Abstract
Antibiotic susceptibility testing (AST), the screening of effective types and dosages of antibiotics, has become significantly important in the antimicrobial- resistance era over the last few decades. In order to overcome the limitations of conventional AST methods, several recent studies have developed AST platforms which exhibit the advantages of microfluidics. They demonstrated the performance of the platforms by determining effective antimicrobials for bacterial strains and their minimum inhibitory concentrations within hours. In this review, we cover recent developments of on-chip approaches for measurements of bacterial growth as well as for dilutions of antibiotic concentrations. We also discuss Point-of-Care AST devices that employ inexpensive materials and simple working principles to operate screenings near the site of care, which can potentially bring current laboratory- limited assays to clinical standards. All thing considered, emerging microfluidic AST devices have the potential to be decent alternatives to commonplace macro-scale AST methods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Antibiotic resistance of pathogenic bacteria has been recognized as a significant challenge in recent decades1,2. In particular, problems associated with antibiotics, such as inappropriate prescriptions, prolonged treatment, and excessive consumptions, are exacerbating the severity of this challenge3,4. Thus, in vitro antibiotic susceptibility testing (AST) is becoming increasingly important in this post-antibiotic era, since they provide medical information for finding suitable types of antibiotics as well as their effective dosages, which inhibit bacterial growth and retard resistance emergence5.
Conventional phenotypic AST methods mostly fulfill their role in screening proper antibiotic medications, particularly complemented with genotypic AST methods (e.g. polymerase chain reaction-based techniques and DNA microarrays)6,7. The traditional phenotypic assays such as micro-dilution on 96-well plates and disk diffusion on agar plates provide feasibility measurements without the need of biochemistry tools8,9. While these standard methods have become the staple of in vitro phenotypic ASTs, they nevertheless have several limitations. These macro-scale methods are labor-intensive to prepare the wide range of dosages, and are time-consuming to differentiate drug-susceptible cells from healthy cells (16–20 hours)10. They also require large specimen samples which don’t allow them to be used routinely. These drawbacks may hinder the ideal implementation of AST, which potentially results in weakened surveillance of resistance trends11.
In recent years, microfluidic AST platforms for phenotypic screening have emerged, with superior performance in certain aspects compared to the conventional methods12,13. The advantages are namely: lower cost, smaller amount of resources, reduced turnaround time, possible automation, more rapid evaluation, and higher sensitivity, comparatively12,13. Although most of the AST platforms have tested common types of antibiotics (e.g. Ampicillin, Gentamicin, and Ciprofloxacin, etc) against wild-type strains (e.g. Escherichia coll, Pseudomonas aeruginosa, and Staphylococcus aureus, etc.), these novel chips demonstrated their improved performance and validated their technical characteristics14. Here, we review the on-chip approaches for AST purposes, which are categorized into methods of observing growth levels and methods of diluting antibiotic concentrations. We also cover applications of Point-of-Care (POC) devices in AST, since the promising POC assays have demonstrated improved simplicity and utility over other laboratory-based microfluidic AST chips.
Measurement Methods of Cell Growth Level and Susceptibility in the Microfluidic Chips
Information about the growth level of bacteria, both in the presence and absence of antibiotics, should be provided for determining the baseline growth and susceptibility. In a microfluidic platform, while small volumes of samples could allow the entire duration of testing to be reduced to a matter of hours, they conversely make the measurements of cell growth level to be technically more difficult. Due to these small volumes and their consequent low accessibility of the samples loaded on the chip, on-chip growth level cannot be precisely measured using simple equipment, or using the naked eyes as in the conventional macroscale methods. Thus, most microfluidic studies have employed specific equipment, assays, or original techniques for obtaining the on-chip growth levels of bacteria. In this section, we review the chip-based measurements in four main categories: fluorescence imaging, metabolic activity indicators, optical imaging without labeling, and magnetic beads rotation measurement (Table 1). All the aforementioned categories are highlighted with their general characteristics and example on-chip studies showing their benefits over conventional methods.
Fluorescence Imaging for Cell Density and Viability
To monitor bacterial cell growth, many microbiologists have utilized bacteria strains that constitutively express plasmid-encoded green fluorescence protein (GFP)15. However, this approach involves a preliminary process of genetic medication at the molecular level, which may be complex for some strains and interferes with rapid AST16. In addition, GFP plasmid that is transformed to bacteria confers resistance to certain antibiotics and requires the antibiotic- supplemented media for constitutively expressing GFP. Despite these drawbacks, fluorescence imaging has been frequently utilized even in the microfluidic phenotypic AST (Table 1), because a significant linear correlation between fluorescence intensities and cell densities is stably presented for GFP-expressing strains, which allows for real-time monitoring of on-chip cell growth17. As a representative example, Golchin et al. monitored the on-chip growth and viability of GFP-expressing Mycobacterium smegmatis, whose susceptibility to Rifampicin was highlighted by the uptake of the nucleic acid stain such as propidium iodide, using a confocal laser microscope with high resolution (Figure 1a)18.
Representative on-chip bacterial growth measurement methods. (a) During 11 hours of Rifampicin treatment, GFP- expressing M. smegmatis gradually lost the GFP signal (green) and showed propodium iodide signal (red)18. Scale bars 10 μm. (b) Decreasing intensity of the fluorescence signal (red) from the metabolic product of resazurin indicates inhibited metabolism of E. coli by Kanamycin in a dose-dependent manner20. Scale bars 100 μm. (c) Brightfield microscope images show the filamentary formation of single cells of P. aeruginosa induced by Ceftazidime treatment23. Scale bar 25 μm. (d) Rotational periods of E. coli-bound magnetic beads gradually increase due to cell division, but increasing concentration of Gentamicin brings about constant rotational periods of the cell-bound bead25. Scale bar 2 μm. Images reproduced from the references with permission.
Metabolic Activity Indicators
Metabolic activity is involved in the life cycle of bacteria cells, and the magnitude of the activity is significantly correlated to cell growth or cell susceptibility to antibiotics. Resazurin is one of the most commonly used metabolism markers, and when it is irreversibly metabolized by bacteria, it is converted to resorufin, which exhibits strong fluorescence that can be observed using fluorescence microscopes even in the microfluidic chips (Table 1)19. For instance, Azizi et al. measured the fluorescence change from the metabolic product of resazurin and confirmed the inhibitory effect of Kanamycin on E. coli that was confined within nanoliter-sized chambers (Figure 1b)20.
The adenosine triphosphate (ATP) bioluminescence assay, another common indicator of metabolism, estimates the metabolic activity of bacteria by measuring photons (550–570 nm) from which luciferin substrate is catalyzed by luciferase in the presence of ATP and oxygen21. However, since the intensities of bioluminescence signals are significantly weaker than fluorescence intensities, the assay generally require more sensitive microplate readers that can record photons. For that reason, bioluminescence assays have not been employed frequently in microfluidic AST studies. Exceptionally, Dong et al. utilized them to quantify the bacterial growth in a multi-layered device that could be compatible with standard microplate readers22. They could measure the ATP bioluminescence signal generated from the metabolism of a pathogen sample with high sensitivity, and verified antimicrobial effects of eight drugs in 3-6 hours.
Optical Imaging Without Labeling
Several microfluidic-based studies have utilized bright-field microscope for single cell analysis or grayscale-based analysis without engineered strains and prestaining process, demonstrating their applicability and simplicity (Table 1). As a representative study of single cell imaging, Choi et al. mixed agarose gel with the bacterial sample and injected the mixture into the microchannel, resulting in fixed bacteria in the same location, without altering the properties of the cell (Figure 1c)23. They determined the minimum inhibitory concentrations (MICs) of antibiotics by screening changes in morphology (e.g. filamentary formation, swelling, dividing) and numbers of individual cells using a brightfield microscope. In the research by Hou et al., which employed grayscale- based analysis, they also used agarose gel to immobilize the bacteria cells24. They obtained bacterial densities from grayscale intensity changes of the images (from black to white) originated from the cell growth inside the gel, which is similar to measuring the turbidity of bacteria samples in optical density measurements. Although this approach requires a relatively higher initial cell density than in the single cell analysis method, the simple working principle enables a wide range of researchers to easily measure the growth level of a broad spectrum of strains.
Magnetic Beads Rotation Measurement for Cell Growth Level
The Kopelman group developed a biosensor that quantifies antibiotic toxicity in 15 – 30 minutes by measuring the rotation rate of cell-bound magnetic beads (Figure 1d, Table 1)25,26. They coated a single or small population of bacteria on an 8-μm-diameter magnetic bead by an immunoassay (off-chip), and generated nanoliter-volume droplets encapsulating the bacteria-bound beads in a microfluidic chip23,24. Under an external magnetic field, the rotational rate of the beads changed due to the growth and division of the bacteria, which was observed via a microscope. Although there were drawbacks, such as the immunoassay process and the time-consuming pre-incubation, their approach exhibited high sensitivity and rapid evaluation in the growth measurements.
Microfluidic Approaches for Diluting Antibiotic Concentrations
Some studies have employed microfluidic chips that enable the dilution of concentrations of an antibiotic, taking advantage of the precise manipulation of micro-volume fluids. On-chip dilution of antibiotic concentrations allows automation, reproducibility, and reduced turn-around time of preparing dilutions, in spite of employing complex designs of chips and requiring external equipment. In this section, we review on-chip dilution of antibiotics by three main methods: diffusion-based gradient formation, micro-droplet, and micro-array (Table 2).
Diffusion-based Formation of Concentration Gradients
Several studies have developed microfluidic devices which can generate a continuous and linear concentration gradient of the drug onto bacteria-trapped hydrogel between parallel channels by using a syringe pump (Table 2). Due to the constant flow in the channels by the pump, molecules of the drug collectively diffuse from the drug-supplemented channel to the drug-free channel. The complete formation of concentration gradient takes 30 to 90 minutes depending on the characteristic length of diffusion of the chip, and the gradient is maintained during the AST using the pump. For instance, Kim et al. injected a mixture of an agar gel and bacteria suspensions into a 1300 μm-wide channel and then applied the antibiotic gradient using an external pump for 6 hours to determine the MIC against P. aeruginosa (Figure 2a)27. They obtained the bacterial susceptibility to specific concentrations of the antibiotic by dividing the gel region into twenty-one portions along the gradient, and compared the MICs of the antibiotics tested in the chips with the standard MICs presented by the Clinical Laboratory Standards Institute. Furthermore, for on-chip combinatory AST in particular, Kim et al. utilized a microfluidic platform enabling formation of two orthogonal concentration gradients onto the bacteria-trapping gel within 35 minutes28. They tested the combinatory effects of several antibiotic pairs against P. aeruginosa on the chips, and they categorized the pairs into synergy or antagonism in 7 hours after injecting the antibiotic-supplemented media.
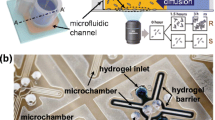
Representative on-chip antibiotic concentration diluting methods. (a) The flowing media both with and without an antibiotic in two parallel channels creates a linear concentration gradient of the antibiotic in the region of bacteria-trapped gel27. Scale bar 1.2 mm. (b) The mixing module passively generates a series of droplets with dilutions of a drug and mixes them with the bacteria suspension. Scale bar 5 mm. Reproduced from ref. 31 – Published by The Royal Society of Chemistry. (c) The valve-integrated micro-array automatically operates dilution functions of a drug within 9 minutes using an external pneumatic pump34. Images reproduced from the references with permission.
Dilution Using Micro-droplets
Micro-droplet systems automatically enable the creation of water-in-oil droplets whose antibiotic concentrations are serially diluted29,30. Individual droplets (volume of pL to mL) act as miniaturized incubators for the encapsulated target cells, which could additionally include drugs or viability-indicating dyes. In spite of having the disadvantages such as the addition of a surfactant to prevent coalescence, a potential risk of cross-contamination, and evaporation of the plug, this system is capable of precise control with small volumes of reagents and bacteria suspensions, enabling rapid AST against pathogens (Table 2). Interestingly, Derzsi et al. developed a passive-diluting platform that could make droplets using five manual pipettes, and then screen antibiotic toxicity in 5 hours (Figure 2b)31. The mixing module of the chip successively made aliquots of injected bacteria suspension, antibiotics, and pure medium (containing resazurin) with pre-defined volumes using the Rayleigh-Plateau instability. The measurement of fluorescence intensity confirmed that the final eleven droplets have serially diluted concentrations of an agent (approximately 160-fold between the lowest and highest concentrations). After incubating the droplets for 4 hours at 37 °C, they measured the resofurin fluorescence intensity of individual droplets and obtained the dose-response profile of Ampicillin against E. coli, consequently determining the MIC.
Dilution on the Micro-array Chips
Other systems featuring micro-arrays were also employed for generating various sets of antibiotic concentrations (Table 2)32. For instance, a tree-shape array causes the injected drugs-supplemented media to be separated and merged repeatedly, which in turn results in the drug concentrations to be gradually diluted, eventually generating a continuous profile of drug concentration onto a chamber that contains the bacteria suspension33. Another example is a valve-integrated array which operates the loading and mixing of the reagents by controlling the flow in the channels using a pneumatic pump (Figure 2c)17. Lee et al. utilized the valve-integrated array to treat various concentrations of Vancomycin against wild-type or Vancomycin-resistant Enterococcus faecalis in the chamber, and determined the MICs through pH-dependent colorimetric changes of the broth34. The system automatically mixed and diluted the bacteria suspension and antibiotic broth in 9 minutes, using the pump to press membrane-shape valves onto the channels.
Point-of-Care Microfluidics for AST
Point-of-Care tests, or POC tests in short, refer to the experiments carried outside the laboratory at or near the site of patient care, using equipment that can be easily delivered and that produce instantaneous (or at the very least, quick) results35–37. POC tests have found major applications in diagnostics38 and antimicrobial resistance measurement39–42, areas in which microfluidic devices are widely employed due to their patent advantages, such as being small, portable, and being able to manipulate small volumes of reagents. A representative example is the research conducted by Cira et al., where they introduced a poly-dimethylsiloxane (PDMS) microfluidic chip in which sample liquids were loaded and isolated in individual chambers by the presence of air or oil (Figure 3a)39. The device was particularly useful in determining the MIC of antibiotics since it isolated the bacteria chamber from the experimenter. Bacterial growth was easily identified by the change of color using a pH indicator. In addition to the usual advantages that microfluidic devices provide, this device was highly portable, avoiding nonessential equipment.
Representative Point-of-Care (POC) microfluidic devices for antibiotic research. (a) POC chip is assembled by connecting the channels to the chambers with aligning the microfluidic channels in one layer of PDMS and the chambers in another. The two top-view images show the determination of MIC values of wild-type E. coli and Kanamycin-resistant E. coli in two parallel devices39. (b) Immuno-strip biosensor is composed of numerous pads and a membrane. The device with 3 branches was used for testing specificity using S. aureus, P. aeruginosa, and S. aureus + P. aeruginosa40. (c) The device was used to compare results of Tetracycline and Kanamycin AST with those of AST on agar-filled Petri dishes using different concentrations of E. coli41. (d) Serial dilutions tests were conducted of E. coli that were both positive and negative (control) for expressing β-lactamase in order to exhibit specificity. Only β-lactamase expressing E. coli resulted in color change42. Images reproduced from the references with permission.
Most microfluidic AST devices are fabricated using cheap and accessible soft polymers (including PDMS) or glass, but these have certain disadvantages, such as being complex to operate and requiring oxygen plasma treatment to make channels. Furthermore, PDMS requires the careful consideration of uncured oligomers leaching into media, as well as the rapid partitioning of small hydrophobic molecules, including drug molecules, into the bulk, both of which adversely affect the data collected43. To overcome these inconvenient barriers, a market for a different category of microfluidic devices has emerged; this is where paper-based microfluidic chips enter the fray. The paper-based devices have recently undergone a soar in popularity due to their low-cost, low weight, and easy disposability while overcoming the limitations of the previously mentioned polymer materials38,44,45. For instance, in the research undertaken by Li et al., they used a multiplex immuno-sensing paper-disc for the detection using capillary action based antibody conjugated gold nanoparticles, and the subsequent analysis of whole cell bacteria, namely P. aeruginosa and S. aureus (Figure 3b)40. The detection was rapid, highly specific, and had a considerable detection range. In addition, Deiss et al. introduced a paper-based AST device in which the susceptibility of E. coli and Salmonella typhimurium were measured using the changing of colors from blue to pink to indicate growth (Figure 3c)41. The device was cheap and the fabrication and use were easy to follow. Furthermore, it produced results comparable to that of the conventional Kirby-Bauer AST, which is done with antibiotic-permeated paper disks atop a slab of agar46. Moreover, Boehle et al. also used low-cost paper-based devices to detect β-lactamase mediated resistance in E. coli by observing color changes in the colorimetric assay (Figure 3d)42. In all representative examples, the devices were cheap and easy to fabricate and use while needing a small volume of samples and reagents in relatively short amounts of time.
Conclusions and Future Perspectives
In this paper, we cover four prevalent methods of determining cell growth level for baseline and susceptibility of microfluidic phenotypic AST studies, and also review three on-chip methods of diluting antibiotic concentrations for alleviating burdens of experimental preparations. Lastly, we focus on the POC microfluidic chips for their use in AST because these devices have many advantages over alternative macro- or micro-scale AST platforms, such as low use of resources, low cost of materials, user-friendly handling, production of rapid results, and tailoring for specific user needs. All things considered, while most of the studies for microfluidic AST have screened general types of antibiotics against wild-type bacteria, focusing on the technical validation of their developed devices, those technologies have provided promising results which could make us consider them as standard methods applicable to clinical cases in the near future. Thus, it can be concluded that multifarious approaches are being used for different facets of microfluidic AST research.
Microfluidics for AST, and microfluidics in general, is an ever-growing field, with the gap between the conventional testing platforms and microfluidic chips getting smaller by the day. More novel antibiotic growth level measurements and antibiotic dilution methods are being studied at this very moment. In order to bring microfluidic chips for AST into the mainstream, collaborations are needed between the engineers who make the chips and the clinicians who would use them commercially. As microfluidic systems become simpler to use, and superfluous components such as active pumps are replaced by passive ones, the world would be more conducive to the aforementioned collaboration. As the microfluidic-based AST chips continue to be developed, we expect them to be widely utilized for their convenience and versatility in the near future.
References
Ventola, C.L. The antibiotic resistance crisis: part 1: causes and threats. Pharm. Ther. 40, 277–283 (2015).
Brown, E.D. & Wright, G.D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343 (2016).
Blair, J.M., Webber, M.A., Baylay, A.J., Ogbolu, D.O. & Piddock, L.J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Levy, S.B. & Marshall, B. Antibacterial resistance worldwide: causes, challenges and responses. Nat. Med. 10, S122–129 (2004).
van Belkum, A. et al. Developmental roadmap for antimicrobial susceptibility testing systems. Nat. Rev. Microbiol. 17, 51–62 (2019).
Jorgensen, J.H. & Ferraro, M.J. Antimicrobial susceptibility testing: a review of general principles and contemporary practices. Clin. Infect. Dis. 49, 1749–1755 (2009).
Balouiri, M., Sadiki, M. & Ibnsouda, S.K. Methods for in vitro evaluating antimicrobial activity: A review. J. Pharm. Anal. 6, 71–79 (2016).
Jenkins, S.G. & Schuetz, A.N. Current concepts in laboratory testing to guide antimicrobial therapy. Mayo Clin. Proc. 87, 290–308 (2012).
Reller, L.B., Weinstein, M., Jorgensen, J.H. & Ferraro, M.J. Antimicrobial susceptibility testing: a review of general principles and contemporary practices. Clin. Infect. Dis. 49, 1749–1755 (2009).
Wiegand, I., Hilpert, K. & Hancock, R.E.W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163–175 (2008).
Ayukekbong, J.A., Ntemgwa, M. & Atabe, A.N. The threat of antimicrobial resistance in developing countries: causes and control strategies. Antimicrob. Resist. Infect. Control 6, 47 (2017).
Liu, Z., Banaei, N. & Ren, K. Microfluidics for Combating Antimicrobial Resistance. Trends Biotechnol. 35, 1129–1139 (2017).
Dai, J., Hamon, M. & Jambovane, S. Microfluidics for Antibiotic Susceptibility and Toxicity Testing. Bioengineering 3 (2016).
Campbell, J. et al. Microfluidic advances in phenotypic antibiotic susceptibility testing. Biomed. Microdevices 18, 103 (2016).
Shaner, N.C., Steinbach, P.A. & Tsien, R.Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Errampalli, D. et al. Applications of the green fluorescent protein as a molecular marker in environmental microorganisms. J. Microbiol. Methods 35, 187–199 (1999).
Dai, J., Suh, S.J., Hamon, M. & Hong, J.W. Determination of antibiotic EC50 using a zero-flow microfluidic chip based growth phenotype assay. Biotechnol. J. 10, 1783–1791 (2015).
Golchin, S.A., Stratford, J., Curry, R.J. & McFadden, J. A microfluidic system for long-term time-lapse microscopy studies of mycobacteria. Tuberculosis (Edinb) 92, 489–496 (2012).
González-Pinzón, R., Haggerty, R. & Myrold, D.D. Measuring aerobic respiration in stream ecosystems using the resazurin-resorufin system. J. Geophys. Res.: Biogeosci. 117, G3 (2012).
Azizi, M. et al. Nanoliter-Sized Microchamber/Microarray Microfluidic Platform for Antibiotic Susceptibility Testing. Anal. Chem. (2018).
Mirasoli, M., Guardigli, M., Michelini, E. & Roda, A. Recent advancements in chemical luminescencebased lab-on-chip and microfluidic platforms for bioanalysis. J. Pharm. Biomed. Anal. 87, 36–52 (2014).
Dong, T. & Zhao, X. Rapid identification and susceptibility testing of uropathogenic microbes via immunosorbent ATP-bioluminescence assay on a microfluidic simulator for antibiotic therapy. Anal. Chem. 87, 2410–2418 (2015).
Choi, J. et al. A rapid antimicrobial susceptibility test based on single-cell morphological analysis. Sci. Transl. Med. 6, 267ra174 (2014).
Hou, Z. et al. Time lapse investigation of antibiotic susceptibility using a microfluidic linear gradient 3D culture device. Lab Chip 14, 3409–3418 (2014).
Sinn, I. et al. Asynchronous magnetic bead rotation (AMBR) biosensor in microfluidic droplets for rapid bacterial growth and susceptibility measurements. Lab Chip 11, 2604–2611 (2011).
Kinnunen, P. et al. Monitoring the growth and drug susceptibility of individual bacteria using asynchronous magnetic bead rotation sensors. Biosens. Bioelectron. 26, 2751–2755 (2011).
Kim, S., Lee, S., Kim, J.-K., Chung, H.J. & Jeon, J.S. Microfluidic-based observation of local bacterial density under antimicrobial concentration gradient for rapid antibiotic susceptibility testing. Biomicrofluidics 13, 014108 (2019).
Kim, S., Masum, F., Kim, J.-K., Chung, H.J. & Jeon, J.S. On-chip phenotypic investigation of combinatory antibiotic effects by generating orthogonal concentration gradients. Lab Chip (2019).
Cao, J. & Köhler, J.M. Droplet-based microfluidics for microtoxicological studies. Eng. Life Sci. 15, 306–317 (2015).
Theberge, A.B. et al. Microdroplets in microfluidics: an evolving platform for discoveries in chemistry and biology. Angew. Chem. Int. Ed. Engl. 49, 5846–5868 (2010).
Derzsi, L., Kaminski, T.S. & Garstecki, P. Antibiograms in five pipetting steps: precise dilution assays in sub-microliter volumes with a conventional pipette. Lab Chip 16, 893–901 (2016).
Wu, J., Wu, X. & Lin, F. Recent developments in microfluidics-based chemotaxis studies. Lab Chip 13, 2484–2499 (2013).
Kim, S.C., Cestellos-Blanco, S., Inoue, K. & Zare, R.N. Miniaturized Antimicrobial Susceptibility Test by Combining Concentration Gradient Generation and Rapid Cell Culturing. Antibiotics (Basel, Switz.) 4, 455–466 (2015).
Lee, W.B. et al. A microfluidic device for antimicrobial susceptibility testing based on a broth dilution method. Biosens. Bioelectron. 87, 669–678 (2017).
Bissonnette, L. & Bergeron, M.G. Diagnosing infections-current and anticipated technologies for pointofcare diagnostics and home-based testing. Clin. Microbiol. Infect. 16, 1044–1053 (2010).
Park, S., Zhang, Y., Lin, S., Wang, T.H. & Yang, S. Advances in microfluidic PCR for point-of-care infectious disease diagnostics. Biotechnol. Adv. 29, 830–839 (2011).
Syedmoradi, L. et al. Point of care testing: The impact of nanotechnology. Biosens. Bioelectron. 87, 373–387 (2017).
Hu, J. et al. Advances in paper-based point-of-care diagnostics. Biosens. Bioelectron. 54, 585–597 (2014).
Cira, N.J., Ho, J.Y., Dueck, M.E. & Weibel, D.B. A self-loading microfluidic device for determining the minimum inhibitory concentration of antibiotics. Lab Chip 12, 1052–1059 (2012).
Li, C.Z. et al. Paper based point-of-care testing disc for multiplex whole cell bacteria analysis. Biosens. Bioelectron. 26, 4342–4348 (2011).
Deiss, F., Funes-Huacca, M.E., Bal, J., Tjhung, K.F. & Derda, R. Antimicrobial susceptibility assays in paper-based portable culture devices. Lab Chip 14, 167–171 (2014).
Boehle, K.E. et al. Utilizing Paper-Based Devices for Antimicrobial-Resistant Bacteria Detection. Angew. Chem. Int. Ed. Engl. 56, 6886–6890 (2017).
Regehr, K.J. et al. Biological implications of polydimethylsiloxanebased microfluidic cell culture. Lab Chip 9, 2132–2139 (2009).
Yetisen, A.K., Akram, M.S. & Lowe, C.R. Paperbased microfluidic point-of-care diagnostic devices. Lab Chip 13, 2210–2251 (2013).
Park, M., Kang, B.-H. & Jeong, K.-H. Paper-Based Biochip Assays and Recent Developments: A Review. Biochip J. 12, 1–10 (2018).
Schwalbe, R., Steele-Moore, L. & Goodwin, A.C. Antimicrobial susceptibility testing protocols. (Crc Press, 2007).
Liu, Z., Sun, H. & Ren, K. A Multiplexed, Gradient-Based, Full-Hydrogel Microfluidic Platform for Rapid, High-Throughput Antimicrobial Susceptibility Testing. ChemPlusChem 82, 792–801 (2017).
Sabhachandani, P. et al. Integrated microfluidic platform for rapid antimicrobial susceptibility testing and bacterial growth analysis using bead-based biosensor via fluorescence imaging. Microchimica Acta 184, 4619–4628 (2017).
Dai, J. et al. Charting microbial phenotypes in multiplex nanoliter batch bioreactors. Anal. Chem. 85, 5892–5899 (2013).
Sun, P. et al. High-throughput microfluidic system for long-term bacterial colony monitoring and antibiotic testing in zero-flow environments. Biosens. Bioelectron. 26, 1993–1999 (2011).
Mohan, R. et al. A microfluidic approach to study the effect of bacterial interactions on antimicrobial susceptibility in polymicrobial cultures. RSC Adv. 5, 35211–35223 (2015).
Mohan, R. et al. A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens. Bioelectron. 49, 118–125 (2013).
Baraban, L. et al. Millifluidic droplet analyser for microbiology. Lab Chip 11, 4057–4062 (2011).
Puchberger-Enengl, D., van den Driesche, S., Krutzler, C., Keplinger, F. & Vellekoop, M.J. Hydrogel-based microfluidic incubator for microorganism cultivation and analyses. Biomicrofluidics 9, 014127 (2015).
Kalashnikov, M., Lee, J.C., Campbell, J., Sharon, A. & Sauer-Budge, A.F. A microfluidic platform for rapid, stress-induced antibiotic susceptibility testing of Staphylococcus aureus. Lab Chip 12, 4523–4532 (2012).
Kalashnikov, M. et al. Rapid phenotypic stressbased microfluidic antibiotic susceptibility testing of Gram-negative clinical isolates. Sci. Rep. 7, 8031 (2017).
Kalashnikov, M., Lee, J.C. & Sauer-Budge, A.F. Optimization of Stress-Based Microfluidic Testing for Methicillin Resistance in Staphylococcusaureus Strains. Diagnostics 8 (2018).
DiCicco, M. & Neethirajan, S. An in vitro microfluidic gradient generator platform for antimicrobial testing. Biochip J. 8, 282–288 (2014).
Avesar, J. et al. Rapid phenotypic antimicrobial susceptibility testing using nanoliter arrays. Proc. Natl. Acad. Sci. U.S.A. 114, E5787–E5795 (2017).
Churski, K. et al. Rapid screening of antibiotic toxicity in an automated microdroplet system. Lab Chip 12, 1629–1637 (2012).
Boedicker, J.Q., Li, L., Kline, T.R. & Ismagilov, R.F. Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 8, 1265–1272 (2008).
Tang, Y., Zhen, L., Liu, J. & Wu, J. Rapid antibiotic susceptibility testing in a microfluidic pH sensor. Anal. Chem. 85, 2787–2794 (2013).
Li, B. et al. Gradient microfluidics enables rapid bacterial growth inhibition testing. Anal. Chem. 86, 3131–3137 (2014).
Choi, J. et al. Rapid antibiotic susceptibility testing by tracking single cell growth in a microfluidic agarose channel system. Lab Chip 13, 280–287 (2013).
Choi, J. et al. Rapid drug susceptibility test of Mycobacterium tuberculosis using microscopic timelapse imaging in an agarose matrix. Appl. Microbiol. Biotechnol. 100, 2355–2365 (2016).
Choi, J. et al. Direct, rapid antimicrobial susceptibility test from positive blood cultures based on microscopic imaging analysis. Sci. Rep. 7, 1148 (2017).
Baltekin, O., Boucharin, A., Tano, E., Andersson, D.I. & Elf, J. Antibiotic susceptibility testing in less than 30 min using direct single-cell imaging. Proc. Natl. Acad. Sci. U.S.A. 114, 9170–9175 (2017).
Kim, K., Kim, S. & Jeon, J.S. Visual Estimation of Bacterial Growth Level in Microfluidic Culture Systems. Sensors (Basel) 18 (2018).
Kinnunen, P. et al. Self-assembled magnetic bead biosensor for measuring bacterial growth and antimicrobial susceptibility testing. Small 8, 2477–2482 (2012).
Malmberg, C. et al. A Novel Microfluidic Assay for Rapid Phenotypic Antibiotic Susceptibility Testing of Bacteria Detected in Clinical Blood Cultures. PLoS ONE 11, e0167356 (2016).
Acknowledgments
This work was supported by the Climate Change Research Hub (Grant No. N11180109) of the Korea Advanced Institute of Science and Technology (KAIST), Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2017R1 D1A1B03030428) and the BK 21 Plus program.
Author information
Authors and Affiliations
Corresponding author
Conflict of Interests
Conflict of Interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kim, S., Masum, F. & Jeon, J.S. Recent Developments of Chip-based Phenotypic Antibiotic Susceptibility Testing. BioChip J 13, 43–52 (2019). https://doi.org/10.1007/s13206-019-3109-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-019-3109-7