Abstract
Black scurf/stem canker disease, caused by the basidiomycetous fungus Rhizoctonia solani Kühn, became one of the major constraints to potato production worldwide. R. solani isolates of AG-3 are considered the main causal organism of black scurf, characterized by the presence of sclerotial bodies on the surface of potato tubers. R. solani limits the potato plants growth by developing cankers on sprouts, stems and tubers which make tubers ugly due to the appearance of corky spots and elephant hide symptoms on the tubers. To stop the establishment of disease, early detection and precise identification of pathogens are important components of an integrated disease management system. The present review summarizes the current knowledge about symptomology and epidemiology of black scurf, methods for early and accurate detection of black scurf pathogen/s, and molecular basis of potato–R. solani interaction. Elaborative and up-to-date information on various management options including cultural, chemical, biological, genetic manipulation and nanotechnological approaches and their effectiveness for managing black scurf are discussed. Genetic approaches that show promise for the control of black scurf include the development of transgenic lines by overexpressing or silencing pathogenesis-related (PR) genes and genome editing to develop lines with lower susceptibility to the disease is discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Potato (Solanum tuberosum L.) is an important non-grain vegetable food crop and ranked fourth after maize, paddy and wheat in total production and consumption (Lal et al. 2019). Worldwide total potato production was estimated at 359.07 million tonnes in 2020 and India placed second after China with 51.3 million tons (FAOSTAT 2022). The world population increasing exponentially is putting further pressure on agricultural lands, water and other resources. Therefore, farmers have to increase their output to feed the growing population (Chaudhary et al. 2020a). In that scenario, potatoes have great importance in the global food system, strengthening global food security and alleviating poverty. The potato crop is susceptible to various fungal, bacterial, viral diseases and many other disorders (Chaudhary et al. 2020b). Among the fungal diseases, black scurf/stem canker caused by the ubiquitous fungus Rhizoctonia solani Kühn (teleomorph: Thanatephous cucumeris Frank (Donk)) is a serious problem in various potato growing regions in the world, including India. Rhizoctonia diseases in potatoes can cause a reduction in yield as well as quality (Das et al. 2014). The quantitative yield losses resulted from the infection of stems and underground portions that reduce the size and number of potato tubers (Carling et al. 1989). In contrast, qualitative losses occur mainly owing to mishappening of the tubers and sclerotial formation on the tuber surface (James and McKenzie 1972). The estimated yield loss due to Rhizoctonia diseases was reported up to 25% in India (Sharma 2015), 30% in Canada and 50% in other countries (Woodhall et al. 2008). The marketable yield loss of potatoes due to Rhizoctonia spp. reached up to 30% (Tsror 2010). Black scurf/stem canker disease of potatoes is difficult to control due to the prolonged survivability of the fungus as a dormant structure called sclerotia and its wide host range. In the present review, we provide up-to-date information regarding black scurf/stem canker disease caused by Rhizoctonia spp. Information is arranged under headings: pathogen, disease symptoms, anastomosis groups (AGs), detection & diagnosis and current control practices. The information can be useful for the better management of the disease.
The pathogen
The soil-borne pathogen Rhizoctonia solani AG-3 is considered the main causal organism of Rhizoctonia diseases of potatoes (Banville et al. 1996; Virgen-Callaros et al. 2000). Based on rDNA internal transcribed spacer (ITS) sequences variation the members of AG-3 were divided into three subgroups: AG-3PT (potato type), AG-3TB (tobacco type) and AG-3TM (tomato type) (Kuninaga et al. 2000; Misawa and Kuninaga 2010). R. solani is unable to produce asexual structures and exists in the form of mycelia, sclerotia, or basidiospores (sexual spores) (Keijer 1996). Anamorphic classification of Rhizoctonia spp. is based on the cell’s nuclear condition (multi, bi, or uni-nucleate) and the ability of hyphal anastomosis with tester strains of designated anastomosis groups (AGs) (Sneh et al. 1991). To date, thirteen AGs (AG1-AG13) and AGB1 have been identified based on phenotypic and genotypic characteristics including cultural, morphological, host range, virulence, nutritional requirements, molecular and biochemical characteristics (González et al. 2016). Generally, on potato dextrose agar (PDA) medium R. solani AG-3 isolates grew slower than AG1-I subgroup isolates (Chaudhary et al. 2023a) with whitish to light brown mycelial colour during early growth which turn brown as colony become aged (Fig. 1a). Microscopically, hyphal branch originates from distal dolipore septum with a characteristic constriction at the branching point (Ajayi-Oyetunde and Bradley 2018; Chaudhary et al. 2023b).
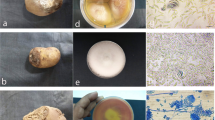
Symptoms and causal organism of black scurf and stem canker disease of potato (a) Culture petriplate of Rhizoctonia solani, the causal organism of black scurf/stem canker disease; (b) Characteristic symptom of black scurf; (c) Corky spot symptom/ Eyeless or blind tuber; (d) Elephant hide symptom; (e) Damage on underground main stem; (f) Rhizoctonia lesions on stolons (g) Stunting and rosettingof plant tops; curled leaves; (h) White sheet of mycelia on stem; (i) Formation of Aerial tubers
Disease symptoms
Shortly after planting, necrosis on germinating sprouts are the typical symptoms of stem canker which results in the late emergence of potato plants in the field. Black scurf symptoms (Fig. 1b) appear later in the cropping season when sclerotial bodies start to cover the progeny potato tubers (Banville 1989). Generally, R. solani does not penetrate or damage the potato tubers; however, tuber mishappening may occur (Weinhold et al. 1982). Additionally, in severe infection, atypical symptoms including cracking, corky lesions, and elephant hide may also be observed on tubers (Campion et al. 2003; Muzhinji et al. 2014) (Fig. 1c, d). Reddish-brown to sunken grey lesions are formed on the newly developing sprouts, resulting girdle the young sprout (Fig. 1e). Below the affected area secondary sprouts are formed and if these secondary sprouts also infected, tertiary sprouts may be developed from non-affected lower buds. This process may be repeated several times. Sprouts will fail to emerge or wilt after emergence, resulting in uneven or irregular emergence and in severe cases may lead to a poor crop stand. Reddish-brown to brown lesions appear underneath the stems and on the stolons (Fig. 1f). As these lesions mature, they become rough and brown cankers and have craters, cracks or cracks or both (Banville 1978).
Infection of the stem causes stunting and rosetting of plant tops resulting in curling of the upper leaves which sometimes turn red or yellow (Fig. 1g) (Wharton et al. 2007). In a recent study, Ito et al. (2017) observed that leaf curling is not a direct symptom of Rhizoctonia. Still, prior infection with the Potato leaf roll virus enhanced the severity of Rhizoctonia diseases. At the base of stems and on the plant parts that are in contact with soil a greyish-white, felt-like mycelium mat can be observed (Fig. 1h), which is caused by the perfect stage (T. cucumeris) of the fungus (Banville and Carling 2001). Aerial tubers could be formed in the leaf axils of stems due to the interference of carbohydrate movement (Beukema and van der Zang 1990). These are green to reddish-purple round to bottle-shaped transformations of lateral shoots in the axils, with a few small leaves at the top (Fig. 1i).
Occurrence of R. solani anastomosis group (AGs) in potato
Rhizoctonia solani AG-3 is considered the most prevalent AG causing black scurf/stem canker in potatoes (Woodhall et al. 2007; Lehtonen et al. 2008a). However, other AGs such as AG2-1 (Woodhall et al. 2007; Lehtonen et al. 2008a), AG4 (Virgen-Calleros et al. 2000), AG4 HG-I, AG4 HG-III (Muzhinji et al. 2014; 2015), AG4 HG-II (Woodhall et al. 2012) and AG5 (Bandy et al. 1988), AG-8 (Balali et al. 1995), AG-9 (Yanar et al. 2005) have also been reported in potato fields at a lower frequency around the world. Besides, binucleate Rhizoctonia (BNR) isolates were also recovered from potato plants (Carling et al. 1986a). Farrokhi-Nejad et al. (2007) collected 12 BNR isolates (out of 58) that cause mild symptoms in potato sprouts. BNR AG A and AG R causing stem canker, black scurf and tuber defects were reported in South Africa (Muzhinji et al. 2015; Zimudzi et al. 2017). Recently, Shuai et al. (2022) reported that AG2-2IV causes black scurf in potatoes in Heilongjiang province, China.
Disease cycle and epidemiology
Rhizoctonia infection on potato crops can be initiated through seed-borne or soil-borne inoculums viz., either sclerotia or runner hyphae from the plant debris. R. solani may survive as dormant sclerotia for over the years in soil and stubbles and can re-infect healthy potato plants in the subsequent crop season (Keijer et al. 1996) (Fig. 2). After successful attachment of vegetative growing hyphae to the surface of plant the T-shaped branches are formed within 12 h (Lehtonen et al. 2008b). R. solani enters the plant tissue and produces RS toxin, a mixture that includes N-acetyl glucosamine, N-acetyl galactosamine, glucose and mannose (Vidhyasekaran et al. 1997) along with pathogen effectors (Zheng et al. 2013). Penetration into the epidermis and cortex takes place with lobate appressoria or infection cushion or both from which the infection peg grows and enters the host (Marshall and Rush 1980; Singh and Subramanian 2017). Further, inter- or intra-cellular growth of mycelium triggers extracellular enzyme secretion, resulting in the infected tissue’s collapse and forming brownish lesions called stem canker (Banville et al. 1996). This condition develops mainly before the formation of daughter tubers. The mycelium continues to grow on stolons and roots and develops sclerotial structures on them, which stimulated the senescence at the end of the growing season. Consequently, sclerotia are developed on daughter tubers known as black scurf. At the maturity of the potato crop, sclerotia remaining in the soil serve as the source of primary inoculum which infects the host plants in the next growing season (Scholte 1989). Environmental conditions like temperature and relative humidity are important for the infection and initiation of Rhizoctonia disease in potatoes. Low temperatures with high soil moisture and neutral to acidic soil (pH 7 or less) are favourable for stem cankers. Initiation of sclerotia formation on daughter tubers started late in the cropping season, mainly after harm cutting, but sclerotia can be seen at mid of the cropping season.
Molecular mechanism of Potato - R. solani AG3 interaction
Presently, there is limited information about the molecular responses of R. solani AG3-PT during pathogenic interaction with potato plants (Zrenner et al. 2020). The published complete whole genome sequence assemblies of AG-3 R. solani isolates (Cubeta et al. 2014; Wibberg et al. 2017; Patil et al. 2017) and potatoes (Xu et al. 2011) can be utilized for understanding the key mechanisms of R. solani infection and disease development (Table 1).
There is some information on phytotoxins (3-methylthiopropionic acid (3-MTPA) and 3-methylthioacrylic acid (3-MTAA)) produced by R. solani AG3 with relation to disease symptoms, and concentration of the phytotoxin was correlated with pathogenicity (Kankam et al. 2016a, b). In rice sheath blight, three potential secreted effectors (such as glycosyltransferase, cytochrome C oxidase CtaG/cox11 and peptidase inhibitor I9) correlate with the virulence of R. solani AG1-IA (Zheng et al. 2013). In a study, Rioux et al. (2011) isolated and compared ESTs from AG1-IA infected rice leaves and AG-3 infected potato sprouts. Of 25 mRNAs from AG1-IA and AG3 showed significant similarity, 12 were associated with the pathogenesis processes. The six putative pathogenesis-related genes viz., pyruvate carboxylase (PC), ABC-transporter (ABC), glycosyl-transferase (GTF), kappa-family glutathione-S-transferase (GLU), Rab-type GTPase (RAB), and Nic96-type nucleoporin (NIC) had similar expression patterns in the AG1/rice and AG3/potato pathosystems. However, expression patterns of the putative AAA-type ATPase gene (AAA) and MFS were quite different between AG1 and AG3 which underscores the potential differences in R. solani pathogenesis mechanisms utilized in these two pathosystems (Subterranean vs. Foliar). In a recent transcriptomic analysis of AG3- potato interaction, various genes transcribed proteins with diverse hydrolase and peptidase activities have been predicted that were expressed differentially with due course of time response (Zrenner et al. 2020), while an additional increase of expression of hydrolases and genes coding various integral membrane proteins with transporter function was lined to interaction progression.
R. solani produces phenylacetic acid (PAA), three hydroxy (OH-) and a 3-methoxy (3-MeO-) derivative of PAA, which are important in the parasitism and infection process in plants (Mandava et al. 1980). In a study, Bartz et al. (2012) demonstrated the involvement of the PAA metabolic complex in Rhizoctonia disease development in tomatoes and also suggested that the production of these compounds is not the primary or the only determinant of pathogenicity. Reactive Oxygen Species (ROS) are very active and highly toxic to biological molecules and important growth regulators which are involved in limiting pathogen spread, induction of cell death and cell signalling in host-plant interactions (Torres 2010; Barna et al. 2012). It was also reported that fungi also produce ROS during pathogenic interactions (Daub and Ehrenshaft 2000; Samsatly et al. 2018). Therefore, the regulation of ROS in fungal cells and tolerance to external ROS produced by the host plant represent a balanced control and detoxification by both partners which can govern the fate of disease development (Heller and Tudzynski 2011). To maintain this balance, plant and fungal cells possess a complex array of protective mechanisms such as oxalic acid or the NADPH oxidase RBohD (Torres et al. 2005; Kadota et al. 2015), or ROS-quenching molecules including vitamin B6 (VB6) and various antioxidant enzymes such as catalase (CAT), superoxide dismutase (SOD), ascorbate peroxidase (APX), glutathione-S-transferase (GST), and glutathione reductase (Zhang et al. 2015; Girard et al. 2017). Little information are reported on ROS scavenging systems in R. solani and to date the up-regulation of R. solani genes particularly, PYRIDOXALREDUCTASE (PLR AKR8; DW520695) and PYRIDOXAL-5-PHOSPHATASES and TRANSAMINASES of the vitamin B6 (VB6) salvage biosynthetic pathway was reported in R. solani hyphae in association with a mycoparasite or an antagonistic bacterium, respectively (Chamoun and Jabaji 2011; Gkarmiri et al. 2015). Samsatly et al. (2015) characterized two genes of the de-novo VB6 biosynthetic pathway; RsolPDX1 (KF620111.1) and RsolPDX2 (KF620112.1), and one gene RsolPLR (KJ395592.1) of the VB6 salvage biosynthetic pathway of AG3. Recently, in a study Samsatly et al. (2020) provided indirect evidence on the functionality of RsolPDX1 and RsolPDX2 of AG3 and their involvement in VB6 de-novo biosynthesis pathway of the yeast Saccharomyces cerevisiae and showed that the antioxidant genes encoding VB6 (i.e., PDX, PLR), CAT and, GST of AG3 and potato are differentially induced and transcriptionally regulated at the infection site (i.e., necrotic tissues, and surrounding areas) during AG3- potato sprout interaction.
To defend themselves against phytopathogens like bacteria, fungi, viruses and insect herbivores, a complex defence system is induced in plants (Glazebrook 2005). Defence mechanisms can either be performed or induced. In response to necrotrophic fungi like R. solani, defence mechanisms attributed to ethylene (ET) and jasmonic acid (JA) signalling are known to be induced but not the salicylic acid (SA) signalling which plays an important role in plant resistance against biotrophic/hemibiotrophic pathogens (Tsuda et al. 2013). Recently, Kouzai et al. (2018) reported on the discovery of SA-dependent resistance of Oryza sativa and Brachypodium distachyon towards R. solani suggesting the existence of a pseudo biotrophic phase during the interaction with these two host species. The importance of SA-mediated defences plant defences in the AG3-potato pathosystem was further underlined by Genzel et al. (2017). Currently, no qualitative resistance has been reported, it assumed that a general response to AG3- potato infection is more probable and different pathways are involved in pathogen-associated molecular patterns (PAMP)-triggered immunity (PTI), previously identified as being induced by necrotrophs (Genzel et al. 2017).
It is well documented that pathogenesis-related (PR) genes become activated by induction of the systemic acquired resistance (SAR) pathway (Sanz-Alférez et al. 2008). The PR proteins form a group of at least 17 structurally and functionally distinct protein families that are ubiquitous in plants (Liu and Ekramoddoullah 2006). The expression levels of five PR genes viz., PR-1 (1,3-β-glucanase), PR-3 (chitinase), PR-10, glutathione-S-transferase (GST) and phenylalanine ammonia-lyase (PAL) gene were found to be induced in potatoes at different time points of P. infestans infection indicating the involvement of SAR activation (Vleeshouwers et al. 2000; Gallou 2011). Further, microarray analysis revealed a systemic transcriptional induction of PR-2, PAL, and PI2 (PR-6) associated with JA and abscisic acid (ABA) pathways in potato sprouts in response to AG3- potato infection (Lehtonen et al. 2008b). The expression of PR-10 is also described to be induced by the plant hormones JA and ABA (Liu and Ekramoddoullah 2006). In addition, while attacking the host plants, several fungi produce extracellular proteinases and protease inhibitors (PIs) (Kim et al. 2005). In turn, plants synthesize proteases and PIs as a way to defend themselves and recognise fungal derived proteinases (Jashni et al. 2015). It has been reported that Solanaceous plants have high contents of proteinase inhibitors. In a study, Gvozdeva et al. (2006) revealed that potato plants synthesize various proteinases which can suppress trypsin-like extracellular proteinases of R. solani in vitro.
For R. solani-potato interaction study, another critical area is the investigation of adaptive alterations in the metabolic profile of potato plants upon R. solani AG-3 infection. Although, limited studies have been conducted to reveal the involvement of metabolic changes during the period of potato response to the R. solani attack. In a study, Aliferis and Jabaji (2012) studied the accumulation of metabolites in potato sprouts at the R. solani infection site and they observed differential accumulation of phenolics, amino acids, alkaloids, fatty acids and organic acids in infected and mock infected sprout tissue. Further, carboxylic acids and sugars were increased during AG-3 infection, whereas host cell wall precursors and protein amino acids decreased. Also, the R. solani-derived virulence factor phenylacetic acid was quantified in infected sprouts (Bartz et al. 2012).
Detection and diagnosis
Detection and diagnosis of phytopathogens in crop plants and other host plant species may be required to monitor the presence and quantitative level of the pathogen(s) in a crop for preventive and curative measures. Various methods used for R. solani detection are outlined in Fig. 3.
Conventional approaches
Plant infection or the presence of phytopathogens can be determined by the visual inspection of disease symptoms followed by isolation of putative pathogen on a suitable nutrient medium. The isolated pathogen can be identified and characterized based on microscopic observations and taxonomic characteristics. However, conventional methods are nonspecific and not so much reliable due to their inability to differentiate among closely related species (Narayanasamy 2001). Additionally, the detection of pathogens becomes more difficult when disease symptoms are indefinite, low pathogen levels, absence of fruiting bodies, latent infection, etc. (Agrios 2005). Various detection techniques i.e., bait method using susceptible host material (Weinhold 1977; Paulitz and Schroeder 2005; Spurlock et al. 2015), culture plating (Anderson and Huber 1965; Ko and Hora 1971; Vincelli and Beaupre 1989), wet sieving and direct microscopic observation (Boosalis and Scharen 1959), incubating immersion tubes in soil (Martinson 1963), wooden toothpicks (Paulitz and Schroeder 2005) and anastomosis test (Ogoshi 1987) have been developed for monitoring R. solani in soil. The addition of tannic acid (300 ppm) as a marker to water agar was useful for the isolation, identification and quantification of R. solani (Hsieh et al. 1996). Although quite effective, these methods are time-consuming, labour-intensive and require considerable knowledge of fungal taxonomy.
Molecular approaches
The advancement in molecular biology techniques has provided new insights into the detection and cataloguing the soil-borne fungal pathogens like R. solani and can identify unknown species or strains from their DNA sequences.
PCR (Polymerase chain Reaction) based
For the detection of R. solani AG1-IA (Matsumoto 2002; Lal et al. 2020), AG-1-IB (Grosch et al. 2007), AG-2 and subgroups (Salazar et al. 2000), AG-3 (Bounoua et al. 1999), AG-4 and AG-8 (Brisbane et al. 1995) PCR-based methods have been used. Bounoua et al. (1999) used restriction endonuclease, XhoI to construct PCR-based restriction map for the detection of AG-3 from plants and soil samples. Recently, Irandukunda et al. (2022) used a multiplex PCR for the rapid detection of R. solani AG-3PT from potato tubers and soil.
rRNA-ITS (Internal Transcribe Spacer) sequence-based method
The internal transcribed spacer (ITS) region of nuclear DNA (rDNA) has been widely used for evolutionary studies and phylogenetics of fungal genus (Cubeta et al. 1996). The ITS region presents in several hundred copies in the genome and each unit is comprised of three genes viz., 18S (Small Subunit Ribosomal DNA, or SSU), 5.8S and 28S (Large Subunit Ribosomal DNA, or LSU) (Capote et al. 2012). Because ITS1 and ITS2 regions have not transcribed any protein, they are less affected by evolutionary pressure and therefore, highly variable among different isolates. By analyzing sequence differences of these regions, Rhizoctonia spp. can be grouped into clades having phylogenetic relationships. The 5.8S region of R. solani rDNA gene is highly conserved. The 18S and 28S subunits are used to differentiate high taxonomic levels such as family and genera while ITS allows the characterization of organisms at the species level (Gardes and Bruns 1993). The ITS sequence (ITS1-5.8S-ITS2) database of R. solani is extensively available at the NCBI GenBank (http://www.ncbi.nlm.nih.gov/genbank) which facilitates phylogenetic analysis. The rDNA-ITS sequences available at NCBI contained a conserved 5.8S region but showed variation in ITS1 and ITS2 regions (Carling et al. 2002; Amaradasa et al. 2013). Kanetis et al. (2016) performed an rDNA-ITS sequence analysis of 68 isolates collected from potato tubers. Sequence analysis of ITS regions of rDNA confirmed the prevalence of AG3. Additionally, the phylogenetic analysis found that AG3 isolates were of the potato type, distinctly separated from the AG3 tobacco type.
Quantitative real-time PCR
Currently, the qPCR (Quantitative PCR) technique is an important advanced method used for diagnosing and detecting phytopathogens. Visual examination of Rhizoctonia disease at the early stages of infection does not provide a reliable estimation of the level of disease infection. A real-time quantitative (Q) PCR method was developed for the detection and quantification of R. solani AG3-PT DNA from tuber and soil samples. A specific primer based on the rDNA-ITS region of R. solani was designed. The assay produced amplicons with AG3-PT, and non-specific for the isolates from other AGs or subgroups of AG3 (Lees et al. 2002; Woodhall et al. 2013). Similarly, Zhao et al. (2014) designed specific primers for qPCR from rDNA-ITS region of R. solani and detected DNA as low as 100 fg from the infected tobacco tissues and soil samples. Shen et al. (2017) established the SYBR Green-I qPCR detection assay for the quantification of R. solani AG3 sclerotia from the soil. They reported that detecting sensitivity for the wet-sieving qPCR method was 10-fold higher than that of the conventional PCR.
DNA fingerprinting approaches
DNA fingerprinting is a technique that simultaneously detects many minisatellites in the genome to produce patterns unique to an organism. These methods are widely used to amplify the tandem repeats present in the random regions of the genome of an organism which helps to identify species-specific patterns when conserved genes do have not enough to differentiate the species (McCartney et al. 2003). In R. solani to assess the genetic variations among AG subgroups DNA fingerprinting techniques like Randomly amplified polymorphic DNA (RAPD) and DNA amplification fingerprinting (DAF) markers have been widely used (Stodart et al. 2007). Similarly, UP-PCR (Universally Primed PCR markers were also very resembled RAPD (Bulat et al. 1998) which differentiates R. solani isolates belonging to different AGs and different subgroups (Lübeck and Lübeck 2005). Various DNA fingerprinting assays use only based on the same principle of DNA polymerase mediated amplification of DNA fragments to generate multiple copies of target genome sites. These techniques' difference depends primarily on the design or choice of primers and the level of stringency (Patil and Solanki 2016). However, AFLP (Amplified Fragment Length Polymorphism) technique is different from the above mentioned assays (Vos et al. 1995; Lübeck and Lübeck 2005). Ceresini et al. (2002) used AFLP analysis to differentiate AG-3 isolates from potato (AG-3 PT) and tobacco (AG-3 TB). The findings revealed that analysed isolates from both hosts had distinct AFLP phenotypes. Thus, the AFLP technique has a very high discriminatory ability to facilitate intra-group variation.
SCAR (Sequence Characterized Amplified Region) approach
RAPD (Randomly amplified polymorphic DNA) markers may be used to differentiate target organisms from those of non-target organisms and unique specific bands with the target organism genomic DNA could be produced and cloned. Once the unique bands have been amplified and detected, they can be used as probes for the presence of similar DNA fragments in the related species. Further, after analysis, if the amplicons do not match, sequenced them, and species-specific SCAR (Sequence Characterized Amplified Region) markers could be developed (Ma and Michialides 2005) that selectively amplifies the marker and acts as a target site in diagnostic assays. Since R. solani is a species complex, SCAR markers are necessary for identification at strain or AG subgroup level. Grosch et al. (2007) designed and used SCAR markers to produce species-specific probes and PCR primers in R. solani.
Loop-Mediated Isothermal Amplification (LAMP) assay
LAMP technique can be performed onsite in the field, resulting in a significant reduction in time required to detect and diagnose the diseases. The LAMP method can be carried out inexpensively using simple water or heating block. A positive reaction is recognized by the accumulation of a visible product appearing as a white precipitate (Notomi et al. 2000). The products can be detected by agarose gel electrophoresis, by the use of spectrophotometry to measure turbidity (Mori et al. 2004), in RT-LAMP using intercalating fluorescent dyes (Oscorbin et al 2016), or by visualize the turbidity through naked eyes or colour changes (Iwamtoo et al. 2003; Mori et al. 2001). The LAMP method was integrated with lateral flow devices (LFDs) to improve the efficiency for in‐field detection of R. solani in plant tissues, seeds, and propagules. LAMP primers based on the internal transcribed spacer (ITS) DNA sequences were used for the detection of anastomosis groups of R. solani. The LAMP‐LFD procedure effectively detected R. solani in several infected plant species belonging to diverse families and has the potential for onsite diagnosis of R. solani in plants, seeds, propagules, and soils.
The detection limit of the LAMP‐LFD protocol (10 fg) was comparable to that of qRT‐PCR format (Patel et al. 2015). LAMP assay was utilized by Lu et al. (2015) for the detection and diagnosis of R. solani (ITS-Rs-LAMP) and Macrophomina phaseolina (ITS-Mp-LAMP) in diseased soybean tissues in the field. The detection limit of the ITS-Rs-LAMP assay was 10 pg/μl of genomic DNA, and that of the ITS-Mp-LAMP assay was 100 pg/μl of genomic DNA.
Serological diagnostics
In the serological techniques, specific antibodies are used to detect their respective antigens in the test samples. Every antibody is specific for a particular antigen and binds to it, usually foreign proteins, complex carbohydrates, polynucleotides, or lipopolysaccharides. Enzyme-Linked Immuno Sorbent Assay (ELISA) is a valuable serological technique used for plant fungal pathogens detection (Johnson et al. 1982). Polyclonal antibodies (PAbs) and monoclonal antibodies (MAbs) have been produced against fungal antigens present in whole cells, cell fractions, extracellular components, and culture filtrates. Methods of production of PAbs and MAbs, principles of immunological reactions, and applications of various immunoassays have been discussed (Narayanasamy 2001, 2005). Using PAbs and MAbs in immunodiffusion tests attempts have been made to distinguish anastomosis groups of R. solani (Benson 1992; Thornton et al. 1999). For the detection of R. solani and related species, a one-step immuno-chromatographic lateral flow device (LFD) was developed. Antigens from representative isolates of R. solani AGs 1, 2-1, 2-3, 2-t, 3, 4, 5, 6, 7, 8, 9, 10, 11, and BI gave a positive response in LFD tests (Thornton et al. 2004).
Isozymes-based method
Isozymes are defined as multiple molecular forms of a single enzyme and these forms have similar enzymatic properties but slightly different amino acid sequences. The genetic locus may be monomorphic (expressed in a single allele). When the genetic locus is polymorphic the isozymes formed by the expression of different alleles are called allozymes. Isozyme analysis is a powerful biochemical technique that can be used to detect, differentiate and identify morphologically similar or closely related species, varieties and forma specialis. Liu et al. (1990) studied the genetic relationship among 14 isolates of R. solani AG-2 group by evaluating data derived from 11 enzyme systems. Pannecoucque et al. (2008) used pectic zymograms to group and subgroup R. solani isolates from Belgian cauliflower fields. Isozyme polymorphism was also profiled to analyze the genetic diversity of Indian R. solani isolates of AG1-IA (Neeraja et al. 2003), and Iranian R. solani isolates of AG1 subgroups infecting cotton (Mohammadi et al. 2003). Its use for population genetics investigation is limited in a predominantly asexual organism like R. solani.
Biosensor
Biosensors are analytical devices that use a biological sensing element integrated into a physiochemical transducer and produce an electrical signal when in contact with the analyte (pathogen). During the last decade, numerous biosensors have been reported, and many have shown high sensitivity and low detection limits (Ray et al. 2017). The specificity and sensitivity of the biosensors can be enhanced by the use of enzymes, antibodies, DNA probes and bacteriophages as the specific recognition elements (Fang and Ramasamy 2015). Nanomaterials like nanoparticles and quantum dots (QDs) etc. have emerged as essential tools for the faster detection of particular biological entities with extreme accuracy. Presently, on-site detection is gaining importance for plant disease diagnosis. The need for on-site detection has led to development the advance, rapid and sensitive detection devices and kits which can be used in-field for disease detection. Singh et al. (2010, 2014) developed a nano-Au-based dipstick to detect Karnal bunt disease in wheat rapidly. Few devices or kits are currently available as commercial products such as Alert test kits and Pocket diagnostic test kits from Neogen Corp. and La Chandra Bioscience, respectively, are available for pathogenic fungi, such as Pythium, Phytophthora and R. solani detection (Ray et al. 2017). Biosensors would become a promising and attractive alternative to other time-consuming and tedious assays such as ELISA, but there is a need for some modifications, improvements and proper validation for in-field application.
As the black scurf organism is a widespread seed and soilborne pathogen. R. solani has a large host range and survives in the soil for a long time; therefore, it is difficult to manage with any single practice. Integrated disease management (IDM) strategies and knowledge of each stage are required for the effective control of this disease. Current management approaches are discussed below and can be considered related to cultural practices, chemical and biological control, crop improvement and nanotechnological approach (Fig. 4).
Cultural practices
Cultural practices i.e., planting diseased free seed tubers, soil disinfection, non-host crop rotation, haulm cutting, tuber harvesting time, soil management, plant residues and irrigation influence the development of Rhizoctonia diseases in potatoes (Lal et al. 2022a). Black scurf can be managed by planting certified seeds free from any R. solani inoculums, therefore black scurf incidence monitoring is the first line of prevention of the disease. Furthermore, it would minimize the chance of establishing pathogens in the field. Non-host crop rotation is important for reducing the inoculum level of pathogenic microbes that require living hosts for survival (Peters et al. 2003). The crop rotation strategy is less effective with pathogens such as Pythium spp., Sclerotinia sclerotiorum, Sclerotium rolfsii and R. solani having broad host ranges and long term survival characteristics. Even though, R. solani can survive through multi-year rotations, increasing the time between potato crops can lower the inoculum level in the soil resulting in less disease severity and incidence (Hopkins et al. 2004). Honeycutt et al. (1996) observed that Rhizoctonia diseases were observed more severe in continuous potato cropping than in potatoes cultivated in rotation with non-host crops. Rotations of 3-5 years are often recommended to effectively reduce the disease severity of black scurf. The use of crops with known disease-suppressive capabilities, such as Brassica spp., cereals, millets, Sunhemp and non-solanaceous crops may provide additional resources for reducing disease through improved cropping systems. Field crops belonging to the Brassica family used in crop rotations and as green manures have been associated with reductions in soil-borne pathogens. These reductions could be due to the volatile sulfur compounds production through a process known as biofumigation and to change the structure of soil microbial density (Larkin and Griffin 2007). The mustard mixture reduced Rhizoctonia and common scab diseases of potatoes (Larkin et al. 2011). Maize, green gram, sun hemp and cowpea were evaluated as green manure crops for managing black scurf. Various other plant species (including weeds) have been shown to sustain R. solani (Jager et al. 1982; Carling et al. 1986b) and should be considered in crop rotation and weed control. Crop rotation may have some beneficial effects, but the fungus has such a wide host range and so easily reintroduced as sclerotia on seed potatoes that it is not very effective. In three cropping sequences viz., potato-wheat-paddy, potato-onion-maize & potato-green gram-groundnut, the highest incidence of black scurf was recorded in the potato-onion-maize cropping sequence (CPRI 2019).
Chemical control
The application of chemical fungicides is the most frequently used and effective method for managing the Rhizoctonia diseases in potatoes. Fungicides are chemically toxic compounds having unique mechanisms of action applied to eliminate or inhibit the growth of pathogens (Gullino et al. 2000). Fungicides prevent the Rhizoctonia disease development by several means like damaging the cell membrane of fungus, acts as enzyme inhibitors, disrupts the processes such as respiration or energy production or altered the metabolic pathways regulates the cell wall synthesis (Singh et al. 2019). As there is more than one AG responsible for Rhizoctonia disease of potatoes, these AGs have varying sensitivity to fungicides. Therefore, the identification of the group(s) causing disease in any particular field is crucial to fungicide selection (Kataria and Gisi 1996, 1999). Isolates of Rhizoctonia AGs 1, 3 and 5 were affected moderately by fungicides having aromatic hydrocarbon, whereas AGs 2-1, 4, 7 and 8 isolates were least sensitive. R. solani isolates showed high sensitivity levels against pencycuron, flutolanil and iprodione, except isolates of AG-5 (Campion et al. 2003). Commonly used available fungicides against R. solani in potatoes with active ingredients and action mechanisms are presented in Table 2.
Tuber-borne R. solani is easily manageable as compared to its soil-borne counterpart due to its accessibility to control agents. For controlling the black scurf of potatoes, seed tubers treatment with 3% acetic acid was found effective. (Dutt 1979). Potato variety ‘Kurfi Chandramukhi’ dipped in a mixture of acetic acid 1% + zinc sulphate 0.05% for 15 min before or after cold storage successfully controlled the R. solani (Somani 1986). Seed tubers treated with boric acid (3%) as dip treatment before cold storage (Singh et al. 2002) and with pencycuron as spray and dip treatments at planting time (Arora 2013; Thind et al. 2002) for controlling seed inoculums was followed to manage Rhizoctonia disease. Two chemicals viz., boric acid and pencycuron are frequently used by Indian farmers to control the potato black scurf disease (Khurana et al. 2001). In an in-vitro study, R. solani AG-3 was inhibited completely by tolclofos-methyl and Pencycuron, whereas in-field experiment, pencycuron and azoxystrobin controlled the sclerotial development on potato tubers (Virgen-Calleros et al. 2000). In India, few fungicide products viz., Penflufen 22.43% FS, Pencycuron 22.9% SC, Thifluzamide 24% SC, Carbendazim 25% + Mancozeb 50% WP, Carbendazim 12% +Mancozeb 63% WP, Carboxin 37.5% + Thiram 37.5% WS, Thiophanate-methyl 450g/l + Pyraclostrobin 50g/l w/v FS and Tebuconazole 15% + Zineb 57% WDG are registered under Central Insecticide Board and Registration Committee (CIB and RC) for control of Rhizoctonia disease in potato. Fluxapyroxad 333 FS is also registered in India for the management of potato black scurf disease (http://crop-protection.basf.in/en/fungicide). Recently, Arora et al. (2022) evaluated the efficacy of fluxapyroxad 333 FS at 0.08, 0.1 and 0.12% as tuber dip treatment and reported that these doses were statistically at par for managing black scurf. Some naturally derived fungicides like β-methoxyacrylates (also known as strobilurins) or QoI (Quinone outside Inhibitors) extracted from Strobilurus tencellus (wild mushroom) were found effective against R. solani (Bag et al. 2016). Tuber seeds treated with a mixture of sodium hypochlorite and thiophanatemethyl at the pre-planting stage reduced black scurf severity at harvesting and after storage (Errampalli and Johnston 2001).
Dusting seed tubers with tolclofos-methyl, or pencycuron spraying, gave control equal to that achieved by dipping in formaldehyde (Wicks et al. 1995). Acetaldehyde (5.0 ml/L) and Benzaldehyde (10.0 ml/L) in addition to fungicide (Basamid) @ 50 g/m2 of soil significantly reduced the disease incidence of black scurf (Abd-Alla et al. 2013). Chemical control is the highly effective and most widely used method for controlling field crop disease caused by fungus. However, the regular and continued application of a chemical fungicide increases the risk of the evolution of new highly pathogenic and fungicide-resistant races (Materatski et al. 2019). Mutations may occur in the genome of the fungus resulting in the alteration of the target site for molecular binding, target protein production can be increased and reduced uptake or metabolic breakdown of the fungicide may also increase (Gullino et al. 2000). Therefore, farmers and producers must either choose more specialized and long-lasting fungicides that increase the production expense or increase the frequency of fungicide application to control the fungal diseases. Another concern over fungicide application is related to hazards to the environment and human health (Kim et al. 2017). The fungicides application may also lead to adverse impacts on the terrestrial and aquatic ecosystems, soil organisms (e.g., earthworms, microorganisms) and poses a risk to the long-term fertility of the soil (Komarek et al. 2010). Therefore, research and development activities have to be established for searching the best alternatives to chemical fungicides such as the introduction of biocontrol agents to control Rhizoctonia diseases in potato crop.
Biological control
Biocontrol is the action of microbes, predators or parasites to minimize the population density of pathogenic organisms and is considered an eco-friendly and cost-effective component of an integrated disease management program (Verma et al. 2019; Kumar et al. 2022). Microbes such as PGPRs (plant growth promoting rhizobacteria) are the residents of the rhizosphere that are known to be involved in the synthesis of phytohormones, enhance nitrogen uptake, cause phosphorus/zinc/potassium solubilization and induced systemic resistance (Mustafa et al. 2019). The PGPRs that were found effective against R. solani included Pseudomonas spp., Bacillus spp. and Enterobacter spp. (Tabassum et al. 2017). In a greenhouse experiment, the interaction of potato seeds with Bacillus spp. showed 30-41.45% disease reduction of black scurf and 28.50-40.25% of stem canker caused by R. solani (Kumar et al. 2012). Pseudomonas sp. strain (S8.Fb11) reduced the proportion of infected tubers by R. solani to 40% for cv. Spunta and to 74% for cv. Nicola (Mrabet et al. 2013). Recently, Lal et al. (2022b) reported that talc formulation of Pseudomonas sp. strain (Pf14) enhanced agronomical characters and inhibited black scurf severity by up to 68% in a field experiment. In an in-vitro study, B. subtilis (V26) strain was found effective against R. solani and reduced disease incidence up to 63% and 81% of root canker and black scurf, respectively as well as enhanced plant growth in-planta (Khedher et al. 2015). Trichoderma and Gliocladium have also been reported as biocontrol agents against plant pathogens. Trichoderma spp. and Gliocladium spp. reduce the growth of R. solani employing different mechanisms, such as competition for nutrients and space, antibiosis and by mycoparasitism (Harman 2007). Volatile antibiotics (e.g. 6-pentyl-α-pyrone and isocyanide derivatives), hydrophilic compounds (e.g. heptelidic acid or koningic acid) and amphipathic polypeptides (e.g. peptaibiotics and peptaibols) produced by Trichoderma spp. are the major antifungal secondary metabolites (Lorito et al. 2010, Bailey and Lumsden 2014). Tsror et al. (2001) reported that the application of T. harzianum to the soil surface had a relatively small effect compared to the in-furrow treatments. Wilson et al. (2008) documented that application of T. harzianum, either in-furrow or in combination with flutolanil applied to seed tubers, increased marketable tuber yield (from 35% to 60%), and reduced black scurf incidence on progeny tubers from 31% to 11%, which could not be achieved using flutolanil alone. In a study, Hicks et al. (2014) reported that isolates of Trichoderma spp. (T. virens, T. atroviride and T. barbatum) reduced the percentage of diseased stolon by 41-46% in-planta. Recently, in a pot experiment, Walid et al. (2022) treated the potato seed with T. harzianum and observed the reduced severity of black scurf. Rahman et al. (2014) evaluated Trichoderma spp. against R. solani on potatoes and suggested that integrated or combination approaches could be effective for controlling black scurf. Brewer and Larkin (2005) demonstrated that a mixed formulation of B. subtilis and T. virens control stem canker well than each organism alone. In a field trial study, tuber treatment with 2% boric acid along with T. viride @ 10 g/kg seed recorded the lowest disease incidence (15.33%) and index (0.38) with the highest yield (324.68 q/ha) (Patel and Singh 2021). Despite the promising results with antagonists, the introduction of new biocontrol agents involves various considerations such as the tedious work of selection and screening, optimization of the mode of application to achieve the best results (Tabassum et al. 2017), the shelf life of the bioagents, efficacy in the field experiments, eco-friendly measures, and registration to be used as a PGPR (Etesami and Maheshwari 2018).
Hypovirulent R. solani strains show potential as new biocontrol agents against soil-borne potato diseases. Hypovirulent properties in these isolates are due to the presence of specific M2 cytoplasmic double-stranded (ds) RNA elements (Liu et al. 2003). In these strains, the dsRNA elements might be involved in the up-regulation of quinic acid pathway or down-regulation of the shikimic acid pathway, resulting in the drastic reduction in the phenyl acetic acid (PAA) production responsible for pathogenicity and virulence of R. solani (Bartz et al. 2012). In a field experiment, Rhs1AI, a hypovirulent strain of R. solani AG-3 had the potential to reduce Rhizoctonia disease incidence and severity up to 65% (Bandy and Tavantzis 1990; Larkin and Tavantzis 2013). However, in another study, where hypovirulent isolate Rhs1A1 did not show any reduction in black scurf severity when it was applied in combination with compost amendment with crop rotation (Bernard et al. 2014). More recently, Larkin (2020) tested two R. solani hypovirulent isolates (Rhs1AI and Bs69) combined with B. subtilis (GB03) and reported a reduction in disease incidence and severity of black scurf by 25–30% and 30–47%, respectively. The hypovirulent R. solani isolates may endow comparable or superior control of Rhizoctonia diseases of potatoes, than the existing bioagents, however, conjugations of hypovirulent strains with compatible bioagents having different modes of action are a matter of concern.
Nanotechnological approaches
The use of nanomaterials in plant disease management has created great interest (Kah et al. 2019), which may be very effective in the future with the progress of the application aspect of nanotechnology. Carbon, silver, silica and alumina-silicates-based nanoparticle were used for controlling plant disease. Recently, nanotechnology has a great effectiveness against numerous phytopathogens using silver nanoparticles (AgNPs). Interactions of AgNPs with microbes increase because of their larger surface area-to-volume ratio and thus more ability to permeate (Liao et al. 2019). When aqueous silver (Ag+) ions were exposed to a filtrate of Vitis vinifera, they reduced in solution resulting in the formation of stable AgNPs with 10-80 nm size which inhibited the growth of pathogenic bacteria (Chaudhary et al. 2012). Min et al. (2009) assayed the fungistatic and fungicidal effect of AgNPs against sclerotium-forming phytopathogenic fungi, R. solani, Sclerotinia sclerotiorum & Sclerotinia minor and documented that the AgNPs strongly inhibited the growth of fungal colonies and sclerotial germination. Nanosized silica-silver was effectual in the suppression of the growth of many fungi including R. solani and showed 100% growth inhibition at 10 ppm concentration (Park et al. 2006). Silica–silver nanoparticles are potentially effective against B. cinerea, B. sorokiniana, C. gloeosporioides, M. grisea, and R. solani (Jo et al. 2009). Kaur et al. (2012) examined the fungicidal properties of nano-size silver/chitosan nanoformulation against seed-borne plant pathogens viz., R. solani, A. flavus and A. alternata. It was documented that AgNPs concentration @ 15 mg exhibited excellent growth inhibition potential against A. alternata, S. sclerotiorum, Macrophomina phaseolina, R. solani, B. cenerea and Curvularia lunata (Krishnaraj et al. 2012). Elgorban et al. (2015) evaluated the different concentrations of AgNPs against six anastomosis groups (AGs) of R. solani infecting cotton and reported the antifungal properties to control R. solani AGs. Nejad et al. (2017) documented that AgNPs @ 50 ppm were effective against R. solani causing sheath blight in rice in both in-vitro and in-vivo conditions and showed the highest inhibition of sclerotia formation and mycelia growth and supressed the lesion development on leaves.
The in-vitro antifungal potential of various nanoparticles has been examined against phytopathogenic fungi namely A. alternata, M. phaseolina and R. solani. Among the various formulations of nanoparticles, Cu-chitosan nanoparticles were found most effective at 0.1% concentration (Saharan et al. 2013). Recently, Cui et al (2020) developed dual-functionalized polylactide (PLA) nanocapsules loaded with two fungicides validamycin and thifluzamide which showed better spreading performance on foliage application against R. solani compared with commercial fungicide formulations. However, several aspects of nanoparticles with relation to plants and the environment viz., their half-life in soil, their toxic effect on plants and animals and the optimum dosage for in-field application need to be determined. There are a few questions remaining to be addressed, viz., the exact mechanism of interaction of nanoparticles with fungal cells and how the surface area of nanoparticle influences the killing mechanism.
Genetic improvement of potato for Rhizoctonia resistance
Resistant germplasm is the most effective and environmental friendly way to control plant diseases. The conventional breeding approaches in potatoes can be coupled with modern biotechnology techniques to develop improved disease-resistant germplasms. Here the various strategies which can be implemented in the genetic improvement of potatoes against R. solani are discussed.
Selection and breeding
For determining the degree of host plant resistance against a pathogen, the maximum plant response to the pathogen must occur over a sufficient period under uniform selection pressure (Nelson and MacKenzie 1973). Monogenic host plant resistance, controlled by a single dominant gene is easily backcrossed into existing cultivars; however, this type of resistance may not be as durable as a resistance controlled by multiple genes. Resistance to Rhizoctonia diseases has existed in several wild Solanum species (Wastie 1994) and crosses with these wild cultivars have led to the conclusion that resistance to Rhizoctonia is under polygenic control and recessive (Li et al. 1995; Zeng et al. 2011). Screening tetraploid Solanum clones for resistance to R. solani has resulted in varied degrees of resistance to Rhizoctonia, which suggests that, even though not specifically selecting for resistance, breeders have incorporated some resistance by selecting away from damage caused by this pathogen (Leach and Webb 1993). To date, the availability of potato germplasms showing high resistance to R. solani is very limited. Few potato varieties e.g., Portage, Mainestay, AC Belmont and AC Brador, are moderately resistant to Rhizoctonia infection (Reeves et al. 1995, 1997; Tarn et al. 1995a, b), but varietal resistance is not regarded as a solution for long term to black scurf and stem canker. In a study, Khandaker et al. (2011) reported 6 out of 25 potato germplasms show moderate resistance against black scurf in Bangladesh. In India, mostly the commercially cultivated varieties are susceptible to black scurf. However, varieties like Kufri Dewa and K. Bahar showed moderate susceptibility and K. Sherpa found resistance to Rhizoctonia disease (CPRI 1989, 1999). Recently, Singh et al. (2021) screened 18 potato varieties against black scurf, among them, K. Ashok and K. Pukhraj exhibited moderately and highly susceptible reactions, respectively. To date, no quantitative trait loci (QTL) has been well characterized for black scurf resistance. Furthermore, identification and annotation of black scurf and stem canker resistance genes in QTL loci, functional characterization and application in marker-assisted breeding will help to develop resistant potato cultivars against Rhizoctonia. However, resistance breeding for Rhizoctonia in potatoes is difficult due to the presence of two phases of the disease (black scurf and stem canker), pathogen population diversity, environmental factors and soil conditions (Leach and Webb 1993). This coupled with the limited availability of resistant germplasm has led to the search for alternatives like manipulating plant genomes to enhance resistance.
Genetic manipulation through biotechnology: Defense-related proteins
Besides traditional agricultural practices and integrated disease management (IDM), developing resistant cultivars either by genetic alteration or conventional breeding would be the best alternative for controlling plant diseases. Nowadays, the development of genetically modified plants is an easier and preferred strategy to the complex pre-breeding approaches, especially in potatoes for expressing the gene of interest for a particular desired phenotypic/genotypic trait/s. The non-availability of complete resistant germplasm of potato against R. solani, conventional breeding for this trait has not succeeded. Published references on rice and potato carrying active transgene/s against Rhizoctonia in-planta are listed in Table 3. Each transgene construct contained a promoter that controls the gene expression in plants fused to a coding region for a protein expected to have direct antifungal properties, activate host defense response, and inhibited the fungal enzymes and virulence factors. Co-expression of two or more foreign genes was used in many studies.
On pathogen invasion, the accumulation of pathogenesis-related (PR) proteins is one important plant defence response. Transformation and expression of glycoside hydrolase proteins, which can degrade or lyse the cell wall of fungus and cell membrane, has been the most used method to develop fungal-resistant plants (Molla et al. 2020). Chitinases and glucanases are important antifungal proteins that hydrolyze or degrade chitin and glucan components of the fungal cell walls. Datta et al. (2001) introduced and overexpressed the PR genes i.e., chitinase 11 (PR3 family) which hydrolyse and degrade the fungal cell wall resulting inhibited the growth of R. solani in rice plants. In potatoes, the transformation of chitinase (ChiC), from Streptomyces griseus along with a bialaphos resistance (bar) gene conferred resistance against Alternaria solani (Khan et al. 2008). However, in a study, Moravčíková et al. (2004) concluded that the cucumber class III (ChiC) gene could not enhance resistance against R. solani AG-3 to any considerable level. A class I chitinase gene i.e., AF153195 from potato, was introduced into the tea genome and its overexpression resulted in an increased resistance against Exobasidium vexans (Singh et al. 2015). Similarly, in another study, the overexpression of chitinase gene LOC_Os11g47510 showed improved resistance against R. solani in rice plants (Richa et al. 2017). Other proteins with antifungal activity conferring enhanced tolerance to necrotrophic phytopathogens include small antimicrobial peptides (AMP) that disrupt fungal membrane integrity. The thaumatin-like proteins (TLP), osmotins, lysine-rich dermaseptins, cysteine-rich defensins and thionins, are acted by forming pores in the fungal membranes and causing cell lysis. Defensins attack fungal plasma membrane ceramide components and inhibit the transport of K+ and Ca+, with host-specific effects on hyphal branching and tip extension (Jha & Chattoo 2010). Snakin-1 (SN-1) is a basic, cysteine-rich AMP encoded by a small gene family that confer tolerance to R. solani when transferred in potato (Almasia et al. 2008). Transgenic potato minitubers with genes encoding dermaseptin, the osmotin AP24 and lysozyme gave rise to foliage showing reduced necrosis against R. solani in detached leaf assays (Rivero et al. 2012). In an experiment, M’hamdi et al. (2013) integrated a ribosome-inactivating protein (rip30) gene from barley into the potato genome and observed that transgenic clones showed reduced black scurf disease incidence and severity.
Recent gene editing techniques can provide platforms for precise transgene-free genome editing. CRISPR/Cas9 (clustered regularly interspaced short palindromic repeats) technique has been successfully implemented in potatoes for targeted mutagenesis to generate knockout mutations (using nonhomologous end-joining) and gene targeting to edit an endogenous gene (by homologous recombination) (Wang et al. 2015; Butler et al. 2016). CRISPR/Cas9 system has been utilized to install mutation in OsSWEET11 gene, leading to improved tolerance against rice sheath blight (Gao et al. 2018). More recently, González et al (2020) used CRISPR/Cas9 system to induce mutation in the StPPO2 gene to produce potato tubers with reduced PPO activity and enzymatic browning. Rapid advancements in technologies would ease genome modification and subsequently aid in developing disease-resistant potato plants.
Conclusion and recommendations
Globally, black scurf is an important disease in potatoes having economic importance. Various inoculum sources like; soil, infected seed tubers, crop residues and wider host range, diverse genetic and pathogenic variability contribute to the difficulty in successful control of black scurf. Modern molecular techniques permitting accurate detection and identification of Rhizoctonia solani at anastomosis sub-group levels in field soil and seed tubers allows to develop a decision-making system to support the growers in selecting seed materials and fields for planting potato crop. Assays such as RT-PCR, multiplex PCR, nested PCR, repetitive PCR, and LAMP are among the detection alternatives that endow rapid data analysis with specificity. As black scurf spreads through seed and soil-borne inoculums, the development of an integrated disease management (IDM) strategy that includes agronomic practices i.e. planting disease-free tubers, postharvest drying and field disinfection, rotating to a non-host crop, and utilization of a recommended dose of registered chemicals can control the disease. However, continued application of chemicals has negative effects on human health and the environment as well as induces pathogen resistance. An eco-friendly sustainable approach for controlling Rhizoctonia disease in potatoes is using biological agents such as PGPRs, Trichoderma spp., Gliocladium spp., and hypovirulent R. solani strains. Planting black scurf-resistant cultivars is another economical, effective and eco-friendly approach to managing the disease. Further, the identification and validation of pathogenicity factors in R. solani and defense-related genes in host plants associated with molecular interaction between R. solani and the potato will be a reference for developing black scurf resistant varieties. Modern biotechnological approaches such as Host derived dsRNA mediated silencing and CRISPR/Cas9 mediated knockout/knockdown are additional approaches that may be included to achieve eco-friendly and efficient disease management over synthetic fungicides.
Various biosynthetic and chemically synthesized nanomaterials and inorganic compounds have been tested to explore their efficacy in nano-fungicide formulations for black scurf management. However, improved nano-formulations need to develop for their potency and stability, considering the safety of the environment and human health. When it comes to the transportation of seed tubers, the phytosanitary certificate should be issued following a careful examination of potato bags to limit the movement of inoculums. Additionally, information about disease epidemiology is required for integrated disease management (IDM) programs.
References
Abd-Alla MA, El-Mougy NS, Abd-El-Kader MM, Abd-El-Kareem F, El-Gamal NG, El-Mohamedy RR (2013) Aldehydes compounds for controlling black scurf disease of potato (Solanum tuberosum L.) under field conditions. International Journal of Agriculture and Forestry 3:34–39
Agrios GN (2005) Plant pathology, 3rd edn. Academic, New York, p 803
Ajayi-Oyetunde OO, Bradley CA (2018) Rhizoctonia solani: Taxonomy, population biology and management of rhizoctonia seedling disease of soybean. Plant Pathology 67:3–17
Aliferis KA, Jabaji S (2012) FT-ICR/MS and GC-EI/MS metabolomics networking unravels global potato sprout’s responses ton Rhizoctonia solani infection. PLoS ONE 7:e42576
Almasia NI, Bazzini AA, Hopp HE, Vazquez-Rovere C (2008) Overexpression of snakin-1 gene enhances resistance to Rhizoctonia solani and Erwinia carotovora in transgenic potato plants. Molecular Plant Pathology 9:329–338
Amaradasa BS, Horvath BJ, Lakshman DK, Warnke SE (2013) DNA fingerprinting and anastomosis grouping reveal similar genetic diversity in Rhizoctonia spp. infecting turf grasses in the transition zone of USA. Mycologia 105:1190–1201
Anderson AL, Huber DM (1965) The plate-profiling technique for isolating soil fungi and studying their activity in the vicinity of roots. Phytopathology 55:592–594
Arora JK, Gupta S, Singh R, Chopra S, Choudhary S (2022) Fluxapyroxad 333FS: A novel systemic fungicide for effective management of black scurf of potato. The Pharma Innovation Journal 11:1253–1256
Arora RK (2013) Comparative efficacy of boric acid and pencycuron for management of black scurf of potato. Potato Journal 40:60–64
Bag MK, Yadav M, Mukherjee AK (2016) Bioefficacy of strobilurin based fungicides against rice sheath blight disease. Transcriptomics 4:128
Bailey BA, Lumsden RD (2014) Direct effect of Trichoderma and Gliocladium on plant growth and resistance to pathogens. In: Harman Gary E, Kubicek CP (eds) Trichoderma and Gliocladium, vol 2. enzymes, biological control commercial application. CRC Press, London, pp 185–204
Balali GR, Neate SM, Scott ES, Whisson DL, Wicks TJ (1995) Anastomosis group and pathogenicity of isolates of Rhizoctonia solani from potato crops in South Australia. Plant Pathology 44:1050–1057
Bandy BP, Leach SS, Tavantzis SM (1988) Anastomosis group 3 is the major cause of Rhizoctonia disease of potato in Maine. Plant Disease 72:596–598
Bandy BP, Tavantzis SM (1990) Effect of hypovirulent Rhizoctonia solani on rhizoctonia disease, growth, and development of potato plants. American Potato Journal 67:189–199
Banville GB, Carling DE (2001) Rhizoctonia canker and black scurf. In: Stevenson WR, Loria R, Franc G, Weingartner DP (eds) Compendium of potato diseases. APS Press, St Paul, pp 36–37
Banville GJ (1978) Studies on the Rhizoctonia disease of potatoes. American Potato Journal 55:56
Banville GJ (1989) Yield losses and damage to potato plants caused by Rhizoctonia solani Kühn. American Potato Journal 66:821–834
Banville GJ, Carling DE, Otrysko BE (1996) Rhizoctonia disease on potato. In: Sneh B, Jabaji-Hare S, Neate S, Dijst G (eds) Rhizoctonia species: Taxonomy, molecular biology, ecology, pathology and disease control. Kulwer Academic Publishers, Dordrecht, pp 321–330
Barna B, Fodor J, Harrach BD, Pogany M, Kiraly Z (2012) The Janus face of reactive oxygen species in resistance and susceptibility of plants to necrotrophic and biotrophic pathogens. Plant Physiology and Biochemistry 59:37–43
Bartz FE, Glassbrook NJ, Danehower DA, Cubeta MA (2012) Elucidating the role of the phenylacetic acid metabolic complex in the pathogenic activity of Rhizoctonia solani anastomosis group 3. Mycologia 104:793–803
Benson DM (1992) Detection by enzyme-linked immunosorbent assay of Rhizoctonia species in poinsettia cuttings. Plant Disease 76:578–581
Bernard E, Larkin RP, Tavantzis S, Erich MS, Alyokhin A, Gross SD (2014) Rapeseed rotation, compost and bicontrol amendments reduce soil-borne diseases and increase tuber yield in organic and conventional potato production systems. Plant Soil 374:611–627
Beukema HP, van der Zang DE (1990) Introduction to potato production. Centre for Agriculture Publishing and Documentation, The Netherlands
Bhandari P, Meenakshi R, Rai MK (2017) Management of black scurf disease of potato (Solanum tuberosum L.) with effective fungicide thifluzamide 24% SC. Annals of Horticulture 9(2):211–215
Boosalis MG, Scharen AL (1959) Methods for microscopic detection of Aphanomyces euteiches, Rhizoctonia solani and for isolation of Rhizoctonia solani associated with plant debris. Phytopathology 49:192–8
Bounoua S, Jabaji-Harec SH, Hogueb R, Charesta PM (1999) Polymerase chain reaction-based assay for specific detection of Rhizoctonia solani AG-3 isolates. Mycological Researsh 103:1–8
Brewer MT, Larkin RP (2005) Efficacy of several potential biocontrol organisms against Rhizoctonia solani on potato. Crop Protection 24:939–950
Brisbane PG, Neate SM, Pankhurst CE, Scott NS, Thomas MR (1995) Sequence-tagged site markers to identify Rhizoctonia solani AG4 or AG8 infecting wheat in South Australia. Phytopathology 85:1423–1427
Bulat SA, Lübeck M, Mironenko NV, Jensen DF, Lübeck PS (1998) UP-PCR analysis and ITS1 ribotyping of Trichoderma and Gliocladium fungi. Mycological Research 102:933–943
Butler NM, Baltes NJ, Voytas DF, Douches DS (2016) Gemini virus-mediated genome editing in potato (Solanum tuberosum L.) using sequence-specific nucleases. Frontiers in Plant Science 7:1045
Campion C, Chatot C, Perraton B, Andrivon D (2003) Anastomosis groups, pathogenicity and sensitivity to fungicides of Rhizoctonia solani isolates collected on potato crops in France. European Journal of Plant Pathology 109:983–992
Capote N, Pastrana AM, Aguado A, Sanchez-Torres P (2012) Molecular tools for detection of plant pathogenic fungi and fungicide resistance. In: Cumagun CJ (ed) Plant Pathology. InTechOpen, London, pp 151–202
Carling DE, Baird RE, Gitaitis RD, Brainard KA, Kuninaga S (2002) Characterization of AG–13, a newly reported anastomosis group of Rhizoctonia solani. Phytopathology 92:893–899
Carling DE, Kebler KM, Leiner RH (1986) Interactions between Rhizoctonia solani AG-3 and 27 plant species. Plant Disease 70:577–578
Carling DE, Leiner RH, Kebler KM (1986) Characterisation of Rhizoctonia solani and binucleate Rhizoctonia-like fungi collected from Alaskan soils with varied crop histories. Canadian Journal of Plant Pathology 8:305–310
Carling DE, Leiner RH, Westphale PC (1989) Symptoms, signs and yield reduction associated with Rhizoctonia disease of potato induced by tuber-borne inoculum of Rhizoctonia solani AG-3. American Potato Journal 66:693–701
Ceresini PC, Shew HD, Vilgalys RJ, Cubeta MA (2002) Genetic diversity of Rhizoctonia solani AG-3 from potato and tobacco in North Carolina. Mycologia 94:437–449
Chamoun R, Jabaji S (2011) Expression of genes of Rhizoctonia solani and the biocontrol Stachybotrys elegans during mycoparasitism of hyphae and sclerotia. Mycologia 103:483–93
Chaudhary S, Paul S, Sagar S (2012) Biosynthesis of silver nanoparticles using Vitis vinifera extract and evaluation of their antimicrobial activity. International Journal of Bio-Technology Research 2:1–12
Chaudhary S, Sagar S, Lal M, Tomar A, Kumar J, Kumar V, Kumar M (2023) Morpho-genetic variability of Rhizoctonia solani population causing sheath blight disease in rice (Oryza sativa L.). Journal of Environmetal Biology 44:108–121
Chaudhary S, Sagar S, Lal M, Tomar A, Kumar V, Kumar M (2020a) Biocontrol and growth enhancement potential of Trichoderma spp. against Rhizoctonia solani causing sheath blight disease in rice. Journal of Environmental Biology 41:1034–1045
Chaudhary S, Sharma S, Lal M, Sagar S, Shrama S, Kumar M (2023) Morphological and pathological variability of Rhizoctonia solani isolates from dhaincha-potato crop rotation and their mycelial compatibility relationship. Indian Pahytopathology 76:355–369
Chuadhary S, Lal M, Sagar S, Tyagi H, Kumar M, Shrama S, Chakrabarti SK (2020b) Genetic diversity studies based on morpho-pathological and molecular variability of the Sclerotinia sclerotiorum population infecting potato (Solanum tuberosum L.). World Journal of Microbiology and Biotechnology 36:177
Chye M, Zhao K, He Z, Ramalingam S, Fung K (2005) An agglutinating chitinase with two chitin-binding domains confers fungal protection in transgenic potato. Planta 220:717–730
Anonymous (1989) Annual Scientific Report. Central Potato Research Institute, Shimla, pp 74–76
Anonymous (1999) Annual Scientific Report. Central Potato Research Institute, Shimla, pp 132–133
Anonymous (2019) Annual Scientific Report, Central Potato Research Institute, Shimla, pp 68
Cubeta MA, Thomas E, Dean RA, Jabaji S, Neate SM, Tavantzis S, Toda T, Vilgalys R, Bharathan N, Abrams NF, Pakala SB, Pakala SM, Zafar N, Joardar V, Losada L, Nierman WC (2014) Draft genome sequence of the plant-pathogenic soil fungus Rhizoctonia solani anastomosis group 3 strain Rhs-1AP. Genome Announcements 2:e1072–e1014
Cubeta MA, Vilgalys R, Gonzalez D (1996) Molecular analysis of ribosomal RNA genes in Rhizoctonia fungi. In: Sneh B, Jabaji-Hare S, Neate S, Dijst G (eds) Rhizoctonia Species: Taxonomy molecular biology, ecology, pathology and disease control. Kulwer Academic Publishers, Dordrecht, pp 81–86
Cui J, Sun C, Wang A, Wang Y, Zhu H, Shen Y, Li N, Zhao X, Cui B, Wang C, Gao F, Zeng Z, Cui H (2020) Dual-functionalized pesticide nanocapsule delivery system with improved spreading behaviour and enhanced bioactivity. Nanomaterials 10:220
Das S, Shah FA, Butler RC, Falloon RE, Stewart A, Raikar S, Pitman AR (2014) Genetic variability and pathogenicity of Rhizoctonia solani associated with black scurf of potato in New Zealand. Plant Pathology 63:651–666
Datta K, Koukolikova-Nicola Z, Baisakh N, Oliva N, Datta SK (2000) Agrobacterium mediated engineering for sheath blight resistance of indica rice cultivars from different ecosystems. Theoretical and Applied Genetics 100:832–839
Datta K, Tu J, Oliva N, Ona I, Velazhahan R, Mew TW, Muthukrishnan S, Datta SK (2001) Enhanced resistance to sheath blight by constitutive expression of infection-related rice chitinase in transgenic elite indica rice cultivars. Plant Science 160:405–414
Datta K, Velazhahan R, Oliva N, Ona I, Mew T, Khush GS, Muthukrishnan S, Datta SK (1999) Over-expression of the cloned rice thaumatin-like protein (PR-5) gene in transgenic rice plants enhances environmental friendly resistance to Rhizoctonia solani causing sheath blight disease. Theoretical and Applied Genetics 98:1138–1145
Daub ME, Ehrenshaft M (2000) The photoactivated cercospora toxin cercosporin: contributions to plant disease and fundamental biology. Annual Review of Phytopathology 38:461–490
Dutt BL (1979) Bacterial and fungal diseases of potato. ICAR, New Delhi, p 109 (Tech. Bull)
Elgorban AM, El-Samawaty AEM, Yassin MA, Sayed SR, Adil SF, Elhindi KM, Bakri M, Khan M (2015) Antifungal silver nanoparticles: synthesis, characterization and biological evaluation. Biotechnology & Biotechnological Equipment 30:56–62
Errampalli D, Johnston HW (2001) Control of tuber-borne black scurf (Rhizoctonia solani) and common scab (Streptomyces scabies) of potatoes with a combination of sodium hypochlorite and thiophanate-methyl preplanting seed tuber treatment. Canadian Journal of Plant Pathology 23:68–77
Esfahani K, Motallebi M, Zamani MR, Sohi HH, Jourabchi E (2010) Transformation of potato (Solanum tuberosum cv. Savalan) by chitinase and β-1,3-glucanase genes of mycoparasitic fungi towards improving resistance to Rhizoctonia solani AG-3. Iranian Journal of Biotechnology 8:73–81
Etesami H, Maheshwari DK (2018) Use of plant growth promoting rhizobacteria (PGPRs) with multiple plant growth promoting traits in stress agriculture: action mechanisms and future prospects. Ecotoxicology and Environmental Safety 156:225–246
Fang Y, Ramasamy RP (2015) Current and prospective methods for plant disease detection. Biosensors 5:537–561
FAOSTAT (2022) FAO Statistical data, http://faostst.fao.org/
Farrokhi-Nejad R, Cromey MG, Moosawi-Jorf SA (2007) Determination of the anastomosis grouping and virulence of Rhizoctonia spp. associated with potato tubers grown in Lincoln, New Zealand. Pakistan Journal of Biological Sciences 10:3786–3793
Gallou A (2011) Impact of Rhizophagus sp. (syn. Glomus sp.) and Trichoderma harzianum on the potato resistance against Rhizoctonia solani and Phytophthora infestans, two major potato pathogens. PhD thesis. Louvain-la-Neuve, Belgium: Université catholique de Louvain, p 387
Gao Y, Zhang C, Han X, Wang ZY, Ma L, Yuan DP, Wu JN, Zhu XF, Liu JM, Li DP, Hu YB, Xuan YH (2018) Inhibition of OsSWEET11 function in mesophyll cells improves resistance of rice to sheath blight disease. Molecular Plant Pathology 19:2149–2161
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes -application to the identification of mycorrhizae and rust. Molecular Ecology 2:113–118
Genzel F, Franken P, Witzel K, Grosch R (2017) Salicylic acid-related plant defences are systemically induced in potato in response to Rhizoctonia solani AG3PT. Plant Pathology 67:337–348
Girard IJ, Tong C, Becker MG, Mao X, Huang J, Kievit T, Fernando WGD, Liu S, Belmonte MF (2017) RNA sequencing of Brassica napus reveals cellular redox control of Sclerotinia infection. Journal of Experimental Botany 68:5079–5091
Gkarmiri K, Finlay RD, Alstrom S, Thomas E, Cubeta MA, Hogberg N (2015) Transcriptomic changes in the plant pathogenic fungus Rhizoctonia solani AG-3 in response to the antagonistic bacteria Serratia proteamaculans and Serratia plymuthica. BMC Genomics 16:630
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annual Review of Phytopathology 43:205–27
González D, Rodríguez-Carres M, Boekhoutc T, Stalpers J, Kuramae EE, Nakatani AK, Vilgalys R, Cubeta MA (2016) Phylogenetic relationships of Rhizoctonia fungi within the Cantharellales. Fungal Biology 120:603–619
González MN, Massa GA, Andersson M, Turesson H, Olsson N, Falt AS, Storani L, Oneto CAD, Hofvander P, Feingold SE (2020) Reduced enzymatic browning in potato tubers by specific editing of a polyphenol oxidase gene via ribonucleoprotein complexes delivery of the CRISPR/Cas9 system. Frontiers in Plant Science 10:1649
Goswami SK, Kumar S, Singh V, Thind TS (2018) Efficacy of fungicides against Rhizoctonia solani causing black scurf of potato. Life Science Leaflets 106:5–9
Grosch R, Schneider JHM, Peth A, Waschke A, Franken P, Kofet A, Jabaji-Hare SH (2007) Development of a specific PCR assay for the detectionof Rhizocotonia solani AG1-IB using SCAR primers. Journal of Applied Microbiology 102:806–819
Gullino ML, Leroux P, Smith CM (2000) Uses and challenges of novel compounds for plant disease control. Crop Protection 19:1–11
Gvozdeva E, Volotskaya A, Sof’in A, Kudryavtseva N, Revina T, Valueva T (2006) Interaction of proteinases secreted by the fungal plant pathogen Rhizoctonia solani with natural proteinase inhibitors produced by plants. Applied Biochemistry and Microbiology 42:502–507
Hane JK, Anderson JP, Williams AH, Sperschneider J, Singh KB (2014) Genome sequencing and comparative genomics of the broad host-range pathogen Rhizoctonia solani AG8. PLoS Genetics 10:e1004281
Harman GE (2007) Overview of mechanisms and uses of Trichoderma spp. Phytopathology 96:190–194
Heller J, Tudzynski P (2011) Reactive oxygen species in phytopathogenic fungi: signaling, development, and disease. Annual Review of Phytopathology 49:369–390
Hicks E, Bienkowski D, Braithwaite M, McLean K, Falloon R, Stewart A (2014) Trichoderma strains suppress Rhizoctonia diseases and promote growth of potato. Phytopathologia Mediterranea 53:502–514
Honeycutt CW, Clapham WM, Leach SS (1996) Crop rotation and N fertilization effects on growth, yield, and disease incidence in potato. American Potato Journal 73:45–61
Hopkins BG, Hutchinson PJS, Patterson P, Miller J, Thornton M, Hafez S, Alvarez J (2004) Cropping sequence and rotation: impact on potato production and soil condition. Presented in two parts at the Idaho Seed Potato Conference, January 20, and the Idaho Potato Conference, pp 12
Hsieh SPY, Huang RZ, Wang TC (1996) Application of tannic acid in qualitative and quantitative growth assay of Rhizoctonia spp. Plant Pathology Bulletin 5:100–106
Irudukunda L, Wang YP, Nkurikiyimfura O, Wang T, Yang LN, Zhan J (2022) Establishment and application of a multiplex PCR assay for the rapid detection of Rhizoctonia solani anastomosis group (AG)-3PT, the pathogen causing black scurf and stem canker. Pathogen 11:627
Ito M, Meguro-Maoka A, Maoka T, Akino S, Masuta C (2017) Increased susceptibility of potato to Rhizoctonia diseases in Potato leafroll virus-infected plants. Journal of General Plant Pathology 83:169–172
Iwamtoo T, Sonobe T, Hayashi L (2003) Loop mediated isothermal amplification for the direct detection of Mycobacterium tuberculosis complex, M. Avium, and M. Intracellular in sputum samples. Journal of Clinical Microbiology 41:2616–2622
Jager G, Hekman W, Deenen A (1982) The occurrence of Rhizoctonia solani on subterranean parts of wild plants in potato fields. Netherland Journal of Plant Pathology 88:155–161
James WC, McKenzie AR (1972) The effect of tuberborne sclerotia of Rhizoctonia solani Kühn on the potato crop. American Potato Journal 49:296–301
Jashni MK, Mehrabi R, Collemare J, Mesarich CH, De Wit PJ (2015) The battle in the apoplast: further insights into the roles of proteases and their inhibitors in plant-pathogen interactions. Frontiers in Plant Science 6:584
Jha S, Chattoo BB (2010) Expression of plant defensin in rice confers resistance to fungal phytopathogens. Transgenic Research 19:373–384
Jo YK, Kim BH, Jung G (2009) Antifungal activity of silver ions and nanoparticles on phytopathogenic fungi. Plant Disease 93:1037–1043
Johnson MC, Pirone TP, Siegel MR, Varney DR (1982) Detection of Epichloe typhina in tall fescue by means of enzyme linked immunoassay. Phytopathology 72:647–650
Kadota Y, Shirasu K, Zipfel C (2015) Regulation of the NADPH oxidase RBOHD during plant immunity. Plant Cell Physiology 56:1472–1480
Kah M, Tufenkji N, White JC (2019) Nano-enabled strategies to enhance crop nutrition and protection. Nature Nanotechnology 14:532–540
Kanetis L, Tsimouris D, Christoforou M (2016) Characterization of Rhizoctonia solani associated with black scurf in Cyprus. Plant Disease 100:1591–1598
Kankam F, Long HT, He J, Zhang C, Zhang HX, Pu L, Qiu H (2016) 3-Methylthiopropionic acid of Rhizoctonia solani AG-3 and its role in the pathogenicity of the fungus. Plant Pathology Journal 32:85–94
Kankam F, Qiu H, Pu L, Long HT, Zhang C (2016) Isolation, purification and characterization of phytotoxins produced by Rhizoctonia solani AG-3, the cause agent of potato stem canker. American Journal of Potato Research 93:321–330
Kataria HR, Gisi U (1996) Chemical control of Rhizoctonia species. In: Sneh B, Jabaji-Hare S, Neate S, Dijst G (eds) Rhizoctonia Species: Taxonomy molecular biology, ecology, pathology and disease control. Kulwer Academic Publishers, Dordrecht, pp 537–547
Kataria HR, Gisi U (1999) Selectivity of fungicides within the genus Rhizoctonia. In: Lyr H, Russell PE, Dehne HW, Sisler HD (eds) Modern fungicides and antifungal compounds. Intercept, Andover, pp 421–429
Kaur P, Thakur R, Choudhary A (2012) An in vitro study of the antifungal activity of silver/chitosan nanoformulations against important seed borne pathogens. International Journal of Scientific & Technology Research 1:83–86
Keijer J (1996) The initial steps of the infection process in Rhizoctonia solani. In: Sneh B, Jabaji-Hare S, Neate S, Dijst G (eds) Rhizoctonia Species: Taxonomy molecular biology, ecology, pathology and disease control. Kulwer Academic Publishers, Dordrecht, pp 149–162
Keijer J, Houterman PM, Dullemans AM, Korsman MG (1996) Heterogeneity in electrophoretic karyotype within and between anastomosis groups of Rhizoctonia solani. Mycological Research 100:789–797
Khan RS, Sjahril R, Nakamura I, Mii M (2008) Production of transgenic potato exhibiting enhanced resistance to fungal infections and herbicide applications. Plant Biotechnology Reports 2:13–20
Khandaker MM, Khair A, Bhuiyan MKA (2011) Disease reaction of potato germplasms and true potato seeds against Rhizoctonia solani. Bangladesh Journal of Botany 40(2):193–196
Khedher SB, Kilani-Feki O, Dammak M, Jabnoun-Khiareddine H, Daami-Remadi M, Tounsi S (2015) Efficacy of Bacillus subtilis V26 as a biological control agent against Rhizoctonia solani on potato. Comptes Rendus Biologies 338:784–792
Khurana SMP, Thind TS, Mohan C (2001) Diseases of potato and their management. In: Thind TS (ed) Diseases of fruits and vegetables and their management. Kalyani Publisher, New Delhi, pp 237–265
Kim JK, Jang IC, Wu R, Zuo WN, Boston RS, Lee YH, Ahn IP, Nahm BH (2003) Co-expression of a modified maize ribosome-inactivating protein and a rice basic chitinase gene in transgenic rice plants confers enhanced resistance to sheath blight. Transgenic Research 12:475–484
Kim JY, Park SC, Kim MH, Lim HT, Park Y, Hahm KS (2005) Antimicrobial activity studies on a trypsin-chymotrypsin protease inhibitor obtained from potato. Biochemical and Biophysical Research Communication 330:921–7
Kim KH, Kabir E, Jahan SA (2017) Exposure to pesticides and the associated human health effects. Science of The Total Environment 575:525–535
Ko W, Hora FK (1971) A selective medium for the quantitative determination of Rhizoetonia solani in soil. Phytopathology 61:707–710
Komarek M, Cadkova E, Chrastny V, Bordas F, Bollinger JC (2010) Contamination of vineyard soils with fungicides: A review of environmental and toxicological aspects. Environment International 36:138–151
Kouzai Y, Kimura M, Watanabe M, Kusunoki K, Osaka D, Suzuki T, Matsui H, Yamamoto M, Ichinose Y, Toyoda K, Matsuura T, Mori IC, Hirayama T, Minami E, Nishizawa Y, Inoue K, Onda Y, Mochida K, Noutoshi Y (2018) Salicylic acid-dependent immunity contributes to resistance against Rhizoctonia solani, a necrotrophic fungal agent of sheath blight, in rice and Brachypodium distachyon. New Phytologist 217:771–783
Krishnaraj C, Ramachandran R, Mohan K, Kalaichelvan PT (2012) Optimization for rapid synthesis of silver nanoparticles and its effect on phytopathogenic fungi. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy 93:95–99
Kulkarni S, Chavhan T (2017) Management of black scurf disease caused by Rhizoctonia solani Kühn through research and farmers participatory trials in major potato growing regions of Northern Karnataka. International Journal of Agriculture Innovations and Research 6:2319–1473
Kumar KK, Poovannan K, Nandakumar R, Thamilarasi K, Geetha C (2003) A high throughput functional expression assay system for a defence gene conferring transgenic resistance on rice against the sheath blight pathogen, Rhizoctonia solani. Plant Science 165:969–976
Kumar SS, Rao MRK, Kumar RD, Panwar S, Prasad CS (2012) Biocontrol by plant growth promoting rhizobacteria against black scurf and stem canker disease of potato caused by Rhizoctonia solani. Archives of Phytopathology and Plant Protection 46:487–502
Kumar V, Srivastava A, Jain L, Chuadhary S, Kaushal P, Soni R (2022) Harnessing the potential of genetically improved bioinoculants for sustainable agriculture: recent advances and perspectives. In: Soni R, Suyal DC, Yadav AN, Goel R (eds) Developments in applied microbiology and biotechnology, trends of applied microbiology for sustainable economy. Academic Press, pp 319–341
Kuninaga S, Carling DE, Takeuchi T, Yokosawa R (2000) Comparison of rDNA-ITS sequences between potato and tobacco strains in Rhizoctonia solani AG3. Journal of Geneneral Plant Pathology 66:2–11
Lal M, Chaudhary S, Kumar M, Shrama S, Chakrabarti SK (2020) First report of collar and stem rot caused by Rhizoctonia solani AG1-IA on Sesbania sesban in India. Plant Disease 104(12):3251
Lal M, Chaudhary S, Sharma S, Subhash S, Kumar M (2022) Bio-intensive management of fungal diseases of potatoes. In: Chakrabarti SK, Sharma S, Shah MA (eds) Sustainable management of potato pests and diseases. Springer, Singapore, pp 453–493
Lal M, Chaudhary S, Yadav S, Sharma S, Chakrabarti SK, Kumar M (2019) Development of spray schedules for management of late blight of potato using new chemicals. Journal of Mycology and Plant Pathology 49:405–412
Lal M, Kumar A, Chaudhary S, Singh RK, Sharma S, Kumar M (2022) Antagonistic and growth enhancement activities of native Pseudomonas spp. against soil and tuber-borne diseases of potato (Solanum tuberosum L.). Egyptian Journal of Biological Pest Control 32:22
Lal M, Sharma S, Chakrabarti SK, Kumar M, Singh N (2018) Bio-efficacy and phytotoxicity of carboxin 37.5% + thiram 37.5% WS against black scurf of potato. International Journal of Agricultural and Statistical Sciences 14:617–621
Lal M, Sharma S, Chakrabarti SK, Kumar M (2017) Thifluzamide 24% SC: A new molecule for potato tubers treatment against black scurf disease of potato caused by Rhizoctonia solani. International Journal of Current Microbiology & Applied Sciences 6:370–375
Lal M, Sharma S, Yadav S, Kaushik SK (2014) Bioefficacy of new molecule: penflufen 240 FS against black scurf of potato. International Journal of Agricultural and Statistical Sciences 10:63–66
Lal M, Yadav S, Chand S (2017) Thiophanate methyl 45% + Pyraclostrobin 5% FS: A new molecule for potato tubers treatment against black scurf disease of potato caused by Rhizoctonia solani. Indian Journal of Plant Protection 45:177–180
Larkin RP (2020) Biological control of soilborne diseases in organic potato production using hypovirulent strains of Rhizoctonia solani. Biological Agriculture & Horticulture 36:119–129
Larkin RP, Griffin TS (2007) Control of soil-borne potato diseases using Brassica green manures. Crop Protection 26:1067–1077
Larkin RP, Honeycutt CW, Griffin TS, Olanya OM, Halloran JM, He Z (2011) Effect of different cropping system approaches and water management on soil borne diseases and soil microbial communities. Phytopathology 101:58–67
Larkin RP, Tavantzis (2013) Use of biocontrol organisms and compost amendments for improved control of soilborne diseases and increased potato production. American Journal of Potato Research 90:261–270
Leach SS, Webb RE (1993) Evaluation of potato cultivars, clones and a true seed population for resistance to Rhizoctonia solani. American Potato Journal 70:317–328
Lees AK, Cullen DW, Sullivan L, Nicolson MJ (2002) Development of conventional and quantitative real-time PCR assays for the detection and identification of Rhizoctonia solani AG-3 in potato and soil. Plant Pathology 51:293–302
Lehtonen MJ, Ahvenniemi P, Wilson PS, German-Kinnari M, Valkonen JPT (2008) Biological diversity of Rhizoctonia solani (AG-3) in a northern potato-cultivation environment in Finland. Plant Pathology 57:141–151
Lehtonen MJ, Somervuo P, Valkonen JP (2008) Infection with Rhizoctonia solani induces defense genes and systemic resistance in potato sprouts grown without light. Phytopathology 98:1190–8
Li ZK, Pinson SRM, Marchetti MA, Stansel JW, Park WD (1995) Characterization of quantitative trait loci (QTL) in cultivated rice contributing to field resistance to sheath blight (Rhizoctonia solani). Theoretical and Applied Genetics 91:382–388
Liao C, Li Y, Tjong SC (2019) Bactericidal and cytotoxic properties of silver nanoparticles. International Journal of Molecular Sciences 20:449
Liu C, Lakshman DK, Tavantzis SM (2003) Expression of a hypovirulence-causing double-stranded RNA is associatedwith up-regulation of quinic acid pathway in Rhizoctonia solani. Current Genetics 42:284–291
Liu JJ, Ekramoddoullah AK (2006) The family 10 of plant pathogenesis-related proteins: their structure, regulation, and function in response to biotic and abiotic stresses. Physiological and Molecular Plant Pathology 68:3–13
Liu Z, Nickrent DL, Sinclair JB (1990) Genetic relationship among isolates of Rhizoctonia solani anastomosis group-2 based on isozyme analysis. Candian Journal of Plant Pathology 12:376–382
Lorito M, Woo SL, Harman GE, Monte E (2010) Translational research on Trichoderma: from ‘omics to the field. Annual Review of Phytopathology 48:395–417
Lu C, Song B, Zhang H, Wang Y, Zheng X (2015) Rapid diagnosis of soybean seedling blight caused by Rhizoctonia solani and soybean charcoal rot by Macrophomina phaseolina using LAMP assay. Phytopathology 105:1612–1617
Lübeck M, Lübeck PS (2005) Universally primed PCR (UP–PCR) and its applications in mycology. In: Deshmukh SK, Rai MK (eds) Biodiversity of fungi: their role in human life. Science Publishers, Enfield, pp 409–438
M’hamdi M, Chikh-Rouhou H, Boughalleb N, Ruiz de Galarreta JI (2012) Enhanced resistance to Rhizoctonia solani by combined expression of chitinase and Ribosome Inactivating Protein in transgenic potatoes (Solanum tuberosum L.). Spanish Journal of Agricultural Research 10:778–785
M’hamdi M, Chikh-Rouhou H, Boughalleb N, Ruiz de Galarreta JI (2013) Ribosome inactivating protein of barley enhanced resistance to Rhizoctonia solani in transgenic potato cultivar ‘Desiree’ in greenhouse conditions. Biotechnology, Agronomy and Society and Environment 17:20–26
Ma Z, Michialides TJ (2005) Advances in understanding molecular mechanisms of fungicide resistance and molecular detection of resistant genotypes in phytopathogenic fungi. Crop Protection 24:853–863
Malik O, Chohan S, Naqvi SAH (2014) Occurrence of black scurf disease of potato in Multan (Punjab) along with its in-vitro chemical and biotic elicitor mediated management. Journal of Agricultural Science 6(9):134–143
Mandava NB, Orellana RG, Warthen JD Jr, Worley JF, Dutky SR, Finegold H, Weathington BC (1980) Phytotoxins in Rhizoctonia solani: isolation and biological activity of M-hydroxy- and M-methoxyphenylacetic acids. Journal of Agricultural and Food Chemistry 28:71–75
Marshall D, Rush M (1980) Infection cushion formation on rice sheaths by Rhizoctonia solani. Phytopathology 70:947–950
Martinson CA (1963) Inoculum potential relationships of Rhizoctonia solani measured with soil microbiological sampling tubes. Phytopathology 58:634–8
Maruthasalam S, Kalpana K, Kumar KK, Loganathan M, Poovannan K, Raja JAJ, Kokiladevi E, Samiyappan R, Sudhakar D, Balasubhramanian P (2007) Pyramiding transgenic resistance in elite indica rice cultivars against the sheath blight and bacterial blight. Plant Cell Report 26:791–804
Materatski P, Varanda C, Carvalho T, Dias AB, Campos MD, Gomes L, Nobre T, Rei F, Felix MR (2019) Effect of long-term fungicide applications on virulence and diversity of Colletotrichum spp. associated to Olive anthracnose. Plants (Basel) 8(9):311
Matsumoto M (2002) Trials of direct detection and identification of Rhizoctonia solani AG1 and AG2 subgroups using specifically primed PCR analysis. Mycoscience 43(2):185–189
McCartney HA, Foster SJ, Fraaije BA, Ward E (2003) Molecular diagnostics for fungal plant pathogens. Pest Management Science 59:129–142
Min JS, Kim KS, Kim SW, Jung JH, Lamsal K, Kim SB, Jung M, Lee YS (2009) Effects of colloidal silver nanoparticles on sclerotium-forming phytopathogenic fungi. Plant Pathology Journal 25:376–380
Misawa T, Kuninga S (2010) The first report of tomato foot rot caused by Rhizoctonia solani AG-3PT and AG-2-Nt and its host range and molecular characterization. Journal of General Plant Pathology 76:310–319
Mohammadi M, Banihashemi M, Hedjaroude GA, Rahimian H (2003) Genetic diversity among Iranian isolates of Rhizoctonia solani Kühn anastomosis group1 subgroups based on isozyme analysis and total soluble protein pattern. Journal of Phytopathology 151:162–170
Molla KA, Karmakar S, Molla J, Bajaj P, Varshney RK, Datta SK, Datta K (2020) Understanding sheath blight resistance in rice: the road behind and the road ahead. Plant Biotechnology Journal 18:895–915
Moravčíková J, Matusikova I, Libantova J, Bauer M, Mlynarova L (2004) Expression of cucumber class III chitinase and Nicotiana plumbaginifolia class I glucanase genes in transgenic potato plants. Plant Cell, Tissue and Organ Culture 79:161–168
Mori Y, Kitao M, Tomita N, Notomi T (2004) Real time turbidimetry of LAMP reaction for quantifying template DNA. Journal of Biochemical and Biophysical Methods 59:145–157
Mori Y, Nagamine K, Tomita N, Notomi T (2001) Detection of loop-mediated isothermal amplification reaction by turbidity derived from magnesium pyrophosphate formation. Biochemical and Biophysical Research Communications 289:150–154
Mrabet M, Djebali N, Elkahouri S, Miloud Y, Saidi S, Tarhouni B, Mhamdi R (2013) Efficacy of selected Pseudomonas strains for biocontrol of Rhizoctonia solani in potato. Phytopathologia Medeterranea 52:449–456
Mustafa S, Kabir S, Shabbir U, Batool R (2019) Plant growth promoting rhizobacteria in sustainable agriculture: from theoretical to pragmatic approach. Symbiosis 78:115–123
Muzhinji N, Truter M, Woodhall JW, van der Waals JE (2015) Anastomosis groups and pathogenicity of Rhizoctonia solani and Binucleate Rhizoctonia from potato in South Africa. Plant Disease 99:1790–1802
Muzhinji N, Woodhall JW, Truter M, van der Waals JE (2014) Elephant hide and growth cracking on potato tubers caused by Rhizoctonia solani AG 3-PT in South Africa. Plant Disease 98:570
Muzhinji N, Woodhall JW, Truter M, van der Waals JE (2018) Variation in fungicide sensitivity among Rhizoctonia isolates recovered from potatoes in South Africa. Plant Disease 102:1520–1526
Nadarajah K, Razali NM, Cheah BH, Sahruna NS, Ismail I, Tathode M, Bankar K (2017) Draft genome sequence of Rhizoctonia solani anastomosis group 1 subgroup 1a strain 1802/KB isolated from rice. Genome Announcements 5:e01188-17
Nandakumar R, Babu S, Kalpana K, Raguchander T, Balasubramanian P, Samiyappan R (2007) Agrobacterium-mediated transformation of indica rice with chitinase gene for enhanced sheath blight resistance. Biologia Plantarum 51:142–148
Narayanasamy P (2001) Plant pathogen detection and disease diagnosis, 2nd edn. Marcel Dekker, New York
Narayanasamy P (2005) Immunology in plant health and its impact on food safety. The Haworth Press, New York
Neeraja CN, Shenoy VV, Reddy CS, Sarma NP (2003) Isozyme polymorphism and virulence of Indian isolates of the rice sheath blight fungus. Mycopathologia 156:101–108
Nejad MS, Bonjar GHS, Khatami M, Amini A, Aghighi S (2017) In vitro and in vivo antifungal properties of silver nanoparticles against Rhizoctonia solani, a common agent of rice sheath blight disease. IET Nanobiotechnology 11:236–240
Nelson RR, MacKenzie DR (1973) The detection and stability of disease resistance. In: Nelson RR (ed) Breeding plants for disease resistance. University Park, The Pennsylvania State University Press, Pennsylvania, pp 12–39
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Hase AN (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Research 28:e63
Ogoshi A (1987) Ecology and pathogenicity of anastomosis and intraspecific groups of Rhizoctonia solani Kühn. Annual Review of Phytopathology 25:125–143
Oscorbin IP, Belousova EA, Zakabunin AI, Boyarskikh UA, Filipenko ML (2016) Comparison of florescent intercalating dyes for quantitative loop-mediated isothermal amplification (qLAMP). Biotechniques 61:20–25
Pannecoucque J, Van Beneden S, Höfte M (2008) Characterization and pathogenicity of Rhizoctonia isolates associated with cauliflower in Belgium. Plant Pathology 57:737–746
Park HJ, Kim SH, Kim HJ, Choi SH (2006) A new composition of nanosized silica-silver for control of various plant diseases. Journal of Plant Pathology 22:295–302
Patel JS, Brennan MS, Khan A, Ali GS (2015) Implementation of loop-mediated isothermal amplification methods in lateral flow devices for the detection of Rhizoctonia solani. Canadian Journal of Plant Pathology 37:118–129
Patel VM, Singh N (2021) Management of black scurf (Rhizoctonia solani) of potato through organic approaches. Indian Journal of Agricultural Research 55:157–162
Patil HJ, Solanki MK (2016) Microbial inoculant: modern era of fertilizers and pesticides. In: Singh D, Singh H, Prabha R (eds) Microbial inoculants in sustainable agricultural productivity: research perspectives, 1st edn. Springer, New Delhi, pp 319–343
Patil VU, Girimalla V, Sagar V, Bhardwaj V, Chakrabarti SK (2017) Draft genome sequencing of Rhizoctonia solani anastomosis group 3 (AG3-PT) causing stem canker and black scurf of potato. American Journal of Potato Research 95:87–91
Paulitz TC, Schroeder KL (2005) A new method for the quantification of Rhizoctonia solani and R. oryzae from soil. Plant Disease 89:767–772
Peters RD, Sturz AV, Carter MR, Sanderson JB (2003) Developing disease-suppressive soils through crop rotation and tillage management practices. Soil and Tillage Research 72:181–192
Rahman M, Ali MA, Dey TP, Islam MM, Naher L, Ismail A (2014) Evolution of disease and potential biocontrol activity of Trichoderma spp. against Rhizoctonia solani on potato. Bioscience Journal 30:1108–1117
Rajesh T, Maruthasalam S, Kalpana K, Poovannan K, Kumar KK, Kokiladevi E, Sudhakar D, Samiyappan R, Balasubramanian P (2016) Stability of sheath blight resistance in transgenic ASD16 rice lines expressing a rice chi11 gene encoding chitinase. Biologia Plantarum 60:749–756
Ray M, Ray A, Dash S, Mishra A, Achary KG, Nayak S, Singh S (2017) Fungal disease detection in plants: Traditional assays, nival diagnostic techniques and biosensors. Biosensors and Bioelectronics 87:708–723
Reeves AF, Porter GA, Cunningham CE, Nickeson RJ, Manzer FE, Work TM, Davis AA, Plissey ES (1995) Portage: A new early-maturing, round white table potato variety. American Potato Journal 72:681–688
Reeves AF, Porter GA, Work TM, Lambert DH, Davis AA, Plissey ES (1997) Mainestay: A high-yielding, round white potato variety for fresh markets. American Potato Journal 74:255–263
Richa K, Tiwari IM, Devanna BN, Botella JR, Sharma V, Sharma TR (2017) Novel chitinase gene LOC_Os11g47510 from Indica rice Tetep provides enhanced resistance against sheath blight pathogen Rhizoctonia solani in rice. Frontiers in Plant Science 8:596
Rioux R, Manmathan H, Singh P, Reyes B, Jia Y, Tavantzis S (2011) Comparative analysis of putative pathogenesis-related gene expression in two Rhizoctonia solani pathosystems. Current Genetics 57:391–408
Rivero M, Furman N, Mencacci N, Picca P, Toum L, Lentz E, Almonacid FB, Mentaberry A (2012) Staking of antimicrobial genes in potato transgenic plants confers increased resistance to bacterial and fungal pathogens. Journal of Biotechnology 157:334–343
Saharan V, Mehrotra A, Khatik R, Rawal P, Sharma SS, Pal A (2013) Synthesis of chitosan based nanoparticles and their in vitro evaluation against phytopathogenic fungi. International Journal of Biological Macromolecules 62:677–683
Salazar O, Julian MC, Rubio V (2000) Primers based on specific rDNA–ITS sequences for PCR detection of Rhizoctonia solani, R. solani AG–2 subgroups and ecological types, and binucleate Rhizoctonia. Mycological Research 104:281–285
Samsatly J, Bayen S, Jabaji SH (2020) Vitamin B6 is under a tight balance during disease development by Rhizoctonia solani on different cultivars of potato and on Arabidopsis thaliana mutants. Frontiers in Plant Science 11:875
Samsatly J, Chamoun R, Gluck-Thaler E, Jabaji S (2015) Genes of the de novo and salvage biosynthesis pathways of vitamin B6 are regulated under oxidative stress in the plant pathogen Rhizoctonia solani. Frontiers in Microbiology 6:1429
Samsatly J, Copley TR, Jabaji SH (2018) Antioxidant genes of plants and fungal pathogens are distinctly regulated during disease development in different Rhizoctonia solani pathosystems. PLoS One 13:e0192682
Sanz-Alférez S, Mateos B, Alvarado R, Sánchez M (2008) SAR induction in tomato plants is not affective against root-knot nematode infection. European Journal of Plant Pathology 120:417–425
Scholte K (1989) Effect of soil-borne Rhizoctonia solani Kühn on yields and quality of ten potato cultivars. Potato Research 32:367–376
Sharma S (2015) Black Scurf. In: Singh BP, Nagesh M, Sharma S, Sagar V, Jeevlatha A, Sridhar J (eds) A manual on diseases and pest of potato, ICAR-Central Potato Research Institute, Shimla, pp 11–13 (Tech Bull. No.: 101)
Shen YM, Guo CJ, Wang XG, Shen RQ, Chen AC, Hu XP (2017) Rapid detection of Rhizoctonia solani AG3 sclerotia in soil by quantitative real-time PCR. Mycosystema 36:1383–1391
Shuai Y, Yu K, Mei G, Xuezhi D, Fanxiang M, Qi W, Wenzhong W, Yanzhi M, Xin G, Ling W (2022) First report of black scurf caused by Rhizoctonia solani AG2-2IV on potato tubers in Heilongjiang province, China. Plant Disease 106:2996
Singh BP, Arora RK, Khurana SMP (2002) Soil and Tuber Borne Diseases of Potato. CPRI, Shimla, p 74 (Tech Bull. No.: 41)
Singh HR, Deka M, Das S (2015) Enhanced resistance to blister blight in transgenic tea (Camellia sinensis [L.] o. Kuntze) by overexpression of class I chitinase gene from potato (Solanum tuberosum). Functional & Integrative Genomics 15:461–480
Singh P, Mazumdar P, Harikrishna JA, Babu S (2019) Sheath blight of rice: a review and identification of priorities for future research. Planta 250:1387–1407
Singh P, Subramanian B (2017) Responses of rice to Rhizoctonia solani and its toxic metabolite in relation to expression of Osmyb4 transcription factor. Plant Protection Science 53:208–215
Singh PK, Patidar JK, Singh R, Roy S (2021) Screening of potato varieties against black scurf caused by Rhizoctonia solani Kühn. International Journal of Current Microbiology and Applied Sciences 10:1444–1449
Singh S, Gupta AK, Gupta S, Gupta S, Kumar A (2014) Surface plasmon resonance (SPR) and cyclic voltammetry based immunosensor for determination of teliosporic antigen and diagnosis of Karnal Bunt of wheat using anti-teliosporic antibody. Sensors and Actuators B: Chemical 191:866–73
Singh S, Singh M, Agrawal VV, Kumar A (2010) An attempt to develop surface plasmon resonance based immunosensor for Karnal bunt (Tilletia indica) diagnosis based on the experience of nano-gold based lateral flow immuno-dipstick test. Thin Solid Films 519:1156–59
Sneh B, Burpee L, Ogoshi A (1991) Identification of Rhizoctonia Species. The American Phytopathological Society, Minnesota
Somani AK (1986) Non-hazardous chemical control of black scurf of potato. Indian Journal of Agricultural Sciences 56:366–369
Spurlock TN, Rothrock CS, Monfort WS (2015) Evaluation of methods to quantify populations of Rhizoctonia in soil. Plant Disease 99:836–841
Sridevi G, Parameswari C, Sabapathi N, Raghupathy V, Veluthambi K (2008) Combined expression of chitinase and β-1,2-glucanase genes in indica rice (Oryza sativa L.) enhances resistance against Rhizoctonia solani. Plant Science 175:283–290
Sridevi G, Sabapathi N, Meena P, Nandakumar R, Samiyappan R (2003) Transgenic indica rice variety Pusa Basmati 1 constitutively expressing a rice chitinase gene exhibits enhanced resistance to Rhizoctonia solani. Journal of Plant Biochemistry and Biotechnology 12:93–101
Stodart BJ, Harvey PR, Neate SM, Melanson DL, Scott ES (2007) Genetic variation and pathogenicity of anastomosis group 2 isolates of Rhizoctonia solani in Australia. Mycological Research 111:891–900
Tabassum B, Khan A, Tariq M, Ramzan M, Khan MS, Shahid N, Aaliya K (2017) Bottlenecks in commercialisation and future prospects of PGPR. Applied Soil Ecology 121:102–117
Tarn TR, De Jong H, Murphy AM, Tai GCC, Arsenault WJ, Thorpe HE, Bangall RH, Platt HW, Young DA, Davies HT (1995) AC Belmont: A new early-maturing potato cultivar with short dormancy. American Potato Journal 72:409–415
Tarn TR, Tai GCC, Murphy AM, De Jong H, Platt HW, Bagnall RH, Arsenault WJ, Thorpe JHE, Young DA, Davies HT (1995) AC Brador: A new late-maturing cultivar with a high degree of field resistance to late blight. American Potato Journal 72:401–408
Thind TS, Mohan C, Kaur S (2002) Promising activity of pencycuron, a phenylurea-based fungicide, for effective management of black scurf of potato. Indian Phytopathology 55:39–44
Thornton CR, Andrew CG, Forrest R, Lamotte R (2004) A one-step, immuno-chromatogarphic lateral flow device specific to Rhizoctonia solani and certain related species, and its use to detect and quantify R. solani in soil. Phytopathology 94:280–288
Thornton CR, O’Neill TM, Hilton G, Gilligan CA (1999) Detection and recovery of Rhizoctonia solani in naturally infested glasshouse soils using a combined baiting double monoclonal antibody ELISA. Plant Pathology 48:627–634
Torres MA (2010) ROS in biotic interactions. Physiologia Plantarum 138:414–429
Torres MA, Jones JD, Dangl JL (2005) Pathogen-induced, NADPH oxidase–derived reactive oxygen intermediates suppress spread of cell death in Arabidopsis thaliana. Nature Genetics 37:1130
Tsror L, Barak R, Sneh B (2001) Biological control of black scurf on potato under organic management. Crop Protion 20:145–150
Tsror L (2010) Biology, epidemiology and management of Rhizoctonia solani on potato. Journal of Phytopathology 158:649–658
Tsuda K, Mine A, Bethke G, Igarashi D, Botanga CJ, Tsuda Y, Glazebrook J, Sato M, Katagiri F (2013) Dual regulation of gene expression mediated by extended MAPK activation and salicylic acid contributes to robust innate immunity in Arabidopsis thaliana. PLoS One Genetics 9(12):e1004015
Verma DK, Srivastava S, Mohapatra B, Prakash R, Kumar V, Talukdar D, Yulianto R, Pandey AK, Kumar A, Zuan ATK, Jobanputra AH, Hwang HM, Sahu M, Asthir B (2019) Microbial control: A potential solution for plant disease management in sustainable environments and agriculture. In: Verma DK (ed) Microbiology for sustainable agriculture, soil health, and environmental protection. Apple Academic Press, USA, pp 107–188
Vidhyasekaran P, Ponmalar TR, Samiyappan R, Velazhahan R, Vimala R, Ramanathan A, Paranidharan V, Muthukrishnan S (1997) Host-specific toxin production by Rhizoctonia solani, the rice sheath blight pathogen. Phytopathology 87:1258–1263
Vincelli PC, Beaupre CM-S (1989) Comparison of media for isolating Rhizoctonia solani from soil. Plant Disease 13:1014–1017
Virgen-Calleros G, Olalde-Portugal V, Carling DE (2000) Anastomosis groups of Rhizoctonia solani on potato in central Mexico and potential for biological and chemical control. American Journal of Potato Research 77:219–224
Vleeshouwers VG, Van Dooijeweert W, Govers F, Kamoun S, Colon LT (2000) Does basal PR gene expression in Solanum species contribute to non-specific resistance to Phytophthora infestans? Physiological and Molecular Plant Pathology 57:35–42
Vos P, Hogers R, Bleeker M, Reijans M, Lee T, Hornes M, Frijters A, Pot J, Peleman J, Kuiper M, Zabeau M (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acid Research 23:4407–4414
Walid N, Al-Jaramany L, Elbenay A, Al-Mhethawi R (2022) Biological control of tomato damping off and potato black scurf by seed treatment with Trichoderma harzianum. Jordan Journal of Biological Sciences. 15:373–380
Wang S, Zhang S, Wang W, Xiong X, Meng F, Cui X (2015) Efficient targeted mutagenesis in potato by the CRISPR/Cas9 system. Plant Cell Reports 34:1473–1476
Wastie RL (1994) Inheritance of resistance to fungal diseases of tubers. In: Bradshaw JE, MacKay GR (eds) Potato Genetics. CAB International, Walingford, pp 411–429
Weinhold AR (1977) Population of Rhizoctonia solani in agricultural soils determined by screening procedure. Phytopathology 67:566–569
Weinhold AR, Bowman T, Hall DH (1982) Rhizoctonia disease of potato: effect on yield and control by seed tuber treatment. Plant Disease 66:815–818
Wharton P, Kirk W, Berry D, Snapp S (2007) Rhizoctonia stem canker and black scurf of potato. Michigan Potato Diseases series, Michigan State University, Lansing, MI, pp 1–4 (Ext Bull. E-2994)
Wharton P, Wood E (2013) Rhizoctonia stem canker and black scurf of potato. Agricultural Experiment & UI Extension Publications, Special Collections Idaho S 53 (Between E3 -E415), University of Idaho Library, pp 1–5 (Ext Bull)
Wibberg D, Genzel F, Verwaaijen B, Blom J, Rupp O, Goesmann A, Zrenner R, Grosch R, Puhler A, Schluter A (2017) Draft genome sequence of the potato pathogen Rhizoctonia solani AG3-PT isolate Ben3. Archives of Microbiology 199:1065–1068
Wibberg D, Jelonek L, Rupp O, Hennig M, Eikmeyer F, Goesmann A, Hartmann A, Borriss R, Grosch R, Puhler A, Schluter A (2013) Establishment and interpretation of the genome sequence of the phytopathogenic fungus Rhizoctonia solani AG1-IB isolate 7/3/14. Journal of Biotechnology 167:142–155
Wibberg D, Rupp O, Jelonek L, Krober M, Verwaaijen B, Blom J, Winkler A, Goesmann A, Grosch R, Puhler A, Schluter A (2015) Improved genome sequence of the phytopathogenic fungus Rhizoctonia solani AG1-IB 7/3/14 as established by deep mate-pair sequencing on the MiSeq (Illumina) system. Journal of Biotechnology 203:19–21
Wibberg D, Rupp O, Blom J, Jelonek L, Krober M, Verwaaijen B, Goesmann A, Albaum S, Grosch R, Puhler A, Schluter A (2015) Development of a Rhizoctonia solani AG1-IB specific gene model enables comparative genome analyses between phytopathogenic R. solani AG1-IA, AG1-IB, AG3 and AG8 isolates. PloS One 10:e0144769
Wicks TJ, Morgan B, Hall B (1995) Chemical and biological control of Rhizoctonia solani on potato seed tubers. Australian Journal of Experimental Agriculture 35:661–664
Wilson PS, Ketola EO, Ahvenniemi PM, Lehtonen MJ, Valkonen JPT (2008) Dynamics of soilborne Rhizoctonia solani in the presence of Trichoderma harzianum: effects on stem canker, black scurf and progeny tubers of potato. Plant Pathology 57:152–61
Woodhall JW, Adams IP, Peters JC, Harper G, Boonham N (2013) A new quantitative real-time PCR assay for Rhizoctonia solani AG3-PT and the detection of AGs of Rhizoctonia solani associated with potato in soil and tuber samples in Great Britain. European Journal of Plant Pathology 136:273–280
Woodhall JW, Belcher AR, Peters JC, Kirk WW, Wharton PS (2012) First report of Rhizoctonia solani AG2-2IIIB infecting potato stem and stolon in the United Sates. Plant Disease 96:460
Woodhall JW, Lees AK, Edwards SG, Jenkinson P (2007) Characterization of Rhizoctonia solani from potato in Great Britain. Plant Pathology 56:286–295
Woodhall JW, Lees AK, Edwards SG, Jenkinson P (2008) Infection of potato by Rhizoctonia solani: effect of anastomosis group. Plant Pathology 57:697–905
Xu X, Pan S, Cheng S, Zhang B, Mu D, Ni P, Zhang G, Yang S, Li R, Wang J, Orjeda G, Guzman F, Torres M, Lozano R, Ponce O, Martinez D, Cruz GD, Chakrabarti SK, Patil VU (2011) Genome sequence and analysis of the tuber crop potato. Nature 475:189–195
Yanar Y, Yilmaz G, Cesmeli I, Coskum S (2005) Characterization of Rhizoctonia solani isolates from potatoes in Turkey and screening potato cultivars for resistance to AG-3 isolates. Phytoparasitica 33:370–376
Zeng YX, Ji ZJ, Li XM, Yang CD (2011) Advances in mapping loci conferring resistance to rice sheath blight and mining Rhizoctonia solani resistant resources. Rice Science 18:56–66
Zhang Y, Jin X, Ouyang Z, Li X, Liu B, Huang L, Hong Y, Zhang H, Song F, Li D (2015) Vitamin B6 contributes to disease resistance against Pseudomonas syringae pv. tomato DC3000 and Botrytis cinerea in Arabidopsis thaliana. Journal of Plant Physiology 175:21–25
Zhao C, Zhang X, Hua H, Han C, Wu X (2019) Sensitivity of Rhizoctonia spp. to flutolanil and characterization of the point mutation in succinate dehydrogenase conferring fungicide resistance. European Journal Plant Pathology 155:13–23
Zhao YQ, Wu YH, Zhao XX, An MN, Chen JG (2014) Study on the Taq man real-time PCR to the detection and quantification of Rhizoctonia solani AG3 of Tobacco target spot. Advanced Material Research 1010:80–83
Zheng A, Lin R, Zhang D, Qin P, Xu L, Ai P, Ding L, Wang Y, Chen Y, Liu Y, Sun Z, Feng H, Liang X, Fu R, Tang C, Li Q, Zhang J, Xie Z, Deng Q, Li S, Wang S, Zhu J, Wang L, Liu H, Li P (2013) The evolution and pathogenic mechanisms of the rice sheath blight pathogen. Nature Communication 4:1424
Zimudzi J, Coutinho TA, van der Waals JE (2017) Pathogenicity of fungi isolated from atypical skin blemishes on potatoes in South Africa and Zimbabwe. Potato Research 60:119–144
Zrenner R, Genzel F, Verwaaijen B, Wibberg D, Grosch R (2020) Necrotrophic lifestyle of Rhizoctonia solani AG3-PT during interaction with its host plant potato as revealed by transcriptome analysis. Scientific Report 10:12574
Funding
The authors have no relevant financial or non-financial interests to disclose.
Author information
Authors and Affiliations
Contributions
ML and SC suggested the research topic, planned the research methodology and wrote the original draft. SS and SC investigated the articles and drew the figures. SS and MK participated in data representation and article revising and editing. All authors read and approved the final article.
Corresponding author
Ethics declarations
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chaudhary, S., Lal, M., Sagar, S. et al. Black scurf of potato: Insights into biology, diagnosis, detection, host-pathogen interaction, and management strategies. Trop. plant pathol. 49, 169–192 (2024). https://doi.org/10.1007/s40858-023-00622-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40858-023-00622-4