Abstract
Spider dragline fibers are predominantly made out of the major ampullate spidroins (MaSp) 1 and 2. The assembly of dissolved spidroin into a stable fiber is highly controlled for example by dimerization of its amino-terminal domain (NRN) upon acidification, as well as removal of sodium chloride along the spinning duct. Clustered residues D39, E76 and E81 are the most highly conserved residues of the five-helix bundle, and they are hypothesized to be key residues for switching between a monomeric and a dimeric conformation. Simultaneous replacement of these residues by their non-titratable analogues results in variant D39N/E76Q/E81Q, which is supposed to fold into an intermediate conformation between that of the monomeric and the dimeric state at neutral pH. Here we report the resonance assignment of Latrodectus hesperus NRN variant D39N/E76Q/E81Q at pH 7.2 obtained by high-resolution triple resonance NMR spectroscopy.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Biological context
The terminal domains of major ampullate spidroins control the assembly of dragline spider silks intermolecular protein contacts (Hinman and Lewis 1992; Xu and Lewis 1990). Upon lowering the pH and decreasing the sodium chloride concentration of the protein solution along the spinning duct, the amino-terminal domain forms a homodimer and, thereby, initiates spidroin assembly (Gaines et al. 2010; Hagn et al. 2010; Landreh et al. 2010). The sequence of NRN is highly conserved between different silk types and spider species, underlining the crucial role of NRN for controlling spidroin assembly (Bini et al. 2004; Chen et al. 2012; Garb et al. 2010; Rising et al. 2006). X-ray crystal and NMR solution structures of Euprosthenops australis NRN revealed separation of acidic and basic amino acids, prearranging the dimer in antiparallel (Askarieh et al. 2010). The negative charge cluster is composed of three highly conserved residues—D39, E76 and E81—that are hypothesized to control the pH-dependent dimerization and the simultaneous conversion into a tighter conformation, as evidenced by NMR and crystals structures. Neutralization of the acidic charge cluster suppresses the electrostatic repulsion between helices 2 and 3, which leads to their rearrangement and subsequent flattening of the dimerization interface (Bini et al. 2004; Kronqvist et al. 2014). To study the role of the acidic cluster during the conformational change, the clustered aspartic and glutamic acid residues were simultaneously substituted by their non-titratable analogues asparagine and glutamine, resulting in the NRN variant D39N/E76Q/E81Q. This variant mimics a protonated state of NRN at low pH, which reflects conditions as found close to the end of the spider’s spinneret. The majority of published research was done on E. australis MaSp1. For NRN from the black widow spider Latrodectus hesperus, solely monomeric wild type resonance assignments, but no structural coordinates or NMR distance restraints were published [Hagn et al. 2011, BioMagResBank (BMRB) accession code 17131]. Here we collected three-dimensional NMR data of D39N/E76Q/E81Q and assigned backbone as well as sidechain resonances.
Methods and experiments
Protein expression and purification
The variant L.h. MaSp1 NRN D39N/E76/E81Q was obtained by cloning MaSp1-NRN cDNA—mutated by using QuikChange® Site-Directed Mutagenesis Kit (Agilent, Santa Clara, CA, US)—together with a His6-SUMO-tag in vector pET28a (Novagen, Merck, Darmstadt, Germany). Genes were transformed to and expressed in Escherichia coli BL21 (DE3). Before IPTG induction the cells were grown to an OD600 = 0.7 in M9 minimal medium at 37 °C containing kanamycin, 15N-ammonium sulphate and 13C-glucose as exclusive nitrogen and carbon source, respectively. After 5 h of protein production, the cells were harvested by centrifugation for 12 min at 12,100g and 4 °C. Cells were opened using a Microfluidizer M-110S (Microfluidics, Westwood, MA, US) at 6.5 bar twice. Proteins were purified by Ni-NTA chromatography (HisTrap FF, GE Healthcare, Little Chalfont, UK) and size exclusion chromatography (HiLoad™ 26/60 Superdex™ 75 pg, GE Healthcare). After SUMO protease cleavage another Ni-NTA chromatography was performed to separate the tag from the protein. The protein was freeze-dried and stored at −20 °C.
NMR experiments
The NMR samples were prepared by dissolving the freeze-dried protein in 22 mM sodium phosphate buffer at a concentration of 0.6 mM, the addition of 10 % (v/v) D2O and final pH adjustment to 7.2. All NMR data were recorded on a Bruker Avance 700 and Avance II+ 600 MHz NMR spectrometer equipped with a 5 mm TCI cryo and TXI probe with Z-axis gradients, respectively. For sequential backbone assignment standard HNCA, HNCACB, CBCA(CO)NH and 15N-resolved NOESY-HSQC spectra were recorded (Sattler et al. 1999). For side chain assignment 13C-resolved aliphatic and aromatic NOESY-HSQC, as well as (H)CCH-TOCSY and NOESY spectra were acquired (Marion et al. 1989; Zuiderweg and Fesik 1989). Additionally two-dimensional 15N- and 13C-HSQC experiments were recorded regularly to check for protein stability (Mori et al. 1995). Spectral analysis, resonance assignment and imaging was done with the CCPNMR software package (Vranken et al. 2005), NMRViewJ (One Moon Scientific, Westfield, NJ, US), qtiplot (IONDEV SRL, Bucarest, Romania) and Adobe Illustrator CS3 (Adobe Systems, San Jose, CA, US).
Resonance assignment, secondary structure prediction and data deposition
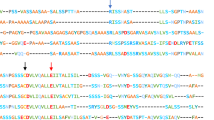
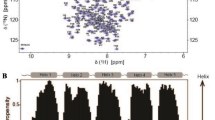
The 15N-HSQC spectrum of D39N/E76Q/E81Q showed well-dispersed signals as typical for a well-folded folded protein (Fig. 1). TALOS-N was used to perform a secondary structure estimation that indicated high helix probability for regions 12–29, 34–42, 44–54, 61–80, 85–103 and 108–125 (Shen and Bax 2013); Fig. 2). Sequential backbone assignment using standard experiments could be achieved for the majority of the chain. In sequence regions in helices 2 and 3 (39–78), where signal intensities were very weak, the assignments turned out to be more challenging. Additionally Met2, Pro8, Ile43, Pro60 and Ser61 could not be assigned. The dimerization interface could be anticipated by highlighting of region 39–78 on the homologue dimeric NRN from E. australis (PDB 2LTH, (Kronqvist et al. 2014). The modelled dimerization interface coincides with a sequence region of D39N/E76Q/E81Q that showed weak signal intensities in the NMR spectra, probably resulting from an intermediate chemical exchange of partial dimerization. C-terminal amino acids 134-137 neither showed inter-residual NOE nor sidechain proton signals. Through examination of all available types of spectra a total of 90.3 % of the backbone (HN, CA, N, HA) and 71.5 % of the total sidechain, 83 % of aromatic and 85.6 % of aliphatic side chain atoms could be assigned. 13Cα chemical shifts were compared to deposited wild type resonances (BioMagResBank accession code 17131) and summarized in Fig. 3. For most sequence positions of the protein 13Cα resonances of both variants were in good agreement. However some isolated residues showed unusual big shift differences. After careful re-evaluation of the assigned and deposited resonances with the recorded spectra and with the BMRB chemical shift statistics, some unusual chemical shifts could be observed for BMRB entry 17131. However, given that there is no access to the wild type spectra, a direct comparison of the wild type and variant three-dimensional data is currently not possible. The resonances and assignments of NRN D39N/E76Q/E81Q have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu) under the accession code 25643.
a Amino acid sequence of NRN variant D39N/E76Q/E81Q with predicted secondary structure from a chemical shift index using assigned HA, HN, CA resonances. b TALOS-N predicted helix probabilities (black bars) and secondary structure (top bar c = coil, L = loop, H = helix), estimated from assigned chemical shifts with associated confidence levels as grey vertical lines (Shen and Bax 2013)
References
Askarieh G et al (2010) Self-assembly of spider silk proteins is controlled by a pH-sensitive relay. Nature 465:236–238
Bini E, Knight DP, Kaplan DL (2004) Mapping domain structures in silks from insects and spiders related to protein assembly. J Mol Biol 335:27–40
Chen GF et al (2012) Full-length minor ampullate spidroin gene sequence. PLoS One 7(12):e52293
Gaines WA, Sehorn MG, Marcotte WR (2010) Spidroin N-terminal domain promotes a pH-dependent association of silk proteins during self-assembly. J Biol Chem 285:40745–40753
Garb JE, Ayoub NA, Hayashi CY (2010) Untangling spider silk evolution with spidroin terminal domains. BMC Evol Biol 10:243
Hagn F, Eisoldt L, Hardy JG, Vendrely C, Coles M, Scheibel T, Kessler H (2010) A highly conserved spider silk domain acts as a molecular switch that controls fibre assembly. Nature 465:239–242
Hagn F, Thamm C, Scheibel T, Kessler H (2011) pH-dependent dimerization and salt-dependent stabilization of the N-terminal domain of spider dragline silk—implications for fiber formation. Angew Chem Int Ed 50:310–313
Hinman MB, Lewis RV (1992) Isolation of a clone encoding a second dragline silk fibroin. Nephila clavipes dragline silk is a two-protein fiber. J Biol Chem 267:19320–19324
Kronqvist N et al (2014) Sequential pH-driven dimerization and stabilization of the N-terminal domain enables rapid spider silk formation. Nat Commun 5:3254
Landreh M et al (2010) A pH-dependent dimer lock in spider silk protein. J Mol Biol 404:328–336
Marion D, Driscoll PC, Kay LE, Wingfield PT, Bax A, Gronenborn AM, Clore GM (1989) Overcoming the overlap problem in the assignment of 1H NMR spectra of larger proteins by use of three-dimensional heteronuclear 1H-15 N Hartmann-Hahn-multiple quantum coherence and nuclear Overhauser-multiple quantum coherence spectroscopy: application to interleukin 1 beta. Biochemistry 28:6150–6156
Mori S, Abeygunawardana C, Johnson MO, Vanzijl PCM (1995) Improved sensitivity of Hsqc spectra of exchanging protons at short interscan delays using a new fast Hsqc (Fhsqc) detection scheme that avoids water saturation. J Magn Reson Ser B 108:94–98
Rising A, Hjalm G, Engstrom W, Johansson J (2006) N-terminal nonrepetitive domain common to dragline, flagelliform, and cylindriform spider silk proteins. Biomacromolecules 7:3120–3124
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution empolying pulsed field gradients. Prog NMR Spectrosc 34:93–158
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241
Vranken WF et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Xu M, Lewis RV (1990) Structure of a protein superfiber: spider dragline silk. Proc Natl Acad Sci 87:7120–7124
Zuiderweg ER, Fesik SW (1989) Heteronuclear three-dimensional NMR spectroscopy of the inflammatory protein C5a. Biochemistry 28:2387–2391
Acknowledgments
Funding was obtained from the Deutsche Forschungsgemeinschaft DFG (SFB 840).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical standards
The authors declare that the experiments in this study comply with the current laws of the country in which they were performed.
Rights and permissions
About this article
Cite this article
Schaal, D., Bauer, J., Schweimer, K. et al. Resonance assignment of an engineered amino-terminal domain of a major ampullate spider silk with neutralized charge cluster. Biomol NMR Assign 10, 199–202 (2016). https://doi.org/10.1007/s12104-016-9666-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-016-9666-y