Abstract
Agrobacterium-mediated transformation (AMT) of sugarcane has been limited by low transformation efficiency, high variability between experiments and genotype specificity. We tested combinations of parameters that have been useful in other recalcitrant plant systems, aiming to develop an efficient and reproducible method. Applied to elite sugarcane cultivar Q117, key parameters were (i) minimal handling of callus near the time of co-cultivation, (ii) use of a super-binary helper vector with additional virB,C,G gene copies, and (iii) use of Agrobacterium strain AGL1. Transformation efficiency was in the range 0.5 to 3.5 stably transformed, embryogenic-callus-forming lines per gram fresh weight of co-cultivated callus, over six independent callus batches. Addition of 5 μM copper sulphate to the callus-growth medium appeared beneficial in a single further test. Following selection for aminoglycoside resistance conferred by PUbi-aphA, 87 % of transformed lines that formed embryogenic callus were regenerable to plants. Southern blot analysis of 24 transgenic lines showed 21 % with a single-copy insertion of an intact T-DNA without vector backbone, and a mean transgene copy number of 2.5. Over multiple batches, the AMT protocol approached the transformation efficiency from our routine conditions for particle bombardment of Q117. However, the same parameters were ineffective for AMT of cultivars Q208 and Q172, and yielded a lower transformation efficiency (0.02) with KQ228. As experienced in other systems such as rice, high-efficiency transformation of one recipient genotype may provide useful starting parameters for work towards AMT of additional genotypes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transformation of sugarcane, by particle bombardment of embryogenic callus and selection for antibiotic resistance conferred by an aminoglycoside phosphotransferase, is routine for many research groups internationally (Brumbley et al. 2008; Lakshmanan et al. 2005). It has proven efficient, reproducible, and substantially genotype-independent; with minor variations to selection and culture conditions for a workable efficiency of transgenic plant production (Basnayake et al. 2011). There is also substantial interest in Agrobacterium-mediated transformation (AMT) due to the potential advantages of simpler methodology and a higher proportion of simple transgene integration events.
Although grasses are not natural hosts of Agrobacterium, conditions for reliable AMT have been developed for several species including some of the major cereal crops (Cheng et al. 2004; Shrawat and Lorz 2006), Brachypodium (Vain et al. 2008) and switchgrass (Li and Qu 2011). Factors important for transformation of multiple monocot species include: (1) inoculation of actively dividing cells (typically in immature embryos or embryogenic callus); (2) use of super-binary vectors containing extra vir gene copies; and (3) optimised co-cultivation conditions, including pH, temperature and presence of a vir gene inducer such as acetosyringone (Shrawat and Lorz 2006). Recipient genotype specificity is common, but more than 40 rice genotypes have been transformed by tailoring of parameters (Hiei and Komari 2008; Nishimura et al. 2006).
The first reported transgenic sugarcane plants generated by AMT were from variety Ja 60–5 after co-cultivation of embryogenic cell suspensions, or leaf spindle sections pre-treated to reduce necrosis (Arencibia et al. 1998; Enriquez-Obregon et al. 1998). It is difficult to discern broadly applicable parameters from subsequent reports using additional sugarcane recipient genotypes. The efficiency for production of non-chimeric transgenic plants has generally been either low, not reported, or not verified by molecular confirmation of independent integrative transformants (Arvinth et al. 2010; Elliott et al. 1998; Joyce et al. 2010; Manickavasagam et al. 2004; Santosa et al. 2004; Zhangsun et al. 2007). No method for AMT of sugarcane has reportedly been extended across independent research groups or sugarcane cultivars. It is challenging to determine optimal parameters in systems with low transformation efficiency and substantial variability between batches, because large-scale and repeated experiments are required to draw reliable conclusions.
As a contribution to this effort, we set out to test combinations of parameters that have facilitated AMT in other recalcitrant plant systems. Initially we chose to work with the elite cultivar Q117. This cultivar is readily amenable to embryogenic tissue culture and widely used in studies of transgenes in sugarcane following microprojectile-mediated gene transfer. AMT of Q117 was reported recently (Joyce et al. 2010), using selection on MS medium for antibiotic resistance conferred by a PUbi-aphA transgene, as widely applied in microprojectile-mediated transformation (Basnayake et al. 2011; Bower et al. 1996). The efficiency was 0.1 to 0.8 transgenic plant lines per gram fresh weight (/gFW) of inoculated callus, across replicates of 100 callus pieces or 6 gFW callus (Joyce et al. 2010; P. Joyce, personal communication). Among the variables that we tested, minimal handling of callus around the time of inoculation, and the use of suitable super-binary helper vectors with Agrobacterium strain AGL1 had the greatest impact. They were combined in a method for which efficiency and reproducibility were confirmed across six callus batches tested over 16 months: yielding 0.5–3.5 (mean 1.7) independent transgenic lines/gFW of co-cultivated embryogenic callus.
Results
Preliminary Testing of Selection and Reporter Systems in Cultivar Q117
AMT systems involving plant cell culture require antibiotic treatment after co-cultivation to prevent bacterial overgrowth of the plant cultures. Timentin (containing ticarcillin and the β-lactamase inhibitor clavulanic acid) has been used as a bactericide without adverse effects on plants in culture (Costa et al. 2000; Ling et al. 1998; Nauerby et al. 1997). We found that timentin concentrations above 100 mg/L eliminated Agrobacterium from at least 90 % of co-cultivated calli within 8 weeks, provided excessive Agrobacterium concentrations that caused rapid overgrowth after co-cultivation were avoided.
Also, we found that in the presence of timentin there were more surviving non-transgenic calli after 12 weeks on some geneticin concentrations. For example, the proportion of non-transgenic calli that survived to produce green shoots after 12 weeks on media containing geneticin at 40 mg/L (G40) increased from 10 % to 68 % in the presence of timentin at 150 mg/L (T150) (P < 0.0001, t-test). At 50 mg/L geneticin, the time needed for ‘escape-free’ selection based on completely inhibited growth or necrosis of non-transformed calli was extended by 2 weeks to 10 weeks in the presence of 150 mg/L timentin. These observations are consistent with a stimulatory effect of timentin on some plant cell cultures as reported previously (Ling et al. 1998; Robert et al. 1989). Nevertheless, faster-emerging transformants were reliably selected after 6 to 10 weeks on G50 + T150 medium as described below.
In several preliminary experiments, we tested permutations of two Agrobacterium strains (LBA4404 or AGL1 at 109 cfu/mL); three helper vectors (differing in the presence of additional vir genes) based on the pGreenII/pSoup system; three callus pre-treatment regimens varying the presence of acetosyringone as used for maize AMT (D’Halluin et al. 2008) or antinecrosis compounds as used for sugarcane AMT (Enriquez-Obregon et al. 1998); and two co-cultivation periods (3 or 5 days). Luciferase (Luc) activity, from expression in sugarcane cells of a PUbi-intron-luc + gene transferred on the T-DNA, was detected in up to 16 % of calli at 3 days after co-cultivation. A single geneticin-resistant, Luc-positive, stably-transformed callus line was obtained from each of three (out of 18) treatments tested on 150–175 callus pieces per treatment. There was no apparent relationship between transient Luc activity and production of stable transformants, and no apparent advantage from any of the parameters listed above, except that all three of the stable lines were from treatments using vir-supplemented (super-binary) helper vectors.
Inoculum densities for subsequent experiments were reduced to 0.5 × 109 cfu/mL for LBA4404 and 0.25 × 109 cfu/mL for AGL1 derivatives, to minimise visible overgrowth by Agrobacterium. Because stable transformation efficiencies were very low and poorly predicted by transient expression, we designed subsequent experiments with larger amounts of treated callus and only tested for stable transgenic lines. Each treatment was typically tested on 500–1,000 embryogenic callus pieces of about 5 mm diameter.
Key Parameters from Large-Scale Experiments
In a series of 11 large-scale experiments using cultivar Q117, we further explored the effects of Agrobacterium strain, binary vector, co-cultivation medium and callus handling around the time of inoculation. The latter was motivated initially by ideas to reduce the amount of work needed to handle large numbers of callus pieces individually through the co-cultivation process. An early experiment tested inoculation by flooding of (i) undisturbed calli on their growth medium, versus (ii) calli after transfer to a sterile dish. In both treatments, 10 min after mixing with Agrobacterium, the inoculated callus pieces were transferred to co-cultivation medium. Inoculation of undisturbed calli resulted in less browning after co-cultivation and more transgenic lines after selection.
The most effective protocol from our experiments involved inoculation by flooding undisturbed calli on their growth medium, with no transfer of calli until after the 3 day co-cultivation period (‘minimal handling’). In this method, acetosyringone is supplied only in the inoculation medium carrying the Agrobacterium. Figure 1 shows significant increases in transformation efficiency obtained through minimal handling, compared to transfer of calli for co-cultivation, in two identical experiments using separately-initiated callus batches. The callus used in these experiments was last subcultured 10–18 days before inoculation. In particle bombardment and AMT protocols, it is common to subculture callus a few days before gene transfer, to obtain actively-growing material. We observed a reduction in transformation efficiency when callus was subcultured 4 days prior to inoculation. Subculturing callus immediately prior to inoculation (required when a specialised co-cultivation medium is used) reduced transformation efficiency to almost zero (Fig. 2).
Effect of handling during co-cultivation, on transformation efficiency (stably transformed, embryogenic callus-forming lines/gFW of co-cultivated embryogenic callus). Calli were inoculated by flooding with transfer of individual calli for co-cultivation (white bars) or without transfer at this time (grey bars; ‘minimal handling’). Results shown from two experiments are means with standard errors from 10 replicates of 100 callus pieces per treatment, using Agrobacterium strain EHA105 and helper vector pAL154. Treatments were significantly different at P < 0.01 (t-test)
Effect of handling before co-cultivation, on transformation efficiency (stably transformed, embryogenic callus-forming lines/gFW of co-cultivated callus). Treatments involved minimal handling, with subculture 16 days before inoculation (MH) or 4 days before inoculation (MH-4) or immediately before inoculation (MH0). Results are means with standard errors from five replicates of 100 callus pieces per treatment, using Agrobacterium strain AGL1 and helper vector pAL154. Treatment MH0 was significantly inferior to MH (P < 0.05, ANOVA with Dunnett’s post-test)
With minimal handling, the use of super-binary helper vector pAL154 containing additional virB,C,G genes significantly increased transformation efficiency relative to pSoup helper vector (P < 0.0001), as did the use of Agrobacterium strain AGL1 compared with EHA105 (P < 0.0001). There was no significant interaction between these variables by 2-way ANOVA (Fig. 3). However, other parameters may interact. For example: (i) the advantage of super-binary vectors was not apparent when calli were handled after inoculation by flooding (P > 0.05, Mann–Whitney test); (ii) EHA105 outperformed AGL1 in an experiment using immersion of calli after transfer (P < 0.05, Mann–Whitney test).
Effects of Agrobacterium strain and binary vector type on transformation efficiency (stably transformed, embryogenic callus-forming lines/gFW of co-cultivated embryogenic callus). Treatments involved co-cultivation with Agrobacterium strain EHA105 (white bars) or AGL1 (grey bars); using binary helper pSoup or super-binary version pAL154. Results are means with standard errors from 10 replicates of 100 callus pieces per treatment. Both strain and vector effects were significant at P < 0.0001, and there was no significant interaction (two-way ANOVA)
Reproducibility
Over six experiments using separately-initiated callus batches and various genes in the T-DNA, the same combination of parameters (minimal handling, Agrobacterium strain AGL1 and super-binary helper vector pAL154) yielded an average transformation efficiency of 1.7 (±0.4 se) stably transformed, embryogenic callus-forming lines/gFW of co-cultivated embryogenic callus. Expressed another way, the stable transformation efficiency over six experiments ranged from 3 % to 20 % of co-cultivated, 5 mm diameter callus pieces. In total, these experiments yielded more than 600 independent embryogenic callus lines after geneticin selection.
Additional Parameters
Use of filter-sterilised, callus-growth medium supplemented with 5 μM CuSO4, did not alter callus growth rate or appearance (data not shown), but transformation efficiency was increased by 70 % (Fig. 4A). A non-significant (22 %) decrease in transformation efficiency was observed after co-cultivation at 20 °C (data not shown). We have not tested these additional parameters in repeated experiments with independent callus batches.
Effect of CuSO4 supplementation of the callus growth medium on transformation efficiency (stably transformed lines/gFW of co-cultivated callus), and appearance of transformed callus on selection. A. Transformation efficiency is based on production at 10 weeks of geneticin-resistant embryogenic callus (EC) or non-embryogenic callus (NC) from co-cultivated callus produced on routine MSC1.5 medium (white bars) or medium supplemented with 5 μM CuSO4 (grey bars). Results are means with standard errors from six replicates of 100 callus pieces per treatment. * indicates treatments significantly different at P < 0.01 (t-test). B. Two actively-growing, geneticin-resistant, embryogenic calli (indicated by arrows) and the dying (non-transformed) calli from which they have emerged after 6 weeks of selection (scale bars = 5 mm)
Timing of Selection and Regeneration
Transgenic lines emerged at different rates on selection medium. In a typical experiment, of 731 geneticin-resistant, embryogenic lines selected by 10 weeks, 66 % were identified by 8 weeks, and 43 % by 6 weeks on selection medium (Fig. 4B). The additional time for regeneration of shoots with roots adequate for transfer to soil ranged from approximately 6 to 12 weeks, and 87 % of embryogenic calli regenerated plants.
Molecular Characteristics of Transgenic Lines from AMT Versus Particle Bombardment
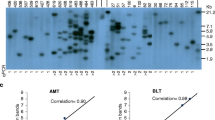
Transgene integration complexity and T-DNA integrity were evaluated by Southern blot analysis on 24 transgenic lines from AMT. Hybridisation to a luc+ probe revealed that 29 % contained single copy insertions, 79 % had 3 or fewer copies, and the mean copy number was 2.5 (Fig. 5). Six of the seven single-copy lines contained intact T-DNA. Vector backbone integration was detected in one of the six single-copy lines with intact T-DNA, and in 53 % of multi-copy lines (Fig. 6).
Southern blot analysis of sugarcane lines for T-DNA integration and integrity, probed for the luc + transgene. Genomic DNA was from non-transformed Q117 (U), and transgenic lines from AMT (A1-A24) or particle bombardment (B1-6). Digestion using HindIII or EcoRI is expected to yield a luc-hybridizing fragment of variable size depending on integration site in the plant genome. SbfI and EcoRV cut the T-DNA just inside the R and L borders to yield a 7,198 bp fragment (indicated by dashed white lines) from intact T-DNA integration. Molecular weights in bp are from standards run on the same gels
Southern blot analysis of sugarcane lines for vector backbone. Blots from Fig. 5 were stripped and reprobed using a 2,495 bp BglII fragment comprising the entire pGreenII region outside of the T-DNA borders. The restriction enzymes used to digest the genomic DNA do not cut within this vector backbone, so each hybridizing band represents a separate integration site in the plant genome. In some cases these are distinct from the T-DNA integration sites. The expected SbfI/EcoRV fragment size for intact plasmid integration after particle bombardment is 3,248 bp (no matching bands observed). Molecular weights in bp are from standards run on the same gels
For comparison, among six lines produced by particle bombardment using 3 μg of the intact binary vector DNA per shot, the mean copy number was 5.8, with one line containing a single copy insertion. All six lines contained vector backbone and showed more evidence of fragmentation or rearrangement of the transferred DNA than the AMT lines (Figs. 5 and 6).
Attempted Transformation of Additional Sugarcane Varieties
We attempted to transform additional genotypes, initially focusing on elite cultivars Q172, Q208 and KQ228 which produce different proportions of embryogenic, friable and soft callus (Basnayake et al. 2011). Using minimal handling and EHA105, one Luc-positive regenerable transgenic line was recovered from 1,000 inoculated callus pieces for KQ228. In subsequent experiments involving 2,700 KQ228 callus pieces inoculated using AGL1 or EHA105, five non-regenerable lines following AGL1 treatment were confirmed by PCR assay for the aphA selected marker or luminometer assay for Luc activity. No transgenic lines were obtained in parallel experiments involving 2,700 Q208 callus pieces inoculated using AGL1 or EHA105, or 1,560 Q172 callus pieces inoculated using AGL1.
Discussion
Factors Influencing Transformation Efficiency for Q117
Minimal handling of callus around the time of co-cultivation, combined with use of a super-binary helper vector containing additional virB,C,G genes in Agrobacterium strain AGL1 allowed reproducible and efficient AMT of elite sugarcane cultivar Q117. We obtained 0.5–3.5 independent stably transformed, embryogenic callus-forming lines/gFW of co-cultivated embryogenic callus across six independently-initiated callus batches over a period of 16 months. This is the highest reproducible efficiency reported for AMT of sugarcane, with molecular confirmation of stable independent transformants. It approaches the transformation efficiency of about 4–6 transgenic lines/gFW routinely obtained using our conditions for particle bombardment of Q117 embryogenic callus followed by geneticin selection.
The key factors confirmed over repeated experiments to significantly increase AMT efficiency in Q117 were minimal handling before and during co-cultivation (8-fold, Figs. 1 and 2), the use of a super-binary helper relative to an isogenic vector without additional vir genes (1.3 to 3.0-fold), and Agrobacterium strain AGL1 over EHA105 (1.3 to 4.2-fold). The strain effect was less compelling given contradictory results in initial experiments using different conditions. CuSO4 supplementation gave a further significant benefit (1.7-fold) that has not been retested with independent callus batches.
Minimal handling is a substantial time-saver for practitioners, but its benefit may be considered surprising as it eliminates treatments such as vacuum infiltration, deliberate wounding, transfer of callus for rapid growth, specialised co-cultivation media, and washing or blotting, which have been useful in some AMT systems. Rothrock et al. (2007) explored the necessity for wounding during AMT of American chestnut somatic embryos. They found that for one genotype minimal handling was 100-fold better than desiccation, whereas these treatments were equally effective in a second genotype. One interpretation of the increased callus browning after co-cultivation of handled callus is that mechanical stimulation potentiates the hypersensitive response against Agrobacterium in some sugarcane genotypes such as Q117.
Effects of super-binary vectors and Agrobacterium strains have been documented in other monocot crops (Shrawat and Lorz 2006). In our experiments, the helper plasmid pAL154 carrying virB,C,G was significantly better than isogenic pSoup with no added vir genes. A similar effect was observed with this helper vector in wheat (Wu et al. 2008). Strains AGL1 and EHA105 are very similar (Table 1), but it is possible that the recA mutation in AGL1 enhances stability of the binary and helper plasmids, or that the different T-region deletions in these strains have undocumented effects. We also obtained Q117 transformants using Agrobacterium strain LBA4404 and we recommend testing several Agrobacterium strains on any additional sugarcane genotype of interest.
Bartlett et al. (2008) found that copper-supplemented, filter-sterilised media improved regeneration and increased the efficiency of AMT in barley. We noticed that Agrobacterium grew faster on CuSO4-supplemented medium and it may be necessary to alter other parameters to avoid overgrowth of callus in experiments to explore the repeatability of increased transformation efficiency using this supplement.
Transfer to Additional Varieties
The ability to transform multiple sugarcane cultivars is important for biotechnological application of AMT. We repeatedly transformed the elite variety KQ228, albeit at lower efficiency than Q117. Failure to transform Q172 and Q208 shows that formation of compact embryogenic callus is not a reliable indicator of whether a cultivar is amenable to AMT. This contrasts with the experience using particle bombardment (Basnayake et al. 2011), and presumably reflects the greater biological complexity of the Agrobacterium-plant interaction (Birch 1997).
In both maize and rice, the establishment of efficient AMT protocols for one genotype provided a useful basis for experimentation with factors including Agrobacterium strain, co-cultivation conditions and culture media that led to success with other cultivars (Hiei and Komari 2008; Ishida et al. 2007; Nishimura et al. 2006). Similar approaches might be justified in sugarcane if molecular characteristics of transformants obtained using AMT prove advantageous over those obtained using tailored direct gene transfer parameters.
Transgene Integration Complexity
Transgene copy number in Agrobacterium-generated lines can be influenced by the strain, transformation method and plant target tissue (De Buck et al. 2009; Oltmanns et al. 2010). Among the 24 transgenic lines we tested by Southern blot analyses, the mean transgene copy number was 2.5, and 21 % of lines had intact, single-copy T-DNA with no vector backbone detectable by hybridisation.
Reported frequencies of vector backbone integration following AMT vary widely, depending on the resolution of the probe, the T-DNA border sequence context (De Buck et al. 2000), and possibly the vector size (Oltmanns et al. 2010). The frequency we measured using a full-length backbone probe (46 %) is similar to transgenic rice produced using the pGreenII/pSoup vector system (Vain et al. 2003; Vain et al. 2004). Like others (Vain et al. 2004; Ye et al. 2008) we observed more frequent vector backbone integration in lines containing multiple T-DNA insertions.
If a stringent requirement for transferred DNA integration simplicity were to be adopted, the efficiency for production of qualifying transformants via AMT in Q117 would still be in the range 0.1–0.7 lines/gFW of co-cultivated callus, which is sufficient for transfer of genes of commercial interest. By comparison, sugarcane lines produced via particle bombardment using 3 μg of intact binary vector DNA per shot had a higher average copy number and integration complexity (Fig. 5). The results were comparable with those in previous reports that used similar bombardment parameters (Bower et al. 1996). More work is required with particle bombardment parameters designed for simpler integration patterns (Kim et al. 2012) and in comparison of insertional sites and transgene expression stabilities, to weigh the merits of these alternative approaches.
Materials and Methods
Agrobacterium Strains and Vectors
Tables 1 and 2 summarise the key properties of tested Agrobacterium strains and vectors. We used the pGreenII/pSoup binary and helper vector system (Hellens et al. 2000) with ‘super-binary’ derivatives of the replication-helper vector pSoup modified to express additional copies of virB,C,G (pAL154) or virG (pAL155) (Wu et al. 2008).
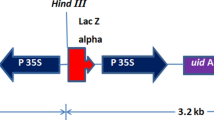
Into the T-DNA region of the minimal binary vector pGreenII we introduced an intron-containing firefly luc + gene (Bourdon et al. 2001) under the control of the maize ubi-1 promoter (PUbi; Christensen et al. 1992) and ocs terminator; and the aphA-2 (syn. nptII) selectable marker under the control of the Emu promoter (Last et al. 1991) and nos terminator (Depicker et al. 1982). This reporter vector, designated pG2LK (Fig. 7) has a T-region of 7921 nt and confers no detectable Luc activity in Agrobacterium.
Schematic map of pG2LK. Relevant restriction enzyme sites are shown. PUbi, promoter and leader including first intron from the maize ubi-1 gene; intron-luc+, luciferase+ gene containing intron 4 from the maize rpoT gene; ocs3′, octopine synthase terminator; RB, right border; ColE1 ori, pColE1 origin of replication; aphA-1, aminoglycoside phosphotransferase 1 gene; pSa ori, pSa origin of replication; LB, left border; nos3′, nopaline synthase terminator; aphA-2, aminoglycoside phosphotransferase 2 gene; PEmu, Emu promoter
In experiments to confirm that the recommended method is applicable for transfer of various expression cassettes and sequences, we used other derivatives from pGreenII with a PUbi-aphA selectable marker and various reporter or potential commercial gene constructs, with T-regions ranging from 4,143 to 8,754 nt. Similar efficiencies were obtained in all cases.
Agrobacterium strains listed in Table 1 were transformed using electroporation (Nagel et al. 1990) and grown on YEP medium (An et al. 1988) containing relevant antibiotics. For plant transformation experiments, Agrobacterium broth cultures were grown at 28 °C with shaking to an optical density at 660 nm of 0.5–1.0. Cells were pelleted by centrifugation at 6,000 × g for 10 min, then resuspended to the required density in liquid ‘co-cultivation medium’ (detailed below) supplemented with 100 μM acetosyringone.
Sugarcane Callus Growth, Transformation, Selection and Regeneration
Sugarcane tissue culture and particle bombardment for comparative analyses were performed as described previously (Basnayake et al. 2011; Bower et al. 1996). The primary callus-culture medium designated MSC1.5 comprised Murashige and Skoog basal medium (Murashige and Skoog 1962) supplemented with 1.5 mg/L 2,4-dichlorophenoxy acetic acid, 500 mg/L casein hydrolysate, 15 g/L1 sucrose, 0.1 mg/L thiamine-HCl, 0.5 mg/L pyridoxine-HCl, 0.5 mg/L nicotinic acid, 2 mg/L glycine, 100 mg/L myo-inositol, and 10 % (v/v) coconut water, solidified where relevant using 9 g/L agar (pH 6.0 before autoclaving). CuSO4-supplemented media included an additional 5 μM CuSO4, with the liquid component filter-sterilised as a 2x concentrate. The agar component was made as a 2x concentrate, autoclaved, cooled to 60 °C and combined with the liquid component pre-heated to 50 °C.
Where specified in the results, ‘co-cultivation medium’ consisted of MSC1.5 medium supplemented with 100 μM acetosyringone (pH 5.6 before autoclaving). In our recommended ‘minimal handling’ method, co-cultivation occurs on unmodified MSC1.5 medium. Resting medium included 150 mg/L timentin, freshly prepared for each batch. Selection medium included 150 mg/L timentin plus 50 mg/L geneticin for Q117 or 25 mg/L geneticin for KQ228. Regeneration medium lacked 2,4-D.
Callus pieces counted at the time of co-cultivation for subsequent calculation of transformation efficiencies were typically 5 mm in diameter, with an average weight of 57 mg. Therefore amounts of co-cultivated callus in each replicate and treatment based on visual estimation of the number of ‘standard callus piece equivalents’ can also be expressed on a fresh weight basis. Results are presented on a fresh weight basis as the best option for comparison across genotypes that vary in callus morphology or research groups using various techniques. For genotypes Q117 and Q172 the cultures comprised primarily embryogenic callus. For genotypes Q208 and KQ228, the cultures were enriched by selective subculture of embryogenic callus, but they included 20–60 % of friable and soft callus (Basnayake et al. 2011). The time from callus initiation to inoculation ranged from 8.3 to 12.3 weeks, with no discernible influence on transformation efficiency.
Various inoculation methods were tested. For ‘immersion’, callus pieces were gently separated to the required size range using fine forceps and transferred to a sterile dish before adding Agrobacterium inoculum to cover the callus. For ‘plate flooding’ the inoculum was added to immerse the callus without transfer from the callus-growth medium. In both of the above methods, the plates were then gently swirled and incubated for 10 min. The inoculum was then removed by pipette and calli were transferred to co-cultivation medium. ‘Minimal handling’ was as for ‘plate flooding’ except that the callus pieces were not transferred after removal of Agrobacterium inoculum by pipette. Variations in subculture around the time of co-cultivation are described with the results. In our recommended ‘minimal handling’ method, the callus is not subcultured within 10 days before co-cultivation.
All co-cultivations were performed in the dark at 25 °C for 3 days unless specified. Following co-cultivation, calli were transferred to resting medium for 4 days, then to selection medium and subcultured every 2 weeks. No washing or blotting steps were used. Putative transgenic lines (callus outgrowths greater than 5 mm in diameter after at least 6 weeks on selection, Fig. 4B) were transferred to regeneration medium in the light under a 16 h photoperiod of approximately 1,000 Lux provided by fluorescent lamps. A single primary shoot was taken per selected callus piece for subsequent analyses.
Luciferase Assays
Luc assays were performed by immersing tissues in a 0.4 mM luciferin solution (in filter-sterilised MSC1.5 for non-destructive assays), and recording light emission using a cooled CCD camera (Pixis, Princeton Instruments), essentially as described by Mudge et al. (1996).
DNA Extractions and Southern Blots
Genomic DNA was extracted from callus or young leaves using a CTAB method (Rogers and Bendich 1988). Digested DNA was separated by electrophoresis on a 0.8 % agarose gel (20 μg per lane) and transferred onto a nylon membrane (Hybond N+, GE Healthcare) by capillary blotting (Sambrook and Russell 2001). Probes were labelled using 32P-dCTP and random primers (Rediprime II kit, GE Healthcare), followed by removal of unincorporated nucleotides (Illustra Microspin columns, GE Healthcare). Hybridisation was performed at 42 °C in Ambion Ultrahyb solution (Applied Biosystems). Membranes were washed at high stringency (0.1 × SSC, 0.1 % SDS) for 15 min at 65 °C. Non-specific background from vector backbone probe was removed by further immersing blots in 500 ml of 0.1 % SDS heated to 95 °C and cooling to room temperature. Images captured using a Molecular Dynamics 820 Storm phosphorimager were processed using ImageQuantTL software (GE Healthcare).
The probes were (i) a 1,899 bp NotI/PacI fragment of pG2LK, comprising intron-luc+; and (ii) a 2,495 bp BglII fragment of pG2LK comprising the complete vector backbone up to 5 nt outside the predicted left border nick site and 25 nt outside the right border nick site.
Statistical Analyses
Statistical analyses were performed using GraphPad Prism software version 5.04.
Abbreviations
- AMT:
-
Agrobacterium-mediated transformation
- LB:
-
Left border
- RB:
-
Right border
- LUC:
-
Luciferase (encoded by the luc gene)
- nt:
-
Nucleotide
References
An G, Ebert PR, Mitra A, Ha SB (1988) Binary vectors. In: Gelvin SB, Schilperoort RA, Verma DPS (eds) Plant molecular biology manual. Kluwer Academic Publishers Dordrecht, pp A3/1–19
Arencibia AD, Carmona ER, Tellez P, Chan MT, Yu SM, Trujillo LE, Oramas P (1998) An efficient protocol for sugarcane (Saccharum spp. L.) transformation mediated by Agrobacterium tumefaciens. Transgenic Res 7:213–222
Arvinth S, Arun S, Selvakesavan RK, Srikanth J, Mukunthan N, Kumar PA, Premachandran MN, Subramonian N (2010) Genetic transformation and pyramiding of aprotinin-expressing sugarcane with cry1Ab for shoot borer (Chilo infuscatellus) resistance. Plant Cell Rep 29:383–395
Bartlett JG, Alves SC, Smedley M, Snape JW, Harwood WA (2008) High-throughput Agrobacterium-mediated barley transformation. Plant Methods 4:22
Basnayake SWV, Moyle R, Birch RG (2011) Embryogenic callus proliferation and regeneration conditions for genetic transformation of diverse sugarcane cultivars. Plant Cell Rep 30:439–448
Birch RG (1997) Plant transformation: problems and strategies for practical application. Annu Rev Plant Physiol Plant Mol Biol 48:297–326
Bourdon V, Harvey A, Lonsdale DM (2001) Introns and their positions affect the translational activity of mRNA in plant cells. EMBO Rep 2:394–398
Bower R, Elliott AR, Potier BAM, Birch RG (1996) High-efficiency, microprojectile-mediated cotransformation of sugarcane, using visible or selectable markers. Mol Breed 2:239–249
Brumbley SM, Snyman SJ, Gnanasambandam A, Joyce P, Hermann SR, da Silva JAG, McQualter RB, Wang ML, Egan BT, Patterson AH, Albert HH, Moore PH (2008) Sugarcane. In: Kole C, Hall TC (eds) Compendium of transgenic crop plants: sugar, tuber and fiber crops. Blackwell London, pp 1–58
Cheng M, Lowe BA, Spencer TM, Ye XD, Armstrong CL (2004) Factors influencing Agrobacterium-mediated transformation of monocotyledonous species. In Vitro Cell Dev Biol - Plant 40:31–45
Christensen AH, Sharrock RA, Quail PH (1992) Maize polyubiquitin genes—structure, thermal perturbation of expression and transcript splicing, and promoter activity following transfer to protoplasts by electroporation. Plant Mol Biol 18:675–689
Costa MGC, Nogueira FTS, Figueira ML, Otoni WC, Brommonschenkel SH, Cecon PR (2000) Influence of the antibiotic timentin on plant regeneration of tomato (Lycopersicon esculentum Mill.) cultivars. Plant Cell Rep 19:327–332
De Buck S, De Wilde C, Van Montagu M, Depicker A (2000) T-DNA vector backbone sequences are frequently integrated into the genome of transgenic plants obtained by Agrobacterium-mediated transformation. Mol Breed 6:459–468
De Buck S, Podevin N, Nolf J, Jacobs A, Depicker A (2009) The T-DNA integration pattern in Arabidopsis transformants is highly determined by the transformed target cell. Plant J 60:134–145
Depicker A, Stachel S, Dhaese P, Seurinck J, Deboeck F, De Greve H, Lemmers M, Van Montagu M, Schell J (1982) Nopaline Synthase: transcript mapping and DNA sequence. J Mol Appl Genet 1:561–573
D’Halluin K, Vanderstraeten C, Stals E, Cornelissen M, Ruiter R (2008) Homologous recombination: a basis for targeted genome optimization in crop species such as maize. Plant Biotechnol J 6:93–102
Elliott AR, Campbell JA, Brettell RIS, Grof CPL (1998) Agrobacterium-mediated transformation of sugarcane using GFP as a screenable marker. Aust J Plant Physiol 25:739–743
Enriquez-Obregon GA, Vazquez-Padron RI, Prieto-Samsonov DL, De la Riva GA, Selman-Housein G (1998) Herbicide-resistant sugarcane (Saccharum officinarum L.) plants by Agrobacterium-mediated transformation. Planta 206:20–27
Hellens RP, Edwards EA, Leyland NR, Bean S, Mullineaux PM (2000) pGreen: a versatile and flexible binary Ti vector for Agrobacterium-mediated plant transformation. Plant Mol Biol 42:819–832
Hiei Y, Komari T (2008) Agrobacterium-mediated transformation of rice using immature embryos or calli induced from mature seed. Nat Protoc 3:824–834
Hood EE, Gelvin SB, Melchers LS, Hoekema A (1993) New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res 2:208–218
Ishida Y, Hiei Y, Komari T (2007) Agrobacterium-mediated transformation of maize. Nat Protoc 2:1614–1621
Joyce P, Kuwahata M, Turner N, Lakshmanan P (2010) Selection system and co-cultivation medium are important determinants of Agrobacterium-mediated transformation of sugarcane. Plant Cell Rep 29:173–183
Kim JY, Gallo M, Altpeter F (2012) Analysis of transgene integration and expression following biolistic transfer of different quantities of minimal expression cassette into sugarcane (Saccharum spp. hybrids). Plant Cell Tissue Organ Cult 108:297–302
Lakshmanan P, Geijskes RJ, Aitken KS, Grof CLP, Bonnett GD, Smith GR (2005) Sugarcane biotechnology: the challenges and opportunities. In Vitro Cell Dev Biol-Plant 41:345–363
Last DI, Brettell RIS, Chamberlain DA, Chaudhury AM, Larkin PJ, Marsh EL, Peacock WJ, Dennis ES (1991) PEmu - an improved promoter for gene expression in cereal cells. Theor Appl Genet 81:581–588
Lazo GR, Stein PA, Ludwig RA (1991) A DNA transformation-competent Arabidopsis genomic library in Agrobacterium. Bio/Technology 9:963–967
Li RY, Qu RD (2011) High throughput Agrobacterium-mediated switchgrass transformation. Biomass Bioenergy 35:1046–1054
Ling HQ, Kriseleit D, Ganal MW (1998) Effect of ticarcillin potassium clavulanate on callus growth and shoot regeneration in Agrobacterium-mediated transformation of tomato (Lycopersicon esculentum Mill.). Plant Cell Rep 17:843–847
Manickavasagam M, Ganapathi A, Anbazhagan VR, Sudhakar B, Selvaraj N, Vasudevan A, Kasthurirengan S (2004) Agrobacterium-mediated genetic transformation and development of herbicide-resistant sugarcane (Saccharum species hybrids) using axillary buds. Plant Cell Rep 23:134–143
Mudge SR, Lewis-Henderson WR, Birch RG (1996) Comparison of Vibrio and firefly luciferases as reporter gene systems for use in bacteria and plants. Aust J Plant Physiol 23:75–83
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15:473–497
Nagel R, Elliott A, Masel A, Birch RG, Manners JM (1990) Electroporation of binary Ti plasmid vector into Agrobacterium tumefaciens and Agrobacterium rhizogenes. FEMS Micro Lett 67:325–328
Nauerby B, Billing K, Wyndaele R (1997) Influence of the antibiotic timentin on plant regeneration compared to carbenicillin and cefotaxime in concentrations suitable for elimination of Agrobacterium tumefaciens. Plant Sci 123:169–177
Nishimura A, Aichi I, Matsuoka M (2006) A protocol for Agrobacterium-mediated transformation in rice. Nat Protoc 1:2796–2802
Oltmanns H, Frame B, Lee LY, Johnson S, Li B, Wang K, Gelvin SB (2010) Generation of backbone-free, low transgene copy plants by launching T- DNA from the agrobacterium chromosome. Plant Physiol 152:1158–1166
Ooms G, Hooykaas PJJ, Vanveen RJM, Vanbeelen P, Regensburgtuink TJG, Schilperoort RA (1982) Octopine Ti-plasmid deletion mutants of Agrobacterium tumefaciens with emphasis on the right side of the T-region. Plasmid 7:15–29
Robert ML, Flores MR, Loyolavargas VM (1989) Growth-promoting activity of certain penecillins on cultivated cells of Bouvardia ternifolia. Phytochemistry 28:2659–2662
Rogers S, Bendich A (1988) Extraction of DNA from plant tissues. In: Gelvin SB, Schilperoort RA, Verma DPS (eds) Plant molecular biology manual. Kluwer Academic Publishers Dordrecht, pp A6/1–10
Rothrock RE, Polin-McGuigan LD, Newhouse AE, Powell WA, Maynard CA (2007) Plate flooding as an alternative Agrobacterium-mediated transformation method for American chestnut somatic embryos. Plant Cell Tissue Organ Cult 88:93–99
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Santosa DA, Hendroko R, Farouk A, Greiner R (2004) A rapid and highly efficient method for transformation of sugarcane callus. Mol Biotechnol 28:113–119
Shrawat AK, Lorz H (2006) Agrobacterium-mediated transformation of cereals: a promising approach crossing barriers. Plant Biotechnol J 4:575–603
Vain P, Afolabi AS, Worland B, Snape JW (2003) Transgene behaviour in populations of rice plants transformed using a new dual binary vector system: pGreen/pSoup. Theor Appl Genet 107:210–217
Vain P, Harvey A, Worland B, Ross S, Snape JW, Lonsdale D (2004) The effect of additional virulence genes on transformation efficiency, transgene integration and expression in rice plants using the pGreen/pSoup dual binary vector system. Transgenic Res 13:593–603
Vain P, Worland B, Thole V, McKenzie N, Opanowicz M, Fish LJ, Bevan MW, Snape JW (2008) Agrobacterium-mediated transformation of the temperate grass Brachypodium distachyon (genotype Bd21) for T-DNA insertional mutagenesis. Plant Biotechnol J 6:236–245
Wu HX, Doherty A, Jones HD (2008) Efficient and rapid Agrobacterium-mediated genetic transformation of durum wheat (Triticum turgidum L-var. durum) using additional virulence genes. Transgenic Res 17:425–436
Ye XD, Williams EJ, Shen JJ, Esser JA, Nichols AM, Petersen MW, Gilbertson LA (2008) Plant development inhibitory genes in binary vector backbone improve quality event efficiency in soybean transformation. Transgenic Res 17:827–838
Zhangsun D, Luo S, Chen R, Tang K (2007) Improved Agrobacterium-mediated genetic transformation of GNA transgenic sugarcane. Biologia 62:386–393
Acknowledgments
This work was undertaken through a collaboration between CSR Sugar Limited (Sucrogen) and The University of Queensland with support from the Australian Research Council. The authors acknowledge the excellent technical assistance of Andrea Gray, Peter Bakker, Kerrin Henderson, and Alam Chen.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by: Paul Moore
Rights and permissions
About this article
Cite this article
Anderson, D.J., Birch, R.G. Minimal Handling and Super-Binary Vectors Facilitate Efficient, Agrobacterium-Mediated, Transformation of Sugarcane (Saccharum spp. hybrid). Tropical Plant Biol. 5, 183–192 (2012). https://doi.org/10.1007/s12042-012-9101-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12042-012-9101-1