Abstract
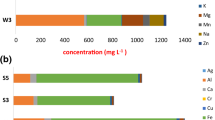
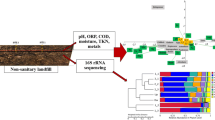
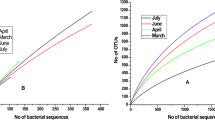
Eleven acid mine drainage (AMD) samples were obtained from southeast of China for the analysis of the microbial communities diversity, and the relationship with geochemical variables and spatial distance by using a culture-independent 16S rDNA gene phylogenetic analysis approach and multivariate analysis respectively. The principle component analysis (PCA) of geochemical variables shows that eleven AMDs can be clustered into two groups, relative high and low metal rich (RHMR and RLMR) AMDs. Total 1 691 clone sequences are obtained and the detrended correspondence analysis (DCA) of operational taxonomic units (OTUs) shows that, γ-Proteobacteria, Acidobacteria, Actinobacteria, Cyanobacteria, Firmicutes and Nitrospirae are dominant species in RHMR AMDs. In contrast, α-Proteobacteria, β-Proteobacteria, Planctomycetes and Bacteriodetes are dominant species in RLMR AMD. Results also show that high-abundance putative iron-oxidizing and only putative sulfur-oxidizing microorganisms are found in RHMR AMD. Multivariate analysis shows that both geochemical variables (r=0.429 3, P=0.037 7) and spatial distance (r=0.321 3, P=0.018 1) are significantly positively correlated with microbial community and pH, Mg, Fe, S, Cu and Ca are key geochemistry factors in shaping microbial community. Variance partitioning analysis shows that geochemical variables and spatial distance can explain most (92%) of the variation.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
BAKER B J, BANFIELD J F. Microbial communities in acid mine drainage [J]. Fems Microbiol Ecol, 2003, 44(2): 139–152.
GERHARDT A, DE BISTHOVEN L J, GUHR K, SOARES A M V M, PEREIRA M J. Phytoassessment of acid mine drainage: Lemna gibba bioassay and diatom community structure [J]. Ecotoxicology, 2008, 17(1): 47–58.
TREMAINE S C, MILLS A L. Impact of water column acidification on protozoan bacterivory at the lake sediment-water interface [J]. Appl Environ Microb, 1991, 57(3): 775–784.
DSA J V, JOHNSON K S, LOPEZ D, KANUCKEL C, TUMLINSON J. Residual toxicity of acid mine drainage-contaminated sediment to stream macroinvertebrates: Relative contribution of acidity vs. metals [J]. Water Air Soil Poll, 2008, 194(1/2/3/4): 185–197.
LEVINGS C D, BARRY K L, GROUT J A, PIERCEY G E, MARSDEN A D, COOMBS A P, MOSSOP B. Effects of acid mine drainage on the estuarine food web, Britannia Beach, Howe Sound, British Columbia, Canada [J]. Hydrobiologia, 2004, 525(1/2/3): 185–202.
NORDSTROM D K, ALPERS C N. Negative pH, efflorescent mineralogy, and consequences for environmental restoration at the Iron Mountain Superfund site, California [J]. Proc Natl Acad Sci USA, 1999, 96(7): 3455–3462.
TIWARY R K. Environmental impact of coal mining on water regime and its management [J]. Water Air Soil Poll, 2001, 132(1/2): 185–199.
JOHNSON D B, HALLBERG K B. Acid mine drainage remediation options: A review [J]. Sci Total Environ, 2005, 338(1/2): 3–14.
VALENZUELA L, CHI A, BEARD S, ORELL A, GUILIANI N, SHABANOWITZ J, HUNT D F, JEREZ C A. Genomics, metagenomics and proteomics in biomining microorganisms [J]. Biotechnol Adv, 2006, 24(2): 197–211.
CID C, GARCIA-DESCALZO L, CASADO-LAFUENTE V, AMILS R, AGUILERA A. Proteomic analysis of the response of an acidophilic strain of Chlamydomonas sp (Chlorophyta) to natural metal-rich water [J]. Proteomics, 2010, 10(10): 2026–2036.
KNOB A, CARMONA E C. Purification and characterization of two extracellular xylanases from penicillium sclerotiorum: A novel acidophilic xylanase [J]. Appl Biochem Biotech, 2010, 162(2): 429–443.
WAN Ming-xi, YANG Yu, QIU Guan-zhou, XU Ai-ling, QIAN Lin, HUANG Zhi-ying, XIA Jin-lan. Acidophilic bacterial community reflecting pollution level of sulphide mine impacted by acid mine drainage [J]. Journal of Central South University of Technology, 2009, 16(2): 223–229.
RAMETTE A, TIEDJE J M. Multiscale responses of microbial life to spatial distance and environmental heterogeneity in a patchy ecosystem [J]. Proc Natl Acad Sci USA, 2007, 104(8): 2761–2766.
HORNER-DEVINE M C, LAGE M, HUGHES J B, BOHANNAN B J M. A taxa-area relationship for bacteria [J]. Nature, 2004, 432(7018): 750–753.
MARTINY J B H, BOHANNAN B J M, BROWN J H, COLWELL R K, FUHRMAN J A, GREEN J L, HORNER-DEVINE M C, KANE M, KRUMINS J A, KUSKE C R, MORIN P J, NAEEM S, OVREAS L, REYSENBACH A L, SMITH V H, STALEY J T. Microbial biogeography: Putting microorganisms on the map [J]. Nat Rev Microbiol, 2006, 4(2): 102–112.
ZHOU Ji-zhong, KANG S, SCHADT C W, GARTEN C T. Spatial scaling of functional gene diversity across various microbial taxa [J]. Proc Natl Acad Sci USA, 2008, 105(22): 7768–7773.
XIE Jian-ping, HE Zhi-li, LIU Xin-xing, VAN NOSTRAND J D, DENG Ye, WU Li-you, ZHOU Ji-zhong, QIU Guan-zhou. GeoChip-based analysis of the functional gene diversity and metabolic potential of microbial communities in acid mine drainage [J]. Appl Environ Microbiol, 2011, 11(3): 991–999.
QIU Guan-zhou, WANG Jun. Bio-leaching of low-grade large porphyry chalcopyrite-containing ore [J]. Transactions of Nonferrous Metals Society of China, 2000, 10(s1): 19–22.
ZHOU Ji-zhong, BRUNS M A, TIEDJE J M. DNA recovery from soils of diverse composition [J]. Appl Environ Microb, 1996, 62(2): 316–322.
AHN S J, COSTA J, EMANUEL J R. PicoGreen quantitation of DNA: Effective evaluation of samples pre- or post-PCR [J]. Nucleic Acids Res, 1996, 24(13): 2623–2625.
SCHLOSS P D, HANDELSMAN J. Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness [J]. Appl Environ Microb, 2005, 71(3): 1501–1506.
HUMAYOUN S B, BANO N, HOLLIBAUGH J T. Depth distribution of microbial diversity in Mona Lake, A mermictic soda lake in California [J]. Appl Environ Microb, 2003, 69(2): 1030–1042.
TERBRAAK C J F. Canoco-An extension of decorana to analyze species-environment relationships [J]. Vegetatio, 1988, 75(3): 159–160.
VAN NOSTRAND J D, WU Wei-min, WU Li-you, DENG Ye, CARLEY J, CARROLL S, HE Zhi-li, GU Bao-hua, LUO Jian, CRIDDLE C S, WATSON D B, JARDINE P M, MARSH T L, TIEDJE J M, HAZEN T C, ZHOU Ji-zhong. GeoChip-based analysis of functional microbial communities during the reoxidation of a bioreduced uranium-contaminated aquifer [J]. Environ Microbiol, 2009, 11(10): 2611–2626.
MANTEL N. Detection of disease clustering and a generalized regression approach [J]. Cancer Res, 1967, 27(2p1): 209–220.
HOTELLING H. Relations between two sets of variates [J]. Biometrika, 1936, 28(3/4): 321–377.
OKLAND R H, EILERTSEN O. Canonical correspondence-analysis with variation partitioning-Some comments and an application [J]. J Veg Sci, 1994, 5(1): 117–126.
YIN Hua-qun, CAO Lin-hui, XIE Ming, CHEN Qi-jiong, QIU Guan-zhou, ZHOU Ji-zhong, WU Li-you, WANG Dian-zuo, LIU Xue-duan. Bacterial diversity based on 16S rRNA and gyrB genes at Yinshan mine, China [J]. Syst Appl Microbiol, 2008, 31(4): 302–311.
MATLOCK M M, HOWERTON B S, ATWOOD D A. Chemical precipitation of heavy metals from acid mine drainage [J]. Water Res, 2002, 36(19): 4757–4764.
SCHMIDT A, HAFERBURG G, SINERIZ M, MERTEN D, BUCHEL G, KOTHE E. Heavy metal resistance mechanisms in actinobacteria for survival in AMD contaminated soils [J]. Chem Erde-Geochem, 2005, 65(S1): 131–144.
DINELLI E, TATEO F. Different types of fine-grained sediments associated with acid mine drainage in the Libiola Fe-Cu mine area (Ligurian Apennines, Italy) [J]. Appl Geochem, 2002, 17(8): 1081–1092.
ALLEN E E, BANFIELD J F. Community genomics in microbial ecology and evolution [J]. Nat Rev Microbiol, 2005, 3(6): 489–498.
JOHNSON D B. Biodiversity and ecology of acidophilic microorganisms [J]. Fems Microbiol Ecol, 1998, 27(4): 307–317.
YIN Hua-qun, QIU Guan-zhou, WU Li-you, XIE Ming, ZHOU Ji-zhong, DAI Zhi-min, WANG Dian-zuo, KELLOGG L, CAO Lin-hui, LIU Xue-duan. Microbial community diversity and changes associated with a mine drainage gradient at the Dexing copper mine, China [J]. Aquat Microb Ecol, 2008, 51(1): 67–76.
EDWARDS K J, GIHRING T M, BANFIELD J F. Seasonal variations in microbial populations and environmental conditions in an extreme acid mine drainage environment [J]. Appl Environ Microb, 1999, 65(8): 3627–3632.
SINGER P C, STUMM W. Acidic mine drainage: The rate-determining step [J]. Science, 1970, 167(3921): 1121–1127.
FINNEY L A, O’HALLORAN T V. Transition metal speciation in the cell: Insights from the chemistry of metal ion receptors [J]. Science, 2003, 300(5621): 931–936.
Author information
Authors and Affiliations
Corresponding author
Additional information
Foundation item: Project(2010CB630901) supported by the National Basic Research Program of China; Project(50621063) supported by Creative Research Group of China; Projects(51104189, 50321402, 50774102) supported by the National Natural Science Foundation of China; Project (1343-77341) supported by the Graduate Education Innovative Program of Central South University, China; Project(DOE-ER64125) supported by the Department of Energy, Office of Science under the Environmental Remediation Science Program of USA
Rights and permissions
About this article
Cite this article
Xie, Jp., Jiang, Hc., Liu, Xx. et al. 16s rDNA based microbial diversity analysis of eleven acid mine drainages obtained from three Chinese copper mines. J. Cent. South Univ. Technol. 18, 1930–1939 (2011). https://doi.org/10.1007/s11771-011-0925-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11771-011-0925-x