Abstract
Wastewater represents the main reusable water source after being adequately sanitized by wastewater treatment plants (WWTPs). In this sense, only bacterial quality indicators are usually checked to this end, and human pathogenic viruses usually escape from both sanitization procedures and controls, posing a health risk on the use of effluent waters. In this study, we evaluated a protocol based on aluminum adsorption–precipitation to concentrate several human enteric viruses, including norovirus genogroup I (NoV GI), NoV GII, hepatitis A virus (HAV), astrovirus (HAstV), and rotavirus (RV), with limits of detection of 4.08, 4.64, 5.46 log genomic copies (gc)/L, 3.31, and 5.41 log PCR units (PCRU)/L, respectively. Furthermore, the method was applied in two independent laboratories to monitor the presence of NoV GI, NoV GII, and HAV in effluent and influent waters collected from five WWTPs at two different sampling dates. Concomitantly, a viability PMAxx-RT-qPCR was applied to all the samples to get information on the potential infectivity of both influent and effluent waters. The ranges of the titers in influent waters for NoV GI, NoV GII, RV, and HAstV were 4.80–7.56, 5.19–7.31 log gc/L, 5.41–6.52, and 4.59–7.33 log PCRU/L, respectively. In effluent waters, the titers ranged between 4.08 and 6.27, 4.64 and 6.08 log gc/L, < 5.51, and between 3.31 and 5.58 log PCRU/L. Moreover, the viral titers detected by viability RT-qPCR showed statistical differences with RT-qPCR alone, suggesting the potential viral infectivity of the samples despite some observed reductions. The proposed method could be applied in ill-equipped laboratories, due to the lack of a requirement for a specific apparatus (i.e., ultracentrifuge).
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Water represents the primary production source for almost all agricultural and industrial processes. Given the limitation of water resources and the global climate changes, many concerns have been raised in the recent decades to reduce its use and develop strategies to reuse it. In this sense, wastewater has been pointed out as the main reusable water source, and wastewater treatment plants (WWTPs) are the most common systems used for wastewater management worldwide, where influent waters are treated and effluent waters are intended to be reused (Becerra-Castro et al. 2015; Haramoto et al. 2018). Thereby, wastewater discharge to the environment or its reuse after sanitization procedures poses a great issue, given the increasing public health risk of human infections and illnesses associated with viral contamination of drinking water, coastal waters (i.e., shellfish), irrigation waters (i.e., vegetables and berries), and recreational activities (e.g., swimming). In fact, human enteric viruses are at high concentrations in feces excreted by infected people and, hence, can be present in untreated and treated wastewaters (Sano et al. 2016).
While physical and chemical parameters, together with microbiological indicators (i.e., fecal indicator bacteria (FIB) and Escherichia coli counts), have been widely used to assess water quality and its foreseeable use, the presence of human enteric viruses has not been routinely considered to this purpose by legal authorities. The presence of human enteric viruses in effluent waters has been well documented (Sano et al. 2016; Gerba et al. 2018), posing a public health risk-related concern and questioning the efficiency of WWTP in virus removal (National Research Council 2012; Verbyla and Mihelcic 2015). Among others, the viruses most commonly detected in influent and effluent waters include adenovirus (AdV), enterovirus (EV), hepatitis A and E viruses (HAV and HEV), norovirus (NoV), sapovirus (SaV), astrovirus (HAstV), and rotavirus A (RV) (Ashbolt 2015). To the best of our knowledge, only the USA included caliciviruses, a virus family that comprises the genera, NoV and SaV, in the list of water contaminants that need to be regulated (EPA 2016).
Despite the agreement between the scientific community and governments on controlling the viral population in various types of waters, a state of uncertainty rests on the analytical methods for quantitation and, thus, on reduction levels required to ensure minimal risk to the exposed population (WHO 2017; Gerba et al. 2018). It is evident that methodologies to concentrate and quantitate human enteric viruses in environmental waters need to be improved to finally design suitable water reclamation systems. Cell-culture methods have been used for a long time to detect infectious enteric viruses in water samples. However, these methods are impaired by the low levels of environmental contamination, the availability of a single cell-culture system for each targeted virus, and by the absence of reliable cell-culture assays for some viruses (Hamza et al. 2011; Condit 2013; Gerba et al. 2018). Alternatively, molecular detection methods have emerged as rapid, sensitive, and reliable tools for enteric virus detection and quantitation in water samples (Katayama et al. 2008; Simmons and Xagoraraki 2011; Farkas et al. 2018). However, despite the huge progress in viral detection due to the development of molecular assays based on real-time polymerase chain reaction (qPCR), water concentration procedures are cumbersome, and most of them require special equipment (Nordgren et al. 2009). In addition, molecular-based methods cannot discriminate between inactivated and potentially infectious viruses. Thus, the use of viability markers has been incorporated into qPCR-based methods for assessing infectivity of enteric viruses in several types of water samples in the past few years (Parshionikar et al. 2010; Kim et al. 2011; Coudray-Meunier et al. 2013; Prevost et al. 2016; Randazzo et al. 2016; López-Gálvez et al. 2018; Randazzo et al. 2018a, b).
To overcome these issues, in this study, a rapid and user-friendly protocol based on aluminum adsorption–precipitation previously approved by the Standard Methods Committee (2011) was evaluated to recover, detect, and quantify NoV GI, NoV GII, and HAV in influent and effluent waters. Moreover, the limits of detection were determined for the above-mentioned viruses as well as for HAstV and RV in effluent waters.
Ultimately, an interlaboratory study involving two laboratories was designed to validate the proposed method. To reduce the uncertainty and control the factors influencing virus estimation, the following variables were included: (i) two different sampling dates (November 2017 and May 2018); (ii) five WWTP facilities applying different reclaiming treatments, and (iii) at least two sampling points (influent and effluent waters) for each WWTP. In addition, to obtain information on the potential infectivity of the samples, a viability treatment previously optimized for water samples (Randazzo et al. 2016; López-Gálvez et al. 2018; Randazzo et al. 2018a, b) was run in parallel for all the samples.
Materials and Methods
Viral Stocks
Feces positive for NoV GI, NoV GII, and HAstV (courtesy of Dr. Buesa from Hospital Clínico Universitario, University of Valencia, Spain) were resuspended (10%, wt/vol) in phosphate-buffered saline (PBS) containing 2 M NaNO3 (Panreac), 1% beef extract (Conda), and 0.1% Triton X-100 (Fisher Scientific) (pH 7.2), vortexed and centrifuged at 1000×g for 5 min. The supernatant was stored at − 80 °C in aliquots.
The cytopathogenic HM-175 strain of HAV (ATCC VR-1402), the human RV strain Wa (ATCC VR-2018), and mengovirus (CECT 100,000) were propagated in FRhK, MA-104, and HeLa cell monolayers, respectively. Semipurified stocks were thereafter produced in the same cells by low-speed centrifugations of infected cell lysates (3000×g for 20 min).
WWTPs and Sample Collection
Influent and effluent waters were collected from five different municipal wastewater treatment plants located in Valencia (Spain), a region with high population density where agriculture and food processing are important economic activities (Table 1). For each WWTP, grab samples were taken from the flow at the influent and effluent in November 2017 and May 2018. Water samples were immediately placed on ice, sent to laboratories, and immediately processed or, alternatively, stored at − 80 °C.
Comparison of Virus Concentration Methods in Influent Waters
Initially, the performance of two different concentration methods was evaluated in five influent water samples by an ultracentrifugation-based protocol (Method A) and by an aluminum hydroxide adsorption–precipitation procedure (Method B). For Method A, 35 mL of influent water was centrifuged at 140,000×g for 2 h 30 min at 4 °C. The pellet was incubated on ice for 30 min with 5 ml of 0.25 mol/L glycine buffer (pH 9.5) and then the solution neutralized with 5 mL of PBS 2X. Suspended solids were removed by centrifugation at 12,000×g for 15 min, and, viruses from supernatant were recovered by ultracentrifugation at 229,600×g for 1 h at 4 °C and, finally eluted in 1 mL of PBS (Rodríguez-Díaz et al. 2009).
For Method B, 35 mL of influent water was adjusted to pH 6.0 and Al(OH)3 precipitate formed by adding 1 part 0.9 N AlCl3 solution to 100 parts of sample. The pH was readjusted to 6.0, and the sample was mixed slowly for 15 min at room temperature. Then, viruses were collected by centrifugation at 1700×g for 20 min. The pellet was resuspended into 1.75 mL of 3% beef extract pH 7.4, and samples were shacked for 10 min at 150 rpm. The concentrate was recovered by centrifugation at 1900×g for 30 min and the pellet resuspended in 1 mL of PBS (“Standard Methods For the Examination of Water and Wastewater,” 2011). Moreover, to determine the efficacy of the procedures, and thus validate the results, water samples were spiked with approx. 106 PCRU of MgV as process control as suggested by the ISO 15216-1:2017 (2017) guidelines and Gerba et al. (2018). Experiments were performed in triplicate.
Detection Limits of Enteric Viruses in Effluent Waters
Effluent water samples were concentrated by Method B, and the limit of detection of each virus was determined. Experiments were performed in triplicate using 200 mL of effluent water samples that previously tested negative for the viruses under study. Water samples were artificially inoculated with different concentrations of viral inocula (approximately 6, 5, 4, and 3 log genomic copies (gc)/L) and concentrated according to Method B described in “Comparison of Virus Concentration Method in Influent Waters” section. The limit of detection (LoD95%) was calculated for each virus according to Wilrich and Wilrich (2009).
Interlaboratory Comparison Study of Influent and Effluent WWTP Waters
An interlaboratory study involving two laboratories was designed to validate the proposed method. Thus, Method B was applied by Lab1 and Lab2 to determine NoV GI, GII and HAV levels in influent and effluent water samples collected in November 2017 and May 2018 from five WWTPs. The two laboratories independently concentrated and analyzed single samples, and RT-qPCRs were run in duplicate. Moreover, to obtain information on the potential infectivity of the samples, a viability RT-qPCR was run in parallel for all the samples by both laboratories. In addition, to obtain further information on enteric virus population present in influent and effluent waters, Lab1 screened all influent and effluent waters for RV and HAstV by both RT-qPCR and viability RT-qPCR. For viability RT-qPCR, previously optimized protocol was applied (Randazzo et al. 2016; López-Gálvez et al. 2018; Randazzo et al. 2018b). In brief, the photoactivatable dye PMAxx™ (Biotium) was added to concentrated samples at 50 μM together with 7.7 mmol/L Triton 100-X (Fisher-Scientific) and incubated in the dark at room temperature for 10 min at 150 rpm. Then, samples, in DNA LoBind 1.5 mL tubes (Eppendorf), were exposed to photoactivation using a photoactivation system (Led-Active Blue, GenIUL) for 15 min, and RNA was extracted as described in “RNA Extraction and RT-qPCR” section.
RNA Extraction and RT-qPCR
RNA from water sample concentrates was extracted using the NucleoSpin® RNA virus kit (Macherey-Nagel GmbH & Co.) according to the manufacturer’s instructions including the Plant RNA Isolation Aid (Ambion) pretreatment as previously described (Randazzo et al. 2016; Randazzo et al. 2018b). Primers, probes, and reverse transcription-qPCR (RT-qPCR) conditions used in this study are listed in Table S1.
RT-qPCRs were carried out in 96-well plates using the RNA UltraSense One-Step quantitative RT-PCR system (Invitrogen SA) with a half-scale modification of the manufacturer’s protocol and the LightCycler 480 (Roche Diagnostics at Lab1) or QuantStudio 5 (Applied Biosystems at Lab2) instruments. Each viral RNA was analyzed at least in duplicate. Undiluted and tenfold diluted RNA was tested to determine inhibition of the RT-qPCRs. Different controls were used: positive and negative extractions and RT-qPCR controls, and MgV as a whole process control was spiked prior to concentration, and detected in downstream RT-qPCR (Gerba et al. 2018).
Virus Quantitation
Standard curves were determined according to the Public Health England (PHE) Reference Materials for Microbiology for NoV GI (batch number 0122-17), NoV GII (batch number 0247-17) and HAV (batch number 0261-2017), while standard curves for RV, MgV, and HAstV were generated by amplifying tenfold serial dilutions of viral suspensions in quintuplicates and calculating the numbers of PCR units (PCRU).
Statistical Analysis
To statistically compare the performance of the evaluated concentration methods, data were subjected to the analysis of variance (ANOVA) to test the impact of the method followed by the Tukey’s HSD as post hoc test to obtain homogenous groups (Table 2). Moreover, to explore the effects of the five variables considered (WWTP, sampling date, influent/ effluent water, use of PMAxx and laboratory facility), ANOVA was applied to the data matrix of quantitation values of each targeted virus. To this end, a hypothetical value of half of the detection limit was assigned to viral loads under the detection limit (SEPA 2008). Furthermore, a principal component analysis (PCA) explored the input matrix based on titers of NoV GI and GII replicates introduced as cases together with the explanatory variables WWTP, type of water (influent or effluent water), laboratory facility (Lab1 and Lab2), and viability marker pretreatment. Raw data were collected and preliminarily analyzed in Excel spreadsheets (Microsoft), statistical data processing was performed using STATISTICA software version 7 (StatSoft Inc., Tulsa, OK, USA), and graphic constructions executed on R (R Core Team 2014) by means of Scatterplot3d Package (Ligges and Mächler 2003). In all cases, values of p < 0.05 were deemed significant.
Results and Discussion
Comparison of Concentration Methods for Enteric Viruses from Influent Waters
Several protocols for the concentration, detection, and the quantitation of human enteric viruses from wastewater have been published (reviews by Cashdollar and Wymer 2013; Haramoto et al. 2018). The surveillance of water supplies for the presence of enteric viruses requires procedures that have a high likelihood for adaptation to different laboratory facilities around the world. Initially, an ultracentrifugation-based methodology (Method A) (Rodríguez-Díaz et al. 2009), which requires equipment which is not always available at the water analysis laboratories, and an AlCl3-based method (Method B) were compared by using naturally contaminated influent water samples. The titers of NoV GI, NoV GII, HAV, RV, and HAstV from five influent water samples using the two different methods are reported in Table 2, together with MgV recoveries. Viral concentrations ranged between 4.10 and 4.81 and 4.41 and 5.66 for NoV GI; and between 4.64 and 5.30 and 4.13 and 4.94 for NoV GII log gc/L for Methods A and B, respectively. HAV was detected in a total of 3 out of 5 samples at levels of < 5.46 log gc/L applying Method B, while only two positive samples resulted from Method A. RV titers ranged between 5.19 and 6.01 and 6.22 and 6.73 log PCRU/L for Methods A and B, respectively. Similarly, concentrations in the ranges of of 5.00–5.65 and 5.44-6.00 log PCRU/L were detected for HAstV by Methods A and B, respectively.
Significant differences (p < 0.05) were shown for NoV GI, NoV GII, HAV, RV, and HAstV for the two methods under comparison. In particular, slightly higher NoV GI, RV, and HAstV titers were reported for Method B than for Method A. On the contrary, a sharp pattern cannot be defined for NoV GII and HAV, since differences were detected in samples with the lowest viral titers, close to the detection limits, as in the cases of IW4 and IW5 for NoV GII and IW2 for HAV.
The recovery of spiked MgV was also determined, and greater efficiency was detected for Method A (8.04–25.72%) compared with Method B (0.02–4.30%) for all samples (n = 5). Only one sample, IW4, concentrated with Method B, did not comply with the recovery efficiency indicated in the ISO 15216-1:2017 to validate viral concentration in bottled water (> 1% of MgV recovery).
Despite the discordance on MgV recoveries, targeted viral quantitations were similar when comparing the two methods. An explication of such difference between the process control recoveries (MgV) and the similarity of targeted genome titers maybe related with the pH changes in Method B, which may interfere with MgV stability. These pH changes may not affect the viral population already present in the water. These viral particles are commonly aggregated among themselves and together with organic material finally resulting duly protected by pH changes (Gerba et al. 2017). Nevertheless, Table 4 shows that MgV recoveries analyzed by Lab1 reported higher recoveries, most likely due to being more familiar with the procedure. These are in line with the previously reported MgV recoveries in influent waters (Miura et al. 2016).
Moreover, when considering concentration methods, several factors (such as the volume of the sample, organic matters, and chemicals) can affect RT-qPCR outcomes by inhibiting RNA extractions and amplification steps (Ikner et al. 2011, 2012; Cashdollar and Wymer 2013; Borgmästars et al., 2017). Thus, tenfold diluted RNAs were also analyzed to check for inhibitors according to ISO 15216-1:2017, and the results confirmed the absence of interfering substances (data not shown). This approach excludes the possibility of false negative and/or subestimation due to complete or partial inhibition of PCRs, confirming the reliability of obtained results, especially in sensitive samples as influent and effluent waters.
To sum up, our results suggest that the aluminum-based procedure (Method B) is an alternative concentration method useful in the absence of an ultracentrifuge or when greater volumes of samples need to be processed, as in the case of effluent waters. Taking these reasons together, Method B was further used to determine the LoD95% in effluent waters and to compare two independent laboratories’s outcomes in analyzing naturally contaminated influent and effluent water samples.
Detection Limit of Enteric Viruses in Effluent Waters
Low virus levels in effluent water samples are a major analytic challenge, thus the LoD95% values were assessed. Method B resulted in LoD95% of 4.08, 4.64, 5.46, 3.31, and 5.41 log gc/L for NoV GI, NoV GII, HAV, HAstV, and RV, respectively, calculated according to the method of Wilrich and Wilrich (2009). Similar limits of detection have been previously reported for NoV GI and GII (around 4 log gc/L) applying an ultracentrifugation-based concentration method (Nordgren et al. 2009), suggesting the comparable efficacies of both methods.
Lower LoDs could have been reached by concentrating larger water volume, even though this may impair in coextraction of interfering substances that can affect RT-qPCR outcomes. As example, Hill et al. (2010), by concentrating 50 L ground water by ultrafiltration coupled to PEG precipitation, achieved lower LoDs for GI (200–3000 particles/L), but not for GII (1000–10,000 particles/L), while D’Ugo et al. (2016) was able to detect 4 log gc/L for NoV GII and HAV in 50 L river water samples. A wider comparison with other available studies is restricted by the fact that only RT-qPCR detection limits are usually reported, often excluding other molecular process controls (i.e., extraction control) and/or a whole process control (Haramoto et al. 2018).
Improvements on the LoDs can be also achieved by increasing the volume of concentrate extracted or the volume of the master-mix reaction, although the latter will increase the cost of the analysis.
Viral recoveries of 13.13, 7.52, 0.65, 71.40, and 0.97% were detected for NoV GI, NoV GII, HAV, HAstV, and RV, respectively, using the aluminum-based method. Given an acceptable level of viral recovery for the whole process of 1% (Haramoto et al. 2018), HAV and RV were not recovered efficiently (Table 3). This may be due to the heterogeneities of the RV and HAV suspensions given an estimation that 1 TCID50 corresponds to 50 genome copies and 1000 PCRU for HAV and RV, respectively. This would lead to an underestimation of the RV and HAV recoveries. Free RNA molecules and defective particles were detected in the positive control submitted only to the RNA extraction, whereas in water samples, those free RNAs and defective particles were most likely lost during the concentration steps.
Interlaboratory Study for the Detection and Quantitation of Potentially Infectious Enteric Viruses in Influent and Effluent Waters
The spread of qPCR has allowed the detection of human enteric viruses in environmental waters (Aw and Rose 2012), including those that cannot be detected by routine cell culture. Moreover, monitoring viruses in influent and effluent waters could be an appropriate approach for determining the prevalence, the epidemiology, and, finally, the associated human health risks (Sinclair et al. 2008; Prevost et al. 2015; Kazama et al. 2016, 2017). Thus, to further validate the proposed methodology, influent and effluent water samples were collected at each WWTP (n = 5, Table 1) in November 2017 and May 2018 and assayed by two independent laboratories (Lab1 and Lab2) to detect NoV GI, NoV GII, and HAV. Moreover, a viability RT-qPCR was run concomitantly to evaluate the potential infectivity of each sample (Table 4).
In influent samples, NoV GI ranged between 4.80 and 6.54 (for Lab1) and between 5.29 and 7.56 log gc/L (for Lab2). In effluent waters, titers were in the ranges of 4.08–6.01 (for Lab1) and 4.08–6.27 log gc/L (for Lab2), together with some samples resulting negative. NoV GII varied between 5.19 and 7.16 (Lab1) and 5.00 and 7.31 (Lab2) in influent water samples and between 4.64 and 5.43 (Lab1) and 4.72 and 6.08 log gc/L (Lab2) in effluent water samples. Only three samples were positive for HAV by Lab1 at quantitation values of < 5.46 log gc/L.
In influent waters, NoV GII showed, on average, slightly greater titers than NoV GI, similar to previous publications reporting ranges from 104 to 108 for NoV GI and from 105 to 109 gc/L for NoV GII (da Silva et al. 2007; Katayama et al. 2008; da Silva et al. 2008; La Rosa et al. 2010; Simmons and Xagoraraki 2011; Kitajima et al. 2014; Montazeri et al. 2015; Qiu et al. 2015; Schmitz et al. 2016; Haramoto et al. 2018).
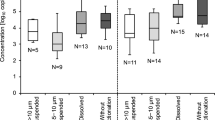
For better describing the viral population, Lab1 additionally determined the levels of RV and HAstV in influent and effluent waters (Fig. 1; Table 5). RV titers ranged between 5.51 and 6.52 log PCRU/L in influent waters resulting below the LoD95% (< 5.51 log PCRU/L) in effluent water samples. Similar concentrations (around 4.5 log gc/L) have been previously described in both influent and effluent water samples (Haramoto et al. 2018). Furthermore, levels of HAstV ranging between 4.59 and 7.33 and between 3.31 and 5.58 log PCRU/L were detected in influent and effluent waters, respectively.
Overview of detected enteric viruses in influent and effluent waters. Boxplots show median concentrations (log genome copies/L) with the 25th and 75th percentile values of NoV GI (blue), NoV GII (orange), RV (green), and HAstV (yellow) in influent (dark colors) and effluent (light colors) waters from five different wastewater treatment plants as detected by Lab1 (Color figure online)
As a general statement, given an efficiency of the concentration and extraction method below 100%, all of these values should be considered as a conservative estimation of the virus load, being the corrected titer as high as 1010 log genomic copies/L (Gerba et al. 2017). Recoveries of spiked whole process control were in the ranges of 3.6–40.9% (Lab1) and 3.2–66.6% (Lab2) (Table 4) suggesting that the method was suitable for viral concentration estimation (Haramoto et al. 2018; ISO 15216-1:2017). Overall, previous studies showed similar wide recovery ranges in surface and wastewaters using different viruses as process control such as MgV (Farkas et al. 2018; Miura et al. 2016), PP7 bacteriophage, or AdV (Kundu et al. 2013; Prevost et al. 2015; Barrios et al. 2018). Average recovery percentages ranging from 38 to 49% were obtained in different studies using adsorption–elution on electrocharged (either positive or negative) filters and summarized in the review by Cashdollar and Wymer (2013). Regardless, considering that the back-calculation is not recommended (Haramoto et al. 2018), all reported values have not been adjusted in this sense.
Comparing viral titers before and after the treatment at WWTP, differences were registered for both NoV GI and GII, as well for RV and HAstV. Reductions were observed in all samples at different extents with the highest removal rates shown by WWTP1, WWTP2, and WWTP4 (Fig. 1). Other authors reported viral reductions between influent and effluent waters due to WWTP treatments, suggesting a higher persistence of NoV GI compared to GII (Haramoto et al. 2006; da Silva et al. 2007; Nordgren et al. 2009; Haramoto et al. 2015). In contrast, Hewitt et al. (2011) found similar NoV GI and GII concentrations in influent and effluent water samples. Our results clearly suggest that reductions are mainly due to the type of reclamation treatments applied at WWTP (secondary vs. tertiary), given the different reductions registered among the WWTPs (Table 1).
In this sense, the determination of virus infectivity in waters is required to assess the real risk of using these waters. Thus, a viability RT-qPCR procedure was applied in parallel in both influent and effluent waters by both laboratories. Even if PMAxx showed to significantly affect the virus titers (Table S2), the high loads of viruses in the PMAxx-treated samples indicate the potential infectivity of detected viruses. Similarly, Gyawali and Hewitt (2018) evaluated the performance of PEMAX-RT-qPCR on influent and effluent waters and found a high proportion of potentially infectious NoV GI and GII (i.e., 21–89% in influent, and 24–59% in effluent).
On the contrary, a recent study by Prevost et al. (2016) demonstrated the suitability of EMA to differentiate infectious and inactivated NoV GI, NoV GII, RV, and HAstV in effluent waters from drinking water plants with UV and chlorination treatments in which none of the samples resulted positive following the viability RT-qPCR.
Our results are not surprising when taking into account that (i) in influent water, detected viruses originated from recently excreted feces, being expected as infectious; (ii) the viability RT-qPCR has been mainly fostered to discriminate heat-inactivated viruses, and is still not being optimized and validated with other inactivation techniques, such as chlorination or UV; (iii) altered, noninfectious viruses and their free RNAs degrade rapidly, being unlikely in their detection (Limsawat and Ohgaki 1997); and iv) the water concentration procedure most likely targets infectious viruses rather than free RNAs or altered capsids.
Thus, the infectivity profiles obtained by viability RT-qPCR may reflect either an ineffective virus inactivation at WWTP and/or that the capsid damage by secondary/tertiary water treatments may not be enough to allow viability marker penetration. In this sense, a recent study by López-Gálvez et al. (2018) demonstrated that chlorination with chlorine dioxide (ClO2) is not able to significantly reduce the NoV GI, NoV GII, and HAstV loads in effluent waters, according to previous research by Kingsley et al. (2014) that suggested that NoVs are quite resistant to ClO2. The differences in the final outcomes of such studies are justified by different parameters such as pH, turbidity, temperature, dissolved organic matter, and ionic strength, which all play a respective key role in inactivation kinetics, and especially in chlorination-sanitizing strategies (Carvajal et al. 2017; López-Gálvez et al. 2018).
Thus, the effectiveness of viability RT-qPCR should be investigated and validated for different inactivation mechanisms (i.e., UV treatment, chlorination, ozone treatment, etc.) for each virus of interest and for each type of water supply. Despite limitations, viability RT-qPCR may improve public health risk evaluations by providing more realistic datasets than RT-qPCR alone.
Variation is observed in viral titers independently detected by two laboratories (Table 4). Significant roles in determining such differences in measurements could have been played by the different levels of expertise in virus analysis among laboratories and homogeneity of the sample. A more robust comparison of the dataset generated in this study should take advantage of water-quality parameters (Borgmästars et al. 2017), that unfortunately were not available for the tested samples.
Statistical Analysis
The effects of the five variables considered in this interlaboratory comparison study (WWTP, sampling date, influent/ effluent water, use of PMAxx and laboratory facility) tested by ANOVA for each virus are shown in Table S2. The results, indicated as p values, show significant differences within the parameters of WWTP, the type of water (influent/ effluent water), and the use of viability marker (PMAxx) for all the investigated enteric viruses. The effect of the sampling date was significant for RV and HAstV, but not for NoV GI and GII. In this sense, NoVs fluctuation over the seasons has been described by Farkas et al. (2018) and Haramoto et al. (2006), but not always confirmed (Nordgren et al. 2009). Seasonal profiles of human caliciviruses concentrations in water environments are not surprising, since higher concentrations in colder months reflect the epidemic period for those viruses. Despite this, Katayama et al. (2008), Kitajima et al. (2014) reported that constant concentrations of AdV, EVs, and Aichi virus in influent water are without a clear seasonal pattern. Despite all these factors, the results of this study cannot support any robust conclusion of seasonal pattern due to the limited number of samples analyzed.
Regarding the effect of the laboratory facilities, significant differences were detected only for NoV GI, but not for NoV GII. A PCA was applied to the data matrix of NoV GI and GII titers, and the results are shown in Fig. 2; Tables S3 and S4. The correlation analysis among variables (Tables S3 and S4) showed significant relationships, and the data matrixes were appropriated to be subjected to the PCA to condense the information within factors. Thus, the three main factors, representing up to 72.11 and 73.66% of the total variance for NoV GI and GII, respectively, were plotted in a 3D scatterplot showing the relationship among them and samples. For NoV GI (Fig. 2a), samples were grouped mainly according to Factor1 and Factor2, correlated with viability treatment, laboratory facilities, and WWTPs. Factor3 contributed only marginally to discriminate samples according to the type of water, influent or effluent. For NoV GII (Fig. 2b), all the factors contributed to represent the total variance resulting in a wide spread of samples in the plot. In particular, Factor1 and Factor2 mainly discriminated samples being correlated with WWTP, viability treatment, and type of water.
Scatterplots show relationship between Factors and samples based on principal component analysis of the detected titers of NoV GI (a) and NoV GII (b) in influent and effluent water samples by two independent laboratory facilities. Influent (filled square, open square, filled diamond, open diamond) and effluent (filled circle, open circle, filled triangle, open triangle) water samples tested by Lab1 (filled square, open square, filled circle, open circle) and Lab2 (filled diamond, open diamond, filled triangle, open triangle) with (non-solid symbols) and without (solid symbols) viability marker pretreatment. Symbols are colored according to wastewater treatment plant (WWTP1 in red, WWTP2 in blue, WWTP3 in green, WWTP4 in yellow, WWTP5 in black) (Color figure online)
Finally, the PCA estimations of titers of both NoV GI and GII showed that the main factors that contributed in discriminating samples were correlated to WWTP and type of water.
Conclusions
The demand for rapid and reliable protocols to define virological water quality is increasing due to the spread of alternative uses of regenerated/recycled water given the limitation of water resources. In such a scenario, we tested a rapid and low-cost method based on aluminum hydroxide adsorption–precipitation. We determined the enteric viral pathogen population, including NoV GI, NoV GII, HAV, RV, and HAstV, in influent and effluent water samples. Even MgV recoveries varied greatly across samples, the method, evaluated by an interlaboratory comparative study, provided good mean recovery of the process control virus in both influent and effluent waters. Titers of up to 7 log gc/L were detected for NoV GI and GII in influent waters, in line with previous studies. Moreover, three samples were positive for HAV. In general, reductions in viral loads were detected in effluent water samples compared to influent waters, posing a health risk concern since the viability pretreatment applied showed their potential viral infectivity.
A PCA analysis applied to NoVs data matrix confirmed the correlations among WWTP, type of water, viability pretreatment, and sample variability. Thus, despite the limitations, this proposed approach could be useful not only to control virus loads in influent and effluent water samples, but also to evaluate the efficacy of sanitation procedures applied in WWTPs and, thus, to better predict the risk by quantitative microbial risk assessment (QMRA) analysis (Van Abel et al. 2017; Dias et al. 2019).
However, further improvements should be considered before adapting the method for routine use, such as lowering the limit of detections by analyzing full-scale RT-qPCR and/or validating the viability PMAxx-RT-qPCRs with viral inactivation by UV treatment and chlorination.
References
Ashbolt, N. J. (2015). Microbial contamination of drinking water and human health from community water systems. Current Environmental Health Reports,2, 95–106. https://doi.org/10.1007/s40572-014-0037-5.
Aw, T. G., & Rose, J. B. (2012). Detection of pathogens in water: From phylochips to qPCR to pyrosequencing. Current Opinion in Biotechnology,23(3), 422–430. https://doi.org/10.1016/j.copbio.2011.11.016.
Barrios, M. E., Blanco Fernández, M. D., Cammarata, R. V., Torres, C., & Mbayed, V. A. (2018). Viral tools for detection of fecal contamination and microbial source tracking in wastewater from food industries and domestic sewage. Journal of Virological Methods,262, 79–88. https://doi.org/10.1016/j.jviromet.2018.10.002.
Becerra-Castro, C., Lopes, A. R., Vaz-Moreira, I., Silva, E. F., Manaia, C. M., & Nunes, O. C. (2015). Wastewater reuse in irrigation: A microbiological perspective on implications in soil fertility and human and environmental health. Environment International,75, 117–135. https://doi.org/10.1016/j.envint.2014.11.001.
Borgmästars, E., Jazi, M. M., Persson, S., Jansson, L., Rådström, P., Simonsson, M., et al. (2017). Improved detection of norovirus and hepatitis A virus in surface water by applying pre-PCR processing. Food and Environmental Virology,9(4), 395–405.
Carvajal, G., Roser, D. J., Sisson, S. A., Keegan, A., & Khan, S. J. (2017). Bayesian belief network modelling of chlorine disinfection for human pathogenic viruses in municipal wastewater. Water Research,109, 144–154. https://doi.org/10.1016/j.watres.2016.11.008.
Cashdollar, J. L., & Wymer, L. (2013). Methods for primary concentration of viruses from water samples: A review and meta-analysis of recent studies. Journal of Applied Microbiology,115(1), 1–11. https://doi.org/10.1111/jam.12143.
Condit, R. C. (2013). Principles of virology. In D. M. Knipe & P. M. Howley (Eds.), Fields virology (pp. 21–25). Philadelphia: Wolters Klewer/Lippincott Williams & Wilkins.
Coudray-Meunier, C., Fraisse, A., Martin-Latil, S., Guillier, L., & Perelle, S. (2013). Discrimination of infectious hepatitis A virus and rotavirus by combining dyes and surfactants with RT-qPCR. BMC Microbiology,13(1), 216. https://doi.org/10.1186/1471-2180-13-216.
D’Ugo, E., Marcheggiani, S., Fioramonti, I., Giuseppetti, R., Spurio, R., Helmi, K., et al. (2016). Detection of human enteric viruses in freshwater from European countries. Food and Environmental Virology,8(3), 206–214.
da Silva, Le, Guyader, F. S., Le Saux, J. C., Pommepuy, M., Montgomery, M. A., & Elimelech, M. (2008). Norovirus removal and particle association in a waste stabilization pond. Environmental Science and Technology,42(24), 9151–9157. https://doi.org/10.1021/es802787v.
da Silva, Le, Saux, J.-C., Parnaudeau, S., Pommepuy, M., Elimelech, M., & Le Guyader, F. S. (2007). Evaluation of removal of noroviruses during wastewater treatment, using real-time reverse transcription-PCR: Different behaviors of genogroups I and II. Applied and Environmental Microbiology,73(24), 7891–7897. https://doi.org/10.1128/aem.01428-07.
Dias, E., Ebdon, J., & Taylor, H. (2019). Estimating the concentration of viral pathogens and indicator organisms in the final effluent of wastewater treatment processes using stochastic modelling. Microbial Risk Analysis,11, 47–56. https://doi.org/10.1016/j.mran.2018.08.003.
EPA. (2016). Drinking Water Contaminant Candidate List 4-Final. (2016-27667). Retrieved from https://www.federalregister.gov/documents/2016/11/17/2016-27667/drinking-water-contaminant-candidate-list-4-final.
Farkas, K., Marshall, M., Cooper, D., McDonald, J. E., Malham, S. K., Peters, D. E., et al. (2018). Seasonal and diurnal surveillance of treated and untreated wastewater for human enteric viruses. Environmental Science and Pollution Research,25(33), 33391–33401. https://doi.org/10.1007/s11356-018-3261-y.
Gerba, C. P., Betancourt, W. Q., & Kitajima, M. (2017). How much reduction of virus is needed for recycled water: A continuous changing need for assessment? Water Research,108, 25–31. https://doi.org/10.1016/j.watres.2016.11.020.
Gerba, C. P., Betancourt, W. Q., Kitajima, M., & Rock, C. M. (2018). Reducing uncertainty in estimating virus reduction by advanced water treatment processes. Water Research,133, 282–288. https://doi.org/10.1016/j.watres.2018.01.044.
Gyawali, P., & Hewitt, J. (2018). Detection of infectious noroviruses from wastewater and seawater using PEMAXTM treatment combined with RT-qPCR. Water,10(7), 841. https://doi.org/10.3390/w10070841.
Hamza, I. A., Jurzik, L., Überla, K., & Wilhelm, M. (2011). Methods to detect infectious human enteric viruses in environmental water samples. International Journal of Hygiene and Environmental Health,214, 424–436. https://doi.org/10.1016/j.ijheh.2011.07.014.
Haramoto, E., Fujino, S., & Otagiri, M. (2015). Distinct behaviors of infectious F-specific RNA coliphage genogroups at a wastewater treatment plant. Science of the Total Environment,520, 32–38. https://doi.org/10.1016/j.scitotenv.2015.03.034.
Haramoto, E., Katayama, H., Oguma, K., Yamashita, H., Tajima, A., Nakajima, H., et al. (2006). Seasonal profiles of human noroviruses and indicator bacteria in a wastewater treatment plant in Tokyo, Japan. Water Science and Technology,54(11–12), 301–308. https://doi.org/10.2166/wst.2006.888.
Haramoto, E., Kitajima, M., Hata, A., Torrey, J. R., Masago, Y., Sano, D., et al. (2018). A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Research,135, 168–186. https://doi.org/10.1016/j.watres.2018.02.004.
Hewitt, J., Leonard, M., Greening, G. E., & Lewis, G. D. (2011). Influence of wastewater treatment process and the population size on human virus profiles in wastewater. Water Research,45(18), 6267–6276. https://doi.org/10.1016/j.watres.2011.09.029.
Hill, V. R., Mull, B., Jothikumar, N., Ferdinand, K., & Vinjé, J. (2010). Detection of GI and GII noroviruses in ground water using ultrafiltration and TaqMan real-time RT-PCR. Food and Environmental Virology,2(4), 218–224.
Ikner, L. A., Gerba, C. P., & Bright, K. R. (2012). Concentration and recovery of viruses from water: A comprehensive review. Food and Environmental Virology,4(2), 41–67. https://doi.org/10.1007/s12560-012-9080-2.
Ikner, L. A., Soto-Beltran, M., & Bright, K. R. (2011). New method using a positively charged microporous filter and ultrafiltration for concentration of viruses from tap water. Applied and Environmental Microbiology,77(10), 3500–3506. https://doi.org/10.1128/AEM.02705-10.
ISO 15216–1:2017. (2017). Microbiology of food and animal feed—horizontal method for determination of Hepatitis A virus and norovirus in food using real-time RT-PCR—Part 1: Method for quantification. Geneva: Switzerland.
Katayama, H., Haramoto, E., Oguma, K., Yamashita, H., Tajima, A., Nakajima, H., et al. (2008). One-year monthly quantitative survey of noroviruses, enteroviruses, and adenoviruses in wastewater collected from six plants in Japan. Water Research,42(6–7), 1441–1448. https://doi.org/10.1016/j.watres.2007.10.029.
Kazama, S., Masago, Y., Tohma, K., Souma, N., Imagawa, T., Suzuki, A., et al. (2016). Temporal dynamics of norovirus determined through monitoring of municipal wastewater by pyrosequencing and virological surveillance of gastroenteritis cases. Water Research,92, 244–253. https://doi.org/10.1016/j.watres.2015.10.024.
Kazama, S., Miura, T., Masago, Y., Konta, Y., Tohma, K., Manaka, T., et al. (2017). Environmental surveillance of norovirus genogroups I and II for sensitive detection of epidemic variants. Applied and Environmental Microbiology,83(9), e03406-16. https://doi.org/10.1128/aem.03406-16.
Kim, K., Katayama, H., Kitajima, M., Tohya, Y., & Ohgaki, S. (2011). Development of a real-time RT-PCR assay combined with ethidium monoazide treatment for RNA viruses and its application to detect viral RNA after heat exposure. Water Science and Technology,63(3), 502–507. https://doi.org/10.2166/wst.2011.249.
Kingsley, D. H., Vincent, E. M., Meade, G. K., Watson, C. L., & Fan, X. (2014). Inactivation of human norovirus using chemical sanitizers. International Journal of Food Microbiology,171, 94–99. https://doi.org/10.1016/j.ijfoodmicro.2013.11.018.
Kitajima, M., Iker, B. C., Pepper, I. L., & Gerba, C. P. (2014). Relative abundance and treatment reduction of viruses during wastewater treatment processes—Identification of potential viral indicators. Science of the Total Environment,488–489, 290–296. https://doi.org/10.1016/j.scitotenv.2014.04.087.
Kundu, A., McBride, G., & Wuertz, S. (2013). Adenovirus-associated health risks for recreational activities in a multi-use coastal watershed based on site-specific quantitative microbial risk assessment. Water Research,47(16), 6309–6325. https://doi.org/10.1016/j.watres.2013.08.002.
La Rosa, G., Iaconelli, M., Pourshaban, M., & Muscillo, M. (2010). Detection and molecular characterization of noroviruses from five sewage treatment plants in central Italy. Water Research,44(6), 1777–1784. https://doi.org/10.1016/j.watres.2009.11.055.
Ligges, U., & Mächler, M. (2003). Scatterplot3d—An R package for visualizing multivariate data. Journal of Statistical Software,8, 1–20.
Limsawat, S., & Ohgaki, S. (1997). Fate of liberated viral RNA in wastewater determined by PCR. Applied and Environmental Microbiology,63(7), 2932–2933.
López-Gálvez, F., Randazzo, W., Vásquez, A., Sánchez, G., Tombini Decol, L., Aznar, R., et al. (2018). Irrigating lettuce with wastewater effluent: Does disinfection with chlorine dioxide inactivate viruses? Journal of Environmental Quality,47(5), 1139–1145. https://doi.org/10.2134/jeq2017.12.0485.
Miura, T., Lhomme, S., Le Saux, J. C., Le Mehaute, P., Guillois, Y., Couturier, E., et al. (2016). Detection of Hepatitis E virus in sewage after an outbreak on a French Island. Food and Environmental Virology,8(3), 194–1999. https://doi.org/10.1007/s12560-016-9241-9.
Montazeri, N., Goettert, D., Achberger, E. C., Johnson, C. N., Prinyawiwatkul, W., & Janes, M. E. (2015). Pathogenic enteric viruses and microbial indicators during secondary treatment of municipal wastewater. Applied and Environmental Microbiology,81(18), 6436–6445. https://doi.org/10.1128/aem.01218-15.
National Research Council. (2012). Water reuse. Washington, DC: The National Academies Press.
Nordgren, J., Matussek, A., Mattsson, A., Svensson, L., & Lindgren, P. E. (2009). Prevalence of norovirus and factors influencing virus concentrations during one year in a full-scale wastewater treatment plant. Water Research,43(4), 1117–1125. https://doi.org/10.1016/j.watres.2008.11.053.
Parshionikar, S., Laseke, I., & Fout, G. S. (2010). Use of propidium monoazide in reverse transcriptase PCR to distinguish between infectious and noninfectious enteric viruses in water samples. Applied and Environmental Microbiology,76(13), 4318–4326. https://doi.org/10.1128/aem.02800-09.
Prevost, B., Goulet, M., Lucas, F. S., Joyeux, M., Moulin, L., & Wurtzer, S. (2016). Viral persistence in surface and drinking water: Suitability of PCR pre-treatment with intercalating dyes. Water Research,91, 68–76. https://doi.org/10.1016/j.watres.2015.12.049.
Prevost, B., Lucas, F. S., Goncalves, A., Richard, F., Moulin, L., & Wurtzer, S. (2015). Large scale survey of enteric viruses in river and waste water underlines the health status of the local population. Environment International,79, 42–50. https://doi.org/10.1016/j.envint.2015.03.004.
Qiu, Y., Lee, B. E., Neumann, N., Ashbolt, N., Craik, S., Maal-Bared, R., et al. (2015). Assessment of human virus removal during municipal wastewater treatment in Edmonton, Canada. Journal of Applied Microbiology,119(6), 1729–1739. https://doi.org/10.1111/jam.12971.
R Core Team. (2014). R: A language and environment for statistical computing. Vienna: R Core team.
Randazzo, W., Khezri, M., Ollivier, J., Le Guyader, F. S., Rodríguez-Díaz, J., Aznar, R., et al. (2018a). Optimization of PMAxx pretreatment to distinguish between human norovirus with intact and altered capsids in shellfish and sewage samples. International Journal of Food Microbiology,266, 1–7. https://doi.org/10.1016/j.ijfoodmicro.2017.11.011.
Randazzo, W., López-Gálvez, F., Allende, A., Aznar, R., & Sánchez, G. (2016). Evaluation of viability PCR performance for assessing norovirus infectivity in fresh-cut vegetables and irrigation water. International Journal of Food Microbiology,229, 1–6. https://doi.org/10.1016/j.ijfoodmicro.2016.04.010.
Randazzo, W., Piqueras, J., Rodríguez-Díaz, J., Aznar, R., & Sánchez, G. (2018b). Improving efficiency of viability-qPCR for selective detection of infectious HAV in food and water samples. Journal of Applied Microbiology,124(4), 958–964. https://doi.org/10.1111/jam.13519.
Rodríguez-Díaz, J., Querales, L., Caraballo, L., Vizzi, E., Liprandi, F., Takiff, H., et al. (2009). Detection and characterization of waterborne gastroenteritis viruses in urban sewage and sewage-polluted river waters in Caracas, Venezuela. Applied and Environmental Microbiology,75(2), 387–394. https://doi.org/10.1128/AEM.02045-08.
Sano, D., Amarasiri, M., Hata, A., Watanabe, T., & Katayama, H. (2016). Risk management of viral infectious diseases in wastewater reclamation and reuse: Review. Environment International,91, 220–229. https://doi.org/10.1016/j.envint.2016.03.001.
Schmitz, B. W., Kitajima, M., Campillo, M. E., Gerba, C. P., & Pepper, I. L. (2016). Virus reduction during advanced Bardenpho and conventional wastewater treatment processes. Environmental Science and Technology,50(17), 9524–9532. https://doi.org/10.1021/acs.est.6b01384.
SEPA. (2008). Guidance to the Swedish Environmental Protection Agency’s regulation for environmental reports (Vägledning om Naturvårdsverkets föreskrifter om miljörapport).
Simmons, F. J., & Xagoraraki, I. (2011). Release of infectious human enteric viruses by full-scale wastewater utilities. Water Research,45(12), 3590–3598. https://doi.org/10.1016/j.watres.2011.04.001.
Sinclair, R. G., Choi, C. Y., Riley, M. R., & Gerba, C. P. (2008). Chapter 9—Pathogen surveillance through monitoring of sewer systems. In A. I. Laskin, S. Sariaslani, & G. M. Gadd (Eds.), Advances in applied microbiology (Vol. 65, pp. 249–269). New York: Academic Press.
Standard Methods For the Examination of Water and Wastewater. (2011). In 9510 detection of enteric viruses: American Public Health Association.
Van Abel, N., Schoen, M. E., Kissel, J. C., & Meschke, J. S. (2017). Comparison of risk predicted by multiple norovirus dose-response models and implications for quantitative microbial risk assessment. Risk Analalysis,37(2), 245–264. https://doi.org/10.1111/risa.12616.
Verbyla, M. E., & Mihelcic, J. R. (2015). A review of virus removal in wastewater treatment pond systems. Water Research,71, 107–124. https://doi.org/10.1016/j.watres.2014.12.031.
WHO. (2017). Potable reuse: Guidance for producing safe drinking-water. Geneva: WHO.
Wilrich, C., & Wilrich, P. T. (2009). Estimation of the pod function and the LOD of a qualitative microbiological measurement method. Journal of AOAC International,92(6), 1763–1772.
Acknowledgements
This study was supported by the “VIRIDIANA” Project AGL2017-82909 (AEI/FEDER, UE) funded by Spanish Ministry of Science, Innovation and Universities; the APOTI Grant (APOTIP/2018/007) from the Generalitat Valenciana; and the CSIC internal Project 201770I088. W. Randazzo was supported by a postdoctoral fellowship from Generalitat Valenciana (APOSTD/2018/150).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Randazzo, W., Piqueras, J., Evtoski, Z. et al. Interlaboratory Comparative Study to Detect Potentially Infectious Human Enteric Viruses in Influent and Effluent Waters. Food Environ Virol 11, 350–363 (2019). https://doi.org/10.1007/s12560-019-09392-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12560-019-09392-2