Abstract
MicroRNAs (miRNAs) are small non-coding RNAs which regulate gene expressions post-transcriptionally. Nowadays, various miRNAs have been found to be sensitive and specific biomarkers for the early diagnosis of colorectal cancer (CRC); however, there are different, even conflicting results in different publications concerning the diagnostic accuracy of miRNA. Therefore, we aim to conduct a meta-analysis of the relevant publications to comprehensively evaluate the diagnostic value of miRNAs in CRC detection. Several public databases such as PubMed, Embase, and Google Scholar were retrieved up to July 13, 2014. Sensitivity was applied to plot the summary receiver operator characteristic (SROC) curve against specificity. The area under the SROC curve (AUC) was calculated to assess the classified effects. STATA 12.0 software was used to perform all statistic analyses. A total of 29 articles, including 80 studies, were involved in our meta-analysis, 55 of which focus on single-miRNA assays and the other 25 on multiple-miRNA assays. Our results suggested that multiple-miRNA assays show a better diagnostic accuracy compared with single-miRNA assays. In addition, blood-based miRNA assays were more accurate than feces-based miRNA assays in CRC diagnosis. Our results also showed that miRNA diagnosis appear to be more accurate in Asians than in Caucasians. However, further researches are needed to validate our results and the feasibility of miRNAs as biomarkers in routine clinical diagnosis of CRC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As the third most common cancer in men and the second one in women, colorectal cancer (CRC) accounts for approximately 8 % of all the cancer-related deaths [1]. The overall incidence of CRC is 5 % in the general population, and the 5-year survival rate ranges from 40 to 60 % [2]. In spite of advances in the CRC treatment, early diagnosis is still really crucial to improve the survival rate of cancer patients. Early detection of CRC depends on traditional screening technology, but with several defects. For instance, although colonoscopy is regarded as gold-standard test for CRC, it has a bad experience [3]. The diagnostic method of fecal occult blood test (FOBT) is accepted widely, but has a low sensitivity and specificity. For the method of fecal immunochemical tests (FIT), it is proven to be more sensitive in CRC, but the sensitivity is not high enough to detect CRC [4]. The fecal DNA and messenger RNA (mRNA) testing are non-invasive techniques, but they need more optimization to be applied in the clinical examination [5]. Therefore, it is necessary to explore new efficient and non-invasive methods for CRC detection.
MicroRNAs (miRNAs) are small non-coding RNAs (18–25 nucleotides long), and regulate the gene expressions by targeting the mRNA post-transcriptionally and binding mRNA in the 3’ untranslated regions (UTR) [6]. Further studies demonstrate that miRNAs play crucial roles in cell proliferation, differentiation, and apoptosis [7]. Increasing evidences show the different expression profile of miRNAs in blood was observed in the patients who have prostate cancer, lymphoma, ovarian cancer, lung cancer, gastric cancer, or CRC. In addition, miRNAs in feces have also been tested as an indicator to detect the CRC non-invasively, and a number of miRNAs have been reported to be closely associated with CRC, such as miR-106a, miR-143, miR-145, and miR-9 [3]. Therefore, the miRNAs in blood or feces have great potential to be biomarker for CRC diagnosis.
There are several researches that evaluate the feasibility of miRNA as biomarkers for CRC detection. An overexpression of miR-221 is found in cancer patients’ blood, which implies that miR-221 may play an important role in tumorigenesis. Therefore, it is possible to use miR-221 as a biomarker to diagnose CRC. An increasing number of miRNAs, like miR-21 miR-129-3p, miR-767-3p miR-877, miR-60, miR-760, and miR-92a, etc., are reported to have similar effect like miR-221. However, most of the researches were performed on a relatively small sample. Thus, we conduct a comprehensive meta-analysis of miRNAs for CRC diagnosis based on the former publications.
Materials and methods
Literature search strategy
We performed a meta-analysis according to diagnostic meta-analysis guidelines. Eligible studies were retrieved from several online public databases, including PubMed, Embase, Google Scholar, the Cochrane Library, and the Chinese National Knowledge Infrastructure (CNKI), up to July 13, 2014. The search terms include: (“microRNAs” or “miRNA” or “miR”) and (“colorectal neoplasms” or “colorectal cancer” or “colorectal malignancy”) and (“diagnosis” or “sensitivity” or “specificity” or “ROC curve”). In addition, the reference lists of relevant reviews were manually searched to obtain additional articles.
Publications inclusion and exclusion criteria
Studies qualified to be included have to fulfill the following inclusion criteria: (1) studies related to diagnostic value of miRNAs for CRC; (2) studies using the gold standard to confirm the diagnosis of CRC patients; and (3) studies providing sufficient data to construct two-by-two tables, including true positive (TP), false positive (FP), true negative (TN), and false negative (FN). Exclusion criteria were (1) studies not related to the diagnostic values of miRNAs for CRC; (2) studies with duplicate data from other studies; and (3) studies in the form of letters, editorials, case reports, or reviews.
Data extraction
The following data were extracted from all eligible studies using a standardized form: (1) basic characteristics of publications, name of the first author, year of publication, country of publication, ethnicity, sample sizes, age, gender ratio, and specimen types included; (2) diagnostic outcomes, including sensitivity, specificity, TP, FP, FN, and TN, respectively.
Quality assessment
Each article was evaluated systematically by reviewers according to updated Quality Assessment of Diagnostic Accuracy Studies 2 (QUADAS-2) criteria [8]. This score system is comprised of four basic ingredients: patient selection, the index test, the reference standard, and flow and timing. There are seven questions required to be answered with “yes”, “no”, or “unclear”. The answer “yes” gets one score, while the answer “no” or “unclear” answers are given zero score. All questions have equal weight that the maximum score is 7.
Statistic analysis
All analyses were performed using the STATA 12.0 software. We calculated the pooled sensitivity, specificity, positive likelihood ratio (PLR), negative likelihood ratio (NLR), and diagnostic odds ratio (DOR) of all included publications with bivariate meta-analysis model. Since the area under the SROC curve (AUC) was a pooled evaluation index of the diagnosis, it was calculated by applying the sensitivity and specificity of each publication to plot the summary receiver operator characteristic (SROC) curve [9]. Heterogeneity among included studies was evaluated by the Q test and I 2 statistic [10]. If the P value was smaller than 0.05 for the Q test or the I 2 value ≥ 50 %, it means that there is significant heterogeneity, and then the random-effects model should be applied. In addition, we also conducted subgroup and meta-regression analyses to explore potential sources of between-study heterogeneity. The Fagan’s nomogram was used to estimate the post-diagnostic effect after pooled analysis [11]. In addition, we performed Deek’s funnel plot asymmetry test to evaluate the publication bias, P < 0.10 imply significant publication bias [12].
Results
Literature search
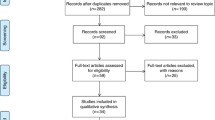
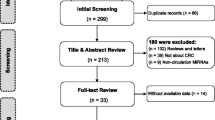
The results of our literature search were listed in Fig. 1. The initial database search includes 397 potential relevant articles, of which 46 duplicates were excluded. Then, 351 study articles are left for further examination. After reviewing abstract and keywords, 138 letters, reviews, and meta-analysis were further excluded, the study subjects of 5 articles were not human, and 136 of them were not related to our research. Then, 72 articles remain for the full-text review. After the full-texts review, another 43 studies were excluded, including 14 studies irrelevant to diagnosis, 19 studies without sufficient data, 5 studies about metastatic disease, and other 5 studies with duplicate data. Finally, 29 eligible articles were included in our meta-analysis, including 19 articles about blood-based miRNAs assays [13–31], and 10 articles about feces-based miRNAs assays [3, 5, 32–39].
Basic characteristics of included studies
The characteristics of the included studies are shown in Table 1. Finally, 80 trials from 29 articles were included in the present meta-analysis, including 55 single-miRNA assays, and 25 multiple-miRNA assays. The participants of the selected articles are Asian and Caucasian. The number of total CRC patients was 2626, and the number of all healthy controls was 2017. The specimens used in these 29 articles including serum, plasma, blood, and feces, which can be categorized into two general class of specimens, “blood” and “feces”. The expression levels of miRNA are measured by the real-time reverse transcription polymerase chain reaction (RT-PCR) in most of the selected articles. The qualities of these articles were assessed with QUADAS-2 system, and the assessment results, which were shown in Fig. 2, turned out to be moderate to high.
Diagnostic accuracy of miRNA for CRC
There is significant heterogeneity among the sensitivity and specificity results in the 80 studies of the 29 articles. For the sensitivity, the I 2 was 81.96 % (95 % CI, 78.14–85.78), and Q value was 376.54 (P < 0.01); for the specificity, the I 2 was 83.21 (95 % CI, 79.72–86.69), and Q value was 404.91 (P < 0.01). So, the random-effects model was selected in the next steps. The pooled estimates results are displayed in Table 2. The assessment results for overall studies are presented as follows: sensitivity, 0.74 (95 % CI, 0.70–0.77); specificity, 0.81 (95 % CI, 0.77–0.84); PLR, 3.9 (95 % CI, 3.2–4.7); NLR, 0.32 (95 % CI, 0.28–0.37); DOR, 12 (95 % CI, 9–16); and AUC, 0.84 (95 % CI, 0.81–0.87), suggesting a moderate diagnostic accuracy. The likelihood ratio was moderate as well as post-test probabilities, shown in the Fagan’s plot (Fig. 3). Likelihood ratios (LRs) provide a pooled effect by combining the sensitivity and specificity. In detail, for any people with a pre-test probability of 25 % to have CRC, if a positive test was given by miRNA assay with PLR value at 3, the post-test probability to have NDs would rise to 53 %; by contrast, for the same people, a negative result of miRNA assay would lower the post-test probability to 10 %. Hence, miRNA assay may play an important role as initial screening method for CRC.
Subgroup analysis
The pooled results for diagnostic accuracy in different subgroups were displayed in Table 2. There are three subgroups, and the basic criteria for clarification were ethnicity, miRNA profiling (single- or multiple-miRNA), and sample types, respectively. For the ethnicity subgroups, the sensitivity, specificity, PLR, NLR, and DOR of Caucasian-based miRNA assays were 0.66 (95 % CI, 0.59–0.73), 0.74 (95 % CI, 0.69–0.79), 2.6 (95 % CI, 2.0–3.3), 0.45 (95 % CI, 0.35–0.59), and 6 (95 % CI, 3–9), respectively, with AUC of 0.77 (95 % CI, 0.73–0.80) (Fig. 4a). The diagnostic sensitivity, specificity, PLR, NLR, and DOR of Asian-based miRNA assays were 0.77 (95 % CI, 0.73–0.81), 0.84 (95 % CI, 0.79–0.88), 4.8 (95 % CI, 3.7–6.1), 0.28 (95 % CI, 0.24–0.33), and 17 (95 % CI, 13–24), respectively, with AUC of 0.87 (95 % CI, 0.84–0.89) (Fig. 4b). It seemed to be that the diagnostic accuracy of Asian-based miRNA assays might be higher than Caucasian-based miRNA assays.
For the diagnostic accuracy of single-miRNA assays, the sensitivity was 0.69 (95 % CI, 0.64–0.74), the specificity was 0.80 (95 % CI, 0.75–0.84), the PLR was 3.5 (95 % CI, 2.7–4.4), the NLR was 0.38 (95 % CI, 0.33–0.45), the DOR was 9 (95 % CI, 6–13), and the AUC was 0.77 (95 % CI, 0.73–0.80) (Fig. 4c). For multiple-miRNA assays, the sensitivity was 0.82 (95 % CI, 0.79–0.85), the specificity was 0.83 (95 % CI, 0.78–0.86), the PLR was 4.7 (95 % CI, 3.7–6.0), the NLR was 0.21 (95 % CI, 0.18–0.26), the DOR was 22 (95 % CI, 15–31), and the AUC was 0.89 (95 % CI, 0.86–0.92) (Fig. 4d). The data shows multiple-miRNA assays were more accurate than single-miRNA assays for CRC diagnosis.
Similarly, for the subgroups of specimens, the blood-based miRNA assays exhibit a better performance to feces-based miRNA assays, with the AUCs of 0.84 versus 0.82. However, in terms of DOR, the feces-based miRNA assays of 15 (95 % CI, 8–31) had a better performance in CRC diagnosis than blood-based assays of 11 (95 % CI, 8–16). In addition, blood-based assays had a higher diagnostic sensitivity of 0.76 (95 % CI, 0.71–0.80) than feces-based assays of 0.68 (95 % CI, 0.60–0.75); on the opposite, feces-based assays had a higher diagnostic specificity of 0.88 (95 % CI, 0.78–0.93) than blood-based assays of 0.78 (95 % CI, 0.75–0.81). In conclusion, blood-based assays, with a lower specificity, were more sensitive than feces-based assays in CRC diagnosis.
Meta-regression and robustness analyses
Meta-regression analysis was performed to further explore potential sources of the heterogeneity and to testify the results of subgroup analysis (Fig. 4). For the sensitivity, Asian (P < 0.01), multiple-miRNA assay (P < 0.001), and blood-based specimen (P < 0.001) were found as the main factors that contributed to the heterogeneity. For diagnostic specificity, Asian (P < 0.001), multiple-miRNA assay (P < 0.001), and blood-based specimen (P < 0.001) were also essential elements to affect the heterogeneity. The conclusion can be drawn that ethnicity, miRNA profiling, and types of specimen were significant factors affecting the heterogeneity of both the diagnostic sensitivity and specificity. Goodness of fit and bivariate normality analyses indicated that the random-effects bivariate model was robust for the pooled analyses. Eleven deviated studies were found in influence analysis, and eight outliers were discovered in outlier detection. Some duplicate outliers were found in the two different methods. In summary, there were 11 deviated studies that may affect the robustness of the meta-analysis. After excluding the deviated studies or outliers, the pooled diagnostic sensitivity, specificity, PLR, NLR, DOR, and AUC were 0.73 (95 % CI, 0.70–0.76), 0.78 (95 % CI, 0.75–0.81), 3.4 (95 % CI, 2.9–3.9),0.34 (95 % CI, 0.30–0.39), 10 (95 % CI, 8–13), and 0.82 (95 % CI, 0.79–0.86), respectively, without significant differences relative to overall analysis with outliers, indicating that our meta-analysis was robust.
Discussion
CRC is a kind of malignant disease, with increasing incidence worldwide. The low accuracy of early diagnosis and high prevalence of late diagnosis lead to high mortality rate. So, it is crucial to find an effective early-stage diagnostic method for CRC patients to prevent the cancer deterioration, thus the patients will have a greater chance to survive [40]. Since miRNAs are widely reported to be dysregulated in cancer patients, it is proposed to be a potential biomarker for CRC detection without any invasive procedures involved [41]. MiR-223 and miR-451, measured in the feces samples, appear to have an aberrant expression profile between the CRC patients and healthy people [39]. The significant difference of expression level was also discovered in miR-21, let-7g, miR-31, miR-92a, miR-181b, and miR-203. Thus, miRNAs could be an ideal biomarker for CRC detection with less invasion and more accuracy, as well as lower costs, compared with the other diagnostic methods [29].
However, most of these studies just focused on the diagnostic value of only one single miRNA or a combination of miRNAs. Also, the different specimens such as blood, serum, plasma, or feces are analyzed among the selected specimens. Therefore, this meta-analysis was necessary to summarize and evaluate the accuracy of miRNA as biomarker based the included studies. The pooled sensitivity was 0.74 (95 % CI, 0.70–0.77), specificity was 0.81 (95 % CI, 0.77–0.84), and AUC was 0.84 (95 % CI, 0.81–0.87). Therefore, miRNAs are proven to be highly sensitive and specific diagnostic biomarkers. What is more, the DOR was 12 (95 % CI, 9–16), indicating that miRNA test-positive patients have 12-times higher chance of CRC than those healthy persons.
Our meta-analysis also has several interesting results. Firstly, it is more accurate to use miRNAs as CRC diagnostic biomarkers in the Asian group than in the Caucasian group. What is more, multiple-miRNA assay as biomarker is more accurate than single-miRNA assay. Therefore, multiple-miRNA assays tend to be given a more reliable diagnostic result, thus it is important to test multiple useful miRNAs to increase the credibility of results in clinical examination. According to the results in the individual study, panels of miR-203, −146b-5p, −215, −192, and -30e-3p (100 % sensitivity and 93 % specificity) in the Sheinerman el al. study and miR-21,-31, −92a, −181b, −203, and let-7 g (96.4 % sensitivity and 88.1 % specificity) in the Wang et al. study are the most predictive miRNA among all. Thirdly, CRC diagnosis with blood-based specimen is more sensitive but less specific than feces-based specimen.
To some extent, heterogeneity may undermine the reliability of this meta-analysis. Thus, in the beginning, we have tested the heterogeneity by a forest analysis (Fig. 3). According to the I 2 and Q value, a random-effects model was selected. Then, we performed subgroup analysis to detect the main factors that contributed to the heterogeneity. As a result, ethnicity, miRNA profiling (single or multiple miRNA), and specimen types are verified to be the main elements leading to heterogeneity. Also, eleven deviated studies were found that may contribute to heterogeneity. Then, we adopted these eleven studies and made a SROC curve again. However, the result without deviated studies was not significantly different from that with these deviated studies. It means that deviated studies had few impacts on our overall CRC meta-analysis.
Although there was existing meta-analysis that investigate the diagnostic value of miRNAs as biomarker to detect CRC, this present meta-analysis is the most comprehensive one. Only 13 studies were included in the previous meta-analysis by Zhou et al. (2013), with the pooled sensitivity of 0.81 (95 % CI, 0.79–0.84), specificity of 0.78 (95 % CI, 0.75–0.82), and AUC of 0.89 [42]. And their subgroup analysis was conducted based on the specimen only. However, a relatively more comprehensive analysis with 16 publications was performed in our meta-analysis. Apart from the specimen, the subgroup analysis of our study was also conducted based on the ethnicity and miRNA profiling. A conflict of result was found between the meta-analysis of Zhou et al. (2013) and ours. Zhou et al. (2013) had a conclusion that blood-based miRNA tests were more accurate than feces-based tests. However, our result was implied that blood-based CRC diagnosis had a higher sensitivity but lower specificity than feces-based diagnosis. In addition, in our present meta-regression, three main factors were found to contribute to the heterogeneity, including ethnicity, miRNA profiling, and specimen.
Despite our great efforts to perform an accurate analysis, there were also several limitations that should be noticed: (1) some useful publications may be missed in the process of retrieving and screening; (2) there was statistical heterogeneity in our analysis attributed to ethnicity, miRNA profiling, and specimen types; (3) no articles were found to report data relevant to African populations; and (4) the cut-off values were different or deficient in retrieved publications so that we did not extract the cut-off values.
In conclusion, we comprehensively assessed the value of miRNAs as biomarkers to detect CRC, and supported miRNAs as biomarkers in our meta-analysis. Moreover, the miRNA diagnostic analysis of CRC was more accurate in Asian populations than Caucasian populations. In addition, multiple-miRNA assay shows a higher accuracy than single-miRNA array.
References
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer. 2010;127:2893–917. doi:10.1002/ijc.25516.
Weitz J, Koch M, Debus J, Hohler T, Galle PR, Buchler MW. Colorectal cancer. Lancet. 2005;365:153–65. doi:10.1016/S0140-6736(05)17706-X.
Wu CW, Ng SC, Dong Y, Tian L, Ng SS, Leung WW, et al. Identification of microRNA-135b in stool as a potential noninvasive biomarker for colorectal cancer and adenoma. Clin Cancer Res. 2014. doi:10.1158/1078-0432.CCR-13-1750.
Morikawa T, Kato J, Yamaji Y, Wada R, Mitsushima T, Shiratori Y. A comparison of the immunochemical fecal occult blood test and total colonoscopy in the asymptomatic population. Gastroenterology. 2005;129:422–8. doi:10.1016/j.gastro.2005.05.056.
Wu XD, Song YC, Cao PL, Zhang H, Guo Q, Yan R, et al. Detection of miR-34a and miR-34b/c in stool sample as potential screening biomarkers for noninvasive diagnosis of colorectal cancer. Med Oncol. 2014;31:1–6.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31. doi:10.1038/nrg1379.
Whiting PF, Rutjes AW, Westwood ME, Mallett S, Deeks JJ, Reitsma JB, et al. QUADAS-2: a revised tool for the quality assessment of diagnostic accuracy studies. Ann Intern Med. 2011;155:529–36. doi:10.7326/0003-4819-155-8-201110180-00009.
Deville WL, Buntinx F, Bouter LM, Montori VM, de Vet HC, van der Windt DA, et al. Conducting systematic reviews of diagnostic studies: didactic guidelines. BMC Med Res Methodol. 2002;2:9.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–60. doi:10.1136/bmj.327.7414.557.
Yang X, Guo Y, Du Y, Yang J, Li S, Liu S, et al. Serum MicroRNA-21 as a diagnostic marker for lung carcinoma: a systematic review and meta-analysis. PLoS ONE. 2014;9:e97460. doi:10.1371/journal.pone.0097460.
Deeks JJ, Macaskill P, Irwig L. The performance of tests of publication bias and other sample size effects in systematic reviews of diagnostic test accuracy was assessed. J Clin Epidemiol. 2005;58:882–93. doi:10.1016/j.jclinepi.2005.01.016.
Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58:1375–81. doi:10.1136/gut.2008.167817.
Huang Z, Huang D, Ni S, Peng Z, Sheng W, Du X. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer. 2010;127:118–26.
Pu XX, Huang GL, Guo HQ, Guo CC, Li H, Ye S, et al. Circulating miR-221 directly amplified from plasma is a potential diagnostic and prognostic marker of colorectal cancer and is correlated with p53 expression. J Gastroenterol Hepatol. 2010;25:1674–80. doi:10.1111/j.1440-1746.2010.06417.x.
Kanaan Z, Rai SN, Eichenberger MR, Roberts H, Keskey B, Pan J, et al. Plasma MiR-21: a potential diagnostic marker of colorectal cancer. Ann Surg. 2012;256:544–51.
Chen X, Hu Z, Wang W, Ba Y, Ma L, Zhang C, et al. Identification of ten serum microRNAs from a genome-wide serum microRNA expression profile as novel noninvasive biomarkers for nonsmall cell lung cancer diagnosis. Int J Cancer. 2012;130:1620–8. doi:10.1002/ijc.26177.
Wang Q, Huang Z, Ni S, Xiao X, Xu Q, Wang L, et al. Plasma miR-601 and miR-760 are novel biomarkers for the early detection of colorectal cancer. PLoS ONE. 2012;7:e44398. doi:10.1371/journal.pone.0044398.
Feng L, Pang Z, Sha S, Wu B, Xu F, Yin SP. The value of diagnosis and prognosis prediction of serum miR-29a and miR-92a for colorectal cancer. Chin J Clin Oncol Rehabil. 2013:1313–1315.
Giraldez MD, Lozano JJ, Ramirez G, Hijona E, Bujanda L, Castells A, et al. Circulating microRNAs as biomarkers of colorectal cancer: results from a genome-wide profiling and validation study. Clin Gastroenterol Hepatol. 2013;11:681–688.e683.
Kanaan Z, Roberts H, Eichenberger MR, Billeter A, Ocheretner G, Pan J, et al. A plasma microRNA panel for detection of colorectal adenomas: a step toward more precise screening for colorectal cancer. Ann Surg. 2013;258:400–8. doi:10.1097/SLA.0b013e3182a15bcc.
Liu GH, Zhou ZG, Chen R, Wang MJ, Zhou B, Li Y, et al. Serum miR-21 and miR-92a as biomarkers in the diagnosis and prognosis of colorectal cancer. Tumour Biol. 2013;34:2175–81. doi:10.1007/s13277-013-0753-8.
Luo X, Stock C, Burwinkel B, Brenner H. Identification and evaluation of plasma microRNAs for early detection of colorectal cancer. PLoS ONE. 2013;8:e62880. doi:10.1371/journal.pone.0062880.
Sheinerman KS, Tsivinsky VG, Umansky SR. Analysis of organ-enriched microRNAs in plasma as an approach to development of universal screening test: feasibility study. J Transl Med. 2013; 11.
Toiyama Y, Takahashi M, Hur K, Nagasaka T, Tanaka K, Inoue Y, et al. Serum miR-21 as a diagnostic and prognostic biomarker in colorectal cancer. J Natl Cancer Inst. 2013;105:849–59.
Wang S, Xiang J, Li Z, Lu S, Hu J, Gao X, et al. A plasma microRNA panel for early detection of colorectal cancer. Int J Cancer. 2013. doi:10.1002/ijc.28136.
Yong FL, Law CW, Wang CW. Potentiality of a triple microRNA classifier: MiR-193a-3p, miR-23a and miR-338-5p for early detection of colorectal cancer. BMC Cancer. 2013; 13.
Zhang GJ, Zhou T, Liu ZL, Tian HP, Xia SS. Plasma miR-200c and miR-18a as potential biomarkers for the detection of colorectal carcinoma. Mol Clin Oncol. 2013;1:379–84.
Hu J, Liu C, Yin Q, Ying M, Li J, Li L, et al. Association between the CYP1A2-164 A/C polymorphism and colorectal cancer susceptibility: a meta-analysis. Mol Genet Genomics. 2014;289:271–7. doi:10.1007/s00438-013-0806-0.
Zanutto S, Pizzamiglio S, Ghilotti M, Bertan C, Ravagnani F, Perrone F, et al. Circulating miR-378 in plasma: a reliable, haemolysis-independent biomarker for colorectal cancer. Br J Cancer. 2014;110:1001–7. doi:10.1038/bjc.2013.819.
Zhang L, Meng L, Fan Z, Liu B, Pei Y, Zhao Z. Expression of plasma miR-106a in colorectal cancer and its clinical significance. Nan Fang Yi Ke Da Xue Xue Bao. 2014;34:354–7.
Koga Y, Yasunaga M, Takahashi A, Kuroda J, Moriya Y, Akasu T, et al. MicroRNA expression profiling of exfoliated colonocytes isolated from feces for colorectal cancer screening. Cancer Prev Res. 2010;3:1435–42.
Kalimutho M, Del Vecchio Blanco G, Di Cecilia S, Sileri P, Cretella M, Pallone F, et al. Differential expression of miR-144*as a novel fecal-based diagnostic marker for colorectal cancer. J Gastroenterol. 2011;46:1391–402.
Koga Y, Yasunaga M, Takahashi A, Kuroda J, Matsumura Y. Fecal RNA test using microRNA expressions of exfoliated colonocytes for colorectal cancer screening. Cancer Res. 2011; 71.
Wu CW, Dong YJ, Ng S, Leung WW, Wong CY, Sung JJ, et al. Identification of a panel of MicroRNAs in stool as screening markers for colorectal cancer. Gastroenterology. 2011;140:S73.
Kuriyama S, Hamaya Y, Yamada T, Sugimoto M, Osawa S, Sugimoto K, et al. Fecal microRNA assays as a marker for colorectal cancer screening. Gastroenterology. 2012;142:S770.
Wu CW, Ng SSM, Dong YJ, Ng SC, Leung WW, Lee CW, et al. Detection of miR-92a and miR-21 in stool samples as potential screening biomarkers for colorectal cancer and polyps. Gut. 2012;61:739–45.
Koga Y, Yamazaki N, Yamamoto Y, Yamamoto S, Saito N, Kakugawa Y, et al. Fecal miR-106a is a useful marker for colorectal cancer patients with false-negative results in immunochemical fecal occult blood test. Cancer Epidemiol Biomarkers Prev. 2013;22:1844–52.
Phua LC, Chue XP, Koh PK, Cheah PY, Chan EC, Ho HK. Global fecal microRNA profiling in the identification of biomarkers for colorectal cancer screening among Asians. Oncol Rep. 2014. doi:10.3892/or.2014.3193.
Andreoli SC, Gasparini NJ, Carvalho GP, Garicochea B, Pogue RE, Andrade RV. Use of microRNAs in directing therapy and evaluating treatment response in colorectal cancer. Einstein (Sao Paulo). 2014;12:256–8.
Schou JV, Rossi S, Jensen BV, Nielsen DL, Pfeiffer P, Hogdall E, et al. miR-345 in metastatic colorectal cancer: a non-invasive biomarker for clinical outcome in non-KRAS mutant patients treated with 3rd line cetuximab and irinotecan. PLoS ONE. 2014;9:e99886. doi:10.1371/journal.pone.0099886.
Zhou XJ, Dong ZG, Yang YM, Du LT, Zhang X, Wang CX. Limited diagnostic value of microRNAs for detecting colorectal cancer: a meta-analysis. Asian Pac J Cancer Prev. 2013;14:4699–704.
Acknowledgments
This work was supported in part by the grant from National Natural Science Foundation of China (Grant # 81272251), a grant for the Innovative Team of Science and Technology, Henan Province, China.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, X., Zhong, J., Ji, Y. et al. The expression and clinical significance of microRNAs in colorectal cancer detecting. Tumor Biol. 36, 2675–2684 (2015). https://doi.org/10.1007/s13277-014-2890-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2890-0