Abstract
Plant communities growing in metal-contaminated areas can develop resistance mechanisms by establishing symbiotic associations with endophytic microorganisms. The functionality and diversity of endophytic communities depend on the amount and type of metal present in the soil. To characterise the response of endophytic bacterial communities to mercury-induced abiotic stress, we analysed the colonization frequency and number of bacterial isolates in the roots of Aeschynomene fluminensis (Joint Vetch) and Polygonum acuminatum (Smartweed), which represent the families Fabaceae and Polygonaceae, respectively. These two plant species are found in many mercury-contaminated areas. The isolates were characterised by morpho- and genotyping and identified by 16S rDNA gene sequencing. The bacteria belonged to the phyla Actinobacteria, Bacteriodetes, Firmicutes, and Proteobacteria. The Hill series and Venn diagram provided evidence that mercury affects the composition, diversity, and richness of the endophytic bacterial communities. Inoculation with Bacillus_sp_BacI34, Burkholderia_sp_BacI45, Enterobacter_sp_BacI14, Enterobacter_sp_BacI26, Enterobacter_sp_BacI18, Klebsiella_pneumoniae_BacI20, Lysobacter_soli_BacI39, Pantoea_sp_BacI16, and Pantoea_sp_BacI23 promoted the growth of corn (Zea mays) plants in mercury-supplemented substrata. It is noteworthy that Pantoea sp_BacI23 increased the host plant length (root and shoot) by 117.09 ± 0.28%. Endophytic bacterial strains may well provide important inoculants for plant growth promotion on metal-contaminated sites and in metal bioremediation programs.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Mercury is a class B metal (Nieboer and Richardson 1980) naturally found in the earth crust that occurs in soil, water, and air in several chemical forms, including metallic (Hg0), ionic (Hg+, Hg2+), organometallic ((CH3)2Hg, CH3Hg+) forms (Carrasco-Gil et al. 2013). Transformation of this metal via methylation, demethylation, and reduction depends on its distribution and the environment conditions (Asaduzzaman et al. 2019). Mercury is among the 20 substances that the United States Environmental Protection Agency and the Agency for Toxic Substances and Disease Registry classify as highly toxic to humans and aquatic organisms (Ullah et al. 2015; Darko et al. 2016). It therefore threatens not only human and animal health, but also ecosystems (Román-Ponce et al. 2016). The high toxicity of mercury has prompted the search for strategies that minimise its detrimental effects or contamination levels to the environment (Farias et al. 2012; Seccatore et al. 2014; Oliveira et al. 2015).
The Pantanal is the largest tropical wetland in the world and comprises one of the largest and most biodiverse biomes in Brazil (Junk et al. 2014). Anthropic influences, such as deforestation, erosion, and gold mining have led to severe mercury contamination in many parts of the Pantanal (Ceccatto et al. 2016), where the mercury concentration in suspended sediments has ranged from 0.02 to 0.61 mg.kg−1 (Lacerda et al. 1991). The Brazilian legislation (BRASIL 2009) and World Health Organization (WHO, 2003) recommend that mercury concentrations should be lower than 0.5 mg.kg−1 for urban areas. The main problem is illegal mining activity that has increased the mercury contamination of soil, water, and biota (Ceccatto et al. 2016; Cebalho et al. 2017). Mercury levels in the soil samples near a gold-mining site of Poconé, Mato Grosso State, Brazil, are 6.48 times greater than the limit established by the Brazilian legislation (Pietro-Souza et al. 2017). Mercury contamination is of public interest because it threatens humans via various exposure routes (Seccatore et al. 2014), and this metal has the capacity to bioaccumulate in the food chain (Vishnivetskaya et al. 2011; Zhang et al. 2013; Mani and Kumar 2014; Alanoca et al. 2016).
Mercury contamination of natural areas exerts a strong selective pressure for the development of mercury-resistant plants (Fidalgo et al. 2016). The association with endophytes enables the host plants to adapt and face adverse conditions including attack by phytopathogens (Soares et al. 2016a), high levels of metal contamination (Shen et al. 2013; Manohari and Yogalakshmi 2016), and other physical and chemical stresses (Rodriguez et al. 2008; Soares et al. 2016c). Endophytic microorganisms inhabit the internal organs of the host plant without causing any disease or infection (Schulz and Boyle 2005; White et al. 2014). These microorganisms have functional traits that promote host plant growth, such as phosphate solubilization and the production and release of ammonia, cyanuric acid, hydrocyanic acid, indoleacetic acid (IAA), nitrogen, siderophores, and enzymes (amylase, cellulase, esterase, and protease) (Cuzzi et al. 2011; Glick 2015; Mathew et al. 2015; Manohari and Yogalakshmi 2016; Soares et al. 2016b).
Plant-associated endophytes can remove, transform, and even assimilate the contaminants present in sediments, soil, water, and air as a strategy to mitigate the toxic effects of metals (Zhang et al. 2013, 2016). Bacteria have mercury resistance mechanisms mediated by enzymes encoded in the operon mer that are capable of reducing this metal (Harichová et al. 2012; Yu et al. 2014). Other mechanisms of bacterial resistance to toxic metals involve alteration of plasma membrane permeability, cell morphology, and efflux systems; biosorption, complexation, demethylation, oxidation, precipitation, reduction, and volatilization of metals; and production of exopolysaccharides (Yu et al. 2014; Ullah et al. 2015; Xie et al. 2015; Naik and Dubey 2016).
Plants can host mercury-resistant endophytic bacteria (Pérez et al. 2016; Durand et al. 2018) and fungi (Pietro-Souza et al. 2017). Soil mercury contamination influences the composition and structure of root endophytic fungal communities of Aeschynomene fluminensis Vell and Polygonum acuminatum Kunth. that colonize wetland environments (Pietro-Souza et al. 2017). We hypothesise that plants growing in mercury-contaminated environments host endophytic bacterial communities. The objectives of the present study were a) to characterise the endophytic bacterial community isolated from roots of Aeschynomene fluminensis Vell. and Polygonum acuminatum Kunth. collected at environments contaminated or not with mercury; b) to identify the mercury-resistant community in the collected samples; c) to characterise functional traits important for bioremediation and host plant growth; and d) to examine to what extent bacteria inoculation improves plant growth.
2 Materials and methods
2.1 Sampling and processing
The biological material and soil samples were collected in September 2014, at three sites of Poconé, a typical Pantanal region from the State of Mato Grosso, Brazil: Site 1: S 16°15′42.7” W 056°38′43.6″; Site 2: S 16°21′19.7” W 056°20′13.9″; and Site 3: S 16°15′51.3 “W056°38′54.3″. This area is characterised by a rainy period from October to April, a drought period from June to December, a long flooding period from December to May (Junk et al. 2016), annual average rainfall of 1239 mm, and temperature of 26 °C (Alvares et al. 2013). The climate is classified as Aw (Köppen 1930). Data from previous chemical analyses that determined the total soil mercury concentration were used to select the collection sites (Pietro-Souza et al. 2017).
Endophytic bacteria were isolated from roots of five adult plants of A. fluminensis (named Asc) and P. acuminatum (named Pol) collected at areas contaminated or not with mercury (named HgY and HgN, with Hg2+ levels of 3.24 and < 0.0017 mg/kg, respectively). These plant species were chosen due to their capacity to colonize HgY environments abundantly (Pietro-Souza et al. 2017). The soil and vegetal material were packaged in plastic bags, identified according to the collection site, and stored at 4 °C until processing. The samples were superficially cleaned with neutral detergent, washed with tap water, and further superficially disinfected with ethanol 70% (1 min) and sodium hypochlorite 2.5% (5 min). They were then rinsed five times with sterile distilled water (Pietro-Souza et al. 2017).
Three bacterial isolation procedures (de Souza et al. 2013; Franchi et al. 2017) were then used: fragmentation, maceration, and enrichment. 1) Fragmentation: 120 root fragments of each sample were transferred to Petri dishes (N = 10) containing Luria Bertani (LB) medium supplemented with 30 μg.mL−1 of HgCl2 (LB + Hg). 2) Maceration: the disinfected roots were macerated and diluted (10−1 to 10−3) in 0.87% NaCl, and further plated in triplicate in solid LB medium not supplemented with HgCl2. 3) Enrichment: 5 mL of the macerate were inoculated in 45 mL of LB + Hg broth and shaken (100 rpm; 72 h); 5 mL of this culture were inoculated in 45 mL of LB + Hg broth and incubated under the same conditions; finally, the culture was diluted (10−1 to 10−8) in 0.87% NaCl and plated in triplicate in solid LB + Hg medium (Cabral et al. 2013). In the three, the Petri dishes were incubated at 28 °C and analysed daily. The colonies were characterised macroscopically and grouped morphologically after purification. The strains were stored in 20% glycerol, at −20 °C.
2.2 Identification of root endophytic bacteria
DNA was extracted from the isolated strains using the Wizard Genomic DNA Purification Kit (Promega) following the manufacturer’s protocol. The morphological groups were confirmed by ERIC-PCR fingerprinting of the products using the initiator oligos ERIC-1R (5′-ATGTAAGCTCCTGGGGATTCAC-3′) and ERIC-2 (5′-AAGTAAGTGACTGGGGTGAGCG-3′) (Vandamme et al. 1993).
One lineage from each ERIC-PCR group was identified through 16S rDNA gene sequencing, using the primers 27F and 1492R to amplify the 16S gene region (Lane 1991). The amplicons were enzymatically purified using ExoSap-it PCR Product Cleanup Reagent (GE Healthcare) and sequenced by the Sanger method using the BigDye™ Terminator Cycle Sequencing kit. The sequences were edited using BioEdit software (version 7.2.5) and compared to sequences deposited at GenBank using the nBLAST tool (http://www.ncbi.nlm.nih.gov/). The nucleotide sequences were deposited at GenBank under the accession numbers KX641492 to KX641588.
2.3 Plant growth-promoting properties and mercury resistance of the isolates
The isolated bacterial strains were characterised with respect to their capacity to solubilize inorganic phosphates (Podile and Kishore 2007), fix nitrogen (Cavalcante and Dobereiner 1988), synthesize ammonia (Pandey et al. 2015) and IAA (Cuzzi et al. 2011), and secrete hydrocyanic acid (HCN) (Lorck 1948), siderophores (Milagres et al. 1999), and the hydrolytic enzymes amylase, cellulase, esterase, and protease (Carrim et al. 2006). The presence of halo, color change and/or colony growth was analysed for each methodology. The minimal inhibitory concentration (MIC) of mercury was determined in LB broth containing serial concentrations of Hg2+ (0–500 μg/mL) (El-deeb et al. 2012).
2.4 Mitigation of mercury toxicity to host plants by endophytic bacteria
Asc and Pol seed germination and growth in the greenhouse are very irregular, making it very difficult to use them in bioremediation assays. Corn (Zea mays hybrid maize AG 1051) plants were chosen due to their agronomic importance to this region of Brazil (Duarte and Pasa 2016) and their effectiveness for bioremediation of contaminants and metals (Mani and Kumar 2014; Dixit et al. 2015; Ullah et al. 2015; Shinwari et al. 2015), including mercury remediation (Pietro-Souza et al. 2017).
Endophytic bacterial strains isolated using the fragmentation and enrichment methods in mercury-supplemented medium were selected for the assays of corn plant growth promotion. First, corn seeds were disinfected by immersion in 70% ethanol (1 min) and 2.5% sodium hypochlorite (5 min), rinsed in sterile distilled water, and submerged for 1 h in the test bacterial suspension previously activated in LB broth (OD: 108 CFU.mL−1). Next, the seeds were transferred to 1.0 dm3 vessels containing vermiculite:sand 1:1 (m:m) supplemented with 40 mg.kg−1 of HgCl2. Seven days after sowing, bacteria were reinoculated by adding 1 mL of bacterial inoculum (OD: 108 CFU.mL−1) to the soil near the plants. The field capacity of the substratum was maintained at 70%, and it was irrigated weekly with 70% ionic strength Hoagland solution (Hoagland and Arnon 1950). The control groups consisted of plants not inoculated with endophytic bacteria (P), and plants inoculated with endophytic bacteria grown in vessels with (CHgY) or without (CHgN) addition of mercury. After 20 days of cultivation, the length of aerial shoots and roots was measured. The growth promotion efficiency was calculated to determine how effectively endophytic bacteria promoted plant growth (Almoneafy et al. 2014).
2.5 Data analysis
The colonization frequency in root fragments inoculated with bacteria was calculated and data were analysed using the F and Student’s t tests (p < 0.05) (Harris and Sommers 1968). The diversity of bacterial communities was analysed using the Hill Series (Hill 1973). The species composition of the communities (AscHgN, AscHgY, PolHgN, PolHgY) was visualized in the Venn diagram constructed with the aid of the online software DrawVenn (http://bioinformatics.psb.ugent.be/webtools/Venn/).
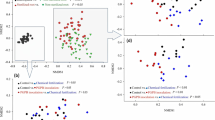
The co-occurrence patterns of microbial taxa within the host and contaminated soil was explored by Network analysis. A Spearman’s correlation between two genera was considered statistically robust if p < 0.05 (Vegan package on R). Bacterial modules or sub-communities of the community were calculated using the Louvain algorithm (Blondel et al. 2008) and network properties were calculated using the statistics tools implemented in Gephi 0.9.1 (Bastian et al. 2009).
Results from the qualitative functional characterisation were expressed as positive (+) or negative (−) when the production of functional traits were detected or not, respectively. Differences between treatments in the corn growth parameters were analysed by the Dunnett’s test, using the softwares R (version 3.2.5) and Assistant 7.7.
3 Results
3.1 Structure of the endophytic bacterial community of A. fluminense and P. acuminatum
This section comprises data from four endophytic bacterial communities isolated from roots of A. fluminense (Asc) and P. acuminatum (Pol) collected at areas contaminated (HgY) or not (HgN) with mercury, which were named as AscHgN, AscHgY, PolHgN, and PolHgY.
Fragmentation of root tissues allowed estimation of the extent of plant tissue colonization by endophytes. The percentage of root fragments colonized by endophytic bacteria in the HgY area (41.88 ± 34.56%) was higher than the percentage found in the HgN area (14.12% ± 13.49) (p < 0.05), regardless the plant species. Colonization rate in Pol roots (48.10% ± 29.59) was greater than colonization rate in Asc roots (7.92% ± 7.79) collected at both sites (p < 0.05).
A total of 207 bacterial strains were isolated from root fragments of AscHgN, AscHgY, PolHgN, and PolHgY inoculated in mercury-supplemented medium: 34, 44, 59, and 70 strains, respectively. The isolated strains belonged to phyla Proteobacteria (PolHgY = AsHgY = 100%, PolHgN = 98.31%, and AscHgN = 76.47%) and Firmicutes (PolHgN = 1.69% and AscHgN = 23.53%) (Table 1); the last phylum was exclusively found in HgN areas. The classes identified were: Alphaproteobacteria, Bacilli, Betaproteobacteria, and Gammaproteobacteria; the two last ones were isolated from all the communities evaluated, and they were abundant in Pol and Asc roots, respectively. The class Bacilli was exclusively found in HgN areas, especially in Asc (23.53%). Nine genera distributed in 23 species were identified, among which Enterobacter was the most frequent in AscHgY (47.73%) and PolHgY (51.43%), and Burkholderia was the most frequent in AscHgN (66.7%) and PolHgN (44.12%). The dominant species were Burkholderia_sp_BacI41 (32.35%) in AscHgN, Burkholderia_cepacia_BacI47 and Pantoea_sp_BacI16 (64.40%) in AscHgY, Enterobacter_sp1_X (29.55%) in PolHgN, and Enterobacter_cloacae_X (51.43%) in PolHgY (Table 1).
The Hill series demonstrated that richness and diversity indices depended on the plant species and the presence or absence of mercury in the soil (Fig. 1). Endophytic bacterial communities from the HgN site had richness (when a = 0, AscHgN = 7 and PolHgN = 8) and diversity α greater than communities from the HgY site (AscHgY = 4 and PolHgY = 7). The Shannon diversity indice confirmed these parameters (eH’ when a = 1; AscHgN = 1.54, AscHgY = 1.38, PolHgN = 1.63, and PolHgY = 1.28). The Simpson diversity indice (1/D when a = 2) evidenced that Pol was represented by dominant species and low diversity as compared with Asc, regardless mercury contamination (AscHgN and AscHgY = 0.75, PolHgN = 0.76, PolHgY = 0.64) (Fig. 1a).
Pol roots had lower number of cultivable endophytic bacteria per gram of macerated root tissue (PolHgN = 1.04 × 105 ± 5.02 × 103 UFC/g and PolHgY = 3.27 × 105 ± 1.36 × 104 UFC/g) than Asc roots (AscHgN = 1.11 × 105 ± 6.82 × 103 UFC/g and AscHgY = 5.72 × 105 ± 3.23 × 104 UFC/g) (p < 0.05). Fifty-one morphotypes were differentiated on the basis of the growth characteristics in culture medium. They belonged to phyla Bacteriodetes (0.39%), Actinobacteria (8.05%), Proteobacteria (21.58%), and Firmicutes (69.98%), and were distributed in 18 genera and 39 species (Table 1), as determined by ERIC-PCR fingerprinting and 16S rDNA gene sequencing. Bacteriodetes was exclusively found in the PolHgN community while Actinobacteria was exclusively found in Pol, independently of the environment. The most abundant species in AscHgN, AscHgY, PolHgN, and PolHgY communities were Burkholderia_kururiensis_BacI100 (20.65%), Ralstonia_sp_X (22.49%), Bacillus_subtilis_BacI75 (17.96%), and Enterobacter_sp3_X (23.08%), respectively (Table 1). The Hill Series of diversity indices provided evidence that endophytic bacterial communities from plants grown in HgY areas had richness, diversity, and dominance indices (AscHgY: richness = 8, Shannon = 1.99, Simpson = 0.85; PolHgY: richness = 18, Shannon = 2.67, Simpson = 0.91) greater than communities from plants grown in HgN areas (AscHgN: richness = 6, Shannon = 1.78, Simpson = 0.83; PolHgN: richness = 13, Shannon = 2.14, Simpson = 0.86). Pol had richness, diversity, and dominance indices greater than Asc, regardless the collection site (Fig. 1b).
The four endophytic bacterial communities differed markedly with respect to their composition (Fig. 2). Data from both isolation methods revealed that more species were specific to one host than were shared (Fig. 2). Enterobacter_sp1_X specifically colonized Asc roots while Enterobacter_cloacae_X and Klebsiella_pneumoniae_X specifically colonized Pol roots, as evidenced by fragmentation of root tissues (Table 1). Enterobacter_ludwigii_X was isolated from Asc and Pol roots collected at contaminated sites (AscHgY and PolHgY), using the maceration technique (Table 1).
The endophytic bacterial communities of plants collected at HgY areas were enriched after three cycles of passage in culture media supplemented with mercury. Inoculation with root samples from hosts collected at the HgN site did not result in microbial growth, but inoculation with root samples collected at the HgY site resulted in bacterial isolates from the phyla Proteobacteria (84.61%) and Firmicutes (7.69%), that included 8 genera and 12 species (Table S1). This method also resulted in the isolation of the yeast Rhodotorula mucilaginosa_X from Asc roots.
The Asc roots had higher species richness than Pol roots: 8 and 5 species, respectively. The relative abundance of these species in AscHgY was Rhodotorula mucilaginosa_X (71.60%), Pseudomonas_sp_BacI38 (10.76%), Klebsiella_sp_BacI31 (8.52%), Enterobacter_sp_BacI32 (3.14%), Pseudomonas_stutzeri_BacI36 (1.94%), Bacillus_sp_X (1.79%), Sphingomonas_sp_X (1.79%), and Lysobacter_soli_BacI39 (0.45%), while the relative abundance of species in PolHgY was Enterobacter_sp_BacI14 (97.16%), Klebsiella_pneumoniae_BacI15 (2.65%), Enterobacter_sp_BacI12 (0.06%), Novosphingobium_sp_BacI10 (0.06%), and Pantoea_agglomerans_BacI11 (0.06%).
A correlation matrix was constructed using qualitative data from the presence or absence of the species identified through the three isolation methods – fragmentation, maceration, and enrichment. The data were also used to construct interaction networks to compare the hosts and collection sites (Table 2). A total of 66 species were present in the communities AscHgN, AscHgY, PolHgN, and PolHgY: 11, 20, 20, and 28, respectively. The parameters of the interaction network of endophytic bacteria (Table 2) evidenced that (i) Pol had a more structured network; (ii) the presence or absence of mercury had a determining force on the interaction and connectivity among endophytic species; and (iii) endophytic bacterial communities from plants collected at HgY environments were centralized with less modularity (Table 2).
3.2 Functional characterisation of endophytic bacterial communities
Thirteen endophytic bacterial strains had score = 7 for the promising plant growth-promoting functional traits (Table S1), while six strains had low score (2) for ammonia and IAA. Strains isolated from plants collected at HgY environments exhibited greater proportion of functional traits than strains isolated from plants collected at HgN areas (Table 3). PolHgY roots hosted three amylase-secreting strains (Bacillus_nanhaiensis_BacI69, Enterobacter_sp_BacI14, and Klebsiella_sp_BacI2), and three siderophore-producing strains (Bacillus_megaterium_BacI64, Enterobacter_sp_BacI14, and Kosakonia_cowanii_BacI60). AscHgY and PolHgN roots hosted the cyanide acid-producing bacterial strains Burkholderia_seminalis_BacI48 and Enterobacter_sp_BacI22, respectively. Ammonia was produced by 27.63%, 26.80%, 22.68%, and 35.50% of the bacterial strains isolated from roots of AscHgN, AscHgY, PolHgN, and PolHgY, respectively (Table 3). IAA-secreting and nitrogen-fixing bacterial strains predominated in plants collected at HgY areas (Table 3).
Maceration of root tissues provided isolation of endophytic bacterial strains that were more sensitive to mercury, with MIC values ranging from 0 to 62 μg/mL of Hg2+; MIC values of most of the strains ranged from 0 to 7.5 μg/mL of Hg2+ (Fig. 3 and Table S1). Endophytic bacterial strains isolated using the fragmentation and enrichment techniques exhibited broader ranges of MIC values: 0–250 μg/mL and 15–500 μg/mL of Hg2+, respectively. The last technique provided isolation of mercury-resistant strains with high MIC values.
3.3 Host growth promotion in the presence of mercury
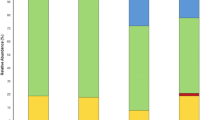
Addition of 40 mg.kg−1 of HgCl2 to the substrate reduced the corn plant length (CHgY = 25.16 ± 2.65 cm) by approximately 40% relative to the plants grown in the absence of this metal (CHgN = 43.03 ± 4.80 cm) (Dunnett’s test, p < 0.05) (Fig. 4). Growth reduction was more pronounced in the shoot (43.5% reduction) than in the root (38.4% reduction) (Fig. 4). Corn plant inoculation with 27 endophytic bacterial strains promoted growth of plants seeded in the HgCl2-supplemented substrate (Fig. 4); 36.36% and 63.64% of such strains were isolated using the enrichment and fragmentation techniques, respectively. Inoculation with B. cereus_BacI42 and Pantoea_sp_BacI23 increased the plant length by 57.48 ± 5.45 and 117.09 ± 0.28%, respectively. Inoculation with Bacillus_sp_BacI34, Burkholderia_sp_BacI45, Enterobacter_sp_BacI14, Enterobacter_sp_BacI26, Enterobacter_sp_BacI18, K. pneumoniae_BacI20, L. soli_BacI39, Pantoea_sp_BacI23, and Pantoea_sp_BacI16 increased the plant length by more than 70% in HgCl2-supplemented substrate when compared with non-inoculated plants grown in the same substrate.
Growth rate of corn plants (Zea mays) inoculated with endophytic microorganisms and seeded in mercury-supplemented substrate. Data are presented as the mean ± standard deviation of four replicates of plants. CNHg = non-inoculated corn plants grown in substrate non-supplemented with mercury; CYHg = non-inoculated corn plants grown in mercury-supplemented substrate. *p < 0.05 (analysis of variance followed by the Dunnett’s test - control CYHg)
4 Discussion
We examined how mercury contamination influenced the diversity of endophytic bacteria in A. fluminensis (Asc) and P. acuminatum (Pol) roots. The predominance of these two plant species in the community grown in the selected HgY area suggests that they have developed mechanisms to limit soil mercury toxicity, and root endophytic fungi communities appear to play important roles (Pietro-Souza et al. 2017). It is also clear that, in the case of Asc and Pol, the roots host endophytic bacteria, regardless the site of plant collection. Endophytic bacteria colonize root tissues, can migrate to other plant organs (Jha et al. 2013), and play roles in plant adaptation and growth in contaminated soils (Afzal et al. 2017). Analysis of endophyte colonization of host plants growing in environments contaminated with elements such as arsenic, copper, chrome, nickel, and zinc has shown the presence of metal-resistant strains with potential in bioremediation (Sun et al. 2010; Fidalgo et al. 2016; Román-Ponce et al. 2016; Sánchez-López et al. 2018).
The endophytic bacteria population density in plants collected at HgY sites ranged from 5.72 × 105 to 3.27 × 105 CFU.g−1 of tissue, which is smaller than the range reported in the literature: 2.7 × 107 to 1.2 × 108 CFU.g−1 (Pérez et al. 2016). Plant roots collected at HgY sites had greater colonization frequency and richness than plant roots collected at HgN sites. Such variations indicate that environment composition determines the associated community more strongly than the plant species (Teixeira et al. 2010). The host roots usually have greater richness and diversity of endophytes than the leaves, bark, flowers, and fruits (Gaiero et al. 2013), which are determined by edaphic factors (Hardoim et al. 2008). Soil mercury contamination increased the richness of the root endophytic bacterial community as has been shown for fungal communities (Pietro-Souza et al. 2017).
The high soil mercury concentration influenced the diversity and structure of endophytic bacterial communities (Figs. 1and 2), corroborating another report on the composition and diversity of endophytes (Sun et al. 2010). The interaction among endophytes, host plants, and the environment promotes diversity variation, increases richness, and provides competitive advantages to the host plant over native species (Mallon et al. 2015).
Analysis of the interaction network identified alterations in the co-occurrence patterns from microorganisms undergoing different treatments (Long et al. 2018). The most compact and complex networks – from Pol and HgY areas – indicate that the species keep a microbial community that is more stable, with strong correlation and that respond to mercury presence in the environment (Stegen et al. 2012; Jiao et al. 2016). The networks from Asc and HgN areas maintained weaker interspecific cooperation, which can be associated with the lower number of positive correlations found among the analysed species.
Isolation of cultivable bacteria represents only a small fraction of the real diversity that exists in the plant (Tanaka et al. 2014; Fidalgo et al. 2016). Actinobacteria, Bacteriodetes, Firmicutes, and Proteobacteria were the predominant phyla in Asc and Pol roots (Table 1). These phyla are often associated with the two host plants studied (Pereira and PML 2014; Maida et al. 2015; Maropola and Ramond 2015; Fidalgo et al. 2016; Román-Ponce et al. 2016; Sánchez-López et al. 2018), including those growing in metal-contaminated environments (Luo et al. 2011; Mesa et al. 2017; Durand et al. 2018; Gu et al. 2018). The species Bacillus_cereus_X, Burkholderia_cepacia_BacI47, and Enterobacter_cloacae_X were detected with high abundance; they are usually found in endophytic bacterial communities. The exclusive presence of the genus Enterobacter in communities from HgY areas (Table 1) was probably associated with mercury resistance mechanisms (Mosa et al. 2016) (see Table S1) that could include increasing the solubility, reducing or oxidizing the metal to less toxic forms (Mani and Kumar 2014).
Endophytic bacteria isolated from host plants collected at HgY sites produce more plant growth-promoting functional traits than those collected at HgN sites (Table 3; Table S1). Bacillus, Burkkolderia, Enterobacter, Klebsiella, Pantoea, and Pseudomonas are bacterial genera that bear a variety of plant growth-promoting functional traits (da Costa et al. 2014; Ullah et al. 2015; Meng et al. 2015).
Endophytic bacteria that carry plant growth-promoting traits and are resistant to metals can be used to enhance plant growth (Sun et al. 2010). It is noteworthy that 60.47% of the isolated bacterial strains mitigated mercury toxicity in corn plants (Fig. 4). Our data and those of others show that host plants and endophytes probably established a mutualistic symbiotic relationship that increases plant growth in the presence of mercury (Rodriguez et al. 2008). In conclusion, we demonstrated that mercury-resistant endophytic bacterial strains – especially Pantoea sp_BacI23 – promote host plant growth. However, further research on mercury remediation under field conditions, as well as the elucidation of resistance mechanisms are still required.
References
Afzal S, Begum N, Zhao H, Fang Z, Lou L, Cai Q (2017) Influence of endophytic root bacteria on the growth cadmium tolerance and uptake of switchgrass (Panicum virgatum L.). J Appl Microbiol 123:498–510. https://doi.org/10.1111/jam.13505
Alanoca L, Amouroux D, Monperrus M, Tessier E, Goni M (2016) Diurnal variability and biogeochemical reactivity of mercury species in an extreme high-altitude lake ecosystem of the Bolivian Altiplano. Environ Sci Pollut Res 23:6919–6933. https://doi.org/10.1007/s11356-015-5917-1
Almoneafy AA, Kakar KU, Nawaz Z, Li B, Chun-lan Y, Xie GL (2014) Tomato plant growth promotion and antibacterial related-mechanisms of four rhizobacterial Bacillus strains against Ralstonia solanacearum. Symbiosis 63:59–70. https://doi.org/10.1007/s13199-014-0288-9
Alvares CA, Stape JL, Sentelhas PC, De Moraes Gonçalves JL, Sparovek G (2013) Kooppen’s climate classification map for Brazil. Meteorol Zeitschrift 22:711–728. https://doi.org/10.1127/0941-2948/2013/0507
Asaduzzaman A, Riccardi D, Afaneh AT, Cooper SJ, Smith JC, Wang F, Schreckenbach G (2019). Environmental mercury chemistry–in silico. Acc Chem Res. 52(2):379-388. https://doi.org/10.1021/acs.Accounts.8b00454
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. Paper Presented at the International AAAI Conference on Weblogs and Social Media AAAI 8:361–362
Blondel VD, Guillaume JL, Lambiotte R, Lefebvre E (2008) Fast unfolding of communities in large networks. J Stat Mech Theory Exp 10:P10008. https://doi.org/10.1088/1742-5468/2008/10/P10008
Brasil, Ministerio de Meio Ambiente. Comissão Nacional do Meio Ambiente, Conama. Resolução 420 - Estabelecimento de valores orientadores para elementos-traço em solos e água subterrânea. Brasília, 2009. http://www.mma.gov.br/port/conama/legiabre.cfm?codlegi=620/. Accessed 10 October 2018
Cabral L, Giovanella P, Gianello C, Bento FM, Andreazza R, Camargo FAO (2013) Isolation and characterization of bacteria from mercury contaminated sites in Rio Grande do Sul, Brazil, and assessment of methylmercury removal capability of a Pseudomonas putida V1 strain. Biodegradation 24:319–331. https://doi.org/10.1007/s10532-012-9588-z
Carrasco-Gil S, Siebner H, Leduc DL, Webb SM, Milla R, Andrews JC, Hernández LE (2013) Mercury localization and speciation in plants grown hydroponically or in a natural environment. Environ Sci Technol 47(7):3082–3090. https://doi.org/10.1021/es303310t
Carrim AJI, Barbosa EC, Vieira JDG (2006) Enzymatic activity of endophytic bacterial isolates of Jacaranda decurrens Cham. (Carobinha-do-campo). Braz Arch Biol 49:353–359. https://doi.org/10.1590/S1516-89132006000400001
Cavalcante VA, Dobereiner J (1988) A new acid-tolerant nitrogen-fixing bacterium associated with sugarcane. Plant Soil 108:23–31. https://doi.org/10.1007/BF02370096
Cebalho EC, Díez S, Santos Filho M, Muniz CC, Lázaro W, Malm O, Ignácio ARA (2017) Effects of small hydropower plants on mercury concentrations in fish. Environ Sci Pollut Res 24:22709–22716. https://doi.org/10.1007/s11356-017-9747-1
Ceccatto AP, Testoni MC, Ignácio AR, Santos-Filho M, Malm O, Díez S (2016) Mercury distribution in organs of fish species and the associated risk in traditional subsistence villagers of the Pantanal wetland. Environ Geochem Health 38(3):713–722. https://doi.org/10.1007/s10653-015-9754-4
Cuzzi C, Link S, Vilani A, Onofre SB (2011) Enzimas extracelulares produzidas por fungos endofíticos isolados de Baccharis Dracunculifolia DC (Asteraeceae). Glob Sci Technol 4:47–57 ISSN 1984-3801
da Costa PB, Granada CE, Ambrosini A, Moreira F, De Souza R, Dos Passos JFM, Arruda L, Passaglia LMP (2014) A model to explain plant growth promotion traits: a multivariate analysis of 2,211 bacterial isolates. PLoS One 9:1–25. https://doi.org/10.1371/journal.pone.0116020
Darko G, Azanu D, Logo NK (2016) Accumulation of toxic metals in fish raised from sewage-fed aquaculture and estimated health risks associated with their consumption. Cogent Environ Sci 2:1–37. https://doi.org/10.1080/23311843.2016.1190116
de Souza LT, Cnossen-Fassoni A, Pereira OL, Mizubuti ESG, de Araújo EF, de Queiroz MV (2013) Novel and highly diverse fungal endophytes in soybean revealed by the consortium of two different techniques. J Microbiol 51:56–69. https://doi.org/10.1007/s12275-013-2356-x
Dixit R, Wasiullah E, Malaviya D, Pandiyan K, Singh U, Sahu A, Shukla R, Singh B, Rai J, Sharma P, Lade H, Paul D (2015) Bioremediation of heavy metals from soil and aquatic environment: an overview of principles and criteria of fundamental processes. Sustainability 7:2189–2212. https://doi.org/10.3390/su7022189
Duarte GSD, Pasa MC (2016) Agricultural biodiversity and ethnobotany at São Benedito community Poconé, Mato Grosso, vol 17. Interações (Campo Grande), Brazil, pp 247–256. https://doi.org/10.20435/1984042X2016208
Durand A, Maillard F, Alvarez-lopez V, Guinchard S, Bertheau C, Valot B, Blaudez D, Chalot M (2018) Bacterial diversity associated with poplar trees grown on a hg-contaminated site: community characterization and isolation of hg-resistant plant growth-promoting bacteria. Sci Total Environ 622:1165–1177. https://doi.org/10.1016/jscitotenv201712069
El-deeb B, Gherbawy Y, Hassan S (2012) Molecular characterization of endophytic bacteria from metal hyperaccumulator aquatic plant (Eichhornia crassipe) and its role in heavy metal removal. Geomicrobiol J 29(10):906–915. https://doi.org/10.1080/014904512011635764
Farias L, Fávaro DIT, Pessoa A, Aguiar JPL, Yuyama LKO (2012) Mercury and methylmercury concentration assessment in children’s hair from Manaus, Amazonas state, Brazil. Acta Amaz 42:279–286. https://doi.org/10.1590/S0044-59672012000200015
Fidalgo C, Henriques I, Rocha J, Tacão M, Alves A (2016) Culturable endophytic bacteria from the salt marsh plant Halimione portulacoides: phylogenetic diversity, functional characterization, and influence of metal(loid) contamination. Environ Sci Pollut Res 23:10200–10214. https://doi.org/10.1007/s11356-016-6208-1
Franchi E, Rolli E, Marasco R, Agazzi G, Borin S, Cosmina P, Pedron F, Rosellini I, Barbafieri M (2017) Phytoremediation of a multi contaminated soil: mercury and arsenic phytoextraction assisted by mobilizing agent and plant growth promoting bacteria. J Soils Sediments 17(5):1224–1236. https://doi.org/10.1007/s11368-015-1346-5
Gaiero JR, McCall CA, Thompson KA, Day NJ, Best AS, Dunfield KE (2013) Inside the root microbiome: bacterial root endophytes and plant growth promotion. Am J Bot 100:1738–1750. https://doi.org/10.3732/ajb.1200572
Glick BR (2015) Stress control and acc deaminase. Princ Plant-Microbe Interact Microbes Sustain Agric:257–264. https://doi.org/10.1007/978-3-319-08575-3_27
Gu Y, Wang Y, Sun Y, Zhao K, Xiang Q, Yu X, Chen Q (2018) Genetic diversity and characterization of arsenic-resistant endophytic bacteria isolated from Pteris vittata, an arsenic hyperaccumulator. BMC Microbiol 18:1–42. https://doi.org/10.1186/s12866-018-1184-x
Hardoim PR, van Overbeek LS, van Elsas JD (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends in microbiol 16(10):463–471. https://doi.org/10.1016/j.tim.2008.07.008
Harichová J, Edita K, Pangallo D, Peter F (2012) Structure analysis of bacterial community and their heavy-metal resistance determinants in the heavy-metal-contaminated soil sample. Biologia 67(6):1038–1048. https://doi.org/10.2478/s11756-012-0123-9
Harris RF, Sommers LE (1968) Plate-dilution frequency technique for assay of microbial. Ecology Appl Microbiol 16:330–334 ISSN: 0099-2240
Hill MO (1973) Diversity and evenness: a unifying notation and its consequences. Ecology 54:427–432. https://doi.org/10.2307/1934352
Hoagland DR, Arnon DI (1950) The water-culture method for growing plants without soils. Circular. Calif Agric Exp Stn 347(2nd edit)
Jha PN, Gupta G, Jha P, Mehrotra R (2013) Association of rhizospheric / endophytic bacteria with plants: a potential gateway to sustainable agriculture. Greener J Agric Sci 3:73–84
Jiao S, Chen W, Wang E, Wang J, Liu Z, Li Y (2016) Microbial succession in response to pollutants in batch-enrichment culture. Nat Publ Gr:1–11. https://doi.org/10.1038/srep21791
Junk WJ, Piedade MTF, Lourival R, Wittmann F, Kandus P, Lacerda LD, Bozelli RL, Esteves FA, Nunes da Cunha C, Maltchik L, Schöngart J, Schaeffer-Novelli Y, Agostinho AA (2014) Brazilian wetlands: their definition, delineation, and classification for research, sustainable management, and protection. Aquat Conserv Mar Freshw Ecosyst 24:5–22. https://doi.org/10.1002/aqc2386
Junk WJ, Piedade MT, Schoengart J, Wittmann F, da Cunha CN (2016) Brazilian wetlands: classification The Wetland Book: I: Structure and Function. Management and Methods:1–7. https://doi.org/10.1007/978-94-007-6172-8_333-1
Köppen W (1930) Geiger R Handbuch der klimatologie. Berlin, Gebrüder Borntraeger
Lacerda LD, Pfeiffer WC, Marins RV, Rodrigues S, Souza CMM, Bastos WR (1991) Mercury dispersal in water, sediments and aquatic biota of a gold mining tailing deposit drainage in Poconé Brazil. Water Air Soil Pollut 55:283–294. https://doi.org/10.1007/BF00211194
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stachebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester
Long XE, Yao H, Huang Y, Wei W, Zhu YG (2018) Phosphate levels influence the utilisation of rice rhizodeposition carbon and the phosphate-solubilising microbial community in a paddy soil. Soil Biol Biochem 118:103–114. https://doi.org/10.1016/j.soilbio.2017.12.014
Lorck H (1948) Veterinary R. Production of hydrocyanic acid by bacteria Physiologia Plantarum 1:1–6. https://doi.org/10.1111/j1399-30541948tb07118x
Luo SL, Chen L, Chen JL, Xiao X, Xu TY, Wan Y, Rao C, Liu CB, Liu YT, Lai C (2011) Zeng GM. Analysis and characterization of cultivable heavy metal-resistant bacterial endophytes isolated from Cd-hyperaccumulator Solanum nigrum L and their potential use for phytoremediation Chemosphere 85:1130–1138. https://doi.org/10.1016/jchemosphere201107053
Maida I, Chiellini C, Mengoni A, Bosi E, Firenzuoli F, Fondi M, Fani R (2015) Antagonistic interactions between endophytic cultivable bacterial communities isolated from the medicinal plant Echinacea purpurea. Environ Microbiol 18(8):2357–2365. https://doi.org/10.1111/1462-292012911
Mallon CA, Van Elsas JD, Salles JF (2015) Microbial invasions: the process, patterns, and mechanisms trends. Microbiol 23:719–729. https://doi.org/10.1016/jtim201507013
Mani D, Kumar C (2014) Biotechnological advances in bioremediation of heavy metals contaminated ecosystems: an overview with special reference to phytoremediation international journal of environ Sci Technol 11(3):843–872. https://doi.org/10.1007/s13762-013-0299-8
Manohari R, Yogalakshmi KN (2016) Optimization of copper (II) removal by response surface methodology using root nodule endophytic bacteria isolated from Vigna unguiculata. Water, Air, & Soil Pollution 227-285. https://doi.org/10.1007/s11270-016-2964-2
Maropola MKA, Ramond JB (2015) Trindade M. Impact of metagenomic DNA extraction procedures on the identifiable endophytic bacterial diversity in Sorghum bicolor (L Moench) J Microbiol Methods 112:104–117. https://doi.org/10.1016/jmimet201503012
Mathew DC, Ho YN, Gicana RG, Mathew GM, Chien MC, Huang CC (2015) A rhizosphere-associated symbiont, Photobacterium spp strain MELD1, and its targeted synergistic activity for phytoprotection against mercury. PLoS One 10:1–18. https://doi.org/10.1371/journalpone0121178
Meng X, Bertani I, Abbruscato P, Piffanelli P, Licastro D, Wang C, Venturi V (2015) Draft genome sequence of rice endophyte-associated isolate Kosakonia oryzae KO348. Genome Announc 3:2164. https://doi.org/10.1128/genomeA00594-15
Mesa V, Navazas A, González-Gil R, González A, Weyens N, Lauga B, Gallego JLR, Sánchez J (2017) Use of endophytic and rhizosphere bacteria to improve phytoremediation of arsenic-contaminated industrial soils by autochthonous Betula celtiberica. Appl Env Microbiol 83:e03411–e03416. https://doi.org/10.1128/AEM03411-16
Milagres AMF, Machuca A, Napoleao D (1999) Methods detection of siderophore production from several fungi and bacteria by a modification of chrome azurol S (CAS) agar plate assay. J Microbiol Methods 37:1–6. https://doi.org/10.1016/S0167-7012(99)00028-7
Mosa KA, Saadoun I, Kumar K, Helmy M, Dhankher OP (2016) Potential biotechnological strategies for the cleanup of heavy metals and metalloids. Front Plant Sci 7:303. https://doi.org/10.3389/fpls201600303
Naik MM, Dubey SK (2016) Marine pollution and microbial remediation springer https://doi.org/10.1007/978-981-10-1044-6
Nieboer E, Richardson DHS (1980) The replacement of the nondescript term ‘heavy metals’ by a biologically and chemically significant classification of metal ions. Environmental Pollution Series B, Chemical and Physical 1 (1):3-26
Oliveira KF, Lacerda LD, Peres TF, Bezerra MF, da Silva Dias FJ (2015) Emission factor and balance of mercury in fish farms in an artificial reservoir in NE Brazil. Environ Sci Pollut Res 22:18278–18287. https://doi.org/10.1007/s11356-015-5102-6
Pandey PK, Samanta R, Narain R, Yadav S (2015) Plant beneficial endophytic bacteria from the ethnomedicinal Mussaenda roxburghii (Akshap) of eastern Himalayan Province, India Hindawi Publ Corp ID 580510:1–8. https://doi.org/10.1155/2015/580510
Pereira SIA, PML C (2014) Diversity and characterization of culturable bacterial endophytes from Zea mays and their potential as plant growth-promoting agents in metal-degraded soils. Environ Sci Pollut Res Int 21:14110–14123. https://doi.org/10.1007/s11356-014-3309-6
Pérez A, Martínez D, Barraza Z, Marrugo J (2016) Bacterias endófitas asociadas a los géneros Cyperus y Paspalum en suelos contaminados con mercurio Revista UDCA. Actualidad Divulgación Científica 19(1):67–76 ISSN 0123-4226
Pietro-Souza W, Mello IS, Vendruscullo SJ, GFd S, CNd C, White JF, Soares MA (2017) Endophytic fungal communities of Polygonum acuminatum and Aeschynomene fluminensis are influenced by soil mercury contamination. PLoS ONE 12:e0182017. https://doi.org/10.1371/journalpone0182017
Podile A, Kishore G (2007) Plant growth-promoting rhizobacteria Gnanamanickam SS plant-associated Bact springer, Netherlands, pp 195–230. https://doi.org/10.1094/Phyto-71-642
Rodriguez RJ, Henson J, van Volkenburgh E, Hoy M, Wright L, Beckwith F, Redman RS (2008) Stress tolerance in plants via habitat-adapted symbiosis. The ISME journal 2(4):404–416. https://doi.org/10.1038/ismej2007106
Román-Ponce B, Ramos-Garza J, Vásquez-Murrieta MS, Rivera-Orduña FN, Chen WF, Yan J, Estrada-de los Santos P, Wang ET (2016) Cultivable endophytic bacteria from heavy metal(loid)-tolerant plants. Arch Microbiol 198:941–956. https://doi.org/10.1007/s00203-016-1252-2
Sánchez-López AS, Thijs S, Beckers B, González-Chávez MC, Weyens N, Carrillo-González R, Vangronsveld J (2018) Community structure and diversity of endophytic bacteria in seeds of three consecutive generations of Crotalaria pumila growing on metal mine residues. Plant Soil 422:51–66. https://doi.org/10.1007/s11104-017-3176-2
Schulz B, Boyle C (2005) The endophytic continuum. Mycol Res 109:661–686. https://doi.org/10.1017/S095375620500273X
Seccatore J, Veiga M, Origliasso C, Marin T, De Tomi G (2014) An estimation of the artisanal small-scale production of gold in the world. Sci Total Environ 496:662–667. https://doi.org/10.1016/jscitotenv201405003
Shen M, Liu L, Li D, Zhou W, Zhou Z, Zhang C, Luo Y, Wang H, Li H (2013) The effect of endophytic Peyronellaea from heavy metal-contaminated and uncontaminated sites on maize growth, heavy metal absorption and accumulation fungal. Ecol 6:539–545. https://doi.org/10.1016/jfuneco201308001
Shinwari KI, Shah AU, Afridi MI, Zeeshan M, Hussain H, Hussain J, Ahmad O (2015) Application of plant growth promoting rhizobacteria in bioremediation of heavy metal polluted soil. Asian J Multidisc Stud 3(4):179–185
Soares MA, Li H, Bergen M, Manoel J, Kowalski KP, White JF (2016a) Functional role of an endophytic Bacillus amyloliquefaciens in enhancing growth and disease protection of invasive English ivy (Hedera helix L) Plant Soil 405(1-2):107–123. https://doi.org/10.1007/s11104-015-2638-7
Soares MA, Li HY, Kowalski KP, Bergen M, Torres MS, White JF (2016b) Functional role of bacteria from invasive Phragmites australis in promotion of host growth. Microb Ecol 72:407–417. https://doi.org/10.1007/s00248-016-0793-x
Soares MA, Li HY, Kowalski KP, Bergen M, Torres MS, White JF (2016c) Evaluation of the functional roles of fungal endophytes of Phragmites australis from high saline and low saline habitats. Biol Invasions 18:2689–2702. https://doi.org/10.1007/s10530-016-1160-z
Stegen JC, Lin X, Konopka AE, Fredrickson JK (2012) Stochastic and deterministic assembly processes in subsurface microbial communities. ISME J 6:1653–1664. https://doi.org/10.1038/ismej201222
Sun L, Zhang Y, He L, Chen Z, Wang Q, Qian M, Sheng X (2010) Genetic diversity and characterization of heavy metal-resistant-endophytic bacteria from two copper-tolerant plant species on copper mine wasteland. Bioresour Technol 101:501–509. https://doi.org/10.1016/jbiortech200908011
Tanaka T, Kawasaki K, Daimon S, Kitagawa W, Yamamoto K, Tamaki H, Kamagata Y (2014) A hidden pitfall in the preparation of agar media undermines microorganism cultivability. Appl Environ Microbiol 80(24):7659–7666. https://doi.org/10.1128/AEM.02741-14
Teixeira LC, Peixoto RS, Cury JC, Sul WJ, Pellizari VH, Tiedje J, Rosado AS (2010) Bacterial diversity in rhizosphere soil from Antarctic vascular plants of Admiralty Bay, maritime Antarctica. ISME J 4:989–1001. https://doi.org/10.1038/ismej201035
Ullah A, Heng S, Munis MFH, Fahad S, Yang X (2015) Phytoremediation of heavy metals assisted by plant growth promoting (PGP) bacteria: a review. Environ Exp Bot 117:28–40. https://doi.org/10.1016/jenvexpbot201505001
Vandamme P, Giesendorf BAJ, Van Belkum A, Pierard D, Lauwers S, Kersters K, Butzler J-P, Goossens H, Quint WGV (1993) Discrimination of epidemic and sporadic isolates of Arcobacter butzleri by polymerase chain reaction-mediated DNA fingerprinting. J Clin Microbiol 31:3317–3319 0095-1137/93/123317-03$02.00/0
Vishnivetskaya TA, Mosher JJ, Palumbo AV, Yang ZK, Podar M, Brown SD, Brooks SC, Gu B, Southworth GR, Drake MM, Brandt CC, Elias DA (2011) Mercury and other heavy metals influence bacterial community structure in contaminated Tennessee streams. J Appl Environ Microbiol 77:302–311. https://doi.org/10.1128/AEM01715-10
White JF, Torres MS, Johnson H, Irizarry I, Tadych M (2014) A functional view of plant microbiomes: endosymbiotic systems that enhance plant growth and survival in: Verma V, Gange a (eds) advances in endophytic research 425-439 springer. New Delhi. https://doi.org/10.1007/978-81-322-1575-2_21
Xie P, Hao X, Herzberg M, Luo Y, Nies DH, Wei G (2015) Genomic analyses of metal resistance genes in three plant growth promoting bacteria of legume plants in northwest mine tailings. China J Environ Sci (China) 27:179–187. https://doi.org/10.1016/jjes201407017
Yu Z, Li J, Li Y, Wang Q, Zhai X, Wu G, Liu P, Li X (2014) A mer operon confers mercury reduction in a Staphylococcus epidermidis strain isolated from Lanzhou reach of the Yellow River. Int Biodeterior Biodegrad 90:57–63. https://doi.org/10.1016/jibiod201402002
Zhang Y, Liu J, Zhou Y, Gong T, Wang J, Ge Y (2013) Enhanced phytoremediation of mixed heavy metal (mercury)-organic pollutants (trichloroethylene) with transgenic alfalfa co-expressing glutathione S-transferase and human P450 2E1. J Hazard Mater 260:1100–1107. https://doi.org/10.1016/jjhazmat201306065
Zhang MQ, Guo Y, Powell CA, Doud MS, Yang CY, Zhou H, Duan YP (2016) Zinc treatment increases the titre of “Candidatus Liberibacter asiaticus” in huanglongbing-affected citrus plants while affecting the bacterial microbiomes. J Appl Microbiol 120:1616–1628. https://doi.org/10.1111/jam13102
Funding
This work was supported by grants from Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq, grant 409,062/2018–9) and Fundação de Amparo à Pesquisa do Estado de Mato Grosso (FAPEMAT, grant 568,258/2014) to Marcos Antônio Soares, Ph.D.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 55 kb)
Rights and permissions
About this article
Cite this article
Mello, I.S., Pietro-Souza, W., Barros, B.M. et al. Endophytic bacteria mitigate mercury toxicity to host plants. Symbiosis 79, 251–262 (2019). https://doi.org/10.1007/s13199-019-00644-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13199-019-00644-0