Abstract
UDP-glucose 4-epimerase (UGE; EC 5.1.3.2) is an enzyme that plays an essential role in the interconverts UDP-d-glucose (UDP-Glc) and UDP-Dgalactose (UDP-Gal). Five members of the Chinese cabbage (Brassica rapa) UGE gene family, designated BrUGE1 to BrUGE5, have been cloned and characterized. Quantitative PCR shows that the BrUGE1and BrUGE4 mRNA are most abundant among other BrUGE genes, accounting for more than 55 % of total BrUGE transcripts in most of the tissues examined. All genes showed organ-specific expression pattern, two of which (BrUGE1 and 4) actively responded after Pectobacterium carotovorum subsp. carotovorum infection, while four genes (BrUGE-1, -3, -4, and -5)were shown to respond considerably against salt, drought and abscisic acid treatments. To better understand the function of the UGE gene, we constructed a recombinant pART vector carrying the BrUGE1 gene under the control of the CaMV 35S promoter and nos terminator and transformed using Agrobacterium tumefaciens. We then investigated BrUGE1 overexpressing rice lines at the physiological and molecular levels under biotic and abiotic stress conditions. Bioassay of T3 progeny lines of the transgenic plants in Yoshida solution containing 120 mM NaCl for 2 weeks, confirmed that the BrUGE1 enhances salt tolerance to transgenic rice plants. Also T3 progeny lines of the transgenic plants, when exposed to infection caused by Xanthomonas oryzae pv. oryzae, showed tolerance to bacterial blight. These results showed that BrUGE1 can be used as potential genetic resource for engineering Brassica with multiple stress resistance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant cell walls are chemically complex, diverse structures that are modified throughout the processes of cell division, cell growth, and differentiation, and in response to abiotic and biotic stresses. During the process of cell wall biosynthesis the different glycosyl transferases use varying nucleoside-diphosphate (NDP) sugars to assemble the different cell wall polysaccharides (Feingold and Avigad 1980). The availability of the NDP-sugars is thus an important factor in determining the final polysaccharide composition. Nevertheless, it is unclear to what extent modifications in the pool size of a particular NDP-sugar can affect the presence or structure of cell wall polysaccharides. Plants possess a sophisticated sugar biosynthetic machinery comprising families of nucleotide sugars that can be modified at their glycosyl moieties by nucleotide sugar interconversion enzymes to generate different sugars (Seifert 2004). UDP- glucose 4-epimerase (also UDP-galactose 4-epimerase, UGE, EC 5.1.3.2) catalyzes the interconversion of UDP-GIc and UDP-Gal (Majumdar et al. 2004). UGEs identified from plants lack transmembrane motifs and signal peptides and appear to exist as soluble entities in the cytoplasm (Barber et al. 2006). Generally, plant UDP-GIc epimerase enzymes are localized to the cytosol, where their substrates UDP-GIc and UDP-Gal are present at high levels (Seifert 2004; Pattathil et al. 2005). UDP-glucose is the most important of the NDP-sugars. It can be synthesized either from glucose 1-phosphate using the reaction NTP + sugar 1 − phosphate ↔ NDP-sugar + pyrophosphate or from sucrose using the enzyme sucrose synthase (Nakai et al. 1999). The gene encoding the UDP-glucose epimerase has been cloned from a variety of organisms (Dörmann and Benning 1998). In Arabidopsis thaliana, five UGEs were present and each UGE had different catalytic properties (Seifert 2004; Barber et al. 2006). AtUGE2 and AtUGE4 cooperate in providing cell wall biosynthesis and growth, AtUGE3 is specialized for pollen development, and AtUGE1 and AtUGE5 might act in stress situations in Arabidopsis (Mayer et al. 1999; Theologis et al. 2000; Rösti et al. 2007). In rice, four UGE genes (OsUGE1-4) were activated after drought, salt, or UV irradiation stress (Liu et al. 2007; Kim et al. 2009; Abdula et al. 2013; Joo et al. 2014). Indeed, drought stress induces increased expression of a UGE gene, which maps to a root thickness quantitative trait locus (QTL) region (Nguyen et al. 2004). Overexpression of OsUGE-1 altered raffinose level and tolerance to abiotic stress but not morphology in Arabidopsis (Liu et al. 2007). Similarly, two potato UGEs increased the galactose content of potato tuber cell walls (Oomen et al. 2004). The expression of the UGEs isolated from different plant species was found variable in response to different stresses. Until recently, no study has been reported about the UGE in Chinese cabbage, which is one of the important crops in Asia. This study investigated the UGE gene subfamily of Chinese cabbage. Five BrUGE genes belonging to the same family were identified from the Brassica database. cDNAs were isolated and expression profiling in different organs was analyzed following application of different biotic and abiotic stresses. In particular, this paper describes the expression analysis of these genes under different biotic and abiotic stresses and the overexpression of BrUGE1 gene in rice plants using the CaMV 35S promoter. The expression patterns and physiological effects in both overexpressing and wild-type plants under biotic and abiotic stress conditions were also investigated.
Materials and methods
Plant tissues sampling
Chinese cabbage (Brassica rapa ‘SUN-3061’) plants were grown at the Department of Horticulture, Hankyong National University, Korea. Fresh roots, stems, leaves, and flower buds of the Chinese cabbage were harvested, immediately frozen in liquid nitrogen, and then stored at −80 °C until subsequent analysis in an organ-specific expression study.
Abiotic stress treatments
Chinese cabbage seeds were aseptically grown on half-strength MS (HMS) agar medium in a culture room under a 16 h light photoperiod at 25 °C. After 3 weeks of growth, the seedlings were transferred to fresh liquid HMS medium containing 250 mM NaCl for salt stress, and 100 μM ABA. In addition, drought stress treatment was applied by keeping the seedling on filter paper for 24 h. For each stress, the leaf samples were collected after 0, 30 min, 1, 2, 4, 8, 12, and 24 h of treatment and each sample was collected three times using two individual plants.
Biotic stress treatments
Two of the most important pathogens that constantly challenge Chinese cabbage production worldwide include Fusarium oxysporum which causes wilt and Pectobacterium carotovorum which causes soft rot. Chinese cabbage (B. rapa ‘SUN-3061’) plants were infected with F. oxysporum f.sp. conglutinans at the Screening Center for Disease Resistant Vegetable Crops, Korea. The root-dip inoculation (RDI) method was used with some modifications to inoculate the cabbage and Chinese cabbage with the fungus (Ospina-Giraldo et al. 2003). Briefly, 3-week-old seedlings were removed from the soil and immersed in the conidial suspension. Samples were then collected from infected and mock-treated plants at 0, 3, 6, 24 h, 6, 8, and 11 days. The local (fourth) and systemic (fifth) leaves were harvested as samples. Chinese cabbage (B. rapa ‘SUN-3061’) plants were grown for 6 weeks under culture room conditions with 16 h light and 8 h dark maintaining 25 °C temperature prior to treatment. The P. carotovorum subsp. carotovorum stock (10 μl) was cultured in 25 ml of liquid YEP medium until OD600 = 1.4 equals 1,170,000 colony forming units (cfu) ml−1 and then diluted it to OD600 = 1.19 equals 1 × 106 CFU ml−1 by adding double-distilled water. For pathogen inoculation, 10 μl of P. carotovorum subsp. carotovorum culture solution (1 × 106 cfu ml−1) was added to the freshly needle wounded site (at the lower 1/3 position) of the midrib of the upper third leaves, and incubated at 25 °C covering with polyvinyl bags to maintain 80–90 % humidity. All inoculations were performed three times, and the infection was confirmed by observing disease lesion in the leaves of Chinese cabbage plants. About one-third parts from the top of the infected leaves were harvested for RNA extraction at 0, 6, 12, 24, and 72 h after inoculation. Upon collection of tissues for both biotic and abiotic treatments, the samples were immediately frozen in liquid nitrogen and stored at −80 °C until RNA isolation.
RNA extraction
Total RNA were extracted using the RNeasy mini kit (Qiagen, USA), after which it was treated with RNase-free DNase (Promega, USA) to remove genomic DNA contaminants. The cDNA was subsequently synthesized using a Superscript® III First-Strand synthesis kit (Invitrogen, USA) according to the manufacturer’s instructions.
Amino acid sequence analysis of BrUGE genes
We constructed a full-length cDNA library of B. rapa cv. Osome (Park et al., 2010) and obtained 3429 expressed sequence tags (ESTs). This dataset was subsequently analyzed for gene ontology and found 1017 ESTs functionally annotated to stress responses. Based on the cluster analysis of these stress responsive genes, 140 are related to biotic stresses, two of which (BrUGE1 and BrUGE2) were annotated as UDP-glucose 4-epimerase (UGE) proteins (unpublished). Using these two UGE proteins as reference, 3 more genes (BrUGE3 to BrUGE5) belonging to the same family were recovered from the Brassica database (http://brassicadb.org/brad/index.php). The primary structure of genes was analyzed using protParam (http://expasy.org/tools/protparam.html) and Augustus (http://augustus.gobics.de/submission). An alignment search was conducted using NCBI BLAST (http://www.ncbi.nlm.nih.gov/BLAST/) and the program BLASTp, with the “nr” database. Typical domains were analyzed using the EMBL web tool (http://smart.embl.de/smart/set.mode.cgiGENOMIC=1). Multiple protein sequences were aligned using PIR (http://pir.georgetown.edu/pirwww/search/multialn.shtml), and a phylogenetic tree was constructed according to the neighbor-joining method using the ClustalW program (http://www.genome.jp/tools/clustalw/) (Thompson et al. 1994). Accession numbers are as follows: from A. thaliana At1g12780, At4g23920, At1g63180, At1g64440, At4g10960 coded as AtUGE1 t0 5, respectively; XP_001698706 (Chlamydomonas reinhardtii CrUGE1); O65781 (Cyamopsis tetragonoloba CtUGE1); XP_643834 (Dictyostelium discoideum DdUGE1); NP_001035389 (zebrafish DaniorerioDrUGE1); Q14376 (Homo sapiens HsUGE1); from barley Hordeum vulgare, AAX49504, AAX49505, AAX49503 (HvUGE1–3); from Medicago truncatula, ACJ85116 and ACJ84690 (MtUGE1–2); BAF51705 (apple Malus × domestica MxdUGE1); OsUGE1 from rice (Oryza sativa), Os05g0595100, Os08g0374800, Os09g0526700, Os09g0323000 for OsUGE1–4; CAL54894 (Ostreococcus tauri OtUGE1); from Physcomitrella patens subsp. patens, XP_001768301, XP_001777464, XP_001775163, XP_001751529 for PpUGE1–4; and AAP68981 (Zea mays ZmUGE1).
Expression analysis
RT-PCR was conducted using an AMV one step RT-PCR kit (Takara, Japan). Specific primers for all the genes were used for RT-PCR, and actin primers of B. rapa (FJ969844) were used as a control (Table 1). PCR was performed using 50 ng of cDNA from the roots, leaves, stems and flower buds as templates in master mixes composed of 20 pmol of each primer, 150 μM of each dNTP, 1.2 U Taq polymerase, 1 × Taq polymerase buffer, and double-distilled H2O diluted to a total volume of 20 μl in 0.5 ml PCR tubes. The samples were then subjected to the following conditions: pre-denaturing at 94 °C for 5 min, followed by 30 cycles of denaturation at 94 °C for 30 s, annealing at 58 °C for 30 s and extension at 72 °C for 1 min, with a final extension for 5 min at 72 °C. Real-time quantitative PCR was performed using 1 μl of cDNA in a 25 μl reaction volume employing iTaqTM SYBR® Green Super-mix with ROX (California, USA). The specific primers used for real-time PCR are listed in Table 1. The conditions for real-time PCR were as follows: 10 min at 95 °C, followed by 40 cycles at 94 °C for 30 s, 58 °C for 30 s, and 72 °C for 45 s. The fluorescence was measured following the last step of each cycle, and three replications were used for each sample. Amplification, detection, and data analysis were conducted using a Rotor-Gene 6000 real-time rotary analyzer (Corbett Life Science, Australia).
Vector construction and plant transformation
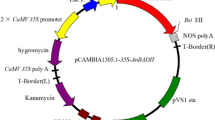
The full-length cDNA of BrUGE1 (Accession No. KF601691) was isolated by PCR from Brassica using the primers 5′-CCGCTCGAGATCAAGCTATGTCCGAGAAGG-3′ (XhoI site underlined) and 50-CGGGGTACCTCAGG CTGGTCTCTGCACATCTCCT-3′ (KpnI site underlined). The product was ligated into the pART vector (Gleave 1992). The recombinant vector carrying BrUGE1 was constructed under the control of the CaMV 35S promoter and nos terminator. The pART-UGE1 construct was electrophoresed into Agrobacterium tumefaciens LBA4404 and then introduced into pre-soaked rice seed using the method of Lee et al. (2011) with minor modifications (Supplemental Fig. 3).
PCR analysis of transgenic rice plants
Four-week-old rice seedlings were analyzed using PCR amplification to select transgenic lines. DNA was extracted from the leaves of the rice seedlings using the cetyltrimethyl ammonium bromide (CTAB) method (Rogers and Bendich 1994). The primer set for the PCR amplification and the probe for the Southern blot analysis were BrUGE-F2 and BrUGE1-R2. The PCR amplification profile consisted of an initial step at 94 °C for 5 min followed by 30 cycles of 94 °C for 1 min, 58 °C for 1 min, 72 °C for 2 min, and a final step at 72 °C for 10 min.
Expression analysis of transgenic lines
Total RNA from leaf tissue was extracted using Trizol reagent (Invitrogen, USA) according to the manufacturer’s protocol. The specific primers for the analysis of BrUGE1 expression of germinating seeds were BrUGE1-F3 5-ATGCCTTCTTCTGGAGATCC-3′ and BrUGE1-R3 5′-GTTCGTTGGACGATGAAGGT-3′. Real-time PCR was performed using a Bio-RAD I Cycler IQ5 machine as previously described using RT pre-mix (TOYOBO Co., Japan) (Ali-Benali et al. 2005). The threshold cycle (Ct) values of PCR reactions from three independent biological replicates were averaged and the relative quantification of the expression levels was performed using the comparative Ct method for all experiments (Livak and Schmittgen 2001). The fold change in total RNA of a target gene relative to the reference gene (actin gene) was determined by the following formula: fold change = 2−ΔΔCt, where ΔΔCt = (Ct target gene − Ct actin gene) transgenic plants − (Ct target gene − Ct actin gene) wild-type plants.
Bioassay for bacteria blight
Six-week-old plants and controls were inoculated with K3a (race 3) using the leaf-clipping method (Kauffman et al. 1973; Jung et al. 2012). Inoculum suspension density was about ~1 × 107 cells per milliliter. The inoculated plants were covered with polythene bag for 24 h and incubated under the following conditions: 4 and 23 h following humidity at night. Plant reaction to bacteria was scored on six leaves by measuring the lesion lengths 14 days after inoculation.
Statistical analysis
Data requiring statistical analysis were computed using the Statistix version 8 (http://www.statistix.com). Significant P values were analyzed using the two-sided Dunnett’s multiple comparisons with the WT Dongjin and MOCK as a reference.
Results
Identification and sequence analysis
In this study, we identified 5 UGE, designated B. rapa UDP-glucose 4-Epimerase (BrUGE 1–5). Gene sequences were analyzed, and are presented in Table 2. The size of the 5 BrUGEs varies from 343 to 351 amino acids (~37.7–39.1 kDa) and the predicted isoelectric points vary from 5.28 to 6.61 (Table 2). All BrUGEs contained the conserved β-NAD+ binding domain (GXXGXXG, where X is any amino acid) at the N terminal. The catalytic triad (YXXXK) conserved in UGEs (Weirenga et al. 1986) was found in all BrUGEs. Genomic DNA sequences of all BrUGE genes were isolated from the B. rapa chromosome sequences and the introns and exons were identified upon sequence analysis. Structural information pertaining to the 5 BrUGE genes is presented in Fig. 1. The sequence data revealed that the identified genes are UGEs which contain conserved β-NAD+ binding domain. For comparison of these BrUGEs with other published UGEs, an alignment search was carried out using a BLAST search of the NCBI database. The deduced amino acid sequences of 5 BrUGEs shared high homology, UGEs of A. thaliana and some other homologous species (Table 3). Amino acid sequence among these 5 BrUGEs of B. rapa was showed 60–80 % homology (Supplemental Fig. 1). We again retrieved 32 UGE sequences of different plant species from NCBI and a phylogenetic tree was constructed with the deduced amino acid sequences of 5 UGEs of B. rapa using the NJ method (Fig. 2). Results revealed two groups; BrUGE1 and 3 together with 16 UGEs from other crops formed Group I, and BrUGE2, 4 and 5 among other 14 UGEs fell under Group II. In addition, phylogenetic relationships of BrUGE1 with the enzymes from other origins including mammals, insects, fungi, and bacteria were analyzed (Fig. 2). Together with AtUGE1, AtUGE3, OsUGE3, and several other plant enzymes, BrUGE1 formed a plant subgroup apart from mammalian and bacterial enzymes.
Phylogenetic tree showing the relatedness of the deduced full-length amino acid sequences of 5 BrUGEs and 32 UGE proteins of other plant species. The phylogenetic tree was generated using ClustalW program. The species names and the GenBank protein accessions numbers are described in materials and methods. Enzymes shown to have high UDP-glucose 4-epimerase activity are boxed. The bootstrap values for the respective branches are shown. The bar indicates 0.1 substitution per site
Organ-specific expression analysis
Although all BrUGE genes showed considerable expressions in roots, stems, leaves, and flower buds, expression pattern of each gene seemingly occurred in an organ-specific manner. In BrUGE1, 2 and 3, transcripts were highest in the leaves. In BrUGE4, gene expression was highest in roots, while in BrUGE5, the expression peaked in flower buds. Among the organs tested, expression in stem was the lowest for BrUGE1, 3 and 5, whereas mRNA of BrUGE2 and 4 were very low in the flower buds (Fig. 3). It is also worth considering that overall expression of these genes was higher in leaves and roots than in stems and flower buds.
Real-time quantitative PCR expression analysis of 5 BrUGE genes in leaves, roots, flower buds, and stems of Brassica rapa plants were determined by qRT-PCR in leaves, roots, flower buds, and stems Brassica rapa “SUN-3061”SUN-3061′ plants. The error bars represent the standard error of the means of three independent replicates. Letters indicate significant differences between tissues for each gene
BrUGE expression in response to abiotic stresses
As UGEs have been shown to express in response to abiotic stress conditions (Rajam et al. 2007; Alam et al. 2014), responses of the 5 BrUGEs identified in this study, were investigated after salt, drought, and ABA stresses on seedling plants (Fig. 4). Among the 5 BrUGEs, four showed differential expression pattern. The genes that were upregulated throughout the whole observation time (0–24 h) include BrUGE1 and 5 during salt stress treatment, BrUGE4 and 5 in response to drought stress, and BrUGE4 in response to ABA stress. On the other hand, expression of BrUGE2 was down regulated during early observation time in three treatments. Specifically, expression response of this gene spiked only after 12 h of treatments both in drought and ABA stress, while it was only expressed mildly against salt stress. Notable strong expression at 12 h of ABA treatment was observed with BrUGE3, 4 and 5, and 12 h of drought condition with BrUGE2 and 5, as indicated by high intensity band (Fig. 4).
BrUGE genes expression under biotic stresses
In addition to abiotic stresses, responses of the 5 BrUGEs were also investigated against the infection of F. oxysporum f.sp. conglutinans in Chinese cabbage at various times after exposure. Here, no responsive expression was observed (Supplemental Fig. 2), indicating that the genes may not be effective against broad range of pathogens. To explore their possible role against phytobacteria, analysis of expression of 5 BrUGEs was performed after inoculating P. carotovorum subsp. Carotovorum, a necrotroph bacterium that causes soft rot, a serious problem in Brassica production. Expression of BrUGE1 and 4 were relatively higher among other BrUGEs, with levels that peaked at 6 h post inoculation (hpi). Low levels of expression were observed for BrUGE2,3, and 5, which initially showed an increasing pattern, reached the plateau at 6–24 h and which eventually declined until 72 h (Fig. 5). The BrUGE1 and 3 in Group I were responsive to the salt stress as sensitive markers, while there was no clear pattern regarding the response of three BrUGEs in Group II to various abiotic and biotic stresses except for drought stress where BrUGE4 and 5 were responsive.
Real-time quantitative PCR expression of 5 BrUGE genes in leaves of Brassica rapa ‘SUN-3061’ plants after infection with Pectobacterium carotovorum subsp. carotovorum. The error bars represent the standard error of the means of three independent replicates. Letters indicate significant differences between tissues for each gene
Characterization and expression of UGE1 transgenic rice
A total of 35 regenerated plants were analyzed by PCR. We confirmed that 86 % of the 35 T0 plants were positive with the UGE1 gene-specific primers. mRNA transcript analysis of CaMV 35S::BrUGE1 plants showed an enhanced expression of the BrUGE1 gene compared to that of the WT, Dongjin. Although the transgenic plants showed BrUGE1 overexpression, variability in the degree of expression was observed. Along with this, we selected five uniform lines with strong expression and named these as BrUGE1-OX5. These selected lines were used in all subsequent experiments. Furthermore, we investigated mRNA expression in various organs. BrUGE1-OX1-5 was selected for expression analysis in young leaves among the transgenic lines. The results showed that the BrUGE1 mRNA transcript was highly expressed in young seedlings except for BrUGE-OX1 and BrUGE-OX5 (Fig. 6). As expected, Dongjin (WT) also showed an mRNA transcription was not expressed in young seedling (Fig. 6). To examine NaCl treatment’s effect in UGE1-overexpressing (TG#1 and TG#2) and WT plants, 2-week-old seedlings were grown in hydroponics treated with 300 mM NaCl and supplied with nutrient solution. No significant differences were observed in the survival rates between the WT and transgenic plants, although yellowing of the leaf margin was observed in the WT plants after the NaCl treatment. When growth on media was monitored 7 days after planting, a growth reduction was noticed in all plants, including the WT, which showed a higher reduction than the BrUGE-OX plants (TG#1 and TG#2) (Fig. 7). The growth reduction of shoots was 80 % for WT and 30–45 % for TG#1 and TG#2 plants. In the case of the roots, the WT plants exhibited a 40 % growth reduction, whereas the TG#1 and TG#2 plants showed only 35 % reductions in growth. To analyze the resistance to pathogenic bacteria, TG#1, TG#2, and WT plants were inoculated with Xoo (Xanthomonas oryzae pv. oryzae, K3a). The K3a diluted with ~1 × 107 cultured in an LB liquid medium. Rice was infected with the three leaf stage. T3 progeny lines compared with WT and MOCK showed many symptoms noticeable against infection disease. Three weeks later, the leaves of WT and MOCK were significantly increased the size of the lesion to 8.3 and 8.8 cm, respectively, the transgenic lines (TG#1 and TG#2) showed a lesion of the low value to 5.8–6.8 cm, which indicated enhanced tolerance to bacterial blight disease (Fig. 8).
Identification of transferred gene in transgenic rice lines. PCR amplification using primer set of hygromycin (a) and BrUGE1 gene (b). c Expression analysis of BrUGE1 gene. The amplification products were separated using a 1.5 % agarose gel. Lane M DNA ladder, Lane P PCR product generated from the DNA template of the vector plasmid that contains the gene, WT Wild-type, Lane 1–5 independent transgenic lines
Salt tolerance assays of BrUGE1 transgenic rice. 3-week-old seedlings were watered with 250 mM NaCl solution for 24 h (a), and then recovered for 20 days (b). Survival rates of the WT and transgenic rice lines (c). WT; Dongjin rice, Mock; transferred only vector plasmid, TG #1–2; transgenic plant lines. The data are the mean ± SE of three independent experiments. The values with significant differences according to t tests are P < 0.05
Discussion
Enzyme catalyzing the reversible conversion/epimerization of UDP-glucose into UDP-galactose is the UDP-glucose 4-epimerase (UGE; EC.5.1.3.2) (Majumdar et al. 2004). The reversible epimerization is catalyzed via an enzyme-bound UDP-4-keto-hexose intermediate (Maitra and Ankel 1971). The gene encoding the UDP-glucose epimerase has been cloned from a variety of organisms (Dörmann and Benning 1998). UGE freely interconverts UDP-glucose and UDP-galactose, and a family of five UGE isoforms is encoded in the Arabidopsis genome. UGE1 to UGE5 show in vitro variations in substrate affinity, cofactor requirement, and metabolite inhibition profile (Barber et al. 2006). Their global expression pattern suggests that UGE1 and UGE3 are coregulated with carbohydrate catabolic enzymes, while UGE2, UGE4, and UGE5 are coregulated with carbohydrate biosynthetic enzymes. All isoforms can act in vivo in both directions, as indicated by overexpression experiments (Barber et al. 2006). Expression levels and experiments with UGE1 antisense lines both suggest that UGE1 might be the dominant isoform in green plant parts (Dörmann and Benning 1998). Antisense interference of UGE1, however, neither induces a morphological phenotype nor alters cell wall polymers or any other galactose-containing carbohydrate. This suggests that other isoforms are required for the biosynthesis of glycoconjugates. In rice, four UGE genes (OsUGE1-4) were activated after drought, salt or UV irradiation stress (Liu et al. 2007; Kim et al. 2009). Indeed, drought stress induces increased expression of a UGE gene, which maps to a root thickness QTL (quantitative trait locus) region (Nguyen et al. 2004). The 5 BrUGE genes isolated from B. rapa was showed specificity expression in different tissues, cluster with 2 groups (Fig. 2) and could play distinct catabolic functions for Arabidopsis (Barber et al. 2006). For example, as BrUGE2 and BrUGE4 are very homologous to AtUGE2 and AtUGE4 of Arabidopsis (Rösti et al. 2007). The transcriptional analysis in different tissues and after different stresses can provide clues about the in vivo functions of BrUGEs. UDP-sugars are sugar donors for various UDP-dependent glycosyltransferase (UGTs). UGTs from plants are highly specific for UDP-sugars, but microbial UGTs have broad UDP-sugar ranges (Vogt and Jones 2005). In addition, depending on the sugar, the physical and biological activities of some antibiotics and flavonoids are changed (Hollman et al. 1999). Thus, expression analysis of BrUGE genes during biotic and abiotic stresses supports the idea that PR-proteins belong to a general plant stress response pathway rather than being specific to distinct stresses, as often hypothesized (Van Loon et al. 2006). The BrUGE1 and 3 in Group I showed to associate with tolerance to salt stress, while there was no clear pattern regarding the response of three BrUGE genes in Group II to various abiotic and biotic stresses except for drought stress where BrUGE4 and 5 were responsive (Figs. 3, 4). Recent studies showed that UGEs are involved in tolerance against abiotic stresses (Liu et al. 2007; Kim et al. 2009). Overexpression of OsUGE1 in A. thaliana increased the content of raffinose, which resulted in tolerance against abiotic stresses (Liu et al. 2007). The increased level of UDP-galactose resulted in an increased level of raffinose because raffinose is synthesized from UDP-galactose and sucrose (Pridham and Hassid 1965). Transgenic rice overexpressing a putative UDP-galactose epimerase from Paspalum vaginatum also showed tolerance against salt (Endo et al. 2005). To better understand the function of the BrUGE1 gene, we constructed a recombinant pART vector carrying the BrUGE1 gene under the control of the CaMV 35S promoter and nos terminator and transformed using A. tumefaciens. We then investigated BrUGE1 overexpressing rice lines at the physiological and molecular levels under biotic and abiotic stress conditions (Figs. 7, 8). Bioassay of T3 progeny lines of the transgenic plants in Yoshida solution containing 120 mM NaCl for 2 weeks, confirmed that the BrUGE1 enhances salt tolerance to transgenic rice plants (Fig. 7). Also T3 progeny lines of the transgenic plants, when exposed to infection caused by Xanthomonas oryzae pv. oryzae, showed enhanced tolerance to bacterial blight (Fig. 8). These results showed that BrUGE1 can be used as potential genetic resource for engineering Brassica with multiple stress resistance.
References
Abdula SE, Lee HJ, Jee MG, Jung YJ, Kang KK, Nou IS, Lee SB, Yang WH, Cho YG (2013) Development and identification of transgenic rice lines with abiotic stress tolerance by using a full-length overexpression gene hunting system. Plant Breed Biotechnol 1:33–48
Alam MM, Nahar K, Hasanuzzaman M, Fujita M (2014) Exogenous jasmonic acid modulates the physiology, antioxidant defense and glyoxalase systems in imparting drought stress tolerance in different Brassica species. Plant Biotechnol Rep 8:279–293
AliBenali MA, Alary R, Joudrier P, Gautier MF (2005) Comparative expression of five lea genes during wheat seed development and in response to abiotic stresses by real-time quantitative RT-PCR. Biochim Biophys Acta 1730:56–65
Barber C, Rösti J, Rawat A, Findlay K, Roberts K, Seifert GJ (2006) Distinct properties of the five UDP-d-glucose/UDP-D-galactose 4-epimerase isoforms of Arabidopsis thaliana. J Biol Chem 281:17276–17285
Dörmann P, Benning C (1998) The role of UDP-glucose epimerase in carbohydrate metabolism of Arabidopsis. Plant J 13:641–652
Endo N, Yoshida K, Akiyoshi M, Yoshida Y, Hayashi N (2005) Putative UDP-galactose epimerase and metallothionein of Paspalum vaginatum enhanced the salt tolerance of rice, Oryza sativa L. from transplanting to harvest stages. Breed Sci 55:163–173
Feingold DS, Avigad G (1980) Sugar nucleotide transformations in plants. In: Stumpf PK, Conn EE (eds) The biochemistry of plants: a comprehensive treatise, vol 3. Academic Press, New York, pp 101–170
Gleave AP (1992) A versatile binary vector system with a T-DNA organizational-structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20:1203–1207
Hollman P, Buysman MP, van Gameren Y, Cnossen E, de Vries J, Katan MB (1999) The sugar moiety is a major determinant of the absorption of dietary flavonoid glycosides in man. Free Radic Res 3:569–573
Joo JS, Lee YH, Song SI (2014) Overexpression of the rice basic leucine zipper transcription factor OsbZIP12 confers drought tolerance to rice and makes seedling hypersensitive to ABA. Plant Biotechnol Rep 8:431–441
Jung YJ, Lee SY, Moon YS, Kang KK (2012) Enhanced resistance to bacterial and fungal pathogens by overexpression of a human cathelicidin antimicrobial peptide (hCAP18/LL-37) in Chinese cabbage. Plant Biotechnol Rep 6:39–46
Kauffman HE, Reddy APK, Hsieh SPY, Merca SD (1973) An improved technique for evaluating resistance of rice varieties to Xanthomonas oryzae. Plant Dis Rep 57:537–541
Kim SK, Kim DH, Kim BG, Jeon YM, Hong BS, Ahn JH (2009) Cloning and characterization of the UDP glucose/galactose epimerases of Oryza sativa. J Korean Soc Appl Biol Chem 52(4):315–320
Lee HJ, Abdula SE, Jee MG, Jang DW, Cho YG (2011) High-efficiency and rapid Agrobacterium-mediated genetic transformation method using germinating rice seeds. J Plant Biotechnol 38:251–257
Liu H, Da X, Xu Y, Chong K (2007) Over-expression of OsUGE-1 altered raffinose level and tolerance to abiotic stress but not morphology in Arabidopsis. J Plant Physiol 164:1384–1390
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408
Maitra US, Ankel H (1971) Uridine diphosphate-4-keto-glucose, an intermediate in the uridine diphosphate-galactose 4-epimerase reaction. Proc Natl Acad Sci USA 68:2660–2663
Majumdar S, Ghatak J, Mukherji S, Bhattacharjee H, Bhaduri A (2004) UDP galactose 4-epimerase from Saccharomyces cerevisiae: a bifunctional enzyme with aldose 1-epimerase activity. Eur J Biochem 271:753–759
Mayer K, Schuller C, Wambutt R, Murphy G, Volckaert G, Pohl T, Dusterhoft A, Stiekema W, Entian KD, Terryn N et al (1999) Sequence and analysis of chromosome 4 of the plant Arabidopsis thaliana. Nature 402:769–777
Nakai T, Tonouchi N, Konishi T, Kojima Y, Tsuchida T, Yoshinaga F, Sakai F, Hayashi T (1999) Enhancement of cellulose production by expression of sucrose synthase in Acetobacter xylinum. Proc Natl Acad Sci USA 96:14–18
Nguyen TT, Klueva N, Chamareck V, Aarti A, Magpantay G, Millena AC, Pathan MS, Nguyen HT (2004) Saturation mapping of QTL regions and identification of putative candidate genes for drought tolerance in rice. Mol Genet Genomics 272:35–46
Oomen RJFJ, Bang DT, Tzitzikas EN, Bakx EJ, Schols HS, Visser RGF (2004) Overexpression of two different potato UDP-Glc 4-epimerases can increase the galactose content of potato tuber cell walls. Plant Sci 166:1097–1104
Ospina-Giraldo MD, Mullins E, Kang S (2003) Loss of function of the Fusarium oxysporum SNF1 gene reduces virulence on cabbage and Arabidopsis. Curr Genet 44:49–57
Park JI, Kumar TS, Ahmed NU, Nou IS (2010) Construction of full-length cDNA library and investigation of potential genes in Brassica rapa. J Basic Life Res Sci 10:6–11
Pattathil S, Harper AD, Bar-Peled M (2005) Biosynthesis of UDP-xylose: characterization of membrane-bound AtUxs2. Planta 221:538–548
Pridham JB, Hassid WZ (1965) Biosynthesis of raffinose. Plant Physiol 40:984–986
Rajam MV, Chandola N, Goud P, Singh D, Kashyap V, Choudhary ML, Sihachakr D (2007) Thaumatin gene confers resistance to fungal pathogens as well as tolerance to abiotic stresses in transgenic tobacco plants. Biol Plant 51:135–141
Rogers S, Bendich A (1994) Extraction of total cellular DNA from plants, algae and fungi. In: Gelvin SB, Schilperoort S (eds) Plant molecular biology manual. Section D. Springer, Netherlands, pp 183–190
Rösti J, Barton CJ, Albrecht S, Dupree P, Pauly M, Findlay K, Roberts K, Seifert GJ (2007) UDP-glucose 4-epimerase isoforms UGE2 and UGE4 cooperate in providing UDP-galactose for cell wall biosynthesis and growth of Arabidopsis thaliana. Plant Cell 19:1565–1579
Seifert GJ (2004) Nucleotide sugar interconversions and cell wall biosynthesis: how to bring the inside to the outside. Curr Opin Plant Biol 7(3):277–284
Theologis A, Ecker JR, Palm CJ, Federspiel NA, Kaul S, White O, Alonso J, Altafi H, Araujo R, Bowman CL et al (2000) Sequence and analysis of chromosome 1 of the plant Arabidopsis thaliana. Nature 408:816–820
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Van Loon LC, Rep M, Pieterse CM (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Vogt T, Jones P (2005) Glycosyltransferases in plant natural product synthesis: characterization of a supergene family. Trends Plant Sci 5:380–386
Weirenga RK, Terpstra P, Hol WGJ (1986) Prediction of the occurrence of the ADP-binding beta alpha beta fold in proteins, using an amino acid sequence fingerprint. J Mol Biol 187:101–107
Acknowledgments
This research was supported by Golden Seed Project (Center for Horticultural Seed Development, No. 213003-04-1-SBC10), Ministry of Agriculture, Food and Rural Affairs (MAFRA), Ministry of Oceans and Fisheries (MOF), Rural Development Administration (RDA) and Korea Forest Service (KFS).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.

11816_2015_370_MOESM1_ESM.jpg
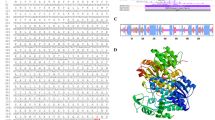
Supplemental Fig. 1. Alignment of deduced amino acid sequences of UGE (UDP-glucose 4-epimerase) proteins of phylogenetic tree using the Clustal W program in multiple sequence alignment mode. The residues conserved between all eight UGEs are marked with asterisks, and those conserved between at least five UGEs are marked with colons. The GGXGXXG motif involved in NAD + -binding is underlined, and a stretch of amino acid residues encompassing the catalytic site (including the completely conserved YXXXK motif) is doubly boxed. Supplementary material 1 (JPEG 258 kb)

11816_2015_370_MOESM2_ESM.jpg
Supplemental Fig. 2. Real-time quantitative PCR expression analysis of 5 UGE (UDP-glucose 4-epimerase) genes in Brassica rapa after infection with Fusarium oxysporum f.sp. conglutinans. The error bars represent the standard error of the means of three independent replicates. Supplementary material 2 (JPEG 306 kb)

11816_2015_370_MOESM3_ESM.jpg
Supplemental Fig. 3. Introduction of BrUGE1 gene through Agrobacterium-mediated transformation. A; seeding in the N6D media, B; callus growing after inoculation. C; shoot formation and elongation, D; root formation and elongation.. Supplementary material 3 (JPEG 42 kb)
Rights and permissions
About this article
Cite this article
Jung, Y.J., Kyoung, J.H., Nou, I.S. et al. Molecular characterization of the UDP-glucose 4-epimerase (BrUGE) gene family in response to biotic and abiotic stress in Chinese cabbage (Brassica rapa). Plant Biotechnol Rep 9, 339–350 (2015). https://doi.org/10.1007/s11816-015-0370-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11816-015-0370-7