Abstract
Amyotrophic lateral sclerosis (ALS) is a disease caused by the degeneration of motor neurons (MNs) leading to progressive muscle weakness and atrophy. Several molecular pathways have been implicated, such as glutamate-mediated excitotoxicity, defects in cytoskeletal dynamics and axonal transport, disruption of RNA metabolism, and impairments in proteostasis. ALS is associated with protein accumulation in the cytoplasm of cells undergoing neurodegeneration, which is a hallmark of the disease. In this review, we focus on mechanisms of proteostasis, particularly protein degradation, and discuss how they are related to the genetics of ALS. Indeed, the genetic bases of the disease with the implication of more than 30 genes associated with familial ALS to date, together with the important increase in understanding of endoplasmic reticulum (ER) stress, proteasomal degradation, and autophagy, allow researchers to better understand the mechanisms underlying the selective death of motor neurons in ALS. It is clear that defects in proteostasis are involved in this type of cellular degeneration, but whether or not these mechanisms are primary causes or merely consequential remains to be clearly demonstrated. Novel cellular and animal models allowing chronic expression of mutant proteins, for example, are required. Further studies linking genetic discoveries in ALS to mechanisms of protein clearance will certainly be crucial in order to accelerate translational and clinical research towards new therapeutic targets and strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) is the most frequent motor neuron disease with an incidence of 2 per 100,000 persons per year and a prevalence of 5 per 100,000 people [1]. The mean age for ALS diagnosis is 64 years old. Signs of spasticity, resistance to movement, and brisk reflexes occur with upper motor neuron degeneration, while lower motor neurons damage include muscular weakness, muscular atrophy along with fasciculations. The consequences are progressive paralysis and difficulties in swallowing and respiration, leading to death in patients three to 5 years after the diagnosis. Ninety percent of cases are sporadic (SALS), while the other 10% comprise familial amyotrophic lateral sclerosis (FALS). ALS remains incurable, but genetic mutations implicated in the disease point out several impaired cellular and molecular mechanisms and, thus, provide clues for potential therapeutic strategies. These mechanisms include the abnormal response to oxidative stress, glutamate-mediated excitotoxicity, defects in cytoskeletal dynamics and axonal transport, and impaired protein homeostasis [2, 3].
Most neurodegenerative diseases are characterized by an accumulation of misfolded proteins and by the presence of protein aggregates in cells [4]. This is largely due to malfunctions in proteostasis. The term proteostasis refers to the biological pathways involved in the biogenesis of proteins including ribosomal translation, folding and trafficking, and protein degradation in cells. In ALS, the dysfunction of these mechanisms has been suspected for many years. In effect, post-mortem studies of ALS patients have shown protein aggregates in upper and lower motor neurons [5]. Many of these aggregates have been ubiquitin-positive, linking these histological observations to a mechanism of protein degradation, the ubiquitin proteasome system (UPS) [6, 7]. These aggregates have become a cornerstone of the disease. Different types of aggregates have been observed. Ubiquitinated aggregates are classified as either Lewy body-like hyaline inclusions or skein-like inclusions. These last inclusions appear as randomly oriented filaments covered by fine granules. In some other cases, ubiquitination is not present. These aggregates are called Bunina bodies, which are small, eosinophilic, and round hyaline inclusions [8]. In ALS, protein inclusions are mainly ubiquitinated and co-localized with the TDP-43 protein, as initially described in 2006 [9]. These cytoplasmic inclusions are present not only in motor neurons but also in astrocytes in ALS [10, 11]. Subsequently, mutations in the TARDBP gene encoding TDP-43 were described in FALS and SALS patients [12, 13]. The aggregation of several proteins in ALS has also been proposed to proceed via the formation of SGs. SGs are part of ribonucleoprotein (RNP) granules, which are cellular sites dedicated to RNA processing [14]. These structures, whose formation is favored by various types of stress, have no membrane and are visible under the light microscope. They contain several species and, in particular, RNA and ribonucleoproteins.
An increasing amount of misfolded or unfolded proteins is toxic for cells, generating stress mechanisms, such as the response to ER stress. These proteins must be efficiently removed from cells by quality control mechanisms linked to protein degradation pathways. This is particularly true for neurons that are not able to dilute their cytoplasmic contents, including damaged substrates, through cellular division. These mechanisms and pathways also participate in a continuous turnover of proteins and release of amino acids after protein degradation. Certain steps of these degradation pathways are known to be downregulated due to decreased expression levels and activity with age, which pose concern for their implication in several neurodegenerative diseases, such as ALS [15, 16]. Therefore, it has been championed that the presence of protein aggregates in motor neurons and glial cells in FALS and SALS reflects dysfunctions in these quality control protein degradation pathways. The discoveries in ALS patients of mutations in several genes encoding proteins directly linked to these pathways support this hypothesis (Table 1).
In this review, we present defects in proteostasis with respect to ALS, focusing on disturbances in quality control/degradation of proteins as a central mechanism in ALS pathophysiology. We first describe two of the main protein degradation pathways in cells, the UPS and autophagy. Researchers have shown an increased interest in proteostasis studies pertaining to ALS since 1989, when the first article linking ubiquitin and ALS was published (Fig. 1). More than 30 genes associated with familial ALS have been described, 15 of which are directly involved in UPS and/or autophagy mechanisms that will be described further below. Other susceptible genes, which increase the risk of ALS, also play roles in UPS and/or autophagy.
The Ubiquitin Proteasome System
The UPS was associated with protein degradation in the 1980s [33]. Ubiquitination of proteins in cells occurs through the coordinated activity of a unique ubiquitin activating enzyme (E1), a conjugating enzyme (E2), and a ligase (E3) [34]. E1 activates ubiquitin by an ATP-dependent mechanism. Ubiquitin (Ub) is then transferred to an E2 by a transthiolation reaction. We have identified 38 genes encoding E2 enzymes in the human genome [35]. E3 ligases determine the specificity of the ubiquitination process by two main mechanisms: a E3 recognizes the E2-Ub complex and transfers Ub to the target protein, or Ub is transferred from the E2-Ub complex to an E3, and this E3 transfers Ub to the target protein. The human genome encodes more than 500 E3 enzymes [36]. These E3 ligases are implicated in many cellular pathways by acting on protein homeostasis. But they also directly regulate the function of many proteins involved in key mechanisms, such as DNA repair for example [37].
The consequence of the binding of a lysine-48 linked poly-ubiquitin chain to a target protein is its degradation by the 26S proteasome, a multi-subunit complex containing one 20S domain and two 19S subunits (Fig. 2).
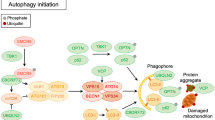
Schematic representation of mechanisms implicated in ubiquitin proteasome system (UPS) and links with ALS. Misfolded proteins are poly-ubiquitinated (Un) through the coordinated activity of an ubiquitin activating enzyme (E1), a conjugating enzyme (E2), and a ligase (E3). UBD (ubiquitin-binding domain) containing proteins (such as UBLQN2, VCP) bind ubiquitin-tagged substrates before degradation by the proteasome 26S. Chaperone-assisted proteasomal degradation (CAP) pathway contributes to improve the presentation of ubiquitinated proteins to the proteasome thanks to co-chaperone and an E3 ligase called CHIP. CHIP is able to bind Hsc70/Hsp70 which are associated with misfolded or aggregated proteins during protein quality control. Other proteins are implicated in CAP: BAG1 recruitment leads to proteasomal degradation and BAG2 binding leads to correct folding and stops degradation. In red, genes mutated in ALS and their implication in UPS: chromosome 9 open reading frame 72 (C9ORF72), ubiquilin 2 (UBQNL2), valosin-containing protein (VCP), fused in sarcoma (FUS), cyclin F (CCNF), superoxide dismutase 1 (SOD1)
UPS and ALS
Several studies illustrated or suggested deregulation of UPS in ALS [38,39,40,41]. The presence of ubiquitin in intracellular inclusions has been found in motor neurons of FALS and SALS patients [42, 43]. This positive immune-reactivity for ubiquitin has become a neuropathological feature of ALS. Inclusions also contain components of the proteasome [44], and impaired proteasomal function in motor neurons has been observed in ALS [45]. The decrease in proteasome subunits was observed during disease progression in the spinal cord of these transgenic mice, as well [46]. Another observation was the locomotive alteration accompanied by progressive motor neuron loss detected in the conditional knockout of the murine proteasome subunit Rpt3 in a motor neuron-specific manner [29]. Proteasome inhibition, using lactacystin, induced selective motor neuron death in organo-typic slice cultures [47]. Importantly, genetic evidence has also revealed an implication of defective UPS in ALS pathogenesis. For example ubiquilin 2, whose mutated form causes X-linked ALS/frontotemporal dementia (FTD), physically associates with ubiquitin ligases E3 and the proteasome to mediate protein degradation [48]. Furthermore, mutated valosin-containing protein (VCP) has been discerned in ALS [49]. One of the suggested functions of VCP is to maintain the solubility or to reverse the aggregation of insoluble, misfolded proteins prior to their proteasomal degradation [50]. The best characterized function of VCP is to participate in the endoplasmic reticulum-associated protein degradation (ERAD), a system linked to UPS [51].
ERAD and ALS
Proteins of the secretory pathway represent one third of the proteins synthesized in cells. These proteins enter the ER in order to be properly folded. If proteins remain unfolded or are misfolded in the ER lumen, they enter the endoplasmic reticulum-associated protein degradation (ERAD) pathway. ERAD consists of the translocation of these proteins from the ER to the cytosol to be ubiquitinated and, in turn, degraded by the proteasome machinery [52]. An accumulation of unfolded/misfolded proteins in the ER results in a stress response, which activates the unfolded protein response (UPR). The UPR stimulates the production of chaperones and components of the ERAD to handle this protein accumulation (Fig. 3). The implication of the ER and, particularly, ER stress in ALS has been questionable for many years [53, 54]. A thorough review on this particular point has been previously published [55]. Causes of ER stress in motor neurons are several. Briefly, however, as protein folding is calcium-dependent, an imbalance of calcium levels in the ER may be at play [56]. Inhibition of ER-Golgi transport by mutant SOD1, TDP-43, or FUS can also result in ER stress [57]. The most studied causes of ER stress in ALS are genetic mutations leading to accumulations of proteins in the ER and consequently to a stress.
Schematic representation of endoplasmic reticulum-associated protein degradation (ERAD) and links with ALS. After synthesis, proteins are translocated in the inner face of the ER to be folded and undergo protein quality control. If the folding is correct, proteins are sent to the Golgi apparatus, where they will undergo post-translational modifications. If proteins are misfolded and accumulated, ER is subjected to stress. Unfolded protein response (UPR) up-regulates molecular chaperones involved in protein folding, allowing misfolded protein to be correctly folded. If not, this response encourages degradation of misfolded protein by UPS (ERAD). Because proteasomes are in the cytoplasm, ubiquitinated substrates are translocated back in cytosol through transmembrane proteins cooperating with protein such as UBQLN2 in order to present substrates to degradation. In red, genes mutated in ALS and their implication in ERAD: protein disulfide isomerase family A member (PDIA), sigma non-opioid intracellular receptor 1 (SIGMAR1), vesicle-associated membrane protein-associated protein B/C (VAPB), fused in sarcoma (FUS), valosin-containing protein (VCP), ubiquilin 2 (UBQNL2)
SUMO Pathway
The UPS system has a tight relationship with the SUMO pathway. Like ubiquitin, SUMO proteins are expressed as precursors and, after being cleaved by SENPs (sentrin-specific proteases), they enter the SUMO pathway. The main contribution of the SUMO pathway in proteostasis is its ability to cooperate with or balance the UPS system [58]. Indeed, these two systems work cooperatively [59]. Nevertheless, they can compete if a certain lysine residue on the target protein can be either ubiquitinated or SUMOylated. Proteins encoded by genes mutated in ALS patients, such as SOD1, VCP, and TDP-43, are targets of the SUMO pathway. SOD1 mutants co-localize in aggregates in cells with ubiquitin but also with SUMO proteins. Studies have reported that SUMO1 and SUMO3 increase aggregation of SOD1 mutants [60, 61]. Remarkably, we observed an inhibition of mutant SOD1 aggregation in motor neurons through the prevention of its SUMOylation on lysine 75 [62]. We have also reported that other proteins encoded by ALS-related genes, such as OPTN or VAPB, contain SUMO consensus sites and, thus, could be SUMOylated [63]. Finally, an additional observation relating the SUMO pathway to ALS is the fused in sarcoma protein (FUS), a SUMO1 E3 ligase encoded by an ALS causative gene [64].
Chaperone-Assisted Proteasomal Degradation
Molecular chaperones are facilitators of protein folding and assembly. They also play a central role in protein degradation by facilitating the sorting of misfolded proteins to the proteasome or the lysosomal compartment. The chaperone-assisted proteasomal degradation (CAP) is one of these facilitating systems. CAP improves the access of ubiquitinated proteins to the 26S proteasome using a co-chaperone ubiquitin ligase called CHIP. CHIP binds Hsc70/Hsp70 and Hsp90, which are associated with misfolded or aggregated proteins during protein quality control (Fig. 2). In ALS, CHIP promotes the proteasomal degradation of ALS-linked mutant SOD1 by ubiquitinating Hsp/Hsc70 [40]. CHIP-mediated CAP is also linked to the degradation of several proteins that aggregate in neurodegenerative diseases such as Alzheimer’s and Parkinson’s diseases [65]. Studies have revealed that Hsps are sequestered within SOD1-positive aggregates in ALS, reducing their capacity to take care of with misfolded proteins [66] (Fig. 2).
Autophagy
Autophagy plays a significant role in neurodegenerative diseases. For instance, the inactivation of constitutive autophagy, in Atg7 (autophagy-related 7)-deficient mice, results in the formation of Ub-positive, cytoplasmic protein inclusions in neurons and drives behavioral defects and neurodegeneration [67]. Its implication in ALS is critical as demonstrated by the fact that several proteins encoded by genes mutated in FALS participate in autophagy processes. The term autophagy entails three protein-presentation mechanisms on lysosomal vesicles for degradation: macro-autophagy (including chaperone-assisted selective autophagy, CASA), chaperone-mediated autophagy (CMA), and micro-autophagy. Macro-autophagy comprises in the removal of aggregated proteins via the formation of a bilayer membrane autophagosome that encapsulates substrates in the cytosol, such as misfolded proteins, before fusing with the lysosome to allow degradation and recycling (Fig. 4). Chaperone-mediated autophagy (CMA) degrades cytosolic proteins containing the pentapeptide motif, KFERQ, recognized by Hsp70 and sent to lysosomes (Fig. 4).
Schematic representation of autophagy and links with ALS. Chaperone-mediated autophagy (CMA) pathway is dedicated to misfolded proteins containing a specific motif: KFERQ (sequence present in 30% of cytoplasmic proteins). These misfolded proteins are recognized by Hsc70/co-chaperones complex and delivered to the lysosome through lysosomal membrane-associated protein 2A (LAMP-2A), a CMA adaptor. Concerning macro-autophagy, poly-Ub chains on cytoplasmic proteins triggers autophagy receptor recruitment (such as OPTN, p62/sequestosome1, UBLQN2) thanks to their ubiquitin-binding domain. Cargos are associated with a double membrane vacuole (phagophore) through the interaction of autophagic receptor with light chain 3 II (LC3-II). LC3-II proteins are activated by cleavage and lipidation of LC3 form by autophagy-related proteins 5 and 7 (Atg5/7) before incorporation in the phagophore membrane. The phagophore extend his membrane to engulf cargos. Next, the autophagosome can merge with lysosomes to create an autolysosome where protein degradation occurs. Chaperone-assisted selective autophagy (CASA) is a particular macro-autophagy pathway: Hsc/Hsp70 and HspB8 chaperones with the association of BAG3, a co-chaperone, improves the recognition of cargos and attracts components of the autophagy machinery. BAG 3 facilitates binding with p62, an autophagic receptor. In red, genes mutated in ALS and their implication in autophagy: superoxide dismutase 1 (SOD1), optineurin (OPTN), TANK-binding kinase 1 (TBK1), TAR DNA-binding protein (TARDBP), ubiquilin 2 (UBQNL2), p62/sequestosome1 (SQSTM1), chromosome 9 open reading frame 72 (C9ORF72), valosin-containing protein (VCP), fused in sarcoma (FUS), FIG4 phosphoinositide 5-phosphatase (FIG4)
P62/sequestosome1 is an autophagy receptor that binds ubiquitinated proteins and recruits them to the autophagosome [68]. Several observations directly link P62/sequestosome1 to ALS. For example, the depletion in C9ORF72 protein, which is one of the possible pathogenic mechanisms in C9ORF72-linked ALS, causes the accumulation of p62/sequestosome1-positive inclusions in cell lines and primary neurons. Notably, C9ORF72 controls the Rab1a-dependent trafficking of the ULK1 autophagy initiation complex [69]. Another example is that mutant SOD1 can be recognized by p62/sequestosome1 in an ubiquitin-independent manner and targeted to for autophagy [70].
Moreover, optineurin is another important autophagy receptor presenting ubiquitinated substrates to autophagic processes [71]. Target components of autophagy containing p62/sequestosome1 or optineurin, called cargo or substrates, are then associated with a bilayer membrane vacuole, called phagophore, through the interaction with autophagic receptor with light chain 3 II (LC3-II). Then, the phagophore extends its membrane to engulf cargo. This autophagosome can merge with the lysosome to create an autolysosome, where cargo degradation occurs. These formations of vesicles, from phagophores to autolysosomes, are regulated by several proteins, one of which is TANK-binding kinase 1, a cargo recruiter in autophagy [31]. Recently, mutations in its gene TBK1 were found in ALS [72].
Autophagy and ubiquitin proteasome system and have been viewed as two separate pathways with no or little interplay. Recent works indicate that this is not the case. Crosstalk exists from the UPS to the autophagy and from the autophagy to the UPS [73]. For example, bortezomib, a proteasome inhibitor, induces autophagy [74]. Autophagic inhibition can also impair proteasomal function, resulting in protein accumulation and aggregation [75]. Cooperation seems also very important between UPS and autophagy. These two systems act together for protein degradation in cells, with the UPS mainly implicated in degradation of soluble proteins, and autophagy in degradation of insoluble and/or aggregated proteins. For example, CHIP protein is directly implicated in the choice between UPS and autophagy. Indeed, as previously mentioned, CHIP acts in CAP, leading to proteasomal degradation of misfolded proteins. But it can also mediate autophagic degradation of misfolded and/or aggregated proteins [76].
Causative Genes in ALS and Protein Degradation Pathways
Significant advances have been made in the genetic footprint of ALS since the discovery of mutations in the SOD1 gene in ALS patients in 1993 [26, 27]. To date, more than 30 genes have been demonstrated to be involved in familial ALS. They are listed in the Amyotrophic Lateral Sclerosis Online Genetics Database website (http://alsod.iop.kcl.ac.uk/). These genes have mostly been identified through exome screening analysis, candidate gene strategies and linkage approaches have also rendered important results. For instance, a hexanucleotide repeat expansion (HRE) in the first exon of C9ORF72 was discovered in ALS patients thanks to linkage analyses followed by sequencing [77, 78]. These discoveries indicated the involvement of several molecular mechanisms in the disease, including those regulating proteins degradation in cells.
C9ORF72
The pathogenic hexanucleotide repeat expansion (HRE) (GGGGCC)n found in C9ORF72 is the most common genetic cause of ALS [77, 78]. It is present in up to 50% of FALS cases and around 10% of SALS cases [79]. This mutation in C9ORF72 is associated with neuronal inclusions [80, 81]. The function of C9ORF72 protein is still unclear. Conditional knockout of C9ORF72 in mice does not result in motor neuron degeneration [82]. Bioinformatics studies have indicated the presence of a DENN domain (differentially expressed in normal and neoplastic cells) in C9ORF72 protein. This particular domain has been ascertained in more than 20 proteins, whose function is often to act as guanine nucleotide exchange factors (GEFs) for Rab GTPases or Rag GTPases [83, 84]. The consequences of the HRE mutations could be multiple. Three mechanisms have been proposed: loss of function caused by a decreased expression of the protein, production of toxic GGGGCC repeat-containing RNAs, and accumulation of dipeptide repeat proteins (DPR) produced by non-canonical translation (repeat-associated non-ATG translation, RAN) [2, 85, 86].
Ubiquitin-positive inclusions are present in post-mortem brains and in skeletal muscles of C9ORF72 ALS patients, suggesting an implication of the ubiquitin pathway in the pathogenesis [87]. Transcriptional studies in C9ORF72 expansion carriers have displayed significant dysregulation of ubiquitination-related genes such as E2 genes UBE2I, UBE2Q1, UBE2E1, and UBE2N [88] (Fig. 2). The two gain-of function hypotheses, i.e., the production of toxic hexanucleotide repeat-containing RNAs and toxic dipeptide repeat proteins (DPR), could impair the ubiquitin proteasome pathway. Concerning the DPR hypothesis, a recent in vitro study on the effects of proline/arginine dipeptide repeats (20 repeats) in rat primary spinal cord cultures demonstrated a reduction in flux through both the proteasomal and autophagic degradation pathways [89]. A recent study challenges the impact of DPR in neurodegeneration since there is not much DPR aggregation in motor neurons from C9ORF72 mutated ALS patient’s spinal cord [90]. DPR accumulation may be not responsible for motor neuron loss; nonetheless, potential contribution of DPR in the disease could go through other mechanisms than aggregation. In vitro studies related an UPS impairment and evidence of ER stress in primary neurons expressing DPR poly-GA [91].
Neuronal, cytoplasmic inclusions are also positive for p62 protein (sequestosome 1, SQSTM1) in post-mortem brain tissues from C9ORF72 ALS patients [92]. Levels of p62 and LC3, another protein directly involved in autophagy, are increased in mice lacking C9ORF72 [93]. Abnormalities in autophagy pathways have been found in patient-derived neurons [94]. This might be a consequence of a defect in autophagosome formation, because C9ORF72 controls the Rab1a-dependant trafficking of the ULK1 autophagy initiation complex. Cells expressing endogenous tagged C9ORF72, using a CRISPR-Cas9 approach, revealed its localization on the surface of lysosomes when cells were starved of amino acids [17]. Surprisingly, recent studies have indicated the implication of C9ORF72 in a protein complex containing SMCR8 (another DENN domain-containing protein) and WDR41 [18]. These proteins have also been found on lysosomes [19]. This complex regulates the autophagy-lysosome pathway (Fig. 4). Altogether, current knowledge supports a defect of both ubiquitin proteasome pathway and autophagic pathway in ALS pathogenesis linked to C9ORF72 mutation.
SOD1
The superoxide dismutase 1 (SOD1) gene is altered in 20% of FALS cases and in 2–7% of SALS cases (http://alsod.iop.kcl.ac.uk). More than 180 mutations have been described. The enzyme encoded by SOD1 is part of the defense mechanisms against oxidative stress. SOD1 inactivates superoxide radicals by generating dioxygen and hydrogen peroxide [95]. Several transgenic murine lines for SOD1 studies have been generated—knockout mice deficient in SOD1 and mice bearing a human transgene of FALS-associated mutant SOD1 [96, 97]. These mice, which exhibit a reduced life span, together with in vitro experiments, support a gain-of-function mechanism for ALS-linked SOD1 mutations.
SOD1/ubiquitin-positive aggregates are found in ALS patients carrying a mutation in the SOD1 gene [26, 98]. These protein aggregates can be observed both in animal and in cellular models expressing mutants SOD1 [99, 100]. Several mechanisms have proposed to explain the cause of toxicity by mutant SOD1 proteins, but the main hypothesis states that mutations induce structural destabilization, prompting the improper folding of SOD1 followed by its aggregation [101, 102]. The accumulation of mutant, unfolded/misfolded SOD1 proteins in the ER triggers a stress signal that activates the UPR. For example, mutants SOD1 interact with Derlin-1, a component of the ERAD machinery, and trigger ER stress [103]. This ER stress has been observed in human-transgenic animals and cellular models expressing mutants SOD1 [104]. SOD1 mutations can also impaired proteasome activity. Indeed the level of the 20S proteasome was reduced in lumbar spinal motor neurons relative to the surrounding neuropil in the spinal cord of transgenic mice expressing the familial ALS superoxide dismutase 1 (SOD1) G93A mutant. Several strategies have been employed to prevent or diminish the formation of SOD1 positive aggregates. We recently presented that inhibition of SOD1 SUMOylation inhibits mutant SOD1 aggregation in vitro [62]. In neurons, the E3 ubiquitin ligase dorfin ubiquitinates mutant SOD1 proteins and targets them for proteasomal degradation [105]. Interestingly, human Dorfin overexpression in SOD1G93A mice decreases the amount of mutant SOD1 protein in the spinal cord and improves neurological phenotypes [106]. Another mechanism for mutant SOD1 protein toxicity could be a direct effect on proteasomal activity. Double transgenic mice expressing a fluorescent reporter substrate of the proteasome (UbG76V-GFP) and SOD1G93A revealed a lowered expression of proteasomal subunits, followed by a build-up of the reporter substrate [46].
Autophagic processes also appear to play an important role in toxicity generated by mutants SOD1. Parkin, an E3 ubiquitin ligase related to Parkinson’s disease, allows the poly-ubiquitination of mutants SOD1, promoting their degradation by the autophagy-lysosome system [107]. The clearance of aggregates, mutated SOD1 by macro-autophagy is mediated by Hsp70 and its co-chaperone BAG3 [108]. Not long ago, the E3 ligase Mahogunin ring finger 1 (MGRN1) was shown to contribute to the clearance of toxic mutant SOD1 aggregates likely through autophagy [109]. SQSTM1 and ALS2, two ALS-linked factors, have displayed additive protective roles against mutant SOD1-mediated toxicity through modulating proteostasis perhaps by way of autophagic processes [110]. The reason behind the selective degeneration of motor neurons in SOD1-linked ALS is still not understood. One proposed idea argues that these neurons are less able to correctly degrade unfolded/misfolded proteins. It is intriguing to note that comparisons between the NSC-34 motor neuronal cell line and the the C2C12 muscle cell line revealed that the efficiency of activation of the autophagic system in the context of SOD1 mutants expression is reduced in NSC-34 cells. All these data support an implication of both the UPS and autophagy in SOD1-assoiated ALS.
TARDBP
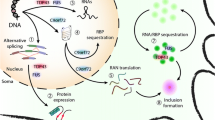
The TARDBP (trans-activation element DNA-binding protein) gene is mutated in 5% of FALS cases and 1% in SALS cases [111, 112]. More than 50 different mutations have been identified. A majority of these mutations are located in the 3′ region encoding a glycin-rich domain in its product, TDP-43. TDP-43 is for the most part expressed in the nucleus. It participates in RNA metabolism in many ways—transcriptional regulation, splicing, mRNA stabilization (including its own transcript), and miRNA processing. TDP-43 also regulates axonal transport and neuronal plasticity [113]. In ALS, TDP-43 is often observed in the cytoplasm, which corresponds to its mislocalization. Post-mortem studies on ALS patients associated with studies on animal and cellular models expressing mutant human TDP-43 support the role of two mechanisms in TDP-43’s pathogenesis [114]: A cytoplasmic build-up of hyper-phosphorylated TDP-43 and a clearance of nuclear TDP-43.
The observations of TDP-43-positive aggregates in post-mortem studies of ALS patients preceded the discovery of mutations in its gene TARDBP [9]. These aggregates were also ubiquitin-positive. Notably, TDP-43-positive aggregates are present even in the absence of mutations in the TARDBP gene. Mutations in other ALS-related genes, such as C9ORF72 or FUS for example are associated with cytoplasmic aggregates containing TDP-43 [115, 116]. A strong connection between TDP-43 and UPS has been proposed. Cells treated with proteasome inhibitors demonstrated an accumulation of ubiquitinated and insoluble TDP-43 particles [117]. Several enzymes participate in TDP-43 ubiquitination, such as UBE2E3 and UBPY [118]. TDP-43 aggregates consist of the adaptor proteins sequestosome 1 and ubiquilin 2, two proteins associated with UPS and autophagy [119]. The precise role of UPS in TDP-43 proteostasis still remains to be determined. Research efforts in cell lines showed that the UPS primarily acts on the degradation of soluble TDP-43 proteins, whereas aggregated TDP-43 requires autophagic clearance [120].
Several observations support an important role for autophagy in TDP-43-linked pathogenesis. RNAi knockdown of TDP-43 in Neuro2A cells shows downregulation of autophagy-related protein 7 (ATG7) at both the mRNA and protein level [121]. ATG7 is an integral constituent of the autophagy machinery (Fig. 4). Several chemical compounds that stimulate autophagy improved TDP-43 clearance and enhanced survival of primary murine neurons and human stem cell-derived neurons harboring mutant TDP-43 [122]. TDP-43 could also influence or interfere with autophagy by an effect on stress granules (SGs). TDP-43 mutants perturb SG dynamics, engendering their persistence in the cytoplasm and a possible toxic gain-of-function [123]. SG breakdown is dependent on selective autophagy [22]. In fact, administration of rapamycin, an inducer of autophagy, promotes SG clearance. It is now clear that TDP-43 is a major factor in motor neuron death in ALS by directly or indirectly disturbing the dynamics of several cellular machineries, such as the UPS, the breakdown of stress granules, and autophagy. Further studies on the mechanisms of TDP-43 aggregation and its localization in the nucleus versus the cytoplasm in motor neurons are necessary. Remarkably, a recent study showed that a physiological oligomerization of TDP-43 through its N-terminal domain antagonizes its pathological aggregation [124].
FUS
The implication of the FUS gene (fused in sarcoma) mutations in ALS was recognized in 2009 [23, 125]. Mutant FUS is observed in 4% of FALS patients and 1% of SALS patients [23, 112, 125]. More than 79 mutations have been described at present, predominantly in the 3′ region encoding an arginine/glycine-rich region and a NLS domain (nuclear localization signal). FUS, which is essentially localized in the nucleus, regulates RNA processing, splicing, and mRNA trafficking [126]. FUS can also bind DNA, taking part in genomic integrity. Furthermore, FUS possesses SUMO E3 ligase activity [64] (Fig. 2). In contrast to Tdp-43, the homozygous knockout of Fus is lethal in embryos [127]. FUS’s connection to ALS pathogenesis could be related to both loss and gain-of-function mechanisms [128].
Pathogenic mutations of FUS are associated with aggregates immune-reactive for FUS within the cytoplasm of neurons and glial cells [129]. These aggregates and the mislocalization of FUS appear essential in the processes of degeneration in ALS. In Drosophila, the ectopic expression of human FUS mutant influences motor neuron degeneration. Strikingly, if mutations are introduced into the RNA-binding region of FUS, aggregation and degeneration are abridged, suggesting that the RNA-binding activity of FUS is critical for aggregate formation [130]. Several studies have exhibited a co-localization of FUS and ubiquitin in some aggregates, but not all [24, 131]. This suggests that ubiquitination of FUS could be a late event in the formation of aggregates and that the UPS may not be crucial in FUS-associated neurodegeneration [132].
A recent work on transgenic mice overexpressing FUS without a NLS domain showed neuronal loss in the motor cortex and the presence of ubiquitin/p62-positive aggregates. RNA-seq analysis revealed specific transcriptome alterations, such as genes regulating endoplasmic reticulum stress [133]. Studies also showed an implication of FUS in SGs dynamics. ALS-associated mutants of FUS increased the lifetime of SGs in the cytoplasm [134]. These mutants also disrupt the autophagy mechanisms by inhibiting the formation of autophagosomes [135] (Fig. 4). Overexpression of Rab1, a protein involved in autophagosome formation, restores the autophagy defects induced by mutant FUS. Despite these recent efforts, the role of autophagy in degeneration associated with mutant FUS remains enigmatic. Nevertheless, many findings advocate that abnormalities in RNA metabolism encompass the cause of neurodegeneration in FUS-linked ALS. Altered RNA metabolism promotes the expression of atypical RNAs and proteins that form aggregates, disrupting the ubiquitin pathway and autophagy.
OPTN
Mutations in the OPTN gene coding for optineurin were first linked to glaucoma [136]. The mutation in OPTN was then described in ALS [137]. Mutations were found in approximately 1% (2/161) of FALS and 3.5% (4/113) of SALS in an Italian cohort [138]. In a Japanese cohort, 3.8% of FALS patients and 0.29% of SALS patients carried OPTN mutations [28]. Optineurin plays a role in the maintenance of Golgi apparatus and vesicular and membrane trafficking. It contains a ubiquitin-binding domain (UBD) and a LC3-interacting region (LIR), indicating functions in both the UPS and autophagy.
Optineurin-positive aggregates are present in FALS and less frequently in SALS. These inclusions are also composed of ubiquitin and other proteins involved in ALS, such as TDP-43 and FUS [131, 137]. A recent work showed that optineurin is mainly degraded by the UPS [139]. The E3 ubiquitin ligase Hdr1 might be a key protein in this pathway because its overexpression enhances the degradation of optineurin. This study also suggested that the formation of optineurin-containing aggregates is dependent on the UPS. Autophagy seems to play a less significant role in this particular function. However, optineurin has paramount functions in autophagy in physiological conditions. It is a ubiquitin-binding receptor that interacts, through its UBD, amino acid 474–479, in a complex with TBK1 (TANK-binding kinase 1), another protein whose gene is mutated in ALS [140]. TBK1 phosphorylates optineurin, enhancing its interaction (amino acid region 412–520 containing UBD) with LC3 [71] (Fig. 4). Optineurin also regulates autophagy through its interaction with myosin VI, which promotes the fusion of autophagosomes with lysosomes [141, 142] (Fig. 4). Genetic analyses of ALS patients pinpoint a mutation within the UBD (p.E478G) which provokes alteration in LC3 level and turnover thereupon decreased autophagy and displayed weakened interaction with myosin VI, engendering autophagosomes’ poor maturation [143]. Conjointly, this mutant and another ALS-related mutation (p.Q398X) disrupt myosin VI-mediated autophagosome-lysosome fusion and intracellular trafficking pathways and induce ER stress [144]. Recent studies support several major roles for optineurin in neuroprotection: regulation of autophagy, participation in inflammatory signaling, blockade of necroptosis, and regulation of apoptosis [145]. A recent study reports that OPTN regulates apoptosis via linear ubiquitin binding [146]. Additional research will consequently be necessary to precisely describe which functions of optineurin are particularly involved in neurodegenerative processes in ALS.
SQSTM1/p62
Mutations in SQSTM1 have been described in 1 to 3% of patients with FALS [25, 147]. SQSTM1 encodes the protein p62/SQSTM1 (sequestosome 1) which regulates various biological processes such as transcription, degradation by UPS and autophagy, and apoptosis.
Aggregates harboring p62 have been identified in patients with ALS and ALS/FTD [148]. p62 is a ubiquitin-binding protein and, thus, may be involved in the regulation of the UPS [149]. p62 is better known for its role in autophagy. Indeed, as optineurin, p62 is an autophagy receptor that interacts with LC3, a key protein in autophagy [150] (Fig. 4). ALS-associated mutations located in the LC3-interacting region of p62 have been characterized in ALS [151]. Inhibition of p62 expression utilizing RNA interference has been seen to lead to autophagy defects. In vivo studies on a zebrafish knockdown model on a sqstm1 ortholog have showed locomotive impairment and have supported an association between ALS and mutated SQSTM1 through a loss-of-function mechanism [152]. Yet, gain-of-function mechanism is also fathomable, because p62 is related to the UPS and the proteostatic and redox balance. p62 participates in one of the mechanisms controlling redox stress, the Keap1-Nrf2 pathway that activates the expression of antioxidant enzymes [153, 154]. In fact, a recent study proposed that the E3 ligase TRIM21 ubiquitinates p62 and that the binding of p62 to ubiquitin plays an essential role in p62-regulated redox homeostasis [155].
VCP
Mutations in the VCP gene (valosin-containing protein, p97) were initially observed in inclusion body myopathy associated with Paget Disease of bone and frontotemporal dementia [156]. VCP’s pathology is characterized by aggregates containing ubiquitin and TDP-43. Neuroanatomic results have revealed the presence of VCP in ubiquitin-staining inclusions in several other neurodegenerative diseases, including ALS [157]. A mutation in VCP has been also identified in familial case of ALS with dominant inheritance [49]. Today, mutations in the VCP gene are found in approximately 2% of FALS case studies [49]. This gene codes for a protein from the AAA+ ATPase family involved in protein homeostasis and protein structure remodeling [30]. Loss of VCP in mouse models is lethal prior to implantation, indicating a crucial role for this protein [158].
VCP is involved in many cellular functions, including regulation of transcription and cell proliferation. Its participation in ALS pathogenesis may be related to its role in mitochondrial energy production [159]. But VCP is best recognized for its role in proteostasis, particularly for the UPS. It binds to ubiquitinated proteins through its N-terminal domain [160] (Fig. 2). This function overlaps with the ERAD mechanism, which allows the release of aberrant proteins from the ER to be degraded by the proteasome [161] (Fig. 3). VCP has links with ER stress. The mutation in the domain of VCP that encodes a hexameric ATPase crucial for protein degradation during ER stress. Also, ER stress caused by protein aggregation activates the SUMOylation of VCP, a post-translational modification, generating its assembly into a hexameric form and its capacity to participate in the ERAD response [162]. The SUMOylation of VCP at its N-terminal domain is reduced by ALS-causing mutations. Another link between VCP and the UPS is its interaction with PI31 protein (PMSF1), a regulator of proteasome complex activity [163]. It is very likely that there is a strong relationship between defects in the UPS caused by mutated VCP and motor neuron degeneration in ALS, but defects in autophagy may also be at play. Indeed, depletion of VCP in cell cultures disrupts autophagosome maturation [164, 165] (Fig. 4). Also, a recent study showed that VCP regulates the clearance of lysosomes by autophagy [166] (Fig. 4). In motor neurons, many endogenous factors, such as protein aggregates and reactive oxygen species (ROS), induce lysosomal damage. A loss of this particular function of VCP in autophagy could, as a result, influence ALS pathogenesis, by way of defects in protein degradation by the UPS.
UBQLN2
The UBQLN2 gene is currently the only X-linked gene involved in ALS. Its mutation results in a dominant inheritance pattern [48]. UBQLN2 encodes for the ubiquilin 2 protein containing an N-terminal ubiquitin-like domain (UBL) and a C-terminal ubiquitin-associated domain (UBA) [167]. The UBA domain binds poly-ubiquitinated proteins, while the UBL domain binds the cap of the 26S proteasome. Thus, ubiquilin 2 delivers poly-ubiquitinated proteins to the proteasome for degradation (Fig. 2). It is also involved in autophagy, cell signaling, cell cycle progression, and cytoskeletal association. [20].
Histopathological analyses on familial and sporadic ALS cases have showed aggregates with ubiquilin 2, regardless of UBQLN2 mutation. These aggregates are sometimes also positive for TDP-43, FUS, p62, and ubiquitin [21, 48]. A recent study showed that ubiquilin 2 associates with the chaperone HSP70 to clear protein aggregates via the proteasome, and that ALS-related mutants ubiquilin 2 are defective in chaperone binding [168]. Moreover, ubiquilin 2 appears to be involved in the ERAD response [169] (Fig. 3). Concerning autophagy, ubiquilin 2 interacts with LC3, suggesting that it participates in delivering cargo to autophagosomes [170, 171] (Fig. 4). This could mean that mutant UBQNL2 fails to deliver cargo to the proteasome for degradation [172] (Fig. 4). Altogether, these findings indicate that ubiquilin 2 has a significant function in regulating protein homeostasis by mediating protein degradation by UPS and autophagy and thus acts as a neuroprotective protein. A deficit in functional ubiquilin 2 would therefore be a risky situation for motor neurons. Current efforts in rodent models, such as transgenic rats, have advocated that an excess of ubiquilin 2 instills a toxic gain-of-function in motor neurons [173]. In contrast, another study showed that mice expressing ALS/FTD-linked UBQLN2 mutants demonstrate cognitive deficits and develop motor neuron disease, but mice overexpressing WT UBQLN2 were mostly devoid of clinical and pathological signs of ALS [174]. In conclusion, the relation between UBQLN2 mutations and defects in proteostasis appears to be paramount in ALS, but the precise mechanisms have not been unequivocally identified. A possible solution could result from studies done directly on patients’ cells. For example, a recent work on lymphoblasts from ALS patients carrying mutations in UBQLN2 reported increased LC3-II and p62 levels, supporting a deregulated proteasome and lysosome in ALS [175]. Further investigations on these cells could further shed extensive information on this matter.
VAPB
VAMP/synaptobrevin-associated protein (VAPB) gene is mutated in rare cases of ALS. The first mutation was discovered in a Brazilian family in 2004 [32]. This study, together with others, indicated that mutations in VAPB are associated with the following phenotypes with dominant, autosomal inheritance: classic, adult form of ALS, atypical form with postural tremor, and adult late-onset proximal SMA [32]. VAPB participates in the UPR [176] (Fig. 3). However, mechanisms by which VAPB mutations cause motor neuron degeneration are still unclear.
The most studied mutation, P56S, causes aggregation of VAPB in transfected cells and animal models [177, 178]. Overexpression of wild-type or mutant VAPB in primary motor neuronal cultures leads to cytosolic aggregates and ER stress [179]. Chen and collaborators also performed functional studies on neuronal cells expressing mutant VAPB (VAPBT46I). They observed intracellular protein aggregates containing ubiquitin ultimately resulting in cell death [180]. Co-transfection experiments have suggested that mutant VAPB inhibits the ability of wild-type VAPB from mediating the unfolded protein response. These data support the argument that mutations in VAPB in ALS stimulate ER stress and produce defects in the UPS, contributing to weakened protein homeostasis and, consequently, to motor neuronal loss. Interestingly, cellular models expressing moderate levels of mutant VAPB contain cytoplasmic aggregates that are not only cytosolic but are also associated with the ER. In these models, VAPB mutants cause dramatic ER restructuring [181, 182].
TBK1
TBK1 (TANK1-binding kinase 1) has recently emerged as a new gene related to ALS, ALS-FTD, and FTD. Causal mutations could be responsible for 0.4 to 4% of ALS cases [72, 183, 184]. This protein is involved in two pathological mechanisms in ALS: inflammation and autophagy [185, 186]. TBK1 participates in the autophagosomes engulfment of poly-Ub targets by phosphorylating autophagic receptors, such as OPTN and p62/SQSTM1 [71, 187, 188] (Fig. 4). Phosphorylation aids to increase the affinity for LC3 [189].
Neuropathological studies of FTD patient carrying mutations in TBK1 display moderate to high amount of TDP-43 and p62 neuronal, cytoplasmic inclusions in agreement with a TDP-43 proteinopathy [72, 190]. These observations support the hypothesis that a loss of TBK1 function induces flaws in protein clearance in neurons. The relationship between TBK1 and the autophagy adapters OPTN and p62 also enforces the notion that a defect in mitophagy leading to neurodegeneration [191, 192]. It is worthwhile to note that ALS/FTD-linked mutations in TBK1 result in a reduction in the amount of TBK1 and/or in modifications in its C-terminal region capable of binding OPTN [72]. TBK1 is also implicated in the innate immune response by regulating the production of IFNα and β [193]. These various roles of TBK1 in the central nervous system suggest a pathogenic mechanism. TBK1 mutations probably generate malfunctions in protein clearance and mitochondrial turnover, which would lead to neuronal damage triggering innate responses by surrounding neurons.
FIG4
Mutations have also been discovered in the FIG4 gene in ALS [194]. However, its role remains debatable with respect to ALS [195]. A recent finding, nonetheless, suggests a role specifically in ALS with longer disease duration and upper motor neuron predominance [196]. FIG4 is a member of the SAC domain-containing protein family with a phosphoinositide 5-phosphatase activity. FIG4 regulates the cellular level of PI(3,5)P2 maintaining endomembrane homeostasis and endosomal trafficking. PI(3,5)P2 is recognized by lysosomes and merges to gain access to the lysosomal machinery. Inactivation of FIG4 in spontaneous mutant mice reveals neurodegeneration and large vacuoles containing lysosome receptor LAMP-2A, suggesting perturbations in the endosome-lysosome network (Fig. 4). Autophagic substrates (p62, Ub, LAMP-2A, and LC3-II) are upregulated and accumulate in neurons in the spinal cord of these mutant mice.
SIGMAR1
The SIGMAR1 gene encodes for the transmembrane chaperone called sigma non-opioid intracellular receptor 1 (Sig-R1) that localizes predominantly on the mitochondrion-associated ER membrane [197]. Sig-R1 not only facilitates the proper folding of newly synthesized proteins, but also prevents the accumulation of misfolded proteins by consigning them to the ERAD [198] (Fig. 3). Sig-R1 has also a role in autophagy through its participation in autophagosome maturation [199]. It is also involved in the ER stress response, Ca2+ metabolism, and chaperone activity, all implicated in neurodegeneration [200]. In 2011, a missense mutation in SIGMAR1 leading to an amino acid change in the transmembrane domain of Sig-R1 was associated with a juvenile ALS incidences [201]. This mutation creates an abnormal subcellular distribution and modification of Sig-R1 provoking modifications in the ER structure, formation of ER-derived autophagic vacuoles, and the induction of ER stress [202]. A contemporary study on an ALS-linked mutant, E102Q of SIGMAR1, proposed a synergistic mechanism of both a gain in toxic function and a loss of function involving altered ER activity, a lack of in protein homeostasis, and a dysregulation of RNA-binding proteins [203].
CCNF
Another current study by whole exome sequencing of an ALS-FTD family identified the CCNF gene as a new ALS-generating gene [204]. CCNF encodes the ubiquitously expressed cyclin F protein member of cyclin protein family that acts in cell cycle, mainly by activating cyclin-dependent kinases (CDK) enzymes. Compared to others cyclins, cyclin F is the only one that cannot bind CDK enzymes. Actually, it is part of a Skp1-Cul1-F-box (SCF) E3 ubiquitin ligase complex enabling ubiquitination of target proteins for proteasome-directed degradation [205, 206] (Fig. 2). Later studies identified other mutations in CCNF in FALS and SALS cases from various countries. Functional studies on some of these mutant proteins in NSC-34 and Neuro-2A cell lines showed perturbations in the UPS and increased ubiquitination of TDP-43.
PDIA1 and PDIA3
PDIA1 (P4HB) and PDIA3 (ERp57) are genes encoding the proteins disulfide-isomerases 1 and 3, respectively [207]. These proteins are mostly present in the ER where proteins are synthesized. They catalyze the rearrangement of disulfide bond between cysteine residues in proteins (Fig. 3). This confers to PDIA proteins their role as chaperones in protein folding. In 2015, a novel study described variants in these two PDIA genes in ALS patients [208]. Functional studies in zebrafish modes and in cell lines showed that these ALS-related PDIA mutations cause defects in the function of PDIA1/3 proteins leading to the disruption of motor neuron connectivity and the impairment neuritogenesis [209]. This observation supports ER proteostasis imbalance as a risk factor for ALS.
Proteostasis and Therapeutic Strategies in ALS
Recent genetic discoveries in ALS will have reasons for clinical management [27]. We can also hope that these discoveries lead more rapidly to therapeutics against ALS. Evidently, the mechanisms of proteostasis are attractive targets. Many compounds targeting various elements of the UPS and autophagy pathways are currently being tested, either as single agents or in combination, in various pathologies including cancer and neurodegenerative diseases, such as ALS. It is possible to remove pathogenic proteins by stimulating the expression or function of chaperones engaged in the UPS or in autophagy [210]. Indeed, the specificity of certain heat-proteins to misfolded proteins can be stimulated for degradation by autophagy. For example, the enhanced expression of Hsp22 increases clearance of mutant SOD1 [211]. Directly targeting autophagy in neurodegenerative diseases with aggregates may also be of interest, as shown by several studies employing small molecules such as valproate, rapamycin, or lithium, which are autophagic inducers [212,213,214]. However, they do not have the same impact depending on the disease. Rapamycin, an inhibitor of mTOR, ameliorates clearance of pathogenic aggregates in Parkinson’s and Alzheimer’s mouse models but not in a SOD1G93A model of ALS. The mood stabilizer lithium succeeds in promoting the degradation of SOD1G93A in cytoplasmic inclusions and extends survival and delays the onset of ALS [214]. Targeting heat shock proteins seems also interesting, since these proteins are participants in the unfolded protein response, proteasomal degradation and autophagy [215]. Several studies highlight the neuroprotective effects of arimoclomol, a hydroxylamine derivative that co-induces heat shock proteins expression. Still under investigation, this compound extends motor neuron lifespan, diminishes muscular dysfunction, and protects motor neurons in a mouse model of ALS [216, 217]. In addition, these small molecules can have interesting effects on cells by acting on other pathways, leading to synergistic effects. The development of new monoclonal antibodies could also create a very interesting future for ALS treatment, as in other neurodegenerative diseases [218, 219]. Injected by the intracerebroventricular route, a monoclonal antibody against mutant SOD1 was able to reduce its level in the spinal cord and to prolong the lifespan of transgenic SOD1G93A mice [220]. Another captivating strategy to reduce the formation of misfolded proteins aggregates is, of course, to reduce the synthesis of the mutated protein. Several antisense strategies are currently being examined. For example, an ISIS antisense technology targeting at the genetic level the production of SOD1 protein and its mutant form is presently being tested in ALS patients (ClinicalTrials.gov ID:NCT02623699). The mouse model SOD1 G93A is the most commonly used animal model to test potential therapeutic agents in ALS. Several preclinical studies using these mice reported interesting results as previously described. However, success in human clinical trials following preclinical-studies using these mice was rare, suggesting for example that this animal model is not relevant for all preclinical studies. Indeed, this model could be of interest for only familial forms of ALS with mutation in SOD1 gene (5% of FALS), but not for the majority of ALS patients [221]. Nevertheless, in conclusion, blocking aggregation, modifying aggregate formation dynamics, modifying the location of aggregates in cells, and promoting disaggregation all seem to be promising research for therapeutic strategies in combatting ALS.
Conclusion
The observation of aggregates of misfolded proteins in ALS has argued for more than 20 years in the implication of impaired protein degradation pathways in the disease. In parallel with these histological observations, mutations have been discovered in SOD1, TARDBP, and FUS genes. Interestingly, proteins encoded by these genes turn out to be also present in aggregates. Thirty other genes associated with familial ALS have been linked to ALS to date [222]. Many of them are directly involved in proteostasis and particularly in the protein degradation pathways UPS and autophagy. The UPS plays a major role in the degradation of a plethora of cellular proteins, especially short-lived proteins. Autophagy also plays an essential role in degrading misfolded or abnormally long-lived proteins. The proper activity of these pathways is essential to remove misfolded proteins that are naturally produced in cells. These pathways are not independent; several connections exist such as their links with ER stress. These two mechanisms, UPS and autophagy, become less efficient with age. In ALS, genetic mutations and environmental factors such as oxidative stress and excitotoxicity promote the production of misfolded proteins by acting directly on mechanisms of proteostasis [223]. The UPS and/or autophagy dysfunction induces vulnerability that ultimately lead to cell death. Thus, the misfolding of proteins and certainly their aggregation are directly involved in ALS pathogenesis. One has yet to understand the relationships between these steps. Numerous studies argue to investigate more closely the intracellular localization of these aggregates and their formation instead of their size [224]. For these experiments, it will be essential to develop innovative cellular and animal models with chronic expression of mutant proteins, for example, and not simply their overexpression. These studies can be performed ideally beginning directly from cells of the patient, such as lymphocytes if the gene is expressed there. The studies can also be done using differentiated motor neurons or glial cells from induced pluripotent stem cells derived from fibroblasts of patients. The CRISPR/CAS 9 system could also be employed in order to create models carrying ALS mutations that would better mirror the conditions of the patients. These new models will certainly aid in better understanding the mechanisms of aggregation, the precise location of these aggregates in cells, and in testing novel therapeutic strategies in the upcoming years.
Abbreviations
- MN:
-
Motor neuron
- ALS:
-
Amyotrophic lateral sclerosis
- UPS:
-
Ubiquitin proteasome system
- ERAD:
-
Endoplasmic reticulum-associated protein degradation
- CAP:
-
Chaperone-assisted proteasomal degradation
- CMA:
-
Chaperone-mediated autophagy
- CASA:
-
Chaperone-assisted selective autophagy
- Ub:
-
Ubiquitin
- SUMO:
-
Small ubiquitin-like modifier
- SG:
-
Stress granules
- C9ORF72:
-
Chromosome 9 open reading frame 72
- SOD1:
-
Superoxide dismutase 1
- TARDBP:
-
TAR DNA-binding protein
- FUS:
-
Fused in sarcoma
- OPTN:
-
Optineurin
- SQSTM1:
-
p62/sequestosome1
- VCP:
-
Valosin-containing protein
- UBQNL2:
-
Ubiquilin 2
- VAPB:
-
Vesicle-associated membrane protein-associated protein B/C
- TBK1:
-
TANK-binding kinase 1
- FIG4:
-
FIG4 phosphoinositide 5-phosphatase
- SIGMAR1:
-
Sigma non-opioid intracellular receptor 1
- CCNF:
-
Cyclin F
- PDIA:
-
Protein disulfide isomerase family A member
References
Marin B, Boumédiene F, Logroscino G et al (2016) Variation in worldwide incidence of amyotrophic lateral sclerosis: a meta-analysis. Int J Epidemiol. https://doi.org/10.1093/ije/dyw061
Taylor JP, Brown RH, Cleveland DW (2016) Decoding ALS: from genes to mechanism. Nature 539:197–206. https://doi.org/10.1038/nature20413
Brown RH, Al-Chalabi A (2017) Amyotrophic lateral sclerosis. N Engl J Med 377:162–172. https://doi.org/10.1056/NEJMra1603471
Forman MS, Trojanowski JQ, Lee VM-Y (2004) Neurodegenerative diseases: a decade of discoveries paves the way for therapeutic breakthroughs. Nat Med 10:1055–1063. https://doi.org/10.1038/nm1113
Oda M, Akagawa N, Tabuchi Y, Tanabe H (1978) A sporadic juvenile case of the amyotrophic lateral sclerosis with neuronal intracytoplasmic inclusions. Acta Neuropathol (Berl) 44:211–216
Murayama S, Ookawa Y, Mori H et al (1989) Immunocytochemical and ultrastructural study of Lewy body-like hyaline inclusions in familial amyotrophic lateral sclerosis. Acta Neuropathol (Berl) 78:143–152
Matsumoto S, Hirano A, Goto S (1990) Ubiquitin-immunoreactive filamentous inclusions in anterior horn cells of Guamanian and non-Guamanian amyotrophic lateral sclerosis. Acta Neuropathol (Berl) 80:233–238
Blokhuis AM, Groen EJN, Koppers M et al (2013) Protein aggregation in amyotrophic lateral sclerosis. Acta Neuropathol (Berl) 125:777–794. https://doi.org/10.1007/s00401-013-1125-6
Neumann M, Sampathu DM, Kwong LK et al (2006) Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314:130–133. https://doi.org/10.1126/science.1134108
Mackenzie IRA, Bigio EH, Ince PG et al (2007) Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann Neurol 61:427–434. https://doi.org/10.1002/ana.21147
Tan C-F, Eguchi H, Tagawa A et al (2007) TDP-43 immunoreactivity in neuronal inclusions in familial amyotrophic lateral sclerosis with or without SOD1 gene mutation. Acta Neuropathol (Berl) 113:535–542. https://doi.org/10.1007/s00401-007-0206-9
Kabashi E, Valdmanis PN, Dion P et al (2008) TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet 40:572–574. https://doi.org/10.1038/ng.132
Sreedharan J, Blair IP, Tripathi VB et al (2008) TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319:1668–1672. https://doi.org/10.1126/science.1154584
Kedersha N, Stoecklin G, Ayodele M et al (2005) Stress granules and processing bodies are dynamically linked sites of mRNP remodeling. J Cell Biol 169:871–884. https://doi.org/10.1083/jcb.200502088
Hara T, Nakamura K, Matsui M et al (2006) Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441:885–889. https://doi.org/10.1038/nature04724
Mizushima N (2010) The role of the Atg1/ULK1 complex in autophagy regulation. Curr Opin Cell Biol 22:132–139. https://doi.org/10.1016/j.ceb.2009.12.004
Amick J, Roczniak-Ferguson A, Ferguson SM (2016) C9orf72 binds SMCR8, localizes to lysosomes, and regulates mTORC1 signaling. Mol Biol Cell 27:3040–3051. https://doi.org/10.1091/mbc.E16-01-0003
Sullivan PM, Zhou X, Robins AM et al (2016) The ALS/FTLD associated protein C9orf72 associates with SMCR8 and WDR41 to regulate the autophagy-lysosome pathway. Acta Neuropathol Commun 4:51. https://doi.org/10.1186/s40478-016-0324-5
Schröder B, Wrocklage C, Pan C et al (2007) Integral and associated lysosomal membrane proteins. Traffic Cph Den 8:1676–1686. https://doi.org/10.1111/j.1600-0854.2007.00643.x
Lee DY, Brown EJ (2012) Ubiquilins in the crosstalk among proteolytic pathways. Biol Chem 393:441–447. https://doi.org/10.1515/hsz-2012-0120
Williams KL, Warraich ST, Yang S et al (2012) UBQLN2/ubiquilin 2 mutation and pathology in familial amyotrophic lateral sclerosis. Neurobiol Aging 33:2527.e3–2527.e10. https://doi.org/10.1016/j.neurobiolaging.2012.05.008
Buchan JR, Kolaitis R-M, Taylor JP, Parker R (2013) Eukaryotic stress granules are cleared by autophagy and Cdc48/VCP function. Cell 153:1461–1474. https://doi.org/10.1016/j.cell.2013.05.037
Vance C, Rogelj B, Hortobágyi T et al (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323:1208–1211. https://doi.org/10.1126/science.1165942
Neumann M, Rademakers R, Roeber S et al (2009) A new subtype of frontotemporal lobar degeneration with FUS pathology. Brain J Neurol 132:2922–2931. https://doi.org/10.1093/brain/awp214
Fecto F, Yan J, Vemula S et al (2011) SQstm1 mutations in familial and sporadic amyotrophic lateral sclerosis. Arch Neurol 68:1440–1446. https://doi.org/10.1001/archneurol.2011.250
Rosen DR, Siddique T, Patterson D et al (1993) Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362:59–62. https://doi.org/10.1038/362059a0
Al-Chalabi A, van den Berg LH, Veldink J (2017) Gene discovery in amyotrophic lateral sclerosis: implications for clinical management. Nat Rev Neurol 13:96–104. https://doi.org/10.1038/nrneurol.2016.182
Iida A, Hosono N, Sano M, et al. (2012) Novel deletion mutations of OPTN in amyotrophic lateral sclerosis in Japanese. Neurobiol Aging 33:1843.e19-24. doi: https://doi.org/10.1016/j.neurobiolaging.2011.12.037
Tashiro Y, Urushitani M, Inoue H et al (2012) Motor neuron-specific disruption of proteasomes, but not autophagy, replicates amyotrophic lateral sclerosis. J Biol Chem 287:42984–42994. https://doi.org/10.1074/jbc.M112.417600
Xia D, Tang WK, Ye Y (2016) Structure and function of the AAA+ ATPase p97/Cdc48p. Gene 583:64–77. https://doi.org/10.1016/j.gene.2016.02.042
Oakes JA, Davies MC, Collins MO (2017) TBK1: a new player in ALS linking autophagy and neuroinflammation. Mol Brain 10:5. https://doi.org/10.1186/s13041-017-0287-x
Nishimura AL, Mitne-Neto M, Silva HCA et al (2004) A mutation in the vesicle-trafficking protein VAPB causes late-onset spinal muscular atrophy and amyotrophic lateral sclerosis. Am J Hum Genet 75:822–831. https://doi.org/10.1086/425287
Ciechanover A, Heller H, Elias S et al (1980) ATP-dependent conjugation of reticulocyte proteins with the polypeptide required for protein degradation. Proc Natl Acad Sci U S A 77:1365–1368
Komander D (2009) The emerging complexity of protein ubiquitination. Biochem Soc Trans 37:937–953. https://doi.org/10.1042/BST0370937
Michelle C, Vourc’h P, Mignon L, Andres CR (2009) What was the set of ubiquitin and ubiquitin-like conjugating enzymes in the eukaryote common ancestor? J Mol Evol 68:616–628. https://doi.org/10.1007/s00239-009-9225-6
Li W, Bengtson MH, Ulbrich A et al (2008) Genome-wide and functional annotation of human E3 ubiquitin ligases identifies MULAN, a mitochondrial E3 that regulates the organelle’s dynamics and signaling. PLoS One. https://doi.org/10.1371/journal.pone.0001487
Natarajan C, Takeda K (2017) Regulation of various DNA repair pathways by E3 ubiquitin ligases. J Cancer Res Ther 13:157–169. https://doi.org/10.4103/0973-1482.204879
Cheroni C, Peviani M, Cascio P et al (2005) Accumulation of human SOD1 and ubiquitinated deposits in the spinal cord of SOD1G93A mice during motor neuron disease progression correlates with a decrease of proteasome. Neurobiol Dis 18:509–522. https://doi.org/10.1016/j.nbd.2004.12.007
Puttaparthi K, Wojcik C, Rajendran B et al (2003) Aggregate formation in the spinal cord of mutant SOD1 transgenic mice is reversible and mediated by proteasomes. J Neurochem 87:851–860
Urushitani M, Kurisu J, Tateno M et al (2004) CHIP promotes proteasomal degradation of familial ALS-linked mutant SOD1 by ubiquitinating Hsp/Hsc70. J Neurochem 90:231–244. https://doi.org/10.1111/j.1471-4159.2004.02486.x
Urushitani M, Kurisu J, Tsukita K, Takahashi R (2002) Proteasomal inhibition by misfolded mutant superoxide dismutase 1 induces selective motor neuron death in familial amyotrophic lateral sclerosis. J Neurochem 83:1030–1042
Leigh PN, Whitwell H, Garofalo O et al (1991) Ubiquitin-immunoreactive intraneuronal inclusions in amyotrophic lateral sclerosis. Morphology, distribution, and specificity. Brain J Neurol 114(Pt 2):775–788
Migheli A, Autilio-Gambetti L, Gambetti P et al (1990) Ubiquitinated filamentous inclusions in spinal cord of patients with motor neuron disease. Neurosci Lett 114:5–10
Hyun D-H, Lee M, Halliwell B, Jenner P (2003) Proteasomal inhibition causes the formation of protein aggregates containing a wide range of proteins, including nitrated proteins. J Neurochem 86:363–373
Kabashi E, Agar JN, Taylor DM et al (2004) Focal dysfunction of the proteasome: a pathogenic factor in a mouse model of amyotrophic lateral sclerosis. J Neurochem 89:1325–1335. https://doi.org/10.1111/j.1471-4159.2004.02453.x
Cheroni C, Marino M, Tortarolo M et al (2009) Functional alterations of the ubiquitin-proteasome system in motor neurons of a mouse model of familial amyotrophic lateral sclerosis. Hum Mol Genet 18:82–96. https://doi.org/10.1093/hmg/ddn319
Tsuji S, Kikuchi S, Shinpo K et al (2005) Proteasome inhibition induces selective motor neuron death in organotypic slice cultures. J Neurosci Res 82:443–451. https://doi.org/10.1002/jnr.20665
Deng H-X, Chen W, Hong S-T et al (2011) Mutations in UBQLN2 cause dominant X-linked juvenile and adult-ons et al. S and ALS/dementia. Nature 477:211–215. https://doi.org/10.1038/nature10353
Johnson JO, Mandrioli J, Benatar M et al (2010) Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68:857–864. https://doi.org/10.1016/j.neuron.2010.11.036
Gallagher PS, Clowes Candadai SV, Gardner RG (2014) The requirement for Cdc48/p97 in nuclear protein quality control degradation depends on the substrate and correlates with substrate insolubility. J Cell Sci 127:1980–1991. https://doi.org/10.1242/jcs.141838
Christianson JC, Ye Y (2014) Cleaning up in the endoplasmic reticulum: ubiquitin in charge. Nat Struct Mol Biol 21:325–335. https://doi.org/10.1038/nsmb.2793
Stolz A, Wolf DH (2010) Endoplasmic reticulum associated protein degradation: a chaperone assisted journey to hell. Biochim Biophys Acta 1803:694–705. https://doi.org/10.1016/j.bbamcr.2010.02.005
Oyanagi K, Yamazaki M, Takahashi H et al (2008) Spinal anterior horn cells in sporadic amyotrophic lateral sclerosis show ribosomal detachment from, and cisternal distention of the rough endoplasmic reticulum. Neuropathol Appl Neurobiol 34:650–658. https://doi.org/10.1111/j.1365-2990.2008.00941.x
Lautenschlaeger J, Prell T, Grosskreutz J (2012) Endoplasmic reticulum stress and the ER mitochondrial calcium cycle in amyotrophic lateral sclerosis. Amyotroph Lateral Scler Off Publ World Fed Neurol Res Group Mot Neuron Dis 13:166–177. https://doi.org/10.3109/17482968.2011.641569
Rozas P, Bargsted L, Martínez F et al (2017) The ER proteostasis network in ALS: determining the differential motoneuron vulnerability. Neurosci Lett 636:9–15. https://doi.org/10.1016/j.neulet.2016.04.066
Grosskreutz J, Van Den Bosch L, Keller BU (2010) Calcium dysregulation in amyotrophic lateral sclerosis. Cell Calcium 47:165–174. https://doi.org/10.1016/j.ceca.2009.12.002
Soo KY, Halloran M, Sundaramoorthy V et al (2015) Rab1-dependent ER-Golgi transport dysfunction is a common pathogenic mechanism in SOD1, TDP-43 and FUS-associated ALS. Acta Neuropathol (Berl) 130:679–697. https://doi.org/10.1007/s00401-015-1468-2
Liebelt F, Vertegaal ACO (2016) Ubiquitin-dependent and independent roles of SUMO in proteostasis. Am J Physiol Cell Physiol 311:C284–C296. https://doi.org/10.1152/ajpcell.00091.2016
Schimmel J, Larsen KM, Matic I et al (2008) The ubiquitin-proteasome system is a key component of the SUMO-2/3 cycle. Mol Cell Proteomics MCP 7:2107–2122. https://doi.org/10.1074/mcp.M800025-MCP200
Fei E, Jia N, Yan M et al (2006) SUMO-1 modification increases human SOD1 stability and aggregation. Biochem Biophys Res Commun 347:406–412. https://doi.org/10.1016/j.bbrc.2006.06.092
Niikura T, Kita Y, Abe Y (2014) SUMO3 modification accelerates the aggregation of ALS-linked SOD1 mutants. PLoS One 9:e101080. https://doi.org/10.1371/journal.pone.0101080
Dangoumau A, Marouillat S, Burlaud Gaillard J et al (2015) Inhibition of pathogenic mutant SOD1 aggregation in cultured motor neuronal cells by prevention of its SUMOylation on lysine 75. Neurodegener Dis. https://doi.org/10.1159/000439254
Dangoumau A, Veyrat-Durebex C, Blasco H et al (2013) Protein SUMOylation, an emerging pathway in amyotrophic lateral sclerosis. Int J Neurosci 123:366–374. https://doi.org/10.3109/00207454.2012.761984
Oh S-M, Liu Z, Okada M et al (2010) Ebp1 sumoylation, regulated by TLS/FUS E3 ligase, is required for its anti-proliferative activity. Oncogene 29:1017–1030. https://doi.org/10.1038/onc.2009.411
Arndt V, Rogon C, Höhfeld J (2007) To be, or not to be--molecular chaperones in protein degradation. Cell Mol Life Sci CMLS 64:2525–2541. https://doi.org/10.1007/s00018-007-7188-6
Matsumoto G, Stojanovic A, Holmberg CI et al (2005) Structural properties and neuronal toxicity of amyotrophic lateral sclerosis-associated Cu/Zn superoxide dismutase 1 aggregates. J Cell Biol 171:75–85. https://doi.org/10.1083/jcb.200504050
Komatsu M, Waguri S, Chiba T et al (2006) Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441:880–884. https://doi.org/10.1038/nature04723
Rea SL, Majcher V, Searle MS, Layfield R (2014) SQSTM1 mutations—bridging Paget disease of bone and ALS/FTLD. Exp Cell Res 325:27–37. https://doi.org/10.1016/j.yexcr.2014.01.020
Webster CP, Smith EF, Bauer CS et al (2016) The C9orf72 protein interacts with Rab1a and the ULK1 complex to regulate initiation of autophagy. EMBO J 35:1656–1676. https://doi.org/10.15252/embj.201694401
Gal J, Ström A-L, Kwinter DM et al (2009) Sequestosome 1/p62 links familial ALS mutant SOD1 to LC3 via an ubiquitin-independent mechanism. J Neurochem 111:1062–1073. https://doi.org/10.1111/j.1471-4159.2009.06388.x
Wild P, Farhan H, McEwan DG et al (2011) Phosphorylation of the autophagy receptor optineurin restricts Salmonella growth. Science 333:228–233. https://doi.org/10.1126/science.1205405
Freischmidt A, Wieland T, Richter B et al (2015) Haploinsufficiency of TBK1 causes familial ALS and fronto-temporal dementia. Nat Neurosci 18:631–636. https://doi.org/10.1038/nn.4000
Ji CH, Kwon YT (2017) Crosstalk and interplay between the ubiquitin-proteasome system and autophagy. Mol Cells 40:441–449. https://doi.org/10.14348/molcells.2017.0115
Selimovic D, Porzig BBOW, El-Khattouti A et al (2013) Bortezomib/proteasome inhibitor triggers both apoptosis and autophagy-dependent pathways in melanoma cells. Cell Signal 25:308–318. https://doi.org/10.1016/j.cellsig.2012.10.004
Matsumoto G, Wada K, Okuno M et al (2011) Serine 403 phosphorylation of p62/SQSTM1 regulates selective autophagic clearance of ubiquitinated proteins. Mol Cell 44:279–289. https://doi.org/10.1016/j.molcel.2011.07.039
Ferreira JV, Soares AR, Ramalho JS et al (2015) K63 linked ubiquitin chain formation is a signal for HIF1A degradation by chaperone-mediated autophagy. Sci Rep 5:10210. https://doi.org/10.1038/srep10210
Renton AE, Majounie E, Waite A et al (2011) A hexanucleotide repeat expansion in C9ORF72 is the cause of chromosome 9p21-linked ALS-FTD. Neuron 72:257–268. https://doi.org/10.1016/j.neuron.2011.09.010
DeJesus-Hernandez M, Mackenzie IR, Boeve BF et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72:245–256. https://doi.org/10.1016/j.neuron.2011.09.011
Majounie E, Renton AE, Mok K et al (2012) Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol 11:323–330. https://doi.org/10.1016/S1474-4422(12)70043-1
Liu Y, Pattamatta A, Zu T et al (2016) C9orf72 BAC mouse model with motor deficits and neurodegenerative features of ALS/FTD. Neuron 90:521–534. https://doi.org/10.1016/j.neuron.2016.04.005
Schipper LJ, Raaphorst J, Aronica E et al (2016) Prevalence of brain and spinal cord inclusions, including dipeptide repeat proteins, in patients with the C9ORF72 hexanucleotide repeat expansion: a systematic neuropathological review. Neuropathol Appl Neurobiol 42:547–560. https://doi.org/10.1111/nan.12284
Koppers M, Blokhuis AM, Westeneng H-J et al (2015) C9orf72 ablation in mice does not cause motor neuron degeneration or motor deficits. Ann Neurol 78:426–438. https://doi.org/10.1002/ana.24453
Chaineau M, Ioannou MS, McPherson PS (2013) Rab35: GEFs, GAPs and effectors. Traffic Cph Den 14:1109–1117. https://doi.org/10.1111/tra.12096
Amick J, Ferguson SM (2017) C9orf72: at the intersection of lysosome cell biology and neurodegenerative disease. Traffic Cph Den. https://doi.org/10.1111/tra.12477
Zu T, Liu Y, Bañez-Coronel M et al (2013) RAN proteins and RNA foci from antisense transcripts in C9ORF72 ALS and frontotemporal dementia. Proc Natl Acad Sci U S A 110:E4968–E4977. https://doi.org/10.1073/pnas.1315438110
Mori K, Arzberger T, Grässer FA et al (2013) Bidirectional transcripts of the expanded C9orf72 hexanucleotide repeat are translated into aggregating dipeptide repeat proteins. Acta Neuropathol (Berl) 126:881–893. https://doi.org/10.1007/s00401-013-1189-3
Türk M, Haaker G, Winter L et al (2014) C9ORF72-ALS: P62- and ubiquitin-aggregation pathology in skeletal muscle. Muscle Nerve 50:454–455. https://doi.org/10.1002/mus.24283
Serpente M, Fenoglio C, Cioffi SMG et al (2015) Profiling of ubiquitination pathway genes in peripheral cells from patients with frontotemporal dementia due to C9ORF72 and GRN mutations. Int J Mol Sci 16:1385–1394. https://doi.org/10.3390/ijms16011385
Gupta R, Lan M, Mojsilovic-Petrovic J et al (2017) The proline/arginine dipeptide from hexanucleotide repeat expanded C9ORF72 inhibits the proteasome. eNeuro. https://doi.org/10.1523/ENEURO.0249-16.2017
Gomez-Deza J, Lee Y-B, Troakes C et al (2015) Dipeptide repeat protein inclusions are rare in the spinal cord and almost absent from motor neurons in C9ORF72 mutant amyotrophic lateral sclerosis and are unlikely to cause their degeneration. Acta Neuropathol Commun 3:38. https://doi.org/10.1186/s40478-015-0218-y
Zhang Y-J, Jansen-West K, Xu Y-F et al (2014) Aggregation-prone c9FTD/ALS poly(GA) RAN-translated proteins cause neurotoxicity by inducing ER stress. Acta Neuropathol (Berl) 128:505–524. https://doi.org/10.1007/s00401-014-1336-5
Al-Sarraj S, King A, Troakes C et al (2011) p62 positive, TDP-43 negative, neuronal cytoplasmic and intranuclear inclusions in the cerebellum and hippocampus define the pathology of C9orf72-linked FTLD and MND/ALS. Acta Neuropathol (Berl) 122:691–702. https://doi.org/10.1007/s00401-011-0911-2
O’Rourke JG, Bogdanik L, Yáñez A et al (2016) C9orf72 is required for proper macrophage and microglial function in mice. Science 351:1324–1329. https://doi.org/10.1126/science.aaf1064
Webster CP, Smith EF, Grierson AJ, De Vos KJ (2016) C9orf72 plays a central role in Rab GTPase-dependent regulation of autophagy. Small GTPases 1–10. https://doi.org/10.1080/21541248.2016.1240495
Radunovíc A, Leigh PN (1996) Cu/Zn superoxide dismutase gene mutations in amyotrophic lateral sclerosis: correlation between genotype and clinical features. J Neurol Neurosurg Psychiatry 61:565
Reaume AG, Elliott JL, Hoffman EK et al (1996) Motor neurons in Cu/Zn superoxide dismutase-deficient mice develop normally but exhibit enhanced cell death after axonal injury. Nat Genet 13:43–47. https://doi.org/10.1038/ng0596-43
Tu PH, Raju P, Robinson KA et al (1996) Transgenic mice carrying a human mutant superoxide dismutase transgene develop neuronal cytoskeletal pathology resembling human amyotrophic lateral sclerosis lesions. Proc Natl Acad Sci U S A 93:3155–3160
Shibata N, Asayama K, Hirano A, Kobayashi M (1996) Immunohistochemical study on superoxide dismutases in spinal cords from autopsied patients with amyotrophic lateral sclerosis. Dev Neurosci 18:492–498
Bruijn LI, Becher MW, Lee MK et al (1997) ALS-linked SOD1 mutant G85R mediates damage to astrocytes and promotes rapidly progressive disease with SOD1-containing inclusions. Neuron 18:327–338
Dangoumau A, Verschueren A, Hammouche E et al (2014) Novel SOD1 mutation p.V31A identified with a slowly progressive form of amyotrophic lateral sclerosis. Neurobiol Aging 35:266.e1–266.e4. https://doi.org/10.1016/j.neurobiolaging.2013.07.012
Valentine JS, Hart PJ (2003) Misfolded CuZnSOD and amyotrophic lateral sclerosis. Proc Natl Acad Sci U S A 100:3617–3622. https://doi.org/10.1073/pnas.0730423100
McAlary L, Aquilina JA, Yerbury JJ (2016) Susceptibility of mutant SOD1 to form a destabilized monomer predicts cellular aggregation and toxicity but not in vitro aggregation propensity. Front Neurosci 10:499. https://doi.org/10.3389/fnins.2016.00499
Nishitoh H, Kadowaki H, Nagai A et al (2008) ALS-linked mutant SOD1 induces ER stress- and ASK1-dependent motor neuron death by targeting Derlin-1. Genes Dev 22:1451–1464. https://doi.org/10.1101/gad.1640108
Hetz C, Mollereau B (2014) Disturbance of endoplasmic reticulum proteostasis in neurodegenerative diseases. Nat Rev Neurosci 15:233–249. https://doi.org/10.1038/nrn3689
Niwa J-I, Ishigaki S, Hishikawa N et al (2002) Dorfin ubiquitylates mutant SOD1 and prevents mutant SOD1-mediated neurotoxicity. J Biol Chem 277:36793–36798. https://doi.org/10.1074/jbc.M206559200
Sone J, Niwa J, Kawai K et al (2010) Dorfin ameliorates phenotypes in a transgenic mouse model of amyotrophic lateral sclerosis. J Neurosci Res 88:123–135. https://doi.org/10.1002/jnr.22175
Yung C, Sha D, Li L, Chin L-S (2016) Parkin protects against misfolded SOD1 toxicity by promoting its aggresome formation and autophagic clearance. Mol Neurobiol 53:6270–6287. https://doi.org/10.1007/s12035-015-9537-z
Stürner E, Behl C (2017) The role of the multifunctional BAG3 protein in cellular protein quality control and in disease. Front Mol Neurosci 10:177. https://doi.org/10.3389/fnmol.2017.00177
Chhangani D, Endo F, Amanullah A et al (2016) Mahogunin ring finger 1 confers cytoprotection against mutant SOD1 aggresomes and is defective in an ALS mouse model. Neurobiol Dis 86:16–28. https://doi.org/10.1016/j.nbd.2015.11.017
Hadano S, Mitsui S, Pan L et al (2016) Functional links between SQSTM1 and ALS2 in the pathogenesis of ALS: cumulative impact on the protection against mutant SOD1-mediated motor dysfunction in mice. Hum Mol Genet 25:3321–3340. https://doi.org/10.1093/hmg/ddw180
Daoud H, Valdmanis PN, Kabashi E et al (2009) Contribution of TARDBP mutations to sporadic amyotrophic lateral sclerosis. J Med Genet 46:112–114. https://doi.org/10.1136/jmg.2008.062463
Millecamps S, Salachas F, Cazeneuve C et al (2010) SOD1, ANG, VAPB, TARDBP, and FUS mutations in familial amyotrophic lateral sclerosis: genotype-phenotype correlations. J Med Genet 47:554–560. https://doi.org/10.1136/jmg.2010.077180
Scotter EL, Chen H-J, Shaw CE (2015) TDP-43 proteinopathy and ALS: insights into disease mechanisms and therapeutic targets. Neurother J Am Soc Exp Neurother 12:352–363. https://doi.org/10.1007/s13311-015-0338-x
Lin G, Mao D, Bellen HJ (2017) Amyotrophic lateral sclerosis pathogenesis converges on defects in protein homeostasis associated with TDP-43 mislocalization and proteasome-mediated degradation overload. Curr Top Dev Biol 121:111–171. https://doi.org/10.1016/bs.ctdb.2016.07.004
Farrawell NE, Lambert-Smith IA, Warraich ST et al (2015) Distinct partitioning of ALS associated TDP-43, FUS and SOD1 mutants into cellular inclusions. Sci Rep 5:13416. https://doi.org/10.1038/srep13416
Edbauer D, Haass C (2016) An amyloid-like cascade hypothesis for C9orf72 ALS/FTD. Curr Opin Neurobiol 36:99–106. https://doi.org/10.1016/j.conb.2015.10.009
Araki W, Minegishi S, Motoki K et al (2014) Disease-associated mutations of TDP-43 promote turnover of the protein through the proteasomal pathway. Mol Neurobiol 50:1049–1058. https://doi.org/10.1007/s12035-014-8644-6
Hans F, Fiesel FC, Strong JC et al (2014) UBE2E ubiquitin-conjugating enzymes and ubiquitin isopeptidase Y regulate TDP-43 protein ubiquitination. J Biol Chem 289:19164–19179. https://doi.org/10.1074/jbc.M114.561704
Lamark T, Johansen T (2010) Autophagy: links with the proteasome. Curr Opin Cell Biol 22:192–198. https://doi.org/10.1016/j.ceb.2009.11.002
Scotter EL, Vance C, Nishimura AL et al (2014) Differential roles of the ubiquitin proteasome system and autophagy in the clearance of soluble and aggregated TDP-43 species. J Cell Sci 127:1263–1278. https://doi.org/10.1242/jcs.140087
Bose JK, Huang C-C, Shen C-KJ (2011) Regulation of autophagy by neuropathological protein TDP-43. J Biol Chem 286:44441–44448. https://doi.org/10.1074/jbc.M111.237115
Barmada SJ, Serio A, Arjun A et al (2014) Autophagy induction enhances TDP43 turnover and survival in neuronal ALS models. Nat Chem Biol 10:677–685. https://doi.org/10.1038/nchembio.1563
Monahan Z, Shewmaker F, Pandey UB (2016) Stress granules at the intersection of autophagy and ALS. Brain Res 1649:189–200. https://doi.org/10.1016/j.brainres.2016.05.022
Afroz T, Hock E-M, Ernst P et al (2017) Functional and dynamic polymerization of the ALS-linked protein TDP-43 antagonizes its pathologic aggregation. Nat Commun 8:45. https://doi.org/10.1038/s41467-017-00062-0
Kwiatkowski TJ, Bosco DA, Leclerc AL et al (2009) Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323:1205–1208. https://doi.org/10.1126/science.1166066
Deng H, Gao K, Jankovic J (2014) The role of FUS gene variants in neurodegenerative diseases. Nat Rev Neurol 10:337–348. https://doi.org/10.1038/nrneurol.2014.78
Kino Y, Washizu C, Kurosawa M et al (2015) FUS/TLS deficiency causes behavioral and pathological abnormalities distinct from amyotrophic lateral sclerosis. Acta Neuropathol Commun. https://doi.org/10.1186/s40478-015-0202-6
Shang Y, Huang EJ (2016) Mechanisms of FUS mutations in familial amyotrophic lateral sclerosis. Brain Res 1647:65–78. https://doi.org/10.1016/j.brainres.2016.03.036
Vance C, Scotter EL, Nishimura AL et al (2013) ALS mutant FUS disrupts nuclear localization and sequesters wild-type FUS within cytoplasmic stress granules. Hum Mol Genet 22:2676. https://doi.org/10.1093/hmg/ddt117
Daigle JG, Lanson NA, Smith RB et al (2013) RNA-binding ability of FUS regulates neurodegeneration, cytoplasmic mislocalization and incorporation into stress granules associated with FUS carrying ALS-linked mutations. Hum Mol Genet 22:1193–1205. https://doi.org/10.1093/hmg/dds526
Ito D, Suzuki N (2011) Conjoint pathologic cascades mediated by ALS/FTLD-U linked RNA-binding proteins TDP-43 and FUS. Neurology 77:1636–1643. https://doi.org/10.1212/WNL.0b013e3182343365
Sun Z, Diaz Z, Fang X et al (2011) Molecular determinants and genetic modifiers of aggregation and toxicity for the ALS disease protein FUS/TLS. PLoS Biol 9:e1000614. https://doi.org/10.1371/journal.pbio.1000614
Shiihashi G, Ito D, Yagi T et al (2016) Mislocated FUS is sufficient for gain-of-toxic-function amyotrophic lateral sclerosis phenotypes in mice. Brain J Neurol 139:2380–2394. https://doi.org/10.1093/brain/aww161
Ryu H-H, Jun M-H, Min K-J et al (2014) Autophagy regulates amyotrophic lateral sclerosis-linked fused in sarcoma-positive stress granules in neurons. Neurobiol Aging 35:2822–2831. https://doi.org/10.1016/j.neurobiolaging.2014.07.026
Soo KY, Sultana J, King AE et al (2015) ALS-associated mutant FUS inhibits macroautophagy which is restored by overexpression of Rab1. Cell Death Discov 1:15030. https://doi.org/10.1038/cddiscovery.2015.30
Rezaie T, Child A, Hitchings R et al (2002) Adult-onset primary open-angle glaucoma caused by mutations in optineurin. Science 295:1077–1079. https://doi.org/10.1126/science.1066901
Maruyama H, Morino H, Ito H et al (2010) Mutations of optineurin in amyotrophic lateral sclerosis. Nature 465:223–226. https://doi.org/10.1038/nature08971
Del Bo R, Tiloca C, Pensato V et al (2011) Novel optineurin mutations in patients with familial and sporadic amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 82:1239–1243. https://doi.org/10.1136/jnnp.2011.242313
Mao J, Xia Q, Liu C et al (2017) A critical role of Hrd1 in the regulation of optineurin degradation and aggresome formation. Hum Mol Genet. https://doi.org/10.1093/hmg/ddx096
Li F, Xie X, Wang Y et al (2016) Structural insights into the interaction and disease mechanism of neurodegenerative disease-associated optineurin and TBK1 proteins. Nat Commun 7:12708. https://doi.org/10.1038/ncomms12708
Tumbarello DA, Waxse BJ, Arden SD et al (2012) Autophagy receptors link myosin VI to autophagosomes to mediate Tom1-dependent autophagosome maturation and fusion with the lysosome. Nat Cell Biol 14:1024–1035. https://doi.org/10.1038/ncb2589
Ying H, Yue BYJT (2015) Optineurin: The autophagy connection. Exp Eye Res. https://doi.org/10.1016/j.exer.2015.06.029
Shen W-C, Li H-Y, Chen G-C et al (2015) Mutations in the ubiquitin-binding domain of OPTN/optineurin interfere with autophagy-mediated degradation of misfolded proteins by a dominant-negative mechanism. Autophagy 11:685–700. https://doi.org/10.4161/auto.36098
Sundaramoorthy V, Walker AK, Tan V et al (2015) Defects in optineurin- and myosin VI-mediated cellular trafficking in amyotrophic lateral sclerosis. Hum Mol Genet 24:3830–3846. https://doi.org/10.1093/hmg/ddv126
Markovinovic A, Cimbro R, Ljutic T et al (2017) Optineurin in amyotrophic lateral sclerosis: Multifunctional adaptor protein at the crossroads of different neuroprotective mechanisms. Prog Neurobiol 154:1–20. https://doi.org/10.1016/j.pneurobio.2017.04.005
Nakazawa S, Oikawa D, Ishii R et al (2016) Linear ubiquitination is involved in the pathogenesis of optineurin-associated amyotrophic lateral sclerosis. Nat Commun 7:12547. https://doi.org/10.1038/ncomms12547
Le Ber I, Camuzat A, Guerreiro R et al (2013) SQstm1 mutations in french patients with frontotemporal dementia or frontotemporal dementia with amyotrophic lateral sclerosis. JAMA Neurol 70:1403–1410. https://doi.org/10.1001/jamaneurol.2013.3849
Teyssou E, Takeda T, Lebon V et al (2013) Mutations in SQSTM1 encoding p62 in amyotrophic lateral sclerosis: genetics and neuropathology. Acta Neuropathol (Berl) 125:511–522. https://doi.org/10.1007/s00401-013-1090-0
Vadlamudi RK, Joung I, Strominger JL, Shin J (1996) p62, a phosphotyrosine-independent ligand of the SH2 domain of p56lck, belongs to a new class of ubiquitin-binding proteins. J Biol Chem 271:20235–20237. https://doi.org/10.1074/jbc.271.34.20235
Pankiv S, Clausen TH, Lamark T et al (2007) p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J Biol Chem 282:24131–24145. https://doi.org/10.1074/jbc.M702824200
Goode A, Butler K, Long J et al (2016) Defective recognition of LC3B by mutant SQSTM1/p62 implicates impairment of autophagy as a pathogenic mechanism in ALS-FTLD. Autophagy 12:1094–1104. https://doi.org/10.1080/15548627.2016.1170257
Lattante S, de Calbiac H, Le Ber I et al (2015) Sqstm1 knock-down causes a locomotor phenotype ameliorated by rapamycin in a zebrafish model of ALS/FTLD. Hum Mol Genet 24:1682–1690. https://doi.org/10.1093/hmg/ddu580
Harder B, Jiang T, Wu T et al (2015) Molecular mechanisms of Nrf2 regulation and how these influence chemical modulation for disease intervention. Biochem Soc Trans 43:680–686. https://doi.org/10.1042/BST20150020
Katsuragi Y, Ichimura Y, Komatsu M (2015) p62/SQSTM1 functions as a signaling hub and an autophagy adaptor. FEBS J 282:4672–4678. https://doi.org/10.1111/febs.13540
Pan J-A, Sun Y, Jiang Y-P et al (2016) TRIM21 ubiquitylates SQSTM1/p62 and suppresses protein sequestration to regulate redox homeostasis. Mol Cell 61:720–733. https://doi.org/10.1016/j.molcel.2016.02.007
Watts GDJ, Wymer J, Kovach MJ et al (2004) Inclusion body myopathy associated with Paget disease of bone and frontotemporal dementia is caused by mutant valosin-containing protein. Nat Genet 36:377–381. https://doi.org/10.1038/ng1332
Ishigaki S, Hishikawa N, Niwa J et al (2004) Physical and functional interaction between dorfin and valosin-containing protein that are colocalized in ubiquitylated inclusions in neurodegenerative disorders. J Biol Chem 279:51376–51385. https://doi.org/10.1074/jbc.M406683200
Müller JMM, Deinhardt K, Rosewell I et al (2007) Targeted deletion of p97 (VCP/CDC48) in mouse results in early embryonic lethality. Biochem Biophys Res Commun 354:459–465. https://doi.org/10.1016/j.bbrc.2006.12.206
Ludtmann MH, Arber C, Bartolome F et al (2017) Mutations in valosin-containing protein (VCP) decrease ADP/ATP translocation across the mitochondrial membrane and impair energy metabolism in human neurons. J Biol Chem. https://doi.org/10.1074/jbc.M116.762898
Meyer HH, Wang Y, Warren G (2002) Direct binding of ubiquitin conjugates by the mammalian p97 adaptor complexes, p47 and Ufd1-Npl4. EMBO J 21:5645–5652
Buchberger A, Schindelin H, Hänzelmann P (2015) Control of p97 function by cofactor binding. FEBS Lett 589:2578–2589. https://doi.org/10.1016/j.febslet.2015.08.028
Wang T, Xu W, Qin M et al (2016) Pathogenic mutations in the valosin-containing protein/p97(VCP) N-domain inhibit the SUMOylation of VCP and lead to impaired stress response. J Biol Chem. https://doi.org/10.1074/jbc.M116.729343
Li X, Thompson D, Kumar B, DeMartino GN (2014) Molecular and cellular roles of PI31 (PSMF1) protein in regulation of proteasome function. J Biol Chem 289:17392–17405. https://doi.org/10.1074/jbc.M114.561183
Ju J-S, Fuentealba RA, Miller SE et al (2009) Valosin-containing protein (VCP) is required for autophagy and is disrupted in VCP disease. J Cell Biol 187:875–888. https://doi.org/10.1083/jcb.200908115
Tresse E, Salomons FA, Vesa J et al (2010) VCP/p97 is essential for maturation of ubiquitin-containing autophagosomes and this function is impaired by mutations that cause IBMPFD. Autophagy 6:217–227
Papadopoulos C, Kirchner P, Bug M et al (2017) VCP/p97 cooperates with YOD1, UBXD1 and PLAA to drive clearance of ruptured lysosomes by autophagy. EMBO J 36:135–150. https://doi.org/10.15252/embj.201695148
Ko HS, Uehara T, Tsuruma K, Nomura Y (2004) Ubiquilin interacts with ubiquitylated proteins and proteasome through its ubiquitin-associated and ubiquitin-like domains. FEBS Lett 566:110–114. https://doi.org/10.1016/j.febslet.2004.04.031
Hjerpe R, Bett JS, Keuss MJ et al (2016) UBQLN2 mediates autophagy-independent protein aggregate clearance by the proteasome. Cell 166:935–949. https://doi.org/10.1016/j.cell.2016.07.001
Xia Y, Yan LH, Huang B et al (2014) Pathogenic mutation of UBQLN2 impairs its interaction with UBXD8 and disrupts endoplasmic reticulum-associated protein degradation. J Neurochem 129:99–106. https://doi.org/10.1111/jnc.12606
N’Diaye E-N, Kajihara KK, Hsieh I et al (2009) PLIC proteins or ubiquilins regulate autophagy-dependent cell survival during nutrient starvation. EMBO Rep 10:173–179. https://doi.org/10.1038/embor.2008.238
Rothenberg C, Srinivasan D, Mah L et al (2010) Ubiquilin functions in autophagy and is degraded by chaperone-mediated autophagy. Hum Mol Genet 19:3219–3232. https://doi.org/10.1093/hmg/ddq231
Chang L, Monteiro MJ (2015) Defective proteasome delivery of polyubiquitinated proteins by ubiquilin-2 proteins containing ALS mutations. PLoS One 10:e0130162. https://doi.org/10.1371/journal.pone.0130162
Huang B, Wu Q, Zhou H et al (2016) Increased Ubqln2 expression causes neuron death in transgenic rats. J Neurochem 139:285–293. https://doi.org/10.1111/jnc.13748
Le NTT, Chang L, Kovlyagina I et al (2016) Motor neuron disease, TDP-43 pathology, and memory deficits in mice expressing ALS-FTD-linked UBQLN2 mutations. Proc Natl Acad Sci U S A 113:E7580–E7589. https://doi.org/10.1073/pnas.1608432113
Teyssou E, Chartier L, Amador M-D-M et al (2017) Novel UBQLN2 mutations linked to amyotrophic lateral sclerosis and atypical hereditary spastic paraplegia phenotype through defective HSP70-mediated proteolysis. Neurobiol Aging. https://doi.org/10.1016/j.neurobiolaging.2017.06.018
Kanekura K, Nishimoto I, Aiso S, Matsuoka M (2006) Characterization of amyotrophic lateral sclerosis-linked P56S mutation of vesicle-associated membrane protein-associated protein B (VAPB/ALS8). J Biol Chem 281:30223–30233. https://doi.org/10.1074/jbc.M605049200
Teuling E, Ahmed S, Haasdijk E et al (2007) Motor neuron disease-associated mutant vesicle-associated membrane protein-associated protein (VAP) B recruits wild-type VAPs into endoplasmic reticulum-derived tubular aggregates. J Neurosci 27:9801–9815. https://doi.org/10.1523/JNEUROSCI.2661-07.2007
Aliaga L, Lai C, Yu J et al (2013) Amyotrophic lateral sclerosis-related VAPB P56S mutation differentially affects the function and survival of corticospinal and spinal motor neurons. Hum Mol Genet 22:4293–4305. https://doi.org/10.1093/hmg/ddt279
Langou K, Moumen A, Pellegrino C et al (2010) AAV-mediated expression of wild-type and ALS-linked mutant VAPB selectively triggers death of motoneurons through a Ca2+-dependent ER-associated pathway. J Neurochem 114:795–809. https://doi.org/10.1111/j.1471-4159.2010.06806.x
Chen H-J, Anagnostou G, Chai A et al (2010) Characterization of the properties of a novel mutation in VAPB in familial amyotrophic lateral sclerosis. J Biol Chem 285:40266–40281. https://doi.org/10.1074/jbc.M110.161398
Genevini P, Papiani G, Ruggiano A et al (2014) Amyotrophic lateral sclerosis-linked mutant VAPB inclusions do not interfere with protein degradation pathways or intracellular transport in a cultured cell model. PLoS One 9:e113416. https://doi.org/10.1371/journal.pone.0113416
Navone F, Genevini P, Borgese N (2015) Autophagy and neurodegeneration: insights from a cultured cell model of ALS. Cell 4:354–386. https://doi.org/10.3390/cells4030354
Le Ber I, De Septenville A, Millecamps S et al (2015) TBK1 mutation frequencies in French frontotemporal dementia and amyotrophic lateral sclerosis cohorts. Neurobiol Aging 36:3116.e5–3116.e8. https://doi.org/10.1016/j.neurobiolaging.2015.08.009
Cirulli ET, Lasseigne BN, Petrovski S et al (2015) Exome sequencing in amyotrophic lateral sclerosis identifies risk genes and pathways. Science 347:1436–1441. https://doi.org/10.1126/science.aaa3650
Corcia P, Beltran S, Vourc’h P et al (2015) TBK1 gene stresses the major role of autophagy in ALS. Rev Neurol (Paris) 171:747–749. https://doi.org/10.1016/j.neurol.2015.10.004
Ahmad L, Zhang S-Y, Casanova J-L, Sancho-Shimizu V (2016) Human TBK1: a gatekeeper of Neuroinflammation. Trends Mol Med 22:511–527. https://doi.org/10.1016/j.molmed.2016.04.006
Weidberg H, Elazar Z (2011) TBK1 mediates crosstalk between the innate immune response and autophagy. Sci Signal 4:pe39. https://doi.org/10.1126/scisignal.2002355
Matsumoto G, Shimogori T, Hattori N, Nukina N (2015) TBK1 controls autophagosomal engulfment of polyubiquitinated mitochondria through p62/SQSTM1 phosphorylation. Hum Mol Genet 24:4429–4442. https://doi.org/10.1093/hmg/ddv179
Stork B, Alers S, S. A, Wesselborg S (2012) Regulation of autophagy by protein phosphorylation. Protein Phosphorylation Hum Health
Gijselinck I, Van Mossevelde S, van der Zee J et al (2015) Loss of TBK1 is a frequent cause of frontotemporal dementia in a Belgian cohort. Neurology 85:2116–2125. https://doi.org/10.1212/WNL.0000000000002220
Richter B, Sliter DA, Herhaus L et al (2016) Phosphorylation of OPTN by TBK1 enhances its binding to Ub chains and promotes selective autophagy of damaged mitochondria. Proc Natl Acad Sci U S A 113:4039–4044. https://doi.org/10.1073/pnas.1523926113
Moore AS, Holzbaur ELF (2016) Dynamic recruitment and activation of ALS-associated TBK1 with its target optineurin are required for efficient mitophagy. Proc Natl Acad Sci U S A 113:E3349–E3358. https://doi.org/10.1073/pnas.1523810113
Chau T-L, Gioia R, Gatot J-S et al (2008) Are the IKKs and IKK-related kinases TBK1 and IKK-epsilon similarly activated? Trends Biochem Sci 33:171–180. https://doi.org/10.1016/j.tibs.2008.01.002
Chow CY, Landers JE, Bergren SK et al (2009) Deleterious variants of FIG4, a phosphoinositide phosphatase, in patients with ALS. Am J Hum Genet 84:85–88. https://doi.org/10.1016/j.ajhg.2008.12.010
Verdiani S, Origone P, Geroldi A et al (2013) The FIG4 gene does not play a major role in causing ALS in Italian patients. Amyotroph Lateral Scler Front Degener 14:228–229. https://doi.org/10.3109/21678421.2012.760605
Osmanovic A, Rangnau I, Kosfeld A et al (2017) FIG4 variants in central European patients with amyotrophic lateral sclerosis: a whole-exome and targeted sequencing study. Eur J Hum Genet EJHG 25:324–331. https://doi.org/10.1038/ejhg.2016.186
Alonso G, Phan V, Guillemain I et al (2000) Immunocytochemical localization of the sigma(1) receptor in the adult rat central nervous system. Neuroscience 97:155–170
Schröder M, Kaufman RJ (2005) The mammalian unfolded protein response. Annu Rev Biochem 74:739–789. https://doi.org/10.1146/annurev.biochem.73.011303.074134
Alemu EA, Lamark T, Torgersen KM et al (2012) ATG8 family proteins act as scaffolds for assembly of the ULK complex: sequence requirements for LC3-interacting region (LIR) motifs. J Biol Chem 287:39275–39290. https://doi.org/10.1074/jbc.M112.378109
Paschen W, Mengesdorf T (2005) Endoplasmic reticulum stress response and neurodegeneration. Cell Calcium 38:409–415. https://doi.org/10.1016/j.ceca.2005.06.019
Al-Saif A, Al-Mohanna F, Bohlega S (2011) A mutation in sigma-1 receptor causes juvenile amyotrophic lateral sclerosis. Ann Neurol 70:913–919. https://doi.org/10.1002/ana.22534
Prause J, Goswami A, Katona I et al (2013) Altered localization, abnormal modification and loss of function of sigma receptor-1 in amyotrophic lateral sclerosis. Hum Mol Genet 22:1581–1600. https://doi.org/10.1093/hmg/ddt008
Dreser A, Vollrath JT, Sechi A et al (2017) The ALS-linked E102Q mutation in sigma receptor-1 leads to ER stress-mediated defects in protein homeostasis and dysregulation of RNA-binding proteins. Cell Death Differ. https://doi.org/10.1038/cdd.2017.88
Williams KL, Topp S, Yang S et al (2016) CCNF mutations in amyotrophic lateral sclerosis and frontotemporal dementia. Nat Commun. https://doi.org/10.1038/ncomms11253
Bai C, Richman R, Elledge SJ (1994) Human cyclin F. EMBO J 13:6087–6098
D’Angiolella V, Esencay M, Pagano M (2013) A cyclin without cyclin-dependent kinases: cyclin F controls genome stability through ubiquitin-mediated proteolysis. Trends Cell Biol 23:135–140. https://doi.org/10.1016/j.tcb.2012.10.011
Galligan JJ, Petersen DR (2012) The human protein disulfide isomerase gene family. Hum Genomics 6:6. https://doi.org/10.1186/1479-7364-6-6
Gonzalez-Perez P, Woehlbier U, Chian R-J et al (2015) Identification of rare protein disulfide isomerase gene variants in amyotrophic lateral sclerosis patients. Gene 566:158–165. https://doi.org/10.1016/j.gene.2015.04.035
Woehlbier U, Colombo A, Saaranen MJ et al (2016) ALS-linked protein disulfide isomerase variants cause motor dysfunction. EMBO J 35:845–865. https://doi.org/10.15252/embj.201592224
Ciechanover A, Kwon YT (2015) Degradation of misfolded proteins in neurodegenerative diseases: therapeutic targets and strategies. Exp Mol Med 47:e147. https://doi.org/10.1038/emm.2014.117
Crippa V, Sau D, Rusmini P et al (2010) The small heat shock protein B8 (HspB8) promotes autophagic removal of misfolded proteins involved in amyotrophic lateral sclerosis (ALS). Hum Mol Genet 19:3440–3456. https://doi.org/10.1093/hmg/ddq257
Wang I-F, Guo B-S, Liu Y-C et al (2012) Autophagy activators rescue and alleviate pathogenesis of a mouse model with proteinopathies of the TAR DNA-binding protein 43. Proc Natl Acad Sci U S A 109:15024–15029. https://doi.org/10.1073/pnas.1206362109
Wang X, Ma M, Teng J et al (2015) Valproate attenuates 25-kDa C-terminal fragment of TDP-43-induced neuronal toxicity via suppressing endoplasmic reticulum stress and activating autophagy. Int J Biol Sci 11:752–761. https://doi.org/10.7150/ijbs.11880
Feng H-L, Leng Y, Ma C-H et al (2008) Combined lithium and valproate treatment delays disease onset, reduces neurological deficits and prolongs survival in an amyotrophic lateral sclerosis mouse model. Neuroscience 155:567–572. https://doi.org/10.1016/j.neuroscience.2008.06.040
Bose S, Cho J (2017) Targeting chaperones, heat shock factor-1, and unfolded protein response: promising therapeutic approaches for neurodegenerative disorders. Ageing Res Rev 35:155–175. https://doi.org/10.1016/j.arr.2016.09.004
Kieran D, Kalmar B, Dick JRT et al (2004) Treatment with arimoclomol, a coinducer of heat shock proteins, delays disease progression in ALS mice. Nat Med 10:402–405. https://doi.org/10.1038/nm1021
Kalmar B, Lu C-H, Greensmith L (2014) The role of heat shock proteins in amyotrophic lateral sclerosis: the therapeutic potential of arimoclomol. Pharmacol Ther 141:40–54. https://doi.org/10.1016/j.pharmthera.2013.08.003
Yu YJ, Watts RJ (2013) Developing therapeutic antibodies for neurodegenerative disease. Neurother J Am Soc Exp Neurother 10:459–472. https://doi.org/10.1007/s13311-013-0187-4
Valera E, Masliah E (2013) Immunotherapy for neurodegenerative diseases: focus on α-synucleinopathies. Pharmacol Ther 138:311–322. https://doi.org/10.1016/j.pharmthera.2013.01.013
Gros-Louis F, Soucy G, Larivière R, Julien J-P (2010) Intracerebroventricular infusion of monoclonal antibody or its derived Fab fragment against misfolded forms of SOD1 mutant delays mortality in a mouse model of ALS. J Neurochem 113:1188–1199. https://doi.org/10.1111/j.1471-4159.2010.06683.x
Benatar M (2007) Lost in translation: treatment trials in the SOD1 mouse and in human ALS. Neurobiol Dis 26:1–13. https://doi.org/10.1016/j.nbd.2006.12.015
Renton AE, Chiò A, Traynor BJ (2014) State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci 17:17–23. https://doi.org/10.1038/nn.3584
Veyrat-Durebex C, Corcia P, Dangoumau A et al (2014) Advances in cellular models to explore the pathophysiology of amyotrophic lateral sclerosis. Mol Neurobiol 49:966–983. https://doi.org/10.1007/s12035-013-8573-9
Holmes WM, Klaips CL, Serio TR (2014) Defining the limits: protein aggregation and toxicity in vivo. Crit Rev Biochem Mol Biol 49:294–303. https://doi.org/10.3109/10409238.2014.914151
Acknowledgments
This work is supported by the Region Centre Val-de-Loire, the Association ARSLA, and the MAbImprove Laboratoire d’excellence (Labex).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maurel, C., Dangoumau, A., Marouillat, S. et al. Causative Genes in Amyotrophic Lateral Sclerosis and Protein Degradation Pathways: a Link to Neurodegeneration. Mol Neurobiol 55, 6480–6499 (2018). https://doi.org/10.1007/s12035-017-0856-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-017-0856-0