Abstract
Key message
Linkage analysis confirmed the association in the region of PHYC in pearl millet. The comparison of genes found in this region suggests that PHYC is the best candidate.
Abstract
Major efforts are currently underway to dissect the phenotype–genotype relationship in plants and animals using existing populations. This method exploits historical recombinations accumulated in these populations. However, linkage disequilibrium sometimes extends over a relatively long distance, particularly in genomic regions containing polymorphisms that have been targets for selection. In this case, many genes in the region could be statistically associated with the trait shaped by the selected polymorphism. Statistical analyses could help in identifying the best candidate genes into such a region where an association is found. In a previous study, we proposed that a fragment of the PHYTOCHROME C gene (PHYC) is associated with flowering time and morphological variations in pearl millet. In the present study, we first performed linkage analyses using three pearl millet F2 families to confirm the presence of a QTL in the vicinity of PHYC. We then analyzed a wider genomic region of ~100 kb around PHYC to pinpoint the gene that best explains the association with the trait in this region. A panel of 90 pearl millet inbred lines was used to assess the association. We used a Markov chain Monte Carlo approach to compare 75 markers distributed along this 100-kb region. We found the best candidate markers on the PHYC gene. Signatures of selection in this region were assessed in an independent data set and pointed to the same gene. These results foster confidence in the likely role of PHYC in phenotypic variation and encourage the development of functional studies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In recent years, association mapping methods have been developed to exploit collections of existing populations for the analysis of the genotype-phenotype relationship (Rafalski 2010; Bergelson and Roux 2010; Myles et al. 2009; Zhu et al. 2008). One advantage of these diverse collections is the presence of recombination events that have accumulated within populations over long periods of time. Such historical recombination provides a mapping resolution that sometimes extends only a few kb (Yu and Buckler 2006). Moreover, association mapping can be used to validate the functional role of polymorphisms associated with selection, bridging the gap between the identification of a molecular selection signature and the identification of the traits shaped by selected polymorphisms (Mariac et al. 2011).
Association studies mainly fall into two groups: candidate gene association studies (CGA, e.g., Camus-Kulandaivelu et al. 2008; Brown et al. 2008) and genome-wide association studies (GWA, e.g., Atwell et al. 2010). The density of markers needed for an efficient GWA study depends on the size of the genome and on the extent of linkage disequilibrium (LD) at the genomic scale. When the LD is relatively extensive, a lower density of markers is sufficient to survey the whole genome. A total of 250,000 SNPs were used for GWA studies in Arabidopsis (Atwell et al. 2010), while 1.6 million SNPs were used in maize (Tian et al. 2011). The candidate gene approach is a more flexible alternative when the LD decreases rapidly (making the need for marker density too high) or for orphan crops with limited genomic information available (e.g., pearl millet).
The control of false positives (type I error) in plant association mapping requires particular attention. The simplest statistical models are often biased because of the confounding effect of a structured background that is shared between individuals (Pritchard et al. 2000). More sophisticated statistical models that account for population structure were developed to limit this bias (Thornsberry et al. 2001; Yu et al. 2006). However, the ability of these models to completely eliminate the bias depends on the characteristics of the data. For example, limiting false positives using mixed models (Yu et al. 2006) was shown to be less efficient for traits that are tightly linked to population differentiation (Saïdou et al. 2009). The use of linkage mapping as a complementary way of confirming associations is one way of overcoming this limitation. Linkage mapping is based on controlled crosses and is thus no longer biased by the effect of population structure. Marker-trait associations highlighted by association mapping can be considered as true positives with the highest confidence when the markers colocalize with QTLs revealed by linkage mapping (Bergelson and Roux 2010).
Association mapping usually identifies genomic regions associated with phenotypes by exploiting the LD between genotyped markers and the unknown causal polymorphisms. Causal polymorphisms shaped by selection often produce extensive LD in their vicinity. This high LD leads to significant statistical associations with a large number of markers in the region (e.g., Ducrocq et al. 2008). Regions showing significant associations need to be examined in great detail to search for the best candidate genes. Depending on the LD, causal polymorphisms may be located far from genotyped markers (Camus-Kulandaivelu et al. 2008). To deal with this issue, larger fragments of the genome surrounding significantly associated markers could be used to analyze the extent of LD and association patterns in order to finally pinpoint the best candidate polymorphisms (Ducrocq et al. 2008; Brown et al. 2008). Such an approach does not replace experimental evidence or the functional validation of the gene effect. However, it could be seen as a step toward the discovery of causal genes, because it allows large numbers of non-causal markers or genes (which are associated with a trait because of the simple effect of LD) to be discarded. Functional studies are generally tedious, so this step is practical and helps focus on the most likely causal genes. Further evidence of selection for these markers would also reinforce confidence in marker-trait associations (Mariac et al. 2011).
In this study, we first used linkage analysis on three F2 families to confirm the association between the genomic region of the PHYTOCHROME C gene (PHYC) and several quantitative traits. We then used an association mapping panel of 90 pearl millet inbreds to examine the pattern of linkage disequilibrium and to assess marker-trait association in this genomic region. In a previous candidate gene study, we examined a shorter fragment of PHYC (866 bp) and observed a significant association with flowering time and morphological variation (Saïdou et al. 2009). Here, we carried out a study that took into account 75 SNPs and insertion-deletion polymorphisms (indels) from a larger genomic region (~100 kb) in the vicinity of PHYC. The full PHYC locus (~6 kb) and fragments of five putative genes surrounding PHYC were sequenced. We assessed the pattern of LD in the region and analyzed marker-trait associations using a mixed model. We then developed a Markov chain Monte Carlo method (MCMC) based on probability and model fit to identify the best candidate loci associated with the phenotype. We also assessed signatures of selection for the five genes surrounding PHYC and six portions of PHYC.
Materials and methods
Linkage association analyses
F2 plant families
F1 plants were derived from three different crosses (Table S1). Parents were selected for having two different alleles at the PHYC gene using a previously developed SNP marker, corresponding to position 5525 at PHYC (Saïdou et al. 2009). The parents of each cross were homozygous plants for a different allele of PHYC. The three crosses included a wild female parent and a cultivated male parent: A7 cross (PE8151 × PE5887), C1 cross (Pe8151 × PE1205) and D1 cross (PE8504 × PE1205). Fifteen microsatellite markers were genotyped on the parents and F1 progenies to identify true F1 hybrids (data not shown). One F1 individual for each cross was selfed to derive the F2 plants used in this study. Two of the 15 microsatellites were used to genotype the F2 plants to confirm self-pollination.
Field trial
The experiment was conducted in the field in Sadoré, Niger. Seeds of F2 plants were sown in pots on 1 June 2009 and transplanted to the field when the young plants reached 4–6 cm in height. For each family, plants were grown in plots of 8 columns and 12 rows with a distance of 1 m between rows and 0.7 m between columns. The trial was conducted under rainfall conditions with supplemental irrigation if necessary. Seven traits were considered: basal primary spike diameter (BSpD), number of days from sowing to heading (FTHE), number of basal tillers at head emergence (NTHE), plant height (PH), stem diameter (SD), primary spike diameter (SpD) and primary spike length (SpL). Each plant was phenotyped individually. The total number of phenotyped F2 plants was 270 for A7, 272 for D1 and 1,182 for C1.
SNP genotyping in PHYC and statistical analysis
DNA was extracted from leaf segments of each individual plant. A restriction enzyme targeting the C/G SNP at position 5525 in PHYC was used to genotype this SNP (protocol described in Saïdou et al. 2009). The genotype was scored CC, GG and CG with respect to the digestion pattern. QTL analysis was performed based on this single marker. We fitted the genotype with each trait using a generalized linear model (R 2.7.2). The significance of the marker effect was assessed using the F test.
Association mapping
Plant material and phenotypic scores
Association analysis was performed using a panel of 90 inbred lines derived from diverse pearl millet material (Saïdou et al. 2009). Seven traits were considered: basal primary spike diameter (BSpD), number of days from sowing to the female flowering stage (FT), number of basal tillers at head emergence (NTHE), plant height (PH), stem diameter (SD), primary spike diameter (SpD) and primary spike length (SpL). Note that the number of days from sowing to the female flowering stage (FT) is highly correlated (R 2 > 0.90) to the number of days from sowing to heading (FTHE) used in the F2 families (FT occurs 2–3 days after FTHE). Field trials and phenotype measurements were described in Saïdou et al. (2009). A total of nine field trials were performed between 2005 and 2008 in Sadoré, Niger. In each trial, each trait was measured in 7–10 plants per inbred line to calculate the average phenotype for each inbred line. The complete design thus consisted of 810 entries per trait, which represents the average inbred scores of the 90 inbreds in the nine trials (except BSpD, which was scored only in five trials).
Sequencing and polymorphism scoring
We identified a single BAC clone containing the PHYC gene from a pearl millet BAC library (Allouis et al. 2001) using primer set PHYC4 (Table S2) and sequenced the clone on a Roche 454 platform (Roche Applied Science) and a PACBIO platform. Sequencing experiments were performed by GATC Biotech (Konstanz, Germany) on 1/8 of a 454 run using a single PACBIO SMRT cell. Based on this BAC sequence, primers were designed to complete sequencing of the entire PHYC gene and to sequence loci in the regions neighboring this gene (Table S2; Fig. 2). The primers were designed to match loci that had good BLASTN hits to predicted or known genes of the whole genome of Sorghum bicolor (BLASTN, near-exact matches, GRAMENE 32, 2010). The corresponding gene fragments were amplified from the entire panel of inbreds and sequenced using the classic Sanger sequencing approach. We also sequenced two additional genes in this panel (hereafter named Pg7840 and Pg7880, corresponding to sorghum genes Sb01g007840 and Sb01g007880). These two genes are located near PHYC in sorghum but were not present on the sequenced pearl millet BAC. Note that while Sb01g007840 is located between Sb01g007830 and PHYC in sorghum, the order appears to be slightly different in pearl millet. The two genes Pg7840 and Pg7880 had previously been shown to cosegregate without recombinants with PHYC in the pearl millet world reference mapping population (81B × ICMP 451) from ICRISAT (India) of 167 F7-RILs individuals (data not shown). All sequences of the 90 inbreds were generated using the standard protocol described in Saïdou et al. (2009). Protocols for DNA extraction and PCR have also been described previously (Mariac et al. 2006; Oumar et al. 2008).
SNP/indel polymorphisms were extracted from the sequence data (Geneious 4.8.5). Entries with ambiguous base calling were checked and, if necessary, designated as missing entries. A third allele present in less than two individuals was found at a few polymorphic sites; these entries were also designated as missing. The finally checked data matrices were then filtered based on a minor allele frequency threshold of 2.5 %.
Association analysis
Marker-trait associations were based on the set of phenotypic data from several trials (up to nine trials) available for the inbred panel. Single markers were fitted to each phenotype using a mixed linear model (see Yu et al. 2006). The mixed linear model considers the effect of population structure as a fixed effect modeled by matrices of ancestry (Q) and uses a matrix of kinship (K) between the inbreds to set random background effects. The panel of 90 inbreds was structured in seven populations (Saïdou et al. 2009) identified using the Bayesian structure model (Pritchard et al. 2000). We used a kinship matrix (K) inferred with AFLP markers using SPAGEDI (Hardy and Vekemans 2002), as previously described (Saïdou et al. 2009; Mariac et al. 2011). Field trial was added as a random factor. The model was fitted using the restricted maximum likelihood method (ASReml-R 2.0/32, 2011). The significance of fixed effects was assessed using an incremental Wald test (Gilmour et al. 2006). We ordered terms in the model so that ancestry fixed effects were adjusted before the fixed effect of the candidate gene marker.
Correlation of p-values across traits
For each pair of traits, the p-values for the marker-trait associations provided by the mixed model analysis were plotted across markers to graphically check which markers were associated with both traits. We also assessed the correlation between the distributions of the p-values using Pearson’s coefficient and Student’s t test. The p-values were transformed as p′ = −log10 (p) before the correlation analyses.
LD analysis and the MCMC approach to identify the best candidate causal SNPs
Measure of linkage disequilibrium
LD between all pairs of polymorphic sites within and among all sequenced genes was calculated as the squared correlation coefficient r 2. The significance of this measure was assessed using a two-sided Fisher exact test. The R package LD tests (Lewin 2008) were used to calculate and test LD, and the R package LD heatmap (Shin 2010) was used to plot the heat map.
MCMC method
To highlight the markers most tightly linked with the trait, we implemented an iterative process of pairwise comparison of markers through the space defined by all 75 available markers. The algorithm was defined as follows:
-
1.
Pick an initial SNP (or indel) at random; this SNP defines the current position and is referred to as SNPcp;
-
2.
Repeat the following steps several times (N iterations);
-
2.1.
Pick a second SNP at random (denoted SNPr);
-
2.2.
Perform association analyses with SNPcp and SNPr by fitting the mixed model to the same panel subset (inbreds with missing entries in either SNP are discarded);
-
2.3.
Compare the log-likelihood associated with SNPcp and SNPr (LSNPcp and LSNPr);
-
2.3.1.
If LSNPr > LSNPcp, SNPr is selected as the more likely causal SNP;
-
2.3.2.
If LSNPcp > LSNPr, the probability to select SNPr is p (SNPr) = 10−ΔL and the probability to select SNPcp is p (SNPcp) = 1 − 10−ΔL; where ΔL = LSNPcp − LSNPr;
-
2.3.1.
-
2.4.
Record the identity of the selected SNP;
-
2.5.
Reset the new current position to the position of the selected SNP and start the next iteration;
-
2.1.
-
3.
Analyze the record of selection over iterations and calculate the frequency corresponding to the choice of each SNP.
Computationally, the probability of 10−ΔL defined in step 2.3.2 was set by comparing ΔL to a random value β sampled from a uniform random distribution in the range 0–1. For each iteration with ΔL ≥ –log10 (β), SNPcp was selected; otherwise, SNPr was selected. Therefore, the probability of selecting SNPcp increased with ΔL, which measures the improvement in fit provided by this marker compared to a randomly selected SNP.
We checked the effect of the total number of iterations and burn-in length on the MCMC result. First, we set the total number of iterations (including burn-in) to 200,000 and varied the length of burn-in from 10,000 to 100,000. Second, we set the burn-in period to 10,000 and varied the total chain length from 20,000 to 200,000 iterations. The selection frequencies obtained for the different markers (selection scores) were compared to the MCMC parameters (Pearson’s correlation and Student’s t test). Markers with a null score for all parameter values were discarded from the correlation test. The analysis for parameter setting was carried out using the FT trait.
Signatures of selection
Plant material, sequencing and diversity analysis
Signatures of selection in the PHYC region were investigated in 33 cultivated pearl millet (Cenchrus americanus syn. Pennisetum glaucum) individuals and 13 wild pearl millet individuals, chosen to cover their respective geographic distribution in West Africa (see Clotault et al. 2012). Ancestral versus derived alleles of detected polymorphisms were defined using two samples of the related Cenchrus polystachion (syn. Pennisetum polystachion) species.
We used previously published data sets of (1) six gene fragments of the PHYC gene and (2) 20 randomly selected genes (Clotault et al. 2012; Table S2). In this study, we sequenced five additional genes around PHYC: Pg7830, Pg7840, Pg7870, Pg7878 and Pg7880 (Table S2). Polymerase chain reactions were performed as described in Saïdou et al. (2009). Sequencing reactions were performed as described in Clotault et al. (2012).
For each segment sequenced, we calculated nucleotide polymorphism θ w (Watterson 1975), nucleotide diversity π (Nei 1987), Tajima’s D (Tajima 1989), and Fay and Wu’s H (Fay and Wu 2000) for both wild and cultivated pearl millet groups using DnaSP 5.10 (Rozas et al. 2003). The differentiation index F ST (Hudson et al. 1992) between wild and cultivated groups was also calculated.
Demographic model
The demographic history of the sample is likely to bias neutrality tests and lead to false negatives and false positives (Vigouroux et al. 2002; Tenaillon et al. 2004). In order to disentangle the effect of demography on genes in the PHYC region, neutrality tests were carried out against a more realistic demographic model than the Wright-Fisher model. The method is described in detail in Clotault et al. (2012). The model used in this study is a model of domestication whereby the cultivated population is derived from the wild population and grows exponentially after domestication; cultivated and wild populations are connected by gene flow (EGGF). It was simulated using msABC software (Pavlidis et al. 2010). Nine summary statistics were used to compare simulated and observed data sets: nucleotide diversity π (Nei 1987), nucleotide polymorphism θ w (Watterson 1975), Tajima’s D (Tajima 1989), ZnS (Kelly 1997) for cultivated and wild samples and F ST (Hudson et al. 1992) between cultivated and wild samples. The rejection-regression procedure described in Beaumont et al. (2002), adapted by a log-tangent transformation of parameters (Hamilton et al. 2005), was used to estimate the posterior distribution of each parameter (Clotault et al. 2012). The rejection-regression step was performed with a threshold of 10−3 using abcreg software (Thornton 2009).
Neutrality tests
For the simulations, Tajima’s D (Tajima 1989) and Fay and Wu’s H (Fay and Wu 2000) were calculated for both cultivated and wild samples and F ST (Hudson et al. 1992) between wild and cultivated samples according to the EGGF model. The different gene fragments were tested for selection according to the rank of observed Tajima’s D and Fay and Wu’s H in their respective expected distribution. The rank of F ST was calculated for simulated data sets sharing a similar θ w per gene ± 0.1. Therefore, the statistical values found at the extremity of the distribution are considered significantly different from what is expected at random (*one-sided test p > 0.95 for FST; *two-sided test p < 0.025 or p > 0.975 for Tajima’s D).
Results
Linkage association analyses
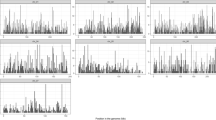
The F2-based study revealed significant associations between the genotype at PHYC and different traits (Fig. 1). Associations of the PHYC polymorphism tested with basal spike diameter (BSpD, A7 F2,234 = 11.1 p < 3 × 10−5; D1 F2,239 = 31.4 p < 8 × 10−13; C1 F2,969 = 103.2 p < 10−40), spike diameter (SpD, A7 F2,234 = 7.9 p < 0.0005; D1 F2,230 = 24.3 p < 3 × 10−10; C1 F2,973 = 112.8 p < 10−44) and spike length (SpL, A7 F2,234 = 12.4 p < 8 × 10−6; D1 F2,239 = 23.3 p < 6 × 10−10; C1 F2,973 = 95.8 p < 10−38) were observed in each cross with a high level of significance. These tests are significant in the three families even after Bonferroni correction (21 tests; p < 0.0024). The PHYC marker was significantly associated with flowering time (FTHE) in the D1 and C1 families (F2,263 = 4.7, p < 0.01; F2,1118, p < 10−20 respectively), but not in the A7 family (F2,260 = 0.29 p = 0.75). A significant association was also found with stem diameter in A7 (F2,260 = 5.31 p < 6 × 10−3) and C1 (F2,1118 = 23.23 p < 10−10), but only the association in C1 remains significant after Bonferroni correction.
Single marker QTL analysis of F2 families. Three F2 families were used for the analysis: A7 (n = 270), D1 (n = 272) and C1 (n = 1182). Histogram showing the effect of the PHYC genotype on each of the seven traits on the x-axis. The p-value (F-test) is given (−log10). The dashed line represents the p-value threshold 0.05. The threshold after Bonferroni correction (21 tests) is 0.0023, corresponding to −log10 (p) = 2.62. Note the change in scale across graphs. See text for the coding of traits
Sequencing
A total of 26,259 454 reads with a median length of 273 bp were obtained for the pearl millet BAC containing PHYC. A total of 21,531 PACBIO reads with an average size of 3,609 bp were obtained and assembled to a single contig. The 454 reads were mapped to the PACBIO assembly, and SNPs were manually corrected if necessary. The total contig size was 80,746 bp (GenBank KF704368).
Pearl millet BAC sequences yielded significant BLASTN hits (Table S3) with genes near the sorghum PHYC gene; three genes, corresponding to sorghum genes Sb01g007830, Sb01g007870 and Sb01g007878, were named Pg7830, Pg7870 and Pg7878, respectively. The prefix Pg refers to the species (Pennisetum glaucum syn. Cenchrus americanus), and the number refers to the last four numbers of the homologous gene in sorghum. Two additional genes (Pg7840 and Pg7880, corresponding to sorghum genes Sb01g007840 and Sb01g007880, respectively) not found in the BAC sequences were also amplified (Table S3). Therefore, including PHYC, we obtained a total of six genes with known or predicted protein coding capability in sorghum. The location of the homologous genes on chromosome 1 in sorghum is shown in Fig. 2a for information.
Signatures of selection and marker-trait association across genes in the vicinity of PHYC. a The genomic region of PHYC in Sorghum bicolor and in Cenchrus americanus. The physical position of sorghum genes is presented at the top according to sequence data from GRAMENE (chromosome 1 of sorghum). A putative order for pearl millet genes (linkage group 2) is proposed at the down, based on sorghum order. Pg7840 was not found on the sequenced BAC and was thus positioned before Pg7830. b F ST between the wild and cultivated samples and Tajima’s D for the cultivated sample were calculated for each amplified fragment of the PHYC region. The significance of these two neutrality statistics was assessed according to the expected distribution of the corresponding statistic in the EGGF model (*p > 0.95 for F ST ; *p < 0.025 or p > 0.975 for Tajima’s D). c The association was fitted on the panel of inbreds using a mixed model. The association analysis was performed on SNPs or indels. The p-value of the marker effect (Wald test) is given at each position (p-value was decimal log transformed). The trait and gene codes are given at the top of each graph. The horizontal line represents the p-value threshold (0.05). a No polymorphism was found for Pg7830 in the germplasm set used to test for signatures of selection
Polymorphism and LD pattern
The complete PHYC locus (6,115 bp) and fragments of 807–923 bp were sequenced for the five other genes in the 90 inbred lines. The amount of sequence information obtained per inbred was 10,423 bp. A total of 75 SNPs/indels with a minor allele frequency >2.5 % were identified. Across the genomic region examined, the LD (r 2) ranged from approximately zero to 1 with an average pairwise measure of r 2 = 0.35 ± 0.01 (Fig. 3). The range and average value of intra-gene LD differed across genes. Polymorphisms within PHYC were the most tightly linked with an average r 2 of 0.74 ± 0.01 (r 2 ranged from 0.15 to 1 for this gene). Pg7830 (and to a lesser extent Pg7840) showed the highest LD with PHYC. The markers in Pg7870, Pg7878 and Pg7880 presented only a weak LD with PHYC.
Linkage disequilibrium in the vicinity of PHYC in pearl millet. a The pairwise squared correlation coefficient (r 2) is plotted across 75 sites (SNPs/indels). b p-value of the F-test. c Summary statistics of r 2 for each individual gene and across all genes. The mean r 2 value, standard error (SE), minimum r 2 value (Min.), maximum r 2 value (Max.) and number of sites (N) underlying each summary statistic are given
Marker-trait association
Each of the 75 markers was fitted to phenotypic traits using a mixed model. Many markers were significantly associated with flowering time (FT), primary spike length (SpL), the number of tillers at head emergence (NTHE) and basal spike diameter (BSpD) (Fig. 2c). No highly significant associations (Figure S1) were found for plant height (PH), stem diameter (SD) and spike diameter (SpD). The significant trait associations observed were limited to polymorphisms in PHYC, Pg7830 and Pg7840. No associations were detected between the traits and the other genes (Pg7870, Pg7878 and Pg7880). Markers associated with BspD were located in PHYC and Pg7880. Some SNPs found in the Pg7830 gene had lower p-values for association with flowering time than the SNP found in the PHYC genes (Table S4). The coefficients of effect estimates in this inbred panel are given for each trait at all markers (Table S6; note that the coefficients for non-significant markers are considered not statistically different from zero).
However, as some genotypic data were missing for a given marker in a fraction of the inbreds, comparing two SNPs based on the strength of the association, i.e., the p value, could be misleading. For example, in terms of their level of significance, the rank of markers Pg7830 767T>G and PHYC 5004C>A could be inversed by changing the subset of inbred analyzed (Table S4). Therefore, to overcome this problem, we compared the SNPs two by two on a similar subset of inbred lines, and for this specific purpose we used an MCMC method.
Comparison of markers using MCMC analysis
We tested different parameter values for the MCMC analysis (see “Materials and methods”). All values in the range concerned gave very similar results. Thus, we set the total number of iterations to 30,000, including a burn-in period of 10,000. The MCMC process was implemented for flowering time, spike length, number of tillers, and basal spike length.
We recorded the frequency of selection of each individual marker with respect to the trait concerned (Fig. 4). The most frequently selected markers were PHYC 5004C>A for FT and SpL traits (respectively 13 and 18 %), PHYC 3219A>C for NTHE (23 %) and PHYC 4967G>A for BSpD (16 %). With respect to these scores, PHYC SNPs are the most frequently selected, and with respect to the other genes sequenced in this study, PHYC appeared to be the best candidate gene for the variation in FT, SpL, NTHE and BSpD.
Comparison of markers using MCMC analysis. The MCMC algorithm was used to iteratively explore the whole set of 75 markers distributed on the six genes. The markers were compared based on log likelihood using a criterion of probability (see text for details). The run consisted of 30,000 iterations per trait. The frequency of choice of each marker (selection score) is presented for flowering time (FT), spike length (SpL), basal spike diameter (BSpD) and number of tillers at head emergence (NTHE). The PHYC gene shows the most frequently chosen markers for all the traits, suggesting a best fit for this gene compared to the other genes
Analysis of shared associations across traits
Some of the polymorphic sites statistically associated with phenotype were shared between two or more traits (FT, SpL, NTHE and BSpD). For each pair of traits, we performed a graphical comparison and a correlation between p-values to highlight markers associated with both traits at the same time (Fig. 5). FT and SpL appeared to be associated with the same set of markers. Markers that provided the highest p-values of FT also provided the highest p-values of SpL. The correlation of paired p-values across markers was positive and significant for these two traits, based on the whole set of markers (R = 0.77, df = 73, p = 4.44 × 10−16) and/or based on PHYC markers (R = 0.81, df = 33, p = 2.79 × 10−9). The remaining combinations of traits produced either no correlation or a weak correlation of p-values, or a negative correlation of p-values. The coefficients of effect estimates for each trait are given with respect to the genotype (Table S7; note that the coefficients for non-significant markers are considered not statistically different from zero).
Correlation of p-values across traits. Marker-trait associations were assessed in the inbred panel using the mixed model for each trait: flowering time (FT), primary spike length (SpL), number of tillers at head emergence (NTHE) and basal spike diameter (BSpD). p-values of the marker effect (Wald test) are correlated for each pair of traits. p-values were decimal log transformed and paired with respect to the marker. Horizontal dashed lines represent the p-value threshold (0.05) for the trait on the first axis, and the vertical line represents the same threshold for the trait on the second axis. Dots located over both thresholds identify markers associated with two traits. The linear regression of the p-values is also plotted (solid line) for each pair of traits. Pearson’s coefficient of correlation is given (R). The significance of each coefficient was assessed using Student’s t test (*p < 0.05, ***p < 0.001 and NS non-significant). The analysis was performed for the 35 markers in the PHYC gene (a) and for all the 75 markers across the six genes sequenced in this study (b). Allelic estimates and log transformed p-values are highly correlated for all of the traits (R values are 0.98, 0.96, 0.97, 0.96, 0.97, 0.96 and 0.93 for the traits BSpD, FT, NTHE, PH, SD, SpD and SpL, respectively). Thus, the correlation based on the estimates of allelic effects rather than the p-values gives consistent results
Signatures of selection
Sequence of six portions of PHYC (Fig. 2: two portions from the first exon, one from the second exon, one from the second intron, one from the third exon and one from the last intron) were already available (Clotault et al. 2012). Five new genes from the PHYC region were amplified, yielding a total of 7,451 bp that were analyzed in a sample of wild and cultivated pearl millet lines (Table S3). A total of 112 SNPs were found in the sequence data. No polymorphisms were found for Pg7830. Values of nucleotide diversity, polymorphisms and neutrality statistical tests were calculated for each gene fragment surrounding the PHYTOCHROME C gene (Table S5). Pg7880 showed a significant positive Fay and Wu’s H for both cultivated (H = 5.198; p = 1) and wild populations (H = 3.384; p = 1, Table S5). A significantly positive Fay and Wu’s H indicates a deficit of moderate- and high-frequency derived SNPs. Other significant neutrality tests were only obtained for portions of PHYC (Table S5). PHYC10 showed a significant positive H for the cultivated polymorphism (H = 0.940; p = 0.982). PHYC4 (Fig. 2a; Table S5) only showed a significant F ST (F ST = 0.311; p = 0.954) suggesting selection along with a significant positive Tajima’s D (D = 2.845; p = 0.980). A positive Tajima’s D suggests the occurrence of balanced selection at PHYC4. The high differentiation suggests a positive signal of selection due to strong changes in allele frequencies between cultivated and wild samples. In addition to the 112 SNPs, indels were also identified. Considering both SNPs and indels gave similar results (data not shown), except for Pg7870, which showed a significant F ST (F ST = 0.357; p = 0.972 with SNPs and Indels compared to F ST = 0.283; p = 0.936 for SNPs).
Discussion
Consistent results in association mapping, QTL analysis and selection signature
Unlike association mapping, the family-based QTL study did not suffer from inflated false positives caused by population structure. Results of the linkage analysis corroborated the presence of QTLs for different traits in this genomic region. All four traits for which we reported significant associations based on association mapping (FT, SpL, NTHE and BSpD) were also significantly associated using linkage mapping. A genotype/phenotype association for flowering time (FTHE) was detected in two families (D1 and C1), but not in the third family (A7). Based on the results observed in the D1 and C1 families, it is likely that a QTL does exist. The absence of association with flowering time suggests that the causal locus underlying this QTL and the diagnostic marker genotyped at PHYC might have recombined in one of the parents during the evolutionary history of pearl millet and resulted in the absence of association in the A7 family. Alternatively, lack of power to detect a QTL effect in the A7 family could explain the absence of association. However, we still clearly detected a spike length QTL, and the family is relatively large and similar in size to D1 (around 260 individuals). The high p-value found for the association with flowering time (F2,260 = 0.29, p = 0.75) did not encourage us to suspect a significant association, which would have been the case if the number of individuals had been higher.
The observed associations were higher in the C1 family. This was expected, as the power for QTL detection depends on sample size (n = 1,182 for C1). Additional genotype-phenotype associations for spike diameter not identified in the inbred panel were observed in the F2 families (Fig. 1). These associations could be due to close linkage of multiple QTLs with the marker analyzed in the PHYC gene or more likely with neighboring genes. The association study revealed stronger effects of the PHYC gene on plant morphology (shorter spike, smaller spike diameter and smaller basal spike diameter) than on flowering time. It seems that the allele of the PHYC identified here has a general effect on inflorescence size that could result in earlier flowering.
In conclusion, the fact that the same associations were identified in both the inbred panel and the F2 families strongly suggests that the PHYC region is indeed associated with flowering time and spike morphology. However, the validation of the association study by QTL analyses should be considered with caution. QTL analyses only allow the identification of a large chromosome region associated with a given trait. It is not excluded that the QTL observed in the F2 mapping population underlies a different genetic locus than the association mapping panel. Further studies such as fine mapping studies are needed to decipher the QTL observed in the F2 family in the vicinity of PHYC. However, the association result was also confirmed by the detection of signatures of selection in wild and cultivated pearl millet samples in the 3′-boundary of the PHYC gene. In a previous study (Saïdou et al. 2009), we found both an association between phenotypic variation and genetic variation and a signature of selection in a restricted region of the PHYC gene. In a more recent work (Clotault et al. 2012) and in the present study, we assessed selection on new cultivated and wild samples and controlled possible bias in the selection signature signal by modeling the evolutionary history of pearl millet. The three studies (Saïdou et al. 2009; Clotault et al. 2012 and this study) with two different samples showed a significant signature and suggest balancing selection at the PHYC gene. This is illustrated by the significant and positive Tajima’s D statistics observed here. Another recent study did not find a strong selection signal at this gene (Lakis et al. 2012). These authors suggest that a difference in the samples might explain the different results (Lakis et al. 2012) and that the PHYC gene could present different selection signatures depending on the area in which the samples are collected. Indeed, there was a major difference between the sample analyzed by Lakis et al. (2012) and our sample (Clotault et al. 2012). In Lakis et al. (2012), the wild sample showed very strong introgression of cultivated alleles: more than half of their wild accessions showed 50 % or more ancestry from the cultivated samples. Similarly, a third of the cultivated samples showed <80 % ancestry in the cultivated cluster (Lakis et al. 2012). None of our samples showed <80 % ancestry in the wild cluster (Clotault et al. 2012), and <10 % of the cultivated cluster showed <80 % ancestry. The signature signals observed in Saïdou et al. (2009) and Clotault et al. (2012) were positive for Tajima D, suggesting balanced selection, and this test is easily compromised if the sample includes highly introgressed material. Indeed, the high gene flow between wild and cultivated forms suggested by the ancestry could easily have blurred this signal by increasing the number of haplotypes and unique SNPs in the cultivated sample. It is also worth noting that the diversity observed in the PHYC gene was higher in the domesticated sample than in the wild sample in Lakis et al. (2012) but not in our studies (Clotault et al. 2012 and the present study). It should also be noted that both studies found evidence of positive selection in the Hd3a pearl millet homolog (Clotault et al. 2012; Lakis et al. 2012) and that detection of positive selection signatures is less influenced by such introgression. The difference between the studies is certainly linked to a difference in the samples and the fact that the sample used was introgressed. In the present study, we not only observed a selection signature, but also confirmed genotype/phenotype association in controlled crosses and in an association mapping population. We fully agree with Lakis et al. (2012) that the PHYC gene could present different selection signatures depending on the sampling area, i.e., in a drier or wetter area. Until now, our studies were performed in dry areas of the Sahel (Saïdou et al. 2009; Vigouroux et al. 2011). In Niger, the frequency of the early flowering allele (Vigouroux et al. 2011) is low (~20 %), while the average flowering time of varieties in Niger is already rather early at ~66.0 days on average for the whole country. Therefore, it is not surprising that the early flowering allele observed in Niger is not present in a wetter area where pearl millet varieties flower later (Haussmann et al. 2006). Flowering time in pearl millet ranged from 40 to 160 days in samples chosen to represent the whole of West and Central Africa (Haussmann et al. 2006). Several genes are expected to be associated with the large variation in flowering time (Clotault et al. 2012; Lakis et al. 2012). The PHYC allelic variation in pearl millet explained only 10 % of the variation (corresponding to an effect of 5–7 days) in the early flowering material composing the inbred panel (Saïdou et al. 2009). However, the early allele identified in the PHYC gene (Saïdou et al. 2009) could help cope with variation in climate conditions in these drier areas in the future (Vigouroux et al. 2011).
Deciphering the statistically most highly associated polymorphisms
Based on the result of the first mixed model (Fig. 2c), several markers at PHYC, Pg7830 and Pg7840 showed a significant association with FT, SpL, BSpD and NTHE. These three genes are tightly linked (high r2 between PHYC and Pg7830 in the inbred panel). In the presence of such LDs, associations would be expected with all SNPs within the haplotype block. A comparison across markers was thus performed to identify the markers yielding the best associations.
We used the MCMC method to iteratively compare two SNPs based on pairwise samples. The probability of selecting a marker as the ‘best’, i.e., that is more closely associated with the trait than the other marker in the pair, was based on the log-likelihood for the model to select this marker. The sample size and the number of parameters were the same in the two markers, which were compared at each step of the MCMC. The frequency with which a marker was selected in this process was expected to increase proportionally to the improvement in the fit provided by this marker compared to other markers. PHYC appeared to be the best candidate for FT, SpL, BSpD and NTHE across the genes examined in this region. However, closer associations may be found with polymorphisms outside the sequences and the genes we examined, for example, upstream of PHYC/Pg7830. Fine mapping of the causative site using segregating populations are needed to clarify and confirm these first results.
It should be noted that, since no genome sequence is available for pearl millet, the genes examined were selected because, in the sorghum genome assembly and BAC sequencing, they are located in the region around PHYC. Comparative analysis with the recently published genome sequence of foxtail millet, a close relative of pearl millet (Bennetzen et al. 2012), showed that in foxtail millet, these genes are located in colinear positions in a 56-kb region. One of the differences between pearl millet and both foxtail millet and sorghum is the gene order. In pearl millet, the Pg7830 gene is located closer to the PHYC gene than Pg7840. In foxtail and sorghum, the Pg7840 homologs are located closer to PHYC than Pg7830. Some rearrangement certainly occurred in the vicinity of the PHYC gene in pearl millet after divergence from the ancestor of sorghum and foxtail millet. At this stage, it is not known whether the rearrangement is shared by different species of the Cenchrus genus or is limited to pearl millet (Cenchrus americanus).
Pleiotropy versus close linkage: MCMC analysis identified the statistically most closely associated polymorphisms in the association mapping population for FT, SpL, NTHE and BSpD within the PHYC gene. These results suggest pleiotropy at the gene scale. However, deciphering the association at the nucleotide level leads to two hypotheses: (1) the association is driven by the same molecular polymorphism; (2) the association is driven by different polymorphisms. Even if the same set of polymorphisms is associated with spike length and flowering time in the inbred sample, the QTL analysis suggests that different polymorphisms are associated with these two traits in the A7 family. However, this interpretation should be considered with caution since it relies on the absence of a statistically significant genotype/phenotype association. At this stage, we propose the hypothesis that the apparently “same molecular polymorphism” observed in the inbred panel is created by a strong linkage disequilibrium in this sample, linkage that is broken in the A7 family. Recent results suggest that a similarly complex pattern is also present in maize with a QTL associated with flowering time (Durand et al. 2012). Further studies are needed to address this particular question.
In this study, we used association mapping combined with linkage analysis and the detection of signatures of selection to provide evidence for the presence of very likely true-positive polymorphisms in the genomic region around PHYC. MCMC analysis suggests that PHYC is still one of the best candidates in the set of sequenced genes at this stage. Further studies, including fine mapping and functional studies in the PHYC region, are suitable to assess whether the identified polymorphisms are causal.
References
Allouis S, Qi X, Lindup S, Gale MD, Devos KM (2001) Construction of a BAC library of pearl millet, Pennisetum glaucum. Theor Appl Genet 102:120–125
ASReml package for R (ASReml-R), version 20/32 (2011) VSN International Ltd, Hemel, Hempstead, HP1 1ES, UK
Atwell S, Huang YS, Vilhjalmsson BJ, Willems G, Horton M et al (2010) Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465:627–631
Beaumont MA, Zhang W, Balding DJ (2002) Approximate Bayesian computation in population genetics. Genetics 162:2025–2035
Bennetzen JL, Schmutz J, Wang H, Percifield R, Hawkins J et al (2012) Reference genome sequence of the model plant Setaria. Nat Biotech 30:555–561
Bergelson J, Roux F (2010) Towards identifying genes underlying ecologically relevant traits in Arabidopsis thaliana. Nat Rev 11:867–879
Brown PJ, Rooney WL, Franks C, Kresovich S (2008) Efficient mapping of plant height quantitative trait loci in a sorghum association population with introgressed dwarfing genes. Genetics 180:629–637
Camus-Kulandaivelu L, Chevin LM, Tollon-Cordet C, Charcosset A, Manicacci D et al (2008) Patterns of molecular evolution associated with two selective sweeps in the Tb1-Dwarf8 region in maize. Genetics 180:1107–1121
Clotault J, Thuillet A-C, Buiron M, De Mita S, Couderc M et al (2012) Evolutionary history of pearl millet (Pennisetum glaucum [L.] R. Br.) and selection on flowering genes since its domestication. Mol Biol Evol 29:1199–1212
Ducrocq S, Madur D, Veyrieras JB, Camus-Kulandaivelu L, Kloiber-Maitz M et al (2008) Key impact of Vgt1 on flowering time adaptation in maize: evidence from association mapping and ecogeographical information. Genetics 178:2433–2437
Durand E, Bouchet S, Bertin P, Ressayre A, Jamin P, Charcosset A, Dillmann C, Tenaillon MI (2012) Flowering time in maize: linkage and epistasis at a major effect locus. Genetics 190:1547–1562
Fay JC, Wu C-I (2000) Hitchhiking under positive darwinian selection. Genetics 155:1405–1413
Gilmour AR, Gogel BJ, Cullis BR, Thompson R (2006) ASReml User Guide Release 20. VSN International Ltd, Hemel Hempstead, HP1 1ES, UK
GRAMENE release 32 (2010) http://www.gramene.org/ (Access Online: November 2010)
Hamilton G, Stoneking M, Excoffier L (2005) Molecular analysis reveals tighter social regulation of immigration in patrilocal populations than in matrilocal populations. Proc Natl Acad Sci USA 102:7476
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyze spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Haussmann BIG, Boubacar A, Boureima SS, Vigouroux Y (2006) Multiplication and preliminary characterization of West and Central African pearl millet landraces. Int Sorghum Millet Newsl 47:110–112
Hudson RR, Slatkin M, Maddison WP (1992) Estimation of levels of gene flow from DNA sequence data. Genetics 132:583–589
Kelly JK (1997) A test of neutrality based on interlocus associations. Genetics 146:1197–1206
Lakis G, Navascués M, Rekima S, Simon M, Remigereau M-S et al (2012) Evolution of neutral and flowering genes along pearl millet (Pennisetum glaucum) domestication. PLoS One 7:e36642
Lewin A (2008) R package: Exact tests for linkage disequilibrium and Hardy-Weinberg equilibrium. http://www.r-project.org
Mariac C, Luong V, Kapran I, Mamadou A, Sagnard F et al (2006) Diversity of wild and cultivated pearl millet accessions (Pennisetum glaucum [L] R. Br.) in Niger assessed by microsatellite markers. Theor Appl Genet 114:49–58
Mariac C, Jehin L, Saïdou AA, Thuillet A-C, Couderc M et al (2011) Genetic basis of pearl millet population adaptation along an environmental gradient investigated by a combination of genome scan and association mapping. Mol Ecol 20:81–91
Myles S, Peiffer J, Brown PJ, Ersoz ES, Zhang Z et al (2009) Association mapping: critical considerations shift from genotyping to experimental design. Plant Cell 21:2194–2202
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Oumar I, Mariac C, Pham J-L, Vigouroux Y (2008) Phylogeny and origin of pearl millet (Pennisetum glaucum [L] R. Br.) as revealed by microsatellite loci. Theor Appl Genet 117:489–497
Pavlidis P, Laurent S, Stephan W (2010) msABC: a modification of Hudson’s ms to facilitate multi-locus ABC analysis. Mol Ecol Resour 10:723–727
Pritchard JK, Stephens M, Rosenberg NA, Donnelly P (2000) Association mapping in structured populations. Am J Hum Genet 67:170–181
Rafalski JA (2010) Association genetics in crop improvement. Curr Opin Plant Biol 13:1–7
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Saïdou A–A, Mariac C, Luong V, Pham J-L, Bezancon G et al (2009) Association studies identify natural variation at PHYC linked to flowering time and morphological variation in pearl millet. Genetics 182:899–910
Shin JH, Blay S, Lewin-Koh N, McNeney B and Graham J (2010) R package: Graphical display of pairwise linkage disequilibria between SNPs. http://www.R-project.org
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tenaillon MI, U’Ren J, Tenaillon O, Gaut BS (2004) Selection versus demography: a multilocus investigation of the domestication process in maize. Mol Biol Evol 21:1214–1225
Thornsberry JM, Goodman MM, Doebley J, Kresovich S, Nielsen D et al (2001) Dwarf8 polymorphisms associate with variation in flowering time. Nat Genet 28:286–289
Thornton K (2009) Automating approximate Bayesian computation by local linear regression. BMC Genet 10:35
Tian F, Bradbury PJ, Brown PJ, Hung H, Sun Q et al (2011) Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat Genet 43:159–162
Vigouroux Y, McMullen M, Hittinger CT, Houchins K, Schulz L et al (2002) Identifying genes of agronomic importance in maize by screening microsatellites for evidence of selection during domestication. Proc Natl Acad Sci USA 99:9650–9655
Vigouroux Y, Mariac C, Pham J-L, Gérard B, Kapran I et al (2011) Selection for earlier flowering crop associated to climatic variations in the Sahel. PLoS One 6:e19563
Watterson GA (1975) On the number of segregating sites in genetical models without recombination. Theor Pop Biol 7:256–276
Yu J, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotech 17:155–160
Yu J, Pressoir G, Briggs WH, BI IV, Yamasaki M et al (2006) A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat Genet 38:203–208
Zhu C, Gore M, Buckler ES, Yu J (2008) Status and prospects of association mapping in plants. Plant Genome 1:5–20
Acknowledgments
This study is funded by a French National Agency for Research (ANR) grant (ANR-07-JCJC-0116-01 to Y.V.). This study is also partly supported by Agropolis Fondation (ARCAD project) and Institut de Recherche pour le Développement PhD grant to A-AS.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The experiments of this article comply with the current laws of the country in which they were performed.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Gore.
A.-A. Saïdou and J. Clotault contributed equally to this work.
Sequence data used in this article are deposited in GenBank under accession numbers KF525941–KF526110 and KF704368.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Saïdou, AA., Clotault, J., Couderc, M. et al. Association mapping, patterns of linkage disequilibrium and selection in the vicinity of the PHYTOCHROME C gene in pearl millet. Theor Appl Genet 127, 19–32 (2014). https://doi.org/10.1007/s00122-013-2197-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-013-2197-3