Abstract
Elite indica rice cultivars were cotransformed with genes expressing a rice chitinase (chi11) and a thaumatin-like protein (tlp) conferring resistance to fungal pathogens and a serine-threonine kinase (Xa21) conferring bacterial blight resistance, through particle bombardment, with a view to pyramiding sheath blight and bacterial blight resistance. Molecular analyses of putative transgenic lines by polymerase chain reaction, Southern Blot hybridization, and Western Blotting revealed stable integration and expression of the transgenes in a few independent transgenic lines. Progeny analyses showed the stable inheritance of transgenes to their progeny. Coexpression of chitinase and thaumatin-like protein in the progenies of a transgenic Pusa Basmati1 line revealed an enhanced resistance to the sheath blight pathogen, Rhizoctonia solani, as compared to that in the lines expressing the individual genes. A transgenic Pusa Basmati1 line pyramided with chi11, tlp, and Xa21 showed an enhanced resistance to both sheath blight and bacterial blight.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice (Oryza sativa L.) is one of the most important food crops, from which nearly one third of world’s population derive its principal source of calories. Rice yield has been affected by several abiotic and biotic factors, of the latter, the pests and diseases are important. Among various diseases affecting rice, sheath blight (ShB) caused by Rhizoctonia solani Kühn and bacterial blight (BB) caused by Xanthomonas oryzae pv. oryzae (Xoo) (Ishyama) Dye are highly pernicious. Estimated yield losses caused by sheath blight ranges from 8 to 50% (Savary et al. 2000), while the losses due to bacterial blight could reach up to 50%, when the pathogen infects the rice plants at maximum tillering stage (Adhikari et al. 1995). Owing to the inherent limitations of conventional breeding and environmentally hazardous nature of chemical pesticides, alternative strategies are being tried globally to enhance the resistance of crops against invading pathogens. Genetic engineering is a promising strategy to generate crop plants with disease resistance against economically important plant diseases like rice sheath blight and bacterial blight.
Conventionally, genetic engineering signifies genetic transformation of the crop with a single gene associated with the trait of interest (Ronald 1997; Zhang et al. 1998). Since disease resistance could rely on highly complex multigenic trait, single gene transformations, in general, result in insufficient and/or narrow spectrum disease resistance (Neuhans et al. 1991; Anand et al. 2003) and there is always a possibility for reversal of resistance because of appearance of resistant strains of pathogens (Mew et al. 1992). Hence, genetic engineering of crop plants with (i) a combination of genes encoding/controlling interdependent or synergistic subcomponents of the disease-resistant trait to realize effective resistance against a particular disease or (ii) a combination of genes associated with different diseases to realize a wide-spectrum disease resistance would be more logical. An ingeniously planned genetic engineering involving a well-balanced expression of transgenes with different modes of action would ensure enhanced and durable resistance against pathogens. Availability of cloned disease resistance (Song et al. 1995) or defence genes (Huang et al. 1991; Velazhahan et al. 1998; Takakura et al. 2000) and enhanced transformation tools and strategies (Christou 1997; Chen et al. 1998; Kim et al. 2003) has made pyramiding of disease resistance genes in crop plants much easier. The capability of rice genome to integrate, coexpress, and inherit several transgenes, without undergoing developmentally and reproductively unfavorable phenotypic changes (Chen et al. 1998), highlights the potential of rice germplasm to be improved toward effective, stable, and inheritable disease resistance without compromising yield significantly. The success of multiple gene transformation or gene pyramiding is largely dependent upon the judicious selection of the genotypes of the crop to be transformed and the genes to be employed. The combination of genes encoding proteins involved in pathogen recognition and/or subsequent activation of signaling pathways leading to defence responses (Van Loon and Van Strien 1999) and those exhibiting direct antimicrobial properties (Broglie et al. 1991; Yun et al. 1998) by virtue of their degradative interaction with pathogen cell wall and cell membrane can be expected to improve the scope for generating crop plants with specific or wide-spectrum resistance against pathogens.
Conventional breeding (Yoshimura et al. 1995) and marker-assisted breeding (Huang et al. 1997; Singh et al. 2001) strategies achieved bacterial blight (BB) resistance genes-pyramided rice lines displaying wide-spectrum resistance against Xoo isolates/races (Kameswara Rao et al. 2002). Conventional crossing of two independent transgenic homozygous rice lines by Datta et al. (2002) pyramided Xa21 gene, a chitinase gene, and a Bt-fusion gene into IR72 to confer multiple resistances against BB, ShB, and yellow stem borer. Combination of marker-assisted breeding (MAS) and genetic transformation yielded rice lines resistant to blast and BB by pyramiding Pi1, Piz5 (major blast resistance genes), and Xa21 (Narayanan et al. 2004). Genetic engineering of rice by Kim et al. (2003) to pyramid a maize ribosome-inactivating protein gene and a rice basic chitinase gene enhanced the ShB resistance of the transgenic rice lines. The pyramiding of Xa21 and gna (Galanthus nivalis agglutinin) genes in rice by genetic transformation produced Xa21 and gna coexpressing rice lines displaying BB and brown plant hopper resistance (Tang et al. 1999). Recently, Kalpana et al. (2006) showed coexpression of transgenic rice chitinase and thaumatin-like protein (TLP) in an elite indica rice line, which showed significantly higher level of ShB resistance than the chitinase or TLP transformants.
The present programme is concerned with pyramiding the disease resistance genes rice chi11, tlp, and Xa21 in a few indica rice cultivars to confer effective, stable, and inheritable resistance against ShB and BB. Xa21, a major dominant gene for BB resistance, originally transferred from the wild rice O. longistaminata into the cultivar IR24 by Khush et al. (1990) and cloned by Ronald and coworkers (Ronald et al. 1992; Song et al. 1995), confers broad-spectrum resistance against diverse Xoo isolates/races in rice (Song et al. 1995; Wang et al. 1996; Zhang et al. 1998; Tu et al. 1998, 2000). Based on models from mammalian systems, Xa21 has been suggested to play an important role in cell surface recognition of a pathogen ligand and subsequent activation of intracellular serine-threonine kinase(s) leading to a defense response (Song et al. 1995).
The other genes used in the study, the rice chi11 and tlp encode, respectively, an enodochitinase and a TLP categorized as important pathogenesis-related (PR)-proteins (Van Loon and Van Strien 1999). PR-proteins accumulate in plant cells along with other defence-related molecules during plant’s hypersensitive response (HR; Heath 2000) against pathogen attack. PR-proteins are classified on the basis of their amino acid sequences, serological relationships, and biochemical functions (Van Loon and Van Strien 1999) and several members of the PR-protein categories are well studied by various groups (Muthukrishnan et al. 2001). Different endochitinases (EC 3.2.1.14) belonging to PR-3, 4, 8, and 11 categories (Van Loon and Van Strien 1999) catalyze the hydrolysis of β-1,4 linkages between N-acetylglucosamine units of chitin, the major polysaccharide component of fungal cell wall (Broglie et al. 1991). A few chitinase cDNAs were isolated from rice (Huang et al. 1991; Takakura et al. 2000) and have been employed to transform rice cultivars to engineer ShB resistance (Lin et al. 1995; Datta et al. 2001; Kumar et al. 2003, Kalpana et al. 2006). TLP, belonging to PR-5 category, causes fungal cell membrane lysis and death by altering the cell membrane permeability (Yun et al. 1998). Several TLP genes were isolated from cereals including rice (Velazhahan et al. 1998). The transgenic rice plants overexpressing a rice tlp transgene have been reported to exhibit an enhanced resistance to ShB pathogen (Datta et al. 1999; Kalpana et al. 2006). A subgroup of TLPs has been shown to be β-1,3-glucan binding proteins (Osmond et al. 2001). Moreover, Grenier et al. (1999) showed the hydrolysis of β-1,3-glucans by some TLPs. The existing literature on the plant antifungal PR-proteins implies that the fungal cell membrane-permeabilizing TLPs and the fungal cell wall hydrolyzing carbohydrases, such as chitinases and β-1,3-glucanases act in concert to kill fungal pathogens. Chitinases, TLPs, and β-1,3-glucanases are coregulated/coexpressed developmentally (Peumans et al. 2002) when the plants are subjected to abiotic stress (Hiilovaara-Teijo et al. 1999) and challenged by pathogens (Jacobs et al. 1999). Hejgaard et al. (1991) reported synergistic activity of barley TLP and chitinase against the fungi Trichoderma viride and Candida albicans. Recent attempt by Kalpana et al. (2006) to genetically engineer elite indica rice cultivars with rice chitinase and TLP suggested their synergistic expression in transgenic context and the resultant enhanced resistance to ShB.
In the present study, high yielding elite indica rice cultivars PB1, ASD16, ADT38, IR72, and White Ponni were cotransformed with chi11, tlp, and Xa21 genes using immature embryos and mature seed-derived calli as explants. To ascertain the integration of transgenes in the host genome and their expression, molecular analyses of transgenic lines were conducted by polymerase chain reaction, Southern Blot hybridization, and Western Blotting analyses. Inheritability of coexpression of the transgenes was monitored till T2 generation of the transgenic lines. The transgenic individuals were rigorously assayed for transgenic resistance against ShB and BB.
Materials and methods
Cotransformation of rice cultivars through particle bombardment
Transformation vectors used
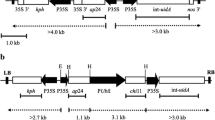
Plant transformation vectors, (i) pMKU-RF2 (Kumar et al. 2003) harboring a 3.2 kbp chitinase gene expression cassette containing a 1.1 kbp rice chi11 gene under the control of ubiquitin promoter and NOS polyA terminator, (ii) pGL2-ubi-tlp (Datta et al. 1999) harboring a 3.1 kbp TLP gene expression cassette containing a 1.1 kbp rice tlp gene under the control of a ubiquitin promoter and NOS terminator, and (iii) pC822 (Song et al. 1995) harboring a 9.6 kbp rice genomic DNA insert containing the Xa21 gene expressing a receptor kinase-like protein conferring bacterial blight resistance were used in the cotransformation experiments. A 10 μg equimolar mixture of the three transformation vectors was used for coating 5 mg of gold particles (Bio-Rad Laboratories, USA), as per the manufacturer’s instructions.
Particle bombardment of immature embryos
Immature seeds of the elite indica rice cultivars ASD16, ADT38, IR72, IR64, and White Ponni, harvested 12–14 days postanthesis from Paddy Breeding Station (PBS), Tamil Nadu Agricultural University (TNAU), Coimbatore, India, were manually dehusked and surface-sterilized with 0.1% mercuric chloride for 5 min. The immature embryos isolated aseptically under a stereomicroscope (Leica, Switzerland) were plated with their scutella up (Datta et al. 2001) onto CC medium (Potrykus et al. 1979) containing 2.0 mg l−l 2,4-D and cultured for 2 days in the dark at 25±2°C. After 2 days, precultured embryos were bombarded twice using PDS-1000/He Biolistic Particle Delivery System (Bio-Rad Laboratories, Hercules, USA) at 1100 psi and 27 in. of Hg vacuum, according to Zhang et al. (1998). The putative transformants were selected and regenerated on a CC-based medium containing 50 mg l−l hygromycin B (Boehringer Mannheim GmbH, Germany).
Particle bombardment of mature seed-derived calli
Manually dehusked seeds of PB1 (obtained from PBS, TNAU) were surface sterilized with 70% ethanol for 3 min followed by 0.1% mercuric chloride for 5 min. After washing with sterile water, the seeds were plated onto MS (Murashige and Skoog 1962) medium containing 2.5 mg l−l 2,4-D and incubated in the dark at 25±2°C for 3 weeks. Embryogenic calli developing from the seeds were subcultured onto fresh medium and allowed to proliferate for another 2 weeks. Hard friable calli were selected and cut into small pieces of 2–3 mm size, 3–4 days prior to bombardment. Calli were bombarded twice following standard protocols (Zhang et al. 1998). After three rounds of selection on hygromycin B (50 mg l−l), plants were regenerated on MS medium supplemented with 3.0 mg l−1 BAP (BA; N6-benzyladenine; Sigma, USA) and 1.5 mg l−l NAA (α-naphthalene acetic acid).
Molecular and biochemical analyses of putative (T0) transgenic lines
Polymerase chain reaction
Genomic DNA was isolated from putative transgenic plants and nontransgenic control plants, following Dellaporta et al. (1983) and PCR was performed to amplify Xa21 gene. Forward primer U1 corresponding to the 3′ untranslated region of the Xa21 gene (5′-CGATCGGTATAACAGCAAAAC-3′) and reverse primer I1 (5′-ATAGCAACTGATTGCTTGG-3′) corresponding to the center of the intron (9.6 kb KpnI fragment of Xa21 gene has a single large ORF of 3075 bp intervened by a 843 bp intron; Song et al. 1995) were used to amplify a 1.4 kbp-long internal fragment of Xa21, as described by Wang et al. (1996). Amplification was performed in a PTC-100TM Programmable Thermal Controller (MJ Research, Inc., USA).
Southern Blot hybridization analysis
Ten micrograms of genomic DNA extracted from putative transgenic plants and nontransgenic control plants and 5 ng of plasmid DNAs with the transgenes of the study (positive control) were digested overnight with appropriate restriction endonucleases (Bangalore Genei Pvt. Ltd., India). The products were electrophoresed on 1.0% (w/v) agarose gels and then transferred onto nylon membranes (Hybond+, Boehringer and Mannheim, UK). The blots were subsequently hybridized overnight with radiolabeled probes prepared by random primer oligo labeling method (Random Primer Labeling Kit, Bangalore Genei Pvt. Ltd., India). Hybridization and autoradiography were carried out following standard procedures (Sambrook et al. 1989)
To demonstrate the transgenic integration of chi11, the genomic DNAs were digested with HindIII to release the 3.2 kbp chitinase expression cassette and blotted onto the membrane. The blot was hybridized with α-32P dCTP-labeled 1.08 kbp chi11 coding sequence obtained by digesting pMKU-RF2 with PstI. To confirm the transgenic integration of tlp gene, the genomic DNAs were digested with HindIII to release the 3.1 kbp TLP expression cassette and hybridized with α-32P dCTP-labeled 1.1 kbp TLP coding sequence obtained by digesting 3.1 kbp TLP expression cassette with BamHI. Similarly, to confirm the transgenic integration of Xa21, the genomic DNAs were digested with EcoRV to release a 3.8 kbp (containing most of the coding sequence of the Xa21; Wang et al. 1996) internal fragment of Xa21 (9.6 kbp) and hybridized with α-32P dCTP-labeled 3.8 kbp Xa21 coding sequence obtained by digesting pC822 with EcoRV.
Western Blotting analysis
Leaves (250 mg) from putative transgenic plants and nontransgenic control plants were ground with 150 μl of ice-cold phosphate-buffered saline. The extracts were centrifuged at 12000 rpm for 10 min at 4°C. The supernatants were collected and the proteins were resolved by sodium dodecyl sulphate-polyacrylamide gel (12%) electrophoresis (Mini-PROTEAN®II cell, Bio-Rad Laboratories, USA), following standard procedure (Laemmli 1970).
Western Blotting analysis was carried out following the method described by Gallagher et al. (1995). Fractionated proteins were transferred onto a nitrocellulose membrane (Protran BAS 5 Cellulosenitrat, Schleicher and Schuell, Germany) using a Trans-Blot® SD Semi-Dry Transfer Cell (Bio-Rad Laboratories, USA). The membranes were then probed with either antibarley chitinase antibody (gifted by Dr. S. Muthukrishnan, Kansas State University, USA) or antitobacco TLP antibody (gifted by Dr. Legrand, Strasbourg, Cedex, France). The polypeptides recognized by the specific antibodies were detected using a goat antirabbit IgG-alkaline phosphatase conjugate by incubating the membrane in the dark with 5-bromo-4-chloro-3-indolyl phosphate/nitroblue tetrazolium (BCIP/NBT) liquid substrate system (Sigma-Aldrich Co., USA).
Cotransformation efficiency
Cotransformation efficiency refers to the ratio of number of putative transgenic lines harboring/expressing more than one gene of interest (at DNA level; as estimated by PCR or Southern analysis or at protein level; as estimated by Western analysis) to the total number of explants bombarded.
Progeny analysis
Of the three T0 PB1 lines expressing all the three transgenes (chi11, tlp, and Xa21), a morphologically normal and fertile line SM-PB1-1 was forwarded to generate T1 progeny. The T0 lines forwarded for T1 generation also included PB1 lines expressing either chi11 or tlp and an ASD16 line expressing Xa21. Nurseries were raised with the seeds collected from five T0 PB1 lines viz., SM-PB1-1 (expressing chi11, tlp, and Xa21), SM-PB1-5 (expressing tlp), SM-PB1-9, 12, and 13 (expressing chi11), and an ASD16 line, SM-ASD16-2 (expressing Xa21). The resulting T1 progenies of these transgenic lines were analyzed by PCR and/or Western Blotting analyses. Plants of T1 progeny showing stable integration and expression of the transgenes were forwarded to generate T2 progeny and the plants were analyzed by PCR and/or Western Blotting analyses.
Evaluation of ShB and BB resistance in T1 and T2 progeny
Bioassay for ShB resistance
The PB1 transgenic lines SM-PB1-1 (expressing chi11, tlp, and Xa21), SM-PB1-5 (expressing tlp), and SM-PB1-9 (expressing chi11) were evaluated for ShB resistance along with suitable nontransgenic control plants. Two different ShB assay methods viz., “bioassay using detached leaves” and “bioassay using leaf sheaths intact” were employed for screening the transgenic lines, as described by Kumar et al. (2003). All the inoculations were carried out with 5 mm mycelial discs obtained from 3-day-old Rs7 (a virulent isolate of R. solani; Krishnamurthy et al. 1999) maintained on potato dextrose agar (250.0 g l−l potato, 20.0 g l−l dextrose, 20.0 g l−l agar, pH 7.0).
In bioassay using detached leaves, observations were made at 24 h intervals and infection cushions were counted under a stereomicroscope, 72 h after inoculation (HAI). In bioassay using leaf sheaths intact, observations were made up to 7 days at 24 h intervals and functional resistance/susceptibility index (FRI) was calculated. At 168 HAI, the highest relative lesion height percentage (HRLH%; http//www.riceweb.org/ses/cd_diseases_d.htm) and the total lesion spread were estimated. HRLH% was calculated using the formula,
Bioassay for BB resistance
Preparation of bacterial suspension
A virulent isolate of Xoo (Tamil Nadu Rice Research Institute, Aduthurai, India) maintained on PSA (5.0 g l−l peptone, 20.0 g l−l sucrose, 0.5 g l−l Ca(NO3)2·4H2O, 0.5 g l−l FeSO4·7H2O, 2.0 g l−l Na2HPO4·2H2O, 20.0 g l−l agar) was used as source of inoculum. A loopful of bacteria was streaked onto a PSA slant and allowed to grow for 48 h at 28°C. Bacterial suspension was prepared by adding sterile distilled water (10 ml) into the 48 h-old culture and adjusting bacterial population to 109 cells/ml. All inoculations were made within 30–60 min after preparing the bacterial suspension.
Inoculation by leaf clipping
The PB1 line SM-PB1-1 (expressing chi11, tlp, and Xa21) and the ASD16 line, SM-ASD16-2 (expressing Xa21) maintained in a greenhouse for transgenic plants were evaluated for BB resistance. Leaf clipping method (Kauffman et al. 1973) was employed for inoculating 60-to-80-day-old plants (maximum tillering stage). After inoculation, the plants were covered with polyethylene bags with inner surface sprinkled with sterile distilled water to maintain plants under higher relative humidity for better symptom development. From each line, five plants (5–10 leaves/plant) were inoculated along with suitable nontransgenic control plants. Lesion length was measured 14 days after inoculation and based on the length of the developed lesions, the plants were categorized as resistant (1–3 cm), moderately resistant (3–6 cm), and susceptible (>9 cm), as described by Mew and Vera Cruz (1979).
Statistical analyses
All the bioassay experiments were carried out in completely randomized design (CRD) with adequate replications. Duncan’s multiple range test (DMRT; Gomez and Gomez 1984) was used to compare treatment means using the software IRRISTAT Version 3.1 (Biometrices Unit, International Rice Research Institute, Manila, The Philippines).
Results and discussion
One of the major challenges in agriculture worldwide is to control the great yield loss caused by pests and pathogens. Realization of this objective, however, in an environmental-friendly way necessitates integrated efforts from plant breeders, pathologists, and genetic engineers. Despite successful genetic engineering of crop plants for the expression of transgene mediated disease resistance, the exhaustive efforts ended up with crops exhibiting insufficient or narrow-spectrum disease resistance or reversal of disease resistance. As a sequel, stacking or pyramiding of genes emerged as an alternative to engineering monogenic resistance with a sole aim of enhancing the durability of resistance against individual members of the pathodeme (Singh et al. 2001; Datta et al. 2002). In the present study, with the view to conferring higher level resistance against sheath blight along with resistance to bacterial blight, a few elite indica rice cultivars have been transformed with rice chi11, tlp, and Xa21 genes. The transformants were molecularly characterized for the stable and inheritable coexpression of the transgenes and rigorously assayed for the associated resistance against ShB and BB.
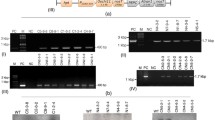
Integration of Xa21 transgene in the putative T0 transgenic lines was ascertained by PCR as well as Southern Blot hybridization analysis. PCR of DNA from transgenic plants with Xa21 kinase domain- and intron-specific primers revealed amplification of the expected 1.4 kbp fragment from four lines of the cultivar PB1 (SM-PB1-1, 2, 3, and 7), one line of the cultivar ASD16 (SM-ASD16-2), one line of the cultivar ADT38 (SM-ADT38-1), and two lines of the cultivar White Ponni (SM-WP-1 and 2) (Fig. 1). Apart from the transgene-specific 1.4 kbp band, a common band of 1.3 kbp was observed in both transgenic and nontransgenic plants. Similar banding pattern was reported earlier from PCRs of Xa21 transgenic lines performed with the same set of primers (Wang et al. 1996; Tu et al. 1998) and the product was attributed to a cross-amplification of a member of the Xa21 multigene family. Southern Blot hybridization analyses of EcoRV-digested DNA of transgenic lines with 3.8 kbp Xa21 coding sequence probe (released from the transformation vector pC822 by EcoRV digestion) revealed a 3.8 kbp hybridization signal from the transgenic lines SM-PB1-1 and 7, SM-ADT38-1, SM-WP-1 and 2, SM-ASD16-2, while such signal was absent from nontransgenic control plants. In addition, nonendogenous transgene-specific bands of different sizes were also observed in some of the transgenic lines (Fig. 2). These transgene-specific bands may be due to transgene rearrangement during integration (Tang et al. 1999; Datta et al. 2002). Moreover, a high molecular weight (>9.0 kbp) band was observed in both the transgenic and nontransgenic lines. This signal may be corresponding to the one reported earlier to represent the polymorphic family members of the Xa21 gene family (Wang et al. 1996; Tu et al. 1998, 2000; Tang et al. 1999).
Southern Blot hybridization analysis of HindIII-digested genomic DNA of transgenic lines with 1.08 kbp chi11 coding sequence detected a 3.2 kbp hybridization signal representing the HindIII-released chi11 gene expression cassette from four transgenic lines (SM-PB1-1, 9, 12, and 13) of the cultivar PB1 and a transgenic line of the cultivar ADT38 (SM-ADT38-1) revealing a stable integration of chi11 transgene expression cassette in their genomes. In addition to this, two low molecular weight bands (<3.2 kbp) were detected in both transgenic and nontransgenic lines (Fig. 3). Lin et al. (1995) used chi11 coding sequence as probe to hybridize the HindIII-digested genomic DNA of transgenic and nontransgenic plants and observed two additional hybridization signals in both transgenic and nontransgenic lines, apart from the expected band in transgenic plants.
Southern Blot hybridization analysis of HindIII-digested DNA of transgenic plants with 1.1 kbp tlp coding sequence identified a hybridization signal of 3.1 kbp representing the 3.1 kbp tlp expression cassette released by HindIII digestion from five independent transgenic lines of the cultivar PB1 (SM-PB1-1, 5, 6, 7, and 8) and two lines of the cultivar White Ponni (SM-WP-1 and 2). Apart from this, a common band was found to be associated with the putative transgenic and nontransgenic plants (Fig. 4).
The expression of chi11 and tlp transgenes in transgenic T0 lines were studied by Western Blotting analyses of their total soluble leaf protein extracts against antibarley chitinase antiserum and antitobacco TLP antiserum, respectively. The antibarley chitinase antiserum detected the expression of a 35 kDa polypeptide in six lines of the cultivar PB1 (SM-PB1-1, 2, 3, 9, 12, and 13), two lines of the cultivar ADT38 (SM-ADT38-1 and 2), and one line of the cultivar ASD16 (SM-ASD16-3). Apart from the expected 35 kDa signal corresponding to chitinase polypeptide, a 30 kDa signal was also found to be associated with all the transgenic lines (Fig. 5). This 30 kDa polypeptide may be a truncated version of the original 35 kDa protein released by proteolytic processing of the latter (Lin et al. 1995). Both these signals were not detected from nontransgenic control plants. Similarly, antitobacco TLP antibody was able to detect the expression of a 23 kDa polypeptide in seven T0 lines of the cultivar PB1 (SM-PB1-1, 2, 3, 5, 6, 7, and 8), one line of the cultivar ASD16 (SM-ASD16-1), and two lines of the cultivar White Ponni (SM-WP-1 and 2) (Fig. 6). Earlier, the expression of TLP (23 kDa) in transgenic lines of the indica rice cultivars Chinsurah Boro II, IR72, and IR501500 was demonstrated by Datta et al. (1999).
Results of the molecular analyses (PCR, Southern Blot hybridization analysis and Western Blotting analysis) of the T0 lines showed the integration and expression of all the three genes (chi11, tlp, and Xa21) in at least three lines of the cultivar PB1 (SM-PB1-1, 2, and 3), tlp and Xa21 in one line of the cultivar PB1 (SM-PB1-7) and two lines of White Ponni (SM-WP-1 and 2) and chi11 and Xa21 in one line of the cultivar ADT38 (SM-ADT38-1) (Fig. 1–6).
In our experiments, cotransformation efficiency (with respect to the genes of interest) of 1.7, 0.20, and 0.54% was observed in PB1, ADT38, and White Ponni, respectively (Table 1). Previously, a cointegration frequency of >70% was reported in japonica cultivars (Chen et al. 1998; Tang et al. 1999). However, low frequency of transformation is common among indica cultivars. Christou et al. (1991) reported a transformation efficiency of 0.80 and 3.20% in IR72 and IR36, respectively, using immature embryos as explants. The low frequency of transformation may be attributed to the recalcitrance posed by indica cultivars in getting transformed (Sivamani et al. 1996) or to the transgene rearrangement during integration (Tang et al. 1999; Datta et al. 2002).
The progeny (T1) analysis of SM-PB1-1, a transgenic line of the cultivar PB1 that showed stable expression of all the three (chi11, tlp, and Xa21) genes was carried out along with four other PB1 (SM-PB1-5, 9, 12, and 13) lines expressing either chi11 or tlp and one line of the cultivar ASD16 (SM-ASD16-2) expressing Xa21. The analysis showed that the transgenes were inherited as a single Mendelian trait (3:1 ratio) in all the lines (data not shown), except the line SM-PB1-1 carrying chi11, tlp, and Xa21 genes. A 3:1 segregation ratio was observed for chi11 and tlp, as against a 1:1 ratio for Xa21 (Table 2). Tang et al. (1999) observed such a distorted segregation pattern in particle bombardment-mediated cotransformation experiments involving four transgenes (gna, Xa21, gusA, and hpt) and correlated it with two independent integration events (integration at two unlinked loci) involving all the four genes. Gahakwa et al. (2000) encountered such an unusual segregation pattern in 15% of the transgenic rice lines analysed. Several other groups have also reported such a distorted segregation ratio in transgenic rice lines (Peng et al. 1995; Fu et al. 2000). Progeny analysis by PCR/Western Blotting of T2 generation of the earlier-described lines showed stable inheritance and expression of transgenes. Though segregation of transgenes in the progeny was evident (Fig. 7; Table 2), no significant variation in the levels of chitinase and TLP expression between T1 and T2 progenies was noticed (data not shown). The previous attempt by Kalpana et al. (2006) to coexpress chi11 and tlp in the indica rice line ended up in silencing of chitinase transgene in the T1 generation.
ShB and BB resistance of the transgenic lines were assessed by bioassays performed with individuals of T1 and T2 generations. For the bioassay, the individuals were selected only on the basis of the level of expression of transgenic chitinase and TLP (as assessed by Western Blotting), since the previous results did not evidence any correlation between homozygosity and disease resistance, though the transgene expression level was found to influence the resistance (Kim et al. 2003).
In the present study, bioassay for ShB resistance using detached leaves showed the appearance of Rs7 infection cushions away from the site of inoculation of Rs7, probably because of the pathogen’s development of new hyphae, 48 HAI. However, the number of infection cushions formed was significantly lesser in the leaves of PB1 lines expressing either chi11 or tlp, as compared to their respective nontransgenic controls. In the PB1 (SM-PB1-1) line expressing both chi11 and tlp, the number of infection cushions was even lesser than that in the lines expressing either chi11 or tlp (Table 3). Apart from the reduction in the number of infection cushions, the leaves of transgenic plants also exhibited reduction in the size of the lesions with characteristic browning around the lesions, signifying effective restriction of pathogenic invasion (Fig. 8A–C). In the leaves of nontransgenic controls, light grayish lesions (1.5–2.0 cm) with thin brown borders were produced. The gradual enlargement of these grayish lesions caused morbid yellowing and subsequent drying of the infected leaf, 72 HAI (Fig. 8D).
In the bioassay on intact leaf sheaths, the transgenic PB1 lines did not show any disease lesion till 48 HAI, whereas in the nontransgenic PB1 plants, blanch lesions with a thin brown border were formed within this time period. These lesions slowly turned grayish and enlarged in size covering the entire leaf sheath within 168 HAI (Fig. 9D). Contrastingly, in all the transgenic lines (SM-PB1-9, 5, and 1) expressing chi11 and/or tlp, much smaller-sized oblong grayish lesions surrounded by thick brown borders appeared (Fig. 9A–C). A precocious browning at the site of inoculation was observed only in the transgenic plants and such a browning might play a major role in arresting the pathogen spread. This was obvious from the drying of infected leaf sheaths in untransformed control plants, while leaf sheaths of transgenic plants expressing chi11 and/or tlp remained green despite ShB infection (data not shown). This clearly demonstrated that the pathogen spread was very much restricted in transgenic lines possibly due to an extensive browning at the site of inoculation, a defense response very similar to hypersensitive reaction. Groth and Nowick (1992) suggested that the resistance to the spread of R. solani in rice could be consequent to production of oxidised phenolics (dark zone around lesions). Yamamato et al. (2000) observed an extensive browning and necrotic symptoms on transgenic grapevine plants expressing a rice chitinase (RCC2) gene leading to a hypersensitive reaction against powdery mildew pathogen Uncinula necator. The rapid cell death consequent to pathogen invasion was considered to be a common expression of resistance/defence in plants against the invading pathogen (Heath 2000).
Evaluation of ShB resistance using FRI revealed that there was a significant rise in FRI of transgenic PB1 (SM-PB1-9, 5, and 1) lines over nontransformed ones at 72, 96, 120, 144, and 168 HAI. Such an increase in FRI over control was maximum (344% in SM-PB1-9, 267% in SM-PB1-5, and 482% in SM-PB1-1) at 120 HAI. Similar increase in FRI of PB1 line SM-PB1-1 expressing both chi11 and tlp was 253%, as compared to 168% and 159% in SM-PB1-9 (chi11) and SM-PB1-5 (tlp) respectively, at 168 HAI (Fig. 10). Kumar et al. (2003) observed a gradual increase in FRI values in untransformed lines, whereas a gradual initial increase followed by steep rise in FRI values in transgenic rice lines expressing chi11 was reported.
Similarly, transgenic PB1 plants expressing chi11 and/or tlp recorded a lower HRLH% while restricting lesion spread, when compared to untransformed PB1 (Table 4). Total lesion spread on leaf sheath of the transgenic line SM-PB1-1 (expressing chi11 and tlp), 168 HAI was 4.90 cm as compared to 7.94 and 8.04 cm in SM-PB1-9 (expressing chi11) and SM-PB1-5 (expressing tlp) respectively, while total lesion spread on nontransgenic PB1 was 15.5 cm. Reduction in HRLH% and total lesion spread was more significant in the transgenic PB1 line (SM-PB1-1) expressing both chi11 and tlp than the lines expressing either chi11 or tlp (Table 4).
In the recent years, there are several successful reports on improved fungal disease resistance of crop plants by the expression of antifungal proteins like chitinases, glucanases, and TLPs (Broglie et al. 1991; Zhu et al. 1994; Lin et al. 1995; Datta et al. 1999; Anand et al. 2003; Kumar et al. 2003; Kalpana et al. 2006). Lin et al. (1995) reported development of lesions within 3–4 days after inoculation with R. solani on both untransformed and transgenic rice plants expressing rice chitinase (chi11) gene. However, they observed a reduction in number and size of lesions in the transgenic plants, when compared to untransformed controls. Though the lesions spread invasively throughout the leaf in control plants, they were restricted to the lower half of the leaf in transgenic plants. Datta et al. (1999) observed a mean sheath infection density of 8.2–19.1% and 36.6% in the transgenic rice plants expressing TLP and control plants, respectively. Similarly, potato plants expressing higher levels of osmotin-like protein (a member of the PR-5 group) were reported to confer an increased tolerance to the late blight fungus, Phytophthora infestans (Zhu et al. 1996).
Transgenic coexpression of more than one PR-proteins could confer enhanced resistance than the expression of single PR-proteins. In the present study, it was observed that the transgenic line SM-PB1-1 expressing both chi11 and tlp showed enhanced resistance to ShB pathogen than the lines expressing either of these genes (Fig. 8–10; Tables 3 and 4). Leah et al. (1991) demonstrated the synergistic antifungal activities of chitinases and β-1,3-glucanases under laboratory conditions. Zhu et al. (1994) studied the constitutive coexpression of a rice chitinase (RCH10) and an alfalfa acidic glucanase (AGLU1) in transgenic tobacco plants and reported that combination of the two PR-proteins enhanced significantly the protection against the fungal pathogen Cercospora nicotianae, causal agent of frogeye leafspot, than by individual transgenes. Similarly, evidence for synergistically enhanced protection in transgenic tomato (Jongedijk et al. 1995) and wheat (Anand et al. 2003) plants by the coexpression of chitinase and β-1,3-glucanase was reported. Kalpana et al. (2006) reported the co-expression of chi11 and tlp in an indica rice line and the resultant enhanced ShB resistance. Enhanced resistance observed in the present transgenic lines coexpressing chitinase and TLP may be attributed to the different modes of action of these proteins. Chitinase degrades the fungal cell wall chitin, whereas TLP targets the fungal plasma membrane (Yun et al. 1998). These proteins could act synergistically (Hejgaard et al. 1991) to effectively kill the fungal pathogens. Moreover, ShB assays using detached leaves and leaf sheaths intact in T2 progeny lines SM-PB1-1, 5, and 9 also evidenced stable inheritance of the resistance. There was no significant variation in the number of infection cushions, HRLH%, and total lesion spread between T1 and T2 progeny lines (Tables 3 and 4). The PB1 line expressing both chi11 and tlp conferred better protection against ShB than the lines expressing individual genes.
In bioassay for BB resistance, consequent to clip inoculation with Xoo, BB symptom appeared within 4–5 days after inoculation in both transgenic (SM-PB1-1 and SM-ASD16-2) and control (PB1 and ASD16) plants. Water-soaked lesion which developed from the cut surface spread along the veins of the inoculated leaf presenting a yellowish lesion with wavy margins. However, on 14 days after inoculation, lesion length of two transgenic T1 lines SM-PB1-1 and SM-ASD16-2 were restricted to 1.67 and 1.40 cm, respectively, while in the untransformed PB1 and ASD16 plants the lesions attained a length of 15.25 and 14.79 cm, respectively (Table 5; Fig. 11A and B). Similar trend was observed in the bioassay performed on transgenic T2 progeny lines expressing Xa21 with BB pathogen, Xoo (Table 5). Xa21 is a major dominant gene for rice BB resistance whose multi-isolate/race resistance potential against Xoo has been well established (Song et al. 1995; Wang et al. 1996). Tu et al. (1998) observed much reduced lesions of <3.1 cm in transgenic BB-resistant plants and a Xa21 donor (IRBB21), in comparison to the larger lesions of 13.3–20.3 cm in control plants. Moreover, their Xa21 transgenic IR72 lines performed well also under field conditions (Tu et al. 2000). As the pyramided line SM-PB1-1 did not segregate for Xa21 in both T1 and T2 progenies, the influence of chi11 and/or tlp on Xa21-mediated bacterial blight resistance could not be assessed. Previously, we developed several Xa21 transgenic PB1 lines, the Xoo inoculation of which resulted in the induction of lesions (Ramesh et al. 2001) comparable in size to the ones exhibited by the presently pyramided line. Hence, at this point, the possible existence of any interaction between the defence (chi11 and tlp) genes and the resistance (Xa21) gene could not be explained.
In summary, present study reports a successful incorporation of multiple disease resistance/defense genes in PB1, an indica rice cultivar, which led to an enhanced resistance to an agronomically important fungal pathogen and a bacterial pathogen. Transgenic lines obtained in this study add variability to the existing germplasm and thus paving way for an environmental-friendly means to achieve a better disease management. Intensification of research in these lines with a view to enriching the available germplasm is sine quo non for future rice breeding programmes that aim at introgressing multiple disease resistance in elite cultivars.
Abbreviations
- BB:
-
bacterial blight
- FRI:
-
functional resistance/susceptibility index
- HRLH%:
-
highest relative lesion height %
- PB1:
-
Pusa Basmati1
- ShB:
-
sheath blight
References
Adhikari TB, Vera Cruz CM, Zhang Q, Nelson RJ, Skinner DZ, Mew TW, Leach JE (1995) Genetic diversity of Xanthomonas oryzae pv. oryzae in Asia. Appl Environ Microbiol 61:966–971
Anand A, Zhou T, Trick HN, Gill BS, Bockus WW, Muthukrishnan S (2003) Greenhouse and field testing of transgenic wheat plants stably expressing genes for thaumatin-like protein, chitinase and glucanase against Fusarium graminearum. J Exp Bot 54:1101–1111
Broglie K, Chet I, Holliday M, Cressman R, Biddle P, Knowlton S, Mauvais CJ, Broglie R (1991) Transgenic plants with enhanced resistance to the fungal pathogen Rhizoctonia solani. Science 254:1194–1197
Chen L, Marmey P, Taylor NJ, Brizard JP, Espinoza C, Cruz P, Huet H, Zhang S, de Kochko A, Beachy RN, Fauquet CM (1998) Expression and inheritance of multiple transgenes in rice plants. Nat Biotechnol 16:1060–1064
Christou P (1997) Rice transformation: bombardment. Plant Mol Biol 35:197–203
Christou P, Ford TL, Kofron M (1991) Production of transgenic rice (Oryza sativa L.) plants from agronomically important indica and japonica varieties via electric discharge particle acceleration of exogenous DNA into immature embryos. Biotechnology 9:957–962
Datta K, Baisakh N, Thet KM, Tu J, Datta SK (2002) Pyramiding transgenes for multiple resistance in rice against bacterial blight, yellow stem borer and sheath blight. Theor Appl Genet 106:1–8
Datta K, Tu J, Oliva N, Ona I, Velazhahan R, Mew TW, Muthukrishnan S, Datta SK (2001) Enhanced resistance to sheath blight by constitutive expression of infection-related rice chitinase in transgenic elite indica rice cultivars. Plant Sci 60:405–414
Datta K, Velazhahan R, Oliva N, Ona I, Mew T, Khush GS, Muthukrishnan S, Datta SK (1999) Over-expression of the cloned rice thaumatin-like protein (PR-5) gene in transgenic rice plants enhances environmental friendly resistance to Rhizoctonia solani causing sheath blight disease. Theor Appl Genet 98:1138–1145
Dellaporta ST, Wood J, Hicks JB (1983) A plant DNA minipreparation: Version II. Plant Mol Biol Rep 1:19–21
Fu X, Kohli A, Twyman RM, Christou P (2000) Alternative silencing effects involve distinct types of non-spreading cytosine methylation at a three-gene single-copy transgenic locus in rice. Mol Gen Genet 263:106–118
Gahakwa D, Maqbool SB, Fu X, Sudhakar D, Christou P, Kohli A (2000) Transgenic rice as a system to study the stability of transgene expression: multiple heterologous transgenes show similar behaviour in diverse genetic backgrounds. Theor Appl Genet 101:388–309
Gallagher S, Winston SE, Fuller SH, Hurre JGR (1995) Immunodetection. In: Auubel F, Bernt R, Kingston EE, DMoore D, Seidman JD, Smith JA, John S (eds) Short protocols in molecular biology. Wiley, New York, pp 1040–1048
Gomez KK, Gomez AA (1984) Statistical procedure for agricultural research. Wiley, New York
Grenier J, Potvin C, Trudel J, Asselin A (1999) Some thaumatin-like proteins hydrolyse polymeric β-1,3-glucan. Plant J 19:473–480
Groth DE, Nowick EM (1992) Selection for resistance to rice sheath blight through number of infection cushions and lesion type. Plant Dis 76:721–723
Heath MC (2000) Hypersensitive response-related death. Plant Mol Biol 44:321–334
Hejgaard J, Jacobsen S, Svendsen I (1991) Two antifungal thaumatin-like proteins from barley grain. FEBS Lett 291:127–131
Hiilovaara-Teijo M, Hannukkala A, Griffith M, Yu XM, Pihakaski-Maunsbach K (1999) Snow mold-induced apoplastic proteins in winter rye leaves lack antifreeze activity. Plant Physiol 121:665–674
Huang N, Angeles ER, Domingo J, Magpantay G, Singh S, Zhang G, Kumaravadivel N, Bennet T, Khush GS (1997) Pyramiding of bacterial blight resistance genes in rice: marker assisted selection using RFLP and PCR. Theor Appl Genet 95:313–320
Huang J, Wen L, Swegle M, Tian H, Thin TH, Naylor HM, Muthukrishnan S, Reeck GR (1991) Nucleotide sequence of a rice genomic clone that encodes class II endo chitinase. Plant Mol Biol 16:479–480
Jacobs AK, Dry IB, Robinson SP (1999) Induction of different pathogenesis-related cDNAs in grapevine infected with powdery mildew and treated with ethephon. Plant Pathol 48:325–336
Jongedijk E, Tigelaar H, van Roekel JSC, Bres-Vloemans SA, Dekker I, Van Den Elzen PJM, Cornelissen BJC, Melchers LS (1995) Synergistic activity of β-1,3-glucanases and chitinases enhances fungal resistance in transgenic tomato plants. Euphytica 85:173–180
Kalpana K, Maruthasalam S, Rajesh T, Poovannan K, Kumar KK, Kokiladevi E, Raja JAJ, Sudhakar D, Velazhahan R, Samiyappan R, Balasubramanian P (2006) Engineering sheath blight resistance in elite indica rice cultivars using genes encoding defense proteins. Plant Sci 170:203–215
Kameswara Rao K, Lakshminarasu M, Jena KK (2002) DNA markers and marker-assisted breeding for durable resistance to bacterial blight disease in rice. Biotech Adv 20:33–47
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Khush GS, Bacalangco E, Ogawa T (1990) A new gene for resistance to bacterial blight from Oryza longistaminata. Rice Genet News L 7:121–122
Kim JK, Jang IC, Wu R, Zuo WN, Boston RS, Lee YH, Ahn IP, Nahm BH (2003) Co-expression of a modified maize ribosome-inactivating protein and a rice basic chitinase gene in transgenic rice plants confers enhanced resistance to sheath blight. Transgenic Res 12:475–484
Krishnamurthy J, Samiyappan R, Vidhyasekaran P, Nakkeeran S, Rajeswari E, Raja JAJ, Balasubramanian P (1999) Efficacy of Trichoderma chitinases against Rhizoctonia solani, the rice sheath blight pathogen. J Biosci 24:207–213
Kumar KK, Poovannan K, Nandakumar R, Thamilarasi K, Geetha C, Jayashree N, Kokiladevi E, Raja JAJ, Samiyappan R, Sudhakar D, Balasubramanian P (2003) A high throughput functional expression assay system for a defence gene conferring transgenic resistance on rice against the sheath blight pathogen, Rhizoctonia solani. Plant Sci 165:969–976
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–683
Leah R, Tommerup H, Svendsen I, Mundy J (1991) Biochemical and molecular characterization of three barley seed proteins with antifungal properties. J Biol Chem 266:1564–1573
Lin W, Anuratha CS, Datta K, Potrykus I, Muthukrishnan S, Datta SK (1995) Genetic engineering of rice for resistance to sheath blight. Biotechnology 13:686–691
Mew TW, Vera Cruz CM (1979) Variability of Xanthomonas oryzae: specificity in infection of rice differentials. Phytopathology 69:152–155
Mew TW, Vera Cruz CM, Medalla ES (1992) Changes in the race frequency of Xanthomonas oryzae pv. oryzae in response to the planting of rice cultivars in the Philippines. Plant Dis 76:1029–1032
Murashige T, Skoog FA (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Muthukrishnan S, Liang GH, Trick HN, Gill BS (2001) Pathogenesis-related proteins and their genes in cereals. Plant Cell Tissue Organ Cult 64:93–114
Narayanan NN, Baisakh N, Oliva NP, VeraCruz CM, Gnanamanickam SS, Datta K, Datta SK (2004) Molecular breeding: marker-assisted selection combined with biolistic transformation for blast and bacterial blight resistance in indica rice (cv. CO39). Mol Breed 14:61–71
Neuhans JM, Ahl-Goy P, Hinz U, Flores S, Meins FJ (1991) High-level expression of a tobacco chitinase gene in Nicotiana sylvestris: susceptibility of transgenic plants to Cercospora nicotianae infection. Plant Mol Biol 16:141–151
Osmond RIW, Hrmova M, Fontaine F, Imberty A, Fincher GB (2001) Barley thaumatin-like proteins: ligand specificity, kinetic and structural analyses and biological implications. Eur J Biochem 268:4190–4199
Peng JY, Wen FJ, Lister RL, Hodges TK (1995) Inheritance of gusA and neo genes in transgenic rice. Plant Mol Biol 27:91–104
Peumans WJ, Proost P, Swennen RL, Van Damme EJM (2002) The abundant class III chitinase homolog in young developing banana fruits behaves as a transient vegetative storage protein and most probably serves as an important supply of amino acids for the synthesis of ripening-associated proteins. Plant Physiol 130:1063–1072
Potrykus I, Harms CT, Lorz H (1979) Callus formation from cell culture protoplasts of corn (Zea mays L.). Theor Appl Genet 54:209–214
Ramesh R, Kumar, KK, Kalpana K, Maruthasalam S, Sudhakar D, Balasubramanian P (2001) Engineering Xa21-mediated resistance in elite indica rice cultivars to Xanthomonas oryzae pv. oryzae. In: Proceedings of the 8th National Rice Biotechnology Network (NRBN) Meeting, Aurangabad, India, p. 89
Ronald PC (1997) The molecular basis of disease resistance in rice. Plant Mol Biol 35:179–186
Ronald PC, Alabani B, Tabien R, Abenes L, Wu KS, McCouch S, Tanksley SD (1992) Genetic and physical analysis of rice bacterial blight disease resistance locus, Xa21. Mol Gen Genet 236:113–120
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning. A laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York
Savary S, Willocquet L, Elazegui FA, Castilla NP, Teng PS (2000) Rice pest constraints in tropical asia: quantification of yield losses due to rice pests in a range of production situations. Plant Dis 84:357–369
Singh S, Sidhu JS, Huang N, Vikal Y, Li Z, Brar DS, Dhaliwal HS, Khush GS (2001) Pyramiding three bacterial blight resistance genes (xa5, xa13 and Xa21) using marker-assisted selection into indica rice cultivar PR106. Theor Appl Genet 102:1011–1015
Sivamani E, Shen P, Opalka N, Beachy RN, Fauquet C (1996) Selection of large quantities of embryogenic calli from indica rice seeds for production of fertile transgenic plants using the biolistic method. Plant Cell Rep 15:322–327
Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Hosten T, Gardner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald PC (1995) A receptor kinase-like protein encoded by the rice disease resistance gene Xa21. Science 270:1804–1806
Takakura Y, Ito T, Saito H, Inoue T, Komari T, Kuwata S (2000) Flower-predominant expression of a gene encoding a novel class I chitinase in rice (Oryza sativa L.). Plant Mol Biol 42:883–897
Tang K, Tinjuangjun P, Xu Y, Sun X, Gatehouse JA, Ronald PC, Qi H, Lu X, Christou P, Kohli A (1999) Particle bombardment-mediated co-transformation of elite chinese rice cultivars with genes conferring resistance to bacterial blight and sap sucking insects. Planta 208:552–563
Tu J, Datta K, Khush GS, Zhang Q, Datta SK (2000) Field performance of Xa21 transgenic indica rice (Oryza sativa L.), IR72. Theor Appl Genet 101:15–20
Tu J, Ona I, Zhang Q, Mew TW, Khush GS, Datta SK (1998) Transgenic rice variety IR72 with Xa21 is resistant to bacterial blight. Theor Appl Genet 97:31–36
Van Loon LC, Van Strien EA (1999) The families of pathogenesis-related proteins, their activities, and comparative analysis of PR-1 type proteins. Physiol Mol Plant Pathol 55:85–97
Velazhahan R, Chen-Cole K, Anuratha CS, Muthukrishnan S (1998) Induction of thaumatin-like proteins (TLPs) in Rhizoctonia solani infected rice and characterization of two new cDNA clones. Physiol Plant 102:21–28
Wang GL, Song WY, Ruan DL, Sideris S, Ronald PC (1996) The cloned gene Xa21 confers resistance to multiple Xanthomonas oryzae pv. oryzae isolates in transgenic plants. Mol Plant Microbe Interact 9:850–855
Yamamato T, Iketani H, Ieki H, Nishizawa Y, Notsuka K, Hayashi T, Matsuta N (2000) Transgenic grapevine plants expressing a rice chitinase with enhanced resistance to fungal pathogens. Plant Cell Rep 19:639–646
Yoshimura S, Yoshimura A, Iwata N, McCouch SR, Abenes MN, Baraoidan MR, Mew TW, Nelson RJ (1995) Tagging and combining bacterial blight resistance genes using RAPD and RFLP markers. Mol Breed 1:375–87
Yun DJ, Ibeas JI, Lee H, Coca MA, Narasimhan ML, Uesono Y, Hasegawa PM, Pardo JM, Bressan RA (1998) Osmotin, a plant antifungal protein, subverts signal transduction to enhance fungal cell susceptibility. Mol Cell 1:807–812
Zhang S, Song WY, Chen L, Ruan D, Taylor N, Ronald PC, Beachy RN, Fauquet CM (1998) Transgenic elite indica rice varieties, resistant to Xanthomonas oryzae pv.oryzae. Mol Breed 4:551–558
Zhu B, Chen THH, Li PH (1996) Analysis of late-blight disease resistance and freezing tolerance in transgenic potato plants expressing sense and antisense genes for an osmotin-like protein. Planta 198:70–77
Zhu Q, Maher EA, Masoud S, Dixon RA, Lamb CJ (1994) Enhanced protection against fungal attack by constitutive co-expression of chitinase and glucanase genes in transgenic tobacco. Biotechnology 12:807–812
Acknowledgements
The authors are grateful to The Rockefeller Foundation, USA [# 99001/225 (PB) and # 99001/227 (DS)] and the Department of Biotechnology, New Delhi, India, for providing the financial support. We greatly acknowledge the help provided by Dr. Legrand, Strasbourg, Cedex, France and Dr. S. Muthukrishnan, Kansas State University, USA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by L. Peña
S. Maruthasalam and K. Kalpana have contributed to this article equally.
Rights and permissions
About this article
Cite this article
Maruthasalam, S., Kalpana, K., Kumar, K.K. et al. Pyramiding transgenic resistance in elite indica rice cultivars against the sheath blight and bacterial blight. Plant Cell Rep 26, 791–804 (2007). https://doi.org/10.1007/s00299-006-0292-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-006-0292-5