Abstract
Our previous work revealed that the soybean GmsSOS1 enhances salt tolerance in Arabidopsis. In this work, we studied the physiological mechanisms by which the GmsSOS1 confers salt and oxidative stress tolerance in Arabidopsis and yeast cells. Under salt stress condition, the GmsSOS1-expressing Arabidopsis plants displayed larger leaf area, lower leaf relative electrolytic leakage, less accumulation of H2O2, superoxide anion radicals (O2 −), and malondialdehyde compared with wild type. In consistent with these observations, the activities of antioxidant enzymes catalase, ascorbate peroxide, and peroxidase in the GmsSOS1-expressing plants were higher than those in wild type under salt stress. Combined salt and oxidative stresses caused more damage and higher accumulation of H2O2 and Na+ than single stress condition in both wild type and the GmsSOS1-expressing plants. However, the GmsSOS1-expressing Arabidopsis plants could maintain significantly lower levels of H2O2 and Na+ and exhibited better growth than wild type under either single or combined stress. The GmsSOS1 complemented the yeast plasma membrane-localized Na+/H+ antiporter and enhanced salt tolerance by reducing Na+ accumulation in yeast cells. Our results suggest that the soybean GmsSOS1 can alleviate the primary Na+ toxicity by limiting Na+ accumulation and mitigate the secondary oxidative stress through improving antioxidant enzyme activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Soil salinization is an increasingly problem for the irrigated lands, and high salinity in soil adversely affects plant growth and development thus reduces crop quality and yield (Yadav et al. 2012; Ma et al. 2014a, b; Volkov 2015). Salt stress causes primary injuries, including osmotic effects at an early phase and ionic toxicity at a later phase of plant growth (Sanadhya et al. 2015). This is often followed by secondary damages involving nutritional imbalance and oxidative stress due to accumulation of reactive oxygen species (ROS), such as O ·−2 and H2O2 (Manchanda and Garg 2008; de Oliveira et al. 2013; He et al. 2015). However, plants have evolved diverse mechanisms to mitigate the damaging effects of salt stress through morphological, physiological, biochemical, and molecular adjustments (Qiu et al. 2014; Schmidt et al. 2013). In soil, sodium (Na+) is a common soluble ion deleterious to plants, including all major crops. Na+ is not an essential ion for most plant growth, thus excessive Na+ outside of or accumulation in plant cells results in disturbance or imbalance of intracellular osmotic, ionic, and oxidative homeostasis (Adem et al. 2014). Therefore, preventing Na+ accumulation and maintaining appropriate K+/Na+ ratio in the cytoplasm, which are largely regulated via Na+ transporters, are crucial for plant survival and growth in saline environments (Xue et al. 2014; Liu et al. 2015; Sanadhya et al. 2015; Volkov 2015; Wu et al. 2015).
The SOS (salt-overly sensitive) pathway consisting of three components, the plasma membrane-localized Na+/H+ antiporter SOS1, the cytoplasmic protein kinase SOS2, and the Ca2+ sensor SOS3, is the most extensively studied salt tolerance mechanisms in plants (Ma et al. 2014a; Zhou et al. 2014). The A. thaliana SOS1 protein (AtSOS1) is the first described plasma membrane-localized Na+/H+ antiporter mediating Na+ efflux and controlling long-distance Na+ transport from roots to shoots (Shi et al. 2002). Oh et al. (2010) further revealed the roles of SOS1 in vacuolar morphology, pH homeostasis, and membrane trafficking, in addition to conferring salt tolerance in roots during early stages of salt stress. Up to now, a number of SOS1 homologous genes have been identified and functionally analyzed from different plant species, including halophytes, such as Thellungiella salsuginea, Populus euphratica, Mesembryanthemum crystallinum, Salicornia europaea, Suaeda salsa and Sesuvium portulacastrum, xerophyte as Zygophyllum xanthoxylum and crop plants, such as rice (Oryza sativa), tomato (Solanum lycopersicum), wheat (Triticum aestivum and Triticum durum), and soybean (Glycine max and Glycine soja) (Ma et al. 2014b; Martínez-Atienza et al. 2007; Olías et al. 2009a; Feki et al. 2014; Zhou et al. 2015; Nie et al. 2015). Transgenic studies suggest that SOS1 as an Na+ efflux transporter could increase salt tolerance in different plant species (Quintero et al. 2002; Yadav et al. 2012; Feki et al. 2014).
SOS1 mRNA in Arabidopsis is inherently instable at normal growth conditions, but its stability is elevated by salt or oxidative stress (Chung et al. 2008; Jiang and Shi 2008). ROS signals were shown to be required for the regulation of Na+/H+ antiporter activity (Zhou et al. 2014). Interestingly, mutations in the gene render atsos1 mutants more tolerant to oxidative stress caused by the treatment with MV (methyl viologen), indicating that AtSOS1 may play a negative role in oxidative stress tolerance (Chung et al. 2008). In addition, under salt or oxidative stress, AtSOS1 could interact through its long cytoplasmic C-terminal tail with RCD1 (Radical-Induced Cell Death1, a regulator of oxidative stress responses), thus the cross-talk between ion homeostasis and oxidative stress response might be involved in plant salt tolerance (Katiyar-Agarwal et al. 2006). Our previous work has shown that the plasma membrane Na+/H+ antiporter gene (named GmsSOS1) from G. max, G. soja, and their hybrid conferred salt tolerance in the Arabidopsis atsos1-1 mutant and wild-type plants (Nie et al. 2015). In this work, we studied the physiological mechanisms of salt tolerance conferred by the GmsSOS1 gene in Arabidopsis and yeast cells by examining Na+ and ROS accumulation and ROS scavenging enzymes. We found that GmsSOS1 not only conferred salt tolerance, but also elevated oxidative stress tolerance in the transgenic plants. Notably, we observed that oxidative stress treatments promoted Na+ accumulation under salt stress condition, while GmsSOS1 expression mitigated such enhanced Na+ accumulation, which suggests a possible link between oxidative stress and Na+ influx and/or efflux transport systems. Our results provided further evidence on the important role of the plasma membrane-localized Na+/H+ antiporter in salt and oxidative stress tolerance in plants.
Materials and methods
Plant materials, growth conditions, and stress treatments
A. thaliana (Columbia glabrous1 ecotype) wild type (WT) and GmsSOS1-expressing line (T3 generation, Nie et al. 2015) were used in this study. For seed germination test, seeds were surface-sterilized and stratified at 4 °C for 2–4 days. The seeds were then germinated and grown on MS medium supplemented with 3% sucrose (w/v) and 0.8% agar with or without 150 mM NaCl. After growth for 7 days, seed germination rate was calculated by counting the percentage of cotyledon greening (Li et al. 2015), 30 seeds per petri plate each with three replications. For stress response at seedling stage, surface-sterilized and stratified seeds were sowed in soil consisting of sterilized peat moss and vermiculite mixture in plastic pots (diameter = 7 cm), cultured with 1/2 Hoagland nutrition solution in a growth chamber (16 h light (22 ± 2 °C)/8 h dark (18 ± 2 °C) cycle at 60–70% relative humidity). The 5-week-old seedlings were divided into two groups for different treatments: one group was used for single salt stress (150 mM NaCl in 1/2 Hoagland nutrition solution) treatment and the other group was used for salt stress (120 mM NaCl), oxidative (3 µM MV or 4 mM H2O2), and combined stress (120 mM NaCl + 3 µM MV or 120 mM NaCl + 4 mM H2O2 in 1/2 Hoagland solution, respectively). The seedlings watered with 1/2 Hoagland nutrition solution only were used as the controls. After 7 days of treatments, the seedlings were photographed and sampled for measurements of leaf area and relative electrolytic leakage (REL), contents of O ·−2 and malondialdehyde (MDA), activities of antioxidant enzymes [catalase (CAT), peroxidase (POD), ascorbate peroxide (APX), and superoxide dismutase, i.e., SOD] in leaves and roots and for DAB (3,5-diaminobenzidine) staining of leaves or plants.
Measurements of leaf area and REL
Leaf area was measured by photographing the leaf samples which were then analyzed using ImageJ according to Maloof et al. (2013). REL in leaves or shoots was assayed as described by Tian et al. (2014). Briefly, the fresh samples (0.5 g) were added to a tube with 20 mL of deionized water, and the conductance of the deionized water (C W), the leaf tissues before boiling (C 1), and the leaf tissue after boiling (C 2) was measured using a conductance meter (DDS-307, Shanghai, China). REL was calculated as (C 1 − C W)/(C 2 − C W).
Determination of O ·−2 and MDA contents
O ·−2 content was measured by following the method by Gajewska and Sklodowska (2007) with some modifications. Fresh leaf and root tissues (0.5 g) were homogenized in 3 mL of 65 mM phosphate buffer (pH 7.8) and then centrifuged at 10,000g for 10 min. The supernatant (0.5 mL) was mixed with 0.5 mL of 65 mM phosphate buffer (pH 7.8) and 0.1 mL of 10 mM hydroxylamine chlorhydrate, and the mixture was incubated at 25 °C for 1 h. 1 mL of 58 mM sulfanilamide and 1 mL of 7 mM α-naphthylamine were then added, and the mixture was further incubated at 25 °C for 20 min. The absorbance was measured at 530 nm, and O ·−2 contents were calculated from a standard curve of NaNO2.
MDA content was measured according to the method described by Jouve et al. (2007). A total of 0.5 g of fresh materials was ground in 5 mL of 5% (w/v) trichloroacetic acid (TCA), and 2 mL of supernatant was mixed with 2 mL of 0.67% (w/v) thiobarbituric acid (TBA) in 5% (w/v) TCA and incubated at 100 °C for 30 min. After centrifuging, the optical density was measured at 450, 532, and 600 nm, respectively. The amount of MDA was calculated from the following formula: C = 6.45(A 532–A 600) − 0.56A 450, C represents the concentration of MDA in supernatant and expressed as µmol L−1, A 532, A 600, and A 450 represent the absorbance values at 532, 600, and 450 nm, respectively. The final MDA content was expressed as nmol g−1 FW.
Assay of antioxidant enzyme activities
Enzyme extractions were performed according to the method of Meng et al. (2014). Fresh shoots and roots (0.5 g) were homogenized in a mortar and pestle with 2 mL of 50 mM ice-cold phosphate buffer (pH 7.0) containing 1 mM EDTA Na2 and 0.5% PVP (w/v). The homogenate was centrifuged at 12,000g for 20 min at 4 °C. The supernatant was used as the enzyme extracts for assays of APX, CAT, POD, and SOD activities. CAT activity was assayed according to Aebi (1984). The 3-mL reaction mixture contained 0.1 mL of enzyme extracts, 0.1 M phosphate buffer (pH 7.0), deionized water, and 20 mM H2O2. The decomposition of H2O2 was measured by following the decrease in absorbance at 240 nm for 3 min and quantified by its molar extinction coefficient (39.4 mmol L−1 cm−1). One unit of CAT activity was defined as a change in absorbance of 0.1 units min−1 caused by addition of the enzyme extracts. POD activity was measured with guaiacol as the substrate according to Nakano and Asada (1981). The reaction mixture (3 mL) consisted of 100 mM sodium acetate buffer (pH 5.4), 10 mM guaiacol solution, 0.1 mM H2O2, and 10 µL enzyme extracts. The increase in absorbance due to oxidation of guaiacol was measured at 470 nm for 1 min. One unit of POD activity was defined as a change in absorbance of 0.1 unit min−1. SOD activity was assayed using the photochemical nitroblue tetrazolium (NBT) method (Beauchamp and Fridovich 1971). The reaction mixture contained 100 mM phosphate buffer (pH 7.8), 130 mM methionine, 750 μM NBT, 20 μM riboflavin, 1 mM EDTA Na2, deionized water, and 80 μL enzyme extracts in a 3 mL volume. One unit of SOD activity was defined as the amount of enzyme required to cause 50% inhibition of NBT reduction monitored at 560 nm. APX activity was assayed in a reaction mixture of 3 mL containing 100 mM phosphate (pH 7.0), 100 mM ascorbic acid, 0.1 mM H2O2, deionized water, and 0.1 mL enzyme extracts. The reaction was started by adding enzyme extracts to the mixture. Enzyme activity was quantified by following the decrease in absorbance at 290 nm for 3 min. One unit of APX activity was defined as an absorbance change of 0.01 units min−1 (Nakano and Asada 1981).
DAB staining
H2O2 in plant tissues was detected in situ by DAB staining as described by Li and Cai (2015). The leaves or plants were vacuum-infiltrated with a solution of DAB (1 mg mL−1, pH 3.8), then placed in a 5-cm erlenmeyer flask wrapped with aluminum foil, and incubated for 6 h at 100 rpm in a shaker. After discarding the DAB staining solution, clearing solution (ethanol:acetic acid:glycerol = 3:1:1) was added to the samples and incubated in a boiling water bath (95 °C) for 15 min, and the clearing process was repeated twice. The DAB stained leaves or plants were then photographed. Each sample with at least ten leaves or six plants was conducted.
Measurements of Na+ and K+ contents
Roots and shoots of seedlings under NaCl, oxidative, or combined stress were fully rinsed in distilled water, fixed at 105 °C for 5 min, and were dried to constant weight at 80 °C. Dry matter was ground and screened with a 60-mesh sieve, then 100 mg of each sample was added to a tube (25 mL) and 20 mL of deionized water was added. Samples were boiled for 2–3 h, then filtered and deionized water was added to make a final volume of 50 mL. K+ and Na+ contents were estimated using a flame spectrophotometer (AP1200 type, Shanghai Aopu Analytical Instrument Co., China), and measurements were calibrated using NaCl or KCl solutions of known concentrations (Wei et al. 2015).
GmsSOS1 expression in yeast mutant and salt treatment test
Yeast (Saccharomyces cerevisiae) strain G19 (∆ena1::HIS3::ena4), and its mutants ANT3 (∆ena1::HIS3::ena4, ∆nha1::LEU2) and GX3 (∆ena1::HIS3::ena4, ∆nhx1::TRP1) have been described elsewhere (Song et al. 2012). The entire soybean GmsSOS1 open reading frame (Nie et al. 2015) was cloned into the yeast expression vector pYES2 under the control of the constitutive promoter GAL1 (Halley et al. 2010). The yeast expression vector containing GmsSOS1 gene was, respectively, transformed into NHA1- or NHX1-deleted yeast mutant strains ena1 nha1 or ena1 nhx1 using PEG/LiAc procedure and transformants were identified by PCR-based method (Gietz 2014). The yeast mutant ena1 was used as a positive control. Ten-fold serial dilutions (starting at OD550 ≈0.5) of each sample were plated on YPD (1% yeast extract, 2% peptone, and 2% dextrose), YPG (1% yeast extract, 2% peptone, and 2% galactose) medium, or YPG medium supplemented with hygromycin B (HygB) (50 mg L−1) or arginine-phosphate medium (AP medium: 10 mM l-arginine, 8 mM H3PO4, 2 mM MgSO4, 0.2 mM CaCl2, 2% glucose, vitamins and trace elements, and pH 6.5) plus NaCl (70 mM) and KCl (1 mM), and the plates were incubated at 28 °C for 60 h and then photographed (Song et al. 2012). For Na+ and K+ content measurements, the yeast cells were grown in the liquid AP medium plus NaCl (70 mM) and KCl (1 mM) and collected at exponential growth phase (OD550 ≈0.2) (Quintero et al. 2002). The Na+ and K+ contents in the yeast cells were determined using a flame spectrophotometer as above-described. Each strain was repeated with three independent cultures.
Statistical analysis
Data were expressed as mean ± SD of at least three replications besides the above special instructions, and were analyzed using one-way analysis of variance (ANOVA) with Duncan’s multiple range test at the P < 0.05 level of significance in the SPSS 19.0 software.
Results
Expression of GmsSOS1 in Arabidopsis enhances seed germination and seedling growth under salt stress
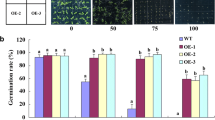
When seeds of WT and GmsSOS1-expressing A. thaliana were grown in MS agar medium without NaCl, the percentages of seed cotyledon greening in both genotypes exceeded 90% and did not show significant difference (P > 0.05). When germinated in MS agar medium plus 150 mM NaCl, the percentages of seed cotyledon greening of both genotypes were remarkably declined as compared to the controls, while the cotyledon greening of GmsSOS1-expressing Arabidopsis was significant higher than that of WT (P < 0.05) (Fig. 1a). When the seedlings grown in soil were treated with 150 mM NaCl solution for 7 days, the growth of both genotypes was visibly inhibited, accompanied with significant decrease in leaf area and increase in REL. However, the salt-induced changes in leaf area and REL in GmsSOS1-expressing Arabidopsis were less than those in WT (Fig. 1b-left, c, d). DAB staining is a common method to visualize in situ H2O2 accumulation in tissues as a reddish-brown precipitate (Fujibe et al. 2004). DAB staining indicated a lower accumulation of H2O2 in the leaves of GmsSOS1-expressing plants than that of WT after salt stress treatment (Fig. 1b-right).
Growth response of GmsSOS1-expressing and WT plants to salt stress. a Effects of 150 mM NaCl treatment on the percentage of cotyledon greening. b Growth phenotype of 5-week-old seedlings after salt stress treatment for 7 days (left panel) and DAB staining of the leaves (right panel, brown color indicates H2O2 accumulation). c Leaf area. d REL
Expression of GmsSOS1 in Arabidopsis reduces O −·2 accumulation and MDA content under salt stress
When cultured under normal conditions, the contents of O2 − and MDA in leaves and roots of WT and GmsSOS1-expressing plants did not show significant differences (P > 0.05). After treated with 150 mM NaCl solution for 7 days, the contents of O −.2 and MDA in leaves of WT increased 2.71 and 2.31 times of the controls, respectively (P < 0.05), while O −.2 and MDA contents in leaves of GmsSOS1-expressing plants were only 1.18 and 1.35 times increases when compared with the controls. In roots, salt stress treatment resulted in a slight elevation of MDA content in wild type but not in GmsSOS1-expressing plants. Salt stress did not cause changes in O −.2 contents in roots of both WT and GmsSOS1-expressing plants (Fig. 2).
Expression of GmsSOS1 in Arabidopsis elevates the activities of antioxidant enzymes under salt stress
When cultured under non-stress condition, the activities of CAT, POD, SOD, and APX did not show significant difference between WT and GmsSOS1-expressing seedlings (P > 0.05) (Fig. 3). When exposed to 150 mM NaCl for 7 days, the activities of CAT in leaves and roots and APX in leaves of WT and GmsSOS1-expressing seedlings were remarkably decreased in comparison with the control plants, but the decreases in GmsSOS1-expressing seedlings were significantly lower than those in WT. The GmsSOS1-expressing seedlings displayed 1.51, 1.71, and 1.97 times higher activities of CAT in leaves and roots and APX in leaves than those in WT. Under salt stress, the POD activity in leaves of GmsSOS1-expressing seedlings increased significantly (P < 0.05) when compared with that in WT. The SOD activity in GmsSOS1-expressing seedlings did not increase when compared with WT under normal growth conditions or after salt treatment (Fig. 3). Overall, stronger activity of antioxidant enzymes could be maintained in GmsSOS1-expressing Arabidopsis seedlings than WT under saline condition.
Effects of NaCl, oxidative and combined stress on growth and H2O2 accumulation in WT and GmsSOS1-expressing Arabidopsis
Under salt treatment with 120 mM NaCl for 7 days, the growth of WT seedlings was more inhibited and the seedlings displayed more severe salt stress phenotypes, such as yellowish and pigmented leaves than GmsSOS1-expressing seedlings. Treatment with MV (3 µM) or H2O2 (4 mM) caused weaker growth inhibition and stress-injury symptoms than NaCl treatment in both genotypes, but GmsSOS1-expressing seedlings displayed higher tolerance to these stress conditions than WT. A combination of salt and oxidative stress treatment (NaCl plus MV or H2O2) for 7 days resulted in more severe growth inhibition and stress damage than the single stress treatment in both genotypes, while the GmsSOS1-expressing seedlings performed visibly better than WT (Fig. 4a, upper panel). Improved tolerance to salt and oxidative stress by GmsSOS1 was also evidenced by lower REL in the shoots of GmsSOS1-expressing seedlings than in WT (Fig. 4b). DAB staining indicated lower H2O2 accumulation in GmsSOS1-expressing seedlings than in WT under single (NaCl, MV, or H2O2) or combined stress conditions (Fig. 4a-lower panel).
Effects of 120 mM NaCl, oxidative stress (3 µM MV or 4 mM H2O2), and the combined stress (120 mM NaCl + 3 µM MV or 120 mM NaCl + 4 mM H2O2) treatment for 7 days on plant growth (a, upper panel), ROS accumulation (DAB staining, a, lower panel), and REL in shoots (b) of 5-week-old GmsSOS1-expressing and WT seedlings
Effects of NaCl, oxidative, and combined stress on Na+ and K+ contents in WT and GmsSOS1-expressing Arabidopsis
NaCl treatment increased the Na+ contents in both leaves and roots of both genotypes, but GmsSOS1-expressing seedlings accumulated significantly lower Na+ than WT. Oxidative stress (MV or H2O2) also caused a slight increase in Na+ content, and GmsSOS1-expressing shoots had lower Na+ content than WT after H2O2 treatment. Interestingly, the combined treatments (NaCl plus MV or H2O2) significantly increased Na+ accumulation than NaCl treatment alone, and GmsSOS1-expressing seedlings accumulated less Na+ than WT under these combined stress conditions (Fig. 5a, b). In addition, greater Na+ reductions in GmsSOS1-expressing shoots than roots under NaCl or NaCl-oxidative combined stress were observed, which may account for improved salt tolerance of GmsSOS1-expressing plants. Overall, GmsSOS1-expressing plants did not show significantly changes in K+ contents when compared with WT under single or combined stress conditions, except that H2O2 treatment resulted in a slight increase in K+ content in the shoots of GmsSOS1-expressing plants (Fig. 5c, d).
GmsSOS1 complements the yeast NHA1
Yeast (S. cerevisiae) cells possess the plasma membrane Na+/H+ antiporter NHA1 for Na+ efflux and the vacuolar membrane Na+/H+ antiporter NHX1 for Na+ sequestration and conferring HygB resistance (Shi et al. 2002). To determine the capacity of GmsSOS1 to substitute for yeast NHA1 or NHX1 antiporters, mutant complementation tests were performed. Yeast mutant strains ena1, ena1 nha1 and ena1 nhx1 could grow normally in YPD or YPG medium, even when transformed with GmsSOS1 (+GmsSOS1). When cultured in the AP medium plus 70 mM NaCl and 1 mM KCl, both ena1 nha1 and ena1 nhx1 mutants exhibited more growth inhibition than ena1 mutant. Heterologous expression of GmsSOS1 in ena1 nha1 or ena1 nhx1 restored cell growth in the presence of NaCl to the level equivalent to that mediated by the native NHA1 and NHX1 antiporters (strain ena1), but it could not rescue the sensitivity of ena1 nhx1 to hygromycin B (Fig. 6a). This result indicates that GmsSOS1 imparts specific tolerance to Na+ through complementing the functions of the plasma membrane Na+/H+ antiporter NHA1. Na+ and K+ measurements revealed that GmsSOS1 overexpression in ena1 nha1 cells reduced cellular Na+ accumulation. While GmsSOS1 overexpression in ena1 nhx1 did not significantly change the cellular Na+ contents (Fig. 6b). GmsSOS1 overexpression in the yeast mutants did not affect cellular K+ accumulation (Fig. 6c).
Complementation of the yeast Na+/H+ antiporter NHA1 by GmsSOS1. a Cells of S. cerevisiae strains ena1, ena1 nha1, ena1 nhx1, and ena1 nha1and ena1 nhx1 transformed with GmsSOS1 (+GmsSOS1) were grown in YPD, YPG, or YPG medium supplemented with HygB (50 mg L−1) or AP medium plus NaCl (70 mM) and KCl (1 mM). Five microliters of serial decimal dilutions were spotted onto agar plates of the same medium. Plates were incubated at 28 °C and photographed after 60 h. b, c Cells of the above-mentioned ena1, ena1 nha1, ena1 nhx1 (control), or GmsSOS1-expressing cells (+GmsSOS1) were grown in liquid AP medium with NaCl (70 mM) and KCl (1 mM). When cultures reached OD550 ≈0.2, cells were collected by filtration, and the Na+ and K+ contents were determined. Data shown are the average ± SD of ion contents of three independent cultures of each strain
Discussion
As an important member of the major salt tolerance SOS pathway, SOS1 has also been implicated in ROS detoxification or ROS-mediated signaling initiated by salt stress (Katiyar-Agarwal et al. 2006; Chung et al. 2008; Jiang and Shi 2008; Zhou et al. 2014). There are a number of research works showing that overexpression of SOS1 in various plant species could confer salt tolerance by reducing Na+ in roots, or retrieving Na+ from the xylem or recirculating from leaves of transgenic plants (Olías et al. 2009a, b; Shi et al. 2003). Post-transcriptional gene silencing of ZxSOS1 gene demonstrated that ZxSOS1 is important not only in long-distance transport and spatial distribution of Na+ and K+, but also in maintaining Na+, K+ homeostasis in salt-treated Z. xanthoxylum plants (Ma et al. 2014b). In this study, the GmsSOS1-expressing Arabidopsis showed clear alleviation in salt injury symptoms and enhanced salt tolerance when compared with WT (Figs. 1, 4). Salt tolerance conferred by GmsSOS1 could be attributed to reduced Na+ accumulation and maintaining ion homeostasis in the transgenic plants under salt stress (Fig. 5). We also observed that GmsSOS1-expressing Arabidopsis seedlings had reduced accumulation of O −.2 and H2O2 (Figs. 2a, 4a), which is coincided with stronger activities of antioxidant enzymes, such as APX, CAT, and POD (Fig. 3). Lower ROS (including H2O2 and O −.2 ) levels may account for reduced damages in GmsSOS1-expression plants under salt stress.
To determine whether GmsSOS1 provided not only salt tolerance but also oxidative stress tolerance, we compared the responses of GmsSOS1-expressing and WT plants to oxidative stress (MV or H2O2) and a combined salt-oxidative stress. Our results indicated that GmsSOS1-expressing plants grew better than WT especially under the combined stress conditions, which is supported by lower REL in the GmsSOS1-expressing plants (Fig. 4). Better performance of GmsSOS1-expressing plants is consistent with reduced Na+ accumulation in shoots and roots of the transgenic plants. Interestingly, oxidative stress treatments (MV or H2O2) resulted in increased accumulation of Na+, and expression of GmsSOS1 alleviated the elevation of Na+ contents caused by oxidative stress (Fig. 5a, b). These results suggest that ROS somehow target and inactivate the Na+ efflux transporters such as the plasma membrane Na+/H+ antiporter SOS1 thus enhances Na+ accumulation in roots and shoots, while overexpression of GmsSOS1 could compensate the plasma membrane Na+/H+ antiporter activity thus reduce Na+ accumulation in the plants. Katiyar-Agarwal et al. (2006) and Chung et al. (2008) have documented the implications of SOS1 in oxidative stress, and suggested that SOS1 and RCD1 might be important in controlling apoplastic ROS accumulation under salt stress and that physical interaction between the cytoplasmic C-terminal tail of plasma membrane-located SOS1 and the nucleus-localized RCD1 could bridge the cross-talk between ionic homeostasis and oxidative stress pathway in plants exposed to salt or oxidative stress. When compared with Arabidopsis WT, sos1, and rcd1 mutant plants were more sensitive to salt stress and apoplastic ROS imposed by H2O2, but more tolerant to chloroplastic ROS imposed by MV (Katiyar-Agarwal et al. 2006). When soybean GmsSOS1 was compared with counterpart genes of the glycophyte Arabidopsis thaliana (AtSOS1, GenBank accession No. HE802928.1) and its close relative and halophytic species, T. halophila (ThSOS1, GenBank accession No. EF207775.1), 68.5 and 65.4% identity were shared, respectively, but 87.1% was displayed between AtSOS1 and ThSOS1. This may indicate that GmsSOS1 is somewhat different with AtSOS1 functioned under oxidative stress (including apoplastic and chloroplastic ROS). This intriguing observation deserves further investigation.
GmsSOS1 expression in the yeast mutants defective of NHA1 or NHX1 did not affect cellular K+ uptake, but significantly reduced Na+ accumulation and restored cell growth of the NHA1 defective mutant under salt stress (Fig. 6). This result further supports that GmsSOS1 functionally complements the yeast plasma membrane Na+/H+ antiporter NHA1. AtSOS1 overexpression in the yeast mutants defective of NHA1 or NHX1 resulted in not only reduction in the Na+ content but also improvement in the K+ status (Quintero et al. 2002; Shi et al. 2002), which is not fully consistent with our work on GmsSOS1-overexpressing yeast mutants. However, AtSOS1-overexpressed yeast mutant cells lacking TRK1 and TRK2 proteins (both comprising the high-affinity K+ uptake system of S. cerevisiae) failed to restore the growth in low external K+ condition, which suggests the role of AtSOS1 on K+ uptake is indirect (Quintero et al. 2002). The previous works have indicated that AtSOS1 may play a negative role in oxidative stress tolerance of Arabidopsis plants (Chung et al. 2008). Salt cress (T. halophila) is a close relative of Arabidopsis but very tolerant to high salinity. The SOS1 homologous gene in salt cress (ThSOS1) is expressed at high levels even in the absence of salt stress, and compared to Arabidopsis, salt cress is also more tolerant to oxidative stress (Taji et al. 2004). In this study, GmsSOS1-overexpressing Arabidopsis plants displayed enhanced salt and oxidative stress tolerance than WT. Certainly, GmsSOS1 gene possesses phylogenetic background different or distant from AtSOS1 or ThSOS1 (Gao et al. 2016), and the capacity of salt tolerance of soybean (Zhang et al. 2011) is between salt cress and Arabidopsis. Thus, the physiological variations between the transgenic Arabidopsis plants harboring GmsSOS1 and AtSOS1, and whether these two homologous genes execute their functions in the Arabidopsis genome in the same or different ways, are worthy to be studied in the future.
Salt tolerance is a complex trait involving multiple genes. Classical and molecular-assisted breeding towards salt tolerance in crops have not been very successful due to the complex feature of soil salinity and plant adaptation to saline environments (Zhang and Shi 2013). However, several key players, such as the SOS genes, have been identified and well characterized in plants. Natural variations of these salt tolerance genes could provide tools for crop improvement. For example, the SOS1 variant genes in soybean varieties and wild soybean species may display distinct capability for salt tolerance. Alternatively, the SOS1 variant genes may have different expression patterns due to evolutionary adaptation to saline environments. Both alternations in the coding regions and regulatory promoters could be utilized for improvement of salt tolerance in soybean by classical or molecular breeding.
In conclusion, compared with WT, GmsSOS1-expressing Arabidopsis exhibited reduced salt injury and enhanced salt and oxidative stress tolerance not only by reducing Na+ accumulation, but also by modulating the antioxidant enzyme activities and maintaining relatively lower ROS levels in plant roots and shoots. Heterologous expression of GmsSOS1 in the yeast mutant defective of NHA1 reduced Na+ accumulation and increased salt tolerance of the mutant cells. Our results further demonstrate that soybean or other plant SOS1 gene can be considered as a superior salt tolerance determinant that could be utilized in molecular breeding for creating salt tolerant crop varieties.
Author contribution statement
Xiufang Zhao, Peipei Wei, and Zhen Liu conducted the experiments, collected, and analyzed the data. BingjunYu and Huazhong Shi designed the experiments. Bingjun Yu, Huazhong Shi, and Xiufang Zhao interpreted the data and wrote the manuscript. All authors read and approved the final version of the manuscript and have no conflict of interest.
References
Adem GD, Roy SJ, Zhou M, Bowman JP, Shabala S (2014) Evaluating contribution of ionic, osmotic and oxidative stress components towards salinity tolerance in barley. BMC Plant Biol 14:113
Aebi H (1984) Catalase in vitro. Methods Enzymol 105:121–126. doi:10.1016/S0076-6879(84)05016-3
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44:276–287. doi:10.1016/0003-2697(71)90370-8
Chung J-S, Zhu J-K, Bressan RA, Hasegawa PM, Shi H (2008) Reactive oxygen species mediate Na+-induced SOS1 mRNA stability in Arabidopsis. Plant J 53:554–565. doi:10.1111/j.1365-313X.2007.03364.x
de Oliveira AB, Alencar NL, Gomes-Filho E (2013) Comparison between the water and salt stress effects on plant growth and development. InTech Open. doi:10.5772/54223
Feki K, Quintero FJ, Khoudi H, Leidi EO, Masmoudi K, Pardo JM, Brini F (2014) A constitutively active form of a durum wheat Na+/H+ antiporter SOS1 confers high salt tolerance to transgenic Arabidopsis. Plant Cell Rep 33:277–288. doi:10.1007/s00299-013-1528-9
Fujibe T, Saji H, Arakawa K, Yabe N, Takeuchi Y, Yamamoto KT (2004) A methyl viologen-resistant mutant of Arabidopsis, which is allelic to ozone-sensitive rcd1, is tolerant to supplemental ultraviolet-B irradiation. Plant Physiol 134:275–285. doi:10.1104/pp.103.033480
Gajewska E, Sklodowska M (2007) Effect of nickel on ROS content and antioxidative enzyme activities in wheat leaves. Biometals 20:27–36. doi:10.1007/s10534-006-9011-5
Gao J, Sun J, Cao P, Ren L, Liu C, Chen S, Chen F, Jiang J (2016) Variation in tissue Na+ content and the activity of SOS1 genes among two species and two related genera of Chrysanthemum. BMC Plant Biol 16:98. doi:10.1186/s12870-016-0781-9
Gietz RD (2014) Yeast transformation by the LiAC/SS carrier DNA/PEG method. In: Xiao W (ed) Yeast protocols, methods in molecular biology, vol 1163, pp 33–44. doi:10.1007/978-1-4939-0799-1_4
Halley JE, Kaplan T, Wang AY, Kobor MS, Rine J (2010) Roles for H2A.Z and its acetylation in GAL1 transcription and gene induction, but not GAL1-transcriptional memory. PLoS Biol 8(6):e1000401. doi:10.1371/journal.pbio.1000401
He Y, Fu J, Yu C, Wang X, Jiang Q, Hong J, Lu K, Xue G, Yan C, James A, Xu L, Chen J, Jiang D (2015) Increasing cyclic electron flow is related to Na+ sequestration into vacuoles for salt tolerance in soybean. J Exp Bot 66:6877–6889. doi:10.1093/jxb/erv392
Jiang J, Shi H (2008) Signaling control of SOS1 mRNA stability. Plant Signal Behav 3:687–688
Jouve L, Jacques D, Douglas GC, Hoffmann L, Hausman J-F (2007) Biochemical characterization of early and late bud flushing in common ash (Fraxinus excelsior L.). Plant Sci 172:962–969. doi:10.1016/j.plantsci.2007.02.008
Katiyar-Agarwal S, Zhu J, Kim K, Agarwal M, Fu X, Huang A, Zhu J-K (2006) The plasma membrane Na+/H+ antiporter SOS1 interacts with RCD1 and functions in oxidative stress tolerance in Arabidopsis. PNAS 103:18816–18821. doi:10.1073/pnas.0604711103
Li J, Cai W (2015) A ginseng PgTIP1 gene whose protein biological activity related to Ser128 residue confers faster growth and enhanced salt stress tolerance in Arabidopsis. Plant Sci 234:74–85. doi:10.1016/j.plantsci
Li ZQ, Li JX, Li HJ, Shi ZH, Zhang GF (2015) Overexpression of TsApx1 from Thellungiella salsuginea improves abiotic stress tolerance in transgenic Arabidopsis thaliana. Biol Plant 2015(59):497–506. doi:10.1007/s10535-015-0533-y
Liu M, Wang T-Z, Zhang W-H (2015) Sodium extrusion associated with enhanced expression of SOS1 underlies different salt tolerance between Medicago falcata and Medicago truncatula seedlings. Environ Exp Bot 110:46–55. doi:10.1016/j.envexpbot
Ma D-M, Xu W-R, Li H-W, Jin F-X, Guo L-N, Wang J, Dai H-J, Xu X (2014a) Co-expression of the Arabidopsis SOS genes enhances salt tolerance in transgenic tall fescue (Festuca arundinacea Schreb.). Protoplasma 251:219–231. doi:10.1007/s00709-013-0540-9
Ma Q, Li Y-X, Yuan H-J, Hu J, Wei L, Bao A-K, Zhang J-L, Wang S-M (2014b) ZxSOS1 is essential for long-distance transport and spatial distribution of Na+ and K+ in the xerophyte Zygophyllum xanthoxylum. Plant Soil 374:661–676. doi:10.1007/s11104-013-1891-x
Maloof JN, Nozue K, Mumbach MR, Palmer CM (2013) LeafJ: an ImageJ plugin for semi-automated leaf shape measurement. J Vis Exp 71:e50028. doi:10.3791/50028
Manchanda G, Garg N (2008) Salinity and its effects on the functional biology of legumes. Acta Physiol Plant 30:595–618. doi:10.1007/s11738-008-0173-3
Martínez-Atienza J, Jiang X, Garciadeblas B, Mendoza I, Zhu J-K, Pardo JM, Quintero FJ (2007) Conservation of the salt overly sensitive pathway in rice. Plant Physiol 143:1001–1012. doi:10.1104/pp.106.092635
Meng J-F, Xu T-F, Wang Z-Z, Fang Y-L, Xi Z-M, Zhang Z-W (2014) The ameliorative effects of exogenous melatonin on grape cuttings under water-deficient stress: antioxidant metabolites, leaf anatomy, and chloroplast morphology. J Pineal Res 57:200–212. doi:10.1111/jpi.12159
Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22:867–880
Nie W, Xu L, Yu B (2015) A putative soybean GmsSOS1 confers enhanced salt tolerance to transgenic Arabidopsis sos1-1 mutant. Protoplasma 252:127–134. doi:10.1007/s00709-014-0663-7
Oh DH, Lee SY, Bressan RA, Yun DJ, Bohnert HJ (2010) Intracellular consequences of SOS1 deficiency during salt stress. J Exp Bot 61:1205–1213. doi:10.1093/jxb/erp391
Olías R, Eljakaoui Z, Li J, De Morales PA, Marín-Manzano MC, Pardo JM, Belver A (2009a) The plasma membrane Na+/H+ antiporter SOS1 is essential for salt tolerance in tomato and affects the partitioning of Na+ between plant organs. Plant Cell Environ 32:904–916. doi:10.1111/j.1365-3040.2009.01971.x
Olías R, Eljakaoui Z, Pardo JM, Belver A (2009b) The Na+/H+ exchanger SOS1 controls extrusion and distribution of Na+ in tomato plants under salinity conditions. Plant Signal Behav 4:973–976
Qiu ZB, Guo JL, Zhu AJ, Zhang L, Zhang MM (2014) Exogenous jasmonic acid can enhance tolerance of wheat seedlings to salt stress. Ecotox Environ Safe 104:202–208. doi:10.1016/j.ecoenv
Quintero FJ, Ohta M, Shi H, Zhu J-K, Pardo JM (2002) Reconstitution in yeast of the Arabidopsis SOS signaling pathway for Na+ homeostasis. PNAS 99:9061–9066. doi:10.1073/pnas.132092099
Sanadhya P, Agarwal P, Agarwal PK (2015) Ion homeostasis in a salt-secreting halophytic grass. AoB Plants 7:plv055. doi:10.1093/aobpla/plv055
Schmidt R, Mieulet D, Hubberten H-M, Obata T, Hoefgen R, Fernie AR, Fisahn J, Segundo BS, Guiderdoni E, Schippers JHM, Mueller-Roeber B (2013) Salt-responsive ERF1 regulates reactive oxygen species–dependent signaling during the initial response to salt stress in rice. Plant Cell 25:2115–2131. doi:10.1105/tpc.113.113068
Shi H, Quintero FJ, Pardo JM, Zhu Jian-Kang (2002) The putative plasma membrane Na+/H+ antiporter SOS1 controls long-distance Na+ transport in plants. Plant Cell 14:465–477. doi:10.1105/tpc.010371
Shi H, Lee BH, Wu SJ, Zhu JK (2003) Overexpressing of a plasma membrane Na+/H+ antiporter gene improves salt tolerance in Arabidopsis thaliana. Nat Biotechnol 21:81–85. doi:10.1038/nbt766
Song A, Lu J, Jiang J, Chen S, Guan Z, Fang W, Chen F (2012) Isolation and characterisation of Chrysanthemum crassum SOS1, encoding a putative plasma membrane Na+/H+ antiporter. Plant Biol 14:706–713. doi:10.1111/j.1438-8677.2011.00560.x
Taji T, Seki M, Satou M, Sakurai T, Kobayashi M, Ishiyama K, Narusaka Y, Narusaka M, Zhu J-K, Shinozaki K (2004) Comparative genomics in salt tolerance between Arabidopsis and Arabidopsis-related halophyte salt cress using Arabidopsis microarray. Plant Physiol 135:1697–1709. doi:10.1104/pp.104.039909
Tian F, Jia TJ, Yu BJ (2014) Physiological regulation of seed soaking with soybean isoflavones on drought tolerance of Glycine max and Glycine soja. Plant Growth Regul 74:229–237. doi:10.1007/s10725-014-9914-z
Volkov V (2015) Salinity tolerance in plants. Quantitative approach to ion transport starting from halophytes and stepping to genetic and protein engineering for manipulating ion fluxes. Front Plant Sci 6:873. doi:10.3389/fpls.2015.00873
Wei P, Chen D, Jing R, Zhao C, Yu B (2015) Ameliorative effects of foliar methanol spraying on salt injury to soybean seedlings differing in salt tolerance. Plant Growth Regul 75:133–141. doi:10.1007/s10725-014-9938-4
Wu H, Shabala L, Liu X, Azzarello E, Zhou M, Pandolfi C, Chen Z-H, Bose J, Mancuso S, Shabala S (2015) Linking salinity stress tolerance with tissue-specific Na+ sequestration in wheat roots. Front Plant Sci 6:71
Xue Z, Zhao S, Gao H, Sun S (2014) The salt resistance of wild soybean (Glycine soja Sieb. et Zucc. ZYD 03262) under NaCl stress is mainly determined by Na+ distribution in the plant. Acta Physiol Plant 36:61–70. doi:10.1007/s11738-013-1386-7
Yadav NS, Shukla PS, Jha A, Agarwal PK, Jha B (2012) The SbSOS1 gene from the extreme halophyte Salicornia brachiata enhances Na+ loading in xylem and confers salt tolerance in transgenic tobacco. BMC Plant Biol 12:188
Zhang JL, Shi H (2013) Physiological and molecular mechanisms of plant salt tolerance. Photosynth Res 115:1–22
Zhang XK, Zhou QH, Cao JH, Yu BJ (2011) Differential Cl−/salt tolerance and NaCl-induced alternations of tissue and cellular ion fluxes in Glycine max, Glycine soja and their hybrid seedlings. J Agron Crop Sci 197:329–339
Zhou H, Lin H, Chen S, Becker K, Yang Y, Zhao J, Kudla J, Schumaker KS, Guo Y (2014) Inhibition of the Arabidopsis salt overly sensitive pathway by 14-3-3 proteins. Plant Cell 26:1166–1182. doi:10.1105/tpc.113.117069
Zhou Y, Yin X, Duan R (2015) SpAHA1 and SpSOS1 coordinate in transgenic yeast to improve salt tolerance. PLoS One 10(9):e0137447. doi:10.1371/journal.pone.0137447
Acknowledgements
This work was funded by the National Natural Science Foundation of China (Nos. 30871462, U1603111).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by H Peng.
X. Zhao and P. Wei contributed equally to this work.
Rights and permissions
About this article
Cite this article
Zhao, X., Wei, P., Liu, Z. et al. Soybean Na+/H+ antiporter GmsSOS1 enhances antioxidant enzyme activity and reduces Na+ accumulation in Arabidopsis and yeast cells under salt stress. Acta Physiol Plant 39, 19 (2017). https://doi.org/10.1007/s11738-016-2323-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-016-2323-3