Abstract
Purpose
Polycyclic aromatic hydrocarbons (PAHs) are a group of environmental pollutants associated with multiple cancers, including female breast cancer. Several xenobiotic metabolism genes (XMGs), including the CYP450 family, play an important role in activating and detoxifying PAHs, and variations in the activity of the enzymes they encode can impact this process. This study aims to examine the association between XMGs and breast cancer, and to assess whether these variants modify the effects of PAH exposure on breast cancer risk.
Methods
In a case–control study in Vancouver, British Columbia, and Kingston, Ontario, 1037 breast cancer cases and 1046 controls had DNA extracted from blood or saliva and genotyped for 138 single nucleotide polymorphisms (SNPs) and tagSNPs in 27 candidate XMGs. Occupational PAH exposure was assessed using a measurement-based job-exposure matrix.
Results
An association between genetic variants and breast cancer was observed among six XMGs, including increased risk among the minor allele carriers of AKR1C3 variant rs12387 (OR 2.71, 95% CI 1.42–5.19) and AKR1C4 variant rs381267 (OR 2.50, 95% CI 1.23–5.07). Heterogeneous effects of occupational PAH exposure were observed among carriers of AKR1C3/4 variants, as well as the PTGS2 variant rs5275.
Conclusion
Our findings support an association between SNPs of XMGs and female breast cancer, including novel genetic variants that modify the toxicity of PAH exposure. These results highlight the interplay between genetic and environmental factors, which can be helpful in understanding the modifiable risks of breast cancer and its complex etiology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Polycyclic aromatic hydrocarbons (PAHs) are one of the most widespread organic pollutants, with exposure occurring through multiple routes, including diet, air pollution, smoking, and the workplace [1]. Evidence from several epidemiological studies suggests that exposure to PAHs is a risk factor for several cancer sites, including breast cancer in women [2,3,4,5].
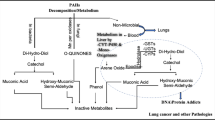
PAH carcinogenicity occurs through the metabolic activation of PAH by cytochrome P450 (CYP450), which consists of a superfamily of hemoproteins that coordinate the metabolism of numerous endogenous and exogenous chemicals. CYP450 enzymes are present in most tissues of the body and function to metabolize potentially toxic compounds [6, 7]. However, this process can produce DNA-binding “ultimate carcinogenic” metabolites that include diol-epoxides, radical cations, and quinones [8,9,10,11,12,13]. PAH exposure can also trigger both estrogenic and antiestrogenic responses [14,15,16] through the increase of estradiol metabolism that in turn increases the formation of quinones [13].
Several studies investigated the association of CYP and other metabolism-related genes with breast cancer risk, many of which are also involved in PAH metabolism [17,18,19,20]. CYP1B1 is an important activator of PAH in mammary glands [17] and certain genotypes have been linked to increased breast cancer risk [21]. Despite their involvement in PAH metabolism, little research has explored the interplay between xenobiotic metabolism genes (XMGs) and PAH exposure and its modifying effects on breast cancer risk. Several studies offered evidence of interactions between PAH–DNA adduct levels and XMGs [22,23,24,25,26], and PAH–DNA adduct levels have been associated with increased breast cancer risk [27]. Among the few studies that have explored interactions between these genes and PAH exposure, the sources of PAH exposure are typically smoking [28, 29] and diet [21, 30]; however, interactions with occupational sources have rarely been studied, despite the fact that exposure levels from occupational sources can be orders of magnitude higher and consist of different mixtures of PAHs. Previous research into the effects of occupational PAH exposure provide support of an interaction between occupational PAH exposure and family history of breast cancer [5].
Our objective was to investigate associations between XMG variants and breast cancer risk in women, and potential interactions between these genetic variants and occupational PAH exposure. We hypothesize that interactions between XMG variants and occupational PAH exposure can modify PAH-related breast cancer risk.
Materials and methods
Study population
A multi-centre, population-based case–control study was conducted in the greater metropolitan area of Vancouver, British Columbia (BC) and Kingston, Ontario (ON) between 2005 and 2010. Ethics approval was provided by the University of British Columbia/BC Cancer Research Ethics Board and the Queen’s University Health Sciences Research Ethics Board. Detailed information on the methods has been previously published [5, 31]. In Vancouver, cases were recruited from the BC Cancer Registry and included women 40–80 years of age, diagnosed with either in situ or invasive breast cancer, no previous history of cancer except for non-melanoma skin cancer, and living in the greater Vancouver metropolitan area at the time of diagnosis. Controls were women recruited from the BC Cancer Breast Screening Program and were frequency matched to cases by 5-year age group. In Kingston, both cases and controls were recruited from the Hotel Dieu Breast Assessment Program. Cases were women 40–80 years diagnosed with either in situ or invasive breast cancer. Controls had either normal mammogram results or benign breast disease and were frequency matched to cases by 5-year age group. Following exclusions according to eligibility criteria, 1130 cases and 1069 controls were included. Participants completed a questionnaire and provided either a blood or saliva sample for genotyping; DNA was extracted from blood (n = 1980) or saliva (n = 204).
Questionnaire and PAH exposure assessment
The questionnaire, which was either self-completed and mailed or administered through telephone interview, included education, ethnicity, medical and reproductive history, lifetime tobacco consumption, and lifetime occupational history. Lifetime work history, including industry, job title, length of employment, work hours (e.g., part-time or full-time), as well as tasks performed and materials handled was collected on all jobs held for at least 6 months. PAH exposure assessment was performed using a job-exposure matrix [5, 32] and the total number of years employed in jobs with risk of exposure above the permissible exposure limit (0.2 mg·m−3) [33] was calculated for each individual.
Gene and variant selection
Twenty-seven genes related to endogenous or exogenous (xenobiotic) metabolism were identified from the literature. The majority of genes are members of the CYP450 superfamily (CYP1A1, CYP1A2, CYP1B1, CYP2C19, CYP2E1, CYP19A1). The remaining genes are grouped by function: modulation of the PAH metabolism response (AHR, AHRR, ARNT, AIP), formulation (or activation) of carcinogenic intermediates during metabolism (AKR1A1, AKR1C1-AKR1C4, DHDH, EPHX1, PTGS2, NAT1, NAT2), and detoxification of metabolites (COMT, NQO1, GSTP1, NRF2, PON1) into their final inactive, excretable forms. Additional genes selected are related to estradiol metabolism (ESR1, ESR2) [34], which is influenced by PAH metabolism [14,15,16, 35]. For each gene, a set of tagSNPs were selected using the CEU (European) population from HapMap release 28 using Tagger [36] and the program Haploview [37] using a minimum minor allele frequency (MAF) of 0.10 and an r2 threshold of 0.8. A total of 158 SNPs associated with the XMGs described above were submitted for genotyping.
Genotyping
SNPs included in the analysis were initially part of a larger Illumina GoldenGate genotyping assay (768 SNPs) that included SNPs related to other potential pathways for breast cancer. SNPs that failed initial assay design were replaced with equivalent tagNPs. Genotyping was performed at the McGill University and Genome Quebec Innovation Centre in Montreal, QC, Canada.
Quality control procedures
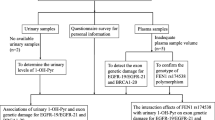
Genotype quality control for the 768 SNP set was performed in Genome Studio v2011.1 (Illumina, San Deigo, CA, USA), PLINK v1.07 [38], GRR [39], and Excel 2007 (Microsoft, Redmond, WA, USA). Figures 1 and 2 summarize the reasons for exclusion of SNPs and samples, respectively.
SNP
SNP exclusion was based on recommendations by Illumina (Illumina User Guide, Illumina, Part #11319113): GenCall Score < 0.25, GenTrain score < 0.40, poor clustering, monoallelic variants, genotype discrepancies in replicate samples in any SNP (n = 124), call rate < 95%, unexpectedly low MAF in European controls compared to HapMap CEU data, or out of Hardy–Weinberg equilibrium (p < 0.001) in European-ancestry controls. Call rate was examined separately for saliva samples. If an SNP had a call rate < 95% for saliva samples, but > 95% for blood samples, participants that provided samples through saliva only were excluded (i.e., the sample size for that SNP was reduced in comparison to the other SNPs). Genotyping of XMGs included 158 SNPs, of which 20 were excluded: 6 by the genotyping center, 8 due to poor clustering, 2 that were monoallelic within the study population, 3 with a low call rate, and 1 with a lower MAF compared to the HapMap CEU population; a total of 138 SNPs were included in the analysis.
Samples
Samples were excluded if heterozygosity was greater than three standard deviations compared to other samples within the same ethnicity, call rate < 0.95, genotypes at Y chromosome markers indicated the sex was male, unrelated samples had identical genotypes, and if there were discrepancies between genotype-estimated and self-reported ethnicity. Comparison between self-reported and genotype-estimated ethnicities was done by calculating the identity by state and multi-dimensional scaling plots [38] with HapMap samples in the CEU, CHB, CHD, JPT, TRI, and TSI populations [40]. Associations between case status and ethnicity were detected through calculation of the genomic inflation factor (λ = 16.99) using ancestral informative markers after SNP quality control [41,42,43]. Reanalysis using European-only samples resulted in no discernible inflation (λ = 1.0), indicating there is no population structure or genotyping error [44]. Nine samples were identified as having a familial relationship, which was verified using questionnaires. If both pairs were cases, the individual with the earlier diagnosis was included (n = 4); if both pairs were controls, the oldest of the pair was included (n = 3); and if one of the pair was a case and the other a control, the case was included (n = 2). Following sample quality control and inclusion criteria, a total of 2083 samples (1037 cases and 1046 controls) were retained.
Statistical analysis
Multivariable logistic regression was used to calculate adjusted odds ratios (OR) and 95% confidence intervals (CI) to examine the relationship between the SNPs and breast cancer risk; all regression models were adjusted for age and study center. To investigate the associations between xenobiotic metabolism-related SNPs and breast cancer, we used an SNP-specific inheritance model (i.e., one of three inheritance models: additive, dominant, or recessive) based on the two-step approach described below. To control for confounding due to population stratification, all analyses were restricted to women of European ethnicity [45, 46]. Results for women of (East) Asian descent (defined as Chinese, Japanese, or Korean ancestry) are in Supplementary Tables (see Appendix); other ethnic groups were excluded due to small sample sizes. Differences in risk by menopausal status were examined through stratified analysis and inclusion of an interaction term in the logistic models.
Multiple testing was corrected through a two-step gene-based process similar to that of Schuetz et al. [47] with modifications for the different inheritance models. Each SNP went through a set-based permutation (10,000 permutations) where, for each permutation, case status was randomly assigned. For each inheritance model, a p-value was calculated for the permutation resulting in three inheritance model-specific p-values for each SNP. An adjusted p-value for each SNP was calculated using the number of times a more extreme (i.e., smaller) p-value, compared to the original inheritance model-specific pvalue, was observed during the 10,000 permutations. Within each gene, the minimum adjusted inheritance model-specific p value was used to select the inheritance model and the gene-representative SNP. Benjamini–Hochberg procedure [48] was applied to control the false discovery rate (FDR) to obtain a corrected p-value for the gene-representative SNP (n = 27).
SNPs that displayed any evidence of association with breast cancer (permutation adjusted p-value < 0.1) were examined for potential gene–environment interactions (GxEs) with PAH exposure. Exposure metrics were defined as: (1) duration at “high” PAH exposure, (2) average probability of exposure, and (3) weighted duration of exposure, as described in Lee et al. [5] G×E analyses were examined through a genotype-exposure interaction term in the logistic models. For some of the genotypes-exposure strata, insufficient sample sizes required PAH exposure to be dichotomized into an ever-never categorization. All interaction term p-values were corrected for the FDR [48]. In the situation where the homozygous minor allele genotype group had insufficient sample size to test for interactions using the recessive inheritance model (minimum requirement: n = 50), the additive model was used to allow stable estimates. Education and smoking were identified as potential confounders of the PAH association in previous analysis [5], therefore all G×E analyses were adjusted for education and smoking (pack-years), in addition to age and center. Analyses involving duration at “high” PAH exposure included a nuisance variable {1 if maximum level of exposure was low or medium, 0 = other} to ensure that the referent exposure group was truly unexposed [5]. Statistical analyses were conducted using the statistical software R (version 2.14.2, R Foundation for Statistical Computing, Vienna, Austria).
Results
Cases were more likely to have ever been pregnant, tended to be older at time of first mammogram, more likely to be overweight or obese, and more likely to have a family history of breast cancer (Table 1). Among current or previous smokers, cases smoked more pack-years than controls. Controls were more likely to be of European descent and to have a higher socioeconomic status (i.e., family income greater than $80,000 and/or have a graduate/professional school degree).
Table 2 shows adjusted ORs from the logistic models for the main genetic analysis involving women of European decent. Following the permutation step of the gene-based approach, 12 SNPs were observed to have associations with breast cancer risk and 6 SNPs still showed evidence of an association after FDR adjustment: rs12387 (AKR1C3), rs3812617 (AKR1C4), rs12248560 (CYP2C19), rs7845127 (NAT1), rs4646243 (NAT2), and rs2813543 (ESR1). Differences in associations between genotype and breast cancer risk among pre- and postmenopausal women were observed for SNPs associated with AKR1A1, AKR1C3, AKR1C4, CYP1B1, and NQ01; however, none remained significant after FDR adjustment (padj value > 0.2) (see Appendix: Supplementary Table A1). Among women of Asian descent, the same 27 SNPs and SNP-specific inheritance models were assessed; no evidence of association with breast cancer was observed (see Appendix: Supplementary Table A2). Minor differences by ethnicity were observed for SNPs associated with COMT, CYP19A1, and NAT2; however, none remained noteworthy following FDR adjustment (padj value > 0.2).
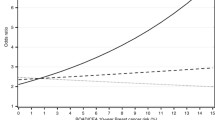
After FDR adjustment, six SNPs that met the threshold for significance, along with six other SNPs that initially showed associations with breast cancer risk [rs5993882 (COMT), rs10046 (CYP19A1), rs2470893 (CYP1A1), rs2854461 (EPHX1), rs854551 (PON1), and rs5275 (PTGS2)], were assessed for interactions with PAH exposure among women of European descent. Table 3 shows adjusted ORs by genotype-exposure stratum for select SNPs with potential modifying effects; for three SNPs, we observed evidence of an interaction with duration at “high” PAH exposure, which remained noteworthy after FDR adjustment (padj-interaction < 0.10). One SNP is a member of the cytochrome c-oxidase (COX) family: rs5275 (PTGS2) and the other two are from the aldo–keto reductase (AKR) superfamily: rs12387 (AKR1C3) and rs3812617 (AKR1C4). As was the case for both AKR SNPs, an increasing risk for breast cancer with increased duration of high exposure was observed within the homozygous major allele genotype stratum; within the heterozygous or homozygous minor strata, no associations were observed. Among the non-exposed group, there was an increasing risk with each minor allele. Similar effects as the AKR SNPs were observed for rs5275 (PTGS2) across duration of exposure within the homozygous major stratum; however, the effects were null among the non-exposed group and, as duration of exposure increased, there was an increasing protective association with the minor allele. For the remaining SNP results, see Supplementary Table A3. Evidence of interactions were observed for rs5275 (PTGS2) using the other two exposure metrics, average probability of PAH exposure and weighted duration of exposure, which remained significant after FDR adjustment (padj-interaction < 0.05); however, no evidence of an interaction was apparent among the AKR SNPs (see Supplementary Table A4 and Supplementary Table A5). Dichotomizing duration at “high” (see Table 4 and Supplementary Table A6) and average probability of PAH exposure to ever-never exposed produced similar results as their original categorization (see Supplementary Table A7). Ever-never categorization for average probability of PAH exposure produces the same results as weighted duration of exposure (data not shown). No evidence of effect modification by PAH exposure on genotype-associated breast cancer risk was observed among women of Asian descent (data not shown).
Tobacco smoke is a known source of PAH exposure, and duration of smoking has been associated with increased risk of breast cancer [49, 50]. We observed no evidence of effect modification by smoking on genotype-associated breast cancer risk (see Supplementary Table A8), nor was there evidence of a three-way or two-way PAH exposure–smoking interactions after FDR adjustment (see Supplementary Table A9 and Supplementary Table A10); due to sample size issues, smoking was dichotomized.
Discussion
In this population-based case–control study, we observed evidence of associations between various XMG variants and breast cancer. Among 12 SNPs that were observed to have associations with breast cancer, there was evidence to suggest an association with 6 SNPs after FDR adjustment. Among these variants, heterogeneous effects of PAH exposure on breast cancer risk were observed for three of these SNPs, including two members of the AKR superfamily (AKR1C3, AKR1C4), which are involved in the production of carcinogenic intermediate o-quinone during PAH metabolism [51,52,53].
AKR1C3 regulates receptor access of androgens and estrogens and is involved in the biosynthesis of prostaglandins, and the overexpression of AKR1C3 in steroid hormone-dependent breast tumors [54] highlights its potential role in the etiology of breast cancer [55]. For the SNP rs12387, which is a missense variant (Lys → Asn), we observed an increased risk of breast cancer among the homozygous major allele genotype that was limited to ever users of HRT (phetero-value = 0.09). Under a similar mode of inheritance, the increased risk with carriers of the G allele was consistent with those observed by Reding [56] (HRT-ever: OR = 1.42 (0.99–2.05), HRT-never: OR = 1.01 (0.76–1.34), phetero-value = 0.16). AKR1C4 is also involved in the metabolic activation of PAHs, and although its role in the breast cancer etiology is less clear because it is predominantly liver specific [57], it is also involved in estrogen metabolism. The SNP rs3812617, which is a splice-region variant, could disrupt RNA splicing by skipping exons, thereby resulting in an altered protein-coding sequence. Alternatively, the SNP is in high linkage disequilibrium (LD) with rs3829125 (r2 = 0.9), which is a missense variant (Cys → Ser), previously associated with prostate cancer through hormone-mediated effects on estrogen receptor α and β [58]. Several PAHs can occupy estrogen receptor (ER) binding sites through a similar endocrine disrupting mechanism, thereby allowing ERs to serve as a pathway to transport an ultimate carcinogen directly to specific DNA regulatory sites [15, 59] that can influence estrogen-mediated breast cancer risk.
An association between breast cancer and CYP2C19 SNP rs12248560 was observed, with the results suggesting an increased risk for minor allele carriers. Justenhoven et al. suggested a protective effect [60]; however, no overall association with breast cancer risk was observed in a follow-up pooled analysis, although there was evidence of an association within the hormone replacement therapy subgroup (≥ 10 years on HRT) [61]. We found no protective association within our study population that received HRT for at least 10 years (data not shown). ESR1 has been shown to play a major role in the development and treatment of breast cancer [62], and, similar to other polymorphisms, a protective effect against breast cancer was observed for carriers of the minor allele for SNP rs2813543 [63]. Given its location downstream of the gene, it is more likely that rs2813543 is in LD with another SNP that may have a functional effect. The NAT1/2 genes have established roles in detoxifying and/or bioactivating a variety of aromatic and heterocyclic amines [64, 65], and certain polymorphisms have demonstrated associations with breast cancer [28, 64]. NAT1 SNP rs7845127 and NAT2 SNP rs4646243, which are both upstream of the gene, were associated with increased risk for breast cancer among the heterozygous and/or homozygous minor allele carriers, respectively.
The study of gene–environment interactions in diseases like cancer may be pivotal in understanding their etiology, especially when risks from certain exposures are only detectable in those with certain genetic susceptibilities, which in turn prevents us from identifying the true impact of either unless both effects are considered together. We observed evidence of interactions between SNPs in XMGs (rs12387, rs3812617, and rs5275) and occupational PAH exposure. The AKR SNPs rs12387 (AKR1C3) and rs3812617 (AKR1C4) are involved in the production of quinones during PAH metabolism [52, 66,67,68]. Exposure to PAHs is thought to trigger estrogenic and antiestrogenic responses [15] through increased metabolism of estradiol, which result in the increased formation of quinones [13]. Women with the homozygous major genotype were found to be at an increased risk for breast cancer in proportion to longer occupational PAH exposure; however, for both heterozygous and homozygous minor allele genotypes, the increased risks were attenuated. PTGS2 SNP rs5275 showed some evidence of a reduced risk of breast cancer for the minor allele carriers; however, similar to the AKR SNPs, women with the homozygous major genotype had an increased risk for breast cancer with duration of occupational PAH exposure. No association was observed in the heterozygous and homozygous minor allele strata. The modifying effect of the PTGS2 SNP on PAH exposure remained consistent across different metrics of PAH exposure. Other studies have also observed a decreased risk for breast cancer with this variant, including a pooled analysis of the Nurses’ Health Study 2 and Harvard Women’s Health Study [69, 70]. Like the AKR superfamily, the PTGS2 SNP may influence breast cancer risk through estrogen metabolism. PTGS2/COX-2 encodes a prostaglandin synthase enzyme (cyclooxygenase) that can increase the production of prostaglandins (e.g., PGE2), which in turn simulates estrogen production through steroidogenesis [71]. High levels of cyclooxygenase have been observed in human mammary tumor tissues [72, 73] and overexpression of the gene is capable of inducing mammary epithelial tumorigenesis in animal models [74].
As our focus when selecting candidate genes was based on the use of tagSNPs, one limitation of this approach is there is no guarantee that the SNP tested for association is the contributing SNP. However, an intent of this study was to identify gene variants that are associated with breast cancer risk, which we have demonstrated. Furthermore, although the identity of the exact causal SNP may not be known, by using a tagSNP it can be surmised that there is a causal SNP in LD that could be identified through functional studies.
Another limitation of the study involves measurement error in assessed PAH exposure. Differential misclassification is a potential limitation of using a job-exposure matrix for classifying exposure status that can either attenuate or accentuate the interaction estimates [75, 76]. This misclassification in inferred exposure status also decreases the efficiency of GxE studies [77]. It is worth noting that there are also circumstances where an observed association with a gene provides evidence of gene–environment interaction, even if the effect estimate of the interaction term in regression models is not strong. This occurs when measurement error in exposure dilutes the power of the test of interaction compared to the test of genetic association alone. In this case, the observed effect of a gene depends on its interaction with the true exposure; thus, without even estimating exposure, the genetic effect can be used to detect (rather than quantify) the interaction [78]. We note that after FDR adjustment, of the six SNPs associated with breast cancer in our gene-only analyses that passed adjustment and of the other six SNPs that showed associations, but failed to meet our threshold after FDR adjustment, three had G × E estimates that passed FDR adjustment. Consequently, it is plausible that several of the other SNPs are providing evidence for gene–environment interactions, in part because of their role in modifying PAH toxicity. If these XMGs contribute to the risk of disease only in the presence of exposure, the existence of a G × E can be inferred from our mis-specified gene-only models. This approach has been effectively used to show evidence of G × Es when exposure is difficult to assess accurately [78].
Tobacco smoke is a known source of PAH exposure and is also a potential confounder, with some studies suggesting that long duration of smoking can result in an increased breast cancer risk among women with certain genotypes [50, 79]. However, there is little evidence of a measurable effect of smoking on breast cancer risk or of an interaction between PAH exposure and smoking within this study population, as we observed null effects from smoking within genotype-specific strata and genotype-exposure strata, i.e., two-way and three-way interactions. Moreover, we previously observed no heterogeneous effect of PAH exposure by smoking or menopausal status within this study population; however, among pre-menopausal women, we did report a large increased risk with PAH exposure, as well as smoking, albeit smoking was only marginally significant. Most importantly, these observations suggest that pre-menopausal occupational exposure to PAH make greater contribution to breast cancer risk [5] (data not shown). Furthermore, the potential effect of active smoking on breast cancer risk has been observed to be only modest in the range of 1–10% excess risk for ever-smoking, and related to smoking before menopause and the first birth, leading to an average 10–20% increase in risk per 20 pack-years in these women [80]; on the other hand, Xue et al. also report inverse association with smoking after the menopause [80]. Among our study population, we observed a similar average excess risk of 9% for ever-smoking, but our study population smoked substantially less than those in Xue et al. Although the lack of effect modification from smoking by PAH-related genes may weaken our argument of a modified breast cancer risk by these XMGs, our results support the observations by Xue et al. [80], concurring with the observation of an increased risk of breast cancer by smoking identified only among pre-menopausal women, indicating that our two studies are in agreement of a window of susceptibility for exposure to mixtures that contain PAHs and their impact on breast cancer. This issue of modification of breast cancer risk by genes depending on timing of environmental exposure, such as PAH and tobacco smoke, appears to be a promising avenue of future research.
In summary, an association between genetic variants and breast cancer was observed among six XMGs: two of the variants belonged to genes from the AKR superfamily, and four were novel variants of genes that have known associations with breast cancer. Modifying effects on breast cancer risk that differed among those exposed to occupational PAH exposure were observed among carriers of three genetics variants. The results of this study support previous evidence observed of interactions between PAH exposure and family history of breast cancer [5], and highlight the interplay of genetic and environmental risk factors, which can be helpful in understanding the modifiable risk factors of breast cancer.
References
IARC Working Group on the Evaluation of Carcinogenic Risks to Humans. Some non-heterocyclic polycyclic aromatic hydrocarbons and some related exposures. In: IARC monographs on the evaluation of carcinogenic risks to humans, vol. 92. Lyon: International Agency for Research on Cancer; 2010.
Petralia SA, Vena JE, Freudenheim JL, Dosemeci M, Michalek A, Goldberg MS, Brasure J, Graham S. Risk of premenopausal breast cancer in association with occupational exposure to polycyclic aromatic hydrocarbons and benzene. Scand J Work Environ Health 1999;25(3):215–221.
Hansen J. Elevated risk for male breast cancer after occupational exposure to gasoline and vehicular combustion products. Am J Ind Med. 2000;37(4):349–52.
Labreche F, Goldberg MS, Valois M-F, Nadon L. Postmenopausal breast cancer and occupational exposures. Occup Environ Med. 2010;67(4):263–9.
Lee DG, Burstyn I, Lai AS, Grundy A, Friesen MC, Aronson KJ, Spinelli JJ. Women’s occupational exposure to polycyclic aromatic hydrocarbons and risk of breast cancer. Occup Environ Med. 2019;76(1):22–9.
Rendic S, Carlo FJD. Human cytochrome P450 enzymes: a status report summarizing their reactions, substrates, inducers, and inhibitors. Drug Metab Rev. 1997;29(1–2):413–580.
Anzenbacher P, Anzenbacherova E. Cytochromes P450 and metabolism of xenobiotics. Cell Mol Life Sci CMLS. 2001;58(5–6):737–47.
Sims P, Grover P. Epoxides in polycyclic aromatic hydrocarbon metabolism and carcinogenesis. Adv Cancer Res. 1974;20:165–274.
Gelboin HV. Benzo [alpha] pyrene metabolism, activation and carcinogenesis: role and regulation of mixed-function oxidases and related enzymes. Physiol Rev. 1980;60(4):1107–66.
Conney AH. Induction of microsomal enzymes by foreign chemicals and carcinogenesis by polycyclic aromatic hydrocarbons: GHA clowes memorial lecture. Can Res. 1982;42(12):4875–917.
Shimada T, Fujii-Kuriyama Y. Metabolic activation of polycyclic aromatic hydrocarbons to carcinogens by cytochromes P450 1A1 and1B1. Cancer Sci. 2004;95(1):1–6.
Baird WM, Hooven LA, Mahadevan B. Carcinogenic polycyclic aromatic hydrocarbon-DNA adducts and mechanism of action. Environ Mol Mutagen. 2005;45(2–3):106–14.
Shimada T. Xenobiotic-metabolizing enzymes involved in activation and detoxification of carcinogenic polycyclic aromatic hydrocarbons. Drug Metab Pharmacokinet. 2006;21(4):257–76.
Chaloupka K, Krishnan V, Safe S. Polynuclear aromatic hydrocarbon carcinogens as antiestrogens in MCF-7 human breast cancer cells: role of the Ah receptor. Carcinogenesis. 1992;13(12):2233–9.
Santodonato J. Review of the estrogenic and antiestrogenic activity of polycyclic aromatic hydrocarbons: relationship to carcinogenicity. Chemosphere. 1997;34(4):835–48.
Arcaro KF, O’Keefe PW, Yang Y, Clayton W, Gierthy JF. Antiestrogenicity of environmental polycyclic aromatic hydrocarbons in human breast cancer cells. Toxicology. 1999;133(2):115–27.
Larsen MC, Angus WG, Brake PB, Eltom SE, Sukow KA, Jefcoate CR. Characterization of CYP1B1 and CYP1A1 expression in human mammary epithelial cells: role of the aryl hydrocarbon receptor in polycyclic aromatic hydrocarbon metabolism. Can Res. 1998;58(11):2366–74.
Huang C-S, Chern H-D, Chang K-J, Cheng C-W, Hsu S-M, Shen C-Y. Breast cancer risk associated with genotype polymorphism of the estrogen-metabolizing genes CYP17, CYP1A1, and COMT A multigenic study on cancer susceptibility. Can Res. 1999;59(19):4870–5.
Williams JA, Phillips DH. Mammary expression of xenobiotic metabolizing enzymes and their potential role in breast cancer. Can Res. 2000;60(17):4667–77.
Kristensen VN, Harada N, Yoshimura N, Haraldsen E, Lønning P, Erikstein B, Kåresen R, Kristensen T, Børresen-Dale A-L. Genetic variants of CYP19 (aromatase) and breast cancer risk. Breast Cancer Res. 2000;2(Suppl 1):1–2.
Zheng W, Xie D-W, Jin F, Cheng J-R, Dai Q, Wen W-Q, Shu X-O, Gao Y-T. Genetic polymorphism of cytochrome P450–1B1 and risk of breast cancer. Cancer Epidemiol Biomark Prev. 2000;9(2):147–50.
Rundle A, Tang D, Zhou J, Cho S, Perera F. The association between glutathione S-transferase M1 genotype and polycyclic aromatic hydrocarbon-DNA adducts in breast tissue. Cancer Epidemiol Biomark Prev. 2000;9(10):1079–85.
Firozi PF, Bondy ML, Sahin AA, Chang P, Lukmanji F, Singletary ES, Hassan MM, Li D. Aromatic DNA adducts and polymorphisms of CYP1A1, NAT2, and GSTM1 in breast cancer. Carcinogenesis. 2002;23(2):301–6.
Terry MB, Gammon MD, Zhang FF, Eng SM, Sagiv SK, Paykin AB, Wang Q, Hayes S, Teitelbaum SL, Neugut AI. Polymorphism in the DNA repair gene XPD, polycyclic aromatic hydrocarbon-DNA adducts, cigarette smoking, and breast cancer risk. Cancer Epidemiol Biomark Prev. 2004;13(12):2053–8.
Shen J, Gammon MD, Terry MB, Wang L, Wang Q, Zhang F, Teitelbaum SL, Eng SM, Sagiv SK, Gaudet MM. Polymorphisms in XRCC1 modify the association between polycyclic aromatic hydrocarbon-DNA adducts, cigarette smoking, dietary antioxidants, and breast cancer risk. Cancer Epidemiol Biomark Prev. 2005;14(2):336–42.
Crew KD, Gammon MD, Terry MB, Zhang FF, Zablotska LB, Agrawal M, Shen J, Long C-M, Eng SM, Sagiv SK. Polymorphisms in nucleotide excision repair genes, polycyclic aromatic hydrocarbon-DNA adducts, and breast cancer risk. Cancer Epidemiol Biomark Prev. 2007;16(10):2033–41.
Gammon MD, Sagiv SK, Eng SM, Shantakumar S, Gaudet MM, Teitelbaum SL, Britton JA, Terry MB, Wang LW, Wang Q. Polycyclic aromatic hydrocarbon-DNA adducts and breast cancer: a pooled analysis. Arch Environ Health Int J. 2004;59(12):640–9.
Ambrosone CB, Freudenheim JL, Graham S, Marshall JR, Vena JE, Brasure JR, Michalek AM, Laughlin R, Nemoto T, Gillenwater KA, et al. Cigarette smoking, N-acetyltransferase 2 genetic polymorphisms, and breast cancer risk. J Am Med Assoc. 1996;276(18):1494–501.
Ishibe N, Hankinson SE, Colditz GA, Spiegelman D, Willett WC, Speizer FE, Kelsey KT, Hunter DJ. Cigarette smoking, cytochrome P450 1A1 polymorphisms, and breast cancer risk in the Nurses’ Health Study. Can Res. 1998;58(4):667–71.
Zheng W, Deitz AC, Campbell DR, Wen W-Q, Cerhan JR, Sellers TA, Folsom AR, Hein DW. N-Acetyltransferase 1 genetic polymorphism, cigarette smoking, well-done meat intake, and breast cancer risk. Cancer Epidemiol Biomark Prev. 1999;8(3):233–9.
Grundy A, Richardson H, Burstyn I, Lohrisch C, SenGupta SK, Lai AS, Lee D, Spinelli JJ, Aronson KJ. Increased risk of breast cancer associated with long-term shift work in Canada. Occup Environ Med. 2013;70(12):831–8.
Lee DG, Lavoue J, Spinelli JJ, Burstyn I. Statistical modelling of occupational exposure to polycyclic aromatic hydrocarbons using osha data. J Occup Environ Hyg. 2014;70(1):14.
Agency for Toxic Substances and Disease Registry. Toxicological profile for polycyclic aromatic hydrocarbons. 1995.
Nilsson S, Mäkelä S, Treuter E, Tujague M, Thomsen J, Andersson G, Enmark E, Pettersson K, Warner M, Gustafsson J-Å. Mechanisms of estrogen action. Physiol Rev. 2001;81(4):1535–65.
Ohtake F, Takeyama K, Matsumoto T, Kitagawa H, Yamamoto Y, Nohara K, Tohyama C, Krust A, Mimura J, Chambon P. Modulation of oestrogen receptor signalling by association with the activated dioxin receptor. Nature. 2003;423(6939):545–50.
de Bakker PI, Yelensky R, Pe’er I, Gabriel SB, Daly MJ, Altshuler D. Efficiency and power in genetic association studies. Nat Genet. 2005;37(11):1217–23.
Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21(2):263–5.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, De Bakker PI, Daly MJ. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81(3):559–75.
Abecasis GR, Cherny SS, Cookson W, Cardon LR. GRR: graphical representation of relationship errors. Bioinformatics. 2001;17(8):742–3.
Consortium IH. Integrating common and rare genetic variation in diverse human populations. Nature. 2010;467(7311):52–8.
Devlin B, Roeder K, Wasserman L. Genomic control for association studies: a semiparametric test to detect excess-haplotype sharing. Biostatistics. 2000;1(4):369–87.
Devlin B, Roeder K, Wasserman L. Genomic control, a new approach to genetic-based association studies. Theor Popul Biol. 2001;60(3):155–66.
Devlin B, Bacanu S-A, Roeder K. Genomic control to the extreme. Nat Genet. 2004;36(11):1129–30.
Devlin B, Roeder K. Genomic control for association studies. Biometrics. 1999;55(4):997–1004.
Thomas DC, Witte JS. Point: population stratification: a problem for case–control studies of candidate-gene associations? Cancer Epidemiol Biomark Prev. 2002;11(6):505–12.
Anderson CA, Pettersson FH, Clarke GM, Cardon LR, Morris AP, Zondervan KT. Data quality control in genetic case-control association studies. Nat Protoc. 2010;5(9):1564–73.
Schuetz JM, Daley D, Graham J, Berry BR, Gallagher RP, Connors JM, Gascoyne RD, Spinelli JJ, Brooks-Wilson AR. Genetic variation in cell death genes and risk of non-Hodgkin lymphoma. PLoS ONE. 2012;7(2):e31560.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (Methodological). 1995;57(1):289–300.
Johnson KC, Miller AB, Collishaw NE, Palmer JR, Hammond SK, Salmon AG, Cantor KP, Miller MD, Boyd NF, Millar J. Active smoking and secondhand smoke increase breast cancer risk: the report of the Canadian Expert Panel on Tobacco Smoke and Breast Cancer Risk (2009). Tob Control. 2010;20(1):e2.
White AJ, Bradshaw PT, Herring AH, Teitelbaum SL, Beyea J, Stellman SD, Steck SE, Mordukhovich I, Eng SM, Engel LS. Exposure to multiple sources of polycyclic aromatic hydrocarbons and breast cancer incidence. Environ Int. 2016;89:185–92.
Palackal NT, Lee SH, Harvey RG, Blair IA, Penning TM. Activation of polycyclic aromatic hydrocarbon trans-dihydrodiol proximate carcinogens by human aldo–keto reductase (AKR1C) enzymes and their functional overexpression in human lung carcinoma (A549) cells. J Biol Chem. 2002;277(27):24799–808.
Penning TM, Burczynski ME, Hung C-F, McCoull KD, Palackal NT, Tsuruda LS. Dihydrodiol dehydrogenases and polycyclic aromatic hydrocarbon activation: generation of reactive and redox active o-quinones. Chem Res Toxicol. 1999;12(1):1–18.
Zhang L, Jin Y, Huang M, Penning TM. The role of human aldo-keto reductases (AKRs) in the metabolic activation and detoxication of polycyclic aromatic hydrocarbons: interconversion of PAH-catechols and PAH o-Quinones. Front Pharmacol. 2012;3:193-204.
Lin H-K, Steckelbroeck S, Fung K-M, Jones AN, Penning TM. Characterization of a monoclonal antibody for human aldo-keto reductase AKR1C3 (type 2 3α-hydroxysteroid dehydrogenase/type 5 17β-hydroxysteroid dehydrogenase); immunohistochemical detection in breast and prostate. Steroids. 2004;69(13):795–801.
Byrns MC, Steckelbroeck S, Penning TM. An indomethacin analogue, N-(4-chlorobenzoyl)-melatonin, is a selective inhibitor of aldo-keto reductase 1C3 (type 2 3α-HSD, type 5 17β-HSD, and prostaglandin F synthase), a potential target for the treatment of hormone dependent and hormone independent malignancies. Biochem Pharmacol. 2008;75(2):484–93.
Reding KW, Li CI, Weiss NS, Chen C, Carlson CS, Duggan D, Thummel KE, Daling JR, Malone KE. Genetic variation in the progesterone receptor and metabolism pathways and hormone therapy in relation to breast cancer risk. Am J Epidemiol. 2009;170:1241–9.
Penning TM, Byrns MC. Steroid hormone transforming aldo–keto reductases and cancer. Ann N Y Acad Sci. 2009;1155(1):33–42.
Multigner L, Ndong JR, Giusti A, Romana M, Delacroix-Maillard H, Cordier S, Jégou B, Thome JP, Blanchet P. Chlordecone exposure and risk of prostate cancer. J Clin Oncol. 2010;28(21):3457–62.
Ebright RH, Wong JR, Chen LB. Binding of 2-hydroxybenzo (a) pyrene to estrogen receptors in rat cytosol. Can Res. 1986;46(5):2349–51.
Justenhoven C, Hamann U, Pierl CB, Baisch C, Harth V, Rabstein S, Spickenheuer A, Pesch B, Brüning T, Winter S. CYP2C19* 17 is associated with decreased breast cancer risk. Breast Cancer Res Treat. 2009;115(2):391–6.
Justenhoven C, Obazee O, Winter S, Couch FJ, Olson JE, Hall P, Hannelius U, Li J, Humphreys K, Severi G. The postmenopausal hormone replacement therapy-related breast cancer risk is decreased in women carrying the CYP2C19* 17 variant. Breast Cancer Res Treat. 2012;131(1):347–50.
Chen GG, Zeng Q, Tse GM. Estrogen and its receptors in cancer. Med Res Rev. 2008;28(6):954–74.
Wang J, Higuchi R, Modugno F, Li J, Umblas N, Lee J, Lui L-Y, Ziv E, Tice JA, Cummings SR, et al. Estrogen receptor alpha haplotypes and breast cancer risk in older Caucasian women. Breast Cancer Res Treat. 2007;106(2):273–80.
Hein DW. N-Acetyltransferase genetics and their role in predisposition to aromatic and heterocyclic amine-induced carcinogenesis. Toxicol Lett. 2000;112:349–56.
Hein DW, Doll MA, Fretland AJ, Leff MA, Webb SJ, Xiao GH, Devanaboyina U-S, Nangju NA, Feng Y. Molecular genetics and epidemiology of the NAT1 and NAT2 acetylation polymorphisms. Cancer Epidemiol Biomark Prev. 2000;9(1):29–42.
Penning TM, Ohnishi ST, Ohnishi T, Harvey RG. Generation of reactive oxygen species during the enzymatic oxidation of polycyclic aromatic hydrocarbon trans-dihydrodiols catalyzed by dihydrodiol dehydrogenase. Chem Res Toxicol. 1996;1:84–92.
Burczynski ME, Harvey RG, Penning TM. Expression and characterization of four recombinant human dihydrodiol dehydrogenase isoforms: oxidation of trans-7, 8-dihydroxy-7, 8-dihydrobenzo [a] pyrene to the activated o-quinone metabolite benzo [a] pyrene-7, 8-dione. Biochemistry. 1998;37(19):6781–90.
Jin Y, Penning TM. Aldo–keto reductases and bioactivation/detoxication. Annu Rev Pharmacol Toxicol. 2007;47:263–92.
Dong LM, Potter JD, White E, Ulrich CM, Cardon LR, Peters U. Genetic susceptibility to cancer: the role of polymorphisms in candidate genes. JAMA. 2008;299(20):2423–36.
Cox DG, Buring J, Hankinson SE, Hunter DJ. A polymorphism in the 3′ untranslated region of the gene encoding prostaglandin endoperoxide synthase 2 is not associated with an increase in breast cancer risk: a nested case–control study. Breast Cancer Res. 2007;9(1):R3.
Brueggemeier RW, Richards JA, Petrel TA. Aromatase and cyclooxygenases: enzymes in breast cancer. J Steroid Biochem Mol Biol. 2003;86(3):501–7.
Hwang D, Byrne J, Scollard D, Levine E. Expression of cyclooxygenase-1 and cyclooxygenase-2 in human breast cancer. J Natl Cancer Inst. 1998;90(6):455–60.
Soslow RA, Dannenberg AJ, Rush D, Woerner B, Khan KN, Masferrer J, Koki AT. COX-2 is expressed in human pulmonary, colonic, and mammary tumors. Cancer. 2000;89(12):2637–45.
Liu CH, Chang S-H, Narko K, Trifan OC, Wu M-T, Smith E, Haudenschild C, Lane TF, Hla T. Overexpression of cyclooxygenase-2 is sufficient to induce tumorigenesis in transgenic mice. J Biol Chem. 2001;276(21):18563–9.
Burstyn I, Lavoué J, Van Tongeren M. Aggregation of exposure level and probability into a single metric in job-exposure matrices creates bias. Ann Occup Hyg. 2012;56(9):1038–50.
Gustafson P. Measurement error and misclassification in statistics and epidemiology: impacts and Bayesian adjustments. Boca Raton: Chapman & Hall/CRC; 2004.
Gustafson P, Burstyn I. Bayesian inference of gene–environment interaction from incomplete data: What happens when information on environment is disjoint from data on gene and disease? Stat Med. 2011;30(8):877–89.
Burstyn I, Kim H-M, Yasui Y, Cherry NM. The virtues of a deliberately mis-specified disease model in demonstrating a gene–environment interaction. Occup Environ Med. 2009;66(6):374–80.
Terry PD, Rohan TE. Cigarette smoking and the risk of breast cancer in women a review of the literature. Cancer Epidemiol Biomark Prev. 2002;11(10):953–71.
Xue F, Willett WC, Rosner BA, Hankinson SE, Michels KB. Cigarette smoking and the incidence of breast cancer. Arch Intern Med. 2011;171(2):125–33.
Acknowledgements
The authors thank Dr. Chris Bajdik for his contributions to the study. The authors also thank Dr. Linda Warren (Screening Mammography Program of BC), Dr. Philip Switzer (Greig Associates), Caroline Speers (Breast Cancer Outcomes Unit, BC Cancer Agency), Agnes Bauzon, Alegria Imperial, Betty Hall, Lina Hsu, Maria Andrews and Teresa Pavlin for their assistance with participant recruitment and data collection in Vancouver. We also thank Dr. Ross Walker, Dr. Ralph George, Celine Morissette, Jane Warner, Hilary Rimmer, Meghan Hamel and Annie Langley for assistance with participant recruitment and data collection in Kingston. Funding for this study was provided by a grant from the Canadian Institutes of Health Research (Funding Reference #: 69036).
Funding
Funding for this study was provided by a grant from the Canadian Institutes of Health Research (Funding Reference #: 69036).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
Ethics approval was provided by the University of British Columbia/BC Cancer Research Ethics Board (R04-0142) and the Queen’s University Health Sciences Research Ethics Board (EPID156-03).
Informed consent
Informed consent was obtained from all individual participants included in the study. All participants provided informed consent prior to their inclusion, which could be withdrawn at any time for any reason.
Human and animal right
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
About this article
Cite this article
Lee, D.G., Schuetz, J.M., Lai, A.S. et al. Interactions between exposure to polycyclic aromatic hydrocarbons and xenobiotic metabolism genes, and risk of breast cancer. Breast Cancer 29, 38–49 (2022). https://doi.org/10.1007/s12282-021-01279-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-021-01279-0