Abstract
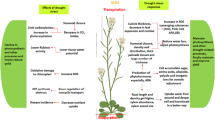
Abiotic stresses such as drought, salinity, frost, etc., affect plant yield manyfold. These stresses can decrease the plant yield of important major crops up to 50%. The abiotic stress-related genes or other transcription factors (TFs) have multiple functions, as it increases proline content, leads closing of stomata to decrease the transpiration rate, enhances the production of some important stress-related protective enzymes, etc. and hence increases abiotic stress tolerance. Many TFs and other stress-related genes have been identified and characterized and transformed to many important cultivated plants against drought and others abiotic stresses. The transformed plants show better morpho-biochemical and physiological performances than non-transgenic plants. Many genetically engineered plants have been developed against drought stress including wheat, rice, tomato, soybean, cotton and many more. The efficiently engineered clustered regulatory interspaced short palindromic repeats (CRISPR)/CRISPR-associated nuclease 9 (Cas9) system is now becoming a preferred choice of researchers to edit plant genomes for introgression natural resistance against a range of abiotic stresses. It leads genome editing by precise manure with minimal or no effect on growth and development of plants. Very limited reports are available to develop drought-tolerant plants using CRISPR/Cas9 system. Here we discuss transgenic plant technology and new [CRISPR Cas9 and Virus-Induced Gene Silencing (VIGS)] techniques to confer drought tolerance in important plant species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Global food security is one of the key issues due to change of world climate condition and by the increase of population. Drought, salinity, heat, etc., stresses significantly affect plant yield and productivity (Nouri et al. 2015; Singh et al. 2018). Drought and salinity affect about one-third of our cultivated land and it cause loss of approximately 1,500,000 ha crop land/year (Peng et al. 2011). In recent years, drought stress significantly decreased the plant yield manyfold by disturbing its morpho-biochemical processes (Azevedo et al. 2011). Even lower heat and drought stress affect plant yield. These stresses reduced crop yield by up to 50%. The plants in reproductive stages are more sensitive to these stresses and thus affect yield of many important plant species (Lamaou et al. 2018).
Plants are more susceptible to these extreme environmental stresses as compared to any other living organisms. Plants respond poorly to high environmental stresses, as it affects both biochemical and physiological processes. So, the development of new engineered plants is important to fight against these stresses (Ramonell and Somerville et al. 2002). Decreasing the water amount up to 40% declines the yield of maize and wheat by as much as 40% and 21% (Daryanto et al. 2016). The important cultivated cowpea yield is affected up to 68% by drought stress (Farooq et al. 2017). About 40% of soybean yield loss occurs due to drought stress (Specht et al. 1999). It also reduces cell division, spreading of leaf surface, retards stem growth and root propagation (Anjum et al. 2015). Long-term exposure of drought stress may cause death of plants (Gill and Tuteja 2010). In sunflower, it affects the chlorophyll a, b and a + b content (Manivannan et al. 2007). Drought stress enhances oxidative damage, stomatal closure, affects other cellular structures and decreases the exchange rate of gases (Hasanuzzaman et al. 2017). It also increases the ROS and leads to breakage of cellular physiological homeostasis (Fernández-Ocana et al. 2011).
The high-level drought and frost stresses in soil affect plants morpho-biochemical processes. Plants’ response to these environmental stresses has been extensively studied through various genetic engineering methods (Gosal et al. 2009). The stress genes regulate important drought stress-related pathways and hence increase drought tolerance in plants. Several drought-resistant genes have been characterized in many plant species whose function is still unknown. Therefore, reverse genetics approaches are useful for proper identification of these abiotic stress-related genes (Azevedo et al. 2011). The conventional breeding techniques are not enough to provide long-term resistance against abiotic stresses. Therefore, identification and cloning of important stress-related gene families is important to provide long-term resistance against abiotic stresses (Shinwari et al. 1998; Narusaka et al. 2003; Jan et al. 2016a, 2017).
Thus in this review, we focus on the negative impact of drought stress on plants and discuss various genetic engineering approaches to confer resistance against drought stress.
Regulons involved in abiotic stress tolerance in plants, transcription factors (TFs) and specific drought-related genes
Abiotic stresses disturb normal morpho-biochemical and physiological processes, hence affect yield. The response to these stresses varies according to genotype (Wang et al. 2018). The plant genes are activated by TFs in combination with other transcription binding sites (Chaves and Oliveira, 2004; Kimotho et al. 2019). These TFs attached the cis-acting elements of upstream regions of all gene promoters (Ciarmiello et al. 2011). In additions, TFs activate or suppress the activity of DNA polymerase enzyme and play a key role in gene expression (Riechmann et al. 2000). TFs activate many stress-related genes and enhance drought tolerance response. The key basic amino acids are involved to confer resistance to plants against abiotic stress (Annunziato 2008). TFs are useful by providing protection to plants in both biotic and abiotic stress conditions (Umezawa et al. 2006). In model Arabidopsis plants, about 34 families have been identified containing approximately 1533 TFs which have been classified (Riechmann et al. 2000). Recently, Wen et al (2019) reported the tolerant response of an important tree species Betula platyphylla (birch). They identified 2917 drought stress-related genes through RNA-Seq method. Among these genes, some drought-related TFs families, i.e., ethylene responsive factor and myeloblastosis oncogene were found to be maximum. In addition, they found that BpERF2 and BpMYB102 TFs enhance plant response against drought. These two TFs further activate several other stress-related genes and provide drought tolerance. Sakuma et al (2002) described various types of DREBs transcription factors in model Arabidopsis plant. They envisaged that DREB1A and DREB2A bind to a specific six-nucleotide sequence (A/GCCGAC) of DRE and increase drought and cold tolerance in Arabidopsis plant. However, the specificity of these transcription factors varies with change in second and third nucleotides in the sequence (A/GCCGAC) of DRE. They further classified these proteins into different classes including AP-2 subfamily, RAV subfamily, DREB subfamily, ERF subfamily, and others. Agarwal et al. (2010) reported that the DREB genes belong to AP2/ERF (apetala2 and ethylene responsive factors) class of TFs and present in different plant species. The DREB1 and DREB2 activate other stress-related genes which leads enhanced tolerance in plants against these environmental extreme conditions. Shinozaki et al. (2003) have clarified that incorporation of a single transcription factor has enhanced tolerance in plants against salt, drought and cold stresses though activation of several stress-tolerant gene against these stresses (Fig. 1). They noted that DREB gene with DRE-containing promoter enhances tolerance in plants against abiotic stresses. The detailed process of activation of biotic and abiotic stress-related genes via different TFs is given in Fig. 2.
Effect of cold, salt loading and dehydration on expression of stress-related genes. The activation of stress-related genes enhanced plant tolerance against drought (Shinozaki et al. 2003)
Activation of transcription factors to improve biotic and abiotic stress tolerance in plants (Fujita et al. 2007)
These TFs are classified into different families, i.e., AP2 transcription factors (Liu et al. 1998), B ZIP transcription factors (Uno et al. 2000), MYB transcription factors ( Fujita et al., 2007) and zinc finger transcription factors (Sugano et al. 2003). Grill and Himmelbach (1998) stated that the two pathways adopted by the plant for the stress response, i.e., abscisic acid-dependent and abscisic acid-independent gene expression are activated by a number of transcription factors such as ABFs proteins (ABRE-binding factor), ABA responsive element binding protein (AREB proteins), MYB/MYC proteins, DREB factors (dehydration responsive element binding factors) and NAC proteins (NAM, ATAF1-2, and CUC domains). The CBF/DREB1 transcription factors act in abscisic acid-independent pathway for the induction of stress-responsive gene by cold stress signal. The bZIP and ABRE transcription family activate stress-responsive gene in abscisic acid-dependent stress signal pathway. Most of the abscisic acid-inducible genes have a nine base pair conserved cis-acting sequence (PyACGTGGC) namely ABA responsive element in their promoter.
Several other DREB-related genes have been characterized and transformed to important cultivated plant species against abiotic stresses. Like, Maruyama et al (2004) reported 38 different classes of DREB genes. They also showed that the transgenic plants express these genes that have multiple functions against drought and salt. Dubouzet et al (2003) isolated and constructed five different CDNAs of DREB1A genes: OsDREB1A, OsDREB1B, OsDREB1C, OsDREB1D, and OsDREB2A from rice and their functions were also characterized. Among these genes, OsDREB1A and OsDREB1B were highly activated in drought and cold conditions while OsDREB2A showed a high level of expression in drought and high salt condition. The resulting OsDREB1A from rice was transformed to Arabidopsis plant and the resulting transgenic plant showed multiple abiotic stress tolerance. From their work, they concluded that OsDREB1A gene show functional similarity with AtDREB1A gene and it is very useful to transform OsDREB1A gene in monocots to produce tolerance against these extreme environmental stresses. The DREBs transcription factors are further classified into three DREB1A and five DREB2A proteins, these two proteins have a main role in the activation of multiple genes against abiotic stresses. Chen et al (2008) isolated GmDREB2 gene from Soybean (Glycin mix) and was transformed to Arabidopsis plant. The resulted transformed Arabidopsis plant showed better response and higher survival rate under drought, salt and cold stress conditions than control plants. Schramm et al (2008) reported that DREB2A transcription factor has a key role in the activation of several heat stress-related proteins especially HsfA3 which provide protection to tested Arabidopsis plant under extreme thermal conditions. Wang et al (2008) reported that DREB gene enhances abiotic stress tolerance in plants. Yamaguchi and Shinozaki (2007) also recorded similar results in Arabidopsis plants by expressing DREB1A gene. The expression analysis revealed that transformed plants showed enhanced tolerance to abiotic stress but also retarded normal plant growth. The DREB1A cDNA has an important role in many agriculturally important crops by gene transfer method. They described that the 35S CaMV promoter also leads to several morphological changes in transgenic Arabidopsis plants. Li et al (2005) isolated three different types of DREB genes from Soybean (Glycin mix) namely GmDREBa, GmDREBb, and GmDREBc under drought, and salt stress conditions. The expression analysis showed that GmDREBa, GmDREBb genes are expressed in leaves under salt and drought stress condition, while GmDREBc is only expressed in roots under extreme drought and salt condition. From their results, they concluded that these three types of DREB genes promote plant response against drought and salt. Kasuga et al. (1999) studied transformation of DREB1A gene to tobacco and found enhanced tolerance against both drought and cold. Details about the function of DREB1A and DREB2B are given in Fig. 3.
The functions of drought stress-inducible genes in stress tolerance and response. Two types of proteins are produced in response to drought stress. The first group includes function proteins that probably function in stress tolerance and the second group includes regulatory proteins that function in signal transduction and gene expression of other abiotic stress-related genes (Shinozaki and Yamaguchi-Shinozaki 2007)
The maize WRKY TF (ZmWRKY4) provides drought, salinity and temperature stress resistance in transgenic Arabidopsis. The gene up-regulates other abiotic stress-tolerant genes in transformed plants. In addition, it also increases important stress-related enzymes, i.e., peroxide dismutase and catalase and increases drought tolerance (Wang et al. 2018). Similarly, wheat TF (TaWRKY33) enhances resistance in transformed Arabidopsis plants against both drought and salt (He et al. 2016). The DREB2 and antioxidant enzyme gene (CAT1) in bread wheat enhance drought resistance. The relative water content (RWC), total chlorophyll content and catalase activity was found to be maximum in wheat cultivar (Kavir) under drought stress condition. So, these two genes are important for drought resistance (Eftekhari et al. 2017). The transgenic wheat lines expressing mutated TF (HaHB4) of sunflower enhance water use efficiency under abiotic stress (González et al. 2019). The high expression of ERF1-V in wheat provided tolerance against powdery mildew disease and salt and drought stresses (Xing et al. 2017).
The rice Rab family proteins, i.e., Rab7 (OsRab7) enhance drought and heat stress response in transformed rice genotypes. The overall morpho-biochemical and physiological responses were higher in transformed plants than wild rice genotypes under both stresses. In addition, malondialdehyde (MDA), hydrogen peroxide and electrolyte leakage was lower in transgenic plants. High yield was noted for transgenic rice than non-transgenic plants (Masood et al. 2005; El-Esawi and Alayafi 2019).The TFs (CUC2, ATAF1-2, and NAM) play a key role in plant developmental stages and abiotic stress conditions. At vegetative stage, the high level of OsNAC14 expression increases tolerance in drought condition in engineered rice plants. RNA-sequencing data showed that OsNAC14 increased the expression of other stress-related genes, plant defense, DNA repair system and biosynthesis of strigolactone (Shim et al. 2018). Also, ZmWRKY58 increases drought tolerance in transformed rice (Cai et al. 2014). The higher expression of SNAC1gene increased drought tolerance in rice. The engineered plants showed higher seed setting (22–34%) at reproductive stage than non-transgenic plants. In addition, no adverse effect was found in plant morphology and yield. The DNA chip data also showed that overexpression of SNAC1 gene activates other TFs (NAM, ATAF, and CUC (NAC) hence, improved drought and salt tolerance (Hu et al. 2006). The transgenic indica rice genotype expressing AtDREBIA gene shows drought tolerance at both vegetative and reproductive stages in T1, T2, and in following generations. The transgenic rice shows better physiological responses than wild plants. The tolerance, spikelet fertility and grain yield response were found to be maximum in homozygous lines than non-transgenic plants (Ravikumar et al. 2014). In another report, the overexpression of OsERF71 enhances drought tolerance in transgenic rice (Lee et al. 2016).
The drought tolerance is found maximum in cotton TM-1 genotype due to high expression of several coding TFs and other regulatory and enzymes controlling genes (ERF, ERFB, DREB, etc.). The TM-1 genotype showed maximum biochemical and physiological performance and healthy chloroplast structure under drought condition (Nakashima et al. 2000; Mosfeq-Ul Hasan et al. 2018). WRKYs in soybean promote drought tolerance. The activation of GmWRKY12 in tissues is minimal under normal conditions than drought condition. In hairy root culture, GmWRKY12 is responsible for higher proline and MDA content production, thus provides tolerance against drought and salt stresses in transgenic plants seedlings (Shi et al. 2018). Transgenic soybean expressing AtABF3 shows both drought and salt tolerance even at the end of 20 days of stress. Various physiological changes were noted during these stress conditions. The chlorophyll amount was found to be higher in transformed plants than control plants. Maximum stomata were found closed at stress condition. In addition, the engineered plants showed normal and stable cell membrane structure and lower transpiration rate at both stress conditions. The transgenic plants showed more total seed weight than NT plants. However, the drought and salt stress responses varied with types of genotype (Kim et al. 2018).
The transgenic tomato genotypes overexpressing the AnnSp2 gene show tolerance to both drought and salinity stresses. In severe stress condition, the AnnSp2 gene enhances ABA content, hence increased rate of closing stomata and decreased water loss. In addition, the transgenic plants showed better biochemical and physiological responses and lower ROS content (Ijaz et al. 2017). The ATAF1 gene from NAC family may increase drought resistance in tomato (Awais et al. 2018).
The SsDREB TFs of Suaeda salsa enhance drought and salt tolerance in tobacco plants. The chlorophyll, proline and soluble sugar content and photosynthetic rate were found to be maximum in transgenic plants as compared to control plants (Zhang et al. 2015). The high expression of Pennisetum glaucum Rab7 in tobacco plant increases both salt and osmotic stress resistance (Agarwal et al. 2008). Cong et al (2008) reported that transgenic Tobacco plant expressed the Brassica juncea DREB1B gene. The resulted transgenic plants showed a high level of proline content as compared to non-transgenic plant which leads to enhanced abiotic stress tolerance in transformed plants.
Many TFs belonging to family AP2/ERF were overexpressed in plants and provide resistance to different types of stresses (Mizoi et al. 2012; Phukan et al. 2017). The high expression of SpERF1 in engineered A. thaliana improves drought resistance (Yang et al. 2016). The AhDREB1gene in Arabidopsis increase ABA level and activate many drought-tolerant genes (RD29A, P5CS2, NCED1, and P5CS1). The resulted transgenic plant showed higher drought tolerance as compared to non-transgenic plants. In addition, at osmotic stress condition, the histone acetylation significantly affects the regulation of AhDREB1 gene, hence increases tolerance against drought (Zhang et al. 2018).The overexpression of AtRabG3e in Arabidopsis increases both salt and osmotic stresses tolerance (Mazel et al. 2004).The engineered Arabidopsis plants having a low amount of lignin and xylan acetylation shows maximum drought tolerance response as compared to control plants. The lower lignin content is due to expression of QsuB, which enhance abscisic acid (ABA) response for germination of seed and stomata closing. In addition, the low xylan-modified plants showed high amount of galactose and sugar (Yan et al. 2018). PYR1/PYL/RCARs play a key role in ABA signal transduction. The ZmPYLs is important in various abiotic stress conditions. Transgenic Arabidopsis-expressing ZmPYL increased sensitivity to ABA. The transformed plants having ZmPYL12, ZmPYL9, and Zmpyl8 show more resistance to drought. The accumulation of a high amount of protective protein (proline) further justifies the drought tolerance capability of ZmPYL genes in plants (He et al. 2018). MtMYBS1 increases both drought and salt tolerance in engineered Arabidopsis plants (Dong et al. 2017). The high expression of GaMYB62L in transformed Arabidopsis plants increases drought tolerance (Butt et al. 2017). Triple mutation of snrk2.2/3/6 removes the ABA response and increases drought susceptibility (Fujii and Zhu 2009). SnRK2.6 enhances the expression of regulating the ubiquitin E3 ligase activity of RZFP34/CHYR1 and promotes drought tolerance (Ding et al. 2015). HAI PP2Cs enhance the content of proline (an important osmo-regulator) and promote drought tolerance (Bhaskara et al. 2012). The high-level expression of epidermal patterning gene (OsEPF1) from rice increased drought tolerance by decreasing stomatal density in transgenic plants. The low stomata density allows rice plant to store about 60% of water between weeks 4 and 5 post-germination. Elevated CO2 level enables transgenic plants to survive in high drought and temperature (40 °C) conditions for a longer time period. In addition, low stomatal density enhances plant yield and helps plant to adapt better under warm climate condition (Caine et al., 2019). The overexpression of Cycling Dof Factor 3 enhances drought, salt and frost tolerance in Arabidopsis (Corrales et al. 2017). Details of different TFs or stress-related genes transformed into important plant species are given in Table 1.
Advanced genomic methods used to confer drought resistance in important plant species
Transgenic technology is very expensive, time consuming and difficult. In addition, this technology is not successful in many important cultivated crops (Slade et al. 2005). Various techniques like acetylation (Kim et al. 2012), methylation (Fu et al. 2013, 2017) and ubiquitination (Chen et al. 2018) are important to produce drought-tolerant genotypes. Some other efficient and quick technologies have been developed recently for crop improvement and for specific gene analysis like VIGS and CRISPR/Cas9 (Senthil-Kumar and Mysore 2014; Khatodia et al. 2016) (Tables 2, 3).
VIGS is an RNA-based antiviral defense technique commonly used to confer resistance against viruses. However, the virus vectors having genes can target the corresponding mRNAs. This method has been exploited in plants for analysis of gene function and has been adapted for high-throughput functional genomics. Various RNA viruses were engineered as viral vectors to check the function of abiotic stress-related genes (Lu et al. 2003). The knock-down of drought-inducible variant (H1-S) in tomato increased drought tolerance and stomata closing (Scippa et al. 2004). The overexpression of TaH2A.7 variant in Arabidopsis increased drought response and decreased water loss efficiency (Xu et al. 2016). The knock-down TaH2B-7D gene via VIGS in common wheat increases the relative electrolyte leakage rate and malondialdehyde amount. In addition, it decreases proline and percent relative water content, hence provides drought tolerance. The knock-down plants shows dwarf phenotype and symptoms of wilting as compared to non-knock-down genotypes. It means that TaH2B-7D gene provides tolerance under drought condition in common wheat (wang et al. 2019). The upregulation of AtHUB2 gene enhances cotton response against drought (Chen et al. 2018). The knock-down of three important genes (SpMAPK1, SpMAPK2, and SpMAPK3) in Solanum pimpinellifolium reduced drought tolerance. The silencing of GhWRKY27a gene enhances drought tolerance in cotton (Yan et al. 2015a, b). The VIGS of two important genes (GhNAC79 and JUB1) decrease drought response in cotton and tomato (Tasaki et al. 2016; Thirumalaikumar et al. 2018). The silencing of two important genes (SISR1L and SlGRX1) led to lower drought tolerance in tomato (Guo et al. 2010). The list of abiotic stress-related genes silenced through VIGS technique is given in Table 2.
The CRISPR/Cas9 system was discovered early in prokaryotic organisms that act as source of defense against the foreign phages by cleaving its genome through RNA-guided DNA nuclease by specialized mechanism (Sorek et al. 2013). Recently, this system has been used as a major source of genome editing for many living organisms including plants (Belhaj et al.2015; Khurshid et al. 2017; Shinwari et al. 2017; Xing et al. 2014). CRISPR/Cas9 technology has been extensively used as a novel technique for conferring drought resistance in plants (Singh et al. 2018).The SlNPR1cis-acting element provides drought tolerance in tomato. The CRISPR/Cas9-based mutation of mutant tomato slnpr1 mutants showed lower resistance to drought with a higher amount of electrolyte leakage and stomatal opening, hydrogen peroxide and MDA content than wild type. These findings showed that SlNPR1 is involved in drought tolerance response in tomato (Li et al. 2019). The maize ARGOS8 variants produced by CRISPR/Cas9-based method showed tolerance to drought and gave maximum yield (Shi et al. 2017). The detailed importance of CRISPR/Cas9 system against drought stress in important plant species is given in Table 3.
Conclusions
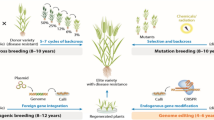
Drought condition has been increased globally over the last few decades and it is one of the major constraints that limit our crop productivity and sustainable agriculture. The drought condition decreased food and feed dramatically especially in the last 2 decades. Therefore, it is vital to develop novel drought resistance varieties for further crop improvement. The transgenic technology can solve this problem by the production of new drought-resistant genotypes with no or minimal effect on plant morpho-biochemical and physiological performances. A large number of environmental stress-related genes have been isolated and transformed to various cultivated plants and the resulted engineered plants showed enhanced drought tolerance. Detailed steps of genetic transformation are given in Fig. 4a. New in planta (tissue culture free)-based transformation protocols should be developed for genotypes having problems with tissue culture protocol and long germination to maturity time periods (Fig. 4b). Many other novel techniques have been developed that are more efficient, precise and quick than genetically modified (GM) technology. VIGS is a quick and robust method to check the function of genes, involved in abiotic stress tolerance in plants. However, this technique has been successfully used for only few model plants and there is a need to extend this technique to check the function of others abiotic stress-related genes in the model and other economically important crop species. Therefore, developing new viral vectors for model and other organisms need to be well optimized. Recently, novel CRISPR/Cas9 plant genome editing protocols have been developed by different researchers. It is one of the novel methods of plant genome editing and provides resistance against drought stress. This system also helps us to know about the gene knock outs/knock in, epigenetic mechanism and gene regulation. However, several modifications are needed to develop some new drought-resistant engineered plants using CRISPR technology. The CRISPR/Cas9 method may produce some new transgenic plants against drought stress with minimal or no biosafety issues in near future. Only few transgenic plants have been developed against drought stress using CRISPR/Cas9 system. Therefore, new drought-resistant plants should be developed using this technology. In addition, new stress models and multiple stress markers data should be developed. The new drought avoidance, escape and tolerance strategies should be used for providing long-term drought tolerance in plants.
a Various steps involved in the development of stable transgenic plants via Agrobacterium-based transformation into different plant species. This protocol is useful for plants having tissue culture protocol optimized. The protocol is expensive and time consuming and needs a long time period. b In planta transformation is a direct transformation method to develop transgenic plants in a short time. The protocol does not need any tissue culture method and is less expensive. The production of chimera plants is a major drawback of this method (Jan et al. 2016b)
References
Agarwal PK, Acharia BJH (2010) Transcription factors in plants and ABA dependent and independent abiotic stress signaling. Biol Plant 54(2):201–212
Agarwal PK, Agarwal P, Jain P et al (2008) Constitutive overexpression of a stress-inducible small GTP-binding protein PgRab7 from Pennisetum glaucum enhances abiotic stress tolerance in transgenic tobacco. Plant Cell Rep 27:105–115. https://doi.org/10.1007/s00299-007-0446-0
Anjum SA, Xie X, Wang L et al (2015) Morphological, physiological and biochemical responses of plants to drought stress. Acta Physiol Plant 37:2026–2032. https://doi.org/10.5897/AJAR10.027
Annunziato A (2008) DNA packaging: nucleosomes and chromatin. Nat Educ 1(1):26
Awais M, Ahmad R, Khan N et al (2018) Transformation of tomato variety rio grande with drought resistant transcription factor gene ATAF1 and its molecular analysis. Pak J Bot 50(5): 1805–1810. https://hdl.handle.net/21.11116/0000-0001-8295-3
Azevedo H, Silva-Correia J, Oliveira J et al (2011) A strategy for the identification of new abiotic stress determinants in Arabidopsis using web-based data mining and reverse genetics. OMICS 15(12):935–947. https://doi.org/10.1089/omi.2011.0083
Bai C, Wang P, Fan Q et al (2017) Analysis of the role of the drought-induced gene DRI15 and salinity-induced gene SI1 in Alternanthera philoxeroides plasticity using a virus-based gene silencing tool. Front Plant Sci 8:1579. https://doi.org/10.3389/fpls.2017.01579
Banavath JN, Chakradhar T, Pandit V et al (2018) Stress inducible overexpression of AtHDG11 leads to improved drought and salt stress tolerance in peanut (Arachis hypogaea L.). Front Chem 6:34. https://doi.org/10.1038/nrmicro280
Belhaj K, Chaparro-Garcia A, Kamoun S et al (2015) Editing plant genomes with CRISPR/Cas9. Curr Opin Biotechnol 32:76–84. https://doi.org/10.1016/j.copbio.2014.11.007
Bhaskara GB, Nguyen TT, Verslues PE (2012) Unique drought resistance functions of the highly ABA-induced clade A protein phosphatase 2Cs. Plant Physiol 160:379–395. https://doi.org/10.1104/pp.112.202408
Butt HI, Yang Z, Chen E et al (2017) Functional characterization of cotton GaMYB62L, a novel R2R3 TF in transgenic Arabidopsis. PLoS ONE 12(1):e0170578. https://doi.org/10.1371/journal.pone.0170578
Cai RH, Zhao Y, Wang YF et al (2014) Overexpression of a maize WRKY58 gene enhances drought and salt tolerance in transgenic rice. Plant Cell Tissue Organ Cult 119:565–577
Caine RS, Yin X, Sloan J et al (2019) Rice with reduced stomatal density conserves water and has improved drought tolerance under future climate conditions. New Phytol 221(1):371–384. https://doi.org/10.1111/nph.15344
Casaretto JA, El-kereamy A, Zeng B et al (2016) Expression of OsMYB55 in maize activates stress-responsive genes and enhances heat and drought tolerance. BMC Genomics 17:312. https://doi.org/10.1186/s12864-016-2659-5
Chaves MM, Oliveira MM (2004) Mechanisms underlying plant resilience to water deficits: prospects for water-saving agriculture. J Exp Bot 55(407):2365–2384. https://doi.org/10.1093/jxb/erh269
Chen H, Feng H, Zhang XY et al (2018) An Arabidopsis E3 ligase HUB2 increases histone H2B monoubiquitination and enhances drought tolerance in transgenic cotton. Plant Biotechnol J 1:1. https://doi.org/10.1111/pbi.12998
Chen QJ, Meng XP, Zhang Y et al (2008) Over-expression of OsDREB genes lead to enhanced drought tolerance in rice. Biotechnol Lett 30:2191–2198. https://doi.org/10.1007/s10529-008-9811-5
Ciarmiello LF, Woodrow P, Fuggi A et al (2011). Plant genes for abiotic stress, abiotic stress in plants–mechanisms and adaptations, Prof. Arun Shanker (Ed.), ISBN: 978-953-307-394-1, InTech. https://www.intechopen.com/books/abioticstress-in-plants-mechanisms-and-adaptations/plant-genes-for-abiotic-stress.
Cong L, Chai TY, Zhang YX (2008) Characterization of the novel gene BjDREB1B encoding a DRE-binding transcription factor from Brassica juncea L. Biochem Biophys Res Com 371:702–706. https://doi.org/10.1016/j.bbrc.2008.04.126
Corrales AR, Carrillo L, Lasierra P et al (2017) Multifaceted role of cycling Dof factor 3(CDF3) in the regulation of flowering time and abiotic stress responses in Arabidopsis. Plant Cell Environ 40:748–764. https://doi.org/10.1111/pce.12894
Daryanto S, Wang L, Jacinthe PA (2016) Global synthesis of drought effects on maize and wheat production. PLoS ONE 11:e0156362. https://doi.org/10.1371/journal.pone.0156362
Ding S, Zhang B, Qin F (2015) Arabidopsis RZFP34/CHYR1, a ubiquitin E3 ligase, regulates stomatal movement and drought tolerance via SnRK2.6- mediated phosphorylation. Plant Cell 27:3228–3244. https://doi.org/10.1105/tpc.15.00321
Dong W, Song Y, Zhao Z et al (2017) The Medicago truncatula R2R3-MYB transcription factor gene MtMYBS1 enhances salinity tolerance when constitutively expressed in Arabidopsis thaliana. Biochem Biophys Res Commun 490:225–230. https://doi.org/10.1016/j.bbrc.2017.06.0251117
Dubouzet JG, Sakuma Y, Ito Y et al (2003) OsDREB genes in rice, Oryza sativa L., encode transcription activators that function in drought, high-salt- and cold-responsive gene expression. Plant J 33:751–763. https://doi.org/10.1046/j.1365-313x.2003.01661.x
Eftekhari A, Baghizadeh A, Yaghoobi MM et al (2017) Differences in the drought stress response of DREB2 and CAT1 genes and evaluation of related physiological parameters in some bread wheat cultivars. Biotechnol Biotechnol Equip 31(4):709–716. https://doi.org/10.1080/13102818.2017.1316214
El-Esawi MA, Alayafi AA (2019) Overexpression of rice Rab7 gene improves drought and heat tolerance and increases grain yield in rice (Oryza sativa L). Genes 10(1):56. https://doi.org/10.3390/genes10010056
Farooq M, Gogoi N, Barthakur S et al (2017) Drought stress in grain legumes during reproduction and grain filling. J Agron Crop Sci 203:81–102. https://doi.org/10.1111/jac.12169
Fernández-Ocana, A, Chaki M, Luque F et al (2011) Functional analysis of superoxide dismutases (SODs) in sunflower under biotic and abiotic stress conditions. Identification of two new genes of mitochondrial Mn-SOD. J Plant Physiol 168(11):1303–1308. https://doi.org/10.1016/j.jplph.2011.01.020
Fu Y, Ma H, Chen S et al (2017) Control of proline accumulation under drought via a novel pathway comprising the histone methylase CAU1 and the transcription factor ANAC055. J Exp Bot 69:579–588. https://doi.org/10.1093/jxb/erx419
Fu YL, Zhang GB, Lv XF et al (2013) Arabidopsis histone methylase CAU1/PRMT5/SKB1 acts as an epigenetic suppressor of the calcium signaling gene CAS to mediate stomatal closure in response to extracellular calcium. Plant Cell 25:2878–2891. https://doi.org/10.1105/tpc.113.113886
Fujii H, Zhu JK (2009) Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc Natl Acad Sci USA 106:8380–8385. https://doi.org/10.1073/pnas.0903144106
Fujita Y, Yamaguchi-Shinozaki K, Shinozaki K (2007) Improving drought and salt stress tolerance in plants by gene transfer. 4th Biomass-Asia, Workshop
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930. https://doi.org/10.1016/j.plaphy.2010.08.016
González FG, Capella M, Ribichich KF et al (2019) Field-grown transgenic wheat expressing the sunflower gene HaHB4 significantly outyields the wild type. J Exp Bot 70(5):1669–1681. https://doi.org/10.1093/jxb/erz037
Gosal SS, Wani SH, Kang MS (2009) Biotechnology and drought tolerance. J Crop Improv 23:19–54. https://doi.org/10.1080/15427520802418251
Grill E, Himmelbach A (1998) ABA signal transduction. Curr Opin Plant Biol 1:412–418. https://doi.org/10.1016/s1369-5266(98)80265-3
Guo Y, Huang C, Xie Y et al (2010) A tomato glutaredoxin gene SlGRX1 regulates plant responses to oxidative, drought and salt stresses. Planta 232(6):1499–1509. https://doi.org/10.1007/s00425-010-1271-1
Hasan M, Ma F, Prodhan Z et al (2018) Molecular and physio-biochemical characterization of cotton species for assessing drought stress tolerance. Int J Mol Sci 19(9):2636. https://doi.org/10.3390/ijms19092636
Hasanuzzaman M, Nahar K, Gill SS et al (2017) Hydrogen peroxide pretreatment mitigates cadmium-induced oxidative stress in Brassica napus L: an intrinsic study on antioxidant defense and glyoxalase systems. Front Plant Sci 8:115. https://doi.org/10.3389/fpls.2017.00115
He Z, Zhong J, Sun X et al (2018) The maize ABA receptors ZmPYL8, 9, and 12 facilitate plant drought resistance. Front Plant Sci 9:422 https://doi.org/10.3389/fpls.2018.00422
He GH, Xu JY, Wang YX et al (2016) Drought-responsive WRKYtranscription factor genes TaWRKY1 and TaWRKY33 from wheat confer drought and/or heat resistance in Arabidopsis. BMC Plant Biol 16:116. https://doi.org/10.1186/s12870-016-0806-4
Hu H, Dai M, Yao J et al (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Nat Acad Sci 103(35):12987–12992. https://doi.org/10.1073/pnas.0604882103
Ijaz R, Ejaz J, Gao S et al (2017) Overexpression of annexin gene AnnSp2, enhances drought and salt tolerance through modulation of ABA synthesis and scavenging ROS in tomato. Sci Rep 7(1):12087. https://doi.org/10.1038/s41598-017-11168-2
Jan SA, Bibi N, Shinwari ZK et al (2017) Impact of salt, drought, heat and frost stresses on morpho-biochemical and physiological properties of Brassica species: an updated review. J Rural Dev Agri 2(1):1–10
Jan SA, Shinwari ZK, Rabbani MA (2016a) Agro-morphological and physiological responses of Brassica rapa ecotypes to salt stress. Pak J Bot 48(4):1379–1384
Jan SA, Shinwari ZK, Shah SH et al (2016b) In-planta transformation recent advances. Rom Biotechnol Lett 21(1):11085–11091
Kasuga M, Liu Q, Miura S (1999) Improving plant drought, salt and freezing tolerance by gene transfer of a single stress-inducible transcription factor. Nat Biotechnol 17:287–291. https://doi.org/10.1038/7036
Khatodia S, Bhatotia K, Passricha N et al (2016) The CRISPR/Cas genome-editing tool: application in improvement of crops. Front Plant Sci 7:506. https://doi.org/10.3389/fpls.2016.00506
Khoudi H, Khemakhem AN, Gouiaa S et al (2009) Optimization and regeneration and transformation parameters in tomato and improvement of its salinity and drought tolerance. Afr J Biotechnol 8(22):6068–6076. https://doi.org/10.5897/AJB09.057
Khurshid H, Jan SA, Shinwari ZK et al (2017) An era of CRISPR/Cas9 mediated plant genome editing. Curr Iss Mol Biol 26:47–54. https://doi.org/10.21775/cimb.026.047
Kim HJ, Cho HS, Pak JH et al (2018) Confirmation of drought tolerance of ectopically expressed AtABF3 gene in soybean. Mol Cells 41(5):413. https://doi.org/10.14348/molcells.2018.2254
Kim JM, To TK, Ishida J et al (2012) Transition of chromatin status during the process of recovery from drought stress in Arabidopsis thaliana. Plant Cell Physiol 53:847–856. https://doi.org/10.1093/pcp/pcs053
Kimotho RN, Baillo EH, Zhang Z (2019) Transcription factors involved in abiotic stress responses in Maize (Zea mays L) and their roles in enhanced productivity in the post genomics era. Peer J 7:e7211. https://doi.org/10.7717/peerj.7211
Kiranmai K, Lokanadha Rao G, Pandurangaiah M et al. (2018). A novel WRKY transcription factor, MuWRKY3 (Macrotyloma uniflorum Lam. Verdc) enhances drought stress tolerance in transgenic groundnut (Arachis hypogaea L) plants. Front Plant Sci 9:346. Doi: 10.3389/fpls.2018.00346.
Kirungu JN, Magwanga RO, Lu P et al (2019) Functional characterization of Gh_A08G1120 (GH3 5) gene reveal their significant role in enhancing drought and salt stress tolerance in cotton. BMC Genet 20(1):62. https://doi.org/10.1186/s12863-019-0756-6
Kornberg RD (1974) Chromatin structure: a repeating unit of histones and DNA. Science 184:868–871. https://doi.org/10.1126/science.184.4139.868
Kuzuoglu-Ozturk D, Cebeci Yalcinkaya O, Akpinar BA et al (2012) Autophagy-related gene, TdAtg8, in wild emmer wheat plays a role in drought and osmotic stress response. Planta 236:1081–1092. https://doi.org/10.1007/s00425-012-1657-3
Lamaoui M, Jemo M, Datla R et al (2018) Heat and drought stresses in crops and approaches for their mitigation. Front Chem 6:26. https://doi.org/10.3389/fchem.2018.00026
Lee DK, Jung H, Jang G et al (2016) Overexpression of theOsERF71 transcription factor alters rice root structure and drought resistance. Plant Physiol 172:575–588. https://doi.org/10.1104/pp.16.00379
Li R, Liu C, Zhao R et al (2019) CRISPR/Cas9-Mediated SlNPR1 mutagenesis reduces tomato plant drought tolerance. BMC Plant Biol 19(1):38. https://doi.org/10.1186/s12870-018-1627-4
Li J, Phan TT, Li YR et al (2018) Isolation, transformation and overexpression of sugarcane SoP5CS gene for drought tolerance improvement. Sugar Tech 20(4):464–473. https://doi.org/10.1007/s12355-017-0568-9
Li P, Li YJ, Zhang FJ et al (2017) The Arabidopsis UDP-glycosyltransferases UGT79B2 and UGT79B3, contribute to cold, salt and drought stress tolerance via modulating anthocyanin accumulation. Plant J 89(1):85–103. https://doi.org/10.1111/tpj.13324
Li L, Zheng M, Deng G et al (2016) Overexpression of AtHDG11 enhanced drought tolerance in wheat (Triticum aestivum L). Mol Breed 36:23. https://doi.org/10.1007/s11032-016-0447-1
Li XP, Tian AG, Luo GZ (2005) Soybean DRE-binding transcription factors that are responsive to abiotic stresses. Theor Appl Genet 110:1355–1362. https://doi.org/10.1007/s00122-004-1867-6
Liang J, Deng G, Long H et al (2012) Virus-induced silencing of genes encoding LEA protein in Tibetan hulless barley (Hordeum vulgare ssp. vulgare) and their relationship to drought tolerance. Mol Breed 30:441–451. https://doi.org/10.1007/s11032-011-9633-3
Lim CW, Lee SC (2014) Functional roles of the pepper MLO protein gene, CaMLO2, in abscisic acid signaling and drought sensitivity. Plant Mol Biol 85:1–10. https://doi.org/10.1007/s11103-11013-10155-11108
Liu L, White JM, Macrae HT (1998) Transcription factors and their genes in higher plants: functional domains, evolution and regulation. Eur J Biochem 262:247–257. https://doi.org/10.1046/j.1432-1327.1999.00349.x
Liu X, Zhai S, Zhao Y et al (2013) Overexpression of the phosphatidylinositol synthase gene (ZmPIS) conferring drought stress tolerance by altering membrane lipid composition and increasing ABA synthesis in maize. Plant Cell Environ 36:1037–1055. https://doi.org/10.1111/pce.12040
Lou D, Wang H, Liang G et al (2017) OsSAPK2 confers abscisic acid sensitivity and tolerance to drought stress in rice. Front Plant Sci 8:993. https://doi.org/10.3389/fpls.2017.00993
Lu R, Martin-Hernandez AM, Peart JR et al (2003) Virus-induced gene silencing in plants. Methods 30(4):296–303. https://doi.org/10.1016/s1046-2023(03)00037-9
Lu Y, Li Y, Zhang J et al (2013) Overexpression of Arabidopsis molybdenum cofactor sulfurase gene confers drought tolerance in maize (Zea mays L). PLoS ONE 8:e52126. https://doi.org/10.1371/journalpone0052126
Manivannan P, Jaleel CA, Sankar B et al (2007) Growth, biochemical modifications and proline metabolism in Helianthus annuus L. as induced by drought stress. Colloids Surf B Biointerfaces 59:141–149. https://doi.org/10.1016/j.colsurfb.2007.05.002
Manmathan H, Shaner D, Snelling J et al (2013) Virus-induced gene silencing of Arabidopsis thaliana gene homologues in wheat identifies genes conferring improved drought tolerance. J Exp Bot 64:1381–1392. https://doi.org/10.1093/jxb/ert003
Maruyama K, Sakuma Y, Kasuga M et al (2004) Identification of cold inducible downstream genes of the Arabidopsis DREB1A/CBF3 transcriptional factor using two microarray systems. Plant J 38:982–993. https://doi.org/10.1111/j.1365-313X.2004.02100.x
Masood S, Seiji Y, Shinwari ZK, Anwar R (2005) Mapping quantitative trait loci (QTLs) for salt tolerance in rice (Oryza sativa) using RFLPs. Pak J Bot 36(4):825–834
Mazel A, Leshem Y, Tiwari BS et al (2004) Induction of salt and osmotic stress tolerance by overexpression of an intracellular vesicle trafficking protein AtRab7 (AtRabG3e). Plant Physiol 134:118–128. https://doi.org/10.1104/pp.103.025379
Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K (2012) AP2/ERF family transcription factors in plant abiotic stress responses. Biochim Biophys Acta 1819:86–96
Nakashima K, Shinwari ZK, Miura S et al (2000) Structural organization, expression and promoter activity of an Arabidopsis gene family encoding DRE/CRT binding proteins involved in dehydration- and high salinity-responsive gene expression. Plant Mol Biol 42(4):657–665. https://doi.org/10.1023/a:1006321900483
Narusaka Y, Nakashima K, Shinwari ZK et al (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis rd29A gene in response to dehydration and high salinity stresses. Plant J 34(2):137–149. https://doi.org/10.1046/j.1365-313x.2003.01708.x
Nouri MZ, Moumeni A, Komatsu S (2015) Abiotic stresses: insight into gene regulation and protein expression in photosynthetic pathways of plants. Int J Mol Sci1 6(9): 20392–20416. Doi: 10.3390/ijms160920392.
Paixao JFR, Gillet FX, Ribeiro TP et al (2019) Improved drought stress tolerance in Arabidopsis by CRISPR/dCas9 fusion with a histone acetyl transferase. Sci Rep 9(1):8080. https://doi.org/10.1038/s41598-019-44571-y
Pellegrineschi A, Reynolds M, Pacheco M et al (2004) Stress-induced expression in wheat of the Arabidopsis thaliana DREB1A gene delays water stress symptoms under greenhouse conditions. Genome 47:493–500. https://doi.org/10.1139/g03-140
Peng X, Ma X, Fan W et al (2011) Improved drought and salt tolerance of Arabidopsis thaliana by transgenic expression of a novel DREB gene from Leymus chinensis. Plant Cell Rep 30:1493–1502. https://doi.org/10.3390/ijms160920392
Phukan UJ, Jeena GS, Tripathi V et al (2017) Regulation of Apetala2/Ethylene response factors in plants. Front Plant Sci 8:150. https://doi.org/10.3389/fpls.2017.00150
Qin N, Xu W, Hu L et al (2016) Drought tolerance and proteomics studies of transgenic wheat containing the maize C4 phosphoenolpyruvate carboxylase (PEPC) gene. Protoplasma 253:1503–1512. https://doi.org/10.1007/s00709-015-0906-2
Ramonell KM, Somerville S (2002) The genomics parade of defense responses: to infinity and beyond. Curr Opin Plant Biol 5:291–294. https://doi.org/10.1016/s1369-5266(02)00266-2
Ravikumar G, Manimaran P, Voleti SR et al (2014) Stress-inducible expression of AtDREB1A transcription factor greatly improves drought stress tolerance in transgenic indica rice. Transgenic Res 23(3):421–439. https://doi.org/10.1007/s11248-013-9776-6
Riechmann JL, Heard J, Martin G et al (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110. https://doi.org/10.1126/science.290.5499.2105
Sakuma Y, Dubouzet JG, Abe H et al (2002) DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration- and cold-inducible gene expression. Biochem Biophys Res Com 290:998–1009. https://doi.org/10.1006/bbrc.2001.6299
Saravanan S, Kumar KK, Raveendran M et al (2018) Genetic engineering of sugarcane for drought and salt tolerant transgenic plants expressing the BcZAT12 gene. Int J Curr Microbiol App Sci 7(7):1594–1613. https://doi.org/10.20546/ijcmas.2018.707.188
Sarkar T, Thankappan R, Kumar A et al (2014) Heterologous expression of the AtDREB1A gene in transgenic peanut-conferred tolerance to drought and salinity stresses. PLoS ONE 9(12):e110507. https://doi.org/10.1371/journal.pone.0110507
Schramm F, Larkindale J, Kiehlmann E (2008) A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J 53:264–266. https://doi.org/10.1111/j.1365-313X.2007.03334.x
Scippa GS, Di Michele M, Onelli E et al (2004) The histone-like protein H1-S and the response of tomato leaves to water deficit. J Exp Bot 55:99–109. https://doi.org/10.1093/jxb/erh022
Senthil-Kumar M, Mysore KS (2014) Tobacco rattle virus-based virus-induced gene silencing in Nicotiana benthamiana. Nat Protoc 9(7):1549–1562. https://doi.org/10.1038/nprot.2014.092
Shen YG, Zhang WK, He SJ et al (2003) An EREBP/AP2-type protein in Triticum aestivum was a DRE-binding transcription factor induced by cold, dehydration and ABA stress. Theor Appl Genet 106:923–930. https://doi.org/10.1007/s00122-002-1131-x
Shi J, Habben JE, Archibald RL et al (2015) Overexpression of ARGOS genes modifies plant sensitivity to ethylene, leading to improved drought tolerance in both Arabidopsis and maize. Plant Physiol 169:266–282. https://doi.org/10.1104/pp.15.00780
Shi J, Gao H, Wang H et al (2017) ARGOS8 variants generated by CRISPR-Cas9 improve maize grain yield under field drought stress conditions. Plant Biotechnol J 15(2):207–216. https://doi.org/10.1111/pbi.12603
Shi WY, Du YT, Ma J et al (2018) The WRKY Transcription factor GmWRKY12 confers drought and salt tolerance in soybean. Int J Mol Sci 19(12):4087. https://doi.org/10.3390/ijms19124087
Shim JS, Oh N, Chung PJ et al (2018) Overexpression of OsNAC14 improves drought tolerance in rice. Front Plant Sci 9:310. https://doi.org/10.3389/fpls.2018.00310
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6(5):410–417. https://doi.org/10.1016/s1369-5266(03)00092-x
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58(2):221–227. https://doi.org/10.1093/jxb/erl164
Shinwari ZK, Nakashima K, Miura S et al (1998) An Arabidopsis gene family encoding DRE binding protein involved in low temperature—responsive gene expression. Biochem Biophys Res Com 250:161–170. https://doi.org/10.1111/pbi.12603
Shinwari ZK, Tanveer F, Khalil AT (2017) Ethical issues regarding CRISPR mediated genome editing. Curr Iss Mol Biol 26:103–110. https://doi.org/10.21775/cimb.026.103
Shou H, Bordallo P, Wang K (2004) Expression of the Nicotiana protein kinase (NPK1) enhanced drought tolerance in transgenic maize. J Exp Bot 55:1013–1019. https://doi.org/10.1093/jxb/erh129
Singh B, Kukreja S, Goutam U (2018) Milestones achieved in response to drought stress through reverse genetic approaches. F1000 Res 7:1311. Doi: 10.12688/f1000research.15606.1.
Slade AJ, Knauf VC (2005) TILLING moves beyond functional genomics into crop improvement. Transgenic Res 14(2):109–115. https://doi.org/10.1007/s11248-005-2770-x
Sorek R, Lawrence CM, Wiedenheft B (2013) CRISPR-mediated adaptive immune systems in bacteria and archaea. Ann Revol Biochem 82:237–266. https://doi.org/10.1146/annurev-biochem-072911-172315
Specht JE, Hume DJ, Kumudini SV et al (1999) Soybean yield potential-a genetic and physiological perspective. Crop Sci 39(6):1560–1570. https://doi.org/10.2135/cropsci1999.3961560x
Srinivasan T, Kumar KRR, Meur G et al (2009) Heterologous expression of ArabidopsisNPR1 (AtNPR1) enhances oxidative stress tolerance in transgenic tobacco plants. Biotechnol Lett 31(9):1343–1351. https://doi.org/10.1007/s10529-009-0022-5
Sugano S, Kaminaka H, Rybka Z et al (2003) Stress-responsive zinc finger gene ZPT2-3 plays a role in drought tolerance in petunia. Plant J 36:830–841. https://doi.org/10.1046/j.1365-313x.2003.01924.x
Tasaki K, Terada H, Masuta C et al (2016) Virus-induced gene silencing (VIGS) in Lilium leichtlinii using the Cucumber mosaic virus vector. Plant Biotechnol 33(5):373–381. https://doi.org/10.5511/plantbiotechnology.16.1018a
Thirumalaikumar VP, Devkar V, Mehterov N et al (2018) NAC transcription factor JUNGBRUNNEN1 enhances drought tolerance in tomato. Plant Biotechnol J 16(2):354–366. https://doi.org/10.5511/plantbiotechnology.16.1018a
Umezawa T, Fujita M, Fujita Y et al (2006) Engineering drought tolerance in plants: discovering and tailoring genes to unlock the future. Curr Opin Biotechnol 17:113–122. https://doi.org/10.5511/plantbiotechnology.16.1018a
Uno Y, Furihata T, Abe H et al (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97:11632–11637. https://doi.org/10.1073/pnas.190309197
Virk N, Liu B, Zhang H et al (2013) Tomato SlMPK4 is required for resistance against Botrytis cinerea and tolerance to drought stress. Acta Physiol Plant 35(4):1211–1221. https://doi.org/10.1007/s11738-012-1160-2
Wang CT, Ru JN, Liu Y et al (2018) The Maize WRKY transcription factor ZmWRKY40 confers drought resistance in transgenic Arabidopsis. Int J Mol Sci 19(9):2580. https://doi.org/10.3390/ijms19092580
Wang L, Chen L, Zhao R et al (2017) Reduced drought tolerance by CRISPR/Cas9-mediated SlMAPK3 mutagenesis in tomato plants. J Agric Food Chem 65(39):8674–8682. https://doi.org/10.1021/acs.jafc.7b02745
Wang Q, Guan Y, Wu Y et al (2008) Over-expression of a rice OsDREB1F gene increases salt, drought, and low temperature tolerance in both Arabidopsis and rice. Plant Mol Biol 67:589–602. https://doi.org/10.1007/s11103-008-9340-6
Wang X, Ren Y, Li J et al (2019) Knock-down the expression of TaH2B-7D using virus-induced gene silencing reduces wheat drought tolerance. Biol Res 52(1):14. https://doi.org/10.1186/s40659-019-0222-y
Wen X, Wang J, Zhang D et al (2019) A gene regulatory network controlled by BpERF2 and BpMYB102 in birch under drought conditions. Int J Mol Sci 20(12):3071. https://doi.org/10.3390/ijms20123071
Xing HL, Dong L, Wang ZP et al (2014) A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol 14:327. https://doi.org/10.1186/s12870-014-0327-y
Xing L, Di Z, Yang W et al (2017) Overexpression of ERF1-V from Haynaldia villosa can enhance the resistance of wheat to powdery mildew and increase the tolerance to salt and drought stresses. Front Plant Sci 8:1948. https://doi.org/10.3389/fpls.2017.01948
Xu C, Fu X, Liu R et al (2017) PtoMYB170 positively regulates lignin deposition during wood formation in poplar and confers drought tolerance in transgenic Arabidopsis. Tree Physiol 37(12):1713–1726. https://doi.org/10.1093/treephys/tpx093
Xu W, Li Y, Cheng Z et al (2016) A wheat histone variant gene TaH2A.7 enhances drought tolerance and promotes stomatal closure in Arabidopsis. Plant Cell Rep 35:1853–1862. https://doi.org/10.1007/s00299-016-1999-6
Xu XM, Lin H, Maple J et al (2010) The Arabidopsis DJ-1a protein confers stress protection through cytosolic SOD activation. J Cell Sci 123:1644–1651. https://doi.org/10.1242/jcs.063222
Yamaguchi-Shinozaki K, Shinozaki K (2007) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94. https://doi.org/10.1016/j.tplants.2004.12.012
Yan J, Aznar A, Chalvin C et al (2018) Increased drought tolerance in plants engineered for low lignin and low xylan content. Biotechnol Biofuels 11(1):195. https://doi.org/10.1186/s13068-018-1196-7
Yan Y, Jia H, Wang F et al (2015) Overexpression of GhWRKY27a reduces tolerance to drought stress and resistance to Rhizoctonia solani infection in transgenic Nicotiana benthamiana. Front Physiol 6:265. https://doi.org/10.3389/fphys.2015.00265
Yang Y, Dong C, Li X et al (2016) A novel AP2/ERF transcription factor from Stipa purpurea leads to enhanced drought tolerance in Arabidopsis thaliana. Plant Cell Rep 35:2227–2239. https://doi.org/10.1007/s00299-016-2030-y
Zhang X, Liu X, Wu L et al (2015) The SsDREB transcription factor from the succulent halophyte Suaeda salsa enhances abiotic stress tolerance in transgenic tobacco. Int J Genomics. https://doi.org/10.1155/2015/875497
Zhang B, Su L, Hu B et al (2018) Expression of AhDREB1, an AP2/ERF transcription factor gene from peanut, is affected by histone acetylation and increases abscisic acid sensitivity and tolerance to osmotic stress in Arabidopsis. Int J Mol Sci 19(5):1441. https://doi.org/10.3390/ijms19051441
Zhao Y, Zhang C, Liu W et al (2016) An alternative strategy for targeted gene replacement in plants using a dual-sgRNA/Cas9 design. Sci Rep 6:23890. https://doi.org/10.1038/srep23890
Author information
Authors and Affiliations
Contributions
Z-KS and S-AJ outlined and conceived the idea. Z-KS, S-AJ, K-N and K-YS helped in manuscript writing, editing and proof reading.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shinwari, Z.K., Jan, S.A., Nakashima, K. et al. Genetic engineering approaches to understanding drought tolerance in plants. Plant Biotechnol Rep 14, 151–162 (2020). https://doi.org/10.1007/s11816-020-00598-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-020-00598-6