Abstract
Purpose
The response rate of first-line fluoropyrimidine-based regimens for metastatic colorectal cancer (mCRC) is generally less than 50 %. The down-regulation of miR-197 in colorectal cancer cells after exposure to 5-fluorouracil might be related to the mechanism of resistance to fluoropyrimidine-based chemotherapy. So we investigated the regulatory mechanism of miR-197 on 5-FU sensitivity.
Methods
Dual luciferase reporter gene construct and dual luciferase reporter assay were used to identify the target of miR-197. TYMS expression was evaluated by immunohistochemistry staining. 5-Fu resistance of colorectal cancer cell lines was detected by MTS assay. The expression of miR-197 was detected by real time PCR.
Results
A luciferase assay and western blot analysis confirmed that miR-197 directly binds to and negatively regulates TYMS expression. Overexpressing miR-197 could increase the sensitivity of colorectal cancer cells to 5-fluorouracil (5-FU). The expression of miR-197 negatively correlated with TYMS expression in cancerous tissues from patients with stage IV colorectal cancer.
Conclusion
miR-197 mediates the response of colorectal cancer cells to 5-FU by regulating TYMS expression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Colorectal cancer is the most frequently diagnosed malignancy of the gastrointestinal tract. It is the third most common cancer in males and the second most common in females globally, with an annual incidence of more than 1.2 million cases [1]. The 5-year survival rate of patients with advanced colorectal cancer (stage IV and unresectable stage IIIc colorectal cancer) is less than 12 %. Approximately, 20 % of patients with colorectal cancer already bear distant metastases at the time of diagnosis. Moreover, up to 50–60 % of patients eventually develop recurrent disease and metastasis. Despite the rapid development of novel targeted therapies, fluoropyrimidine, either in combination or as a single agent, remains the mainstay of treatment strategies for disseminated, metastatic, colorectal cancer. However, the response rate of combined chemotherapy and fluoropyrimidine-containing regimens is less than 50 %. Several issues need to be elucidated: First, the patients who would benefit most from a 5-fluorouracil (5-FU)-containing regimen need to be identified. Second, the molecular mechanism that is involved in the process of 5-FU resistance needs to be delineated. Because personalized medicine is rapidly being more widely adopted, recognizing differences between patients that present with identical location, histology and stage is crucial. Discovering specific biomarkers of chemotherapy response is a primary step toward implementing individualized treatment.
Thymidylate synthase (TYMS) is one of key enzymes of the 5-FU catabolic pathway, and this enzyme has been associated with the response to 5-FU-based therapy. Among them, the TYMS activity is the best predictor of the sensitivity to 5-FU. In the early 1980s, TYMS over-expression was already found to be involved in the resistance to 5-FU in human cell lines. A meta-analysis of 24 studies that included 1112 patients demonstrated that advanced CRC patients with a low expression of TYMS were more sensitive to fluoropyrimidine-based chemotherapy, with TYMS expression having a better predictive value in metastases than in primary lesions [2].
Recently, microRNAs have drawn considerable attention in cancer biology. MiRNAs are ~22 nucleotide, endogenous, non-coding RNAs that can induce translational repression and mRNA degradation by binding to complementary sequences in the 3′ untranslated regions (3′UTR) of the targeted mRNAs. MiRNAs are not prone to degradation due to their small size, and they are easily isolated and detected in paraffin-embedded tissue using quantitative real-time PCR or in situ hybridization. Thus, they are promising biomarkers for predicting drug efficacy. Our previous studies have identified six miRNAs that were down-regulated in colorectal cancer cell lines after exposure to 5-FU. A further bioinformatic analysis (Targetscan: http://www.targetscan.org/) clarified that only miR-197 contained a binding site at the 3′UTR of the TYMS gene. In this study, we investigated the regulatory role of miR-197 in TYMS expression and its impact on 5-FU sensitivity. Next, we examined its relevance to TYMS expression in the cancerous tissues of colorectal patients.
Method
Tumor cell culture
HCT8 (ileocecal colorectal adenocarcinoma), HCT116 (colorectal carcinoma), and SW480 (Dukes’ type B, colorectal adenocarcinoma) cells were purchased from Cell Resource Center, IBMS, CAMS/PUMC). They were maintained in DMEM medium (Gibco, Paisley, UK) supplemented with 10 % fetal calf serum (FCS, Gibco, Paisley, UK) in a humidified 5 % CO2/95 % atmosphere at 37 °C.
miRNA and siRNA transfection
The synthetic miR-197 mimic miR-197 inhibitor, mimic control and inhibitor control were purchased from GenePharma (GenePharma Inc., Shanghai, China), and their sequences are listed in Supplementary Table 1. The siRNAs for TYMS and siRNA control were synthesized (Invitrogen Inc.), and the sequences are listed in Supplementary Table 2.
5-FU treatment and cytotoxicity assay
Twenty-four hours after transfection with the miR-197 mimic, mimic control, miR-197 inhibitor and inhibitor control or TYMS siRNA and siRNA control, the CRC cells were detached with trypsin and seeded in 96-well plates (2 × 103/well). After 24 h, an additional 100 μL of medium containing 5-FU was added, and the cells were cultured for an additional 5 days. The concentration of 5-FU was 1 μg/mL. The cell viability was measured with a Cell Titer 96 Aqueous assay kit (Promega) according to the manufacturer’s instructions on days 1, 3 and 5.
In vitro cell proliferation assay
HCT8, HCT116, and SW480 cells were transfected with the miR-197 mimic, mimic control, miR-197 inhibitor and inhibitor control, detached with trypsin and seeded in 96-well plates (2 × 103/well). The cell proliferation was assessed with the CellTiter 96® Aqueous kit (Promega, Madison, WI, USA) on days 0, 2 and 5.
In vitro cell cycle assay
HCT8 cells transfected with the miR-197 mimic, mimic control, miR-197 inhibitor and inhibitor control were cultured for 3 days, harvested and quantified. One million tumor cells were fixed with 70 % cold ethanol at 4 °C for 30 min, washed twice with PBS, and stained with 50 μg/ml PI (Sigma, USA) at room temperature for 5 min. The data were analyzed with the ModFIT software.
RNA reverse transcription and qRT-PCR
The total RNA was extracted using the Trizol total RNA isolation reagent (Invitrogen) and purified with the Column DNA Erasol kit (TIANGEN, Beijing, China) according to the manufacturers’ instructions. The mRNA expression levels were assessed with qRT-PCR using SYBR Green I (TaKaRa, Dalian, China). The gene expression level was normalized to the endogenous reference gene GAPDH. The experiments were performed in triplicate. The mRNA primers for qRT-PCRare listed in supplementary Table 3. The primers for miR-197 and U6 were purchased from QIAGEN (The sequence were listed in supplementary Table 4). The miRNAs were reverse transcribed with the miScript Reverse Transcription Kit (QIAGEN, Duesseldorf, Germany). The expression of mature miRNAs was measured using miRNA-specific quantitative qRT-PCR (TaKaRa, Dalian, China). The expression levels were normalized to the U6 endogenous control and calculated using the comparative Ct (△△Ct) method.
Western blot analysis
After being washed twice with PBS, the cells were lysed in ice-cold radio immunoprecipitation assay (RIPA) lysis buffer (Beyotime, Nanjing, China) and manually scraped from the culture plates. The proteins were separated using 10 % sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), transferred to a polyvinylidene difluoride (PVDF) membrane and incubated with anti-TYMS antibody (1/1000; Santa Cruz Biotechnology, Santa Cruz, CA) or anti-GAPDH antibody (1/2000; Santa Cruz Biotechnology, Santa Cruz, CA), followed by incubation with a secondary anti-rabbit or anti-mouse horseradish peroxidase-conjugated antibody (1/3000; Santa Cruz Biotechnology, Santa Cruz, CA). The antibody-antigen complexes were detected using a chemiluminescent ECL reagent (Millipore).
Dual luciferase reporter gene construct and dual luciferase reporter assay
A fragment of the TYMS 3′UTR that contained the predicted binding site for hsa-miR-197 and flanking sequence on each side was synthesized with a short extension that contained cleavage sites for XbaI (5′ end) and NotI (3′ end); a second fragment that contained a mutated sequence of the binding site was also synthesized. The two constructs were termed WT (TYMS-wild type) and MT (TYMS-mutant). The fragments were cloned into the pRL-TK vector (Promega Corporation, Madison, WI) downstream of the renilla luciferase reporter gene. The sequences of the miR-197 binding site and mutant site are shown in Supplementary Table 5. Each vector, along with 100 ng of pGL3 and 200 nmol/L miR-197 mimic or mimic control, was transfected into 293T cells using Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. The cells were harvested 24 h after transfection and assayed for renilla and firefly luciferase activity using the dual-luciferase reporter assay system (Promega, Madison, WI, USA).
Clinical samples
Fourteen paraffin-embedded tissue samples of primary cancer were obtained from patients with stage IV colorectal cancer (according to AJCC Cancer Staging Manual, 7th Edition) [Edge SB, Byrd DR, Compton CC, Fritz AG, Greene FL, Trotti A, editors. AJCC cancer staging manual (7th ed.). New York, NY: Springer; 2010] who underwent palliative surgery at the Peking Union Medical College Hospital between 2004 and 2009. Eligibility included metastatic colorectal cancer with primary tumor resected and measurable disease according to the revised response evaluation criteria in solid tumors (RECIST) guideline (version 1.1) [3] after surgery. Eligible patients were treated with four cycles of the mFOLFOX-6 regimen, which consisted of 85 mg/m2 i.v. oxaliplatin over 2 h concurrent with 400 mg/m2 leucovorin, followed by 400 mg/m2 5-FU as an i.v. bolus and a subsequent administration of 2400 mg/m2 5-FU over 46 h. Each treatment cycle was 2 weeks. The response and progression were evaluated every 8 weeks using the RECIST guidelines. The basic parameters of patients are listed in Supplementary Table 6.
The TYMS expression and microsatellite instability were evaluated using immunohistochemistry. Briefly, the 5-μm sections were deparaffinized, followed by antigen retrieval via heat-induced epitope retrieval with 10 mM citrate buffer. The sections were incubated with anti-TYMS (1/100, Santa Cruz Biotechnology, Santa Cruz, CA), anti-MLH1 (1/100; Santa Cruz Biotechnology, Santa Cruz, CA), anti-PMS2 (1/100; Santa Cruz Biotechnology, Santa Cruz, CA), anti-MSH2 (1/100; Santa Cruz Biotechnology, Santa Cruz, CA), anti-MSH6 (1/100, Gene tex, CA). The first antibody was detected using the avidin–biotin-peroxidase technique (Dako LSAB Kit, Dako). A pathologist determined the expression levels of TYMS, MLH1, MSH2, PMS2 and MSH6. The classification of “− or +” was defined by the percentage of TYMS positive cells at the levels of <10 % or 10–100 %, respectively. The classification of “− or +” was defined by the percentage of MLH1, MSH2, PMS2 and MSH6 at the levels of ≤1 % or 1–100 %, respectively. Negative immunohistochemical expression for at least one of these MMR proteins was defined as microsatellite instability.
Statistical analysis
Comparisons between groups were analyzed using t tests (two-sided). Differences with p values of less than 0.05 were considered significant. The correlation between miR-197 and TYMS expression was assessed with the SPSS assay and correlate bivariate Kendall is tau-b assay.
Result
miR-197 directly inhibited TYMS expression via its 3′UTR
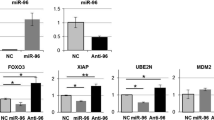
Our previous studies have identified six miRNAs that are significantly down-regulated in colorectal cancer cell lines after exposure to 5-FU [4], but the involvement of these miRNAs in 5-FU sensitivity s undefined. We used bioinformatics analysis to identify microRNA targets. A conserved binding site of miR-197 was observed in the TYMS 3′UTR (Fig. 1a). To test whether TYMS is a target of miR-197, we conducted a standard luciferase reporter assay in 293T cells. The 293T cells were transfected with the luciferase construct TYMS-WT or TYMS-MT, along with the internal control vector pGL3 and either the miR-197 mimic or the mimic control. The cells were harvested after 48 h and analyzed for dual luciferase activity. The results show that the renilla luciferase activity in TYMS-WT-transfected cells decreased by more than 40 % in the presence of co-transfected miR-197 mimic compared to cells that were co-transfected with the mimic control. In addition, the site-directed mutation of the seed region offset the inhibitory effect of the miR-197 mimic (Fig. 1b). To determine the ability of miR-197 to regulate the expression of TYMS in HCC, we measured the protein levels of TYMS in HCT8 and SW480 cells that were transfected with the miR-197 mimic, mimic control, miR-197 inhibitor or the inhibitor control. The results showed that the TYMS protein level increased after transfection with the miR-197 inhibitor, and TYMS was down-regulated after transfection with the miR-197 mimic compared to the control groups (Fig. 1c, d).
TYMS is a direct target of miR-197. a The putative binding site for miR-197 in the 3′UTR of TYMS was revealed by Target Scan (http://www.targetscan.org/). b The miR-197 binding site on TYMS 3′UTR was confirmed with a luciferase assay in 293T cells after co-transfection with (1) a plasmid containing a fragment of TYMS 3′UTR that included either the wild type or mutant predicted miR-197 binding site and (2) the miR-197 mimic or the mimic control. The data represent the mean ± SD of at least three independent experiments. *p < 0.01. c, d Western blot assay showed increased TYMS expression in HCT8 and SW480 cells after transfection with the miR-197 inhibitor and miR-197 mimic (800 nM)
miR-197 sensitize colorectal cancer cells to 5-FU treatment
Because miR-197 modulates TYMS protein expression by directly targeting the 3′UTR of TYMS mRNA, we hypothesized that miR-197 might impact the sensitivity of colorectal cancer cells to 5-FU. 5-FU was added 24 h after the interference of miR-197 expression (the interference efficacy is shown in supplementary Fig 1) in HCT8 cells. Five days after the exposure to 5-FU, the MTS assay showed that a larger number of cells transfected with miR-197 inhibitor survived compared to control-transfected cells, and the OD values were 1.19 ± 0.56 vs. 0.48 ± 0.14, respectively (p < 0.05, Fig. 2a). In contrast, the overexpression of miR-197 enhanced the cytotoxic effect of 5-FU compared to the control, and the OD value was 0.84 ± 0.17 vs. 1.28 ± 0.18 (p < 0.05; Fig. 2b). Similar results were obtained in HCT116 and SW480 cells (Supplementary Figs. 2, 3).
Effect of miR-197 on the sensitivity of HCT8 cells to 5-FU chemotherapy. Transfected cells were detached with trypsin and seeded in 96-well plates (2 × 103/well) after 24 h, an additional 100 μL of medium containing 5-FU was added, and the cells were cultured for an additional 5 days. The cell viability was measured with a CellTiter 96 Aqueous assay kit on days 1, 3 and 5. a HCT8 cells were transfected with miR-197 inhibitor and inhibitor control. b HCT8 cells were transfected with miR-197 mimic and mimic control
miR-197 did not influence proliferation and cell cycle of colorectal cell lines
Boni V et al. revealed that miR-192 and miR-215 not only modulated the expression of TYMS but also played an important role in regulating the cell cycle and proliferation of colorectal cancer cells [5]. Thus, miR-197 may affect cell proliferation and the cell cycle. MiR-197 did not significantly correlate with proliferation and the cell cycle in HCT8 cells transfected with miR-197 mimic, miR-197 inhibitor and respective control [supplementary Fig 4 A, B]. Functional assays conducted in HCT116 and SW480 cell lines produced similar results (As shown in supplementary Fig 4 C, D).
The correlation of TYMS expression with chemosensitivity to 5-FU
Resistance to fluoropyrimidine has been associated with TYMS gene amplification and increased TYMS protein expression. Our studies showed that the inhibition of TYMS expression (the interference efficacy is shown in Fig. 3a) augmented the chemosensitivity of the colorectal cancer cell line HCT8 to 5-FU, which agreed with previous studies (as shown in Fig. 3b).
Effect of TYMS on the sensitivity of HCT8 cells to 5-FU chemotherapy. a The Western blot assay showed that TYMS was down-regulated in HCT8 cells transfected with the TYMS siRNA. b HCT8 cells transfected with TYMS SiRNA or control were detached with trypsin and seeded in 96-well plates (2 × 103/well). After 24 h, an additional 100 μL of medium containing 5-FU was added, and the cells were cultured for an additional five days. The cell viability was measured with a CellTiter 96 Aqueous assay kit on days 1, 3 and 5
The correlation between miR-197 and TYMS expression in colorectal cancer tissues
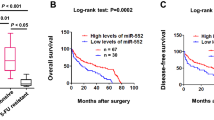
To define the relationship between miR-197 and TYMS expression in colorectal cancer tissues, we immunohistochemically stained formalin-fixed and paraffin-embedded (FFPE) tissues from stage IV colorectal cancer patients who received FOLFOX regimen as a first-line chemotherapy for TYMS (as shown in Fig. 4a, b). Next, we measured the miR-197 expression in cancerous tissues using quantitative PCR assays in seven TYMS-positive and seven TYMS-negative patients. A significantly negative correlation was observed between TYMS and miR-197 expression (p < 0.05). The expression level of miR-197 in TYMS-negative patients was significantly higher than that in TYMS-positive patients (p < 0.05, Fig. 4c).
The relationship between TYMS and miR-197 in CRC patients. a, b Immunohistochemistry of TYMS, a is negative, b is positive; c the relationship between the expression of miR-197(real time PCR data) and TYMS (ISH data). d The relationship between the expression of miR-197 and the response to FOLFOX in CRC patients. PR partial response, PD progressive disease
However, the expression level of miR-197 was not significantly associated with the efficacy of FOLFOX treatment. Although we observed a relatively low level of miR-197 expression in patients with progressive diseases and a higher expression level in patients who partially responded to the FOLFOX regimen, the difference did not reach statistical significance (p > 0.05, Fig. 4d). The conclusion is not sufficiently persuasive, and several aspects need to be clarified. First, the FOLFOX combination regimen, which consists of 5-FU, leucovorin and oxaliplatin, is the standard first-line chemotherapy for adjuvant and metastatic colorectal cancers. The association was likely confounded by the use of oxaliplatin. Second, this study only included a limited number of stage IV colorectal patients. Most cancerous tissues of metastatic colorectal cancer patients were not available because the opportunities to surgically resect the primary site were limited. Moreover, patients with stage II and III cancer who accepted adjuvant chemotherapies of 5-FU-based regimens were excluded due to a lack of measurable or assessable disease sites. Third, miR-197 may target multiple other genes that are involved in the processes of carcinogenesis and drug-related metabolism, leading to a somewhat inconclusive result.
The correlation between microsatellite instability phenotype with miR-197 or TYMS expression in colorectal cancer tissues
We detected microsatellite instability phenotype status in 14 patients using immunohistochemistry for MLH1, MSH2, MSH6 and PMS2 proteins in paraffin-embedded tissues. We found 4 patients with MSI (microsatellite instability) and 10 patients with MSS(microsatellite stability) (Supplementary Table 6 and Supplementary Fig. 5). Microsatellite instability phenotype of these patients was not significantly associated with benefit from chemotherapy. Besides, we also found that MSI was neither associated with the expression of TYMS, nor the expression of miR-197.
Discussion
Recently, several miRNAs have been implicated in the resistance to fluoropyrimidine. miR-21 was found to promote the resistance of tumor cells to 5-FU by targeting human mutS homolog 2(hMSH2), one of the members of the core mismatch repair (MMR) recognition protein complex [6, 7]. MiR-23 expression was also discovered to undermine 5-FU-induced apoptosis by regulating apoptosis-activating factor-1 (APAF-1)/caspase-9 apoptotic pathway [8]. On the contrary, other miRNAs, such as miR-34a [9], miR-122 [10], and miR-129 [11], were revealed to enhance the cytotoxic effect of 5-FU via the down-regulation of Sirt1 and E2F3 [12], PKM2 [13], and B cell lymphoma 2 (BCL2) [14], respectively. These miRNAs mainly exert their functions by affecting the apoptosis signaling pathway. Moreover, clinical studies indicated that miR-19a [15], miR-148a [16], and miR-150 [17] may predict the therapeutic outcome of fluoropyrimidine treatment, but the regulatory mechanisms involved remain unknown. Furthermore, several miRNAs reportedly influence the sensitivity to fluoropyrimidine-based chemotherapy by regulating the expression of TYMS. Boni V et al. identified that miR-192 and miR-215 might play a part in 5-FU sensitization in colorectal cancer cell lines [5]. Gotanda K et al. observed that the overexpression of miR-433 significantly reduced the levels of TYMS mRNA and protein in cervical cancer HeLa cells, which increased the sensitivity to 5-FU [18]. Finally, the association of genetic polymorphisms at the 3′UTR of TYMS gene with chemotherapeutic efficacy offered additional indirect evidence that specific miRNAs likely modulate the expression of TYMS [19].
The study findings regarding the function of miR-197 as an oncogene or tumor suppressor gene in different types of cancer are inconclusive. Recently, miR-197 has been identified as an oncomiR in hepatocellular carcinoma [20], non-small-cell lung cancer [21], pancreatic cancer [22] and male breast cancer [23]. However, other studies revealed that miR-197 was down-regulated in esophageal cancer [24], oral carcinoma [25], gastric carcinoma [26], osteosarcoma [27] and astrocytomas [28], and might exert an anti-cancer effect on these cancers. This discrepancy may be attributed to the distinct miRNA signatures of organ/tissue-specificity and temporal expression patterns. At present, the underlying mechanism of miR-197 involvement in colorectal cancer remains unknown. Regarding the correlation between miR-197 and drug resistance, miR-197 was reported to be significantly down-regulated in head and neck squamous cell lines with multidrug cross-resistance induced by docetaxel, but the molecular mechanism remains unclear [29]. Moreover, the mechanisms underlying the resistance to a single agent may considerably differ from that of multidrug resistance, which is mainly caused by the overexpression of ABC transporters. To the best of our knowledge, this study demonstrated for the first time that TYMS is a novel, direct target of miR-197. Because increased TYMS expression in colorectal cancer is well established to be closely linked to resistance to fluoropyrimidines, we analyzed the impact of miR-197 on the resistance to 5-FU in colorectal cancer cell lines and patients.
Microsatellite instability phenotype was detected in the 14 patients. However, MSS/MSI status of these patients was not significantly associated with benefit from chemotherapy. Some previous studies had reported that patients with MSI and stage II tumor might not benefit from 5-FU single-agent therapy. Patients included in this study were diagnosed with stage IIIC or stage IV colorectal cancer. Our study found no correlation of MSI with efficacy of chemotherapy, which is in accordance with previous studies. Besides, we also found that MSI was neither associated with the expression of TYMS, nor the expression of miR-197.
In conclusion, we demonstrated that miR-197 increases the sensitivity of colorectal cancer cells to the cytotoxic effect of 5-FU by regulating TYMS expression and that the miR-197 level negatively correlated with TYMS expression in cancerous tissues. Our findings highlight the potential value of miR-197 as a predictive tool for assessing the response to 5-FU treatment and also suggest that the overexpression of miR-197 may be a useful therapeutic strategy to reverse the resistance to 5-FU. Additional studies should be conducted to identify other possible targets of miR-197 and their involvement in colorectal cancer.
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61(2):69–90.
Qiu LX, Tang QY, Bai JL, Qian XP, Li RT, Liu BR, et al. Predictive value of thymidylate synthase expression in advanced colorectal cancer patients receiving fluoropyrimidine-based chemotherapy: evidence from 24 studies. Int J Cancer. 2008;123(10):2384–9.
Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer. 2009;45(2):228–47.
Zhou J, Zhou Y, Yin B, Hao W, Zhao L, Ju W, et al. 5-Fluorouracil and oxaliplatin modify the expression profiles of microRNAs in human colon cancer cells in vitro. Oncol Rep. 2010;23(1):121–8.
Boni V, Bitarte N, Cristobal I, Zarate R, Rodriguez J, Maiello E, et al. miR-192/miR-215 influence 5-fluorouracil resistance through cell cycle-mediated mechanisms complementary to its post-transcriptional thymidilate synthase regulation. Mol Cancer Ther. 2010;9(8):2265–75.
Deng J, Lei W, Fu JC, Zhang L, Li JH, Xiong JP. Targeting miR-21 enhances the sensitivity of human colon cancer HT-29 cells to chemoradiotherapy in vitro. Biochem Biophys Res Commun. 2014;443(3):789–95.
Valeri N, Gasparini P, Braconi C, Paone A, Lovat F, Fabbri M, et al. MicroRNA-21 induces resistance to 5-fluorouracil by down-regulating human DNA MutS homolog 2 (hMSH2). Proc Natl Acad Sci USA. 2010;107(49):21098–103.
Shang J, Yang F, Wang Y, Wang Y, Xue G, Mei Q, et al. MicroRNA-23a antisense enhances 5-fluorouracil chemosensitivity through APAF-1/caspase-9 apoptotic pathway in colorectal cancer cells. J Cell Biochem. 2014;115(4):772–84.
Akao Y, Noguchi S, Iio A, Kojima K, Takagi T, Naoe T. Dysregulation of microRNA-34a expression causes drug-resistance to 5-FU in human colon cancer DLD-1 cells. Cancer Lett. 2011;300(2):197–204.
He J, Xie G, Tong J, Peng Y, Huang H, Li J, et al. Overexpression of microRNA-122 re-sensitizes 5-FU-resistant colon cancer cells to 5-FU through the inhibition of PKM2 in vitro and in vivo. Cell Biochem Biophys. 2014;70(2):1343–50.
Karaayvaz M, Zhai H, Ju J. miR-129 promotes apoptosis and enhances chemosensitivity to 5-fluorouracil in colorectal cancer. Cell Death Dis. 2013;4:e659.
Akao Y, Noguchi S, Iio A, Kojima K, Takagi T, Naoe T. Dysregulation of microRNA-34a expression causes drug-resistance to 5-FU in human colon cancer DLD-1 cells. Cancer Lett. 2011;300(2):197–204.
He J, Xie G, Tong J, Peng Y, Huang H, Li J, et al. Over expression of microRNA-122 re-sensitizes 5-FU-resistant colon cancer cells to 5-FU through the inhibition of PKM2 in vitro and in vivo. Cell Biochem Biophys. 2014;70(2):1343–50.
Karaayvaz M, Zhai H, Ju J. miR-129 promotes apoptosis and enhances chemosensitivity to 5-fluorouracil in colorectal cancer. Cell Death Dis. 2013;4:e659.
Chen Q, Xia HW, Ge XJ, Zhang YC, Tang QL, Bi F. Serum miR-19a predicts resistance to FOLFOX chemotherapy in advanced colorectal cancer cases. Asian Pac J Cancer Prev. 2013;14(12):7421–6.
Takahashi M, Cuatrecasas M, Balaguer F, Hur K, Toiyama Y, Castells A, et al. The clinical significance of MiR-148a as a predictive biomarker in patients with advanced colorectal cancer. PLoS One. 2012;7(10):e46684.
Ma Y, Zhang P, Wang F, Zhang H, Yang J, Peng J, et al. miR-150 as a potential biomarker associated with prognosis and therapeutic outcome in colorectal cancer. Gut. 2012;61(10):1447–53.
Gotanda K, Hirota T, Matsumoto N, Ieiri I. MicroRNA-433 negatively regulates the expression of thymidylate synthase (TYMS) responsible for 5-fluorouracil sensitivity in HeLa cells. BMC Cancer. 2013;13:369.
Shen R, Liu H, Wen J, Liu Z, Wang LE, Wang Q, et al. Genetic polymorphisms in the microRNA binding-sites of the thymidylate synthase gene predict risk and survival in gastric cancer. Mol Carcinog. 2014.
Dai W, Wang C, Wang F, Wang Y, Shen M, Chen K, et al. Anti-miR-197 inhibits migration in HCC cells by targeting KAI 1/CD82. Biochem Biophys Res Commun. 2014;446(2):541–8.
Fiori ME, Barbini C, Haas TL, Marroncelli N, Patrizii M, Biffoni M, et al. Antitumor effect of miR-197 targeting in p53 wild-type lung cancer. Cell Death Differ. 2014;21(5):774–82.
Hamada S, Satoh K, Miura S, Hirota M, Kanno A, Masamune A, et al. miR-197 induces epithelial-mesenchymal transition in pancreatic cancer cells by targeting p120 catenin. J Cell Physiol. 2013;228(6):1255–63.
Lehmann U, Streichert T, Otto B, Albat C, Hasemeier B, Christgen H, et al. Identification of differentially expressed microRNAs in human male breast cancer. BMC Cancer. 2010;10:109.
Wang TY, Liu SG, Zhao BS, Qi B, Qin XG, Yao WJ. Implications of microRNA-197 downregulated expression in esophageal cancer with poor prognosis. Genet Mol Res. 2014;13(3):5574–81.
Scapoli L, Palmieri A, Lo ML, Pezzetti F, Rubini C, Girardi A, et al. MicroRNA expression profiling of oral carcinoma identifies new markers of tumor progression. Int J Immunopathol Pharmacol. 2010;23(4):1229–34.
Li X, Zhang Y, Zhang H, Liu X, Gong T, Li M, et al. miRNA-223 promotes gastric cancer invasion and metastasis by targeting tumor suppressor EPB41L3. Mol Cancer Res. 2011;9(7):824–33.
Dai N, Zhong ZY, Cun YP, Qing Y, Chen C, Jiang P, et al. Alteration of the microRNA expression profile in human osteosarcoma cells transfected with APE1 siRNA. Neoplasma. 2013;60(4):384–94.
Yang C, Wang C, Chen X, Chen S, Zhang Y, Zhi F, et al. Identification of seven serum microRNAs from a genome-wide serum microRNA expression profile as potential noninvasive biomarkers for malignant astrocytomas. Int J Cancer. 2013;132(1):116–27.
Dai Y, Xie CH, Neis JP, Fan CY, Vural E, Spring PM. MicroRNA expression profiles of head and neck squamous cell carcinoma with docetaxel-induced multidrug resistance. Head Neck. 2011;33(6):786–91.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (No. 81472785, 61435001, 81101588, 30800429).
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Z. Sun and N. Zhou contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
12094_2015_1318_MOESM1_ESM.tif
Supplementary material 1 (TIFF 1004 kb). The transfection efficiency of miR-197 mimic and inhibitor in HCT8 (a, b) and HCT116 (c, d)
12094_2015_1318_MOESM2_ESM.tif
Supplementary Material 2 (TIFF 530 kb). Effect of miR-197 on the sensitivity of HCT116 cells to 5-FU chemotherapy. Transfected cells were detached with trypsin and seeded in 96-well plates (2 × 103/well) After 24 hours, an additional 100 μL of medium containing 5-FU was added, and the cells were cultured for an additional 5 days. The cell viability was measured with a CellTiter 96 Aqueous assay kit on day 5. a HCT116 cells were transfected with miR-197 inhibitor and inhibitor control. b HCT116 cells were transfected with miR-197 mimic and mimic control
12094_2015_1318_MOESM3_ESM.tif
Supplementary material 3 (TIFF 498 kb). Effect of miR-197 on the sensitivity of SW480 cells to 5-FU chemotherapy. Transfected cells were detached with trypsin and seeded in 96-well plates (2 × 103/well). After 24 h, an additional 100 μL of medium containing 5-FU was added, and the cells were cultured for an additional 5 days. The cell viability was measured with a CellTiter 96 Aqueous assay on day 5. a SW480 cells were transfected with miR-197 inhibitor and inhibitor control. b SW480 cells were transfected with miR-197 mimic and mimic control
12094_2015_1318_MOESM4_ESM.tif
Supplementary material 4 (TIFF 808 kb). miR-197 did not affect CRC cell proliferation. CRC cells were transfected with miR-197 mimic, mimic control, miR-197 inhibitor or inhibitor control. The cell numbers were determined with the MTS assay. Data represent the mean ± SD of six independent experiments. a HCT8; c HCT116; d SW480. b miR-197 had no effect on CRC cell cycle (HCT8)
12094_2015_1318_MOESM5_ESM.tif
Supplementary material 5 (TIFF 19216 kb). Immunohistochemistry of MLH1, MSH2, MSH6 and PMS2 (A, B) Immunohistochemistry of MLH1, A is negative, B is positive (C, D) Immunohistochemistry of MSH2, C is negative, D is positive (E, F) Immunohistochemistry of MSH6, E is negative, F is positive (G, H) Immunohistochemistry of PMS2, G is negative, H is positive
Rights and permissions
About this article
Cite this article
Sun, Z., Zhou, N., Han, Q. et al. MicroRNA-197 influences 5-fluorouracil resistance via thymidylate synthase in colorectal cancer. Clin Transl Oncol 17, 876–883 (2015). https://doi.org/10.1007/s12094-015-1318-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-015-1318-7