Abstract
Waterlogging has increasingly become one of the major constraints to maize (Zea mays L.) production in some maize growing areas as it seriously decreases the yield. Waterlogging tolerance in maize germplasm provides a basis for maize waterlogging improvement. In this study, nine seedling traits, plant height (PH), root length (RL), shoot dry weight (SDW), root dry weight (RDW), adventitious root number (ARN), node number of brace root (BRNN), brace root number (BRN), brace root dry weigh (BRDW), survival rate (SR), and the secondary traits that were defined as relative phenotypic value of seedling traits under waterlogging and control treatments were used in a natural population that contain 365 inbred lines to evaluate the waterlogging tolerance of tropical maize. The result showed that maize waterlogging tolerance was genetically controlled and seedling traits were significantly different between the control and waterlogging treatments. PH, RL, SDW, and RDW are important seedling traits for waterlogging tolerance identification. Some tropical maize inbred lines were identified with extreme waterlogging tolerance that can provide an important germplasm resource for breeding. Population structure analysis showed that two major phylogenetic subgroups in tropical maize could be identified. Genome-wide association study (GWAS) using 39,266 single nucleotide polymorphisms (SNPs) across the whole genome identified 49 trait-SNPs distributed on over all 10 chromosomes excluding chromosome 10. Seventy-one significant SNPs, distributed on all 10 chromosomes excluding chromosome 5, were identified by extend bulked sample analysis (Ext-BSA) based on the inbred lines with extreme phenotypes. GWAS and Ext-BSA identified the same loci on bin1.07, bin6.01, bin2.09, bin6.04, bin7.02, and bin7.03. Nine genes were proposed as potential candidate genes. Cloning and functional validation of these genes would be helpful for understanding the molecular mechanism of waterlogging tolerance in maize.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Waterlogging is one of the most important abiotic stresses for maize (Zea mays L.) in tropical rainfed environments (Prasanna et al. 2021). As the continuous change of the global climate, waterlogging is becoming an important constraint for maize production and can cause numerous economic consequence (Olesen et al. 2011; Bailey-Serres et al. 2012). In South China, waterlogging is one of the most disasters in the maize early seedling growth phase from the second (V2) to the seventh leaf stage (V7) (Zaidi et al. 2004) and the former (V2) is the most susceptible stage (Zaidi et al. 2004; Liu et al. 2010). In this area, heavy rainfall in short periods and poor drainage can lead to waterlogging of soil which results in reduced levels of oxygen in tissues and gas diffusion between cells (Voesenek and Bailey-Serres 2013). The development of waterlogging-tolerant cultivars with high yield potential should be the most economic option to offset the damage caused by waterlogging (Zaidi et al. 2015; Tian et al. 2019). Some tropical or subtropical maize germplasms from International Maize and Wheat Improvement Center (CIMMYT) were tolerant for waterlogging after directive improvement (Zaidi et al. 2010, 2015). The strong photoperiod sensitivity of tropical maize germplasms makes it hard to grow directly in Northern China, particularly for the temperate maize areas. However, the waterlogging tolerance makes it a good resource to broaden the genetic base of China maize through the introgression of favorable genes and alleles.
Maize resistance to waterlogging is a complex trait that is controlled by multiple quantitative trait loci and phenotypic variation for waterlogging resistance explained by single QTL was small. It is hard to incorporate tolerance genes into hybrids (Yu et al. 2018). Therefore, how to select traits that can be easily identified and represent waterlogging tolerance effectively is challenging. For QTL analysis, a major QTL for root dry weight accounting for 36.3% of phenotypic variance was located on chromosome 9 (Qiu et al. 2007). Several QTL associated with root length and root dry weight were identified with moderate effects, each explaining 4.14-18.88% of the phenotypic variation (Osman et al. 2013). Five QTL were mapped on chromosomes 1, 3, 5, 7, and 10, together explaining approximately 30% of phenotypic variance for grain yield (Zaidi et al. 2015). In addition, three QTL controlling adventitious root formation were located on chromosomes 3 (bin3.07-8), 7 (bin 7.04-5), and 8 (bin8.05) (Mano et al. 2005). Two QTL for root aerenchyma formation were located on chromosomes 1 (bin 1.06-7) and 7 (bin7.01), explaining 17% and 12% of the total phenotypic variance (Mano et al. 2012). As linkage analysis exploits the shared inheritance of functional polymorphisms and adjacent markers, we need to create segregating populations (Yu and Buckler 2006). Recently, genome-wide association studies (GWAS) have played an important role in dissecting genetic architecture of complex traits in maize, as maize has numerous molecular markers, abundant genetic variation, and rapid linkage disequilibrium (LD) decay. The development of genotyping technologies has aided the improved resolution of GWAS with tremendous numbers of markers. Unlike linkage mapping, association mapping can be used to explore all the recombination events and mutations in a given population with high resolution (Yan and Warburton 2011). Compared to linkage mapping, however, association mapping has a lower power to detect rare alleles, even those with large effects (Yan and Warburton 2011). GWAS of waterlogging tolerance using 28,791 SNPs and 144 maize inbred lines identified four strong peak signals on chromosomes 5, 6, and 9 (Zhang et al. 2012). A total of 110 SNPs associated with waterlogging tolerance were identified using 368 inbred lines and 558,529 SNPs with single SNP explaining 2.88-10.67% of the phenotypic variance. A key candidate gene, GRMZM2G110141, was identified, which could be used in marker-assisted selection (Yu et al. 2018).
GWAS need to genotype all the individuals from a target population. On the contrary, bulked sample analysis (BSA) just needs to genotype the individuals with extreme phenotypes, significantly reducing the scale and cost of genotyping and simplifying the analysis (Zou et al. 2016). With the development of molecular technologies in recent years, BSA has witnessed many improvements. The pooled DNA analysis can be used for two contrasting groups of individuals with extreme phenotypes from either segregants or variants from any populations that represent wide ranges of phenotypic variation for the target trait as suggested by Sun et al. (2010) and Zou et al. (2016). When the number of individuals contained in each bulk is large enough, for example, more than 500, BSA can be combined with genotyping-by-sequencing (GBS) technologies and used for GWAS (Duncan et al. 2011; Schlötterer et al. 2014). Whole-genome sequencing of pools of individuals (Pool-seq) can facilitate a broad range of genome-wide analyses (Schlötterer et al. 2014). Pool-seq has been used in gene mapping using natural populations (Bastide et al. 2013).
In the present study, 365 tropical or subtropical maize inbred lines were genotyped individually. Since maize brace roots are generally distributed at the soil surface and play an important role in stabling yield under waterlogging (Hochholdinger and Tuberosa 2009; Zaidi et al. 2015). In addition to plant height, root length, shoot dry weight, and root dry weight, we also measured the node number of brace root, brace root number, brace root dry weight under the control and waterlogging treatments. Then, extreme germplasms were selected according to phenotypic data to form two mixed pools for resequencing. Ext-BSA, an extended bulked sample analysis method, combined with GWAS, was used to identify the genomic regions and candidate genes for waterlogging tolerance in tropical maize. Molecular markers associated with waterlogging tolerance provide a support for improving temperate maize germplasm for waterlogging tolerance.
Materials and Methods
Plant materials and waterlogging treatment
A total of 365 tropical /subtropical maize inbred lines were selected from International Maize and Wheat Improvement Center (CIMMYT). Majority of the inbred lines are white kernel types from two major sources, one coming from different gene pools or CIMMYT inbred lines entitled with “CML”, and the other from advanced breeding lines modified from “CML”. These inbred lines have two heterotic patterns, group A with tester CML373 and group B with tester CML418, and they have diverse genetic basis (CIMMYT. 1998). Detailed information about these 365 maize inbred lines is provided in Supplementary Table 1.
Nine seedling traits, including plant height (PH), root length (RL), shoot dry weight (SDW), and root dry weight (RDW), adventitious root number (ARN), node number of brace root (BRNN), brace root number (BRN), brace root dry weight (BRDW), and survival rate (SR), were measured as described previously (Du et al. 2016, 2017). In brief, six uniform seedlings of each line were planted in one silica pot (18 cm in diameter and 18 cm deep). Three and nine pots per line were included as control and waterlogging treatment, respectively. Three pots for waterlogging treatment as a group were treated as one independent biological replicate. Then, the pots for waterlogging treatments were put into three cement pools (each 3 m long, 0.8 m wide, and 0.5 m deep). At three-leaf stage, the cement pools were watered until the water reaches 2 to 3 cm above the pots, maintaining for 8 days. To phenotype, 18 seedlings per inbred lines were taken from each of waterlogging and control treatments with mean value calculated for each trait. PH, RL, ARN, BRNN, and BRN were measured manually. SDW, RDW, and BRDW were measured after dried in an oven (65 °C) for 3 days. SR was determined using 10 waterlogged sensitive and eight tolerant inbred lines. Nine secondary traits that were defined as relative phenotypic value of seedling traits, RPH (e.g., RPH = PHtreatment/PHcontrol), RRL, RSDW, RRDW, RARN, RBRNN, RBRN, RBRDW were calculated based on the phenotypic value of seedling traits under waterlogging treatment and that under control treatment. SR is a seedling trait and is also a secondary trait.
Data analysis and evaluation of waterlogging tolerance
Descriptive statistics were calculated with Excel 2016, and statistical analysis was conducted with Statistical Analysis System (SAS) software version 9.4, including one-way analysis of variance (ANOVA) and Pearson correlation analysis for seedling traits under the waterlogging and control treatments, and principal component analysis (PCA) for the value of nine secondary traits. We retained principal components with eigenvalues more than 1. The sum of scores of the retained principal component was taken as comprehensive waterlogging tolerance index by which 365 inbred lines were ranked. The Student's test was used to test the significance difference of relative phenotypic values between 30 highly tolerant inbred lines and 30 highly susceptible inbred lines. Significant differences of relative phenotypic value are indicated with asterisks in the figures.
Genotyping, population structure, and estimation of Kinship Matrix
For DNA extraction, 15 seeds were grown in one pot and five bulked leaf samples were collected to represent each inbred line using high throughput CTAB method (CAAS-CIMMYT Maize Molecular Breeding Laboratory 2015). For each inbred line, 200 ng genomic DNA was used for genotyping on the Affymetrix@ GeneTitan@ platform with the Axiom@ my Design GW genotyping array according to manufacturer's protocol (Xu et al. 2017). In total, 39,266 informative SNPs with a minor allele frequency (MAF) > 0.05 and missing data < 10% were used in GWAS. Heterozygous loci were considered missing. MAF was estimated using PowerMarker V3.25.
In order to decrease running time, a total of 3000 filtered SNPs with high genetic diversity, low missing rate, and evenly distributed on 10 maize chromosomes were selected to estimate population structure for the 365 inbred lines using Structure V2.3.3 (Hubisz et al. 2009). The K (number of subpopulations) was set from 2 to 10 with five runs for each K and Length of Burnin period and Number of MCMC Reps after Burnin were set to 10,000 and 20,000, respectively. The optimum K was estimated using the log probability lnP (D) from the STRUCTRE output and the ad hoc statistics ΔK (Evanno et al. 2005). All 39,266 SNPs were used to calculate the relative kinship coefficient matrix using TASSEL5.0 (Bradbury et al. 2007). The closer the relative kinship coefficient is to 0, the farther the relationship is between the two inbred lines, and vice visa.
Genome-wide association study (GWAS)
GWAS was performed using values of nine secondary traits by accounting for population structure (Q matrix) and kinship matrix (K) between each pair of inbred lines in the association panel using TASSEL 5.0 and a Bonferroni-corrected threshold set at p = 1/n was used to verify the significance levels, where n is the number of SNPs used. Using “no compression” and “population parameters previously determined” (P3D) algorithms, mixed linear model (MLM) was used to detect the association between phenotype and genotype. The MLM can be expressed as follows:
where Y is the observed value vector, β is the fixed effect vector which includes genetic markers and population structure factors, u is the random effect vector, X and Z are the known design matrices, and ε is the random residual effect vector. Manhattan plots were created in R software using the GWAS results.
LD between pairwise SNPs for the association panel was estimated with TASSEL 5.0 and the average distance of LD decay across the entire genome was estimated using all LD data and SAS Nlin PROC procedure. The candidate genes containing or adjacent to significant SNPs were identified using the maize B73 reference genome assembly v3 available on the MaizeGDB genome database (http://gbrowse.maizegdb.org/gb3/gbrowse/maize_v3/). The closest gene to each peak SNP was proposed to be the most likely candidate gene. All the candidate genes were annotated according to the information available in the MaizeGDB database (http://www.maizegdb.org/gbrowse).
Bulked sample analyses (BSA)
According to the result of PCA scores, 30 highly waterlogging tolerant inbred lines and 30 highly susceptible inbred lines were selected. An equal amount of DNA from each individual is bulked to form tolerant and susceptible pools. The two DNA pools were re-sequenced at Berry Genomics. Four libraries of every pool were constructed for Illumina HiSeqTM 2000 and paired-end sequencing with 150 bp were performed. Base calling and filtering of low-quality bases were done using Fastx-toolkit (v 0.0.14-1) (Li and Durbin 2009). The Genome Analysis Toolkit (GATK) software was employed to identify SNPs (McKenna et al. 2018). Two methods were used to correlation analysis. The first one is SNP-index analysis by which we assessed the loci through ΔSNP-index (Abe et al. 2012; Takagi et al. 2013). The second is Ext-BSA (Extend bulked sample analysis). Ext-BSA contains three parts as follows:
-
(1)
Standardization of S-score. The standardized S-score is the difference between the largest −log(P) upstream and downstream of the marker and the average −log(P) of the whole genome.
$$ \mathrm{S}-\mathrm{score}={\displaystyle \begin{array}{c}s\\ {}\mathit{\max}\\ {}i=1\end{array}}\left(-\log\ {P}_i\right)-\frac{1}{m}\sum \limits_{i=1}^m\left(-\log\ {P}_i\right), $$
where s is the number of markers upstream and downstream of the marker of research, and m is the total number of markers in the whole genome. This method cannot only eliminate false positive markers but also simulate the LD between markers to improve the accuracy of analysis.
-
(2)
Student's test of allele frequencies between susceptible and tolerant pools by using the following formula:
$$ {\displaystyle \begin{array}{l}t=\frac{\mid {f}_1-{f}_2\mid }{\sqrt{\left(\frac{1}{2{n}_1}+\frac{1}{2{n}_2}\right){f}_{exp}\left(1-{f}_{exp}\right)}},\\ {}{f}_{exp}=\frac{f_1{n}_1+{f}_2{n}_2}{n_1+{n}_2},\end{array}} $$
where f1 and f2 were allele frequencies while n1 and n2 were the sample sizes for susceptible and tolerant pools, respectively.
-
(3)
Bonferroni correction of Student's test. Bonferroni method (0.05/N, N is the number of effective markers) was used to correct the test results, and the trait-association markers or intervals were observed.
For the trait association markers, Bayesian was approached to estimate the linkage probability between each significant SNP and the relevant causal gene. SNP index plot regression lines were obtained by averaging SNP indices from a moving of five consecutive SNPs and shifting the window one SNP at a time. The candidate genes containing or adjacent to significant SNPs were identified using the maize B73 reference genome assembly v3 available on the MaizeGDB genome database.
Results
Phenotypic variation in waterlogging stress identified in tropical maize
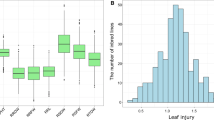
Statistical analysis (Table 1) demonstrated that for nine seedling traits, mean values significantly (P < 0.01) differed among genotypes, indicating rich genetic diversity in waterlogging tolerance. Under waterlogging condition, the mean phenotypic values for PH, RL, SDW, RDW, and ARN decreased significantly (P < 0.01) but increased significantly (P< 0.01) for brace root related traits (BRNN, BRN, BRDW), suggesting that the treatment effects were significant (P < 0.01). The two most phenotypic variations under waterlogging treatment were SR (58.88% decreased) and BRN (74.12% increased). Seedling traits fit a normal distribution and exhibited positive skewness except RL under waterlogging treatment (Table 1; Supplementary Fig. 1). The secondary traits also showed abundant variation, ranging from 0.413 to 1.119 for RPH, 0.116 to 0.982 for RRL, 0.296 to 1.591 for RSDW, 0.229 to 1.739 for RRDW, 0.248 to 1.459 for RARN, 0.487 to 2.038 for RBRNN, 0.313 to 4.960 for RBRN, 0.125 to 7.500 for RBRDW, 0 to 1.000 for SR respectively (Figs. 1, 2).
Distributions of eight secondary traits except SR in the 365 tropical/subtropical maize panel. RPH, relative phenotypic value for plant height (PH); RRL for root length (RL); RSDW for shoot dry weight (SDW); RRDW for root dry weight (RDW); RARN for adventitious root number (ARN); RBRNN for node number of brace root (BRNN); RBRN for brace root number (BRN); RBRDW for brace root dry weight (BRDW); SR: survival rate
All seedling traits except RL were significantly (P < 0.05) positively correlated between waterlogging and control treatments, indicating that there is some common genetic basis under both treatments (Table 2). Under the control condition, SDW was significantly (P < 0.05) positively correlated with other traits except SR, and RDW was significantly (P < 0.01) positively correlated with other traits except SR and BRNN, suggesting that SDW and RDW are two important seedling traits for waterlogging tolerance identification. ARN was significantly (P < 0.05) negatively correlated with BRNN, BRN, and BRDW, suggesting that adventitious roots inhibited the growth of brace roots under the control treatment. ARN was significantly (P < 0.01) positively correlated with BRN under the treatment condition, suggesting waterlogging treatment promoted the growth of adventitious roots and brace roots simultaneously. Under the waterlogging condition, PH and RL was significantly (P < 0.01) positively correlated with other traits except SR, suggesting PH and RL are also important seedling traits. Under both conditions, SR was only significantly (P < 0.01) correlated with BRN, suggesting SR should be an independent trait for waterlogging tolerance identification.
Tropical maize inbreds identified with strong tolerance to waterlogging
PCA for the value of nine secondary traits of 365 tropical inbred lines indicates three principal components should be retained when eigenvalue more than 1 (Fig. 3a,b) and the cumulative proportion of three principal components is 62.80%. Correlation matrix of nine secondary traits showed that there is significantly (P <0.01) positive correlation between RPH and RSDW (Supplementary Table 2). We calculated the sum of the first three PCA and had a rank to the 365 inbred lines (Supplementary Table 3). The results were in line with the former research. For example, CML32 was tolerant to waterlogging (CIMMYT. 1998) while CML311 was susceptible to waterlogging (Zaidi et al. 2010). Except for CML32, other CIMMYT germplasms such as CML130, CML171, CML292, CML325, CML328, CML 343, CML373, CML428, and CML479 were tolerant to waterlogging but CML80, CML127, CML226, CML254, CML373, CML444, and CML511 were susceptible to waterlogging. The extreme waterlogging-tolerant tropical maize germplasms provide an important resource for developing waterlogging-tolerant maize hybrids.
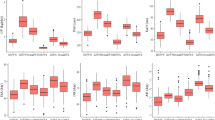
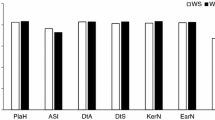
Scree test and value of nine secondary traits for 30 highly tolerant inbred lines and 30 highly susceptible inbred lines. (a) Eigenvalue by numbers of principal components. (b) Proportion and cumulative by numbers of principal. (c) Relative phenotypic value and Student's test of nine secondary traits of 30 highly tolerant inbred lines (T) and 30 highly susceptible inbred lines (S). *, ** indicate significance at 0.05 and 0.01 levels, respectively
Thirty inbred lines with the highest scores and 30 inbred lines with the lowest scores were selected as the extremely tolerant and susceptible materials respectively (Supplementary Table 3). There are significant (P < 0.05 or P < 0.05) difference between the extremely tolerant and susceptible materials in RPH, RRL, RARN, RRDW, RSDW, RBRDW, and SR (Fig. 3c). As an important comprehensive trait, the mean of SR for extremely tolerant materials is 73.5% while is 8.88% for extremely susceptible materials.
Two major phylogenetic subgroups in tropical maize and the suitability of the germplasm panel for GWAS
Population structure analysis for 365 tropical/subtropical inbred lines was implemented for K ranging from 2 to 10. The results indicate that the model overestimated the number of groups, and thus, no suitable K could be chosen from the Ln probability and used directly for group determination (Supplementary Fig. 2). However, when K = 3 to K=9, two major subgroups could be identified, population 1 (pop1) and population 2 (pop2) (Fig. 2a and Supplementary Table 1). The result was line with the previous studies that tropical germplasms can be classified into heterotic groups A and B based on their general combining ability (GCA) (Vasal et al. 1992a, b; CIMMYT. 1998; Mickelson et al. 2001). The frequency distribution showed that 90% of the kinship relative coefficients were between 0 and 0.45 (Fig. 2b). A total of 62.9% of the pairwise kinship estimates were equal to 0, indicating that there was no relatedness between these pairs of lines, and the remaining estimates decreased continuously with the increase of category number. The kinship analysis indicated only few lines with strong similarities. Thus, the maize panel was suitable for GWAS.
Important genetic loci for waterlogging tolerance identified by GWAS
By controlling population structure (Q matrix) and kinship matrix (K), MLM model was fitted to scan 39,266 SNPs across the whole genome. Quantile quantile-quantile (QQ plots) indicated that Q matrix and K were well controlled in GWAS for each trait (Fig. 4 and Supplementary Figs. 3 and 4). With a cutoff of P < 2.52 × 10-5 [−log (1/39,266)], 49 trait-SNP combinations were identified as being associated with waterlogging tolerance (Supplementary Table 4). These significant associated SNPs were distributed on all 10 chromosomes except chromosome 10 with at least one SNP identified for each secondary trait. The genotypic variance explained (PVE) by a single SNP (R2) ranged from 5.3% to 10.1%, and mean of PVE for the 49 SNPs is 7.3%. A total of 17 SNPs were significantly associated with the above-ground traits (RPH, RSDW) and 29 SNPs were significantly associated with root-related traits (RRL, RRDW, RARN, RBRNN, RBRN, RBRDW). The most significant SNPs were associated with RBRNN, which had the highest PVE of 10.1% (Supplementary Fig. 4). Three significant SNPs (3_137228397 on chromosome 3, 4_142418820 on chromosome 4 and 8_101658071 on chromosome 8) were identified by two different secondary traits (RPH and RSDW). Another significant SNP (4_196842052 on chromosome 4) was associated with both RPH and RRL. With a more stringent threshold of 1.27 × 10−6[−log (0.05/39,266)], only one significant SNP (for BRNNR) could be identified on chromosome 4.
Manhattan plot of GWAS and quantile-quantile plots for three seedling traits in the maize association panel. Manhattan plots for RPH, RRDW and SR are shown in a, c and e, respectively, and the quantile-quantile plots for RPH, RRDW, and SR are shown in (b), (d), and (f), respectively. The red lines show genome-wide significance at stringent thresholds of 2.52 × 10−5[−log (1/39,266)]
Significant SNPs identified for waterlogging tolerance by Ext-BAS
A total of 415 million Clean Reads and 125 billion Clean Bases were obtained in the susceptible pool, of which Q30 was 93.30% and 83.79%, respectively, compared with 482 million Clean Reads and 145 billion Clean Bases obtained in the tolerant pool, of which Q30 was 93.58% and 83.50%, respectively. For the susceptible and tolerant pools, 98.75% and 98.56% of the Clean Reads were mapped to the reference genome and the repetition rate was 12.98% and 13.25% respectively. Using GAKT software, 12,384,120 SNPs were generated. These SNPs were filtered according to three standards: SNP-index in both pools was less than 0.3; SNP-index in one of the pools was missing; clean Q > 20, with 2,842,853 SNPs obtained for BSA. For each SNP, SNP index was calculated and plotted on maize genome (Fig. 5a). As expected, the SNP indexes were randomly distributed across the genome around 0.5 for the two extreme pools. Ext-BSA method was used to extract SNP markers (Fig. 5b) with 345,606 effective SNP markers obtained. At the threshold of 6.849 [–log10(0.05/345606)], 71 significant SNPs were detected across all chromosomes except chromosome 5 (Supplementary Table 5). At the threshold of 7.538 [–log10(0.01/345606)], 47 significant SNPs were detected on chromosomes 2, 3, 4, 6, 7, 9, and 10. At a more stringent significant threshold of 8.539 [–log10(0.001/345606)], 21 SNPs were detected with strong signals on bin2.08, bin6.01, bin6.03, bin7.02, and bin10.04.
Nine potential candidate genes identified by GWAS and Ext-BSA
The average distance of LD decay across the entire genome was 212 kb when r2 = 0.1 and this is line with our previous report (Xu et al. 2017). In GWAS, the genes harboring or adjacent to the significant SNPs within the 212 kb region were considered as candidate genes. In Ext-BSA, significant SNPs within 200 kb windows were used for identification of candidate genes. Combined GWAS and Ext-BSA, nine potential candidate genes were identified, locating in six chromosomal regions (bin1.07, bin2.08, bin6.01, bin6.04, bin7.02, and bin7.03) (Table 3). Five candidate genes (GRMZM2G425366, GRMZM2G068672, GRMZM2G085378, GRMZM2G019443, and GRMZM5G851405) identified by Ext-BSA were adjacent to those identified by GWAS. Four candidate genes (GRMZM2G171934, GRMZM2G012717, GRMZM2G106141, and GRMZM2G111309) were identified by both GWAS and Ext-BSA within the same windows (Table 3; Supplementary Tables 4 and 5). Therefore, these four candidate genes could be used to develop markers for marker-assisted selection in maize waterlogging tolerance breeding.
Discussion
Although maize is susceptible to waterlogging at germination and early seedling stage, there is a significant genetic difference among genotypes (Mano et al. 2002; Zaidi et al. 2004; Liu et al. 2010; Zaidi et al. 2015; Yu et al. 2018). CIMMYT has a large number of tropical maize germplasm, most of which are derived from broad germplasm pools, populations and open-pollinated varieties (Xia et al. 2004; Xia et al. 2005). These germplasms have been evaluated for various biological or abiotic stresses including waterlogging and widely used in maize improvement all over the world (Menkir et al. 2004; Goodman 2005; Reif et al. 2010; Prasanna et al. 2021). A large number of tropical maize inbreds from CIMMYT have been intensively evaluated in China (Wu et al. 2007; Chen et al. 2011; Fan et al. 2018) for many traits. In this study, 365 tropical or subtropical maize inbreds were selected from different gene pools and advanced CIMMYT breeding lines with good representativeness, and it is a large-scale evaluation of waterlogging tolerance. Because metabolic mechanism and biochemical basis have not been defined yet it is very difficult to develop a unique index for waterlogging tolerance identification (Liu et al. 2010). Under the waterlogging stress, maize morphological, physiological, biochemical, and transcriptional changes take place. The relative phenotypic value for root dry weight, root fresh weight, and root length decreased significantly (Osman et al. 2013; Yu et al. 2018). Superoxide anion content (\( {O}_2^{.-} \)), hydrogen peroxide content (H2O2), and malondialdehyde (MDA) content increased, causing an up-regulation of superoxide dismutase (SOD), peroxidase (POD), and catalase (CAT) (Tian et al. 2019). Yield is significant positively correlated with brace roots and chlorophyll content (SPAD) and negatively correlated with %stem and root lodging (Zaidi et al. 2015). Compared with above-ground traits like plant height and shoot dry weight, roots were more prone to be affected under waterlogging stress, resulting in reduced root length and root dry weight (Qiu et al. 2007; Zhang et al. 2012; Yu et al. 2018). In this study, nine seedlings traits were evaluated, with PH and SDW as the above-ground traits while RL, RDW, ARN, BRN, BRNN, and BRDW as root characteristics. SR can represent overall maize waterlogging tolerance. SDW, RDW, PH, and RL were significantly positively correlated with other traits, suggesting that they are important seedling traits contributing to waterlogging tolerance in maize. PCA from nine secondary traits was used to evaluate waterlogging tolerance. CIMMYT tropical germplasms CML32, CML130, CML171, CML292, CML325, CML328, CML 343, CML373, CML428, and CML479 were identified as waterlogging tolerant donors and can used in our future breeding. The tropical and subtropical germplasms, which have been adapted to middle area, can be introduced to higher-latitude areas, through marker-assisted gene introgression, and used for breeding temperate maize (Yang et al. 2021). Besides that, we suggested that except PH, RL, RDW, and SDW, SR should be an independent index for waterlogging tolerance identification.
It is estimated that the minimum number of markers needed for a successful GWAS in maize is more than 10 million dues to a much larger genome and a much more rapid LD decay (Myles et al. 2009). Compared with temperate lines, tropical lines are more diverse and contain more rare alleles and the LD decay distance was much shorter (Liu et al. 2003; Yan et al. 2009). Therefore, GWAS in tropical maize lines require more markers to get close to the gene of interest (Yan and Warburton 2011). Nowadays, array-based system can only achieve to one million SNPs (Gupta et al. 2008; Yan et al. 2010). Although the cost of sequencing-based platform decreases recent years, large-scale genotyping is still a bottleneck. Another disadvantage of GWAS is that it is difficult to identify rare alleles in populations, even though they have greater effects. Next generation sequencing (NGS) of pools of individuals is often more effective in gene mapping and provides more accurate allele frequency estimates, even when taking sequencing errors into account (Futschik and Schlötterer 2010). It can be inferred that the plants with extreme phenotypes selected for BSA would be those with accumulated favorable alleles from multiple loci. Therefore, both GWAS-based and BSA-based approaches, which are complementary with each other, should be used for gene mapping and candidate gene discovery. In the present study, four loci (bin1.07, bin6.01, bin6.04, and bin7.03) were colocated by both GWAS and Ext-BSA (Supplementary Tables 4 and 5). Of these, one significant SNP (1_204203096 on chromosome 1) identified by GWAS is very close to the two significant SNPs (1_204240360 and 1_204240418 on chromosome 1) identified by Ext-BSA. Two other SNPs (6_117803575 and 6_117609082 on chromosome 6) identified respectively by GWAS and Ext-BSA are also located very close together.
Up to now, hundreds of QTL have been identified for waterlogging tolerance across the whole maize genome, including those for root aerenchyma formation (Mano and Omori 2008; Mano et al. 2007, 2008, 2012), adventitious root formation (Mano et al. 2005), root length, plant height, and biomass-related traits (Mano et al. 2006; Osman et al. 2013; Qiu et al. 2007; Zhang et al. 2012). Majority of the QTL identified by GWAS were also reported previously. For example, previous studies reported the loci on bin 1.07 (Mano et al. 2007; Mano et al. 2012) and bin 9.07 (Mano et al. 2008) were associated with controlling constitutive root aerenchyma formation. In the present study, QTL in bin 1.07 and bin 9.07 were identified for RBRNN. Both the formation of constitutive root aerenchyma and the increase of brace roots under waterlogging condition promote access to and diffusion of O2 in maize, suggesting these QTL are associated with a low O2 escape strategy (Bailey-Serres and Voesenek 2008). The loci on bin 9.07 was also reported to be associated with controlling root dry weight and root fresh weight (Zhang et al. 2012). The loci on both bin 4.05 and bin 8.04 were identified in two previous reports for controlling shoot dry weight (Qiu et al. 2007; Osman et al. 2013). In the present study, QTL in bin 4.05 and bin 8.04 were also identified for RPH and RSDW. PH was significantly (P < 0.01) positively correlated with SDW under control and waterlogging conditions, suggesting these QTL may control the elongation rate of above-ground part. The loci on bin 1.03, bin 1.07, bin 6.04, and bin 10.04 identified by Ext-BSA have been reported previously for controlling root aerenchyma formation and root fresh weight (Mano et al. 2007; Mano et al. 2012; Zhang et al. 2012).
Previous studies suggest that the genetic architecture of these waterlogging related traits is controlled by a few major QTL plus a large number of minor-effect ones. In the present study, both GWAS and Ext-BSA were used for waterlogging traits with nine candidate genes identified. These genes contain transcription factors, hypothetical protein, histone acetyltransferase1, and amidase isoform 1. The gene GRMZM2G171934 in bin1.07 encodes PLATZ-transcription factor 4 (platz4). Plant AT-rich sequence and zinc-binding proteins (PLATZ) are zinc dependent DNA binding proteins. They bind to AT rich sequences and function in transcriptional repression. The gene GRMZM2G012717 in bin6.04 encodes C2C2-CO-like-transcription factor-10 (col10). Other annotations include intracellular and zinc ion binding. Gene GRMZM2G148056 in bin7.02 encodes two-component response regulator ARR3. Other annotations include phosphorelay response regulator activity, phosphorelay signal transduction system, and regulation of transcription. Further verification of these candidate genes can improve the development of maize molecular breeding for waterlogging tolerance. Considering the polygenic nature of the waterlogging tolerance, whole-genome selection could be the most appropriate breeding method in the future (Das et al. 2020).
Conclusion
Maize waterlogging tolerance was genetically controlled and the waterlogging treatment effects was significant (P <0.01). PH, RL, SDW, and RDW are important seedling traits for waterlogging tolerance identification. CML32, CML130, CML171, CML292, CML325, CML328, CML343, CML373, CML 428, and CML479 were tolerant to waterlogging. Two major phylogenetic subgroups could be identified in tropical maize based on 3000 filtered SNPs. Forty-nine trait-SNP combinations were identified as being associated with waterlogging tolerance by GWAS and 71 significant SNPs were identified by Ext-BSA. Nine candidate genes were identified and four of them were identified by both GWAS and Ext-BSA within the same windows.
References
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30:174–178
Bailey-Serres J, Voesenek LACJ (2008) Flooding stress: acclimations and genetic diversity. Annu Rev Plant Bio 59:313–339
Bailey-Serres J, Lee SC, Brinton E (2012) Waterproofing crops: effective flooding survival strategies. Plant Physiol 160:1698–1709
Bastide H, Betancourt A, Nolte V, Tobler R, Stöbe P, Futschik A, Schlötterer C (2013) A genome-wide, fine-scale map of natural pigmentation variation in Drosophlia melanogaster. PloS Genet 9:e1003534
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635
Chen HM, Wang YF, Yao WH, Luo LM, Li LJ, Xu CX, Fan XM, Guo HC (2011) Utilization potential of the temperate maize inbreds integrated with tropical germplasm. Acta Agronomica Sinica 37:1785–1793 (In Chinese)
CIMMYT (1998) A complete listing of improved maize germplasm from CIMMYT. Maize Program Special Report. Mexico, D.F.
Das RR, Vinayan MT, Patel MB, Phagna RK, Singh SB, Shahi JP, Sarma A, Barua NS, Babu R, Seetharam K, Burgueño JA, Zaidi PH (2020) Genetic gains with rapid-cycle genomic selection for combined drought and waterlogging tolerance in tropical maize (Zea mays L.). Plant Genome 13:e20035
Du H, Shen X, Huang Y, Huang M, Zhang Z (2016) Overexpression of Vitreoscilla hemoglobin increase waterlogging tolerance in Arabidopsis and maize. BMC Plant Biol 16:35
Du H, Zhu J, Su H, Huang M, Wang H, Ding S, Zhang B, Luo A, Wei S, Tian X, Xu Y (2017) Bulked segregant RNA-seq reveals differential expression and SNPs of candidate genes associated with waterlogging tolerance in maize. Front in Plant Sci 8:1022
Duncan EL, Danoy P, Kemp JP, Leo PJ (2011) Genome-wide association study using extreme truncate selection identifies novel genes affecting bone mineral density and fracture risk. PloS Genet 7:e10011372
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fan XM, Zhang YD, Jeffers DP, Bi YQ, Kang MS, Yin XF (2018) Combining ability of yellow lines derived from CIMMYT populations for use in subtropical and tropical midaltitude maize production environments. Crop Sci 58:169–179
Futschik A, Schlötterer C (2010) The next generation of molecular markers from massively parallel sequencing of pooled DNA samples. Genetics 186:207–218
Goodman MM (2005) Broadening the U.S. maize germplasm base. Maydica 50:230–214
Gupta PK, Rustgi S, Mir RR (2008) Array-based high-throughput DNA markers for crop improvement. Heredity 101:5–18
Hochholdinger F, Tuberosa R (2009) Genetic and genomic dissection of maize root development and architecture. Curr Opin Plant Biology 12:172–177
Hubisz MJ, Falush D, Stephens M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332
Li H, Durbin R (2009) Fast and accurate short read and alignment with burrows-wheeler transform. Bioinformatics 25:1754–1760
Liu K, Goodman MM, Muse S, Smith JSC, Buckler ES, Doebley J (2003) Genetic structure and diversity among maize inbred lines as inferred from DNA microsatellites. Genetics 165:2117–2128
Liu Y, Tang B, Zheng Y, Ma K, Xu S, Qiu F (2010) Screening methods for waterlogging tolerance at maize (Zea mays L.) seeding Stage. Agricultural Sciences in China 9(3):365–369
Mano Y, Omori F (2008) Verification of QTL controlling root aerenchyma formation in a maize×teosinte “Zea nicaraguensis” advanced backcross population. Breeding Sci 58:217–223
Mano Y, Muraki M, Komatsu T, Fujimori M, Akiyama F, Takamizo T (2002) Varietal difference in pre-germination flooding tolerance and waterlogging tolerance at the seedling stage in maize inbred lines. Japanese Journal of Crop Science 73(3):361–367
Mano Y, Omori F, Muraki M, Takamizo T (2005) QTL mapping of adventitious root formation under flooding conditions in tropical maize (Zea mays L.) seedlings. Breeding Sci 55:343–347
Mano Y, Masanori M, Tadashi (2006) Identification of QTL controlling flooding tolerance in reducing soil conditions in maize (Zea mays L.) seedlings. Plant Prod Sci 9:176–181
Mano Y, Omori F, Takamizo T, Kindiger B, Bird RM, Loaisiga CH, Takahashi H (2007) QTL mapping of root aerenchyma formation in seedlings of a maize×rare teosinte “Zea nicaraguensis” cross. Plant Soil 295:103–113
Mano Y, Omori F, Kindiger B, Takahashi H (2008) A linkage map of maize × teosinte Zea luxurians and identification of QTLs controlling root aerenchyma formation. Mol Breed 21:327–337
Mano Y, Omori F, Takeda K (2012) Construction of intraspecific linkage maps, detection of a chromosome inversion, and mapping of QTL for constitutive root aerenchyma formation in the teosinte Zea nicaraguensis. Mol Breed 29:137–146
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo A (2018) The genome analysis toolkit: a MapReduce framework for analyzing nest-generation DNA sequencing data. Genome Res 20:1297–1303
Menkir A, Melake-Berhan A, The C, Ingelbrecht I, Adepoju A (2004) Grouping of tropical mid-altitude maize inbred lines on the basis of yield data and molecular markers. Theor Appl Genet 108:1582–1590
Mickelson HR, Cordova H, Pixley KV, Bjarnason MS (2001) Heterotic relationships among nine temperate and subtropical maize populations. Crop sci 41:1010–1020
Myles S, Peiffer J, Brown PJ, Ersoz ES, Zhang Z, Costich DE, Buckler ES (2009) Association mapping: critical considerations shift from genotyping to experimental design. Plant Cell 21:2194–2202
Olesen JE, Trnka M, Kersebaum KC, Skjelvåg AO, Seguin B, Peltonen-Sainio P, Rossi F, Kozyra J, Micale F (2011) Impacts and adaptation of European crop production systems to climate change. Eur J Agron 34:96–112
Osman KA, Tang B, Wang Y, Chen J, Yu F, Li L, Han X, Zhang Z, Yan J, Zheng Y, Yue B, Qiu F (2013) Dynamic QTL analysis and candidate gene mapping for waterlogging tolerance at maize seedling stage. PLoS ONE 8:e79305
Prasanna BM, Cairns J, Zaidi PH, Beyene Y, Makumbi D, Gowda M, Magorokosho C, Zaman-Allah M, Olsen M, Das A, Worku M, Gethi J, Vivek BS, Nair SK, Rashid Z, Vinayan MT, Issa AB, Vicente FS, Dhliwayo T, Zhang X (2021) Beat the stress: breeding for climate resilience in maize for the tropical rainfed environments. Theor Appl Genet. https://doi.org/10.1007/s00122-021-03773-7
Qiu F, Zheng Y, Zhang Z, Xu S (2007) Mapping of QTL associated with waterlogging tolerance during the seedling stage in maize. Ann Bot-London 99:1067–1081
Reif JC, Fisher S, Schrag TA, Lamkey KR, Klein D, Dhillon BS, Utz HF, Melchinger AE (2010) Broadening the genetic base of European maize heterotic pools with US Cornbelt germplasm using field and molecular marker data. Theor Appl Genet 120:301–310
Schlötterer C, Tobler R, Kofler R, Nolte V (2014) Sequencing pools of individuals-mining genome-wide polymorphism data without big funding. Nat Rev Genet 15:749–763
Sun Y, Wang J, Crouch JH, Xu Y (2010) Efficiency of selective genotyping for genetic analysis of complex traits and potential applications in crop improvement. Mol Breed 26:493–511
Takagi H, Abe A, Yoshida K, Kosugi S, Natsume S, Mitsuoka C, Uemura A, Utsushi H, Tamiru M, Takuno S, Innan H, Cano LM, Kamoun S, Terauchi R (2013) QTL-seq: rapid mapping of quantitative trait loci in rice by whole genome resequencing of DNA from two bulked populations. Plant J 74:174–183
Tian L, Bi W, Liu X, Sun L, Li J (2019) Effects of waterlogging stress on the physiological response and grain-filling characteristics of spring maize (Zea mays L.) under field conditions. Acta Physiologiae Plantarum 41:63
Vasal SK, Srinivasan G, Gonzalez F, Han GC, Pandey S, Beck DL, Crossa J (1992a) Heterosis and combining ability of CIMMYT’s tropical * subtropical maize germplasm. Crop sci 32:1483–1489
Vasal SK, Srinivasan G, Crossa J, Bech DL (1992b) Heterosis and combining ability of CIMMT's subtropical and temperate early-maturity maize germplasm. Crop Sci 32:884–890
Voesenek LA, Bailey-Serres J (2013) Flooding tolerance: O2 sensing and survival strategies. Curr Opin Plant Biol 16:647–653
Wu JC, Xu CX, Chen HM, Tan J, Han XR, Huang BH, Fan XM (2007) Studies on combining ability and heterotic grouping of 24 quality protein maize inbreds and four temperate representative inbreds of Chinese major heterotic groups. Acta Agronomica Sinica 40:1288–1296 (In Chinese)
Xia XC, Reif JC, Hoisington DA, Melchinger AE, Frisch M, Warburton ML (2004) Genetic diversity among CIMMTY maize inbred lines investigated with SSR marker: I. lowland tropical maize. Crop Sci 44:2230–2237
Xia XC, Reif AE, Melchinger AE, Frisch M, Hoisington DA, Beck D, Pixley K, Warburton ML (2005) Genetic diversity among CIMMYT maize inbred lines investigated with SSR marker: II. Subtropical, tropical midaltitude, and highland maize inbred lines and their relationships with elite U.S. and European maize. Crop Sci 45:2573–2583
Xu C, Ren Y, Jian Y, Guo Z, Zhang Y, Xie C, Fu J, Wang H, Wang G, Xu Y, Li P, Zou C (2017) Development of a maize 55K SNP array with improved genome coverage for molecular breeding. Mol Breed 37:20
Yan J, Warburton ML (2011) Association mapping for enhancing maize (Zea mays L.) genetic improvement. Crop Sci 51:433–449
Yan J, Shah T, Warburton ML, Buckler ES, McMullen MD, Crouch J (2009) Genetic characterization and linkage disequilibrium estimation of a global maize collection using SNP markers. PLoS ONE 4:e8451
Yan J, Yang X, Shah T, Sánchez-Villeda H, Li J, Warburton M, Zhou Y, Crouch JH, Xu Y (2010) High-throughput SNP genotyping with the GoldenGate assay in maize. Mol Breed 25:441–451
Yang Y, Sang Z, Du Q, Guo Z, Li Z, Kong X, Xu Y, Zou C (2021) Flowering time regulation model revisited by pooled sequencing of mass selection populations. Plant Sci 304:110797
Yu J, Buckler ES (2006) Genetic association mapping and genome organization of maize. Curr Opin Biotech 17:155–160
Yu F, Liang K, Zhang Z, Du D, Zhang X, Zhao H, UI haq B, Qiu F (2018) Dissecting the genetic architecture of waterlogging stress-related traits uncovers a key waterlogging tolerance gene in maize. Theor Appl Genet 131:2299–2310
Zaidi PH, Rafiquea S, Rai PK, Singha NN, Srinivasanb G (2004) Tolerance to excess moisture in maize (Zea mays L.): susceptible crop stages and identification of tolerant genotypes. Field Crop Res 90:189–202
Zaidi PH, Maniselvan P, Srivastava A, Yadav P, Singh RP (2010) Genetic analysis of water-logging tolerance in tropical maize (Zea mays L.). Maydica 55:17–26
Zaidi PH, Rashid Z, Vinayan MT, Almeida GD, Phagna RK, Babu R (2015) QTL mapping of agronomic waterlogging tolerance using recombinant inbred lines derived from tropical maize (Zea mays L.) germplasm. PLoS ONE 10:e0124350
Zhang X, Tang B, Yu F, Li L, Wang M, Xue Y, Zhang Z, Yan J, Yue B, Zheng Y, Qiu F (2012) Identification of major QTL for waterlogging tolerance using genome-wide association and linkage mapping of maize seedlings. Plant Mol Biol Rep 31:594–606
Zou C, Wang P, Xu Y (2016) Bulked sample analysis in genetics, genomics and crop improvement. Plant Biotechnol J 14:1941–1955
Acknowledgements
We would like to thank professor Mingliang Xu from National Maize Improvement Center, China Agricultural University, for his suggestions on the manuscript.
Funding
This study was funded by the National Key Research and Development Program of China (2017YFD0101201), Central Public-interest Scientific Institution Basal Research Fund (Y2020PT20), Agricultural Science and Technology Innovation Program (ASTIP) of Chinese Academy of Agricultural Sciences (CAAS) (CAAS-XTCX2016009), National Natural Science Foundation (31771801, 32072069), the Plan in Scientific and Technological Innovation Team of Outstanding Young (T201704), and Research activities of CIMMYT staff were supported by the Bill and Melinda Gates Foundation and the CGIAR Research Program MAIZE.
Author information
Authors and Affiliations
Contributions
YX, HD, and ZG designed the experiments. ZG, SZ, HD, WXL, and SW performed the experiments. ZG, FC, and HD analyzed the data. ZG and YX wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
Author Zifeng Guo declares that he has no conflict of interest. Author Shuangzhen Zhou declares that she has no conflict of interest. Author Shanhong Wang declares that she has no conflict of interest. Author Wen-Xue Li declares that he has no conflict of interest. Author Hewei Du declares that he has no conflict of interest. Author Yunbi Xu declares that he has no conflict of interest.
Consent for publication
Consent was obtained from all individual participants included in the study
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Communicated by: Izabela Pawłowicz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Fig.1
Distribution of eight seedling traits under the control (blue) and waterlogging conditions (red). PH plant height; RL root length; SDW shoot dry weight; RDW root dry weight; ARN adventitious root number; BRNN node number of brace roots; BRN brace root number; BRDW brace root dry weigh (PDF 246 kb)
Supplementary Fig.2
Estimation of the Ln (probability of data). Delta K was calculated from K =2 to K = 9 (PDF 260 kb)
Supplementary Fig. 3
Manhattan plots of GWAS and Quantile-Quantile plots for RARN, RSDW, and RRL in the maize association panel. Manhattan plots for RBRN, RSDW and RRL are shown in a, c and e, respectively and the Quantile-Quantile plots for RBRN, RSDW and RRL are shown in b, d, f, respectively. The red lines show genome-wide significance at stringent thresholds of 2.52 x10-5[-log (1/39,266)]. (PDF 231 kb)
Supplementary Fig. 4
Manhattan plots of GWAS and Quantile-Quantile plots for RBRN, RBRNN, and RBRDW in the maize association panel. Manhattan plots for RBRN, RBRNN, and RBRDW are shown in a, c and e, respectively and the Quantile-Quantile plots for RBRN, RBRNN, and RBRDW are shown in b, d, f, respectively. The red lines show genome-wide significance at stringent thresholds of 2.52 x10-5[-log (1/39,266)]. (PDF 234 kb)
Supplementary Table 1
(DOCX 90 kb)
Supplementary Table 2
(DOCX 34 kb)
Supplementary Table 3
(DOCX 131 kb)
Supplementary Table 4
(DOCX 66 kb)
Supplementary Table 5
(DOCX 44 kb)
Rights and permissions
About this article
Cite this article
Guo, Z., Zhou, S., Wang, S. et al. Identification of major QTL for waterlogging tolerance in maize using genome-wide association study and bulked sample analysis. J Appl Genetics 62, 405–418 (2021). https://doi.org/10.1007/s13353-021-00629-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-021-00629-0