Abstract
Over the past decade, microRNAs (miRNAs) have become a new paradigm of gene regulation. miRNAs are involved in a wide array of carcinogenic processes. Indeed, increasing evidence has shown the importance of miRNAs in cancer, suggesting their possible use as diagnostic, predictive and prognostic biomarkers, leading to miRNA-based anti-cancer therapies, either alone or in combination with current targeted therapies, with the goal of improving cancer treatment responses and increasing cure rates. The advantage of using a miRNA approach is based on the ability to concurrently target multiple effectors of pathways involved in cell proliferation, migration and survival. This review sheds new light on miRNA regulation of genes that play critical roles in the process of malignant transformation and tumour metastasis, the dysregulation of miRNA expression in cancer development and the development of miRNA-based diagnostics and therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer is the most frequently diagnosed cancer and is the leading cause of cancer death for women worldwide, with 232,340 new cases annually [1]. Although improvements in early detection and treatment have decreased breast cancer mortality rates in recent years, breast cancer prevention and therapy remain to be a major public health concern. Conventional wisdom in the field of oncology indicates that malignant tumour progression involves multiple genetic changes, each of which could result in the dysregulation of important pathways involved in complex cellular processes, such as growth, invasion, and apoptosis. The accumulation of these genetic alterations confers a malignant phenotype. Recently, microRNAs (miRNAs) have received wide attention as crucial regulators of gene expression.
miRNAs are a class of endogenous, small, non-coding RNAs (~22 nt) that negatively regulate protein-coding messenger RNAs (mRNAs) at the posttranscriptional level. Most miRNA genes are derived from primary miRNA transcripts containing a cap and a poly(A) tail and are produced by RNA polymerase II from miRNA genes. The primary miRNAs are further cleaved into 22-nt mature miRNAs by the consecutive function of RNAse III Drosha–DGCR8 and Dicer, present in the nucleus and cytoplasm, respectively. These single-stranded molecules have a length of 19–23 nucleotides and regulate gene expression at the posttranscriptional level, thus controlling crucial physiological processes through three mechanisms: binding to complementary sequences on target messenger RNA transcripts, repressing mRNA translation and cleaving target mRNA. Recently, a study by Mukherji et al. [2] showed that miRNA target genes have an mRNA expression threshold, below which, the gene is efficiently repressed, and above which, it can overwhelm the available miRNA. The threshold level is determined by the available miRNA concentration, whereas the steepness of the transition is determined by the strength and number of miRNA binding sites in the target. However, Dvinge et al. observed that miRNAs act as modulators of mRNA–mRNA interactions rather than as on–off molecular switches in the biology of copy-number aberration (CNA devoid) in breast cancer [3] (Fig. 1).
miRNA expression is altered in cancer cells, with some miRNAs functioning as tumour oncogenes (oncomiRs) and others acting as suppressors, depending on which genes or pathways they regulate. miRNAs are also known to be involved in various biological processes, including cellular proliferation, tumorigenesis and migration, which is indicative of their importance in cellular function [3-5]. This review focuses on recent progress in the understanding of breast cancer regulation by miRNAs. These findings confirm and extend the importance of miRNAs in pathophysiological process and illustrate the value of analysing neoplastic cells for new leads that may improve breast cancer outcomes (Table 1).
ApoptomiRs and microRNA regulation of proliferation
Recent studies have demonstrated that microRNAs can impair cell proliferation or induce apoptosis through oncogene/tumour suppressor targeting. miR-497, a tumour suppressor, can specifically regulate cell growth by targeting cyclin E1 in MDA-MB-231 cells. Furthermore, its overexpression causes a G1 cell cycle arrest [6]. Cui et al. demonstrated that miR-133a, which may act as a tumour suppressor in breast cancer, regulates cell cycle and proliferation in carcinogenesis by targeting epidermal growth factor receptor (EGFR) through the downstream signal molecule Akt [7]. Oestrogen-dependent high levels of miR-191/425 induce proliferation in estrogen receptor alpha (ERα)-positive cells by repressing a strong tumour suppressor gene, such as early growth response protein 1 (EGR1). However, low levels of the miR-191/425 cluster are essential for the high expression of important modulators, such as FSCN1, CCND2 and SATB1, which confer a proliferative advantage to aggressive breast cancer cells [8]. Leivonen et al. reported that miR-342-5p specifically inhibits the growth of human epidermal growth factor receptor 2 (HER2)-positive cells in vitro [9]. Research by Nassirpour et al. demonstrated that miR-221 is specifically overexpressed in triple-negative breast cancer cells and that miR-221 knockdown induces G1 arrest and apoptosis and inhibits cell proliferation and tumour growth, most likely by altering expression levels of p27kip [10]. Interestingly, miR-30c overexpression suppresses the proliferation of hereditary breast cancer cells through inhibition of KARS signalling [11].
Apoptosis is a crucial cellular process that normally serves to maintain cell proliferative homeostasis and prevent unrestricted cell division. A recent report revealed that miR-31 directly targets PRKCE and thereby suppresses nuclear factor kappa B (NF-κB) activity and induces apoptosis and sensitivity to anti-cancer treatment via the downregulation of BCL2 expression in MCF10A breast epithelial and MDA-MB-231 triple-negative breast cancer cells [12]. Anaya-Ruiz et al. found that silencing of miR-153 significantly induces apoptosis in the MDA-MB-231 breast cancer cell line [13]. More recently, miR-26a was shown to be downregulated in breast cancer cells, and its modulation regulated cell proliferation and apoptosis. Myeloid cell leukaemia protein 1 (MCL-1), an anti-apoptotic member of the Bcl-2 family, novel targets of miR-26a, was found to inversely correlate with an ectopic expression of miR-26a, and MCL-1 knockdown phenocopied the effect of miR-26a in breast cancer cell lines [14].
miRNA regulation of metastasis and invasiveness
Metastasis is a multistep process that begins as primary tumour cells invade surrounding tissues and enter the blood and lymphatic vessels. The cells travel through the vasculature, extravagate into distant tissues and finally establish secondary tumours. Substantial direct evidence has demonstrated the links between miRNAs and metastasis. Epithelial–mesenchymal transition (EMT), a key step of metastasis, is an evolutionarily conserved process in which epithelial cells are converted to mesenchymal cells. Adiponectin receptor 1 (ADIPOR1), a direct target of miR-221/222, negatively regulates EMT in breast cancer and provides an additional mode by which miR-221/222 induces EMT [15]. Arora et al. [16] observed that NF-κB binds to the upstream promoter region of miR-506 to suppress transcription. Moreover, miR-506 overexpression inhibits TGFβ-induced EMT signalling in breast cancer cell lines. Certain miRNAs negatively regulate the metastatic potential of cancer cells. For example, Tavazoie et al. [17] reported that restoring the expression of miRNAs in malignant cells, whose expression is specifically lost as human breast cancer cells develop metastatic potential, suppresses lung and bone metastasis in metastatic breast cancer. miR-126 restoration reduced overall tumour proliferation, whereas miR-335 suppressed metastasis. An elegant study by Zhang et al. [18] demonstrated the role of miR-30a in repressing breast cancer migration and invasion, as mediated by metadherin (MTDH), an effective oncogene in metastasis. In addition, miR-30a expression is inversely associated with lymph node or lung metastasis in breast cancer patients, supporting the notion that a loss of miR-30a results in increased expression of the oncogene MTDH, which in turn promotes breast cancer metastasis. Li et al. [19] observed that miR-720 expression is significantly downregulated in primary breast cancer, with greater downregulation in metastatic tumours. Moreover, re-expression of miR-720 in breast cancer cells remarkably suppressed cell invasiveness and migration in vitro and in vivo via direct targeting of Twist-related protein 1 (TWIST1), a highly conserved basic helix-loop-helix transcription factor. It was found that miR-19a-3p, which regulates tumour-associated macrophages (TAMs) in the breast tumour microenvironment, regulates the TAM phenotype by targeting the Fra-1 gene. The downregulation of miR-19a-3p expression in TAMs was likely due to TEM induction, which promotes transformation of M1 to M2 and results in enhanced migration and invasion of breast cancer cells [20]. Additionally, miRNAs are also involved in positively regulating EMT and tumour metastasis. In breast cancer cell lines, EMT and metastasis were promoted by an ectopic overexpression of miR-374a both in vitro and in vivo. Furthermore, miR-374a directly targeted and suppressed multiple negative regulators of the Wnt/β-catenin signalling cascade, including Wnt inhibitory factor 1 (WIF1), phosphatase and tensin homologue (PTEN) and WNT5A [21]. Taylor et al. [22] demonstrated that aberrantly high miR-181a expression enhanced the ability of TGFβ to stimulate breast cancer metastasis via inducing EMT programmes and by promoting resistance to anoikis by downregulating the expression of the pro-apoptotic factor Bim. Moreover, miR-181a expression was highly associated with the development of metastatic disease in triple-negative breast cancers (TNBCs). It has been reported that miR-200a, functioning as an anoikis suppressor, promotes anoikis resistance by targeting Yes-associated protein 1 (YAP1), subsequently leading to distant metastasis in breast cancer [23]. miR-182 promotes tumour cell invasion and metastasis by inhibiting missing in metastasis (MIM) in breast cancer. In addition, MIM represses stress fibre formation and invasion of cancer cells through inactivation of the cytoskeleton regulator RhoA [24]. Moreover, recent reports on the role of miRNAs in breast cancer with brain metastasis have attracted increasing attention. A study by Okuda et al. [25] showed that miR-7 expression significantly suppresses the ability of cancer stem-like cells (CSCs) to metastasise to the brain in an animal model. The authors also found that miR-7 and Kruppel-like factor 4 (KLF4) are significantly down- or upregulated, respectively, in brain-metastatic lesions. Furthermore, the results of experiments in vitro indicate that miR-7 attenuates CSC invasion and self-renewal by modulating KLF4 expression.
Indeed, it is also important to understand the interplay between miRNAs and mRNAs leading to breast cancer. Recently, several studies found that mRNAs regulate tumour progression through targeting miRNAs. For example, Hu et al. [26] reported that elevated BMP-6 expression inhibits G1/S cell cycle progression in MDA-MB-231 cells, resulting in impaired tumorigenesis in a nude mouse xenograft model via the downregulation of miR-192. They also found that the cell cycle regulator, RB1, is a major target of BMP-6/miR-192 signalling in the regulation of MDA-MB-231 cell proliferation. Recently, Chou et al. [27] found that GATA3 promoted differentiation, suppressed metastasis and altered the tumour microenvironment in breast cancer by inducing microRNA-29b expression. Accordingly, loss of miR-29b, even in GATA3-expressing cells, increases metastasis and promotes a mesenchymal phenotype.
miRNA regulation of angiogenesis
Tumour angiogenesis is a crucial process resulting in new blood vessel formation, which plays an important role in tumour growth, invasion and metastasis. Generally, solid tumours growing beyond 1–2 mm3 in diameter require the formation of new vessels to remove metabolic waste and transport nutrients and oxygen, a critical process that allows small developing neoplasias to enter a state of uncontrolled proliferation. Pathways that control the complex processes have been the focus of efforts to use novel therapeutics that inhibit this process. Many of the previously described miRNAs have been found to be vital regulators of angiogenesis.
Zhu et al. [28] revealed that endothelial-specific miR-126 is excised from EGFL7 pre-mRNA without affecting splicing and expression of its host gene. In addition, downregulation of miR-126 in tumours may increase the activity of the VEGF/PI3K/AKT signalling pathway and induce vascular formation. Preliminary work by Siragam et al. [29] investigated the involvement of miR-98 in regulating angiogenesis using a highly aggressive breast cancer model in vitro and in vivo. The authors found that miR-98 functions as a tumour suppressor by inhibiting blood vessel expansion, primarily by targeting matrix metalloproteinase 11 (MMP11) and activin receptor-like kinase 4 (ALK4). Zou et al. [30] showed that miR-145 suppresses tumour angiogenesis through the posttranscriptional regulation of the novel targets N-RAS and VEGF-A in vitro and in vivo. Kong et al. [31] provided in vitro and vivo evidence that miR-155 upregulation promotes breast cancer angiogenesis primarily via targeting Von Hippel–Lindau (VHL). The overexpression of either miR-148a or miR-152 in breast cancer cells is sufficient to inhibit tumour angiogenesis through targeting insulin-like growth factor 1 receptor (IGF1R) and IRS1 and suppressing their downstream AKT and mitogen-activated protein kinase (MAPK)/ERK signalling pathways [32]. It is well known that angiopoietin-2 plays a critical role in angiogenesis. A recent study showed that miR-542-3p represses tumour angiogenesis by targeting angiopoietin-2, a key angiogenesis-promoting protein [33].
miRNA polymorphisms associated with breast cancer
Breast cancer is a consequence of multiple factors, including genetic predisposition. Specific variations in DNA sequence [single nucleotide polymorphisms (SNPs)] of coding genes as well as of non-coding miRNAs have been linked to breast cancer susceptibility. These polymorphisms exist not only in the mature miRNA sequence but also in the pre- and pri-forms of the miRNA, potentially altering the processing of the mature sequence and their binding sites in target genes.
Polymorphisms within miRNAs
Recent years have witnessed an explosion of reports of SNPs within miRNAs. For instance, the pre-mir-27a rs895819 polymorphism was shown to have a protective effect against breast cancer, mainly in those younger than age 50 [34]. However, a further meta-analysis showed that this polymorphism might have some relation to breast cancer susceptibility or cancer development in Caucasians [35]. Lian et al. found that the has-miR-146a rs2910164 polymorphism is associated with increased breast cancer risk among Europeans in a homozygote comparison (CC vs. GG: odds ratio (OR) = 1.29, 95 % confidence interval (CI) = 1.02 − 1.63) [36]. Fan et al. reported that the miR-499 rs3746444 T > C polymorphism is associated with breast cancer, and the C allele could increase cancer susceptibility in Asians [37]. However, these results have been disputed by additional studies that found no significant association between rs2910164 in miR-146a (or rs3746444 in miR-499) and breast cancer susceptibility. A recent meta-analysis demonstrated that the rs11614913 CC genotype of miR-196a2 was revealed to be associated with an increased breast cancer susceptibility compared with the TT + CT genotypes [38-40].
In a case–control study of 1,064 breast cancer cases and 1,073 cancer-free controls, Chen et al. revealed that rs462480 and rs1053872 in the flank regions of pre-miR-101-2 are significantly associated with an increased risk of breast cancer (rs462480 AC/CC vs. AA: adjusted OR = 1.182, 95 % CI = 1.030–1.357, p = 0.017; rs1053872CG/GG vs. CC: adjusted OR = 1.179, 95 % CI = 1.040–1.337, p = 0.010) [41].
Polymorphisms within miRNA target binding sites
miRNAs have been identified as translational factors of protein-coding genes through binding target sites in the 3′ UTRs of mRNAs. Several lines of evidence indicate that genetic variants in the sequence of target sites may affect the miRNA regulation of target gene expression and consequently influence cancer development. For instance, in a hospital-based, case–control study of sporadic breast cancer, the variant rs16430 0 bp at the miRNA binding site in thymidylate synthase (TYMS) gene was associated with a significantly increased risk of breast cancer in non-Hispanic white women aged ≤55 years [42]. The variant rs2867428 in the miR-515-5p binding site may modify breast cancer risk among BRCA1 carriers by regulating the expression of IGF-1R. miR-515-5p was downregulated in tumour tissue compared with non-neoplastic, surrounding tissue, while IGF-1R levels were elevated. This IGF-1R SNP was found to be significantly associated with age at diagnosis of breast cancer in BRCA1 mutation carriers [43]. In a two-stage case–control study, Jiang et al. reported that the variant rs7963551 in the hsa-let-7 binding site may modify breast cancer risk through regulating RAD52 expression [44]. In a study from China, researchers found that DGCR8 rs417309G > A may increase the risk of breast cancer through interrupting miRNA-106b or miRNA-579 binding affinity [45].
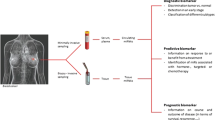
MicroRNAs as diagnostic biomarkers
Because miRNAs have been shown to be stable molecules in body fluids such as blood and cerebrospinal fluids, they are potential biomarkers for non-invasive early detection of breast cancer. Si et al. reported that decreased levels of miR-92a in serum samples of BC are associated with tumour size and a positive lymph node status (p < 0.001) [46]. By comparing miRNA profiles between tumour and serum samples, miR-1, miR-92a, miR-133a and miR-133b were identified as the most important diagnostic markers, and the receiver operating characteristic curves derived from combinations of these miRNAs exhibited areas under the curves of 0.90 to 0.91 [47]. Cuk et al. [48] found that miR-127-3p, miR-148b, miR-376a, miR-376c, miR-409-3p, miR-652 and miR-801 levels are elevated in the plasma of breast cancer patients. These miRNAs could differentiate even women with benign tumours, stage I or II breast cancer from healthy controls. In addition, a panel of these seven circulating miRNAs had a substantial diagnostic potential with an AUC of 0.81 for the detection of benign and malignant breast tumours, which further increased to 0.86 in younger women (up to 50 years of age). Using the product of miR-10b and miR-373 as a combined biomarker, Chen et al. observed that their levels could distinguish breast cancer patients with lymph node metastasis from non-metastatic patients [49]. Kumar et al. showed that circulating levels of miR-21 and miR-146a are significantly higher in plasma samples of breast cancer patients compared with those of healthy controls (p < 0.0004 and p < 0.005, respectively) [50]. Zeng et al. [51] observed that the status of ER and triple-negative breast cancer is significantly associated with miRNA-30a levels (p = 0.007 and p = 0.005, respectively). Moreover, ROC analysis showed the sensitivity and specificity of miRNA-30a for breast cancer diagnosis at 74.0 and 65.6 %, respectively, suggesting that miRNA-30a has great potential for use as a novel biomarker for breast cancer diagnosis. One study reported that the combination of miR-145 and miR-451 could discriminate breast cancer from healthy controls and all other types of cancers; testing with these markers reached a sensitivity and specificity of >90 % for breast cancer prediction, the positive predictive value was approximately 90 %, and the negative predictive value was approximately 92 % [52].
MicroRNAs as prognostic and predictive biomarkers
Identification of patients with a worse prognosis may improve the clinical care of those patients by optimising therapy. The association between miRNA expression signature and prediction of prognosis and survival has recently been evaluated in several studies. Markou et al. [53] revealed that both miR-21 and miR-205 are significantly associated with disease-free interval and only miR-205 with overall survival. Moreover, miR-205 and miR-21 are independent factors associated with early disease relapse, whereas only miR-205 overexpression is associated with overall survival (OS). Leivonen et al. demonstrated that a higher expression of miR-342-5p, a tumour suppressor, is associated with better survival in both breast cancer patient cohorts [9]. A study by Wang et al. [54] found that low miR-497 expression closely correlates with higher differentiation grade, positive HER2 expression, higher incidence of lymph node metastasis and advanced clinical stage. Furthermore, low miR-497 expression correlates with shorter OS and disease-free survival (DFS) in breast cancer. miR-221 was considered to be a highly significant prognostic marker for distinguishing prognostic subgroups particularly in advanced (lymph node+, HER2+) breast cancers, and miR-222 was of prognostic significance in the lymph node-negative tumours and therefore may be of high impact in differentiating between different LN-negative prognostic groups [55]. Tang et al. revealed that reduced miR-1258 expression is associated with a short overall survival and a short relapse-free survival, lymph node status, and late clinical stage by regulating HPSE [56]. Hoppe et al. demonstrated that a higher expression of miR-126 and miR-10a in early breast cancer patients treated with tamoxifen is associated with longer relapse-free time (log-rank p = 0.037 and p < 0.0001, respectively) [57]. A recent study showed that let-7b expression in breast cancer patients is inversely associated with tumour lymph node metastasis (p = 0.001), patient overall survival (p = 0.027), relapse-free survival (p = 0.016) and basigin (BSG) protein expression (p = 0.001). Furthermore, breast cancer patients with low let-7b expression have poor prognoses, indicating that let-7b might act as cancer suppressor gene in BC development and progression by inhibiting BSG expression [58]. Recently, several investigations have emphasised the integrated analysis of miRNAs and mRNAs in clinical outcomes. For example, Volinia et al. [59] performed survival analysis on 466 breast cancer patients by integrating mRNA, miRNA and DNA methylation data from TCGA. The authors found 7 miRNAs and 30 mRNAs that formed an integrated miRNA/mRNA signature with the highest prognostic value in stages 1 and 2 breast cancers. The prognostic RNAs included PIK3CA, one of the two most frequently mutated genes in IDC, and miRNAs such as has-miR-328, has-miR-484 and has-miR-874.
MicroRNAs as therapeutic targets
miRNA regulation of multiple cancer-related pathways provides numerous opportunities for therapeutic intervention. Targeting one miRNA might affect simultaneously a number of vital signalling pathways in the tumour cell. Therefore, it may be expected that miRNA targeting may be less prone to resistance than signal pathway-specific agents. Here, we present some miRNAs that might be useful in breast cancer therapy (Table 2).
Chemotherapy
miRNA-30c, a human breast tumour prognostic marker, is transcriptionally regulated by GATA3 in breast tumours Moreover, miR-30c regulated breast cancer chemoresistance and EMT by direct targeting of the cytoskeleton gene TWF1 as well as by indirect targeting of the cytokine IL-11 [60]. Fang et al. [61] showed that miR-30c restoration sensitises MCF-7/ADR to doxorubicin treatment in vivo and in vitro. Furthermore, the authors document miR-30c as a potential tumour suppressor of the progression to doxorubicin resistance in breast cancer through its ability to target YWHAZ and the p38 MAPK signalling pathway. One study showed that crosstalk between Bmi1 and miR-200c, mediated by p53, and Bmi1 interference would improve chemotherapy response to 5-fluorouracil in breast cancer via susceptive apoptosis induction and cancer stem cell enrichment inhibition [62]. In addition, Chen et al. [63] reported that miRNA-200c increased the drug sensitivity of breast cancer cells to doxorubicin by inactivating the Akt pathway through the E-cadherin-mediated upregulation of PTEN. Zhong et al. identified two miRNAs (miR-222 and miR-29a) involved in acquiring resistance to adriamycin and docetaxel, at least in part via targeting PTEN in breast cancer MCF-7 cells [64]. By using microarray analysis, Jiao et al. [65] found differential expression patterns of miRNAs that targeted breast cancer resistance protein (BCRP) between the parental MCF-7 and its derivative BCRP-overexpressing MCF-7/MX cells. miR-181a is the most significantly downregulated miRNA in MX-resistant MCF-7/MX cells. Importantly, the study revealed that miR-181a, functioning as a BCRP suppressor, reversed BCRP-mediated drug resistance in vitro and in vivo. A study by Hu et al. found that the demethylation of miR-663 promoter mediated miR-663 expression, and miR-663 targeted heparan sulfate proteoglycan 2 (HSPG2) to induce the chemotherapeutic resistance of breast cancer cells [66].
Endocrine therapies
Endocrine therapy has become a significant treatment option for women with ERα-positive breast cancer, with up to 70 % of primary breast cancers expressing ERα. Shibahara et al. found that aromatase inhibitors (AIs) may exert tumour-suppressing effects upon breast cancer cells by repressing aromatase gene expression via restoration of let-7f [67]. Re-expression of microRNA-375 not only sensitises cells to tamoxifen but also reverses EMT-like properties in a tamoxifen-resistant breast cancer model. Moreover, it modulates invasive potential by the direct regulation of the MTDH oncogene [68]. He et al. found that miR-342 expression positively correlates with the expression of ERα mRNA in human breast cancer tissues and that the introduction of miR-342 into oestrogen-dependent breast cancer cells enhances tamoxifen sensitivity [69].
Radiotherapy
miR-200c overexpression may enhance radiosensitivity by inhibiting cell proliferation and by increasing apoptosis and DNA double-strand breaks in breast cancer cells, especially in MDA-MB-231 cells. Additionally, one study revealed that an overexpression of Tank-binding kinase 1 (TBK1), a new miR-200c target, partially rescued miR-200c-mediated apoptosis induced by ionising radiation [70]. miR-34a was upregulated after X-ray irradiation in p53-positive normal and breast cancer cell lines. Moreover, miR-34a might be involved in breast cell responses to low-dose X-ray DNA damage [71].
Targeted therapy
However, in addition to the above therapies, miRNAs can also impact responsiveness to targeted therapies. miR-21 has already been implicated in resistance to trastuzumab (targeting the ERBB2 oncogene) through the downregulation of its protein target PTEN [72]. miR-200c, which is the most significantly downregulated miRNA in trastuzumab-resistant cells, and miR-21 restored trastuzumab sensitivity of breast cancer cells by targeting ZNF217, a transcriptional activator of TGFβ. It has also been observed that ZNF217 exerts a feedback suppression of miR-200c via TGFβ/ZEB1 signalling. Restoration of miR-200c, silencing of ZNF217 or blockade of TGFβ signalling increased trastuzumab sensitivity of breast cancer cells [73]. In addition, miR-221 promotes trastuzumab resistance in HER2-positive breast cancers by regulating PTEN [74]. Moreover, miR-210 appears to be increased in trastuzumab-resistant cell lines and patients [75]. Recently, Ye et al. observed that epigenetic silencing of miR-375 upregulated IGF-1R, which at least partially underlies trastuzumab resistance of HER2-positive breast cancer [76].
Indeed, a profound understanding of miRNAs’ role in oncology is warranted to allow their clinical investigations.
Conclusions
The past decade has witnessed an explosion of research on microRNAs and their functionality. Researchers have identified many miRNA signatures that are clearly associated with diagnosis, prognosis and therapeutic response. Many studies are also illuminating the connections between miRNAs and the progression of carcinogenesis and the basic biological roles that they play in tumour cell survival, invasion and metastasis. There are many issues that need to be addressed, however, before miRNAs can be effectively integrated into the field of clinical oncology. For instance, further evaluation of miRNAs as non-invasive serum markers is required before they can improve upon current methods for cancer diagnosis, such as target validation, the accurate prevention of unwanted off-target effects and the development of efficient methods of a specific drug delivery. Nevertheless, the results obtained to date show great promise.
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2013. CA. 2013;63:11–30.
Mukherji S, Ebert MS, Zheng GX, Tsang JS, Sharp PA, van Oudenaarden A. MicroRNAs can generate thresholds in target gene expression. Nat Genet. 2011;43:854–9.
Dvinge H, Git A, Gräf S, Salmon-Divon M, Curtis C, Sottoriva A, et al. The shaping and functional consequences of the microRNA landscape in breast cancer. Nature. 2013;497:378–82.
Shen J, Xia W, Khotskaya YB, Huo L, Nakanishi K, Lim S-O, et al. EGFR modulates microRNA maturation in response to hypoxia through phosphorylation of AGO2. Nature. 2013;497:383–7.
Song SJ, Poliseno L, Song MS, Ala U, Webster K, Ng C, et al. MicroRNA-antagonism regulates breast cancer stemness and metastasis via TET-family-dependent chromatin remodeling. Cell. 2013;154:311–24.
Luo Q, Li X, Gao Y, Long Y, Chen L, Huang Y, et al. MiRNA-497 regulates cell growth and invasion by targeting cyclin E1 in breast cancer. Cancer Cell Int. 2013;13:95.
Cui W, Zhang S, Shan C, Zhou L, Zhou Z. MicroRNA-133a regulates the cell cycle and proliferation of breast cancer cells by targeting epidermal growth factor receptor through the EGFR/Akt signaling pathway. FEBS J. 2013;280:3962–74.
Di Leva G, Piovan C, Gasparini P, Ngankeu A, Taccioli C, Briskin D, et al. Estrogen mediated-activation of miR-191/425 cluster modulates tumorigenicity of breast cancer cells depending on estrogen receptor status. PLoS Genet. 2013;9:e1003311.
Leivonen S-K, Sahlberg KK, Mäkelä R, Due EU, Kallioniemi O, Børresen-Dale A-L, et al. High-throughput screens identify microRNAs essential for HER2 positive breast cancer cell growth. Mol Oncol. 2014;8:93–104.
Nassirpour R, Mehta PP, Baxi SM. Yin M-J: miR-221 promotes tumorigenesis in human triple negative breast cancer cells. PLoS One. 2013;8:e62170.
Tanic M, Yanowsky K, Rodriguez-Antona C, Andrés R, Márquez-Rodas I, Osorio A, et al. Deregulated miRNAs in hereditary breast cancer revealed a role for miR-30c in regulating KRAS oncogene. PLoS One. 2012;7:e38847.
Körner C, Keklikoglou I, Bender C, Wörner A, Münstermann E, Wiemann S. MicroRNA-31 sensitizes human breast cells to apoptosis by direct targeting of protein kinase C epsilon (PKCepsilon). J Biol Chem. 2013;288:8750–61.
Anaya-Ruiz M, Cebada J, Delgado-López G, Luisa M. miR-153 silencing induces apoptosis in the MDA-MB-231 breast cancer cell line. Asian Pac J Cancer Prev. 2013;14:2983–6.
Gao J, Li L, Wu M, Liu M, Xie X, Guo J, et al. MiR-26a inhibits proliferation and migration of breast cancer through repression of MCL-1. PLoS One. 2013;8:e65138.
Hwang MS, Yu N, Stinson SY, Yue P, Newman RJ, Allan BB, et al. miR-221/222 targets adiponectin receptor 1 to promote the epithelial-to-mesenchymal transition in breast cancer. PLoS One. 2013;8:e66502.
Arora H, Qureshi R, Park W-Y. miR-506 regulates epithelial mesenchymal transition in breast cancer cell lines. PLoS One. 2013;8:e64273.
Tavazoie SF, Alarcón C, Oskarsson T, Padua D, Wang Q, Bos PD, et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature. 2008;451:147–52.
Zhang N, Wang X, Huo Q, Sun M, Cai C, Liu Z, Hu G, Yang Q: MicroRNA-30a suppresses breast tumor growth and metastasis by targeting metadherin. Oncogene 2013
Li L-Z, Zhang CZ, Liu L-L, Yi C, Lu S-X, Zhou X, et al. Yun J-P: miR-720 inhibits tumor invasion and migration in breast cancer by targeting TWIST1. Carcinogenesis. 2014;35:469–78.
Yang J, Zhang Z, Chen C, Liu Y, Si Q, Chuang T, et al. MicroRNA-19a-3p inhibits breast cancer progression and metastasis by inducing macrophage polarization through downregulated expression of Fra-1 proto-oncogene. Oncogene. 2014;33:3014–23.
Cai J, Guan H, Fang L, Yang Y, Zhu X, Yuan J, et al. MicroRNA-374a activates Wnt/β-catenin signaling to promote breast cancer metastasis. J Clin Invest. 2013;123:566.
Taylor MA, Sossey-Alaoui K, Thompson CL, Danielpour D, Schiemann WP. TGF-β upregulates miR-181a expression to promote breast cancer metastasis. J Clin Invest. 2013;123:150.
Yu S-J, Hu J-Y, Kuang X-Y, Luo J-M, Hou Y-F, Di G-H, et al. MicroRNA-200a promotes anoikis resistance and metastasis by targeting YAP1 in human breast cancer. Clin Cancer Res. 2013;19:1389–99.
Lei R, Tang J, Zhuang X, Deng R, Li G, Yu J, et al. Suppression of MIM by microRNA-182 activates RhoA and promotes breast cancer metastasis. Oncogene. 2014;33:1287–96.
Okuda H, Xing F, Pandey PR, Sharma S, Watabe M, Pai SK, et al. Liu Y: miR-7 suppresses brain metastasis of breast cancer stem-like cells by modulating KLF4. Cancer Res. 2013;73:1434–44.
Hu F, Meng X, Tong Q, Liang L, Xiang R, Zhu T, et al. BMP-6 inhibits cell proliferation by targeting microRNA-192 in breast cancer. Biochim Biophys Acta. 1832;2013:2379–90.
Chou J, Lin JH, Brenot A, Kim J-W, Provot S, Werb Z. GATA3 suppresses metastasis and modulates the tumour microenvironment by regulating microRNA-29b expression. Nat Cell Biol. 2013;15:201–13.
Zhu N, Zhang D, Xie H, Zhou Z, Chen H, Hu T, et al. Endothelial-specific intron-derived miR-126 is down-regulated in human breast cancer and targets both VEGFA and PIK3R2. Mol Cell Biochem. 2011;351:157–64.
Siragam V, Rutnam ZJ, Yang W, Fang L, Luo L, Yang X, et al. MicroRNA miR-98 inhibits tumor angiogenesis and invasion by targeting activin receptor-like kinase-4 and matrix metalloproteinase-11. Oncotarget. 2012;3:1370–85.
Zou C, Xu Q, Mao F, Li D, Bian C, Liu L-Z, et al. MiR-145 inhibits tumor angiogenesis and growth by N-RAS and VEGF. Cell Cycle. 2012;11:2137–45.
Kong W, He L, Richards E, Challa S, Xu C, Permuth-Wey J, et al. Upregulation of miRNA-155 promotes tumour angiogenesis by targeting VHL and is associated with poor prognosis and triple-negative breast cancer. Oncogene. 2014;33:679–89.
Xu Q, Jiang Y, Yin Y, Li Q, He J, Jing Y, et al. A regulatory circuit of miR-148a/152 and DNMT1 in modulating cell transformation and tumor angiogenesis through IGF-IR and IRS1. J Mol Cell Biol. 2013;5:3–13.
He T, Qi F, Jia L, Wang S, Song N, Guo L, et al. MicroRNA-542-3p inhibits tumor angiogenesis by targeting angiopoietin-2. J Pathol. 2014;232:499–508.
Yang R, Schlehe B, Hemminki K, Sutter C, Bugert P, Wappenschmidt B, et al. A genetic variant in the pre-miR-27a oncogene is associated with a reduced familial breast cancer risk. Breast Cancer Res Treat. 2010;121:693–702.
Zhong S, Chen Z, Xu J, Li W, Zhao J. Pre-mir-27a rs895819 polymorphism and cancer risk: a meta-analysis. Mol Biol Rep. 2013;40:3181–6.
Lian H, Wang L, Zhang J. Increased risk of breast cancer associated with CC genotype of Has-miR-146a rs2910164 polymorphism in Europeans. PLoS One. 2012;7:e31615.
Fan C, Chen C, Wu D. The association between common genetic variant of microRNA-499 and cancer susceptibility: a meta-analysis. Mol Biol Rep. 2013;40:3389–94.
Wang J, Bi J, Liu X, Li K, Di J, Wang B. Has-miR-146a polymorphism (rs2910164) and cancer risk: a meta-analysis of 19 case-control studies. Mol Biol Rep. 2012;39:4571–9.
Wang J, Wang Q, Liu H, Shao N, Tan B, Zhang G, et al. The association of miR-146a rs2910164 and miR-196a2 rs11614913 polymorphisms with cancer risk: a meta-analysis of 32 studies. Mutagenesis. 2012;27:779–88.
Wang P-Y, Gao Z-H, Jiang Z-H, Li X-X, Jiang B-F, Xie S-Y. The associations of single nucleotide polymorphisms in miR-146a, miR-196a and miR-499 with breast cancer susceptibility. PLoS One. 2013;8:e70656.
Chen J, Qin Z, Jiang Y, Wang Y, He Y, Dai J, et al. Genetic variations in the flanking regions of miR-101-2 are associated with increased risk of breast cancer. PLoS One. 2014;9:e86319.
Guan X, Liu H, Ju J, Li Y, Li P, Wang LE, Brewster AM, Buchholz TA, Arun BK, Wei Q: Genetic variant rs16430 6bp > 0bp at the microRNA-binding site in TYMS and risk of sporadic breast cancer risk in non-Hispanic white women aged ≤55 years. Molecular Carcinogenesis 2013
Gilam A, Edry L, Mamluk-Morag E, Bar-Ilan D, Avivi C, Golan D, et al. Involvement of IGF-1R regulation by miR-515-5p modifies breast cancer risk among BRCA1 carriers. Breast Cancer Res Treat. 2013;138:753–60.
Jiang Y, Qin Z, Hu Z, Guan X, Wang Y, He Y, et al. Genetic variation in a hsa-let-7 binding site in RAD52 is associated with breast cancer susceptibility. Carcinogenesis. 2013;34:689–93.
Jiang Y, Chen J, Wu J, Hu Z, Qin Z, Liu X, et al. Evaluation of genetic variants in microRNA biosynthesis genes and risk of breast cancer in Chinese women. Int J Cancer. 2013;133:2216–24.
Si H, Sun X, Chen Y, Cao Y, Chen S, Wang H, et al. Circulating microRNA-92a and microRNA-21 as novel minimally invasive biomarkers for primary breast cancer. J Cancer Res Clin Oncol. 2013;139:223–9.
Chan M, Liaw CS, Ji SM, Tan HH, Wong CY, Thike AA, et al. Identification of circulating microRNA signatures for breast cancer detection. Clin Cancer Res. 2013;19:4477–87.
Cuk K, Zucknick M, Madhavan D, Schott S, Golatta M, Heil J, et al. Plasma microRNA panel for minimally invasive detection of breast cancer. PLoS One. 2013;8:e76729.
Chen W, Cai F, Zhang B, Barekati Z, Zhong XY. The level of circulating miRNA-10b and miRNA-373 in detecting lymph node metastasis of breast cancer: potential biomarkers. Tumor Biol. 2013;34:455–62.
Kumar S, Keerthana R, Pazhanimuthu A, Perumal P. Overexpression of circulating miRNA-21 and miRNA-146a in plasma samples of breast cancer patients. Indian J Biochem Biophys. 2013;50:210–4.
Zeng RC, Zhang W, Yan XQ, Ye ZQ, Chen ED, Huang DP, et al. Down-regulation of miRNA-30a in human plasma is a novel marker for breast cancer. Med Oncol. 2013;30:1–8.
Ng EK, Li R, Shin VY, Jin HC, Leung CP, Ma ES, et al. Circulating microRNAs as specific biomarkers for breast cancer detection. PLoS One. 2013;8:e53141.
Markou A, Yousef GM, Stathopoulos E, Georgoulias V, Lianidou E. Prognostic significance of metastasis-related microRNAs in early breast cancer patients with a long follow-up. Clin Chem. 2014;60:197–205.
Wang S, Li H, Wang J, Wang D. Expression of microRNA-497 and its prognostic significance in human breast cancer. Diagn Pathol. 2013;8:172.
Falkenberg N, Anastasov N, Rappl K, Braselmann H, Auer G, Walch A, et al. MiR-221/-222 differentiate prognostic groups in advanced breast cancers and influence cell invasion. Br J Cancer. 2013;109:2714–23.
Tang D, Zhang Q, Zhao S, Wang J, Lu K, Song Y, et al. The expression and clinical significance of microRNA-1258 and heparanase in human breast cancer. Clin Biochem. 2013;46:926–32.
Hoppe R, Achinger-Kawecka J, Winter S, Fritz P, Lo W-Y, Schroth W, et al. Increased expression of miR-126 and miR-10a predict prolonged relapse-free time of primary oestrogen receptor-positive breast cancer following tamoxifen treatment. Eur J Cancer. 2013;49:3598–608.
Ma L, Li GZ, Wu ZS, Meng G. Prognostic significance of let-7b expression in breast cancer and correlation to its target gene of BSG expression. Med Oncol. 2014;31:1–5.
Volinia S, Croce CM. Prognostic microRNA/mRNA signature from the integrated analysis of patients with invasive breast cancer. Proc Natl Acad Sci. 2013;110:7413–7.
Bockhorn J, Dalton R, Nwachukwu C, Huang S, Prat A, Yee K, et al. MicroRNA-30c inhibits human breast tumour chemotherapy resistance by regulating TWF1 and IL-11. Nat Commun. 2013;4:1393.
Fang Y, Shen H, Cao Y, Li H, Qin R, Chen Q, et al. Involvement of miR-30c in resistance to doxorubicin by regulating YWHAZ in breast cancer cells. Braz J Med Biol Res. 2014;47:60–9.
Yin J, Zheng G, Jia X, Zhang Z, Zhang W, Song Y, et al. A Bmi1-miRNAs cross-talk modulates chemotherapy response to 5-fluorouracil in breast cancer cells. PLoS One. 2013;8:e73268.
Chen Y, Sun Y, Chen L, Xu X, Zhang X, Wang B, et al. miRNA-200c increases the sensitivity of breast cancer cells to doxorubicin through the suppression of E-cadherin-mediated PTEN/Akt signaling. Mol Med Rep. 2013;7:1579–84.
Zhong S, Li W, Chen Z, Xu J, Zhao J. MiR-222 and miR-29a contribute to the drug-resistance of breast cancer cells. Gene. 2013;531:8–14.
Jiao X, Zhao L, Ma M, Bai X, He M, Yan Y, et al. MiR-181a enhances drug sensitivity in mitoxantone-resistant breast cancer cells by targeting breast cancer resistance protein (BCRP/ABCG2). Breast Cancer Res Treat. 2013;139:717–30.
Hu H, Li S, Cui X, Lv X, Jiao Y, Yu F, et al. The overexpression of hypomethylated miR-663 induces chemotherapy resistance in human breast cancer cells by targeting heparin sulfate proteoglycan 2 (HSPG2). J Biol Chem. 2013;288:10973–85.
Shibahara Y, Miki Y, Onodera Y, Hata S, Chan MS, Yiu CC, et al. Aromatase inhibitor treatment of breast cancer cells increases the expression of let-7f, a microRNA targeting CYP19A1. J Pathol. 2012;227:357–66.
Ward A, Balwierz A, Zhang J, Küblbeck M, Pawitan Y, Hielscher T, et al. Re-expression of microRNA-375 reverses both tamoxifen resistance and accompanying EMT-like properties in breast cancer. Oncogene. 2013;32:1173–82.
He YJ, Wu JZ, Ji MH, Ma T, Qiao EQ, Ma R, et al. Mir‑342 is associated with estrogen receptor‑α expression and response to tamoxifen in breast cancer. Exp Ther Med. 2013;5:813–8.
Lin J, Liu C, Gao F, Mitchel R, Zhao L, Yang Y, et al. miR-200c enhances radiosensitivity of human breast cancer cells. J Cell Biochem. 2013;114:606–15.
Stankevicins L, da Silva APA, dos Passos FV, dos Santos FE, Ribeiro MCM, David MG, et al. MiR-34a is up-regulated in response to low dose, low energy x-ray induced DNA damage in breast cells. Radiat Oncol. 2013;8:231.
Gong C, Yao Y, Wang Y, Liu B, Wu W, Chen J, et al. Up-regulation of miR-21 mediates resistance to trastuzumab therapy for breast cancer. J Biol Chem. 2011;286:19127–37.
Bai WD, Ye XM, Zhang MY, Zhu HY, Xi WJ, Huang X, Zhao J, Gu B, Zheng GX, Yang AG: MiR-200c suppresses TGF-β signaling and counteracts trastuzumab resistance and metastasis by targeting ZNF217 and ZEB1 in breast cancer. International Journal of Cancer 2014
Ye X, Bai W, Zhu H, Zhang X, Chen Y, Wang L, et al. MiR-221 promotes trastuzumab-resistance and metastasis in HER2-positive breast cancers by targeting PTEN. BMB Rep. 2014;47:268–73.
Jung EJ, Santarpia L, Kim J, Esteva FJ, Moretti E, Buzdar AU, et al. Plasma microRNA 210 levels correlate with sensitivity to trastuzumab and tumor presence in breast cancer patients. Cancer. 2012;118:2603–14.
Ye X-M, Zhu H-Y, Bai W-D, Wang T, Wang L, Chen Y, et al. Epigenetic silencing of miR-375 induces trastuzumab resistance in HER2-positive breast cancer by targeting IGF1R. BMC Cancer. 2014;14:134.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
WenCheng Zhang is the first author.
Rights and permissions
About this article
Cite this article
Zhang, W., Liu, J. & Wang, G. The role of microRNAs in human breast cancer progression. Tumor Biol. 35, 6235–6244 (2014). https://doi.org/10.1007/s13277-014-2202-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2202-8