Abstract
Purpose of the Review
Osteoarthritis (OA) is a multifactorial and progressive disease affecting whole synovial joint. The extract pathogenic mechanisms and diagnostic biomarkers of OA remain unclear. In this article, we review the studies related to metabolomics of OA, discuss the biomarkers as a tool for early OA diagnosis. Furthermore, we examine the major studies on the application of metabolomics methodology in the complex context of OA and create a bridge from findings in basic science to their clinical utility.
Recent Findings
Recently, the tissue metabolomics signature permits a view into transitional phases between the healthy and OA joint. Both nuclear magnetic resonance spectroscopy (NMR) and mass spectrometry-based metabolomics approaches have been used to interrogate the metabolic alterations that may indicate the complex progression of OA. Specifically, studies on alterations pertaining to lipids, glucose, and amino acid metabolism have aided in the understanding of the complex pathogenesis of OA.
Summary
The discovery of identified metabolites could be important for diagnosis and staging of OA, as well as for the assessment of efficacy of new drugs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Osteoarthritis (OA) represents the most common musculoskeletal disease worldwide characterised by the degradation of articular cartilage, subchondral bone sclerosis, osteophyte formation, and synovial inflammation leading to pain and limitation of joint movement [1] and increasing disability in late stages [2]. OA has a multifactorial etiology, due to an interplay between systemic and local factors. Old age, female gender, high bodyweight index and obesity, injury, repetitive use, low bone density, muscle weakness, and joint laxity all play roles in the development of OA in joints [3,4,5]. Metabolic syndrome components (abdominal obesity, dyslipidaemia, high blood pressure, insulin resistance, and glucose intolerance) are involved in OA pathophysiology, as seen with the connection of leptin and adipokines with OA in the hand [6]. Genetic predisposition can also influence the appearance of OA in a proportion of the population, with implication of genes associated with the thyroid regulatory pathway (e.g., DIO2) [7], the bone morphogenetic protein pathway (e.g., GDF5) [8], apoptotic pathways [9], inflammation-related genes (e.g., PTGS2 and PLA2G4A) [10], and structural genes such as collagen type VI (COL6A4) [10].
At present, there is no effective drug for treating OA or protecting joints from degradation. However, non-steroidal anti-inflammatory drugs (NSAIDs) are considered as an option for managing the pain and inflammation associated with OA. IL-1 beta and anti-TNF alpha are the two major cytokines involved in the physiopathology of OA [11, 12]. Although, there is clear evidence that blockade of IL-1 beta and TNF-alpha reduces the pathophysiology of OA from in vitro [13] and animal model studies [14], there is only limited data available from clinical studies in human OA showing the efficacy of these anti-cytokine treatments [15]. Among surgical options for OA treatment, total joint replacement [16] provides significant to moderate improvement in physical function and pain reduction within the first 6 months of follow-up [17]. However, a disadvantage of joint replacement is increased incidence of failure, predominantly due to infection. In some cases, revision of total knee surgeries in patients younger than 55 years of age is needed. This is because around third of those patients may experience either a post-operative difficulty or early failure [18].
There is a need for OA prevention and/or cure, due to significant effects on the health and life quality of the affected patients and downstream costs to society. Approaches of identifying early disease processes and tracking the response to treatment are lacking. This situation would be improved if we could identify biomarkers of OA before an irreversible stage is reached [19]. Metabolomics offers one approach with the potential to assist in identification of better biomarkers for facilitating diagnosis of disease and assessment of therapy. Metabolomics analysis provides a “snapshot” of chemical components of an organism [20]. This technique has been used to detect the metabolite profiles from bio-fluids and tissues in a number of different medical conditions. It is a technique that can provide information on health, disease, and the response to environmental factors, including changes during OA disease.
Articular Cartilage
Articular cartilage is a weight-bearing connective tissue that covers the diarthrodial joints such as knees. It is hypocellular and it is free from nerves, lymphatics, and blood vessels [21]. It consists of four zones (superficial, middle, deep and calcified zones), classified according to the shape of the chondrocytes and collagen fibril orientation within the extracellular matrix (ECM), and the tidemark which is the transition between non-calcified cartilage zones and calcified cartilage [21, 22]. Cartilage ECM primarily contains water, proteoglycans, collagens, and non-collagenous proteins. Chondrocytes are found in the articular cartilage within the ECM. It comprises 2 to 5 % of the total tissue. They synthesise and/or degrade collagens, proteoglycans and non-collagenous proteins [23]. There are several types of collagen, with type II being the major form, making up about half of the components of hyaline cartilage ECM, 15–25% of the cartilage weight, and 80–90% of the total collagen [21]. The collagen alpha chain has three major amino acids: glycine, proline and hydroxyproline, with a repetitive gly-X-Y arrangement that permits formation of a triple helix from alpha chains to form a collagen molecule [24]. Proteoglycans (PG) are composed of core protein with glycosaminoglycan side chains. The dominant type that constitutes 90% of the total proteoglycans is aggrecan, consisting of a large core protein attached to hyaluronate polymers through link protein and decorated with keratan sulphate and chondroitin sulphate side chains. The proteoglycan network not only provides elasticity and compressive strength of the cartilage but also attracts cations from the fluid [21].
Biomarkers
The term “biomarker” refers to a characteristic that is objectively measured and evaluated as an indicator of normal or pathogenic biological processes or pharmacological responses to a therapeutic intervention [25]. Ideally, a biomarker will assist in recognizing early-stage abnormalities and could be particularly useful in degenerative diseases such as OA. Despite significant research efforts aimed at detecting an OA biomarker, only a few potential candidates have been identified for clinical use. Those identified in blood, urine and synovial fluid of OA patients include markers of inflammation, metabolism and DNA polymorphisms. The majority of these biomarkers arise from the alteration of the metabolism in OA patients [26].
Metabolomics
Metabolomics is one of the “-omics” family of techniques that is an approach to studying biochemical profiles associated with the anatomy, physiology and reactions of low molecular weight metabolites in an organism [27]. It has been used to detect biomarkers for several disorders such as cardiovascular disease [28], cancer [29] and OA [30], as well as to study autophagy [31], drug intake and/or response [32], and the profiles of sugars [33], lipids [34] and amino acids [35]. Although metabolomics is still an emerging research field, this approach is well suited to study the pathophysiology of OA. OA is known to be a heterogeneous disease with multiple phenotypes and involvement of multiple joint tissue interactions. Considering this tremendous heterogeneity, a single biomarker may not reflect the pathological status of OA. Therefore, identification of multiple markers in to a single panel will be more appropriate to avoid the false positives and to determine the disease severity accurately. The application of metabolomics particularly in context of OA is interesting because subtle changes in genes, transcripts or protein levels can mirror the dynamics of metabolites and can be easily detected in bone [36•], cartilage [37] and synovium or body fluids, including urine [38], serum [39], synovial fluid [30] and plasma [40]. Technologies used to examine the metabolic phenotype include nuclear magnetic spectroscopy [37], liquid chromatography mass spectrometry (LCMS) [41], gas chromatography mass spectrometry (GCMS) [42], enzymatic assays [43], ultraviolet-visible spectrophotometry [44], infrared spectroscopy [45] and RAMAN spectroscopy [46]. Of the analytical techniques that were mentioned in metabolomics, NMR and LCMS are the two most commonly used approaches. Both have low operating cost, capable to investigate various sample type and have high ability for metabolite identification [47]. In this review, we will focus on the NMR spectroscopic- and LCMS-based metabolomics.
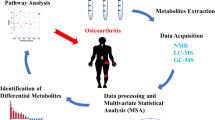
A scheme of the most common metabolomics approaches is shown in Fig. 1. The LCMS technique has two stages: mass spectroscopy (MS) and liquid chromatography (LC). MS needs the ionisation as the first requirement for detecting ions that are released from the sample. It is generally equipped with triple quadrupoles that enable detection of parent ion fragments and is capable of multiple selected reaction monitoring (SRM) in a single run [48]. These two components permit high sensitivity and reliable quantitation of a broad range of targeted metabolites. Retention time plays an integral role in molecular identification and increases the throughput efficiency [49]. Most of the metabolomics methods that use LCMS rely on electrospray ionisation and, therefore, cannot detect non-ionised metabolites such as cholesterol [50]. Reversed-phase (RP) is an effective approach to distinguish imbalances in biological metabolites [49]. An alternative method such as atmospheric pressure chemical ionisation is necessary for detection of non-ionised material [50]. LCMS, which may be the most selective technique for untargeted metabolomics [51], can be used in targeted metabolomics [52].
In particular, LCMS and NMR metabolomics have emerged as the most powerful platform to recognise metabolic anomalies in urine, serum and tissue samples of OA. In particular, by using NMR and GC-MS techniques significant changes in energy metabolism in synovial fluid were revealed in normal vs. OA patient samples [30, 53••, 54, 55••]. Another study has identified the differences between urine NMR spectra of OA cases and controls [56, 57]. Furthermore, metabolomic approaches were employed on OA tissues to explore potential mechanisms of osteophyte formation and have identified phenylalanine metabolism correlative to cartilage degenerative [58•]. The main purpose of chromatography is to decrease ion suppression, which is an alteration of the MS signal due to another co-eluting ion [49]. Identification of metabolites with LCMS relies on the analysis of the retention time and mass of the parent ion derived from the raw data. However, peak mis-annotation remains a problem with LCMS because the retention time can vary between runs and interfering peaks can often elute close to targeted peaks [59].
GCMS is one of the most commonly used techniques for analysis and identification of components of biological samples [60]. It detects low-molecular weight, volatile analytes and uncharged species, which are ionised by electrospray, such as volatile compounds and oils [61]. To obtain broad metabolic profiles, GCMS technique must be combined with a derivatisation procedure. This essential step is used to modify an analyte’s features in order to enable the separations using gas chromatography. Derivatisation aims to enhance the resolution, reduce tailing of polar compounds, reduce the volatility of the compound prior to gas chromatography analysis, increase detectability, improve the analytical efficiency and stabilise the compounds. Derivatisation reactions are classified into three types; alkylation, acylation and silylation. They are used to provide sufficient volatility of organic acids, amino acids, poly-hydroxy compounds, and amides [62]. One potential problem associated with the GCMS is the breakdown of compounds due to the heat during injections, such as the guanidinium group of arginine converting to ornithine [63]. A second issue is the complicated analytical process required for unknown compounds. The third issue is the quantification robustness aimed at some metabolite classes due to the combination of vapour and carrier gas [50].
Both LCMS and GCMS techniques require complicated sample preparation steps that may lead to metabolite loss. Moreover, metabolite profiles could differ from one technique to another based on the ionisation technique used [34, 64]. NMR has been used to detect between 40 and 150 metabolites in biological samples [65•]. NMR is a method that allows the study of atoms within molecules or parts of those molecules within a liquid or intact tissue sample (using high-resolution magic angle spinning (HRMAS), which is explained below) [66]. It is considered a non-destructive method and it is highly appreciated by scientists due to its ability to identify not only known metabolites, but also new metabolites in metabolomics studies, based on analysis of the chemical signal. NMR identifies differences in the metabolite profile that result from a complex sequence of events, beginning with gene translation, based on genotype, through to enzyme interactions that produce the metabolite distribution, resulting in the metabolic phenotype of a sample [67]. NMR is a tool used commonly for studying the metabolic phenotype of articular cartilage [23] and any associated tissue or bio-fluid to identify novel biomarker(s) in OA [60].
NMR spectroscopy (NMR) has several advantages that contribute to providing high throughput metabolite “fingerprints” [50]. It needs minimum sample preparation, has a non-discriminating nature and is non-destructive. It is a reliable, fast and non-invasive technique that can be used in studies in vivo. It is also used for unexpected or unknown compounds [50] (Fig. 1).
NMR Physics Overview
Atoms with asymmetry in the number of protons and/or neutrons result in a magnetic moment that is spinning at a nucleus-specific frequency, thus, making an object behave like a tiny magnet. When an object is placed in an external magnetic field (B0), not only do protons align with or against it, but also they spin with movement called precession around the direction of B0. The rate of precession is proportional to B0. The frequency is calculated using the Larmor equation [68], ω0 = γ B0, where ω0 is the precession frequency, γ is the gyromagnetic ratio (1H has 42.5 MHz/T, which is the highest gyromagnetic ratio among nuclei used in NMR) and B0 is the external magnetic field. Radio frequency (RF) applied at the same frequency of the proton precession results in energy absorption, and when the applied RF is turned off, the resonating nuclei induce a current in a receiver coil that can be measured as an NMR signal. The RF in NMR is most commonly between 60 MHz and 1100 MHz [69, 70].
By nature, the nuclei of atoms in materials spin in a random movement and alignment. Within different metabolites, they are bound to different atoms and are surrounded by different orbital electrons in different shells. This phenomenon produces local magnetic fields that shield and protect each nucleus from the external magnetic field (B0) to different extents. As a consequence, nuclei resonate at diverse frequencies depending on the local environment. This leads to a population of NMR signals [71, 72], which reflect the local environments of the nuclei and provide comprehensive information about the structure of the parent molecule [70]. Several nuclei such as 1H, 13C, 31P and 19F have been used in NMR for this purpose [73]. 1H is the most common nucleus used in NMR due to its high sensitivity compared to other nuclei. High sensitivity is the greatest advantage for application to biological tissues [74].
There are several metabolites that can be identified by using liquid NMR spectroscopy or high-resolution magic angle spinning (HRMAS) NMR spectroscopy for non-liquid samples. Liquid state NMR provides not only well-defined spectral lines but also clear multiplets. This is a result of the interactions between nuclei and electrons. This phenomenon is sometimes observed in solid-state NMR spectroscopy by using HRMAS [75]. Spectra derived from non-liquid state NMR are normally composed of wide lines. In the non-liquid state, NMR spectra are normally composed of wide lines due to chemical shielding because of anisotropic, dipole-dipole coupling and for spin greater than ½, quadrupole coupling. To solve this issue, there are several ways could be applied: magic angle spinning, high-power decoupling, multiple pulse sequences, cross-polarisation and quadrupole echo sequence [72]. HRMAS NMR spectroscopy is a powerful and highly sensitive technique, in which signal-broadening dipolar coupling is minimised in non-solution state NMR spectroscopy [76] and the motion of molecules is restricted [77]. It is characterised by the magic angle (54.74°) spinning (MAS), which is an angle of combined magnetic field resulting from the solenoid coil winding and one dipole winding [78]. HRMAS can be used for detecting and quantifying metabolite profiles in biological tissues for the study of physiological regulation or disease processes [23, 37].
Metabolomics of Serum in OA
It is convenient to use serum samples for OA research [39]. In a comparative study between two groups of mice, the first group consumed a high maize diet of 56% type 2-resistant starch—a diet that resists the bile-acid-enzyme digestion in the small intestine allowing the food to escape to the colon, which provides the bacteria in the colon with additional energy [79]. The second group was fed a high fat diet containing Amioca corn starch; the serum concentrations of branched-chain amino acids (BCAA) were lower in the mice fed high maize diet than the high fat diet group [80]. In another study, the sera of sheep were analysed at 4 weeks and 12 weeks after meniscal destabilisation (MD) or following anterior cruciate ligament transection (ACL) compared to sham-operated. The differences in the metabolite profiles are listed in Table 1 [81].
Metabolomics of Synovial Fluid in OA
Synovial fluid (SF) is considered a key to the study OA because it occupies the space among the tissues in the entire joint [82]. It is a bathing solution that contains metabolites released from and available to the cartilage, as well as from the synovium and other periarticular tissues. The chondrocytes and synoviocytes in the respective tissues synthesise metalloproteases, which play an important role in extracellular matrix remodelling [83]. Metalloproteases (MMPs) are enzymes that play an integral role in extracellular matrix resorption and degradation of cartilage [84]. Although the relative importance of the synovium for MMPs secretion is less clear in OA than in RA, there is an increasing body of evidence that the production of proinflammatory and destructive mediators from the OA synovium are of importance for the symptoms and progression of OA. For example, in a previous study evaluating protein expression profiles in synovial fluid proteins in health and in showed a series of novel proteins differentially expressed in SF from OA patients such as serine protease inhibitors, S100 calcium binding proteins and mitogen-activated protein kinase by using one-dimensional gel electrophoresis and LC-MS/MS, which is contributed to elevation of MMPs and inflammatory mediator expression in OA [85]. A more recent study demonstrated that concordant expression of gelsolin and many complement proteins in synovial tissue and SF suggesting that the synovium are important sources of some of the differentially regulated proteins in OA [86]. Balakrishnan et al. have contributed further evidence of a correlation between synovial fluid proteome and pathophysiology of OA [87]. In this study, 677 proteins from synovial fluid of patients with OA were identified, of these, 545 proteins have not been previously reported, including ADAM-like decysin 1 (ADAMDEC1), alanyl (membrane) aminopeptidase (ANPEP), CD84, fibulin 1 (FBLN1), matrix remodelling associated 5 (MXRA5), secreted phosphoprotein 2 (SPP2) and spondin 2 (SPON2). The proteolytically active metzincin metalloprotease ADAMDEC1 has been found to play a critical role in macrophages-mediated inflammation [88].
Indeed, synovial macrophage has been considered as the key effector for both inflammatory and destructive responses in OA [89]. Infiltration of inflammatory cells especially with macrophages in synovium is the principal characteristic of OA-related synovitis [90,91,92]. Monocyte/macrophages play a major homeostatic role in tissue remodelling, repairing, regeneration and maintenance [93] facilitated by opsonins like CRP, serum amyloid P, C1q, C3b, IgM, ficolin and surfactant proteins [94]. The altered expression of portion of these proteins has been found in OA specimens using proteomic techniques [85] and time-resolved immuno-fluorometric assay [95]. Macrophages have the ability to produce a broad spectrum of matrix metalloproteinase MMPs and may contribute to the erosion of cartilage [84]. In vivo macrophage depletion resulted in decreased production of inflammatory cytokines, growth factors and MMPs which resulted in decreased formation of osteophytes [84]. Moreover, two alarmins of activated macrophages, S100A8 and S100A9 are associated with increased expression and activation of MMPs, thereby inducing cartilage matrix degradation and osteophyte formation in OA [96].
A study using NMR spectroscopy-based metabolomics identified 33 metabolites in SF collected from sheep following ACL and sham surgeries and from non-operated control sheep. Twenty-four metabolites (alanine, lysine, tyrosine, threonine, arginine, phenylalanine, serine, leucine, isoleucine, valine, hypoxanthine, methionine, hydroxyproline, proline, aspartate, asparagine, glutamate 2-hydroxybutyrate, choline, lactate, succinate, formate, tryptophan and acetate) were increased in the samples from anterior cruciate ligament (ACL, experimental OA model) and sham-operated animals. The other metabolites (2-oxovalerate, 2-oxoglutarate, ethanol, glucose, pyruvate and 3-hydroxybutyrate) were decreased compared with the SF from non-operated sheep. Interestingly, glucose and isobutyrate were higher, while serine, hydroxyproline, uridine and asparagine were lower in ACL compared to sham-operated [30].
In an another NMR study, using standard 1D spectroscopy with water pre-saturation at a 600-MHz Bruker, compared SF from OA patients (n = 55) with SF from post mortem samples from subjects without OA (n = 13) and identified 11 metabolites considered as potential biomarkers of OA. The concentrations of methionine, N-phenylacetylglycine, ethanol, creatine, O-acetyl-carnitine and 3-hydroxybutyrate were lower in OA compared to non-OA samples, while citrate concentrations were higher in OA than in non-OA controls [30]. In a different set of human SF samples, saturated triglycerides were detected in SF from 11 patients with traumatic effusions at higher levels than in 10 OA patients [97].
Synovial fluid collected from 28 dogs in which half had been subjected to ACL surgery and the second half were unoperated controls. Alanine, isoleucine, lactate, lipoprotein-associated fatty acid, ketone bodies (3-D-hydroxyisobutyrate and 3-D-hydroxybutyrate), pyruvate, acetate, glycerol and N-acetyl-glycoproteins were increased in ACL-OA SF samples. On the other hand, glucose was the only metabolite that was decreased in ACL-OA compared with unoperated control samples [98]. Another study by the same researchers in seven dogs subjected to ACL and unilateral joint denervation [98] showed that the concentrations of 3-D-hydroxybutyrate, acetate, creatine/creatinine, glutamine/glutamate, N-acetyl-glycoproteins, glycerol lactate, pyruvate citrate, tyrosine and alanine were higher than in sham controls.
Metabolomics of Urine in OA
Metabolomics of urine provide a unique “fingerprint”. It is easy to collect, as it is non-invasive [99], no patient preparation needed [100]. One study examined metabolite variations in human urine samples from knees and hips of both male and female OA patients [98], classified by bilateral radiological assessment of OA using the Kellgren-Lawrence (K-L) grading criteria (K-L) [101]. Because the group represented several K-L grades, samples were separated into two groups based on OA severity, one with K-L grades from 4 to 10 and the control group with K-L grades from 0 to 2 [38]. Hydroxybutyrate, pyruvate, creatine/creatinine and glycerol were detected at higher concentrations in OA, while histidine and methylhistidine were found at lower concentrations in OA, compared to the control group [38]. Another study used NMR spectroscopy of urine from 22 OA patients and 22 control patients [57]. Both groups were exposed to intensive diet and exercise for arthritis (IDEA) [57], a diet and exercise program undertaken for 18 months and designed for arthritis patients. IDEA leads to loss of body weight, reduction of pain [102] and improvement of muscle strength [103] by decreasing the level of plasma pro-inflammatory cytokine IL-6 in arthritis patients [102]. This study described higher levels of hydroxybutyrate, pyruvate, glycerol and an increased creatine/creatinine ratio in OA patients. It also showed lower levels of methylhistidine and histidine in OA patient urine [57].
Metabolomics of Tissue in OA
A study conducted by Shet et al. [37] using 1H HRMAS NMR spectroscopy compared human knee cartilage collected from patients between 57 and 81 years old. Samples were obtained after a total knee replacement for end-stage OA. Control group samples were collected from gender-matched subjects post mortem (n = 5) with an age range of 37–48 years. Differences between the two groups were identified [37]. There was a significant decrease in the concentrations of N-acetyl groups, glycine, alanine, lactate methyne and choline metabolites in OA knees [37]. From another study conducted on the synovial tissues taken from middle to late-stage OA patients, 7 metabolites showed increased concentrations of pro-hydroxyproline, acetylcarntine, myo-inositol, n-acetylornithine, succinate, glutamine and urea compared to control samples from healthy individuals or early OA patients. In contrast, gamma-glutamylleucine, 4-methyl-2-oxopentanoate, 5-oxoproline and phenylacetyglcine decreased in the synovial tissues from OA patients [104].
Metabolite Differences Between OA and Non-OA
In summary, of several types of bio-fluids and tissues that have been examined, all have shown metabolite differences between samples from OA patients and non-OA samples. Both increased and decreased levels and concentrations of metabolites have been observed, as listed in Table 2.
Common Metabolites Detected by Different Metabolomics Approaches
In a study comparing NMR and GCMS to analyse synovial fluid, citrate was increased while 3-hydroxybutyrate was decreased in samples from OA patients compared with control samples [30]. In rabbits subjected to anterior cruciate ligament (ACL) reconstruction after injury, markedly decreased in plasma levels of alanine, threonine and methionine were observed with increasing severity of OA [106]. Analysis of 20 urine samples obtained from OA patients and 17 from non-OA individuals using GCMS-based approach showed significant elevation of histamine, isocitrate, aconitate and citrate while glutamine and histidine were marked decreased, in OA compared to non-OA urine samples [107].
SF were collected from 80 patients undergoing total knee replacement due to OA and examined by targeted LCMS-based metabolomics, which identified a large number of the metabolites. Accordingly, the researchers [108] applied principle component analysis to differentiate the patients and they were able to classify the OA patients in two groups. Higher levels of acylcarnitines were detected in one group, whereas free carnitine levels were lower level, suggesting that there is energy metabolism alteration. [108]. Carnitine plays a role not only in the creation of high energy bonds that result in the acylcarnitine formation but also as transportation of fatty acids in the mitochondria [109]. Accordingly, fatty acids will be oxidised by β-oxidation resulting a major source of energy [108].
Conclusion
The majority of metabolomic profiling approaches in OA reveal changes associated with energy metabolism indicated by perturbation in amino acid metabolism, glycolysis, TCA cycle and ATP biosynthesis. The identified metabolites may be of special clinical relevance for the diagnosis of subtypes of OA, which could lead to a better understanding of, and improvement in, personalised interventions for OA. This approach can potentially serve as a novel tool for metabolic syndrome detection and monitoring, and provide useful information for future interventions targeting obesity-associated OA and also for disease staging. Future investigations are needed with standardised protocols with open access databases for metabolomics to be successful and broadly applied in the clinic setting.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
National Clinical Guideline Centre (UK). Osteoarthritis:Care and Management in Adults. London: National Institute for Health and Care Excellence (UK); 2014 Feb. (NICE Clinical Guidelines, No. 177.) Available from: https://www.ncbi.nlm.nih.gov/books/NBK248069/.
Glyn-Jones S, Palmer AJR, Agricola R, Price AJ, Vincent TL, Weinans H, et al. Osteoarthritis. Lancet. 2015;386(9991):376–87. https://doi.org/10.1016/S0140-6736(14)60802-3.
Messier SP, Gutekunst DJ, Davis C, DeVita P. Weight loss reduces knee-joint loads in overweight and obese older adults with knee osteoarthritis. Arthritis Rheum. 2005;52(7):2026–32. https://doi.org/10.1002/art.21139.
Prieto-Alhambra D, Judge A, Javaid MK, Cooper C, Diez-Perez A, Arden NK. Incidence and risk factors for clinically diagnosed knee, hip and hand osteoarthritis: influences of age, gender and osteoarthritis affecting other joints. Ann Rheum Dis. 2014;73(9):1659–64.
Sturmer T, Gunther KP, Brenner H. Obesity, overweight and patterns of osteoarthritis: the Ulm osteoarthritis study. J Clin Epidemiol. 2000;53(3):307–13.
Yusuf E. Metabolic factors in osteoarthritis: obese people do not walk on their hands. Arthritis Research & Therapy. 2012;14(4):123.
Panicker V. Genetics of thyroid function and disease. Clin Biochem Rev. 2011;32(4):165.
Wang RN, Green J, Wang Z, Deng Y, Qiao M, Peabody M, et al. Bone morphogenetic protein (BMP) signaling in development and human diseases. Genes Dis. 2014;1(1):87–105.
Ramos YF, Bos SD, Lakenberg N, Bohringer S, den Hollander WJ, Kloppenburg M, et al. Genes expressed in blood link osteoarthritis with apoptotic pathways. Ann Rheum Dis. 2014;73(10):1844–53. https://doi.org/10.1136/annrheumdis-2013-203405.
Valdes AM, Spector TD. The genetic epidemiology of osteoarthritis. Curr Opin Rheumatol. 2010;22(2):139–43.
Bondeson J, Wainwright SD, Lauder S, Amos N, Hughes CE. The role of synovial macrophages and macrophage-produced cytokines in driving aggrecanases, matrix metalloproteinases, and other destructive and inflammatory responses in osteoarthritis. Arthritis Res Ther. 2006;8(6):R187. https://doi.org/10.1186/ar2099.
Goldring MB. The role of cytokines as inflammatory mediators in osteoarthritis: lessons from animal models. Connect Tissue Res. 1999;40(1):1–11.
Alaaeddine N, Di Battista JA, Pelletier JP, Kiansa K, Cloutier JM, Martel-Pelletier J. Inhibition of tumor necrosis factor α–induced prostaglandin E2 production by the antiinflammatory cytokines interleukin-4, interleukin-10, and interleukin-13 in osteoarthritic synovial fibroblasts: distinct targeting in the signaling pathways. Arthritis Rheum. 1999;42(4):710–8.
Williams RO, Marinova-Mutafchieva L, Feldmann M, Maini RN. Evaluation of TNF-alpha and IL-1 blockade in collagen-induced arthritis and comparison with combined anti-TNF-alpha/anti-CD4 therapy. J Immunol. 2000;165(12):7240–5.
Zheng W, Chen C, Zhang C, Cai L, Chen H. The protective effect of phloretin in osteoarthritis: an in vitro and in vivo study. Food Funct. 2018;9(1):263–78.
Mont MA, Seyler TM, Ragland PS, Starr R, Erhart J, Bhave A. Gait analysis of patients with resurfacing hip arthroplasty compared with hip osteoarthritis and standard total hip arthroplasty. J Arthroplast. 2007;22(1):100–8.
Bruyère O, Ethgen O, Neuprez A, Zegels B, Gillet P, Huskin J-P, et al. Health-related quality of life after total knee or hip replacement for osteoarthritis: a 7-year prospective study. Arch Orthop Trauma Surg. 2012;132(11):1583–7.
Stambough J, Clohisy J, Barrack R, Nunley R, Keeney J. Increased risk of failure following revision total knee replacement in patients aged 55 years and younger. Bone Joint J. 2014;96(12):1657–62.
Nepple JJ, Thomason KM, An TW, Harris-Hayes M, Clohisy JC. What is the utility of biomarkers for assessing the pathophysiology of hip osteoarthritis? A systematic review. Clin Orthop Relat Res. 2015;473(5):1683–701. https://doi.org/10.1007/s11999-015-4148-6.
Weljie AM, Newton J, Mercier P, Carlson E, Slupsky CM. Targeted profiling: quantitative analysis of 1H NMR metabolomics data. Anal Chem. 2006;78(13):4430–42.
Huber M, Trattnig S, Lintner F. Anatomy, biochemistry, and physiology of articular cartilage. Investig Radiol. 2000;35(10):573–80. https://doi.org/10.1097/00004424-200010000-00003.
Sophia Fox AJ, Bedi A, Rodeo SA. The basic science of articular cartilage: structure, composition, and function. Sport Health. 2009;1(6):461–8.
Borel M, Pastoureau P, Papon J, Madelmont JC, Moins N, Maublant J, et al. Longitudinal profiling of articular cartilage degradation in osteoarthritis by high-resolution magic angle spinning H-1 NMR Spectroscopy: experimental study in the meniscectomized Guinea pig model. J Proteome Res. 2009;8(5):2594–600. https://doi.org/10.1021/pr8009963.
Bhattacharjee A, Bansal M. Collagen structure: the Madras Triple Helix and the current scenario. IUBMB Life. 2005;57(3):161–72. https://doi.org/10.1080/15216540500090710.
Biomarkers Definitions Working Group. Biomarkers and surrogate endpoints: preferred definitions and conceptual framework. Clin Pharmacol Ther. 2001;69(3):89–95. https://doi.org/10.1067/mcp.2001.113989.
Nguyen LT, Sharma AR, Chakraborty C, Saibaba B, Ahn M-E, Lee S-S. Review of prospects of biological fluid biomarkers in osteoarthritis. Int J Mol Sci. 2017;18(3):601.
Deidda M, Piras C, Bassareo PP, Dessalvi CC, Mercuro G. Metabolomics, a promising approach to translational research in cardiology. IJC Metab Endocr. 2015;9:31–8.
Sabatine MS, Liu E, Morrow DA, Heller E, McCarroll R, Wiegand R, et al. Metabolomic identification of novel biomarkers of myocardial ischemia. Circulation. 2005;112(25):3868–75.
Blekherman G, Laubenbacher R, Cortes DF, Mendes P, Torti FM, Akman S, et al. Bioinformatics tools for cancer metabolomics. Metabolomics. 2011;7(3):329–43.
Mickiewicz B, Kelly JJ, Ludwig TE, Weljie AM, Wiley JP, Schmidt TA, et al. Metabolic analysis of knee synovial fluid as a potential diagnostic approach for osteoarthritis. J Orthop Res. 2015;33(11):1631–8. https://doi.org/10.1002/jor.22949.
Stryeck S, Birner-Gruenberger R, Madl T. Integrative metabolomics as emerging tool to study autophagy regulation. Microbial Cell. 2017;4(8):240–58.
Patti GJ, Yanes O, Siuzdak G. Metabolomics: the apogee of the omic triology. Nat Rev Mol Cell Biol. 2012;13(4):263–9.
Kang J, Lee S, Kang S, Kwon HN, Park JH, Kwon SW, et al. NMR-based metabolomics approach for the differentiation of ginseng (Panax ginseng) roots from different origins. Arch Pharm Res. 2008;31(3):330–6.
Dettmer K, Aronov PA, Hammock BD. Mass spectrometry-based metabolomics. Mass Spectrom Rev. 2007;26(1):51–78.
Morris C, O’Grada C, Ryan M, Roche HM, Gibney MJ, Gibney ER, et al. The relationship between BMI and metabolomic profiles: a focus on amino acids. Proc Nutr Soc. 2012;71(4):634–8.
• Marchev AS, Dimitrova PA, Burns AJ, Kostov RV, Dinkova-Kostova AT, Georgiev MI. Oxidative stress and chronic inflammation in osteoarthritis: can NRF2 counteract these partners in crime? Ann N Y Acad Sci. 2017;1401(1):114–35. https://doi.org/10.1111/nyas.13407. This study provides a detailed description of how oxidative stress and inflammation processess are counteracted by transcription factor nuclear factor erythroid p45–related factor 2 (NRF2) during OA progression.
Shet K, Siddiqui SM, Yoshihara H, Kurhanewicz J, Ries M, Li X. High-resolution magic angle spinning NMR spectroscopy of human osteoarthritic cartilage: HR-MAS NMR SPECTROSCOPY OF HUMAN OSTEOARTHRITIC CARTILAGE. NMR Biomed. 2012;25(4):538–44. https://doi.org/10.1002/nbm.1769.
Lamers RJAN, van Nesselrooij JHJ, Kraus VB, Jordan JM, Renner JB, Dragomir AD, et al. Identification of an urinary metabolite profile associated with osteoarthritis. Osteoarthr Cartil. 2005;13(9):762–8. https://doi.org/10.1016/j.joca.2005.04.005.
Beckonert O, Keun HC, Ebbels TM, Bundy J, Holmes E, Lindon JC, et al. Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nat Protoc. 2007;2(11):2692–703. https://doi.org/10.1038/nprot.2007.376.
Jonasdottir HS, Ioan-Facsinay A, Kwekkeboom J, Brouwers H, Zuurmond AM, Toes R, et al. An advanced LC-MS/MS platform for the analysis of specialized pro-resolving lipid mediators. Chromatographia. 2015;78(5–6):391–401. https://doi.org/10.1007/s10337-014-2779-5.
Wilson ID, Plumb R, Granger J, Major H, Williams R, Lenz EM. HPLC-MS-based methods for the study of metabonomics. J Chromatogr B. 2005;817(1):67–76. https://doi.org/10.1016/j.jchromb.2004.07.045.
Halket JM, Waterman D, Przyborowska AM, Patel RK, Fraser PD, Bramley PM. Chemical derivatization and mass spectral libraries in metabolic profiling by GC/MS and LC/MS/MS. J Exp Bot. 2004;56(410):219–43.
Tennessen JM, Barry WE, Cox J, Thummel CS. Methods for studying metabolism in drosophila. Methods. 2014;68(1):105–15.
Moco S, Vervoort J, Bino RJ, De Vos RC, Bino R. Metabolomics technologies and metabolite identification. TrAC Trends Anal Chem. 2007;26(9):855–66.
Yusof NA, Isha A, Ismail IS, Khatib A, Shaari K, Abas F, et al. Infrared-metabolomics approach in detecting changes in Andrographis paniculata metabolites due to different harvesting ages and times. J Sci Food Agric. 2015;95(12):2533–43. https://doi.org/10.1002/jsfa.6987.
Cherney DP, Ekman DR, Dix DJ, Collette TW. Raman spectroscopy-based metabolomics for differentiating exposures to triazole fungicides using rat urine. Anal Chem. 2007;79(19):7324–32. https://doi.org/10.1021/ac070856n.
Roberts MJ, Schirra H, Lavin MF, Gardiner RA. NMR-based metabolomics: global analysis of metabolites to address problems in prostate. cancer. 2014.
Trammell SA, Brenner C. Targeted, LCMS-based metabolomics for quantitative measurement of NAD(+) metabolites. Computational and Structural Biotechnology Journal. 2013;4(5):e201301012. https://doi.org/10.5936/csbj.201301012.
Patti GJ. Separation strategies for untargeted metabolomics. J Sep Sci. 2011;34(24):3460–9.
Lu W, Su X, Klein MS, Lewis IA, Fiehn O, Rabinowitz JD. Metabolite measurement: pitfalls to avoid and practices to follow. Annu Rev Biochem. 2017;86:277–304. https://doi.org/10.1146/annurev-biochem-061516-044952.
Rojo D, Barbas C, Rupérez FJ. LC–MS metabolomics of polar compounds. Bioanalysis. 2012;4(10):1235–43.
Lu W, Bennett BD, Rabinowitz JD. Analytical strategies for LC–MS-based targeted metabolomics. J Chromatogr B. 2008;871(2):236–42.
•• Kim S, Hwang J, Kim J, Ahn JK, Cha HS, Kim KH. Metabolite profiles of synovial fluid change with the radiographic severity of knee osteoarthritis. Joint Bone Spine. 2017;84(5):605–10. https://doi.org/10.1016/j.jbspin.2016.05.018. This study investigates a global metabolite profiles in the synovial fluid of osteoarthritis patients at different disease stages using gas-chromatography/time-of-flight mass spectrometry and revealed that altered metabolites is associated with OA progression.
Blanco FJ, Ruiz-Romero C. Osteoarthritis: metabolomic characterization of metabolic phenotypes in OA. Nat Rev Rheumatol. 2012;8(3):130–2. https://doi.org/10.1038/nrrheum.2012.11.
•• Carlson AK, Rawle RA, Adams E, Greenwood MC, Bothner B, June RK. Application of global metabolomic profiling of synovial fluid for osteoarthritis biomarkers. Biochem Biophys Res Commun. 2018;499(2):182–8. https://doi.org/10.1016/j.bbrc.2018.03.117. This pilot study provides a global metabolic profiles (a total of 1098 metabolites) of OA patient synovial fluid using LC-MS and demonstrates the advantages of global metabolomic profiling for both biomarker discovery and further understanding of OA pathogenesis.
Lamers RJ, van Nesselrooij JH, Kraus VB, Jordan JM, Renner JB, Dragomir AD, et al. Identification of an urinary metabolite profile associated with osteoarthritis. Osteoarthr Cart. 2005;13(9):762–8. https://doi.org/10.1016/j.joca.2005.04.005.
Loeser RF, Pathmasiri W, Sumner SJ, McRitchie S, Beavers D, Saxena P, et al. Association of urinary metabolites with radiographic progression of knee osteoarthritis in overweight and obese adults: an exploratory study. Osteoarthr Cartil. 2016;24(8):1479–86. https://doi.org/10.1016/j.joca.2016.03.011.
• Xu Z, Chen T, Luo J, Ding S, Gao S, Zhang J. Cartilaginous metabolomic study reveals potential mechanisms of osteophyte formation in osteoarthritis. J Proteome Res. 2017;16(4):1425–35. https://doi.org/10.1021/acs.jproteome.6b00676. This study explores that phenylalanine-to-taurine conversion in phenylalanine metabolism might play an important role in the procedure of osteophyte formation and provides a further direction of targeted metabolomic study.
Xu YF, Lu WY, Rabinowitz JD. Avoiding misannotation of in-source fragmentation products as cellular metabolites in liquid chromatography-mass spectrometry-based metabolomics. Anal Chem. 2015;87(4):2273–81. https://doi.org/10.1021/ac504118y.
Sussulini A. Metabolomics: from fundamentals to clinical applications. vol Book, Whole. Cham, Springer; 2017.
Zimmermann D, Hartmann M, Moyer MP, Nolte J, Baumbach JI. Determination of volatile products of human colon cell line metabolism by GC/MS analysis. Metabolomics. 2007;3(1):13–7.
Orata F. Derivatization reactions and reagents for gas chromatography analysis. Advanced gas chromatography-progress in agricultural, biomedical and industrial applications. London: InTech; 2012.
Kaspar H, Dettmer K, Gronwald W, Oefner PJ. Automated GC-MS analysis of free amino acids in biological fluids. J Chromatogr B Anal Technol Biomed Life Sci. 2008;870(2):222–32. https://doi.org/10.1016/j.jchromb.2008.06.018.
Zhang A, Sun H, Wang P, Han Y, Wang X. Modern analytical techniques in metabolomics analysis. Analyst. 2012;137(2):293–300.
• de Sousa EB, Dos Santos GCJ, Duarte MEL, Moura VN, Aguiar DP. Metabolomics as a promising tool for early osteoarthritis diagnosis. Braz J Med Biol Res. 2017;50(11):e6485. https://doi.org/10.1590/1414-431x20176485. This study provides an review of the studies in osteoarthritis animal models and patients regarding the use of metabolomics as a promising tool for early OA diagnosis and detection.
Dale BM, Brown MA, Semelka RC. MRI: basic principles and applications. vol Book, Whole. Hoboken: Wiley Blackwell; 2015.
Kumar D. Nuclear magnetic resonance (NMR) spectroscopy for metabolic profiling of medicinal plants and their products. Crit Rev Anal Chem. 2016;46(5):400–12. https://doi.org/10.1080/10408347.2015.1106932.
Balci M. Basic 1H- and 13C-NMR spectroscopy, vol. Book, Whole. 1st ed. Amsterdam: Elsevier; 2005.
Donald W, McRobbie EAM, Graves MJ, Prince MR. MRI from picture to proton. 2nd ed. Cambridge: Cambridge University Press; 2006.
Günther H. NMR spectroscopy: basic principles, concepts, and applications in chemistry. Third, completely revise and update, vol. Book, Whole. 3rd ed. Weinheim: Wiley-VCH; 2013.
Chatham JC, Blackband SJ. Nuclear magnetic resonance spectroscopy and imaging in animal research. ILAR J. 2001;42(3):189–208.
Duer MJ, Wiley I. Solid-state NMR spectroscopy: principles and applications, vol. Book, Whole. Malden: Blackwell Science; 2002.
Kruk J, Doskocz M, Jodlowska E, Zacharzewska A, Lakomiec J, Czaja K, et al. NMR techniques in metabolomic studies: a quick overview on examples of utilization. Appl Magn Reson. 2017;48(1):1–21. https://doi.org/10.1007/s00723-016-0846-9.
James TL. Fundamentals of NMR. Online Textbook. San Francisco: Department of Pharmaceutical Chemistry, University of California; 1998. p. 1–31.
Bakhmutov VI. NMR spectroscopy in liquids and solids, vol. Book, Whole. Oakville: CRC Press; 2015.
Klinowski J. Magic-angle-spinning NMR. Solid State Ionics. 1985;16:3–14.
Wang Y, Holmes E, Lindon JC, Beckonert O, Keun HC, Ebbels TMD, et al. High-resolution magic-angle-spinning NMR spectroscopy for metabolic profiling of intact tissues. Nat Protoc. 2010;5(6):1019–32. https://doi.org/10.1038/nprot.2010.45.
Li FX, Voccio JP, Sammartino M, Ahn MC, Hahn S, Bascunan J, et al. A theoretical design approach for passive shimming of a magic-angle-spinning NMR magnet. IEEE Trans Appl Supercond. 2016;26(4):1–4. https://doi.org/10.1109/TASC.2015.2512540.
Shortt C, Hasselwander O, Meynier A, Nauta A, Fernandez EN, Putz P, et al. Systematic review of the effects of the intestinal microbiota on selected nutrients and non-nutrients. Eur J Nutr. 2018;57(1):25–49. https://doi.org/10.1007/s00394-017-1546-4.
Barouei J, Bendiks Z, Martinic A, Mishchuk D, Heeney D, Hsieh YH, et al. Microbiota, metabolome, and immune alterations in obese mice fed a high fat diet containing type 2 resistant starch. Mol Nutr Food Res. 2017;61. https://doi.org/10.1002/mnfr.201700184.
Maher AD, Coles C, White J, Bateman JF, Fuller ES, Burkhardt D, et al. 1H NMR spectroscopy of serum reveals unique metabolic fingerprints associated with subtypes of surgically induced osteoarthritis in sheep. J Proteome Res. 2012;11(8):4261–8. https://doi.org/10.1021/pr300368h.
Blanco FJ. Osteoarthritis year in review 2014: we need more biochemical biomarkers in qualification phase. Osteoarthr Cartil. 2014;22(12):2025–32. https://doi.org/10.1016/j.joca.2014.09.009.
Chang C, Werb Z. The many faces of metalloproteases: cell growth, invasion, angiogenesis and metastasis. Trends Cell Biol. 2001;11(11):S37–43.
Blom AB, van Lent PL, Libregts S, Holthuysen AE, van der Kraan PM, van Rooijen N, et al. Crucial role of macrophages in matrix metalloproteinase–mediated cartilage destruction during experimental osteoarthritis: involvement of matrix metalloproteinase 3. Arthritis Rheumatol. 2007;56(1):147–57.
Gobezie R, Kho A, Krastins B, Sarracino DA, Thornhill TS, Chase M, et al. High abundance synovial fluid proteome: distinct profiles in health and osteoarthritis. Arthritis Res Ther. 2007;9(2):R36. https://doi.org/10.1186/ar2172.
Ritter SY, Subbaiah R, Bebek G, Crish J, Scanzello CR, Krastins B, et al. Proteomic analysis of synovial fluid from the osteoarthritic knee: comparison with transcriptome analyses of joint tissues. Arthritis Rheum. 2013;65(4):981–92. https://doi.org/10.1002/art.37823.
Balakrishnan L, Nirujogi RS, Ahmad S, Bhattacharjee M, Manda SS, Renuse S, et al. Proteomic analysis of human osteoarthritis synovial fluid. Clin Proteomics. 2014;11(1):6. https://doi.org/10.1186/1559-0275-11-6.
O'Shea NR, Chew TS, Dunne J, Marnane R, Nedjat-Shokouhi B, Smith PJ, et al. Critical role of the disintegrin metalloprotease ADAM-like Decysin-1 [ADAMDEC1] for intestinal immunity and inflammation. J Crohns Colitis. 2016;10(12):1417–27. https://doi.org/10.1093/ecco-jcc/jjw111.
Manferdini C, Paolella F, Gabusi E, Silvestri Y, Gambari L, Cattini L, et al. From osteoarthritic synovium to synovial-derived cells characterization: synovial macrophages are key effector cells. Arthritis Res Ther. 2016;18(1):83.
Sellam J, Berenbaum F. The role of synovitis in pathophysiology and clinical symptoms of osteoarthritis. Nat Rev Rheumatol. 2010;6(11):625–35.
Benito MJ, Veale DJ, FitzGerald O, van den Berg WB, Bresnihan B. Synovial tissue inflammation in early and late osteoarthritis. Ann Rheum Dis. 2005;64(9):1263–7. https://doi.org/10.1136/ard.2004.025270.
van Lent PL, Blom AB, Schelbergen RF, Sloetjes A, Lafeber FP, Lems WF, et al. Active involvement of alarmins S100A8 and S100A9 in the regulation of synovial activation and joint destruction during mouse and human osteoarthritis. Arthritis Rheum. 2012;64(5):1466–76. https://doi.org/10.1002/art.34315.
Epelman S, Lavine KJ, Randolph GJ. Origin and functions of tissue macrophages. Immunity. 2014;41(1):21–35.
Konttinen YT, Pajarinen J, Takakubo Y, Gallo J, Nich C, Takagi M, et al. Macrophage polarization and activation in response to implant debris: influence by “particle disease” and “ion disease”. J Long-Term Eff Med Implants. 2014;24(4):267–81.
Ammitzboll CG, Thiel S, Ellingsen T, Deleuran B, Jorgensen A, Jensenius JC, et al. Levels of lectin pathway proteins in plasma and synovial fluid of rheumatoid arthritis and osteoarthritis. Rheumatol Int. 2012;32(5):1457–63. https://doi.org/10.1007/s00296-011-1879-x.
Schelbergen RF, de Munter W, van den Bosch MH, Lafeber FP, Sloetjes A, Vogl T, et al. Alarmins S100A8/S100A9 aggravate osteophyte formation in experimental osteoarthritis and predict osteophyte progression in early human symptomatic osteoarthritis. Ann Rheum Dis. 2016;75(1):218–25. https://doi.org/10.1136/annrheumdis-2014-205480.
Williamson MP, Humm G, Crisp AJ. 1H nuclear magnetic resonance investigation of synovial fluid components in osteoarthritis, rheumatoid arthritis and traumatic effusions. Br J Rheumatol. 1989;28(1):23–7.
Damyanovich AZ, Staples JR, Chan ADM, Marshall KW. Comparative study of normal and osteoarthritic canine synovial fluid using 500 MHz H-1 magnetic resonance spectroscopy. J Orthop Res. 1999;17(2):223–31.
Ryan D, Robards K, Prenzler PD, Kendall M. Recent and potential developments in the analysis of urine: a review. Anal Chim Acta. 2011;684(1):8–20. https://doi.org/10.1016/j.aca.2010.10.035.
Zhang A, Sun H, Wu X, Wang X. Urine metabolomics. Clin Chim Acta. 2012;414:65–9. https://doi.org/10.1016/j.cca.2012.08.016.
Kellgren JH, Lawrence JS. Radiological assessment of osteo-arthrosis. Ann Rheum Dis. 1957;16(4):494–502.
Messier SP, Mihalko SL, Legault C, Miller GD, Nicklas BJ, DeVita P, et al. Effects of intensive diet and exercise on knee joint loads, inflammation, and clinical outcomes among overweight and obese adults with knee osteoarthritis: the IDEA randomized clinical trial. JAMA. 2013;310(12):1263–73. https://doi.org/10.1001/jama.2013.277669.
Krishnasamy P, Hall M, Robbins SR. The role of skeletal muscle in the pathophysiology and management of knee osteoarthritis. Rheumatology (Oxford, England). 2018;57:iv22–33. https://doi.org/10.1093/rheumatology/kex515.
Priori R, Scrivo R, Brandt J, Valerio M, Casadei L, Valesini G, et al. Metabolomics in rheumatic diseases: the potential of an emerging methodology for improved patient diagnosis, prognosis, and treatment efficacy. Autoimmun Rev. 2013;12(10):1022–30. https://doi.org/10.1016/j.autrev.2013.04.002.
Damyanovich AZ, Staples JR, Marshall KW. 1H NMR investigation of changes in the metabolic profile of synovial fluid in bilateral canine osteoarthritis with unilateral joint denervation. Osteoarthr Cartil. 1999;7(2):165–72. https://doi.org/10.1053/joca.1998.0205.
Ohnishi A, Osaki T, Matahira Y, Tsuka T, Imagawa T, Okamoto Y, et al. Correlation of plasma amino acid concentrations and chondroprotective effects of glucosamine and fish collagen peptide on the development of osteoarthritis. J Vet Med Sci. 2013;75(4):497–502.
Li X, Yang S, Qiu Y, Zhao T, Chen T, Su M, et al. Urinary metabolomics as a potentially novel diagnostic and stratification tool for knee osteoarthritis. Metabolomics. 2010;6(1):109–18.
Zhang WD, Likhodii S, Zhang YH, Aref-Eshghi E, Harper PE, Randell E, et al. Classification of osteoarthritis phenotypes by metabolomics analysis. BMJ Open. 2014;4(11):e006286. https://doi.org/10.1136/bmjopen-2014-006286.
Reuter SE, Evans AM. Carnitine and acylcarnitines. Clin Pharmacokinet. 2012;51(9):553–72.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Osteoarthritis
Rights and permissions
About this article
Cite this article
Showiheen, S.A.A., Sun, A.R., Wu, X. et al. Application of Metabolomics to Osteoarthritis: from Basic Science to the Clinical Approach. Curr Rheumatol Rep 21, 26 (2019). https://doi.org/10.1007/s11926-019-0827-8
Published:
DOI: https://doi.org/10.1007/s11926-019-0827-8