Abstract
Keratin-rich wastes in the form of feathers, hair, nails, and horn are highly available as byproducts of agroindustrial processing. The increased needs for energy conserving and recycling, summed with the huge increase in poultry industry, have strongly stimulated the search for alternatives for the management of recalcitrant keratinous wastes. Keratinases, which are produced by several bacteria that have been often isolated from soils and poultry wastes, show potential use in biotechnological processes involving keratin hydrolysis. Although these isolates are mostly restricted to the genera Streptomyces and Bacillus, the diversity of keratinolytic bacteria is significantly greater. Bacterial keratinases are mostly serine proteases, although increased information about keratinolytic metalloproteases, particularly from Gram-negative bacteria, became available. These enzymes are useful in processes related with the bioconversion of keratin waste into feed and fertilizers. Other promising applications have been associated with keratinolytic enzymes, including enzymatic dehairing for leather and cosmetic industry, detergent uses, and development of biopolymers from keratin fibers. The use of keratinases to enhance drug delivery in some tissues and hydrolysis of prion proteins arise as novel outstanding applications for these enzymes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Proteolytic enzymes constitute one of the most important groups of commercial enzymes. These enzymes have ample utilization in industrial processes, such as the detergent industry, a major consumer of proteases, as well as food and leather industries (Kumar and Takagi 1999; Gupta et al. 2002). Keratinases are a particular class of proteolytic enzymes that display the capability of degrading insoluble keratin substrates. These enzymes are gaining importance in the last years, as several potential applications have been associated with the hydrolysis of keratinous substrates among other applications. A significant amount of fibrous insoluble protein in the form of feathers, hair, nails, horn, and other are available as byproducts of agroindustrial processing (Onifade et al. 1998). These keratin-rich wastes are difficult to degrade as the polypeptide is densely packed and strongly stabilized by several hydrogen bonds and hydrophobic interactions, in addition to several disulfide bonds. Keratin is the insoluble structural protein of feathers and wool and is known for its high stability (Bradbury 1973). The composition and molecular configurations of its constituent amino acids warrant structural rigidity. At least 30 different keratin polypeptides are known, falling into 2 evolutionary families designated type I and type II (Fig. 1). Within each polypeptide chain, the helical rod domain of about 310 amino acids is flanked by a shorter nonhelical head and tail domains, which are thought to have a flexible conformation (Cohlberg 1993). The keratin chain is tightly packed in the α-helix (α-keratin) or β-sheet (β-keratin) into a supercoiled polypeptide chain (Parry and North 1998; Kreplak et al. 2004), resulting in the mechanical stability and resistance to common proteolytic enzymes such as pepsin, trypsin, and papain. In addition, cross-linking of protein chains by cysteine bridges confers high mechanical stability and resistance to proteolytic degradation of keratins.
a Subdomain structure of epidermal keratin chains showing the basic short end regions E1 and E2, the variable glycine/serine-rich regions V1 and V2, and the homologous regions H1 and H2 (Steinert 1993). b Subdomain structure of hard α-keratin chains showing the basic (NB) and acidic (NA) regions of the N-terminal domain. The C-terminal domain of the type I chains is characterized by a repeated proline–cysteine–X motif. The C-terminal domain of type II chains contains a periodic distribution of hydrophobic residues (Parry and North 1998). c Model structure of keratin coiled-coil dimer, 45 nm in length. The hydrophobic amino acids of the two α-helices are meshed together in a regular interlocking pattern (Cohlberg 1993). d Organization of keratin microfibrils, showing the globular head and tail domains (in black). The terminal domains can interact with segments in the rod domain and with other C domain in an antiparallel neighboring molecule (Parry and North 1998)
Despite their elevated resistance, keratins do not accumulate in nature and can be hydrolyzed by some microorganisms. Keratinolytic enzymes are produced by fungi, actinomycetes, and bacteria and have been frequently isolated from soils where keratinous materials are deposited (Kaul and Sumbali 1997; Riffel and Brandelli 2006). Among fungi, keratinases are particularly described among dermathophytes isolated from human and animal injuries. Keratinases produced by Microsporum, Trychophyton, and Doratomyces microsporum are described (Kushwaha 1983; Grzywnowicz et al. 1989; Gradisar et al. 2000) and some of these enzymes are well characterized as they present great medical relevance (Deschamps et al. 2003). Strains of Aspergillus fumigatus and Aspergillus flavus producing keratinases were described (Santos et al. 1996). Among bacteria, keratinolytic activity has been widely documented for strains from the genera Bacillus and Streptomyces (Lin et al. 1999; Kim et al. 2001; Bressolier et al. 1999). Keratinolytic enzymes from bacteria may have important uses in biotechnological processes involving keratin-containing wastes from poultry and leather industries through the development of nonpolluting processes. Insoluble feather keratins can be converted after enzymatic hydrolysis to feedstuffs, fertilizers, glues, and films or used for the production of the rare amino acids serine, cysteine, and proline (Papadopoulos et al. 1986; Onifade et al. 1998; Gupta and Ramnani 2006).
The huge increase of the poultry industry has generated large amounts of feathers as byproduct. The utilization of agroindustrial residues may represent an added value to the industry and meets the increase in conscientious energy conserving and recycling. These facts have stimulated the investigation for alternatives to convert recalcitrant keratinous waste into valuable products. In this regard, although keratin hydrolysis by microbial enzymes has been described earlier (Noval and Nickerson 1959), research on keratinases has significantly increased from the 1990s. Thus, an important amount of information on keratin hydrolysis became available. However, the mechanism of keratin biodegradation by microorganisms is not yet completely elucidated. Although the reduction of cysteine bridges may have a significant influence on keratin degradation (Kunert and Stransky 1988; Kunert 1992; Böckle and Müller 1997), the current investigation has been intensified on proteolytic microorganisms. Comprehensive reviews about keratinases and their applications have been published (Onifade et al. 1998; Gupta and Ramnani 2006; Shih and Wang 2006). This article presents recent advances on keratinolytic proteases from bacterial origin with emphasis on their biochemical properties and discusses on their current and potential applications.
Keratinolytic Bacteria
Several feather-degrading bacteria have been isolated from soils and poultry wastes (Table 1). Although these isolates are mostly confined to the genera Streptomyces and Bacillus, some studies indicate that the diversity of feather-degrading bacteria is significantly greater (Lucas et al. 2003). Among Gram-positive bacteria, novel feather-degrading isolates have been identified as Arthrobacter sp. (Lucas et al. 2003), Microbacterium sp. (Thys et al. 2004), and Kocuria rosea (Bernal et al. 2006a). Most keratinolytic strains can extensively degrade feather keratin within 48 h, as illustrated for Microbacterium sp. kr10 (Fig. 2).
Keratinolytic activity has been already described for several Streptomyces spp., which are associated with the initial reports on keratin hydrolysis by microbial proteases (Noval and Nickerson 1959; Elmayergi and Smith 1971). Several keratinolytic Streptomyces are thermophilic soil-isolated strains that can grow and degrade keratin at temperatures higher than 50 °C (Chitte et al. 1999; Mohamedin 1999). However, mesophilic strains are also described as keratin degraders, like S. pactum DSM 40530 (Böckle et al. 1995) and S. albidoflavus K1-02 (Bressolier et al. 1999). The facility of several Streptomyces to hydrolyze keratin could be associated with the fact that they synthesize broad range proteases such as pronase, commercially produced by S. griseus (Jurasek et al. 1974). In addition, S. pactum can reduce disulfide bonds during growth on feathers (Böckle and Müller 1997), suggesting that this microorganism may produce disulfide reductases to facilitate keratin hydrolysis.
Keratinolytic activity is well established among Bacillus spp. Several strains of B. licheniformis and B. subtilis are described as keratinolytic (Lin et al. 1999; Suh and Lee 2001), and other species such as B. pumilus and B. cereus also produce keratinases (Kim et al. 2001; Werlang and Brandelli 2005). Keratinolytic B. licheniformis strains are often capable of completely disintegrating feathers, and their proteolytic enzymes present a broad range of activity (Lin et al. 1992; Cheng et al. 1995). Three novel keratinolytic Bacillus species isolated from the Amazon basin have been recently characterized by our laboratory (Giongo et al. 2007). These bacteria shared elevated homology with B. subtilis, B. amyloliquefaciens, and B. velesensis, and produced a mixture of proteolytic activities that showed remarkable dehairing activity on bovine pelts. Some thermophilic and alkaliphilic strains of Bacillus have been also described to show keratin-degrading activity, such as Bacillus sp. P-001A (Atalo and Gashe 1993), B. halodurans AH-101 (Takami et al. 1992a, 1999), B. pseudofirmus AL-89 (Gessesse et al. 2003), and B. pseudofirmus FA30-01 (Kojima et al. 2006).
In addition to these few Bacillus strains, keratinolytic activity has been associated with other extremophiles. Keratinases are described for the thermophiles Fervidobacterium pennavorans (Friedrich and Antranikian 1996), Thermoanaerobacter keratinophilus (Riessen and Antranikian 2001), Fervidobacterium islandicum (Nam et al. 2002), and the alkaliphilic strains Nesternkonia sp. AL-20 (Gessesse et al. 2003) and Nocardiopsis sp. TOA-1 (Mitsuiki et al. 2004). Thermophilic bacteria are promising for the hydrolysis of hard insoluble proteins like keratin, as such proteins tend to gain plasticity at elevated temperatures, resulting in more susceptibility to protease attack (Suzuki et al. 2006). Two actinomycete strains, Streptomyces flavis 2BG and Microbispora aerata IMBAS-11A, isolated from Antarctic soils were shown to produce keratinolytic enzymes during growth on wool waste (Gushterova et al. 2005).
Keratinolytic activity has been more recently associated with Gram-negative bacteria (Table 1). Feather-degrading strains of Vibrio sp. (Sangali and Brandelli 2000a), Xanthomonas maltophilia (De Toni et al. 2002), Stenotrophomonas sp. (Yamamura et al. 2002), and Chryseobacterium sp. (Riffel and Brandelli 2002) have been isolated from chicken feathers in decomposition. The investigation on the diversity of keratinolytic bacteria among isolates from the soil environment under temperature climate revealed that strains of Proteobacteria and Cytophaga–Flavobacterium group are predominant (Lucas et al. 2003). This agrees with another study where keratinolytic bacteria from decomposing feathers were mostly Gram-negative (Riffel and Brandelli 2006).
Biochemical Properties of Keratinases
Most of the keratinases reported to date have been found to be serine proteases (Lin et al. 1992; Böckle et al. 1995; Friedrich and Antranikian 1996; Bressolier et al. 1999; Suh and Lee 2001; Nam et al. 2002), and a few metalloproteases have shown keratinolytic activity (Brouta et al. 2001; Allpress et al. 2002; Farag and Hassan 2004). Reports on keratinolytic metalloproteases, mostly associated with Gram-negative bacteria, have been increased. The characteristics of some keratinases are summarized in Table 2.
The keratinase produced by B. licheniformis PWD-1 is well characterized. This enzyme is a serine-type protease (Lin et al. 1992), and its coding gene kerA presents elevated homology with the subtilisin Carlsberg (Lin et al. 1995). The N-terminal sequence of this enzyme is identical to Carlsberg subtilisin (Table 3). The gene kerA, which encodes a B. licheniformis keratinase, is expressed specifically for feather hydrolysis (Lin et al. 1995), therefore, the presence of feather keratin as the sole carbon and nitrogen source in the culture medium may result in the preferential expression of the keratinolytic protease. This gene has been cloned and expressed in heterologous microorganisms such as E. coli and B. subtilis, but the keratinase yields are lower than the parental strain (Wang and Shih 1999; Wang et al. 2003a). However, increased keratinase yield could be achieved by chromosomal integration of multiple copies of the gene kerA in B. licheniformis and B. subtilis (Wang et al. 2004). The gene kerA was also cloned for extracellular expression in Pichia pastoris, resulting in a recombinant enzyme that was glycosylated and active on azokeratin (Porres et al. 2002). Besides the well-characterized keratinase from strain PWD-1, other keratinolytic proteases of B. licheniformis are described. Rozs et al. (2001) identified a novel B. licheniformis strain that produces a keratinase showing unusual catalytic properties. B. licheniformis strain HK-1 produces a keratinolytic protease that was partially inhibited by EDTA, 1,10-phenanthroline (Zn2+-specific chelator) and PMSF, but notably inhibited by Zn2+ (Korkmaz et al. 2004).
Similarly to B. licheniformis, the main proteolytic activity of keratinases from B. subtilis is often associated with serine–protease activity. Zaghloul (1998) cloned the gene aprA from a feather-degrading strain of B. subtilis. The gene aprA showed significant homology with subtilisins, which are typical members of the serine–protease family. However, few studies have described purified keratinases from B. subtilis. A keratinolytic protease was purified from B. subtilis KS-1. This enzyme showed a molecular mass of 25.4 kDa and its N-terminal sequence was similar to that of other serine proteases of B. subtilis (Suh and Lee 2001). Macedo et al. (2005) also described a new keratinase from B. subtilis that presents an identical N-terminal sequence to subtilisin E (Table 3), showing no activity on collagen and an excellent dehairing activity. This property is in contrast to keratinase from B. licheniformis PWD-1 that presents a broad spectrum of activity, including other fibrous proteins such as collagen and elastin (Lin et al. 1992). Keratinolytic enzymes from three novel Bacillus species have been recently characterized by our laboratory. These enzymes were strongly inhibited by phenylmethylsulfonyl fluoride (PMSF) and benzamidine, indicating that they belong to the serine–protease family (Giongo et al. 2007).
Most keratinases are classified as serine-type proteases as determined by specific substrates and inhibitors. S. albidoflavus produces a chymotrypsin-like keratinase that exhibited specificity for aromatic and hydrophobic amino acid residues, as demonstrated by using synthetic peptides (Bressolier et al. 1999). The P1 specificity of the keratinase NAPase of Nocardiopsis sp. was tested with 11 synthetic p-nitroanilide (pNA) substrates. The enzyme had preference for aromatic and hydrophobic residues at the P1 position (Mitsuiki et al. 2004). The feather-degrading Nesterenkonia sp. AL20 produces an alkaline protease that also exhibited higher activity with tetrapeptides with hydrophobic residues located at the P1 site in the order Tyr > Phe > Leu (Bakhtiar et al. 2005). The keratinolytic B. licheniformis K-508 secreted an unusual trypsin-like thiol protease that is strongly active toward benzoyl-Phe-Val-Arg-pNA and is not inhibited by PMSF (Rozs et al. 2001). The keratinolytic serine protease of S. pactum DSM 40530 showed substrate specificity and stereospecificity to pNA derivatives of basic amino acids lysine and arginine (l-enantiomers), but hydrolysis of benzoyl-d-Arg-pNA was not detected (Böckle et al. 1995). The purified keratinase of Kocuria rosea was strongly inhibited by 4-(2-aminoethyl) benzenesulfonyl fluoride, soybean trypsin inhibitor, and chymostatin, indicating that it belongs to the serine protease family (Bernal et al. 2006a).
The primary sequence of fervidolysin, produced by the thermophilic Fervidobacterium pennavorans, was deduced from the coding gene fls (Kluskens et al. 2002). The active-site region of a subtilisin-like serine protease was identified. The deduced primary sequence showed high homology with the subtilisin-like proteases. The unique crystal structure reported to date for a keratin-degrading enzyme is associated with fervidolysin (Kim et al. 2004). The 1.7-A resolution crystal structure showed that fervidolysin is composed of four domains: a catalytic domain, two β-sandwich domains, and the propeptide domain. The architecture of the catalytic domain closely resembles that of a subtilisin, indicated also by the high amino acid sequence conservation. A calcium-binding site was observed within the catalytic domain, which exactly matches that of the subtilisin E-propeptide domain complex (Kim et al. 2004).
More recently, increased information about keratinolytic metalloproteases became available. Allpress et al. (2002) reported a keratinolytic metalloprotease from Lysobacter that was strongly active toward carboxybenzoyl-Phe-pNA. Microbacterium sp. kr10 produces a Zn2+ metalloprotease that effectively hydrolyses feather keratin (Thys and Brandelli 2006). The keratinase produced by the Gram-negative bacterium Chryseobacterium sp. kr6 appears to belong to the metalloprotease type as it was inhibited by EDTA and 1,10-phenanthroline and lacks hydrolysis of the substrate benzoyl-l-Arg-pNA (Riffel et al. 2003a). These properties are in agreement with the described properties for Flavobacterium/Chryseobacterium proteases. The two major proteases secreted by Flavobacterium meningosepticum are zinc metalloendopeptidases, one of them presenting unusual O-glycosylation (Tarentino et al. 1995). A protease was purified from Chryseobacterium indologenes and was inhibited by EDTA and 1,10-phenanthroline, and atomic absorption analysis showed that the enzyme contained Ca2+ and Zn2+ (Venter et al. 1999). Activation by Ca2+ and inhibition by Zn2+ of Gram-negative keratinases (Sangali and Brandelli 2000b; Riffel et al. 2003a) resembles some calpains (Sorimachi et al. 1997) and typical bacterial metalloproteases like thermolysin and extracellular proteases from Pseudomonas species (Auld 1995). Results obtained from protease inhibitors also suggest that Gram-negative bacteria may possess different keratinases from those previously isolated from Gram-positive bacteria (Riffel et al. 2003a).
A keratinolytic metalloprotease was recently purified from culture supernatant of Chryseobacterium sp. kr6. This enzyme belongs to the M14 family of peptidases, also known as the carboxypeptidase A family, being the first enzyme of this family associated with keratinolytic activity and the genus Chryseobacterium (Riffel et al. 2007). This enzyme also display an O-glycosylation site DS* in the peptide 5 (KGSSADS*PNSEEK), that is usually found in proteins secreted by the related specie Chryseobacterium meningosepticum (Plummer et al. 1995). Flavastacin, an extracellular metalloprotease from C. meningosepticum, presents a heptaglycoside linked to the DS* site (Tarentino et al. 1995), which may be associated with protection against autoproteolysis.
Keratinolytic metalloproteases may have great biotechnological promise; acting as secondary keratinases, they may overcome the limited proteolysis on the surface of insoluble keratin particles because of restricted enzyme–substrate interaction (Allpress et al. 2002). In addition, the metalloenzyme nature presents a potential method of enzyme immobilization. Increased stability of immobilized keratinase has been associated with a reduced autolysis (Wang et al. 2003b), which could be also achieved for a metalloenzyme by temporary inactivation by chelating agents during storage.
The diversity of catalytic mechanisms, including both serine and metalloproteases, may have an important effect on the natural environment, where a consortium of microorganisms may hydrolyze native keratin easier than a pure culture (Ichida et al. 2001). In addition, rupture of disulfide bonds must occur to allow an adequate hydrolysis of keratin (Kunert 1992; Böckle and Müller 1997). Keratinases are primarily endo-proteases showing a broad spectrum of activity (Lin et al. 1992; Gradisar et al. 2000; Brandelli 2005), and their ability to degrade native keratin has been associated with the cooperative action of multiple enzymes (Yamamura et al. 2002; Giongo et al. 2007). The initial attack by keratinases and disulfide reductases must allow other less specific proteases to act, resulting in an extensive keratin hydrolysis.
Bioprocessing Keratin-rich Wastes
The use of keratinolytic microorganisms arose as an important alternative for recycling keratinous byproducts, particularly from the poultry and leather industries. The developments of bioprocesses that can convert the huge amounts of such byproducts into added-value products have been investigated (Zaghloul et al. 2004; Bertsch and Coello 2005; Grazziotin et al. 2007). The ability of most keratinases to hydrolyze diverse substrates (Lin et al. 1992; Böckle et al. 1995; Brandelli 2005) indicates the potential of such enzymes for bioconversion of keratin wastes to feed use or other added-value products.
Keratin represents nearly 90% of feather weight, which constitute up to 10% of the total chicken weight. The increased amount of feathers generated by commercial poultry processing may represent a pollutant problem and needs adequate management (Shih 1993). Currently, feathers are converted to feather meal by steam pressure cooking, which require high-energy input. Feather meal has been used on a limited basis as an ingredient in animal feed, as it is deficient in methionine, histidine, and tryptophan (Papadopoulos et al. 1986; Wang and Parsons 1997). The use of keratinase to upgrade the nutritional value of feathers or feather meal has been described (Onifade et al. 1998; Grazziotin et al. 2006). Comparable growth rate was observed between chickens fed with isolated soybean and those fed with feather meal fermented with Streptomyces fradiae plus methionine supplementation (Elmayergi and Smith 1971). The utilization of a B. licheniformis feather-lysate with amino acid supplementation in test diets to fed growing broilers produced an identical weight gain to that of soybean meal (Williams et al. 1991). The use of crude keratinase significantly increased the amino acid digestibility of raw feathers and commercial feather meal (Lee et al. 1991; Odetallah et al. 2003).
The optimization of experimental conditions for keratin hydrolysis by Doratomyces microsporum keratinase was studied using two-order experimental design. An increased quantity of soluble protein was achieved when thioglycolated nails or hooves were treated with keratinase (Vignardet et al. 2001). Protein hydrolysates obtained from submerged cultivation of keratinolytic bacteria on poultry feathers show upgraded nutritional value of feather keratin (Bertsch and Coello 2005; Grazziotin et al. 2006). The use of keratinolytic bacteria for the production of feather hydrolysates has been the subject of some patented processes (Shih and Williams 1990; Burtt and Ichida 1999), and the keratinase from B. licheniformis PWD-1 is commercially produced under the trade name Versazyme.
Production of keratinases using feather meal or raw feathers as the basis for culture media has been described, and several factors such as pH, feather concentration, inoculum, and temperature can influence the resulting enzyme yield (Wang and Shih 1999; Brandelli and Riffel 2005). More recently, the use of statistical optimization by response surface methodology have been described for the production of bacterial keratinases during growth on raw feathers (Ramnani and Gupta 2004; Thys et al. 2006; Bernal et al. 2006b). Factors like temperature and media composition significantly influence the production of keratinases, which could be improved nearly 40-fold after optimization.
Leather production yields significant quantities of organic wastes, a significant portion of which comes from degraded keratin. Biotechnological options are available for handling effluents and proteinaceous solid waste (Thanikaivelan et al. 2004). Biodegradation of waste from the leather industry has been reported, as the ability of Streptomyces and Bacillus to hydrolyze hair and wool keratins has been already described (Mukhopadhyay and Chandra 1990; Takami et al. 1992b; Lal et al. 1996). The bacterium S. fradiae substantially degrade the complex morphological structure of wool, hair, and feather substrates by a combination of mechanical and enzymatic activity, resulting in the release of soluble protein during fermentation (Hood and Healy 1994). Shama and Berwick (1991) described the construction of a rotating frame bioreactor in which wool substrate was almost totally solubilized by S. fradiae.
Currently, the animal feed industry is the main consumer for keratin hydrolysates from agroindustrial byproducts. Recycling of feathers is a subject of great interest for animal nutrition because of its potential as an inexpensive and alternative protein source. Despite the limited nutritional value of keratin, both the digestibility and amino acid balance of feather protein might be improved by microbial fermentation (Williams et al. 1991; Grazziotin et al. 2006).
Keratin-rich wastes have been also considered as soil fertilizers. Several organic materials for animal origin, such as blood meal, hooves, horn, feathers, bones and manure have been evaluated as slow-release nitrogen fertilizers. However, about 70% of nitrogen is released during the first 30 days in field condition, and except for chicken feathers, the total nitrogen release occurs between 6 and 7 weeks (Williams and Nelson 1992). In the case of feather meal, which suffers an extensive thermal treatment, the total nitrogen release also occurs at 6–7 weeks (Hadas and Kautsky 1994). Feathers have about 15% of nitrogen, possessing great potential to be used as slow-release nitrogen fertilizer. The slow release of nitrogen from raw feathers indicates that the soil microorganisms could not easily digest the keratin structure. However, if keratin structure is modified by the rupture of chemical bonds, the rate of mineralization increases. The modification of keratin structure can be achieved by thermal treatments and enzymatic hydrolysis with cleavage of disulfide bridges and peptide bonds (Williams et al. 1990; Kim et al. 2005). An additional alternative is the formation of a Schiff base between the amino groups of keratin with aldehydes (Means and Feeney 1998), resulting in new chemical bonds that contribute to a slower release of nitrogen (Choi and Nelson 1996). In this regard, the use of keratinolytic microorganisms may represent an alternative for the development of nitrogen sources for fertilizer utilization. The enzymatic capability of the feather-degrading bacteria to accelerate the composting of dead chickens or feather waste could be an economical and environmentally safe method of recycling these organic materials into high-nitrogen fertilizers (Ichida et al. 2001).
Emerging Applications for Keratinases
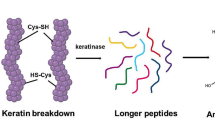
Although biotechnological processes related with the hydrolysis of waste keratin was an early proposition for microbial keratinases, other promising applications have been associated with keratinolytic enzymes. With the advance in the knowledge about keratinases and their action on keratinolysis, a myriad of novel applications have been suggested for these enzymes (Fig. 3).
Enzymatic dehairing is increasingly seen as a reliable alternative to avoid the problem created by sulfide in tanneries (Cantera 2001; Thanikaivelan et al. 2004). The advantages of enzymatic dehairing are a reduction of sulfide content in the effluent, recovery of hair which is of good quality, and elimination of the bate in the deliming. However, this potential benefit remains unfulfilled as enzymes are more expensive than the conventional process chemicals and require careful control (Schraeder et al. 1998). The potential for the commercial use of enzymes in leather production is considerable because of their properties as highly efficient and selective catalysts. The resulting savings in process time may increase the efficiency in leather production, which represents added value to the tanner. A significant feature of the enzymatic dehairing process is the complete hair removal and minimal usage of sulfide and the decomposition products formed from the tannery wastewater with great improvement in wastewater quality as a result. Thus, the substitution of chemical depilatory agents in the leather industry by proteolytic enzymes produced by microorganisms has an important economical and environmental impact. Some microorganisms producing extracellular keratinases showing dehairing activity has been described, and among bacteria, strains of Bacillus are the most studied. Proteolytic strains of Bacillus subtilis and Bacillus amyloliquefaciens have been characterized, presenting desirable properties for leather processing (George et al. 1995; Varela et al. 1997; Macedo et al. 2005; Giongo et al. 2007). The fact that these keratinases can degrade keratin avoiding damage of other structural proteins like collagen, make them exceptional candidates for use in leather industry. Microscopic examination of pelts indicates that they usually hydrolyze the outer sheath rod structure resulting in loose hair (Riffel et al. 2003b), which is easily detached from the skins (Fig. 4).
Some keratinases do not hydrolyze gelatin and synthetic substrate for collagenase, and their use for dehairing bovine pelts cause no collagen damage (Gradisar et al. 2000; Riffel et al. 2003b). These enzymes have attractive characteristics for cosmetic and pharmaceutical purposes where collagen should not be attacked. The use of keratinase for cosmetic application is described as an ingredient in depilatory compositions for shaving (Neena 1993; Slavtcheff et al. 2004). Keratinases may be also useful in topical formulations for the elimination of keratin in acne or psoriasis and removal of human callus (Holland 1993; Vignardet et al. 2001).
Recently, the use of keratinolytic enzymes to enhance drug delivery was investigated. The effectiveness of topical therapy of nail diseases is usually limited by the very low permeability of drugs through the nail plate. The presence of keratinase from Paecilomyces marquandii significantly increases drug permeation through human nail clippings (Mohorcic et al. 2007).
Prion proteins are causative agents of fatal and transmissible neurodegenerative diseases including scrapie, bovine spongiform encephalopathy (mad cow disease), chronic wasting disease, Kuru disease, and Creutzfeldt–Jakob disease (Caughey 2001). The keratinase PWD-1 shows a broad spectrum of activity and also hydrolyses the prion PrPSc protein, the agent responsible for bovine spongiform encephalopathy (Langeveld et al. 2003). Bacillus sp. MSK103 produces a keratinase that exhibited the greatest sequence homology with subtilisin DY and had activity for degrading PrPSc (Yoshioka et al. 2007). Keratinolytic enzymes of selected anaerobic thermophiles were also found to hydrolyze the heat-denatured amyloid prion (Suzuki et al. 2006). The enzymatic hydrolysis of prion proteins constitutes a promising application for bacterial keratinases.
Immobilized proteases often show minimized autolysis and therefore increased stability. However, few attempts to immobilize keratinases are described. The keratinase from B. licheniformis PWD-1 was immobilized on controlled-pore glass beads (Lin et al. 1996). The immobilized keratinase showed improved heat stability and exhibited activity against feather keratin and casein. In another study, a keratinase–streptavidin fusion protein was immobilized onto a biotinylated matrix (Wang et al. 2003b). Heat stability and pH tolerance were greatly improved by immobilization, although the catalytic efficiency was reduced by eightfold. Immobilized keratinases may be useful for developing limited proteolysis processes to modify functional properties of proteins, such as solubility, emulsification, and gelation (Chen et al. 1994).
There is an increased interest in the conversion of keratins into biodegradable films and coatings for both agricultural and biomedical applications. Keratin is chemically or enzymatically treated and the resulting modified keratin is converted into products for compostable packaging, agricultural films, or edible film applications (Yamauchi et al. 1996; Schrooyen et al. 2001; Zaghloul et al. 2004). Polyethylene-based composites were prepared using keratin fibers obtained from chicken feathers (Barone and Schmidt 2005). The keratin fibers can be directly incorporated into the polymer using standard thermomechanical mixing techniques, and keratin feather fiber acts to reinforce the polyethylene polymer matrix. Polymers incorporating feather keratin were shown to have similar properties to many synthetic thermoplastics (Barone and Arikan 2007). One important characteristic of these bio-based polymers is that they biodegrade under natural conditions, reducing significantly the environmental impact of packaging plastics.
Bálint et al. (2005) described a two-stage fermentation system to convert keratin waste into biohydrogen. A Bacillus strain is used to biodegrade keratin-containing materials, converting into fermentation product rich in peptides and amino acids. This mixture could be subsequently used as a nutrient source for the thermophilic anaerobe Thermococcus litoralis, which produces important yields of H2 as a physiological byproduct.
Detergent applications for keratinases have been also suggested (Gupta and Ramnani 2006). These include removal of keratinous dirt that are often encountered in the laundry, such as collars of shirts, and used as additives for cleaning up drains clogged with keratinous waste. Keratinolytic enzymes may have potential application in the woolen textiles industry as a method of shrink proofing wool and to improve wool dyeing (Sousa et al. 2007).
Conclusions
Keratinases are valuable enzymes for bioprocessing keratinous waste. Their ability to degrade the recalcitrant protein keratin constitutes a remarkable property. Increased information on keratinolytic microorganisms and the biochemical properties of their keratinases became available, allowing a better understanding of the biological degradation of keratin. Such amount of information goes toward the development of products and processes linked to proper waste management through recycling keratin-rich agroindustrial byproducts.
References
Allpress, J. D., Mountain, G., & Gowland, P. C. (2002). Production, purification, and characterization of an extracellular keratinase from Lysobacter NCIMB 9497. Letters in Applied Microbiology, 34, 337–342.
Atalo, K., & Gashe, B. A. (1993). Protease production by a thermophilic Bacillus species (P-001A) which degrades various kinds of fibrous proteins. Biotechnology Letters, 15, 1151–1156.
Auld, D. S. (1995). Removal and replacement of metal ions in metallopeptidases. Methods in Enzymology, 248, 228–242.
Bakhtiar, S., Estiveira, R. J., & Hatti-Kaul, R. (2005). Substrate specificity of alkaline protease from alkaliphilic feather-degrading Nesterenkonia sp. AL20. Enzyme and Microbial Technology, 37, 534–540.
Bálint, B., Bagi, Z., Rákhely, G., Perei, K., & Kovács, K. L. (2005). Utilization of keratin-containing biowaste to produce biohydrogen. Applied Microbiology and Biotechnology, 69, 404–410.
Barone, J. R., & Arikan, O. (2007). Composting and biodegradation of thermally processed feather keratin polymer. Polymer Degradation and Stability, 92, 859–867.
Barone, J. R., & Schmidt, W. F. (2005). Polyethylene reinforced with keratin fibers obtained from chicken feathers. Composites Science and Technology, 65, 173–181.
Bernal, C., Cairó, J., & Coello, N. (2006a). Purification and characterization of a novel exocellular keratinase from Kocuria rosea. Enzyme and Microbial Technology, 38, 49–54.
Bernal, C., Diaz, I., & Coello, N. (2006b). Response surface methodology for the optimization of keratinase production in culture medium containing feathers produced by Kocuria rosea. Canadian Journal of Microbiology, 52, 445–450.
Bertsch, A., & Coello, N. (2005). A biotechnological process for treatment and recycling poultry feathers as a feed ingredient. Bioresource Technology, 96, 1703–1708.
Böckle, B., Galunski, B., & Müller, R. (1995). Characterization of a keratinolytic serine protease from Streptomyces pactum DSM40530. Applied and Environmental Microbiology, 61, 3705–3710.
Böckle, B., & Müller, R. (1997). Reduction of disulfide bonds by Streptomyces pactum during growth on chicken feathers. Applied and Environmental Microbiology, 63, 790–792.
Bradbury, J. H. (1973). The structure and chemistry of keratin fibers. Advances in Protein Chemistry, 27, 111–211.
Brandelli, A. (2005). Hydrolysis of native proteins by a keratinolytic strain of Chryseobacterium sp. Annals of Microbiology, 55, 47–50.
Brandelli, A., & Riffel, A. (2005). Production of an extracellular keratinase from Chryseobacterium sp. growing on raw feathers. Electronic Journal of Biotechnology, 8, 35–42.
Bressolier, P., Letourneau, F., Urdaci, M., & Verneuil, B. (1999). Purification and characterization of a keratinolytic serine proteinase from Streptomyces albidoflavus. Applied and Environmental Microbiology, 65, 2570–2576.
Brouta, F., Deschamps, F., Fett, T., Losson, B., & Gerday, C. (2001). Purification and characterization of a 43.5 kDa keratinolytic metalloprotease from Microsporum canis. Medical Mycology, 39, 269–275.
Burtt, E. H., & Ichida, J. M. (1999). Bacteria useful for degrading keratin. US Patent 6214676.
Cantera, C. S. (2001). Hair saving unhairing process. Part 4. Remarks on the evolution of the investigation on enzyme unhairing. Journal of the Society of Leather Technology Chemists, 85, 836–841.
Caughey, B. (2001). Interactions between prion protein isoforms: The kiss of death? Trends in Biochemical Science, 26, 235–242.
Chen, S. X., Swaissgood, H. E., & Foegeding, E., A. (1994). Gelation of β-lactoglobulin treated with limited proteolysis by immobilization trypsin. Journal of Agricultural and Food Chemistry, 42, 234–239.
Cheng, S. W., Hu, H. M., Shen, S. W., Takagi, H., Asano, M., & Tsai, Y. C. (1995). Production and characterization of keratinase of a feather-degrading Bacillus licheniformis PWD-1. Bioscience Biotechnology and Biochemistry, 59, 2239–2243.
Chitte, R. R., Nalawade, V. K., & Dey, S. (1999). Keratinolytic activity from the broth of a feather-degrading thermophilic Streptomyces thermoviolaceus strain SD8. Letters in Applied Microbiology, 28, 131–136.
Choi, J. M., & Nelson, P. V. (1996). Developing a slow release nitrogen fertilizer from organic sources. II. Using poultry feathers. Journal of the American Society of Horticultural Science, 121, 639–643.
Cohlberg, J. A. (1993). The structure of α-keratin. Trends in Biochemical Sciences, 18, 360–362.
De Toni, C. H., Richter, M. F., Chagas, J. R., Henriques, J. A. P., & Termignoni, C. (2002). Purification and characterization of an alkaline serine endopeptidase from a feather-degrading Xanthomonas maltophila strain. Canadian Journal of Microbiology, 48, 342–348.
Deschamps, F., Brouta, F., Vermout, S., Monod, M., Losson, B., & Mignon, B. (2003). Recombinant expression and antigenic properties of a 31.5 kDa keratinolytic subtilisin-like serine protease from Microsporum canis. FEMS Immunology and Medical Microbiology, 38, 29–34.
Elmayergi, H. H., & Smith, R. E. (1971). Influence of growth of Streptomyces fradiae on pepsin-HCl digestibility and methionine content of feather meal. Canadian Journal of Microbiology, 17, 1067–1072.
Farag, A. M., & Hassan, M. A. (2004). Purification, characterization and immobilization of a keratinase from Aspergillus orizae. Enzyme and Microbial Technology, 34, 85–93.
Friedrich, A. B., & Antranikian, G. (1996). Keratin degradation by Fervidobacterium pennavorans, a novel thermophilic anaerobic species of the order Thermatogales. Applied and Environmental Microbiology, 61, 3705–3710.
George, S., Raju, V., Krishnan, M. R. V., Subramanian, T. E., & Jayraman, K. (1995). Production of protease by Bacillus amyloliquefaciens in solid-state fermentation and its application in the unhairing of hides and skins. Process Biochemistry, 30, 457–462.
Gessesse, A., Hatti-Kaul, R., Gashe, B. A., & Mattiasson, B. (2003). Novel alkaline proteases from alkaliphilic bacteria grown on chicken feather. Enzyme and Microbial Technology, 32, 519–524.
Giongo, J. L., Lucas, F. S., Casarin, F., Heeb, P., & Brandelli, A. (2007). Keratinolytic proteases of Bacillus species isolated from the Amazon basin showing remarkable de-hairing activity. World Journal of Microbiology and Biotechnology, 23, 375–382.
Gradisar, H., Friedrich, J., Krizaj, I., & Jerala, R. (2005). Similarities and specificities of fungal keratinolytic proteases: Comparison of keratinases of Paecilomyces marquandii and Doratomyces microsporus to some known proteases. Applied and Environmental Microbiology, 71, 3420–3426.
Gradisar, H., Kern, S., & Friedrich, J. (2000). Keratinase of Doratomyces microsporus. Applied Microbiology and Biotechnology, 53, 196–200.
Grazziotin, A., Pimentel, F. A., de Jong, E. V., & Brandelli, A. (2006). Nutritional improvement of feather protein by treatment with microbial keratinase. Animal Feed Science and Technology, 126, 135–144.
Grazziotin, A., Pimentel, F. A., Sangali, S., de Jong, E. V., & Brandelli, A. (2007). Production of feather protein hydrolysate by keratinolytic bacterium Vibrio sp. kr2. Bioresource Technology, 98, 3172–3175.
Grzywnowicz, G., Lobarzewski, J., Wawrzkiewicz, K., & Wolski, T. (1989). Comparative characterization of proteolytic enzymes from Trichophyton gallinae and Trichophyton verrucosum. Journal of Medical and Veterinary Mycology, 27, 319–328.
Gupta, R., Beg, Q. K., & Lorenz, P. (2002). Bacterial alkaline proteases: Molecular approaches and industrial applications. Applied Microbiology and Biotechnology, 59, 15–32.
Gupta, R., & Ramnani, P. (2006). Microbial keratinases and their prospective applications: An overview. Applied Microbiology and Biotechnology, 70, 21–33.
Gushterova, A., Vasileva-Tonkova, E., Dimova, E., Nedkov, P., & Haertlé, T. (2005). Keratinase production by newly isolated Antarctic actinomycete strains. World Journal of Microbiology and Biotechnology, 21, 831–834.
Hadas, A., & Kautsky, L. (1994). Feather meal, a semi-slow-release nitrogen fertilizer for organic farming. Fertilizer Research, 38, 165–170.
Holland, K. T. (1993). Protease from Micrococcus sedentarius for degrading protein of human callus or corn tissue. US Patent 5213978.
Hood, C. M., & Healy, M. G. (1994). Bioconversion of waste keratins: Wool and feathers. Resources Conservation and Recycling, 11, 179–188.
Huang, Q., Peng, Y., & Li, X. (2003). Purification and characterization of an extracellular alkaline serine protease with dehairing function from Bacillus pumilis. Current Microbiology, 43, 169–173.
Ichida, J. M., Krizova, L., LeFevre, C. A., Keener, H. M., Elwell, D. L., & Burtt, E. H. (2001). Bacterial inoculum enhances keratin degradation and biofilm formation in poultry compost. Journal of Microbiology Methods, 47, 199–208.
Jacobs, M., Elliasson, M., Uhlen, H., & Flock, J. I. (1985). Cloning, sequencing and expression of subtilisin Carlsberg from Bacillus licheniformis. Nucleic Acids Research, 13, 8913–8926.
Jurasek, L., Carpenter, M. R., Smillie, L. B., Getler, A., Levis, S., & Ericson, L. H. (1974). Amino acid sequence of Streptomyces griseus protease B, a major component of pronase. Biochemical and Biophysical Research Communications, 61, 1095–1100.
Kaul, S., & Sumbali, G. (1997). Keratinolysis by poultry farm soil fungi. Mycopathologia, 139, 137–140.
Kim, J. M., Choi, Y. M., & Suh, H. J. (2005). Preparation of feather digests as fertilizer with B. pumilus KHS-1. Journal of Microbiology and Biotechnology, 15, 472–476.
Kim, J. S., Kluskens, L. D., de Vos, W. M., Huber, R., & vand der Oost, J. (2004). Crystal structure of fervidolysin from Fervidobacterium pennivorans, a keratinolytic enzyme related to subtilisin. Journal of Molecular Biology, 335, 787–797.
Kim, J. M., Lim, W. J., & Suh, H. J. (2001). Feather-degrading Bacillus species from poultry waste. Process Biochemistry, 37, 287–291.
Kluskens, L. D., Voorhorst, W. G. B., Siezen, R. J., Schwerdtfeger, R. M., Antranikian, G., van der Oost, J., et al. (2002). Molecular characterization of fervidolysin, a subtilisin-like serine protease from the thermophilic bacterium Fervidobacterium pennivorans. Extremophiles, 6, 185–194.
Kojima, M., Kanai, M., Tominaga, M., Kitazume, S., Inoue, A., & Horikoshi, K. (2006). Isolation and characterization of a feather-degrading enzyme from Bacillus pseudofirmus FA30-01. Extremophiles, 10, 229–235.
Korkmaz, H., Hür, H., & Dinçer, S. (2004). Characterization of alkaline keratinase of Bacillus licheniformis strain HK-1 from poultry waste. Annals of Microbiology, 54, 201–211.
Kreplak, L., Doucet, J., Dumas, P., & Briki, F. (2004). New aspects of the α-helix to β-sheet transition in stretched hard α-keratin fibers. Biophysical Journal, 87, 640–647.
Kumar, C. G., & Takagi, H. (1999). Microbial alkaline proteases: From a bioindustrial viewpoint. Biotechnology Advances, 17, 561–594.
Kunert, J. (1992). Effect of reducing agents on proteolytic and keratinolytic activity of enzymes of Microsporum gypseum. Mycoses, 35, 343–348.
Kunert, J., & Stransky, Z. (1988). Thiosulfate production from cysteine by the keratinophilic prokatyote Streptomyces fradiae. Archives of Microbiology, 150, 600–601.
Kushwaha, R. K. S. (1983). The in vitro degradation of peacock feathers by some fungi. Mykosen, 26, 324–326.
Lal, S., Rajak, R. C., & Hasija, S. K. (1996). Biodegradation of keratin by actinomycetes inhabiting gelatin factory campus at Jablapur: Screening of isolates. Proceedings of the National Academy of Sciences India, 66, 175–180.
Langeveld, J. P. M., Wang, J. J., van de Wiel, D. F. M., Shih, G. C., Garssen, G. J., Bossers, A., et al. (2003). Enzymatic degradation of prion protein in brain stem from infected cattle and sheep. Journal of Infectious Diseases, 188, 1782–1789.
Lee, G. G., Ferket, P. R., & Shih, J. C. H. (1991). Improvement of feather digestibility by bacterial keratinase as a feed additive. FASEB Journal, 59, 1312.
Lin, X., Inglis, G. D., Yanke, L. J., & Cheng, K. J. (1999). Selection and characterization of feather degrading bacteria from canola meal compost. Journal of Industrial Microbiology and Biotechnology, 23, 149–153.
Lin, X., Kelemen, D. W., Miller, E. S., & Shih, J. C. H. (1995). Nucleotide sequence and expression of kerA, the gene encoding a keratinolytic protease of Bacillus licheniformis PWD-1. Applied and Environmental Microbiology, 61, 1469–1474.
Lin, X., Lee, C. G., Casale, E. S., & Shih, J. C. H. (1992). Purification and characterization of a keratinase from a feather-degrading Bacillus licheniformis strain. Applied Environmental Microbiology, 58, 3271–3275.
Lin, X., Shih, J. C. H., & Swaissgood, H. E. (1996). Hydrolysis of feather keratin by immobilized keratinase. Applied and Environmental Microbiology, 62, 4273–4275.
Lucas, F. S., Broennimann, O., Febbraro, I., & Heeb, P. (2003). High diversity among feather-degrading bacteria from a dry meadow soil. Microbial Ecology, 45, 282–290.
Macedo, A. J., Silva, W. O. B., Gava, R., Driemeier, D., Henriques, J. A. P., & Termignoni, C. (2005). Novel keratinase from Bacillus subtilis S14 showing remarkable dehairing capabilities. Applied and Environmental Microbiology, 71, 594–596.
Means, G. E., & Feeney, R. E. (1998). Chemical modifications of proteins: A review. Journal of Food Biochemistry, 22, 399–425.
Mitsuiki, S., Ichikawa, M., Oka, T., Sakai, M., Moriyama, Y., Sameshima, Y., et al. (2004). Molecular characterization of a keratinolytic enzyme from an alkaliphilic Nocardiopsis sp. TOA-1. Enzyme and Microbial Technology, 34, 482–489.
Mohamedin, A. H. (1999). Isolation, identification and some cultural conditions of a protease-producing thermophilic Streptomyces strain grown on chicken feather as substrate. International Biodeterioration and Biodegradation, 43, 13–21.
Mohorcic, M., Torkar, A., Friedrich, J., Kristl, J., & Murdan, S. (2007). An investigation into keratinolytic enzymes to enhance ungual drug delivery. International Journal of Pharmaceutics, 332, 196–201.
Montero-Barrientos, M., Rivas, R., Velazquez, E., Monte, E., & Roig, M. G. (2005). Terrabacter terrae sp. nov., a novel actinomycete isolated from soil in Spain. International Journal of Systematic and Evolutionary Microbiology, 55, 2491–2495.
Mukhopadhyay, R. P., & Chandra, A. L. (1990). Keratinase of a Streptomycete. Indian Journal of Experimental Biology, 28, 575–577.
Nam, G. W., Lee, D. W., Lee, H. S., Lee, N. J., Kim, B. J., Choe, E. A., et al. (2002) Native feather degradation by Fervidobacterium islandicum AW-1, a newly isolating keratinase-producing thermophilic anaerobe. Archives of Microbiology, 178, 538–547.
Neena, K. (1993). Cosmetic treatment of hirsutism. Indian Journal of Dermatology Venerology and Leprology, 59, 109–116.
Noronha, E. F., Lime, B. D., Sá, C. M., & Felix, C. R. (2002). Heterologous production of Aspergillus fumigatus keratinase in Pichia pastoris. World Journal of Microbiology and Biotechnology, 18, 563–568.
Noval, J. J., & Nickerson, W. J. (1959). Decomposition of native keratin by Streptomyces fradiae. Journal of Bacteriology, 77, 251–263.
Odetallah, N. H., Wang, J. J., Garlich, J. D., & Shih, J. C. H. (2003). Keratinase in starter diets improves growth of broiler chicks. Poultry Science, 82, 664–670.
Onifade, A. A., Al-Sane, N. A., Al-Musallam, A. A., & Al-Zarban, S. (1998). Potentials for biotechnological applications of keratin-degrading microorganisms and their enzymes for nutritional improvement of feathers and other keratins as livestock feed resources. Bioresource Technology, 66, 1–11.
Papadopoulos, M. C., El Boushy, A. R., Roodbeen, A. E., & Ketelaars, E. H. (1986). Effects of processing time and moisture content on amino acid composition and nitrogen characteristics of feather meal. Animal Feed Science and Technology, 14, 279–290.
Parry, D. A. D., & North, A. C. T. (1998). Hard α-keratin intermediate filament chains: Substructure of the N- and C-terminal domains and the predicted structure and function of the C-terminal domains of type I and type II chains. Journal of Structural Biology, 122, 67–75.
Plummer, T. H., Tarentino, A. L., & Hauer, C. R. (1995). Novel, specific O-glycosylation of secreted Flavobacerium meningosepticum proteins. Journal of Biological Chemistry, 270, 33192–33196.
Porres, J. M., Benito, M. J., & Lei, X. G. (2002). Functional expression of keratinase (kerA) gene from Bacillus licheniformis in Pichia pastoris. Biotechnology Letters, 24, 631–636.
Ramnani, P., & Gupta, R. (2004). Optimization of medium composition for keratinase production on feather by Bacillus licheniformis RG1 using statistical methods involving response surface methodology. Biotechnology and Applied Biochemistry, 40, 491–496.
Riessen, S., & Antranikian, G. (2001). Isolation of Thermoanaerobacter keratinophilus sp. nov., a novel thermophilic, anaerobic bacterium with keratinolytic activity. Extremophiles, 5, 399–408.
Riffel, A., & Brandelli, A. (2002). Isolation and characterization of a feather-degrading bacterium from the poultry processing industry. Journal of Industrial Microbiology and Biotechnology, 29, 255–258.
Riffel, A., & Brandelli, A. (2006). Keratinolytic bacteria isolated from feather waste. Brazilian Journal of Microbiology, 37, 395–399.
Riffel, A., Brandelli, A., Bellato, C. M., Souza, G. H. M. F., Eberlin, M. N., & Tavares, F. C. A. (2007). Purification and characterization of a keratinolytic metalloprotease from Chryseobacterium sp. kr6. Journal of Biotechnology, 128, 693–703.
Riffel, A., Lucas, F., Heeb, P., & Brandelli, A. (2003a). Characterization of a new keratinolytic bacterium that completely degrades native feather keratin. Archives of Microbiology, 179, 258–265.
Riffel, A., Ortolan, S., & Brandelli, A. (2003b). De-hairing activity of extracellular proteases produced by keratinolytic bacteria. Journal of Chemical Technology and Biotechnology, 78, 855–859.
Rozs, M., Manczinger, L., Vagvolgyi, C., & Kevei, F. (2001). Secretion of a trypsin-like thiol protease from by a new keratinolytic strain of Bacillus licheniformis. FEMS Microbiology Letters, 205, 221–224.
Sangali, S., & Brandelli, A. (2000a). Feather keratin hydrolysis by a Vibrio sp. strain kr2. Journal of Applied Microbiology, 89, 735–743.
Sangali, S., & Brandelli, A. (2000b). Isolation and characterization of a novel feather-degrading bacterial strain. Applied Biochemistry and Biotechnology, 87, 17–24.
Santos, R. M. D., Firmino, A. A. P., Sá, C. M., & Felix, C. R. (1996). Keratinolytic activity of Aspergillus fumigatus Fresenius. Current Microbiology, 33, 364–370.
Schraeder, C. E., Ervin, R. T., & Eberrspacher, J. L. (1998). Economic analysis of the feasibility of using enzymes in the unhairing process. Journal of the American Leather Chemists Association, 93, 265–271.
Schrooyen, P. M. M., Dijkstra, P. J., Oberthur, R. C., Bantjes, A., & Feijen, J. (2001). Partially carboxymethylated feather keratins. 2. Thermal and mechanical properties of films. Journal of Agricultural and Food Chemistry, 49, 221–230.
Shama, G., & Berwick, P. G. (1991). Production of keratinolytic enzymes in a rotating frame bioreactor. Biotechnology Techniques, 5, 359–362.
Shih, J. C. H. (1993). Recent development in poultry waste digestion and feather utilization—A review. Poultry Science, 72, 1617–1620.
Shih, J. C. H., & Wang, J. J. (2006). Keratinase technology: From feather degradation and feed additive, to prion destruction. CAB Reviews: Perspectives in Agriculture, Veterinary Science, Nutrition and Natural Resources, 1(42), 6 pp.
Shih, J. C. H., & Williams, C. M. (1990). Feather-lysate, a hydrolyzed feather feed ingredient and animal feeds containing the same. US Patent 4908220.
Slavtcheff, C. S., Goldberg, J. W., Shiloach, A., Massaro, M., & Kennedy, C. E. (2004). Method and kit for reducing irritation of skin depilatory compositions. US Patent 20040219118.
Sorimachi, H., Ishiura, S., & Suzuki, K. (1997). Structure and physiological function of calpains. Biochemical Journal, 328, 721–732.
Sousa, F., Jus, S., Erbel, A., Kokol, V., Cavaco-Paulo, A., & Gubitz, G. M. (2007). A novel metalloprotease from Bacillus cereus for protein fibre processing. Enzyme and Microbial Technology, 40, 1772–1781.
Stahl, M. L., & Ferrari, E. (1984). Replacement of the Bacillus subtilis subtilisin structural gene with in-vitro derived mutant. Journal of Bacteriology, 158, 411–418.
Steinert, P. M. (1993). Structure, function, and dynamics of keratin intermediate filaments. Journal of Investigative Dermatology, 100, 729–734.
Suh, H. J., & Lee, H. K. (2001). Characterization of a keratinolytic serine protease from Bacillus subtilis KS-1. Journal of Protein Chemistry, 20, 165–169.
Suzuki, Y., Tsujimoto, Y., Matsui, H., & Watanabe, K. (2006). Decomposition of extremely hard-to-degrade animal proteins by thermophilic bacteria. Journal of Bioscience and Bioengineering, 102, 73–81.
Takami, H., Kobayashi, T., Aono, R., & Horikoshi, K. (1992a). Molecular cloning, nucleotide sequence and expression of the structural gene for a thermostable alkaline protease from Bacillus sp. no. AH-101. Applied Microbiology and Biotechnology, 38, 101–108.
Takami, H., Nakamura, F., Aono, R., & Hirishiri, K. (1992b). Degradation of human hair by a thermostable alkaline proteinase from alkalophilic Bacillus sp. no. AH-101. Bioscience Biotechnology and Biochemistry, 56, 1667–1669.
Takami, H., Nogi, Y., & Horikoshi, K. (1999). Reidentification of the keratinase-producing facultatively alkaliphilic Bacillus sp. AH-101 as Bacillus halodurans. Extremophiles, 3, 293–296.
Tarentino, A. L., Quinones, G., Grimwood, B. G., Hauer, C. R., & Plummer, T. H. (1995). Molecular cloning and sequence analysis of flavastascin: An O-glycosylated prokaryotic zinc metalloendopeptidase. Archives of Biochemistry and Biophysisics, 319, 281–285.
Thanikaivelan, P., Rao, J. R., Nair, B. U., & Ramasami, T. (2004). Progress and recent trends in biotechnological methods for leather processing. Trends in Biotechnology, 22, 181–188.
Thys, R. C. S., & Brandelli, A. (2006). Purification and properties of a keratinolytic metalloprotease from Microbacterium sp. Journal of Applied Microbiology, 101, 1259–1268.
Thys, R. C. S., Guzzon, S. O., Cladera-Olivera, F., & Brandelli, A. (2006). Optimization of protease production by Microbacterium sp. on feather meal using response surface methodology. Process Biochemistry, 41, 67–73.
Thys, R. C. S., Lucas, F. S., Riffel, A., Heeb, P., & Brandelli, A. (2004). Characterization of a protease of a feather-degrading Microbacterium species. Letters in Applied Microbiology, 39, 181–186.
Varela, H., Ferrari, M. D., Belobrajdic, L., Vázquez, A., & Loperena, M. L. (1997). Skin unhairing proteases of Bacillus subtilis, production and partial characterization. Biotechnology Letters, 19, 755–758.
Venter, H., Osthoff, G., & Litthauer, D. (1999). Purification and characterization of a metalloprotease from Chryseobacterium indologenes Ix9a and determination of the amino acid specificity with electrospray spectrometry. Protein Expression and Purification, 15, 282–295.
Vignardet, C., Guillaume, Y. C., Michel, L., Friedrich, J., & Millet, J. (2001). Comparison of two hard keratinous substrates submitted to the action of a keratinase using an experimental design. International Journal of Pharmaceutics, 224, 115–122.
Wang, X., & Parsons, C. M. (1997). Effect of processing systems on protein quality of feather meal and hog hair meals. Poultry Science, 76, 491–496.
Wang, J. J., Rojanatavorn, K., & Shih, J. C. H. (2004). Increased production of Bacillus keratinase by chromosomal integration of multiple copies of the kerA gene. Biotechnology and Bioengineering, 87, 459–464.
Wang, J. J., & Shih, J. C. H. (1999). Fermentation production of keratinase from Bacillus licheniformis PWD-1 and a recombinant B. subtilis FDB-29. Journal of Industrial Microbiology and Biotechnology, 22, 608–616.
Wang, J. J., Swaisgood, H. E., & Shih, J. C. H. (2003a). Bioimmobilization of keratinase using Bacillus subtilis and Escherichia coli systems. Biotechnology and Bioengineering, 81, 421–429.
Wang, J. J., Swaisgood, H. E., & Shih, J. C. H. (2003b). Production and characterization of bio-immobilized keratinase in proteolysis and keratinolysis. Enzyme and Microbial Technology, 32, 812–819.
Werlang, P. O., & Brandelli, A. (2005). Characterization of a novel feather-degrading Bacillus sp. strain. Applied Biochemistry and Biotechnology, 120, 71–80.
Williams, C. M., Lee, C. G., Garlich, J. D., & Shih, J. C. H. (1991). Evaluation of a bacterial feather fermentation product, feather-lysate, as a feed protein. Poultry Science, 70, 85–94.
Williams, K. A., & Nelson, P. V. (1992). Low, controlled nutrient availability provided by organic waste materials for Chysanthemum. Journal of the American Society of Horticultural Science, 117, 422–429.
Williams, C. M., Richter, C. S., MacKenzie, J. M., & Shih, J. C. H. (1990). Isolation, identification, and characterization of a feather-degrading bacterium. Applied and Environmental Microbiology, 56, 1509–1515.
Yamamura, S., Morita, Y., Hasan, Q., Yokoyama, K., & Tamiya, E. (2002). Keratin degradation: A cooperative action of two enzymes from Stenotrophomonas sp. Biochemical and Biophysical Research Communications, 294, 1138–1143.
Yamauchi, K., Yamauchi, A., Kusunoki, T., Khoda, A., & Konishi, Y. (1996). Preparation of stable aqueous solutions of keratins, and physicochemical and biodegradational properties of films. Journal of Biomedical Material Research, 31, 439–444.
Yoshioka, M., Miwa, T., Horii, H., Takata, M., Yokoyama, T., Nishizawa, K., et al. (2007). Characterization of a proteolytic enzyme derived from a Bacillus strain that effectively degrades prion protein. Journal of Applied Microbiology, 102, 509–515.
Zaghloul, T. I. (1998). Cloned Bacillus subtilis alkaline protease (aprA) gene showing high level of keratinolytic activity. Applied Biochemistry and Biotechnology, 70/72, 199–205.
Zaghloul, T. I., Haroun, M. A., El-Gayar, K., & Abedalal, A. (2004). Recycling of keratin-containing materials (chicken feather) through genetically engineered bacteria. Polymer–Plastics Technology and Engineering, 43, 1589–1599.
Acknowledgements
The author is a research fellow of Conselho Nacional de Desenvolvimento Cientifico e Tecnologico (CNPq), Brazil.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brandelli, A. Bacterial Keratinases: Useful Enzymes for Bioprocessing Agroindustrial Wastes and Beyond. Food Bioprocess Technol 1, 105–116 (2008). https://doi.org/10.1007/s11947-007-0025-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11947-007-0025-y