Abstract
Treatment of perchlorate or nitrate salt of cadmium(II) with carboxamide derivatives (L) generated four novel mononuclear metal complexes, represented as [Cd(L )4](ClO4)2 (1a and 1b) and [Cd(L)2(ONO2)2] (2a and 2b) in appreciable yields (L = L 1 = N-(furan-2-ylmethyl)-2-pyridine carboxamide and L = L 2 = N-(thiophen-2-ylmethyl)-2-pyridine carboxamide). The complexes have been characterized by FT-IR, UV-Visible, elemental analysis and single crystal X-ray crystallographic analysis which revealed eight coordinated cadmium ions, but in different coordination environments, depending on the counter anion used. In addition, electronic absorption, fluorescence spectroscopy and viscosity measurements revealed a significant interaction of the four complexes with CT-DNA via intercalative/groove binding mode. The intrinsic binding constant K b obtained varies from 0.4 × 10 4 to 1.11 × 10 5 M−1. The results suggest that neutral complexes 2a and 2b bind to DNA in an intercalative mode. On the other hand, cationic complexes 1a and 1b bind with DNA via weak electrostatic/covalent interaction.

Synthesis and characterization of four mononuclear eight coordinated cadmium(II) complexes with newly explored carboxamide derivatives and study of interaction with calf-thymus DNA are reported. The results suggest that neutral complexes 2a and 2b bind to DNA in an intercalative mode. On the other hand, cationic complexes 1a and 1b interact with DNA via weak intercalative or groove binding mode.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The coordination chemistry of carboxamide derivatives, especially towards transition metal ions, has attracted a considerable interest and this attention arises from the fact that pyridine carboxamide complexes have been used in various fields such as in asymmetric catalysis,[1,2]dendrimers,[3] molecular receptor synthesis[4] and also in the synthesis of compounds with possible antitumor properties.[5‐8]The carboxamide [-C(O)NH-] group of the primary structure of proteins represents an important ligand building unit in coordination chemistry, since its chelating nature imparts a unique balance of stability versus reactivity and has allowed for remarkable developments in a variety of catalytic transformations. Cadmium is a highly toxic metal and a potent carcinogen. In fact, even at very low concentrations cadmium can cause extensive damage to kidney, liver, lungs, prostates and hematopoietic systems of humans, and it has been recognized by the International Agency for Research on Cancer as a carcinogenic agent.[9] Recent studies suggest that cadmium-induced carcinogenicity involves direct or indirect interaction of this metal with DNA due to the various binding sites presented in the double helix,[10,11]although the exact molecular mechanism of cadmium-induced carcinogenicity remains mostly ambiguous. Antibacterial activities and binding ability of cadmium complexes with DNA have received a considerable interest.[12‐14]For instance, some Cd(II) complexes exhibit significant antitumor activity on murine melanoma B16 cells and human cervical cancer HeLa cells,[15] showing an activity similar to cis-platin with IC50 less than 10.0 μM for some cell lines such as MDA-361, MDA-453, HeLa, etc.[14] As a result, investigations of interaction of cadmium complexes with DNA have been imperative in order to elucidate the mode and affinity of binding at cellular level.[16]
Herein, we report an account of the synthesis, structural characterization and DNA binding study of cadmium(II) complexes of two carboxamide derivatives (L) formulated as [Cd(L )4](ClO4 )2 (1), [Cd(L)2(ONO2 )2] (2), where L = L 1 = N-(furan-2-ylmethyl)-2-pyridinecarboxamide or L = L 2 = N-(thiophen-2-ylmethyl)-2-pyridinecarboxamide. The binding constants Kb of these complexes with DNA have been obtained from UV-Vis spectra and the quenching constants Ksv were determined from fluorescence displacement experiments using ethidium bromide (EB) as spectral probe. The binding affinity of these complexes with calf-thymus-DNA has also been studied by using viscosity measurements.
2 Experimental
2.1 Materials and Physical measurements
The chemicals and reagents used for the synthesis were obtained from commercial sources. Solvents were distilled from an appropriate drying agent. Calf thymus-DNA (Bangalore Genie, India) and ethidium bromide (Sigma) were used as received. All other chemicals and solvents were of analytical grade. The elemental (C, H, N) analyses were performed on a Perkin Elmer model 2400 elemental analyzer. IR spectra and 1H NMR spectra of complexes were recorded on a Perkin Elmer FTIR model RX1 spectrometer (KBr disc, 4000–400 cm−1) and on a Bruker Avance DPX 500 MHz spectrometer using DMSO-d6 as solvent, respectively. The solution electrical conductivity was measured using Systronics 304 digital conductivity meter with a solution concentration of ca.10 −3 mol dm −3. The electronic absorption spectra were recorded on a JASCO UV-Vis/NIR spectrophotometer model V-570. The fluorescence spectra of the complexes bound to DNA were recorded with an excitation wavelength of 522 nm by using the Hitachi-4500 fluorimeter.
2.2 Synthesis of ligands L 1 and L 2
The organic ligands N-(furan-2-ylmethyl)-2-pyridinecarboxamide (L 1) and N-(thiophen-2-ylmethyl)-2-pyridine-carboxamide (L 2) have been prepared as previously reported and duly characterised by single crystal X-ray crystallography.[17]
2.3 Synthesis of cadmium(II) complexes
The complexes were synthesized following a procedure as described below. The organic compound (4.0 mmol, 816 mg of L 1 or 872 mg of L 2) was dissolved in dry ethanol by stirring the mixture for 15 min. To this solution, an ethanolic solution of cadmium(II) perchlorate hexahydrate (Cd(ClO4)2.6H2O, 419 mg, 1.0 mmol) (for 1) or cadmium(II) nitrate tetrahydrate (Cd(NO3)2.4H2O, 309 mg, 1.0 mmol) (for 2) was added dropwise and the mixture was stirred 30 min and then refluxed for 5 h (Scheme 1). The resulting mixture was filtered off and the clear filtrate was collected. The filtrate was kept aside at room temperature and the volume of the solution was reduced by slow evaporation. The product was collected by washing with cold ethanol and water and dried in vacuo. Pure crystalline products, which were used for elemental analysis and for characterization using physico-chemical and spectroscopic tools, were obtained from ethanol. Single crystals of 1a and 2b suitable for X-ray diffraction study were obtained by slow evaporation of the ethanolic solution.
2.3.1 2.3.1 Complex 1a: [Cd(L 1) 4 ](ClO 4 ) 2 :
Yield: 65–70% (0.772 g); C44H40CdCl2N8O16: Anal. Found: C, 47.11; H, 3.55; N, 9.92; Cd, 9.97% Calc.: C, 47.18; H, 3.60; N, 10.00; Cd, 10.04% IR (cm−1): ν C=N, 1475, \(\nu _{\mathrm {ClO4}}^{\mathrm {-}}\) 1087, 628, ν C=O 1643, ν N−H 3337; Conductivity (Λo, ohm−1 cm2 mol−1) in MeOH: 214; 1H NMR (δ, ppm in Dmso-d6): 9.01 (t, 1H); 8.53 (d, 1H); 7.92 (m, 2H); 7.50 (m, 1H); 7.43 (s, 1H); 6.27 (t, 1H), 6.16 (−NH), 4.38 (s, 2H of CH2).
2.3.2 2.3.2 Complex 1b: [Cd(L 2) 4 ](ClO 4 ) 2 :
Yield: 65–70% (0.807 g); C44H40CdCl2N8O12S4: Anal. Found: C, 40.91; H, 3.25; N, 8.58; Cd, 11.53% Calc.: C, 41.02; H, 3.13; N, 8.70; Cd, 11.64%. IR (cm−1): ν C=N, 1469, \(\nu _{\mathrm {ClO4}}^{\mathrm {-}}\) 1087, 628, ν C=O 1634, ν N−H 3314. Conductivity (Λo, ohm−1 cm2 mol−1) in MeOH: 204. 1H NMR (δ, ppm in dmso-d6): 9.24 (t, 1H); 8.47 (d, 1H); 7.95 (m, 2H); 7.60 (m, 1H); 7.31 (s, 1H); 7.01 (t, 1H), 6.20 (−NH), 4.40 (s, 2H of CH2).
2.3.3 2.3.3 Complex 2a: [Cd(L 1) 2 (ONO 2 ) 2 ]:
Yield: 70–74% (0.467 g); C22H20CdN6O10: Anal. Found: C, 41.16; H, 3.09; N, 13.07; Cd, 17.46% Calc.: C, 41.23; H, 3.15; N, 13.11; Cd, 17.54%. IR (cm−1): \(\nu _{\mathrm {NO3}}^{\mathrm {-}}\) 1381, ν C=O 1643, ν N−H 3336. Conductivity (Λo, ohm−1 cm2 mol−1) in MeOH: 12.2. 1H NMR (δ, ppm in dmso-d6): 9.34 (t, 1H); 8.63 (d, 1H); 7.92 (m, 2H); 7.57 (m, 1H); 7.40 (s, 1H); 6.31 (t, 1H), 6.58 (−NH), 4.71 (s, 2H of CH2).
2.3.4 2.3.4 Complex 2b: [Cd(L 2) 2 (ONO 2 ) 2 ]:
Yield: 70–74% (0.492 g); C22H20CdN6O8S2: Anal. Found: C, 39.13; H, 2.91; N, 12.37; Cd, 16.64% Calc.: C, 39.26; H, 3.00; N, 12.49; Cd, 16.70%; IR (cm−1): ν C=N, 1473, \(\nu _{\mathrm {NO3}}^{\mathrm {-}}\) 1383, ν C=O 1633, ν N−H 3312. Conductivity (Λo, ohm−1 cm2 mol−1) in MeOH: 13.3. 1H NMR (δ, ppm in dmso-d6): 9.40 (t, 1H); 8.60 (d, 1H); 8.01 (m, 2H); 7.60 (m, 1H); 7.35 (s, 1H); 7.01 (t, 1H), 6.90 (−NH), 4.64 (s, 2H of CH2).
2.4 X-ray crystal structure analysis
X-ray single crystal data of 1a and 2b were collected at room temperature (296(2) K) using Mo-K α radiation (λ = 0.71073 Å) on a SMART APEX II diffractometer equipped with CCD area detector. Data collection, data reduction, structure solution/refinement were carried out using the SMART APEX II package.[18] The structures were solved by direct methods of SHELXS-97 and refined by full-matrix least squares refinement methods based on F 2 with SHELXL-97 program.[19] All non-hydrogen atoms were refined anisotropically. In compound 1, one furanyl ring (O6 /C30–33) was fitted by using an input fragment due to a slight disorder of the group. All calculations were performed using the WinGX package.[20] Crystal data and details of the refinement are reported in Table 1, and a selection of bond lengths and angles are shown in Table 2.
2.5 DNA binding experiments: Study with UV spectroscopy
Tris-HCl buffer solution (pH = 7.2) used in all the experiments involving CT-DNA was prepared using deionized and sonicated HPLC grade water (Merck). The CT-DNA used in the experiments was checked to be sufficiently free from protein by using the ratio of UV absorbance of DNA solution in tris-HCl at 260 nm and 280 nm (A260/A 280) that was around 1.9.[21] The concentration of DNA was estimated by using the extinction coefficient (6600 M−1 cm−1) at 261 nm[22] and stock solution of DNA was always stored at 4 °C. The stock solution of each complex was prepared by dissolving it in 2 mL of DMSO and diluted with Tris-HCl buffer to get the required concentration for all the experiments. Absorption spectral titration experiment was performed by maintaining constant complex concentration and varying the CT-DNA concentration. To eliminate the absorbance by DNA itself, equal amount of CT-DNA was added to the reference solution.
2.5.1 2.5.1 EB competitive studies with fluorescence spectroscopy
In the ethidium bromide (EB) fluorescence displacement experiment, 5.0 μL of EB tris-HCl solution (1.0 mM) was added to 1.0 mL of DNA solution (at saturated binding levels)[16] and stored in the dark for 2.0 h. Then, the solution of the complex was titrated into the DNA/EB mixture and diluted with tris-HCl buffer to 5.0 mL to get the solution with the appropriate complex/CT-DNA mole ratio. Before measurements, the mixture was shaken up and incubated at room temperature for 30 min. The fluorescence spectra of EB bound to DNA were obtained in the fluorimeter at λ ex = 522 nm.
2.5.2 2.5.2 Viscosity measurements:
To adjudge the binding mode (groove/intercalative) of complex with DNA, viscosity measurements were performed by employing the Ostwald’s viscometer. Titrations were carried out in the viscometer by adding the complex (0.5–3.5 μM) to the CT-DNA solution (5.0 μM). The viscosity of the solutions were calculated from the observed flow time of CT-DNA-containing solution corrected from the flow time of buffer alone (t0), η = t −t 0. The obtained data were used to plot the (η/η 0)1/3 versus the ratio of the concentration of complex to CT-DNA, where η and η 0 are the viscosity of the CT-DNA solution in the presence of complex and CT-DNA solution only, respectively.
3 Results and Discussion
3.1 Synthesis and characterization
The perchlorate and nitrate salt of cadmium(II) were reacted with two carboxamide derivatives L 1 and L 2 to form four new mononuclear cadmium complexes in ethanol medium at ambient temperature. The complexes, which revealed to be stable in air and soluble in methanol, DMF, DMSO, etc., were characterized by physico-chemical and spectroscopic tools.
3.2 Structure of complexes
3.2.1 3.2.1 Crystal structure of 1a:
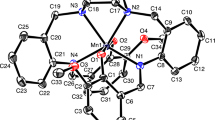
An Ortep view of complex 1a, crystallizing in monoclinic system, space group P21/c, is depicted in Figure 1, and selected bond lengths and angles are listed in Table 2.
The metal ion is coordinated by four chelating L 1 ligands through the pyridine nitrogen and carbonyl oxygen giving a distorted square antiprismatic geometry with a N4O4 chromophore. The asymmetric ligands L 1 chelate the metal with Cd-N and Cd-O bond distances in the range 2.397(5)–2.444(5) and 2.417(4)–2.459(4) Å, respectively. The chelating angle in all cases is very similar, between 66.77(16)–67.53(15 )°. The arrangement of ligands around the metal is such that the trans chelating pyridine-carboxamide fragments are oriented head-head with N(1)-Cd-N(5) and N(3)-Cd-N(7) bond angles of 152.63(17) and 151.08(18 )°, respectively. The present coordination metal-ligand bond distances are as expected for eight-coordinated Cd(II) complexes with four N, O-chelating ligands.[23]
The perchlorate ions are weakly bound to the complexes through H-bonds involving NH groups and perchlorate anions (Table 3 and Figure S1, in Supplementary Information), and these interactions give rise to a 1D chain developed along axis c. In addition, a short ring interaction between thiophene S1 and pyridine N1 of a symmetry related complex (centroid-to-centroid distance = 3.707(4) Å; dihedral angle between planes = 2.3(3 )°) is detected in the crystal packing.
3.2.2 3.2.2 Crystal structure of 2b:
The X-ray diffraction study revealed that the complex crystallizes in monoclinic system, space group P21. The metal is coordinated by two N,O chelating L 2 and by two chelating nitrate anions leading to a neutral molecular species (see Figure 2). The Cd atom exhibits a distorted dodecahedron environment with Cd-O(L 2) and the Cd-N bond distances are slightly shorter (mean 2.357 Å) than those measured in 1a, likely for the lower bulkiness of nitrates, which present an unsymmetrical coordination. In fact, in each nitrate one Cd-O bond distance is shorter by about 0.11-0.13 Å compared to the other (2.424(5) vs 2.530(6) and 2.426(4) vs 2.572(5) Å). The chelating pyridine-carboxamide mean planes form a dihedral angle of 75.78(1 )°, while the two nitrate mean planes define an angle of 64.60(1 )°.
Cadmium(II) complexes with the same coordination environment have been previously reported,[24,25]having coordination bond distance values in agreement with those reported herein. In the crystal lattice, the monomeric units form a 2D polymeric layered supramolecular stucture parallel to the crystallographic bc plane through intermolecular hydrogen bonds between the carboxamide N-H fragments and nitrate anions (Figure 3). The hydrogen bonding parameters are reported in Table 3.
3.3 Spectral characterization
The IR spectrum of the ligand L 1 shows absorption at 3344 and 1660 cm−1 assignable to the ν N−H and ν C=O stretching, respectively. In the spectra of both complexes 1a and 2a, ν C=O band moved towards lower frequency at 1643 cm−1, suggesting the coordination of the metal center via the carbonyl oxygen. Both complexes 1a and 2a show characteristic IR bands at 3347 and 3336 cm−1, which correspond to the characteristic peaks of ν N−H, in addition to a characteristic band at 1475 cm−1 assignable to the ν C=N stretching frequency. The symmetric and asymmetric stretching vibrations of the perchlorate groups of complex 1a appear at 1087 and 628 cm−1, respectively. On the other hand, complex 2a displays a characteristic band at 1381 cm−1 assigned to \(\nu _{\mathrm {NO3}}^{\mathrm {-}}\).
In the IR spectrum of the ligand L 2, the absorption at 3317 and at 1654 cm−1 are assigned to the ν N−H and to the ν C=O, respectively. In the spectra of both complexes 1b and 2b, the ν C=O band is shifted to a lower frequency (1634 cm−1), suggesting the coordination of the carbonyl oxygen to the metal centre. Complex 1b showed broad intense bands at 1087, 628 cm−1, assigned to the symmetric and asymmetric stretching of \(\nu _{\mathrm {ClO4}}^{\mathrm {-}}\), respectively, and bands at 1669 and 3314 cm−1, assignable to the ν C=N and ν N−H stretching frequency, respectively. On the other hand, complex 2b showed a moderate band at 1383 cm−1 assigned to \(\nu _{\mathrm {NO3}}^{\mathrm {-}}\). The bands at 3312, 1472 cm−1 indicate the ν N−H and ν C=N stretching frequency, respectively.
3.4 DNA binding studies
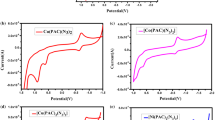
The application of electronic absorption spectra in DNA binding studies is one of the most useful techniques.[26] Generally, binding of the metal complex to the DNA helix is justified by an increase of the n →π* band of cadmium(II) complex due to the involvement of intercalative interactions between the effective chromophore of the complex and the base pairs of DNA.[27‐29]Here, the UV-Visible spectra of the cadmium(II) complexes in DMSO exhibited intense absorption bands at ca. 267 nm, 270 nm, 272 nm and 288 nm for the complexes 1a, 1b, 2a and 2b, respectively, attributable to n– π* transition (Table S1, in Supplementary Information). The absorption spectra of cadmium complexes in the absence and presence of CT-DNA (at a constant concentration of complex) are given in Figures 4, 5, S2 and S6 (Supplementary Information). The extent of the hyperchromism in the absorption band is generally consistent with the strength of intercalative interactions[26,30‐32]and these spectral characteristics suggest that the complexes interact with DNA, most likely through a stacking mode between an heterocyclic aromatic chromophore and the base pairs of DNA.[33,34]
Electronic spectra of titration of complex 1a with CT-DNA (peak at 279 nm) in tris-HCl buffer; [complex] = 2.47 ×10 −5 M; [DNA]: (a) 0.0, (b) 1.01 ×10 −6, (c) 2.18 ×10 −6, (d) 3.364 ×10 6, (e) 4.669 ×10 −6 (f) 6.156 ×10 −6 (g) 7.32 ×10 −6 M. The arrow denotes the gradual increase of DNA concentration.
In order to establish the binding strength of the metal complexes with CT-DNA, the association constant Kb was determined from the spectral titration data using the Wolfe-Shimer equation (1):[35]
where [DNA] is the concentration of DNA, ε f and ε b correspond to the extinction coefficient, respectively, for the free complex and for the cadmium(II) complex in the fully bound form, and ε a is ‘apparent’ extinction coefficient for each addition of DNA to the cadmium(II) complex. Plots of [DNA] /(ε a– ε f) vs [DNA] (Figures 6, 7, S3 and S7 in the Supplementary Information) gave the association constant Kb as the ratio of slope to the intercept. The values of Kb for the complexes were estimated to be: 4.04 × 10 4 M−1 (R = 0.999, n = 5 points) for 1a, 8.18 × 10 4 M−1 (R = 0.988, n = 5 points) for 2a, 4.9 × 10 4 M−1 (R = 0.984, n = 4 points) for 1b and 1.11 × 10 5 M−1 (R = 0.959, n = 5 points) for 2b, indicating the following order 1a <1b <2a <2b. These values are comparable to those previously reported for cadmium(II) complexes.[36‐39]
The binding of the complexes to DNA decreases the emission intensity and the extent of the reduction of the emission intensity gives a measure of the DNA binding propensity of the complexes and stacking interaction (intercalation) between the adjacent DNA base pairs.[40] Although EB does not show any emission in the buffer medium due to fluorescence quenching by solvent molecules, it shows an emission band in the presence of CT-DNA due to intercalative binding to the helix. The decrease in fluorescence intensity of the DNA-bound EB (with excitation wavelength of 522 nm) decreases with the increasing addition of the complexes (see Figures 8, 9, S4 and S8 in the Supplementary Information) because the binding of the complex to DNA promotes the release of the EB molecules from the double helix.[41] This quenching due to the addition of the cadmium complex is in agreement with the Stern-Volmer equation (2):[42]
Emission spectra of the CT-DNA-EB system in tris-HCl buffer upon titration with complex 1a. λ ex = 522 nm; [EB] = 1.03 × 10 −6 M, [DNA] = 1.23 ×10 −6 M; [Complex]: (a) 0.0, (b) 3.02 ×10 −6, (c) 6.05 ×10 −6,(d) 9.07 ×10 −6, (e) 1.21 ×10 −5, (f) 1.51 ×10 −5 , (g) 2.72 ×10 −5 M. The arrow denotes the gradual increase of complex concentration.
Emission spectra of the CT-DNA-EB system in tris-HCl buffer during the titration of complex 2a (λ ex = 522 nm). [EB] = 1.03 ×10 −6 M, [DNA] = 1.23 ×10 −6 M; [Complex]: (a) 0.0, (b) 1.07 ×10 −6, (c) 2.16 ×10 −6, (d) 3.04 ×10 −6, (e) 4.22 ×10 −6 and (f) 5.34 ×10 −6, M. The arrow shows increase in the complex concentration.
where I0 and I represent the fluorescence intensities in the absence and presence of quencher (complex), respectively, Ksv is Stern-Volmer quenching constant, Q is the concentration of the quencher. In the quenching plot (see Figures 10, 11, S5 and S9 in the Supplementary Information) of I0/I vs [complex], Ksv value is given by the slope of the regression line. The Ksv values are 2.99 × 10 4, 2.46 × 10 5, 4.71 × 10 4 and 2.63 × 10 5 M−1 for complexes 1a, 2a, 1b, and 2b, respectively, suggesting a strong affinity of each cadmium(II) complex to CT-DNA. This feature may be attributed to a better ability of the complex to interact with DNA through intercalation binding, thus releasing EB from the EB-DNA complex. It is worth noting that the results obtained from the fluorescence spectra are in agreement with those derived from the absorption spectroscopy.
Furthermore, interactions between the complexes and DNA were investigated by viscosity measurement that is regarded as the most effective means to study intercalative binding mode of DNA in solution.[43,44]The viscosity of DNA is sensitive to changes in DNA length upon interaction with a molecule and the solution viscosity (η/η 0) and DNA length (L /L 0) are related by the equation (L /L 0)= (η/η 0)1/3.[46,47]A classical intercalative mode causes a significant increase of viscosity of the DNA solution due to a larger separation of intercalated base pairs leading to an increase of overall DNA length. In contrast, a molecule that binds exclusively in the DNA groove by partial and/or non-classical intercalation (i.e., electrostatic interaction or external groove-binding), a bend or kink in the DNA helix may be promoted leading to a slight shortening of its effective length. In this case, the change in viscosity of the DNA solution is less pronounced or there is no change at all.[45‐47] As indicated in Figure 12, the viscosity of complexes 2 increases with the increasing ratio of complex to CT-DNA. These results support the hypothesis that complexes 2 bind to CT-DNA by intercalation,[48,49]resembling the binding mode of EB to CT-DNA. In comparison to ethidium bromide, the relative viscosity is slightly lower for complexes 1. The decrease in viscosity of DNA on addition of complexes 1a and 1b is indicative of electrostatic/covalent interaction.[50]It may be due to the presence of four non-planar ligand moieties in complexes (1), which probably play a key role to interact with DNA in electrostatic/covalent binding mode.
4 Conclusions
Carboxamide ligands (L 1 and L 2) obtained in situ from pyridine-2-carboxylic acid with furfuryl amine and 2-thiophenemethyl amine were reacted with perchlorate and nitrate salt of cadmium(II), separately, to afford four mononuclear cadmium complexes of formula [Cd(L )4](ClO4)2 (L = 1a and 1b) and [Cd(L )2(ONO2)2] (L = 2a and 2b). The X-ray structural analysis indicated that the composition is anion dependent: in fact, perchlorate remained uncoordinated while the more coordinating nitrate anion is bound to cadmium(II) ion leading to cationic and neutral complexes, respectively. In this work, the detailed study is reported on the binding of complexes with CT-DNA, investigated by electronic absorption titrations, fluorescence spectroscopy, as well as by viscosity measurements, which indicated that complexes of different composition interfere differently with DNA. Actually weak electrostatic/covalent binding is detected with cationic complexes 1a and 1b, while neutral complexes 2a and 2b bind to DNA via intercalation mode. However, the binding constants of these four eight coordinated cadmium(II) complexes are comparable to those of the previously reported six/five coordinated cadmium(II) complexes,[36‐39]but in some cases, the observed values are somewhat higher than those reported[51‐53](Table S2 in Supplementary Information). The observed values listed in Table S2 clearly indicate that the five/six coordinated cadmium(II) complexes more strongly bound than the seven coordinated (Kb = 3.00 × 10 3 M−1)[52] and eight-coordinated (Kb = 2.1 × 10 3 M−1)[53] cadmium(II) complexes; i.e., with increase of coordinated group ligated with Cd(II) centre, decrease of binding constants (Kb) was observed, but it is not valid in our case though the complexes are eight coordinated cadmium(II) complexes. Here, it is noteworthy to mention that both type of the complexes (1 and 2) are eight coordinated cadmium(II) complexes and they strongly interact with CT-DNA, comparable to the interaction of five and six coordinated cadmium(II) complexes.
References
Lin J, Zhang J Y, Xu Y, Ke X K and Guo Z J 2001 N, N′-(1,2-phenylene)bis(pyridine-2-carboxamide) and N, N′-(1,2-cyclohexanediyl)bis(pyridine-2-carboxamide) toluen hemisolvate Acta Crystallogr. Sect. C 57 192
Trost B M and Hachiya I 1998 Asymmetric Molybdenum-Catalyzed Alkylations J. Am. Chem. Soc. 120 1104
Epperson J D, Ming L J, Baker G R and Newkome G R 2001 Paramagnetic Cobalt(II) as an NMR Probe of Dendrimer Structure: Mobility and Cooperativity of Dendritic Arms J. Am. Chem. Soc. 123 8583
Collinson S R, Gelbrich T, Hursthouse M B and Tucker J H R 2001 Novel ferrocene receptors for barbiturates and ureas Chem. Commun. 555
Zhang J, Liu Q, Duan C, Shao Y, Ding J, Miao Z, You X and Guo Z 2002 Structural evidence for the facile chelate-ring opening reactions of novel platinum(II)–pyridine carboxamide complexes J. Chem. Soc., Dalton Trans. 22 591
Li J, Zheng M L, King I, Doyle T W and Chan S H 2001 Syntheses and Antitumor Activities of Potent Inhibitors of Ribonucleotide Reductase 3-Amino-4- Methylpyridine-2-Carboxaldehyde-Thiosemicarba-zone (3-Amp), 3-Amino-Pyridine-2-Carboxaldehyde-Thiosemicarbazone (3-Ap) and its Water-Soluble Prodrugs Curr. Med. Chem. 8 121
He X M and Carter D C 1992 Atomic structure and chemistry of human serum albumin Nature 358 209
Mukherjee T, Sen B, Zangrando E, Hundal G, Chattopadhyay B and Chattopadhyay P 2013 Palladium(II) and platinum(II) complexes of deprotonated N, N′-bis(2-pyridinecarboxamide)-1,2-benzene: Synthesis, structural characterization and binding interactions with DNA and BSA Inorg. Chim. Acta 406 176 and references therein
Singhal R N and Jain M 1997 Cadmium-Induced Changes in the Histology of Kidneys in Common Carp, Cyprinus carpio (Cyprinidae) Bull. Environ. Contam. Toxicol. 58 456
Hossain Z and Huq F 2002 Studies on the interaction between Cd 2+ ions and nucleobases and nucleotides J. Inorg. Biochem. 90 97
Hossain Z and Huq F 2002 Studies on the interaction between Cd 2+ ions and DNA J. Inorg. Biochem. 90 85
Genova P, Varadinova T, Matesanz A I, Marinova D and Souza P 2004 Toxic effects of bis(thiosemicarbazone) compounds and its palladium(II) complexes on herpes simplex virus growth Toxicol. Appl. Pharmacol. 197 107
Hu W X, Zhou W, Xia C and Wen X 2006 Synthesis and anticancer activity of thiosemicarbazones Bioorg. Med. Chem. Lett. 16 2213
Filipović N R, Bacchi A, Lazić M, Pelizzi G, Radulović S, Sladić D M, Todorović T R and Anđelković K K 2008 Synthesis, structure and cytotoxic activity evaluation of a dinuclear complex of Cd(II) with N ′, N ′2-bis[(1E)-1-(2-pyridyl)ethylidene]propanedihydrazide Inorg. Chem. Commun. 11 47
Bjelogrlic S, Todorovic T, Bacchi A, Zec M, Sladic D, Srdic-Rajic T, Radanovic D, Radulovic S, Pelizzi G and Andelkovic K 2010 Synthesis, structure and characterization of novel Cd(II) and Zn(II) complexes with the condensation product of 2-formylpyridine and selenosemicarbazide: Antiproliferative activity of the synthesized complexes and related selenosemicarbazone complexes J. Inorg. Biochem. 104 673
Bertin G and Averbeck D 2006 Cadmium: Cellular effects, modifications of biomolecules, modulation of DNA repair and genotoxic consequences (a review) Biochimie 88 1549
Mondal B, Sen B, Zangrando E and Chattopadhyay P 2015 Zinc(II) complexes of carboxamide derivatives: Crystal structures and interaction with calf thymus DNA J. Chem. Sci. 127 1747
Altomare A, Cascarano G, Giacovazzo C and Guagliardi A 1993 Completion and refinement of crystal structures with SIR92 J. Appl. Crystallogr. 26 343
Sheldrick G M 2008 A short history of SHELX Acta Crystallogr. Sect. A: Found. Crystallogr. 64 112
Farrugia L J 1999 WinGX suite for small-molecule single-crystal crystallography J. Appl. Crystallogr. 32 837
Marmur J 1961 A procedure for the isolation of deoxyribonucleic acid from micro-organisms J. Mol. Biol. 3 208
Reichmann M E, Rice S A, Thomas C A and Doty P 1954 A Further Examination of the Molecular Weight and Size of Desoxypentose Nucleic Acid J. Am. Chem. Soc. 76 3047
Barszcz B, Jablonska-Wawrzycka A, Stadnicka K and Hodorowicz S A 2005 The synthesis and structural characterization of novel zinc and cadmium complexes of chelating alcohol Inorg. Chem. Commun. 8 951
Hakimi M, Moeini K, Mardani Z and Khorrami F 2012 Crystal Structure and Characterization of a New Eight Coordinated Cadmium Complex J. Kor. Chem. Soc. 57 352
Barszcz B, Hodorowicz S, Jablonska-Wawrzycka A and Stadnicka K 2005 Synthesis, X-ray structure and spectroscopic investigation of an eight-coordinate cadmium(II) complex J. Coord. Chem. 58 203
Barton J K, Danishefsky A and Goldberg J 1984 Tris(phenanthroline)ruthenium(II): Stereoselectivity in binding to DNA J. Am. Chem. Soc. 106 2172
Sun J, Shuo W, An Y, Liu J, Gao F, Ji L N and Mao Z W 2008 Synthesis, crystal structure and DNA-binding properties of ruthenium(II) polypyridyl complexes with dicationic 2,2 ′-dipyridyl derivatives as ligands Polyhedron 27 2845
Lombardo V, Bonomi R, Sissi C and Mancin F 2010 Phosphate diesters and DNA hydrolysis by dinuclear Zn(II) complexes featuring a disulfide bridge and H-bond donors Tetrahedron 66 2189
Dhara K, Ratha J, Manassero M, Wang X Y, Gao S and Banerjee P 2007 Synthesis, crystal structure, magnetic property and oxidative DNA cleavage activity of an octanuclear copper(II) complex showing water–perchlorate helical network J. Inorg. Biochem. 101 95
Dhara K, Roy P, Ratha J, Manassero M and Banerjee P 2007 Synthesis, crystal structure, magnetic property and DNA cleavage activity of a new terephthalate-bridged tetranuclear copper(II) complex Polyhedron 26 4509
Bloomfield V A, Crothers D M and Tinoco I 1974 In Physical Chemistry of Nucleic Acids (New York: Harper and Row) p. 432
Tysoe S A, Morgan R J, Baker A D and Strekas T C 1993 Spectroscopic investigation of differential binding modes of .DELTA.- and.LAMBDA.-Ru(bpy )2(ppz )2+ with calf thymus DNA J. Phys. Chem. 97 1707
Selvi P T, Stoeckli-Evans H and Palaniandavar M 2005 Synthesis, structure and DNA interaction of cobalt(III) bis-complexes of 1,3-bis(2-pyridylimino) isoindoline and 1,4,7-triazacyclononane J. Inorg. Biochem. 99 2110
Kelly J M, Tossi A B, McConnell D J and OhUigin C 1985 A study of the interactions of some polypyridylrutheniumn(I) complexes with DNA using fluorescence spectroscopy, topoisomerisation and thermal denaturation Nucl. Acids. Res. 13 6017
Wolfe A, Shimer G and Meehan T 1987 Polycyclic aromatic hydrocarbons physically intercalate into duplex regions of denatured DNA Biochemistry 26 6392
Lu J, Sun Q, Li J L, Gu W, Tianab J L, Liuab X and Yan S P 2013 Synthesis, characterization, and DNA-binding of two new Cd(II) complexes with 8-[(2-pyridylmethyl)amino]-quinoline J. Coord. Chem. 66 3280
Wu H L, Wang K T, Kou F, Jia F, Liu B, Yuan J K and Bai Y 2011 A six-coordinate picrate cadmium(II) complex with a new V-shaped ligand 1,3-bis(1-ethylbenzimidazol-2-yl)-2-thiapropane: Synthesis, crystal structure, and DNA-binding properties J. Coord. Chem. 64 2676
Wu H L, Wang K, Jia F, Liu B, Kou F, Yuan J and Kong J 2010 Synthesis, characterization, and DNA band of a five-coordinate cadmium(II) complex with 1,3-bis(1-benzylbenzimidazol-2-yl)-2-thiapropane J. Coord. Chem. 63 4113
Wu H L, Yuan J, Huang X C, Kou F, Liu B, Jia F, Wang K T and Bai Y 2012 Two zinc(II) and cadmium(II) complexes based on the V-shaped ligand 2, 6-bis(2-benzimidazolyl)pyridine: Synthesis, crystal structure, DNA-binding properties and antioxidant activities Inorg. Chim. Acta 390 12
Pyle A M, Rehmann J P, Meshoyrer R, Kumar C V, Turro N J and Barton J K 1989 Mixed-ligand complexes of ruthenium(II): Factors governing binding to DNA J. Am. Chem. Soc. 111 3051
Vijayalakshmi R, Kanthimathi M, Subramanian V and Nair B U 2000 Interaction of DNA with [Cr(Schiff base)(H2 O)2]ClO4 Biochim. Biophys. Acta 1475 157
Stern O and Volmer M 1919 Über die Abklingzeit der Fluoreszenz Physik. Zeitschr. 20 183
Kathiravan A and Renganathan R 2009 Photoinduced interactions between colloidal TiO2 nanoparticles and calf thymus-DNA Polyhedron 28 1374
Wang B, Yang Z Y, Crewdson P and Wang D 2007 Synthesis, crystal structure and DNA-binding studies of the Ln(III) complex with 6-hydroxychromone-3-carbaldehyde benzoyl hydrazone J. Inorg. Biochem. 101 1492
Li D, Tian J, Gu W, Liu X and Yan S 2010 A novel 1,2,4-triazole-based copper(II) complex: Synthesis, characterization, magnetic property and nuclease activity J. Inorg. Biochem. 104 171
Garcia-Gimenez J L, Gonzalez-Alvarez M, Liu-Gonzalez M, Macias B, Borras J and Alzuet G 2009 Toward the development of metal-based synthetic nucleases: DNA binding and oxidative DNA cleavage of a mixed copper(II) complex with N-(9H-purin-6-yl)benzenesulfonamide and 1,10-phenantroline. Antitumor activity in human Caco-2 cells and Jurkat T lymphocytes. Evaluation of p53 and Bcl-2 proteins in the apoptotic mechanism J. Inorg. Biochem. 103 923
Tolia C, Papadopoulos A N, Raptopoulou C P, Psycharis V, Garino C, Salassa L and Psomas G 2013 Copper(II) interacting with the non-steroidal antiinflammatory drug flufenamic acid: Structure, antioxidant activity and binding to DNA and albumins J. Inorg. Biochem. 123 53
Satyanarayana S, Daborusak J C and Chaires J B 1993 Tris(phenanthroline)ruthenium(II) enantiomer interactions with DNA: Mode and specificity of binding Biochemistry 32 2573
Satyanarayana S, Daborusak J C and Chaires J B 1992 Neither.DELTA.- nor.LAMBDA.-tris(phenanthroline) ruthenium(II) binds to DNA by classical intercalation Biochemistry 31 9319
Yuan C-X, Wu Y-B, Wei Y-B, Yang P and Zhu M L 2007 Studies on the Interaction of Dinitratobis(phen) Cadmium Complex with DNA Chin. J. Chem. 25 1267
Gao C, Ma X, Tian J, Li D and Yan S 2010 Synthesis, structure, and DNA binding of three reduced amino-acid Schiff-base zinc(II), nickel(II), and cadmium(II) complexes J. Coord. Chem. 63 115
Ma X-F, Li D-D, Tian J-L, Kou Y-Y and Yan S-P 2009 DNA binding and cleavage activity of reduced amino-acid Schiff base complexes of cobalt(II), copper(II), and cadmium(II) Trans. Met. Chem. 34 475
Park H, Kwon J H, Cho T, Kim J M, Hwang I H, Kim C, Kim S, Kim J and Kim S K 2013 Real-time detection of DNA cleavage induced by [M(2,2′-bipyridine )2(NO3)](NO3) (M = Cu(II), Zn(II) and Cd(II)) complexes using linear dichroism technique J. Inorg. Biochem. 127 46
Acknowledgements
Financial support from the Council of Scientific and Industrial Research (CSIR), New Delhi, India [CSIR Scheme No. 01(2685)/12/EMR-II dated 03-10-2012] is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information (SI)
Crystallographic data for the structural analyses of complexes 1a and 2b have been deposited with the Cambridge Crystallographic Data Centre bearing the CCDC Nos. 1438356 and 1438357, respectively. Copies of this information are available on request at free of charge from CCDC, Union Road, Cambridge, CB21EZ, UK (fax: + 44-1223-336-033; e-mail: deposit@ccdc.ac.uk or http://www.ccdc.cam.ac.uk). NMR spectra (Figures S1–S4), plots of DNA binding experiments for complex 2b from the absorbance and emission study and its comparison with the complex 1 and other reported cadmium(II) complexes, and CIF files of complexes 1a and 2b are available at http://www.ias.ac.in/chemsci.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
MONDAL, B., SEN, B., SARKAR, S. et al. Synthesis, characterization, crystal structure and DNA-binding study of four cadmium(II) pyridine-carboxamide complexes. J Chem Sci 129, 45–55 (2017). https://doi.org/10.1007/s12039-016-1215-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12039-016-1215-9
















 ), 1b (
), 1b (
 ), 2a (
), 2a (
 ), 2b (
), 2b (
 ), and EB (
), and EB (
 ) on the relative viscosity of CT-DNA.
) on the relative viscosity of CT-DNA.