Abstract
Background
MicroRNAs (miRNA) are tiny, noncoding, small, endogenous RNAs that play major roles in neoplastic transformation and could therefore offer a better quantitative and noninvasive method for the diagnosis and prognosis of colorectal cancer (CRC) using feces. In the present study, we screened feces for 648 miRNAs and analyzed the role of miR-144* as a potential CRC diagnostic marker.
Methods
Fecal miRNA expression was profiled with RT-pre-amplification-qPCR, and the stability was determined using both endogenous and exogenous miRNA by RT-qPCR. ROC analysis was performed to enhance the diagnosing power of the CRC patients’ fecal specimens.
Results
We detected 39% of all the miRNAs screened in feces. Endogenous miRNAs are more stable over time and temperature, while exogenous miRNAs degraded rapidly. miR-144* was overexpressed in feces, suggesting that it could be a potent candidate diagnostic marker for CRC detection, with a sensitivity of 74% and a specificity of 87% (n = 75, p < 0.0001). Moreover, RT-qPCR analysis showed that miR-144* was also overexpressed in paired CRC tissues, thus suggesting its possible utilization as a diagnostic marker.
Conclusions
We demonstrated that miRNAs are stable in the fecal microenvironment, and that, among them, miR-144* represents a novel fecal-based diagnostic marker for CRC screening. Nevertheless, our data need to be validated in a large cohort of subjects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
MicroRNAs (miRNAs) are a small class of regulatory RNA molecules that modulate the activity of specific mRNA targets and display important roles in a wide range of physiological and pathological processes [1–4]. They guide the RNA-induced silencing complex to target sites, which are thought to be most prevalent in the 3′ untranslated region of some mRNAs [5]. Moreover, they also potentially target up to one-third of all human coding genes, making their role in cellular biology even more important [6]. Initially, miRNAs were discovered in Caenorhabditis elegans as small temporal RNAs (stRNAs) that regulate developmental transitions [7]. Since then, hundreds of members of the class of miRNAs have been found in worms, flies, plants and mammals. Although the mechanisms through which miRNAs regulate their target genes are still gradually being discovered, the finding that at least some miRNAs feed into the RNA interference (RNAi) pathway has provided a starting point for the discovery and elucidation of the biological roles of miRNAs in normal physiology and human diseases, particularly cancers (for more on this topic, refer to [7, 8]).
At the time that this manuscript was written, 1413 human miRNAs had been reported bioinformatically in miRBase (http://www.mirbase.org/cgi-bin/query.pl?terms=hsa-miR), and these require experimental validation. These molecules are implicated in numerous cellular processes, including development, differentiation, proliferation, apoptosis, DNA damage repair and stress responses, and they are frequently deregulated in cancers [9, 10]. miRNA expression varies with tumor type, including colorectal cancer (CRC) [1, 2, 9–12]. Furthermore, miRNA expression signatures have been shown to be clustered according to tumor type, and to predict prognosis [11–14].
Recent reports have proposed the possibility of detecting miRNAs from serum and blood-based bodily fluids as novel targets to interpret disease state and as biomarkers in various malignancies [15–17]. miRNA expression is tissue specific, and it has unusually high stability in formalin-fixed tissues [18–20] and in bodily fluids [15–23]. In view of this observation, and reports published previously, the expression profile of deregulated miRNAs may represent an enhanced quantitative and diagnostic tool for cancer detection. Although various molecular methods have recently been proposed for CRC detection, most of them have presented relatively low sensitivities as screening modalities.
Cells exfoliated from malignant colonocytes undergo genetic changes that confer upon them the ability to escape from programmed cell death and shed more frequently than normal colonocytes in feces [24, 25]. This phenomenon allows us to distinguish the altered expressions of deregulated miRNAs in order to detect CRC using feces (for a detailed explanation, see [24–27]).
In this study, we assessed whether the altered expression of miRNAs detected in feces could serve as a good candidate biomarker for CRC detection. We profiled 648 miRNAs in pooled samples from subsets of diseased and healthy individuals and determined whether miRNAs are stably detected in feces. Moreover, RT-qPCR was performed to determine whether differential miRNA expression can be robustly and reliably measured in feces. With regards to the generated microarray data, we validated a chosen set of miRNAs that could diagnose CRC accurately with acceptable sensitivity and specificity. We showed that endogenous miRNAs are stably detected in feces, and that, among them, miR-144* represents a novel fecal-based diagnostic marker for CRC screening.
Materials and methods
Feces and tissues samples
Fecal specimens were collected from patients who underwent colonoscopy following a standardized procedure, as described previously [26, 27]. Briefly, patients were instructed to collect an aliquot of feces in a sealed plastic container and transport it to the hospital in an ice bag. The samples were stored at −20°C immediately upon receipt with fecal stabilization buffer. The samples were then processed according to the standard protocol, as described previously [26]. During colonoscopy, the colon was visualized until the cecum in all patients, independent of disease state, and the sizes and locations of any lesions were recorded. Later, the diagnosis of cancer was confirmed histologically and pathological stages were scored. Surgical tissues were also collected from another set of cancer patients who underwent colonoscopy and surgery in our institute and kept at −80°C. All subjects gave their informed consent prior to participation in this study. The study was approved by our institutional review board. miRNA analysis was conducted blindly by two technicians who evaluated the pooled and RT-qPCR assays.
Isolation of miRNAs from the feces of CRC and healthy subjects
Total miRNA was isolated from approximately 20–30 mg of feces of CRC and healthy individuals (n = 75) using the mirVana™ miRNA isolation kit with minor modifications from the manufacturer’s (Ambion, Inc.) guidelines. For the initial lysis step, we used 800 μl of lysis/binding solution diluted in feces and vortexed vigorously. Later, the mixture was subjected to centrifugation at 13000 rpm for 1 min. The supernatant was collected and processed according to the instructions that came with the mirVana isolation kit. The concentrations of all miRNA samples were quantified using NanoDrop 1000 (Nanodrop, Wilmington, DE, USA), and an equivalent amount of the total miRNAs of each patient was used for RT-qPCR analysis.
Fecal miRNA stability assay
Feces samples were spiked with synthetic cel-miR-238 from C. elegans, which has no sequence homology to human miRNAs, as described by Mitchell et al. [16] (miRBase accession number: MIMAT000093-UUUGUACUCCGAUGCCAUUCAGA; Primm SRL, Milan, Italy). Molar concentrations to be spiked in were derived empirically to produce C q values that were comparable to those of moderately abundant endogenous miRNAs measured in fecal specimens. Briefly, 20–30 mg of the fecal sample containing 2 μl of 10 μM cel-miR-238 and the negative control were re-suspended in duplicate in 400 μl of 2X denaturing solution (Ambion), which contains RNase inhibitor, or in PBS without RNase inhibitor. Samples were then processed immediately with the mirVana lysis/binding solution. An equivalent amount of total miRNA from each sample was reverse transcribed, and the level of cel-miR-238 was quantified based on RT-qPCR. For the temperature stability analysis, equal amounts of each fecal sample were placed at 4 and 25°C, and the levels of miR-16 and cel-miR-238 were monitored over periods of 0, 2, 4 and 6 days and quantified by standard curve analysis.
RDML data guidelines
Raw miRNA expression, experimental annotation, and sample annotation are available in the RDML data format [28].
Microarray miRNA reverse transcription
Feces from 15 CRC patients and 15 healthy controls were pooled to analyze the expression of 648 miRNAs. Total miRNA was extracted using the mirVana™ miRNA isolation kit, and then 8 μl of the total RNA (50 ng/μl) (the same amount of total miRNA was pooled for each patient, quantified by a NanoDrop 1000) were reverse transcribed using an miRNA reverse transcription kit (Applied Biosystems) in combination with the stem-loop Megaplex primer pool (Applied Biosystems), allowing the simultaneous reverse transcription of 648 miRNAs and 19 endogenous controls [23, 28]. Briefly, 8 μl of the total RNA (50 ng/μl) were supplemented with RT primer mix (10×), RT buffer (10×), MultiScribe reverse transcriptase (10 U/μl), dNTPs with dTTP (0.5 mM each), MgCl2 (3 mM), and AB RNase inhibitor (0.25 U/μl) in a total reaction volume of 80 μl. For RT reactions with subsequent pre-amplification, the reaction volume was proportionally reduced to 5 μl. The concentration of each stem-loop primer in the RT reaction mix was 1 nM, a 50-fold dilution compared with a singleplex RT reaction, ensuring minimal nonspecific interactions between the different stem-loop primers. To increase the reverse transcription efficiency, a pulsed RT reaction was used (40 cycles of 16°C for 2 min, 42°C for 1 min and 50°C for 1 s, followed by a final reverse transcriptase inactivation at 85°C for 5 min).
Pre-amplification of miRNA cDNA
Megaplex RT product (5 μl) was pre-amplified using Applied Biosystems’ TaqMan PreAmp Master Mix (2×) and the PreAmp Primer Mix (5×) in a 25-μl PCR reaction. The primer pool consisted of forward primers (50 nM) that were specific for each of the 667 miRNAs, including 19 endogenous control small RNAs and a universal reverse primer (50 nM) (Applied Biosystems). The pre-amplification cycling conditions were as follows: 95°C for 10 min, 55°C for 2 min and 75°C for 2 min, followed by 14 cycles of 95°C for 15 s and 60°C for 4 min [29, 30].
Real-time qPCR
For each cDNA sample, 648 miRNAs and 19 endogenous control small RNAs were profiled using a gene maximization PCR plate setup in a 384-well plate. This approach allowed us to profile one sample per 384-well plate. The RT product was diluted 400-fold; when pre-amplification was applied, the dilution factor was 1600. PCR amplification reactions were carried out in a total volume of 8 μl, containing 4 μl of TaqMan Master Mix (Applied Biosystems), 1 μl of cDNA and 3 μl of miRNA TaqMan probe and primers (primer details can be accessed through the Applied Biosystems website: http://www.appliedbiosystems.com). Cycling conditions were as follows: 95°C for 10 min, followed by 40 cycles of 95°C for 15 s, and 60°C for 1 min. All PCR reactions were performed on the 7900HT RT-qPCR system (Applied Biosystems). Raw C q values were calculated using SDS software v.2.1 with automatic baseline settings and a threshold of 0.224. The crossing point between the baseline-corrected amplification curve and the threshold line is called the quantification cycle (C q) (according to RDML guidelines, http://www.rdml.org) [28].
Selection of miRNAs/small RNA controls that resemble the mean expression value
In order to perform the robust and unbiased selection of genes with expression levels that had the highest correlations with the mean expression levels, we used the geometric mean calculation method [30, 31]. In brief, for each miRNA and small RNA control, we initially calculated the geometric mean of the RT-pre-amplification-q-PCR microarray data, and then we normalized our miRNA expression with the calculated geometric mean. Next, the standard deviations of the C q values of both groups were determined for every miRNA and small RNA control. The miRNAs and/or small RNA controls with the lowest standard deviations were considered to be good candidate reference genes for RT-qPCR analysis [30].
RT-qPCR analysis of miR-144*, miR-532-3p, and miR-378
We analyzed feces from 35 CRC patients and 40 healthy individuals for the expression of miR-144*, miR-532-3p, and miR-378 using multiplex RT-qPCR. After RNA extraction, the samples were retrotranscribed using a standard ABI RT reaction that contained 4 μl RNA (200 ng), 0.40 μl deoxynucleotide triphosphate mix, 3.0 μl 50 U/μl MultiScribe reverse transcriptase, 2.0 μl 10× RT buffer, 0.25 μl AB RNase inhibitor, 7.35 μl water, and 1.0 μl of each of the three primers in a total reaction mixture of 20.0 μl. The RT reaction was carried out at 16°C for 30 min and 42°C for 30 min. The reverse transcription products were diluted onefold with water (final concentration of 5 ng/μl), and 3.0 μl of cDNA were used for RT-qPCR assays. RT products were processed as follows: 95°C for 10 min, and then 95°C for 15 s and 60°C for 30 s for up to 40 cycles. The experiments were repeated twice with three technical replicates, and then the geometric mean of C q for each sample was calculated for gene expression via the \( 2^{{ - \Updelta \Updelta C_{\text{q}} }} \) method.
Isolation and quantification of miR-144* from CRC paired surgical tissues
Fifteen CRC paired surgical tissues (n = 30) were used for miR-144* expression analysis. Briefly, total RNA was extracted from tissues using the miRNeasy mini kit (Qiagen, Hilden, Germany), following the manufacturer’s instructions. The final elution volume was 30 μl. The concentrations of all miRNA samples were quantified using the NanoDrop 1000. RT-qPCR detection was carried out for each patient sample using the method described above for fecal miRNA detection.
Statistical analysis
Data analysis was performed using the MedCalc (v.9.2.0.1) software package. Receiver operating characteristic (ROC) curve analysis was used to determine the best cutoff value for the highest assay sensitivity and specificity. In addition, contingency table analysis, the independent paired Student’s t test, and one-way ANOVA were performed to evaluate the significance of the data where appropriate. Two-sided P values ≤0.05 were considered significant.
Results
Occurrence of miRNAs in feces
We first determined whether miRNAs could be detected in feces, which contain inhibitors and materials of nonhuman origin, such as DNA, RNA, proteins, polysaccharides and possible noncoding RNAs [26]. We isolated miRNAs from the feces of healthy and CRC individuals and checked the expression levels of miR-16 and miR-21 in the first instance. As expected, we detected the expression of these miRNAs in feces, independent of their disease state (data not shown).
Endogenous miRNAs are stably detected in the fecal microenvironment
Before checking for candidate miRNAs that can discriminate healthy from diseased subjects, we first carried out an experiment to verify the stability of miRNAs in feces. We assumed that endogenous fecal miRNAs are protected from degradation through their association with the macromolecules in feces. To test our assumption, we performed a spiked experiment with a known nonhuman miRNA sequence and measured the degradation pattern caused by endogenous RNase activity. We used cel-miR-238 as a model of exogenous miRNA, as this lacks sequence homology with human miRNAs. We were able to amplify the spiked cel-miR-238 in RT-qPCR analysis after inhibiting the existing RNase activity (Fig. 1A-a). A control experiment was carried out where the RNase activity was not inhibited, and it was found that cel-miR-238 was rapidly degraded within 10 min of exposure to the fecal samples (Fig. 1A-b). Conversely, when RNase inhibitor was added, we observed no decrease in the cel-miR-238 level. In addition to the spiking experiment, we also analyzed the expression of miR-16 in our spiked samples, and found that the miR-144* level was unaltered (data not shown). Thus, we demonstrated that endogenous miRNAs are more stable and resistant to RNases than exogenous miRNA in feces (Fig. 1a).
Analysis of the stability of exogenous cel-miR-238 and endogenous miR-16 in feces over time and at different temperatures. miRNAs were assessed by an RT-qPCR assay. a A Synthetic version of the Caenorhabditis elegans miRNA (cel-miR-238) was spiked into feces after the addition of an equal volume of 2× denaturing solution (to inhibit RNases). The feces were then incubated at room temperature for 10 min and a very high amplification was detected (curve a). The same amount of cel-miR-238 was spiked into feces prior to the addition of an equal volume of 2× denaturing solution, and a very low signal was detected at end of the RT-qPCR cycle (curve b). miR-16 stability analysis at 4°C (b) and 25°C (c) over a period of six days. No changes were observed when the feces were incubated at 4°C, and a slower degradation pattern was detected at 25°C on the fourth and sixth days. d Histogram demonstrating minor changes in the expression of miR-16 over a similar period of incubation (4–6 days) to that shown in b and c, using standard curve analysis
Fecal miRNAs are time and temperature stable
To further investigate the pre-analytical steps involved in fecal collection and transportation, we analyzed miR-16 stability profiles in samples stored at two different temperatures (4°C and 25°C) over a period of six days. We found that feces kept at 4°C (Fig. 1b, d) are more stable than feces stored at room temperature (Fig. 1c, d). We detected a slight degradation pattern of endogenous miR-16 in feces stored at room temperature during the fourth and sixth days relative to 4°C (Fig. 1d). Thus, these data indicate that endogenous miRNAs are relatively time and temperature stable in feces.
The presence of miRNAs in feces and the selection of a putative candidate biomarker
Having demonstrated that miRNAs are detectable and stable in feces, we further evaluated the deregulated expression of miRNAs in a subset of CRC and healthy individuals (n = 30). To do this, we employed an RT-preamp-qPCR microarray technique. A total of 648 miRNAs and 19 endogenous control small RNAs were analyzed, and we observed that 39% (253/648) of the miRNAs in feces. Out of this 39%, 82% (206/252) of the miRNAs were detected in healthy individuals, while 93% (234/252) were detected in CRC subjects. Next, we constructed a linear regression plot to determine the differences in CRC and healthy individuals based on the RT-pre-amp-qPCR microarray data. We found that at least 20% of the miRNAs showed different expression levels in the feces from CRC and healthy individuals (R 2 = 0.80) (Fig. 2a). Following the identification of differentially expressed miRNAs in feces, we identified 21 miRNAs that exhibited greater than fivefold upregulating in the CRC samples, although variations in the samples from individual patients may have influenced this result (Table 1; Fig. 2). We then narrowed down the miRNAs by ordering them from the highest expression levels to the lowest (Table 1). Based on this, miR-144* and miR-532-3p showed the greatest increases in their expression levels in CRC. We then re-evaluated the scatter plot to determine which of the miRNAs could serve as a normalizer, since its expression was not influenced by disease state (Fig. 2b). Thus, the miRNAs that were at the boundary of the scatter plot were possible candidates. In the end, we chose miR-378, which had the lowest standard deviation of C q in the RT-preamp-qPCR microarray experiment (SD; 0.00026) (Fig. 2b).
Regression plot analysis and relative fold changes in expression levels of miRNAs in feces of pooled samples from healthy and CRC subjects. a Linear regression analysis indicated that at least 20% of the miRNAs showed different levels of expression in the feces of CRC and healthy subjects. b A total of 648 miRNAs were profiled with the RT-pre-amplification-qPCR miRNA microarray. The scatter plot represents the relative fold changes in expression levels between control and CRC groups. The arrows show upregulated and downregulated miRNAs. The circles above line A show upregulated miRNAs, and the circles below line B show downregulated miRNAs, while the circles in between line A and B represent nonsignificantly expressed miRNAs. The middle line of the graph represents the boundary between upregulated and downregulated miRNAs
Furthermore, the pooled analysis revealed several miRNAs with differential expression, in accordance with literature observations (Table 1; Fig. 2b). However, we could not consider such miRNAs as candidates, as insignificant data were obtained for them from our preliminary investigation (data not shown). We also avoided selecting endogenous control small RNAs for two main reasons: (1) we could only detect three endogenous control small RNAs (RNU24, RNU44 and U6) out of the 19 endogenous controls analyzed, and these had inconsistent expression levels between CRC and healthy individuals (data not shown); (2) the data for the endogenous controls had relatively high standard deviations and the highest fold changes in expression levels observed in the RT-preamp-qPCR analysis.
Correlation of miR-378 normalization in the feces and tumor samples
The linear regression analysis showed that miR-378 had the lowest R 2 value with miR-144* in both healthy and CRC subjects (Fig. 3a), whereas miR-378 had a slightly higher correlation in CRC with miR-532-3p (Fig. 3b), where 60% of the variance was shared between miR-378 and miR-532-3p. In contrast, only 31% of the variance was shared in the healthy subjects. Then we analyzed whether both of these miRNAs were correlated with our selected normalizer. We found that miR-532-3p was positively correlated (R 2 = 0.53) (Fig. 3c), while miR-144* which had the lowest shared variance (R 2 = 0.15). These data show that the expression of miR-144* is independent of the level of miR-378 in the fecal sample, as only 18% (for the healthy subjects) and 31% (for CRC subjects) of the variance was shared (Fig. 3a). We then analyzed the relationship between miR-378 and miR-144* in tissue samples, and found that miR-144* also had the lowest shared variance, confirming what we observed in fecal specimens (Fig. 3d).
Regression plot analysis of miR-144* and miR-532-3p for biomarker validation, and miR-378 as a normalizer in feces of healthy and CRC subjects. a A nonsignificant positive correlation of miR-144* with miR-378 in healthy and CRC subjects was observed (CRC, R 2 = 0.1835 vs. healthy subjects, R 2 = 0.3115). b A significant positive correlation of miR-532-3p with miR-378 expression was found for the CRC group; R 2 was 0.59. c A significant correlation was not found between the expressions of miR-144* and miR-532-3p when analyzed together with miR-378. d Analysis of tissue specimens gave results similar to those obtained for miR-144* and miR-378 from fecal samples
miR-144* as a novel diagnostic marker for CRC detection
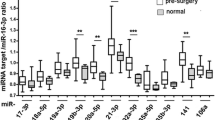
Finally, we evaluated the potential use of our selected miRNAs as a noninvasive fecal-based screening marker for CRC. Based on the C q values obtained, which were normalized to miR-378 [geometric mean expression of all patients (n = 75): 31.97 with SD of 1.75, Fig. 4] in a subset of fecal samples (n = 35 CRC and n = 40 healthy subjects, Table 2), we detected a significant alteration of ΔC q between these two groups, as shown in Fig. 5 (together with the box and whisker plot for miR-144*, Fig. 4). Based on this, we detected CRC with a sensitivity of 74.29% (95% CI 56.7–87.5) and a specificity of 87.20% (95% CI 72.6–95.70), AUC ROC of 0.829 (Fig. 5, left panel). Nevertheless, we could only detect 63% of the CRC cases at a specificity of 95%. Next, the cumulative fold change in miR-144* for the whole group showed an 11-fold upregulation for CRC compared to healthy individuals (Table 2). Taken together, the RT-preamp-qPCR microarray and RT-qPCR data led us to conclude that miR-144* is a novel candidate diagnostic marker for CRC screening. Apart from this, the elevated expression of miR-532-3p could not be used to distinguish CRC patients from healthy individuals, as shown in the box and whisker plot, p = 0.8676 (Figs. 4, 5, right panel). Moreover, the expression of miR-532-3p was positively correlated with that of miR-378 (Fig. 3b, c).
Box and whisker plot analysis of miR-144*, miR-532-3p and miR-378 expression in feces of healthy and CRC patients. The direct C q distribution of the normalizer miR-378 demonstrates that there was no significant difference between the expressions of miR-378 in diseased and healthy subjects (CRC \( \bar{x} \) C q expression: 31.92 ± 1.94 with; healthy subject \( \bar{x} \) C q expression: 32.02 ± 1.60), while the C q distribution of miR-144* clearly distinguishes between CRC and healthy subjects with p < 0.0001. However, the direct C q distribution of miR-532-3p indicates that there is no significant difference between the expression levels of miR-532-3p in healthy and CRC subjects (CRC \( \bar{x} \) C q expression: 33.27 ± 2.96 with; healthy subject \( \bar{x} \) C q expression: 33.34 ± 1.69), p = 0.8676
Receiver operating characteristic (ROC) curve analyses of miR-144* and miR-532-3p in the fecal samples of a subset of CRC and healthy subjects, as derived from ΔC q calculation. The left-hand figure shows the ROC analysis along with the corresponding area under the curve (AUC) for miR-144* expression. The ROC demonstrated a sensitivity of 74% and a specificity of 87%, AUC = 0.829. The right-hand figure shows the sensitivity and specificity of the miR-532-3p expression level when used to detect disease. Solid line represents the actual sensitivity and specificity of the test, while the dashed lines represent the 95% confidence interval
Upregulation of miR-144* expression in CRC tissues
In order to verify that the levels of miR-144* detected in feces are similar to those in tissue samples, we analyzed 15 CRC paired surgical surrounding tissues by RT-qPCR. As expected, a strong upregulation of miR-144* expression in CRC tissue samples (~4.99-fold) was observed compared to the expression in the surrounding tissues (Table 3). However, three patients showed downregulation of miR-144*. Nevertheless, this confirmed that the expression of miR-144* in fecal miRNA is in accord with its expression in tissue samples (Fig. 3a, d). Furthermore, this data strengthens our conclusion that miRNA represents a novel, noninvasive, fecal-based diagnostic marker for CRC screening.
Discussion
Although miRNAs can be detected in many body fluids and in various tissues [15–23], only three reports have been published so far on the presence of miRNAs in feces [32–34]. Thus, further investigations of the properties of miRNAs in feces are of the utmost importance. As we carried out our investigation, Ahmed et al. [32] demonstrated that the detection of specific miRNAs in fecal specimens clearly discriminated between sporadic human colon cancer and active ulcerative colitis in their patient cohort. Furthermore, Link et al. proposed the use of fecal miRNAs as novel biomarkers for CRC screening. They showed that miR-21 and miR-106a are highly overexpressed in adenomas and CRCs compared with individuals who do not have colorectal neoplasia [34]. On the other hand, Koga et al. [33] showed that the expressions of the miR-17-92 cluster and miR-135 were significantly higher in feces of CRC than healthy volunteers. Therefore, miRNA evaluation in feces is a promising and noninvasive approach to CRC screening, although further validation of pre-analytical sampling and miRNA stability are required. Thus, our analysis of the presence and stability of miRNAs and the identification of a putative candidate miRNA strengthens their conclusions. In this study, we mainly focused on determining miRNA stability in feces through methods such as the consistent detection of miRNAs using two different temperatures over time.
We have previously shown that long DNA arising from malignant colonocytes represents a fecal-based diagnostic marker of CRC [26, 27]. To further investigate the diagnostic power of fecal specimens as a noninvasive tool for CRC screening, we profiled feces for 648 miRNAs in a subset of CRC and healthy subjects. Based on our microarray data, we detected 39% of the miRNAs in the feces. To further analyze the stability of the miRNAs, we measured the expressions of endogenous miRNAs and exogenous cel-miR-238 in feces. Chim et al. [15] demonstrated that cell-free maternal and placental origin miRNAs were much more stable than exogenous miRNAs. Furthermore, Park et al. [17] reported that miRNAs could be detected in saliva samples and are much more stable than exogenous miRNAs, and a very slow degradation pattern was observed over time. Nevertheless, Tsui et al. [22] reported that bodily fluid miRNAs are more stable in their microenvironment. Taken together, our data confirm that miRNAs are also stable in the fecal microenvironment.
Next, to evaluate whether miRNAs are stable at different temperatures, we accounted for the fact that fecal mRNA and DNA can be degraded very quickly due to the activities of bacterial RNase and DNase [35]. The complex nature of feces inhibits certain assays like mRNA quantification and/or protein detection for the purposes of disease discrimination. The transportation and processing of feces is a drawback, because the samples need to be stored in fecal stabilization buffer and/or handled as quickly as possible. Furthermore, this degradation is common regardless of the processing method employed, as shown by the reduced sizes of high molecular weight ribosomal bands and the presence of low molecular weight RNA. We found that samples kept for up to 1 year at −20°C could be processed and quantified for miRNA detection (unpublished observation in our laboratory). Furthermore, we found that fecal miRNAs were much more stable at 4°C over a period of six days. Besides, Park et al. [17] showed that salivary miRNA could identify a diseased state in oral cancer detection. They reported that mature miRNAs are bound to a RISC complex, which showed that Ago 2 protein is also present in salivary specimens. This led us to hypothesize that miRNAs could also be associated with macromolecules present in feces, making them approximately as stable in feces as they are in other bodily fluids.
Based on this remarkable finding on the stability of miRNA in feces, we conducted RT-qPCR assays for a selected set of miRNA candidates for CRC diagnosis. Many miRNAs, such as miR-16 and miR-135b, showed upregulation, in accordance with the literature [36], but insignificant results were obtained when calculating the actual sensitivities of these markers in our preliminary investigation. We therefore selected miR-144* and miR-532-3p as markers for further evaluation. We also chose miR-378 as our quality control and as our reference miRNA for data normalization, because it showed a very low standard deviation in C q values in our pooled analysis. We found that miR-378 is consistently expressed across healthy and neoplastic individuals. Moreover, we observed that miR-144* was significantly overexpressed in CRC compared to normal controls. When we calculated the overall fold change between these two groups, we found that this miRNA is upregulated at least 11-fold, and is easily detected in fecal samples. To further strengthen these data, the expression pattern of miR-144* was also analyzed in paired surgical tissues. As expected, we found that the expression was fivefold higher in CRC tissues compared to their surrounding tissues. These data further demonstrate that the levels of miRNA detected in feces strongly reflect the changes occurring in corresponding tissues. However, we observed a significant difference in the fold changes in the fecal and tumor samples (11- vs. 5-fold). This can be explained by the expression analysis, as it was done on two different subpopulations in our pilot study.
Finally, when we calculated the sensitivity and specificity of this assay, we demonstrated that miR-144* represents a novel fecal-based screening marker for CRC. Our findings are in agreement with the results reported by Koga et al. [33], with a sensitivity and a specificity of 74.1 and 79.0%, respectively.
Recently, Ahmed et al. [32] and Link et al. [34] showed that the expressions of miR-21 and miR-106a are strongly upregulated in feces of CRC patients. Their studies were also supported by Schetter and colleagues [14], who observed increased expressions of miR-21 and miR-106a in CRC and adenoma associated tissue from patients with poor survival and therapeutic outcomes. Our data from pooled analysis also confirmed these miRNA increases in CRC, with a borderline fold change for miR-21 (1.8-fold) and a significant fold change in the expression of miR-106a (threefold). Moreover, Xie et al. [23] demonstrated that the expression of miR-21 is significantly higher in sputum samples from patients with non-small cell lung cancer than in those from cancer-free individuals.
Moreover, our chosen miRNA, miR-144*, has not been reported to be correlated with CRC incidence, but we found significant upregulation of miR-144* in both feces and tissue samples from CRC patients. We found that miR-144* may be involved in programmed cell death mechanisms, as this miRNA targets APAF1, a cytoplasmic protein that initiates apoptosis (http://www.microrna.org/), and other potential targets and pathways that are yet to be identified.
The results of our study suggest that further experimentation in this area is needed. The use of a pooled sample approach for candidate miRNA screening is economically very attractive, but our data could be biased due to individual sample variation. Thus, in order to validate these findings, additional experiments using another set of patients who are undergoing CRC screening should be studied to determine the robustness of this approach. Moreover, the overexpression of miR-144* could reflect its involvement in the tumorigenesis of CRC. Therefore, further experimental analysis is needed to rule this out and establish the actual function of miR-144*, in order to show that miR-144* is a viable fecal-based diagnostic marker. The selection of an miRNA as an internal control in our study could be biased due to the unknown function of the miRNA itself. This selection could also differ from laboratory to laboratory and from study to study. Furthermore, in our own experience, endogenous control small RNAs (i.e., RNU19, U6) are not consistently expressed in cells, tissues, or even in fecal samples. Therefore, a better choice for a universal internal control for miRNA is necessary. Finally, an early diagnostic marker should emphasize the presence of premalignant CRC lesions. Thus, the detection of miR-144* in adenoma samples would be ideal, and would add further weight to the idea of using miRNAs as a screening modality.
This proof of principle study has revealed the potential of fecal miRNAs as practical molecular markers for diverse physiological and pathological conditions, such as cancer. We have demonstrated the ease and reliability with which fecal miRNAs can be determined, which may pave the way to their wider application as a possible noninvasive tool for CRC detection. Thus, large randomized control studies are urgently required to validate the results found here, and to discover other specific miRNAs that could represent the new generation of biomarkers for rapid diagnosis.
Abbreviations
- miRNA:
-
MicroRNA
- fDNA:
-
Fecal DNA
- CRC:
-
Colorectal cancer
References
Cummins JM, He Y, Leary RJ, Pagliarini R, Diaz LA Jr, Sjoblom T, et al. The colorectal microRNAome. Proc Natl Acad Sci USA. 2006;103:3687–92.
Akao Y, Nakagawa Y, Naoe T. MicroRNA-143 and -145 in colon cancer. DNA Cell Biol. 2007;26:311–20.
Ventura A, Jacks T. MicroRNAs and cancer: short RNAs go a long way. Cell. 2009;136:586–91.
Aqeilan RI, Calin GA, Croce CM. miR-15a and miR-16-1 in cancer: discovery, function and future perspectives. Cell Death Differ. 2010;17:215–20.
Gartel AL, Kandel ES. miRNAs: little known mediators of oncogenesis. Semin Cancer Biol. 2008;18:103–10.
Fujita S, Iba H. Putative promoter regions of miRNA genes involved in evolutionarily conserved regulatory systems among vertebrates. Bioinformatics. 2008;24:303–8.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Koturbash I, Zemp FJ, Pogribny I, Kovalchuk O. Small molecules with big effects: the role of the microRNAome in cancer and carcinogenesis. Mutat Res. 2010;722:94–105.
Kloosterman WP, Plasterk RH. The diverse functions of microRNAs in animal development and disease. Dev Cell. 2006;11:441–50.
Stefani G, Slack FJ. Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol. 2008;9:219–30.
Lu J, Qian J, Chen F, Tang X, Li C, Cardoso WV. Differential expression of components of the microRNA machinery during mouse organogenesis. Biochem Biophys Res Commun. 2005;334:319–23.
Ng EK, Chong WW, Jin H, Lam EK, Shin VY, Yu J, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut. 2009;58:1375–81.
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE, et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med. 2005;353:1793–801.
Schetter AJ, Leung SY, Sohn JJ, Zanetti KA, Bowman ED, Yanaihara N, et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA. 2008;299:425–36.
Chim SS, Shing TK, Hung EC, Leung TY, Lau TK, Chiu RW, et al. Detection and characterization of placental microRNAs in maternal plasma. Clin Chem. 2008;54:482–90.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105:10513–8.
Park NJ, Zhou H, Elashoff D, Henson BS, Kastratovic DA, Abemayor E, et al. Salivary microRNA: discovery, characterization, and clinical utility for oral cancer detection. Clin Cancer Res. 2009;15:5473–7.
Li J, Smyth P, Flavin R, Cahill S, Denning K, Aherne S, et al. Comparison of miRNA expression patterns using total RNA extracted from matched samples of formalin-fixed paraffin-embedded (FFPE) cells and snap frozen cells. BMC Biotechnol. 2007;7:36.
Nelson PT, Baldwin DA, Kloosterman WP, Kauppinen S, Plasterk RH, Mourelatos Z. RAKE and LNA-ISH reveal microRNA expression and localization in archival human brain. RNA. 2006;12:187–91.
Xi Y, Nakajima G, Gavin E, Morris CG, Kudo K, Hayashi K, et al. Systematic analysis of microRNA expression of RNA extracted from fresh frozen and formalin-fixed paraffin-embedded samples. RNA. 2007;13:1668–74.
El-Hefnawy T, Raja S, Kelly L, Bigbee WL, Kirkwood JM, Luketich JD, et al. Characterization of amplifiable, circulating RNA in plasma and its potential as a tool for cancer diagnostics. Clin Chem. 2004;50:564–73.
Tsui NB, Ng EK, Lo YM. Stability of endogenous and added RNA in blood specimens, serum, and plasma. Clin Chem. 2002;48:1647–53.
Xie Y, Todd NW, Liu Z, Zhan M, Fang H, Peng H, et al. Altered miRNA expression in sputum for diagnosis of non-small cell lung cancer. Lung Cancer. 2009;67:170–6.
Ahlquist DA, Skoletsky JE, Boynton KA, Harrington JJ, Mahoney DW, Pierceall WE, et al. Colorectal cancer screening by detection of altered human DNA in stool: feasibility of a multitarget assay panel. Gastroenterology. 2000;119:1219–27.
Dong SM, Traverso G, Johnson C, Geng L, Favis R, Boynton K, et al. Detecting colorectal cancer in stool with the use of multiple genetic targets. J Natl Cancer Inst. 2001;93:858–65.
Kalimutho M, Del Vecchio Blanco G, Gravina P, Cretella M, Mannucci L, Mannisi E, et al. Quantitative denaturing high performance liquid chromatography (Q-dHPLC) detection of APC long DNA in faeces from patients with colorectal cancer. Clin Chem Lab Med. 2010;48:1303–11.
Kalimutho M, Del Vecchio Blanco G, Cretella M, Mannisi E, Sileri P, Formosa A, et al. A simplified, non-invasive fecal-based DNA integrity assay and iFOBT for colorectal cancer detection. Int J Colorectal Dis. 2011;26:583–92.
Taylor CF, Field D, Sansone SA, Aerts J, Apweiler R, Ashburner M, et al. Promoting coherent minimum reporting guidelines for biological and biomedical investigations: the MIBBI project. Nat Biotechnol. 2008;26:889–96.
Mestdagh P, Feys T, Bernard N, Guenther S, Chen C, Speleman F, et al. High-throughput stem-loop RT-qPCR miRNA expression profiling using minute amounts of input RNA. Nucleic Acids Res. 2008;36:e143.
Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, et al. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 2009;10:R64.
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3:RESEARCH0034.
Ahmed FE, Jeffries CD, Vos PW, Flake G, Nuovo GJ, Sinar DR, et al. Diagnostic microRNA markers for screening sporadic human colon cancer and active ulcerative colitis in stool and tissue. Cancer Genomics Proteomics. 2009;6:281–95.
Koga Y, Yasunaga M, Takahashi A, Kuroda J, Moriya Y, Akasu T, et al. MicroRNA expression profiling of exfoliated colonocytes isolated from feces for colorectal cancer screening. Cancer Prev Res (Phila). 2010;3:1435–42.
Link A, Balaguer F, Shen Y, Nagasaka T, Lozano JJ, Boland CR, et al. Fecal MicroRNAs as novel biomarkers for colon cancer screening. Cancer Epidemiol Biomarkers Prev. 2010;19:1766–74.
Yu YJ, Majumdar AP, Nechvatal JM, Ram JL, Basson MD, Heilbrun LK, et al. Exfoliated cells in stool: a source for reverse transcription-PCR-based analysis of biomarkers of gastrointestinal cancer. Cancer Epidemiol Biomarkers Prev. 2008;17:455–8.
Bartels CL, Tsongalis GJ. MicroRNAs: novel biomarkers for human cancer. Clin Chem. 2009;55:623–31.
Acknowledgments
This work was supported by grants from University Hospital Tor Vergata, University of Rome. MK was supported by a pre-doctoral scholarship for foreign students under the International Italian Government University scholarship.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Kalimutho, M., Del Vecchio Blanco, G., Di Cecilia, S. et al. Differential expression of miR-144* as a novel fecal-based diagnostic marker for colorectal cancer. J Gastroenterol 46, 1391–1402 (2011). https://doi.org/10.1007/s00535-011-0456-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-011-0456-0