Abstract
Cholangiocarcinoma (CCA) remains a deadly disease in part due to its late diagnosis. Non-invasive approaches to early detection are challenging, and pathological confirmation is usually required for final diagnosis. In this chapter, we summarise the biomarkers in clinical use as well as those currently under study for the detection and prognostic classification of CCA.
Access provided by Autonomous University of Puebla. Download chapter PDF
Similar content being viewed by others
Keywords
Introduction
The term “biomarker” in clinical care refers to biomolecules produced by different cell populations in the human body that have been strongly linked with a particular disease or disorder [1, 2]. The detection and quantification of these molecules have become particularly useful to diagnose diseases at an early stage as well as to select the best therapeutic option for personalised medicine. Similarly, some biomarkers can also predict patient prognosis and risk of disease relapse [2, 3]. For clinical use, an ideal biomarker should be highly specific and sensitive for a disease, being able to differentiate the disease from other biologically related conditions (specificity) and detect the disease when low levels of the biomolecule of interest are present (sensitivity) [4, 5].
In the case of CCA, early diagnosis remains an area of critical need, as presently 65% of patients are diagnosed with an advanced stage of the disease when treatment options are very limited [6]. The frequently late diagnosis of CCA is mainly due to the non-specific symptoms and clinical manifestations of CCA, which are common to other biliary obstructive conditions [7, 8]. In contrast, the 35% of the cases diagnosed at early stages have potentially curative options, such as surgical resection or liver transplantation [9].
The diagnosis of CCA is based largely on non-invasive imaging, which may include contrast-enhanced computed tomography (CT) and magnetic resonance imaging (MRI). Endoscopic ultrasonography (EUS), endoscopic retrograde cholangiopancreatography (ERCP), and percutaneous transhepatic cholangiography (PTC) have additional roles in tissue acquisition and stenting of biliary strictures, with a very small risk of peritoneal seeding reported with PTC- and EUS-guided biopsy [10]. Serum biomarkers for diagnosis lack sensitivity and specificity [5, 6, 11], and there are currently no biomarkers established to predict patient outcome for the different disease sites: extrahepatic CCA (eCCA), intrahepatic CCA (iCCA), and perihilar CCA (pCCA) [11].
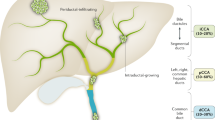
In this chapter we will review current biomarkers used in the clinical setting for the detection of CCA as well as novel strategies still under development for early diagnosis, surveillance, and prognostication and grouped according to tissue source (see Fig. 19.1).
Emerging biomarkers for the detection and prognostication of CCA according to tissue source. (This figure was created using images from Servier Medical Art Commons Attribution 3.0 Unported License (http://smart.servier.com). Servier Medical Art by Servier is licensed under a Creative Commons Attribution 3.0 Unported License)
Established Serum Biomarkers
The most commonly used serum biomarkers for the detection of CCA are carbohydrate antigen 19-9 (CA19-9) and carcinoembryonic antigen (CEA). However, they have low sensitivity and specificity and are not adequate for early diagnosis [3, 7].
CA19-9
CA19-9 is a sialylated Lewis blood group antigen reported for the first time in colorectal cancer cells [12]. Synthesis of this biomolecule is directly linked to the activity of its two precursor enzymes known as fucosyltransferases 2 and 3 (FUT2 and FUT3). Patient genotype regarding these enzymes affects the overall amount of CA19-9 liberated into the bloodstream, and it is estimated that 10% of the population have inactive FUT3, thus being CA19-9 antigen negative (even in the presence of tissue-proven CCA) [13].
The sensitivity and specificity of CA19-9 for CCA vary among studies and patient cohorts analysed but are estimated to be around 50–80% and 40–70%, respectively [14]. Its low specificity arises from the fact that this marker is also detected in the serum of patients with other malignancies, such as pancreatic cancer, colorectal cancer, and hepatocellular carcinoma (HCC), as well as in benign inflammatory conditions including acute cholangitis, pancreatitis, choledocholithiasis, hepatitis, and cirrhosis [15, 16]. Levels of CA19-9 greater than 100 U/mL, in the absence of cholangitis, tend to indicate the presence of malignancy in the biliary tree [14, 15, 17]. Lastly, levels of CA19-9 are also elevated in some respiratory conditions such as bronchiectasis, pulmonary fibrosis, and emphysema. For these reasons, it is estimated that around 10% of patients with elevated CA19-9 levels do not have any cancer of the biliary tree but instead some kind of benign disease [19].
CEA
CEA is a group of 12 glycoproteins involved in cell adhesion that was linked to malignancy for the first time in colorectal cancer specimens [20]. Generally, levels higher than 5 ng/mL are considered abnormal in clinical practice [21]. Levels of these proteins are elevated during foetal development, but their expression is minimal in adults. Although CEA represents a reliable biomarker for colorectal cancer, it is only elevated in 30% of patients with CCA. Like CA19-9, CEA may also be elevated in other conditions such as pancreatic cancer, colorectal cancer, cirrhosis, hepatitis, cholangitis, and inflammatory bowel disease [22]. Thus, detection of CEA in patients with suspected CCA may indicate a different primary malignancy metastatic to the hepatobiliary system.
CA-125
The carbohydrate antigen 125 , also known as mucin 16, is a large cell surface protein encoded by the MUC16 gene generally found in the epithelia of the endometrium, ovaries, bronchi, and cornea [23]. It is elevated in about 65% of CCA patients (normal upper limit of 37 U/mL) but also in pancreatic, colon, ovarian, breast, and lung cancers as well as in benign hepatobiliary conditions like cirrhosis [18, 24]. As with CEA, this serum biomarker is generally only tested when another primary malignancy is suspected.
Emerging Biomarkers
Blood Biomarkers
Among all the non-invasive sources of biomarkers, blood is easily and routinely collected and is rich in biomolecules. Therefore, the majority of published and ongoing research studies have sought to identify more specific and sensitive blood biomarkers for CCA or to combine several biomarkers into a panel to improve their performance (see Table 19.1).
Extracellular Vesicles
Extracellular vesicles (EVs) have emerged as a promising source of biomarkers. EVs consist of a heterogeneous population of lipid bilayer spheres containing different biomolecules such as proteins, nucleic acids (DNA and RNA), lipids, and other metabolites [25]. Cells use these biomolecules for intercellular communication and, in the context of disease, to modulate pathological pathways [26]. EVs have been found in all body fluids including blood [27], urine [28], bile [29], saliva [30], and ascites [31].
Based on their diameter (30 nm–2 μm), EVs have been classified into two main groups: (i) small EVs or exosomes (30–100 nm in diameter) and (ii) large EVs or microvesicles (>100 nm) [27]. Small EVs constitute the most studied group. They have their origin in vesicles derived from the endomembranous system, which accumulate to form multivesicular bodies and merge with the plasma membrane releasing the exosomes to the extracellular space. In contrast, large EVs bud directly from the cell membrane of the parental cell [32, 33].
The amount, content, and surface markers of EVs have proven to reflect the biological features and staging of different types of cancer, including CCA [35]. In one study performed by Arbelaiz et al., the total amount and protein content of EVs isolated from patients with CCA, primary sclerosing cholangitis (PSC), HCC, and healthy individuals were compared [36]. HCC patients had the highest EV density in serum compared to the other groups. A range of different proteins were suggested as potential biomarkers to distinguish CCA from the other groups. Among these proteins, AMPN (aminopeptidase N), VNN1 (pantetheinase), and PIGR (polymeric immunoglobulin receptor) showed the highest diagnostic values when comparing CCA patients to healthy controls (AUC 0.878, 0.876, and 0.844, respectively), whereas FIBG (fibrinogen gamma chain), A1AG1 (alpha-1-acid glycoprotein), and protein S100A8 (AUC 0.796, 0.794, and 0.759, respectively) were the most promising candidates to differentiate PSC from CCA. Lastly, the proteins FCN2 (ficolin 2), ITIH4 (inter-alpha-trypsin inhibitor heavy chain 4), and FIBG (AUC 0.956, 0.881 and 0.881, respectively) showed promise for the differential diagnosis of early-stage CCA (stages I–II) and PSC.
Transcriptomic analysis of EV content revealed messenger RNA (mRNA) and different non-coding RNA molecules (such as microRNAs [miRs], long non-coding RNAs, and small nucleolar RNAs) with potential for CCA diagnosis. In a recent study by Lapitz et al., the serum EV content of CCA patients was compared with benign biliary conditions like PSC and ulcerative colitis (UC) as well as with healthy individuals [37]. A total of 1932 transcripts were differentially expressed in the CCA group compared to the other 3 subgroups combined into a single group, with RFFL (E3 ubiquitin-protein ligase rififylin), ZNF266 (zinc finger protein 266), and OR4F3 (olfactory receptor family 4 subfamily F member 3) being the mRNAs with the highest diagnostic potential (AUC 1.00, 0.976, and 0.960, respectively). The best-performing non-coding RNAs were miR-551B, PMS2L4, and LOC643955 (AUC 0.909, 0.880, and 0.873, respectively).
The fact that large EVs bud from the plasma membrane of the parental cell is particularly interesting because some of the surface markers will remain on the vesicle lipid bilayer and mirror the pathobiological cues of the disease. For instance, tumour-associated microparticles (taMPs) carrying markers such as Annexin V, EpCAM (epithelial cellular adhesion molecule), and ASGPR1 (asialoglycoprotein receptor 1) allowed the differentiation of patients with liver malignancies (including CCA and HCC) from patients bearing non-liver cancers and cirrhosis, with 65.8% sensitivity but only 47% specificity. These taMPs decreased significantly at 7 days after curative resection, demonstrating their potential prognostic value [38, 39].
Nucleic Acid Biomarkers
Similar to EVs, cell-free nucleic acids are released by healthy and cancer cells into different body fluids such as blood, urine, or bile [3, 34]. In terms of circulating DNA, some of the most commonly mutated genes in CCA, including KRAS, NRAS, BRAF, and PIK3CA, were first screened in tumour tissue by multiplex PCR and then in DNA isolated from matched plasma samples from patients. The mutation pattern of the tumour was conserved in plasma suggesting the suitability of the technique for cancer detection [40].
Another group of nucleic acid biomarkers are circulating RNAs, with some studies suggesting their diagnostic value in CCA [41, 42]. Among them, miR-21, a small transcript with important roles in development, inflammation, and cancer invasion [43], was the most commonly upregulated miR in CCA, and in one study, it differentiated iCCA patients from healthy individuals with an AUC 0.91 in serum and 0.94 in plasma [44]. However, it was also upregulated in the blood of patients with other cancers such as HCC, limiting its specificity [45]. Patients with CCA have also shown increased levels of miR-26a [46], miR-150 [47], and miR-192 [48] and decreased levels of miR-106a [49].
Interestingly, a comparison of the differential expression of miRs in the blood of PSC-derived CCA patients (n = 7) and CCA alone patients (n = 63) showed increased levels of miR-222 and miR-483-5p in the latter group, achieving an AUC of 0.770 when combined [50]. Another study reported a panel of five miRs in serum (miR-126, -1281, -26a, -30b, and -122) with an individual maximum AUC of 0.870, though a combination of the different miRs in a logistic regression model did not significantly improve their diagnostic accuracy [51]. However, further studies with larger cohorts are needed to validate these findings.
One of the strongest features that makes cell-free nucleic acids good candidate biomarkers for CCA is their ability to reflect the very heterogeneous pattern of mutations of these tumours. In addition, they can be easily isolated from blood, amplified, and detected by well-established targeted techniques such as qRT-PCR and microarray or by untargeted techniques such as whole genome sequencing (WGS) or whole transcriptome sequencing (also known as total RNA sequencing, RNA-Seq) [52, 53]. Although the above-mentioned publications support the potential of nucleic acid biomarkers for the early detection of CCA and patient stratification, further studies are still needed to validate these findings.
Proteins and Peptides
Proteins are generally more stable and abundant in blood than nucleic acids. In recent years, many studies have taken a proteomic approach aiming at identifying a protein signature for CCA [19].
Osteopontin (OPN) is a potential novel biomarker for CCA. This glycoprotein is involved in normal physiological processes like bone biomineralisation and in pathological conditions such as chronic inflammation [54] and tumour formation [55]. Serum levels of OPN have been reported to be elevated in patients with CCA compared to PSC and healthy controls, with an AUC of 0.964. Persistently high levels post-tumour resection were associated with poor postoperative survival, highlighting its potential role as a prognostic biomarker [56].
Interleukin-6 (IL-6) is an inflammatory cytokine secreted by cholangiocytes upon inflammatory stimuli [57, 58, 59]. In CCA, the cancerous cells also produce this protein, which upregulates Bcl-2, an antiapoptotic cytosolic protein, promoting tumour growth [60]. Serum levels of IL-6 had a 73% sensitivity and 92% specificity (AUC 0.875) for distinguishing CCA patients from healthy controls [61]. Additionally, the concentration of IL-6 in blood decreased after tumour resection. Of note, IL-6 was also elevated in patients with other liver cancers like HCC [62]. Similarly, the cytokine transforming growth factor-β1 (TGF-β1), associated with cell invasion and microenvironment modification, was also found to be elevated in patients with CCA compared to healthy individuals (AUC 0.668) and to other inflammatory conditions (AUC 0.644) [63].
Another proposed CCA biomarker is the soluble fragment of cytokeratin-19 , i.e. CYFRA 21-1 [64]. This marker has been studied in other cancers, e.g. lung cancer [65], and in one study it distinguished CCA from other benign biliary conditions with 75.6% sensitivity and 96.2% specificity [66]. CYFRA 21-1 levels have also been reported to correlate with tumour stage and patient prognosis, with a 3-year survival rate of 76% in patients with low levels of CYFRA 21-1 compared with only 25% in patients with high concentrations [67]. One study from our group by Cuenco et al. reported that a panel of serum protein biomarkers, which included CYFRA21-1, PKM2 (pyruvate kinase M2), MUC5AC (mucin 5AC), and GGT (gamma-glutamyltransferase), was able to differentiate CCA from PSC alone with 81.8% sensitivity and 90.0% specificity (AUC 0.903) [68].
Matrix metalloproteinases 7 and 9 (MMP-7 and MMP-9) are two enzymes responsible for the degradation and remodelling of the extracellular matrix in processes of tissue repair, embryonic development, and angiogenesis [69]. They also play an important role in tumour formation, and as such, increased levels of these proteins in blood have been linked to tumour presence [70,71,72]. Increased serum levels of MMP-7 in CCA patients compared with individuals with benign biliary conditions showed an AUC of 0.730, but MMP-9 did not reliably differentiate between the two groups [73].
Several other proteins for CCA detection have been described. For example, serum S100A6, a calcium-binding protein, had an AUC of 0.909 when comparing CCA to healthy controls [74], while Dickkopf-related protein 1 (DKK1) had an AUC of 0.872 in iCCA versus healthy controls [75]. Spermatogenesis-associated protein 20 (SSP411) showed an AUC of 0.913 in CCA versus benign biliary disorders and healthy controls [76]. A specific glycoprotein known as KL-6, a type of MUC1, has also showed potential when comparing blood levels of CCA patients to healthy individuals, HCC patients, and metastatic liver cancer patients [77]. Serum alpha fetoprotein (AFP) showed good results differentiating HCC from CCA patients, but low specificity [78]. However, when combining AFP with carbohydrate antigen 242 (CA-242), a potential marker of pancreatic cancer [79], the overall sensitivity and specificity increased to 93.4% and 89.7%, respectively [80]. Larger studies are needed to validate these findings.
Serum Metabolites
In addition to the previously mentioned biochemical groups, small molecules have also been reported to be altered in the blood of patients with CCA. Sialic acid (TSA) is a neurotransmitter which has been linked to different cancers including brain tumours, leukaemia, melanoma, and also CCA [81]. In one study by Wongkham et al., the total amount of TSA in CCA patients compared with individuals with benign biliary conditions (including cholangitis, cirrhosis, and gallstones, among others) had an AUC of 0.670, which increased to 0.856 when compared to healthy controls alone [82]. In another study, Kongtawelert et al. compared TSA levels in blood of CCA versus HCC patients (AUC 0.885) and CCA versus cirrhosis and chronic hepatitis patients (AUC 0.964) [83].
Liang et al. performed a metabolomic analysis of the serum of CCA patients and compared it with healthy individuals. During the initial discovery phase, 75 differentially expressed metabolites were found between groups. After the validation phase in 225 CCA and 101 healthy serum samples, the 4 markers that showed the best diagnostic performance were 21-deoxycortisol (under-expressed in CCA), bilirubin (over-expressed), lysophosphatidylcholine 14:0 (lysoPC (14:0), under-expressed), and lysophosphatidylcholine (lysoPC (15:0), over-expressed). The calculated AUCs were 0.918, 0.922, 0.954, and 0.927, respectively, which increased to 0.993 when the four metabolites were combined [84]. Further validation studies of these metabolites are expected.
Circulating Tumour Cells
Circulating tumour cells (CTCs) represent a potential tool for the detection of CCA and have been reported in other cancers including HCC [85], pancreatic cancer [86], and colorectal carcinoma [87]. CTCs have also been studied as a cause of metastasis and tumour relapse, opening the possibility of their use as prognostic biomarkers [88]. To date, the only FDA-approved system for cancer detection is the CellSearch® system, which selects CTCs positive for DAPI, CK-8/18 (hepatocyte antigens), and CK-19 (cholangiocytes antigen) and negative for CD45 staining (leukocyte antigen). However, in one study, only 25% of patients with CCA showed elevated CTC levels [89].
The accuracy of CTCs as prognostic biomarkers has not yet been well-studied in CCA, and only a few studies have reported small subgroups of CCA patients within their cohorts of patients with hepatobiliary disorders. One study isolated CTCs from the blood of CCA patients and showed a high variability in mutations on exon 12 of KRAS gene [88]. Others have reported that CTCs could play an important role in tumour metastasis through their interaction with different T-cell populations in the circulation [86].
Bile Biomarkers
Bile is a complex mix of biomolecules synthesised by the liver and used in the process of digestion. Its major components include bile acids, phospholipids, cholesterol, urea, bilirubin and a variety of hormones and digestive enzymes. In recent years, bile has gained attention as a source of potential novel biomarkers for biliary disorders like CCA [89, 90]. Different groups have used approaches based on omics to identify alterations in the concentration and composition of these organic molecules.
Bile Metabolites
Metabolomic studies performed by several groups have identified alterations in the composition of bile in CCA patients [91]. Bile acids, phospholipids, and cholesterol have shown the biggest differences between disease groups, highlighting their promising diagnostic potential.
Bile acids are steroid acids synthesised from cholesterol by hepatocytes. They regulate levels of cholesterol in the body, assist in fat absorption, and allow phospholipid transport. Analyses of the total content of bile acids in CCA patients have shown a reduction in secondary bile acids, such as deoxycholic and lithocholic acids, compared to patients with biliary tract stones and healthy individuals [92]. A decrease in glycine- and taurine-conjugated bile acids, phospholipids, and cholesterol was also observed in CCA patients compared to benign controls [93]. However, when comparing bile from CCA and PSC patients, the levels of glycine-conjugated acids and phosphatidylcholines were significantly increased in the cancer group [94]. In another study, magnetic resonance spectroscopy of bile had an 88.9% sensitivity and an 87.1% specificity in distinguishing CCA from non-PSC benign biliary conditions [95].
Proteins and Peptides
Analysis of the bile proteome has also been widely explored to identify proteins abnormally expressed in CCA patients. In this context, biliary levels of insulin-like growth factor 1 (IGF-1), also called somatomedin C, a hormone synthesised in the liver with important roles in growth and development, was increased in eCCA patients compared with those with pancreatic cancer or non-malignant disorders (AUC 1.00) [96]. Lipocalin-2 (LCN2), a secreted protein responsible for the transport of some hydrophobic substances, was found elevated in the bile of 30 CCA patients compared to 36 gallstone patients, with a sensitivity of 87% and an AUC of 0.81 [97].
Minichromosome maintenance (MCM) proteins are a family of proteins conserved in all eukaryotic cells due to their role in DNA replication [98] which have been explored as markers of proliferation and cancer progression [99]. Specifically, MCM-7 has been linked to the activation of oncogenes in CCA [100]. In a study conducted in our laboratory by Ayaru et al., biliary MCM-5 as a marker of pancreatobiliary malignancy (including CCA) had a sensitivity of 66% compared to 20% for biliary brush cytology (AUC 0.800) [101].
The ratio of pancreatic elastase (PE) and amylase in bile could be another possible marker of CCA. Low levels of amylase may be associated with complete biliary obstruction caused by a tumour. Patients with CCA had an increased PE/amylase ratio compared with gallstone patients, with a sensitivity of 82% and a specificity of 89% (AUC 0.877) [102].
Lastly, increased biliary levels of Mac-2-binding protein (Mac-2BP), a cell-adhesive protein of the extracellular matrix found upregulated in many cancers, showed promising values for the differential diagnosis of CCA from patients with benign biliary disorders including PSC. The study reported an AUC of 0.700 that increased to 0.750 when Mac-2BP levels were combined with biliary levels of CA19-9 [103].
Extracellular Vesicles
EVs in bile have also been studied for the differential diagnosis of CCA. The total amount of EVs in bile in CCA patients was compared to patients with non-malignant bile duct stenoses. Cancer patients showed an increased total amount of EVs (4.00 × 1015 compared to 1.26 × 1014 nanoparticles/L), being able to differentiate malignant from non-malignant patients with 100% sensitivity and 100% specificity (AUC 1.00) [104]. Another study compared the transcriptomic content of EVs in patients with the same conditions and found a panel of five miRs (miR-191, miR-486-3p, miR-1274b, miR-16, miR-484) able to differentiate CCA with 67% sensitivity and 96% specificity [105].
Genetic Biomarkers
Circulating RNAs have also been reported in bile. In one study by Plieskatt et al., the content of miRs in bile was compared between patients with malignant and benign biliary tract conditions like choledocholithiasis. miR-9 and miR-145 showed the highest diagnostic accuracy (AUC 0.975 in both cases) to differentiate between the two groups [106]. A different research group found that the expression of miR-150-5p was decreased in the bile of CCA patients compared to healthy individuals [47]. In terms of their ability to differentiate PSC-derived CCA from benign PSC, four miRs were identified (miR-412, miR-640, miR-1537, and miR-3189) showing AUC ranging from 0.78 to 0.81. The combination of biliary miR-1537 and CA19-9 levels had an AUC of 0.91 [51].
Changes in DNA methylation in CCA bile samples have been studied by a few groups. Shin et al. described a biomarker panel based on the altered methylation pattern of five genes involved in tumour growth, invasion, migration, and differentiation (CCND2, CDH13, GRIN2B, RUNX3, and TWIST1). The combination was able to differentiate eCCA from a control group of patients with benign disorders like cholecystitis and cholangitis with 83.3% sensitivity and 100% specificity [107]. Thus, the detection of DNA methylation alterations could be a powerful diagnostic strategy for patients with CCA, but further studies are needed to consolidate these data.
Serotonin
Serotonin is a neurotransmitter which also plays an important role in liver regeneration and is overexpressed in CCA [108]. Alpini et al. reported increased levels of TPH-1, an enzyme involved in the route of serotonin synthesis, in 48 tumour biopsies of CCA patients compared to healthy liver tissue. Decreased levels of enzymes responsible for serotonin degradation, such as monoamine oxidase A, were also observed. Finally, increased levels of serotonin were found in the bile of CCA patients but not in patients with intrahepatic stones [109].
Urine Biomarkers
Urine has been shown to be a useful non-invasive source of biomarkers for different cancers, including bladder [110], kidney [111], liver [112], and pancreas [113], with advantages of ease of access and low proteome complexity compared to blood [114]. However, its applicability to biliary tract cancers has been only recently explored.
Volatile Organic Compounds
Urine is a body fluid rich in a variety of volatile organic compounds (VOCs) that have been screened for markers that may differentiate CCA from benign biliary conditions. Navaneethan et al. used selected-ion flow-tube mass spectrometry (SIFT-MS) to measure the concentration of VOCs in urine. Results showed that a combination of ethane and 1-octene levels distinguished CCA from PSC patients with an 80% sensitivity and 100% specificity (AUC 0.900). In addition, by comparing the concentration of 2-propanol and acetonitrile, the authors reported an efficient tool for separating CCA cases from a mixed group of PSC and other benign biliary conditions (AUC 0.862) [115].
Proteomic Profile
The total urinary proteome of 41 patients bearing different biliary conditions including CCA, PSC, and other benign disorders has also been analysed with the aim of finding a singular proteomic signature able to detect and differentiate malignancies. Around 5600 different peptides were selected from the samples with a frequency higher than 20%. Out of these, 43 peptides were chosen as potential biomarkers of CCA showing AUC ranging from 0.630 to 0.890 [116]. Some of these peptides were assigned to well-known proteins such as collagen α-1 and α-2 (extracellular matrix components), osteopontin, uromodulin (an inhibitor of calcium crystallisation in renal fluids), and the antigen CD99 (involved in transmembrane transport). The panel of identified distinct markers was then validated in a different cohort of 123 patients that included CCA, PSC, and other benign biliary disorders, proving efficiency in discriminating CCA cases with a sensitivity of 83% and a specificity of 79% (AUC 0.870) [117].
Extracellular Vesicles
EVs have also been sourced from urine showing particularly good potential when their transcriptomic content was analysed. In a comparative study of the RNA content of EVs from CCA (n = 23), PSC (n = 5), UC (n = 12), and healthy individuals (n = 5), a total number of 27,319 transcripts were identified, out of which 1470 were unique in CCA samples. Messenger RNAs INO80D, MAP6D1, and RRAGD (AUC 1.00 for all) and non-coding RNAs HCG4, MIR200C, and LOC100134868 (AUC 0.930, 0.904, and 0.896, respectively) were the best candidate markers for the differential diagnosis of CCA versus healthy individuals. CLIP3, VCAM1, and TRIM33 messenger RNAs (AUC 0.965 for all) were able to differentiate CCA versus PSC; whereas MT1F, GPX3, and LDHA (AUC 0.915, 0.897, 0.894, respectively) were selected to distinguish CCA from a control group that combined PSC, UC, and healthy patients [37].
Histological Markers
Biopsy confirmation is often required for the clinical diagnosis of CCA [10]. In this regard, biomarkers analysed in resected tissue may provide complementary information for patient stratification, prognosis, and personalised therapies.
Genomic Markers
Considering the high heterogeneity of mutations between the different subtypes of CCA tumours, finding key and specific altered genes can inform treatment options and predict patient outcome after tumour resection [118]. Mutations in DNA repair and cell growth genes (such as TP53 and KRAS, respectively) have been linked to worse patient prognosis than mutations in metabolic genes like IDH-1 and IDH-2 (encoding the enzymes isocitrate dehydrogenases 1 and 2) [7, 119,120,121,122]. The fibroblast growth factor receptor 2 gene (FGFR2) has been frequently reported mutated in iCCA patients and not in other liver cancers, suggesting the diagnostic potential of this genetic marker [118]. However, studies involving larger patient cohorts are required to validate these markers.
CCA tissue samples have also been used to identify novel biomarkers based on epigenetic alterations. A study conducted by Andresen and co-workers identified a four-gene panel (CDO1, CNRIP1, SEPT9, VIM) using tissue from brush cytologies. The panel showed a sensitivity of 85% and a specificity of 98% (AUC 0.944) in discriminating CCA from PSC patients [123]. Similarly, and although less investigated, other genes have also shown abnormal methylation patterns in CCA such as MLH1, DCLK1, CDO1, ZSCAN18, and ZNF331. These genes play key roles in DNA repair, stemness, and tumour growth and invasion [124, 125]. To date, the characterisation of the CCA methylome is still limited. A summary of the main steps towards the implementation of a novel biomarker is represented in Fig. 19.2.
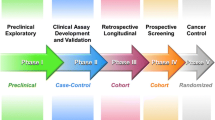
Steps to cancer biomarker discovery. The process starts with the selection of a cohort of patients for biomarker discovery. Samples (blood, bile, urine, tissue) are screened for potential altered biomolecules. Levels of these molecules are then compared between cancer and control groups, and statistical differences and diagnostic potential are calculated. Only a few biomolecules (or a combination) get selected for further validation in larger cohorts. (This figure was created using images from Servier Medical Art Commons Attribution 3.0 Unported License (http://smart.servier.com). Servier Medical Art by Servier is licensed under a Creative Commons Attribution 3.0 Unported License)
Transcriptomic Markers
Whole transcriptome sequencing of resected tumour tissue has allowed researchers to better understand the biology and behaviour of CCA. In an integrative molecular analysis that combined gene expression levels, single-nucleotide polymorphisms (SNPs), and immunohistochemical markers of 153 patients, two subtypes of iCCA were found. In the so-called inflammatory type, an overexpression of inflammation-related genes (such as different cytokines and STAT3) was reported, whereas in the ‘proliferation type’, an activation of oncogenes (KRAS, MAPKs, and MET) was observed and linked to worse patient outcome [126].
A different study examining the recurrence-free interval after resection of CCA and gallbladder cancer linked the overexpression of CTL4 (cytotoxic T-lymphocyte-associated protein 4) and FOXP3 (marker of naturally occurring regulatory T cells) to patients with no CCA recurrence up to 18 months after resection [127]. Recently, the expression of IL-33 in tumour tissue was also associated with better prognosis in both iCCA and pCCA patients [128].
Immunohistochemical Markers
A number of prognostic markers for CCA have been identified using histological tumour samples. In one of the largest meta-studies to date, the histological signature of 4126 CCA patients was compared, identifying 77 potential prognostic biomarkers. Some of these markers were fascin (an actin bundling protein), the epithelial growth factor receptor (EGFR), mucins 1 and 4 (MUC1/4), and p27 (a tumour suppressor protein able to block the proliferation of cancer cells). Over-expression of these five proteins was linked to increased patient survival [129]. Similarly, Suzuki et al. reported that increased expression of the pyruvate kinase type M2 (PKM2) in tumour tissue may enhance tumour cell invasion and promote lymph node metastasis in iCCA cases [130].
micro-RNA
Microarray profiling of CCA resected tumours has allowed the identification of small RNA molecules with the potential for CCA detection and staging [39]. The best characterised is miR-21, which has shown 95% sensitivity and 100% specificity in the detection of CCA versus normal bile duct and liver tissue [131, 132]. This miR has also been linked to worse prognosis. In another study, miR-21 had similar expression in CCA and PDAC tissue compared to normal surrounding tissue, while a panel of seven other miRs were differently expressed [133].
Conclusion and Future Perspectives
CCA remains a deadly disease due in part to the lack of accurate non-invasive diagnostic tests, especially for early-stage disease. The identification and validation of specific markers could not only help discriminate which patients could undergo tumour resection, the only available curative option to date, but also predict the risk of recurrence after surgery. A great number of biomarkers including genetic material (DNA/RNA), circulating proteins, extracellular vesicles, bile acids, and metabolites, among others, have been identified in blood, bile, urine, and tumour tissue of CCA patients. However, in order to be translated to the clinical setting, the diagnostic and prognostic potential of these biological entities need to be validated in large international cohorts and compared with appropriate disease control groups.
Abbreviations
- BBD:
-
benign biliary disorders
- CCA:
-
cholangiocarcinoma
- CT:
-
computed tomography
- CTCs:
-
circulating tumour cells
- eCCA:
-
extrahepatic cholangiocarcinoma
- ERCP:
-
endoscopic retrograde cholangiopancreatography
- EUS:
-
endoscopic ultrasonography
- EVs:
-
extracellular vesicles
- HCC:
-
hepatocellular carcinoma
- iCCA:
-
intrahepatic cholangiocarcinoma
- MRI:
-
magnetic resonance imaging
- pCCA:
-
perihilar cholangiocarcinoma
- PSC:
-
primary sclerosing cholangitis
- PTC:
-
percutaneous transhepatic cholangiography
- UC:
-
ulcerative colitis
- VOCs:
-
volatile organic compounds
References
Ongen Z. What do biomarkers mark? Anatol J Cardiol. 2016;16(2):75.
Diamandis EP. Towards identification of true cancer biomarkers. BMC Med. 2014;12(1):156.
Srivastava A, Creek DJ. Discovery and validation of clinical biomarkers of cancer: a review combining metabolomics and proteomics. Proteomics. 2019;19(10):e1700448.
Henry NL, Hayes DF. Cancer biomarkers. Mol Oncol. 2012;6(2):140–6.
Macias RIR, et al. The search for novel diagnostic and prognostic biomarkers in cholangiocarcinoma. Biochim Biophys Acta Mol basis Dis. 2018;1864(4 Pt B):1468–77.
Banales JM, et al. Expert consensus document: Cholangiocarcinoma: current knowledge and future perspectives consensus statement from the European Network for the Study of Cholangiocarcinoma (ENS-CCA). Nat Rev Gastroenterol Hepatol. 2016;13(5):261–80.
Marin JJG, et al. Chemoresistance and chemosensitization in cholangiocarcinoma. Biochim Biophys Acta Mol basis Dis. 2018;1864(4 Pt B):1444–53.
Banales JM, et al. Cholangiocarcinoma 2020: the next horizon in mechanisms and management. Nat Rev Gastroenterol Hepatol. 2020;17:557.
Rizvi S, Khan SA, Hallemeier CL, Kelley RK, Gores GJ. Cholangiocarcinoma - evolving concepts and therapeutic strategies. Nat Rev Clin Oncol. 2018;15(2):95–111.
Eloubeidi MA, et al. Endoscopic ultrasound-guided fine needle aspiration biopsy of suspected cholangiocarcinoma. Clin Gastroenterol Hepatol. 2004;2(3):209–13.
Macias RIR, et al. Diagnostic and prognostic biomarkers in cholangiocarcinoma. Liver Int. 2019;39(Suppl 1):108–22.
Malaguarnera G, Paladina I, Giordano M, Malaguarnera M, Bertino G, Berretta M. Serum markers of intrahepatic cholangiocarcinoma. Dis Markers. 2013;34(4):219–28.
Wannhoff A, et al. FUT2 and FUT3 genotype determines CA19-9 cut-off values for detection of cholangiocarcinoma in patients with primary sclerosing cholangitis. J Hepatol. 2013;59(6):1278–84.
Khan SA, et al. Guidelines for the diagnosis and treatment of cholangiocarcinoma: an update. Gut. 2012;61(12):1657–69.
Wadsworth CA, Lim A, Taylor-Robinson SD, Khan SA. The risk factors and diagnosis of cholangiocarcinoma. Hepatol Int. 2013;7(2):377–93.
Dumonceau J-M, Delhaye M, Charette N, Farina A. Challenging biliary strictures: pathophysiological features, differential diagnosis, diagnostic algorithms, and new clinically relevant biomarkers - part 1. Ther Adv Gastroenterol. 2020;13:1756284820927292.
Levy C, Lymp J, Angulo P, Gores GJ, Larusso N, Lindor KD. The value of serum CA 19-9 in predicting cholangiocarcinomas in patients with primary sclerosing cholangitis. Dig Dis Sci. 2005;50(9):1734–40.
He X-D, et al. The risk of carcinogenesis in congenital choledochal cyst patients: an analysis of 214 cases. Ann Hepatol. 2014;13(6):819–26.
Galli C, Basso D, Plebani M. CA 19-9: handle with care. Clin Chem Lab Med. 2013;51(7):1369–83.
Beauchemin N, Arabzadeh A. Carcinoembryonic antigen-related cell adhesion molecules (CEACAMs) in cancer progression and metastasis. Cancer Metastasis Rev. 2013;32(3–4):643–71.
Fang T, Wang H, Wang Y, Lin X, Cui Y, Wang Z. Clinical significance of preoperative serum CEA, CA125, and CA19-9 levels in predicting the Resectability of Cholangiocarcinoma. Dis Markers. 2019;2019:6016931.
Rule AH, Goleski-Reilly C, Sachar DB, Vandevoorde J, Janowitz HD. Circulating carcinoembryonic antigen (CEA): relationship to clinical status of patients with inflammatory bowel disease. Gut. 1973;14(11):880–4.
Felder M, et al. MUC16 (CA125): tumor biomarker to cancer therapy, a work in progress. Mol Cancer. 2014;13:129.
Moss EL, Hollingworth J, Reynolds TM. The role of CA125 in clinical practice. J Clin Pathol. 2005;58(3):308–12.
Lapitz A, et al. Extracellular vesicles in hepatobiliary malignancies. Front Immunol. 2018;9:2270.
Yanez-Mo M, et al. Biological properties of extracellular vesicles and their physiological functions. J Extracell Vesicles. 2015;4:27066.
Caby M-P, Lankar D, Vincendeau-Scherrer C, Raposo G, Bonnerot C. Exosomal-like vesicles are present in human blood plasma. Int Immunol. 2005;17(7):879–87.
Pisitkun T, Shen R-F, Knepper MA. Identification and proteomic profiling of exosomes in human urine. Proc Natl Acad Sci U S A. 2004;101(36):13368 LP–13373.
Masyuk AI, et al. Biliary exosomes influence cholangiocyte regulatory mechanisms and proliferation through interaction with primary cilia. Am J Physiol Gastrointest Liver Physiol. 2010;299(4):G990–9.
Ogawa Y, et al. Proteomic analysis of two types of exosomes in human whole saliva. Biol Pharm Bull. 2011;34(1):13–23.
Li X, Wang X. The emerging roles and therapeutic potential of exosomes in epithelial ovarian cancer. Mol Cancer. 2017;16(1):92.
Hirsova P, et al. Extracellular vesicles in liver pathobiology: small particles with big impact. Hepatology. 2016;64(6):2219–33.
Gonzalez E, Falcon-Perez JM. Cell-derived extracellular vesicles as a platform to identify low-invasive disease biomarkers. Expert Rev Mol Diagn. 2015;15(7):907–23.
Raposo G, Stoorvogel W. Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol. 2013;200(4):373–83.
Xie F, Feng S, Yang H, Mao Y. Extracellular vesicles in hepatocellular cancer and cholangiocarcinoma. Ann Transl Med. 2019;7(5):86.
Arbelaiz A, et al. Serum extracellular vesicles contain protein biomarkers for primary sclerosing cholangitis and cholangiocarcinoma. Hepatology. 2017;66(4):1125–43.
Lapitz A, et al. Patients with Cholangiocarcinoma present specific RNA profiles in serum and urine extracellular vesicles mirroring the tumor expression: novel liquid biopsy biomarkers for disease diagnosis. Cell. 2020;9(3):721.
Julich-Haertel H, et al. Cancer-associated circulating large extracellular vesicles in cholangiocarcinoma and hepatocellular carcinoma. J Hepatol. 2017;67(2):282–92.
Olaizola P, et al. MicroRNAs and extracellular vesicles in cholangiopathies. Biochim Biophys Acta Mol basis Dis. 2018;1864(4 Pt B):1293–307.
Andersen RF, Jakobsen A. Screening for circulating RAS/RAF mutations by multiplex digital PCR. Clin Chim Acta. 2016;458:138–43.
Liang Z, Liu X, Zhang Q, Wang C, Zhao Y. Diagnostic value of microRNAs as biomarkers for cholangiocarcinoma. Dig Liver Dis. 2016;48(10):1227–32.
Zhou J, Liu Z, Yang S, Li X. Identification of microRNAs as biomarkers for cholangiocarcinoma detection: a diagnostic meta-analysis. Clin Res Hepatol Gastroenterol. 2017;41(2):156–62.
Kumarswamy R, Volkmann I, Thum T. Regulation and function of miRNA-21 in health and disease. RNA Biol. 2011;8(5):706–13.
Correa-Gallego C, et al. Circulating plasma levels of MicroRNA-21 and MicroRNA-221 are potential diagnostic markers for primary intrahepatic Cholangiocarcinoma. PLoS One. 2016;11(9):e0163699.
Huang C-S, et al. Increased expression of miR-21 predicts poor prognosis in patients with hepatocellular carcinoma. Int J Clin Exp Pathol. 2015;8(6):7234–8.
Wang L-J, et al. Serum miR-26a as a diagnostic and prognostic biomarker in cholangiocarcinoma. Oncotarget. 2015;6(21):18631–40.
Wu X, et al. Profiling of downregulated blood-circulating miR-150-5p as a novel tumor marker for cholangiocarcinoma. Tumour Biol. 2016;37(11):15019–29.
Silakit R, et al. Circulating miR-192 in liver fluke-associated cholangiocarcinoma patients: a prospective prognostic indicator. J Hepatobiliary Pancreat Sci. 2014;21(12):864–72.
Cheng Q, et al. Circulating miR-106a is a novel prognostic and lymph node metastasis Indicator for Cholangiocarcinoma. Sci Rep. 2015;5(1):16103.
Bernuzzi F, et al. Serum microRNAs as novel biomarkers for primary sclerosing cholangitis and cholangiocarcinoma. Clin Exp Immunol. 2016;185(1):61–71.
Voigtlander T, et al. MicroRNAs in serum and bile of patients with primary sclerosing cholangitis and/or Cholangiocarcinoma. PLoS One. 2015;10(10):e0139305.
Elazezy M, Joosse SA. Techniques of using circulating tumor DNA as a liquid biopsy component in cancer management. Comput Struct Biotechnol J. 2018;16:370–8.
Glenn TC. Field guide to next-generation DNA sequencers. Mol Ecol Resour. 2011;11(5):759–69.
Giachelli CM, Steitz S. Osteopontin: a versatile regulator of inflammation and biomineralization. Matrix Biol. 2000;19(7):615–22.
Wai PY, Kuo PC. Osteopontin: regulation in tumor metastasis. Cancer Metastasis Rev. 2008;27(1):103–18.
Loosen SH, et al. Elevated levels of circulating osteopontin are associated with a poor survival after resection of cholangiocarcinoma. J Hepatol. 2017;67(4):749–57.
O’Hara SP, Splinter PL, Trussoni CE, Gajdos GB, Lineswala PN, LaRusso NF. Cholangiocyte N-Ras protein mediates lipopolysaccharide-induced interleukin 6 secretion and proliferation. J Biol Chem. 2011;286(35):30352–60.
Tabibian JH, O’Hara SP, Splinter PL, Trussoni CE, LaRusso NF. Cholangiocyte senescence by way of N-ras activation is a characteristic of primary sclerosing cholangitis. Hepatology. 2014;59(6):2263–75.
Tabibian JH, Trussoni CE, O’Hara SP, Splinter PL, Heimbach JK, LaRusso NF. Characterization of cultured cholangiocytes isolated from livers of patients with primary sclerosing cholangitis. Lab Investig. 2014;94(10):1126–33.
Kumari N, Dwarakanath BS, Das A, Bhatt AN. Role of interleukin-6 in cancer progression and therapeutic resistance. Tumour Biol. 2016;37(9):11553–72.
Cheon YK, et al. Diagnostic utility of interleukin-6 (IL-6) for primary bile duct cancer and changes in serum IL-6 levels following photodynamic therapy. Am J Gastroenterol. 2007;102(10):2164–70.
Wang C-Q, et al. Interleukin-6 enhances cancer stemness and promotes metastasis of hepatocellular carcinoma via up-regulating osteopontin expression. Am J Cancer Res. 2016;6(9):1873–89.
Kimawaha P, Jusakul A, Junsawang P, Loilome W, Khuntikeo N, Techasen A. Circulating TGF-β1 as the potential epithelial mesenchymal transition-biomarker for diagnosis of cholangiocarcinoma. J Gastrointest Oncol. 2020;11(2):304–18.
Chapman MH, et al. Circulating CYFRA 21-1 is a specific diagnostic and prognostic biomarker in biliary tract cancer. J Clin Exp Hepatol. 2011;1(1):6–12.
Edelman MJ, et al. CYFRA 21-1 as a prognostic and predictive marker in advanced non-small-cell lung cancer in a prospective trial: CALGB 150304. J Thorac Oncol. 2012;7(4):649–54.
Huang L, et al. Serum CYFRA 21-1 in biliary tract cancers: a reliable biomarker for gallbladder carcinoma and intrahepatic Cholangiocarcinoma. Dig Dis Sci. 2015;60(5):1273–83.
Uenishi T, et al. Serum cytokeratin 19 fragment (CYFRA21-1) as a prognostic factor in intrahepatic cholangiocarcinoma. Ann Surg Oncol. 2008;15(2):583–9.
Cuenco J, et al. Identification of a serum biomarker panel for the differential diagnosis of cholangiocarcinoma and primary sclerosing cholangitis. Oncotarget. 2018;9(25):17430–42.
Vairaktaris E, et al. High gene expression of matrix metalloproteinase-7 is associated with early stages of oral cancer. Anticancer Res. 2007;27(4B):2493–8.
Štrbac D, Goričar K, Dolžan V, Kovač V. Evaluation of matrix metalloproteinase 9 serum concentration as a biomarker in malignant mesothelioma. Dis Markers. 2019;2019:1242964.
Nanda DP, Sil H, Moulik S, Biswas J, Mandal SS, Chatterjee A. Matrix metalloproteinase-9 as a potential tumor marker in breast cancer. J Environ Pathol Toxicol Oncol. 2013;32(2):115–29.
Lawicki S, Glazewska EK, Sobolewska M, Bedkowska GE, Szmitkowski M. Plasma levels and diagnostic utility of macrophage Colony-stimulating factor, matrix Metalloproteinase-9, and tissue inhibitor of Metalloproteinases-1 as new biomarkers of breast cancer. Ann Lab Med. 2016;36(3):223–9.
Leelawat K, Sakchinabut S, Narong S, Wannaprasert J. Detection of serum MMP-7 and MMP-9 in cholangiocarcinoma patients: evaluation of diagnostic accuracy. BMC Gastroenterol. 2009;9:30.
Onsurathum S, et al. Proteomics detection of S100A6 in tumor tissue interstitial fluid and evaluation of its potential as a biomarker of cholangiocarcinoma. Tumour Biol. 2018;40(4):1010428318767195.
Shi R-Y, et al. High expression of Dickkopf-related protein 1 is related to lymphatic metastasis and indicates poor prognosis in intrahepatic cholangiocarcinoma patients after surgery. Cancer. 2013;119(5):993–1003.
Shen J, et al. Comparative proteomic profiling of human bile reveals SSP411 as a novel biomarker of cholangiocarcinoma. PLoS One. 2012;7(10):e47476–6.
Xu H, et al. Elevation of serum KL-6 mucin levels in patients with cholangiocarcinoma. Hepato-Gastroenterology. 2008;55(88):2000–4.
Li Y, et al. Application of joint detection of AFP, CA19-9, CA125 and CEA in identification and diagnosis of Cholangiocarcinoma. Asian Pac J Cancer Prev. 2015;16(8):3451–5.
Zhang Y, Yang J, Li H, Wu Y, Zhang H, Chen W. Tumor markers CA19-9, CA242 and CEA in the diagnosis of pancreatic cancer: a meta-analysis. Int J Clin Exp Med. 2015;8(7):11683–91.
Tao L-Y, Cai L, He X-D, Liu W, Qu Q. Comparison of serum tumor markers for intrahepatic cholangiocarcinoma and hepatocellular carcinoma. Am Surg. 2010;76(11):1210–3.
Wongkham S, et al. Clinical significance of serum total sialic acid in cholangiocarcinoma. Clin Chim Acta. 2003;327(1–2):139–47.
Wongkham S, Boonla C, Kongkham S, Wongkham C, Bhudhisawasdi V, Sripa B. Serum total sialic acid in cholangiocarcinoma patients: an ROC curve analysis. Clin Biochem. 2001;34(7):537–41.
Kongtawelert P, Tangkijvanich P, Ong-Chai S, Poovorawan Y. Role of serum total sialic acid in differentiating cholangiocarcinoma from hepatocellular carcinoma. World J Gastroenterol. 2003;9(10):2178–81.
Liang Q, Liu H, Zhang T, Jiang Y, Xing H, Zhang H. Serum metabolomics uncovering specific metabolite signatures of intra- and extrahepatic cholangiocarcinoma. Mol BioSyst. 2016;12(2):334–40.
Sun Y-F, et al. Circulating stem cell-like epithelial cell adhesion molecule-positive tumor cells indicate poor prognosis of hepatocellular carcinoma after curative resection. Hepatology. 2013;57(4):1458–68.
Arnoletti JP, et al. Pancreatic and bile duct cancer circulating tumor cells (CTC) form immune-resistant multi-cell type clusters in the portal venous circulation. Cancer Biol Ther. 2018;19(10):887–97.
Tan CRC, Zhou L, El-Deiry WS. Circulating tumor cells versus circulating tumor DNA in colorectal cancer: Pros and Cons. Curr Colorectal Cancer Rep. 2016;12(3):151–61.
Arnoletti JP, et al. Portal venous blood circulation supports immunosuppressive environment and pancreatic cancer circulating tumor cell activation. Pancreas. 2017;46(1):116–23.
Al Ustwani O, Iancu D, Yacoub R, Iyer R. Detection of circulating tumor cells in cancers of biliary origin. J Gastrointest Oncol. 2012;3(2):97–104.
Intuyod K, Armartmuntree N, Jusakul A, Sakonsinsiri C, Thanan R, Pinlaor S. Current omics-based biomarkers for cholangiocarcinoma. Expert Rev Mol Diagn. 2019;19(11):997–1005.
Son KH, Ahn CB, Kim HJ, Kim JS. Quantitative proteomic analysis of bile in extrahepatic cholangiocarcinoma patients. J Cancer. 2020;11(14):4073–80.
Park JY, Park BK, Ko JS, Bang S, Song SY, Chung JB. Bile acid analysis in biliary tract cancer. Yonsei Med J. 2006;47(6):817–25.
Nagana Gowda GA, Shanaiah N, Cooper A, Maluccio M, Raftery D. Bile acids conjugation in human bile is not random: new insights from (1)H-NMR spectroscopy at 800 MHz. Lipids. 2009;44(6):527–35.
Sharif AW, et al. Metabolic profiling of bile in cholangiocarcinoma using in vitro magnetic resonance spectroscopy. HPB (Oxford). 2010;12(6):396–402.
Albiin N, et al. Detection of cholangiocarcinoma with magnetic resonance spectroscopy of bile in patients with and without primary sclerosing cholangitis. Acta Radiol. 2008;49(8):855–62.
Alvaro D, et al. Serum and biliary insulin-like growth factor I and vascular endothelial growth factor in determining the cause of obstructive cholestasis. Ann Intern Med. 2007;147(7):451–9.
Chiang K-C, et al. Lipocalin 2 (LCN2) is a promising target for cholangiocarcinoma treatment and bile LCN2 level is a potential cholangiocarcinoma diagnostic marker. Sci Rep. 2016;6:36138.
Tye BK. MCM proteins in DNA replication. Annu Rev Biochem. 1999;68:649–86.
Ren B, et al. MCM7 amplification and overexpression are associated with prostate cancer progression. Oncogene. 2006;25(7):1090–8.
Kim D-W, et al. Transcriptional induction of minichromosome maintenance protein 7 (Mcm7) in human cholangiocarcinoma cells treated with Clonorchis sinensis excretory-secretory products. Mol Biochem Parasitol. 2010;173(1):10–6.
Ayaru L, et al. Diagnosis of pancreaticobiliary malignancy by detection of minichromosome maintenance protein 5 in bile aspirates. Br J Cancer. 2008;98(9):1548–54.
Chen C-Y, Tsai W-L, Wu H-C, Syu M-J, Wu C-C, Shiesh S-C. Diagnostic role of biliary pancreatic elastase for cholangiocarcinoma in patients with cholestasis. Clin Chim Acta. 2008;390(1–2):82–9.
Koopmann J, et al. Mac-2-binding protein is a diagnostic marker for biliary tract carcinoma. Cancer. 2004;101(7):1609–15.
Severino V, et al. Extracellular vesicles in bile as markers of malignant biliary Stenoses. Gastroenterology. 2017;153(2):495–504.e8.
Li L, et al. Human bile contains microRNA-laden extracellular vesicles that can be used for cholangiocarcinoma diagnosis. Hepatology. 2014;60(3):896–907.
Plieskatt J, et al. A microRNA profile associated with Opisthorchis viverrini-induced cholangiocarcinoma in tissue and plasma. BMC Cancer. 2015;15:309.
Shin S-H, et al. Bile-based detection of extrahepatic cholangiocarcinoma with quantitative DNA methylation markers and its high sensitivity. J Mol Diagn. 2012;14(3):256–63.
Lesurtel M, et al. Platelet-derived serotonin mediates liver regeneration. Science. 2006;312(5770):104–7.
Alpini G, et al. Serotonin metabolism is dysregulated in cholangiocarcinoma, which has implications for tumor growth. Cancer Res. 2008;68(22):9184–93.
Smith ZL, Guzzo TJ. Urinary markers for bladder cancer. F1000Prime Rep. 2013;5:21.
Morrissey JJ, London AN, Luo J, Kharasch ED. Urinary biomarkers for the early diagnosis of kidney cancer. Mayo Clin Proc. 2010;85(5):413–21.
Cartlidge CR, U Abellona MR, Alkhatib AMA, Taylor-Robinson SD. The utility of biomarkers in hepatocellular carcinoma: review of urine-based (1)H-NMR studies - what the clinician needs to know. Int J Gen Med. 2017;10:431–42.
Radon TP, et al. Identification of a three-biomarker panel in urine for early detection of pancreatic adenocarcinoma. Clin Cancer Res. 2015;21(15):3512–21.
Jing J, Gao Y. Urine biomarkers in the early stages of diseases: current status and perspective. Discov Med. 2018;25(136):57–65.
Navaneethan U, et al. Volatile organic compounds in urine for noninvasive diagnosis of malignant biliary strictures: a pilot study. Dig Dis Sci. 2015;60(7):2150–7.
Metzger J, et al. Urine proteomic analysis differentiates cholangiocarcinoma from primary sclerosing cholangitis and other benign biliary disorders. Gut. 2013;62(1):122–30.
Voigtländer T, et al. Bile and urine peptide marker profiles: access keys to molecular pathways and biological processes in cholangiocarcinoma. J Biomed Sci. 2020;27(1):13.
Borad MJ, et al. Integrated genomic characterization reveals novel, therapeutically relevant drug targets in FGFR and EGFR pathways in sporadic intrahepatic cholangiocarcinoma. PLoS Genet. 2014;10(2):e1004135.
Zou S, et al. Mutational landscape of intrahepatic cholangiocarcinoma. Nat Commun. 2014;5:5696.
Wang P, et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene. 2013;32(25):3091–100.
Nakamura H, et al. Genomic spectra of biliary tract cancer. Nat Genet. 2015;47(9):1003–10.
Nepal C, et al. Genomic perturbations reveal distinct regulatory networks in intrahepatic cholangiocarcinoma. Hepatology. 2018;68(3):949–63.
Andresen K, et al. Four DNA methylation biomarkers in biliary brush samples accurately identify the presence of cholangiocarcinoma. Hepatology. 2015;61(5):1651–9.
Limpaiboon T, et al. Promoter hypermethylation is a major event of hMLH1 gene inactivation in liver fluke related cholangiocarcinoma. Cancer Lett. 2005;217(2):213–9.
Vedeld HM, et al. The novel colorectal cancer biomarkers CDO1, ZSCAN18 and ZNF331 are frequently methylated across gastrointestinal cancers. Int J Cancer. 2015;136(4):844–53.
Sia D, et al. Integrative molecular analysis of intrahepatic cholangiocarcinoma reveals 2 classes that have different outcomes. Gastroenterology. 2013;144(4):829–40.
Ghidini M, et al. Characterisation of the immune-related transcriptome in resected biliary tract cancers. Eur J Cancer. 2017;86:158–65.
Sawada R, et al. Interleukin-33 overexpression reflects less aggressive tumour features in large-duct type cholangiocarcinomas. Histopathology. 2018;73(2):259–72.
Ruys AT, Groot Koerkamp B, Wiggers JK, Klumpen H-J, ten Kate FJ, van Gulik TM. Prognostic biomarkers in patients with resected cholangiocarcinoma: a systematic review and meta-analysis. Ann Surg Oncol. 2014;21(2):487–500.
Suzuki H, et al. Relationship between 18-F-fluoro-deoxy-D-glucose uptake and expression of glucose transporter 1 and pyruvate kinase M2 in intrahepatic cholangiocarcinoma. Dig Liver Dis. 2015;47(7):590–6.
Selaru FM, et al. MicroRNA-21 is overexpressed in human cholangiocarcinoma and regulates programmed cell death 4 and tissue inhibitor of metalloproteinase 3. Hepatology. 2009;49(5):1595–601.
He Q, et al. Ars2 is overexpressed in human cholangiocarcinomas and its depletion increases PTEN and PDCD4 by decreasing microRNA-21. Mol Carcinog. 2013;52(4):286–96.
Collins AL, et al. A differential microRNA profile distinguishes cholangiocarcinoma from pancreatic adenocarcinoma. Ann Surg Oncol. 2014;21(1):133–8.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this chapter
Cite this chapter
García-Sampedro, A., Acedo, P., Pereira, S.P. (2021). Established and Emerging Biomarkers for Prediction, Early Detection, and Prognostication of Cholangiocarcinoma. In: Tabibian, J.H. (eds) Diagnosis and Management of Cholangiocarcinoma. Springer, Cham. https://doi.org/10.1007/978-3-030-70936-5_19
Download citation
DOI: https://doi.org/10.1007/978-3-030-70936-5_19
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-70935-8
Online ISBN: 978-3-030-70936-5
eBook Packages: MedicineMedicine (R0)