a candidate gene (ClFS1) for fruit shape in watermelon (Citrulluslanatus L.). TheorAppl Genet 131(4):947–958
Fernandez-Moreno JP, Tzfadia O, Forment J, Presa S, Rogachev I, Meir S, Orzaez D, Aharoni A, Granell A (2016) Characterization of a new pink-fruited tomato mutant results in the identification of a null allele of the SlMYB12 transcription factor. Plant Physiol 171(3):1821–1836
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55(396):307–319
Fu W, Ye X, Ren J, Li Q, Du J, Hou A, Mei F, Feng H, Liu Z (2019) Fine mapping of lcm1, a gene conferring chlorophyll-deficient golden leaf in Chinese cabbage (Brassica rapa ssp. pekinensis). Mol Breed 39(4):1–12
Gao M, Hu L, Li Y, Weng Y (2016a) The chlorophyll-deficient golden leaf mutation in cucumber is due to a single nucleotide substitution in CsChlI for magnesium chelatase I subunit. TheorAppl Genet 129(10):1961–1197
Gao M, Hu L, Li Y, Weng Y (2016b) The chlorophyll-deficient golden leaf mutation in cucumber is due to a single nucleotide substitution in CsChlI for magnesium chelatase I subunit. TheorAppl Genet 129(10):1961–1973
Garcia V, Bres C, Just D, Fernandez L, Tai FWJ, Mauxion JP, Le Paslier MC, Berard A, Brunel D, Aoki K, Alseekh S (2016) Rapid identification of causal mutations in tomato EMS populations via mapping-by-sequencing. Nat Protoc 11(12):2401–2418
Han K, Lee HY, Ro NY, Hur OS, Lee JH, Kwon JK, Kang BC (2018) QTL mapping and GWAS reveal candidate genes controlling capsaicinoid content in capsicum. Plant Biotechnol J 16(9):1546–1558
Hao N, Du Y, Li H, Wang C, Wang C, Gong S, Zhou S, Wu T (2018) CsMYB36 is involved in the formation of yellow green peel in cucumber (Cucumissativus L.). TheorAppl Genet 131(8):1659–1669
Hao N, Han D, Huang K, Du Y, Yang J, Zhang J, Wen C, Wu T (2020) Genome-based breeding approaches in major vegetable crops. TheorAppl Genet 133(5):1739–1752
Hsieh TH, Li CW, Su RC, Cheng CP, Tsai YC, Chan MT (2010) A tomato bZIP transcription factor, SlAREB, is involved in water deficit and salt stress response. Planta 231(6):1459–1473
Hu B (2017) Engineering non-transgenic gynoecious cucumber using an improved transformation protocol and optimized CRISPR/Cas9 system. Mol Plant 10(2):1575–1578
Huang Y, Cao H, Yang L, Chen C, Shabala L, Xiong M, Niu M, Liu J, Zheng Z, Zhou L, Peng Z (2019) Tissue-specific respiratory burst oxidase homolog-dependent H2O2 signaling to the plasma membrane H+-ATPase confers potassium uptake and salinity tolerance in Cucurbitaceae. J Exp Bot 70(20):5879–5893
Hwang I, Kim Y, Han J, Nou IS (2016) Orange color is associated with CYC-B expression in tomato fleshy fruit. Mol Breed 36(4):42
Illa-Berenguer E, Van Houten J, Huang Z, van der Knaap E (2015) Rapid and reliable identification of tomato fruit weight and locule number loci by QTL-seq. TheorAppl Genet 128(7):1329–1342
Jian W, Cao H, Yuan S, Liu Y, Lu J, Lu W, Li N, Wang J, Zou J, Tang N, Xu C (2019) SlMYB75, an MYB-type transcription factor, promotes anthocyanin accumulation and enhances volatile aroma production in tomato fruits. Hort Res 6(1):1–15
Jiang M, Miao L, Zhang H, Zhu X (2020) Over-expression of a transcription factor gene BoC3H4 enhances salt stress tolerance but reduces Sclerotinia stem rot disease resistance in broccoli. J Plant Growth Regul 39(3):1162–1176
Jiang M, Ye ZH, Zhang HJ, Miao LX (2019) Broccoli plants over-expressing an ERF transcription factor gene BoERF1 facilitates both salt stress and sclerotinia stem rot resistance. J Plant Growth Regul 38(1):1–13
Khan MZ, Zaidi SSEA, Amin I, Mansoor S (2019) A CRISPR way for fast-forward crop domestication. Trends Plant Sci 24(4):293–296
Khar A, Singh H, Verma P (2022) Mitigating abiotic stresses in under changing climatic scenario. In: Genomic designing for abiotic stress resistant vegetable crops. Springer, Cham, pp 253–278
Lee YP, Cho Y, Kim S (2014) A high-resolution linkage map of the Rfd1, a restorer-of-fertility locus for cytoplasmic male sterility in radish (Raphanussativus L.) produced by a combination of bulked segregant analysis and RNA-Seq. TheorAppl Genet 127(10):2243–2252
Li S, Pan Y, Wen C, Li Y, Liu X, Zhang X, Behera TK, Xing G, Weng Y (2016) Integrated analysis in bi-parental and natural populations reveals CsCLAVATA3 (CsCLV3) underlying carpel number variations in cucumber. TheorAppl Genet 129(5):1007–1022
Li S, Zhang L, Wang Y, Xu F, Liu M, Lin P, Ren S, Ma R, Guo YD (2017) Knockdown of a cellulose synthase gene BoiCesA affects the leaf anatomy, cellulose content and salt tolerance in broccoli. Sci Rep 7(1):1–14
Li T, Yang X, Yu Y, Si X, Zhai X, Zhang H, Dong W, Gao C, Xu C (2018) Domestication of wild tomato is accelerated by genome editing. Nat Biotechnol 36(12):1160–1163
Li Y, Wen C, Weng Y (2013) Fine mapping of the pleiotropic locus B for black spine and orange mature fruit color in cucumber identifies a 50 kb region containing a R2R3-MYB transcription factor. TheorAppl Genet 126(8):2187–2196
Liang D, Chen M, Qi X, Xu Q, Zhou F, Chen X (2016) QTL mapping by SLAF-seq and expression analysis of candidate genes for aphid resistance in cucumber. Front Plant Sci 7:1000
Liu C, Chen L, Zhao R, Li R, Zhang S, Yu W, Sheng J, Shen L (2019a) Melatonin induces disease resistance to Botrytis cinerea in tomato fruit by activating jasmonic acid signaling pathway. J Agric Food Chem 67(22):6116–6124
Liu G, Mao S, Kim JH (2019b) A mature-tomato detection algorithm using machine learning and color analysis. Sensors 19(9):2023
Lu H, Lin T, Klein J, Wang S, Qi J, Zhou Q, Sun J, Zhang Z, Weng Y, Huang S (2014) QTL-seq identifies an early flowering QTL located near flowering locus T in cucumber. Theor Appl Genet 127(7):1491–1499
Lun Y, Wang X, Zhang C, Yang L, Gao D, Chen H, Huang S (2015) A CsYcf54 variant conferring light green coloration in cucumber. Euphytica 208(3):509–517
Maligeppagol M, Manjula R, Navale PM, Babu KP, Kumbar BM, Laxman RH (2016) Genetic transformation of chilli (Capsicum annuum L.) with Dreb1A transcription factor known to impart drought tolerance
Nimmakayala P, Abburi VL, Saminathan T, Alaparthi SB, Almeida A, Davenport B, Nadimi M, Davidson J, Tonapi K, Yadav L, Malkaram S (2016) Genome-wide diversity and association mapping for capsaicinoids and fruit weight in Capsicum annuum L. Sci Rep 6(1):1–14
Nimmakayala P, Levi A, Abburi L, Abburi VL, Tomason YR, Saminathan T, Vajja VG, Malkaram S, Reddy R, Wehner TC, Mitchell SE (2014) Single nucleotide polymorphisms generated by genotyping by sequencing to characterize genome-wide diversity, linkage disequilibrium, and selective sweeps in cultivated watermelon. BMC Genomics 15(1):1–15
Paudel L, Clevenger J, McGregor C (2019) Chromosomal locations and interactions of four loci associated with seed coat color in watermelon. Front Plant Sci 10:788
Pradhan K, Rout S, Tripathy B, Mishra UN, Sahoo G, Prusty AK, Dash L (2021) Role of biotechnology in vegetable breeding. Turkish Online J Qual Inq 12(3):5092–5102
Ren J, Liu Z, Du J, Fu W, Hou A, Feng H (2019) Fine-mapping of a gene for the lobed leaf, BoLl, in ornamental kale (Brassica oleracea L. var. acephala). Mol Breed 39(3):1–14
Rodríguez GR, Kim HJ, Van Der Knaap E (2013) Mapping of two suppressors of OVATE (sov) loci in tomato. Heredity 111(3):256–264
Rong F, Chen F, Huang L, Zhang J, Zhang C, Hou D, Cheng Z, Weng Y, Chen P, Li Y (2019) A mutation in class III homeodomain-leucine zipper (HD-ZIP III) transcription factor results in curly leaf (cul) in cucumber (Cucumissativus L.). TheorAppl Genet 132(1):113–123
Roy R, Purty RS, Agrawal V, Gupta SC (2006) Transformation of tomato cultivar ‘PusaRuby’withbspA gene from Populustremula for drought tolerance. Plant Cell Tissue Organ Cult 84(1):56–68
Ruangrak E, Su X, Huang Z, Wang X, Guo Y, Du Y, Gao J (2018) Fine mapping of a major QTL controlling early flowering in tomato using QTL-seq. Can J Plant Sci 98(3):672–682
Saha P, Singh S, Bhatia R, Dey SS, Das Saha N, Ghoshal C et al (2022) Genomic designing for abiotic stress resistant brassica vegetable crops. In: Genomic designing for abiotic stress resistant vegetable crops. Springer, Cham, pp 153–185
Sahoo JP, Mohapatra U (2020) Plant signal transduction mechanism against abiotic and biotic stresses. https://doi.org/10.22271/ed.book.1027
Saidou AA, Clotault J, Couderc M, Mariac C, Devos KM, Thuillet AC, Amoukou IA, Vigouroux Y (2014) Association mapping, patterns of linkage disequilibrium and selection in the vicinity of the PHYTOCHROME C gene in pearl millet. TheorAppl Genet 127(1):19–32
Satish Chhapekar S, Singh S, Singh S, Ma Y, Jeevan Rameneni J, Ryun Choi S, Pyo Lim Y (2022) Genomic Design for Biotic Stress Tolerance in vegetable brassicas. In: Genomic designing for biotic stress resistant vegetable crops. Springer, Cham, pp 189–231
Schrager-Lavelle A, Gath NN, Devisetty UK, Carrera E, López-Díaz I, Blázquez MA, Maloof JN (2019) The role of a class III gibberellin 2-oxidase in tomato internode elongation. The Plant J 97(3):603–615
Shang J, Li N, Li N, Xu Y, Ma S, Wang J (2016) Construction of a high-density genetic map for watermelon (Citrullus lanatus L.) based on large-scale SNP discovery by specific length amplified fragment sequencing (SLAF-seq). Sci Hortic 203:38–46
Shang Y, Ma Y, Zhou Y, Zhang H, Duan L, Chen H, Zeng J, Zhou Q, Wang S, Gu W, Liu M (2014) Biosynthesis, regulation, and domestication of bitterness in cucumber. Sci 346(6213):1084–1088
Shu J, Liu Y, Zhang L, Li Z, Fang Z, Yang L, Zhuang M, Zhang Y, Lv H (2018) QTL-seq for rapid identification of candidate genes for flowering time in broccoli × cabbage. Theor Appl Genet 131(4):917–928
Solankey SS, Singh RK, Baranwal DK, Singh DK (2015) Genetic expression of tomato for heat and drought stress tolerance: an overview. Int J Veg Sci 21(5):496–515
Soyk S, Lemmon ZH, Oved M, Fisher J, Liberatore KL, Park SJ et al (2017) Bypassing negative epistasis on yield in tomato imposed by a domestication gene. Cell 169(6):1142–1155
Spaldon S, Samnotra RK, Chopra S (2015) Climate resilient technologies to meet the challenges in vegetable production. Int J Curr Res Aca Rev 3(2):28–47
Subramanyam K, Sailaja KV, Subramanyam K, Rao DM, Lakshmidevi K (2011) Ectopic expression of an osmotin gene leads to enhanced salt tolerance in transgenic chilli pepper (Capsicum annum L.). Plant Cell Tissue Organ Cult 105(2):181–192
Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S, Innan H, Cano LM, Kamoun S, Terauchi R (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74(1):174–183
Tanksley SD, Ganal MW, Prince JP, De Vicente MC, Bonierbale MW, Broun P, Fulton TM, Giovannoni JJ, Grandillo S, Martin GB (1992) High density molecular linkage maps of the tomato and potato genomes. Genet 132(4):1141–1160
Tian S, Jiang L, Cui X, Zhang J, Guo S, Li M, Zhang H, Ren Y, Gong G, Zong M, Liu F (2018) Engineering herbicide-resistant watermelon variety through CRISPR/Cas9-mediated base-editing. Plant Cell Rep 37(9):1353–1356
Tomlinson L, Yang Y, Emenecker R, Smoker M, Taylor J, Perkins S, Smith J, MacLean D, Olszewski NE, Jones JD (2019) Using CRISPR/Cas9 genome editing in tomato to create a gibberellin-responsive dominant dwarf DELLA allele. Plant Biotechnol J 17(1):132–140
Wang L, Chen L, Li R, Zhao R, Yang M, Sheng J, Shen L (2017) Reduced drought tolerance by CRISPR/Cas9-mediated SlMAPK3 mutagenesis in tomato plants. J Agric Food Chem 65(39):8674–8682
Wang Y, Cai S, Yin L, Shi K, Xia X, Zhou Y, Yu J, Zhou J (2015) Tomato HsfA1a plays a critical role in plant drought tolerance by activating ATG genes and inducing autophagy. Autophagy 11(11):2033–2047
Win KT, Zhang C, Silva RR, Lee JH, Kim YC, Lee S (2019) Identification of quantitative trait loci governing subgynoecy in cucumber. TheorAppl Genet 132(5):1505–1521
Xin T, Zhang Z, Li S, Zhang S, Li Q, Zhang ZH, Huang S, Yang X (2019) Genetic regulation of ethylene dosage for cucumber fruit elongation. Plant Cell 31(5):1063–1076
Xu H, Liu Q, Yao TFX (2014) Shedding light on integrative GA signaling. CurrOpin Plant Biol 21:89–95
Xu L, Li T, Wu Z, Feng H, Yu M, Zhang X, Chen B (2018) Arbuscular mycorrhiza enhances drought tolerance of tomato plants by regulating the 14-3-3 genes in the ABA signaling pathway. Appl Soil Ecol 125:213–221
Xu X, Chao J, Cheng X, Wang R, Sun B, Wang H, Luo S, Xu X, Wu T, Li Y (2016) Mapping of a novel race specific resistance gene to phytophthora root rot of pepper (Capsicum annuum) using bulked segregant analysis combined with specific length amplified fragment sequencing strategy. PLoS One 11(3):e0151401
Xu X, Lu L, Zhu B, Xu Q, Qi X, Chen X (2015a) QTL mapping of cucumber fruit flesh thickness by SLAF-seq. Sci Rep 5(1):1–9
Xu X, Xu R, Zhu B, Yu T, Qu W, Lu L, Xu Q, Qi X, Chen X (2015b) A high-density genetic map of cucumber derived from specific length amplified fragment sequencing (SLAF-seq). Front Plant Sci 5:768
Yamaguchi T, Blumwald E (2005) Developing salt-tolerant crop plants: challenges and opportunities. Trends Plant Sci 10(12):615–620
Yan C, An G, Zhu T, Zhang W, Zhang L, Peng L, Chen J, Kuang H (2019) Independent activation of the BoMYB2 gene leading to purple traits in Brassica oleracea. TheorAppl Genet 132(4):895–906
Yang L, Liu H, Zhao J, Pan Y, Cheng S, Lietzow CD, Wen C, Zhang X, Weng Y (2018) LITTLELEAF (LL) encodes a WD40 repeat domain-containing protein associated with organ size variation in cucumber. Plant J 95(5):834–847
Yang X, Li Y, Zhang W, He H, Pan J, Cai R (2014) Fine mapping of the uniform immature fruit color gene u in cucumber (Cucumis sativus L.). Euphytica 196(3):341–348
Yeo AR, Flowers SA, Rao G, Welfare K, Senanayake N, Flowers TJ (1999) Silicon reduces sodium uptake in rice (Oryza sativa L.) in saline conditions and this is accounted for by a reduction in the transpirational bypass flow. Plant Cell Environ 22(5):559–565
Yu D, Gu X, Zhang S, Dong S, Miao H, Gebretsadik K, Bo K (2021) Molecular basis of heterosis and related breeding strategies reveal its importance in vegetable breeding. Horticul Res 8(1):120–120
Zahid G, Iftikhar S, Farooq MU, Soomro SA (2022) Advances in DNA based molecular markers for the improvement of fruit cultivars in Pakistan-a review. Sarhad J Agricult 38(3):812–832
Zhang H, Yi H, Wu M, Zhang Y, Zhang X, Li M, Wang G (2016) Mapping the flavor contributing traits on" Fengwei melon" (Cucumis melo L.) chromosomes using parent resequencing and super bulked-segregant analysis. PLoS One 11(2):e0148150
Zhang J, Zhao J, Xu Y, Liang J, Chang P, Yan F, Li M, Liang Y, Zou Z (2015) Genome-wide association mapping for tomato volatiles positively contributing to tomato flavor. Front Plant Sci 6:1042
Zhang N, Zhang H, Ren Y, Chen L, Zhang J, Zhang L (2019a) Genetic analysis and gene mapping of the orange flower trait in Chinese cabbage (Brassica rapa L.). Mol Breed 39(6):1–11
Zhang X, Zou Z, Gong P, Zhang J, Ziaf K, Li H, Xiao F, Ye Z (2011) Over-expression of microRNA169 confers enhanced drought tolerance to tomato. Biotechnol Lett 33(2):403–409
Zhang XF, Wang GY, Dong TT, Chen B, Du HS, Li CB, Geng SS (2019b) High-density genetic map construction and QTL mapping of first flower node in pepper (Capsicum annuum L.). BMC Plant Biol 19(1):1–13
Zhao J, Sauvage C, Zhao J, Bitton F, Bauchet G, Liu D, Huang S, Tieman DM, Klee HJ, Causse M (2019) Meta-analysis of genome-wide association studies provides insights into genetic control of tomato flavor. Nat Commun 10(1):1–12
Zhao T, Wu T, Pei T, Wang Z, Yang H, Jiang J, Zhang H, Chen X, Li J, Xu X (2021) Overexpression of SlGATA17 promotes drought tolerance in transgenic tomato plants by enhancing activation of the Phenylpropanoid biosynthetic pathway. Front Plant Sci 12
Zhou Q, Wang S, Hu B, Chen H, Zhang Z, Huang S (2015) An accumulation and replication of chloroplasts 5 gene mutation confers light green peel in cucumber. J Integr Plant Biol 57(11):936–942
Zhu M, Chen G, Zhang J, Zhang Y, Xie Q, Zhao Z, Pan Y, Hu Z (2014) The abiotic stress-responsive NAC-type transcription factor SlNAC4 regulates salt and drought tolerance and stress-related genes in tomato (Solanum lycopersicum). Plant Cell Rep 33(11):1851–1863
Zhu WY, Huang L, Chen L, Yang JT, Wu JN, Qu ML, Yao DQ, Guo CL, Lian HL, He HL, Pan JS (2016) A high-density genetic linkage map for cucumber (Cucumis sativus L.): based on specific length amplified fragment (SLAF) sequencing and QTL analysis of fruit traits in cucumber. Front Plant Sci 7:437
Zsögön A, Čermák T, Naves ER, Notini MM, Edel KH, Weinl S, Freschi L, Voytas DF, Kudla J, Peres LEP (2018) De novo domestication of wild tomato using genome editing. Nat Biotechnol 36(12):1211–1216
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Sahoo, J.P., Barik, S., Pathak, M., Tripathy, B., Pradhan, M. (2023). Improvement of Vegetables Through Molecular Breeding in Changing Climate Scenario. In: Solankey, S.S., Kumari, M. (eds) Advances in Research on Vegetable Production Under a Changing Climate Vol. 2. Advances in Olericulture. Springer, Cham. https://doi.org/10.1007/978-3-031-20840-9_13
Download citation
DOI: https://doi.org/10.1007/978-3-031-20840-9_13
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-20839-3
Online ISBN: 978-3-031-20840-9
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)