Abstract
Although microbial reduction of perchlorate (ClO4 −) is a promising and effective method, our knowledge on the changes in microbial communities during ClO4 − degradation is limited, especially when different electron donors are supplied and/or other contaminants are present. Here, we examined the effects of acetate and hydrogen as electron donors and nitrate and ammonium as co-contaminants on ClO4 − degradation by anaerobic microcosms using six treatments. The process of degradation was divided into the lag stage (SI) and the accelerated stage (SII). Quantitative PCR was used to quantify four genes: pcrA (encoding perchlorate reductase), cld (encoding chlorite dismutase), nirS (encoding copper and cytochrome cd1 nitrite reductase), and 16S rRNA. While the degradation of ClO4 − with acetate, nitrate, and ammonia system (PNA) was the fastest with the highest abundance of the four genes, it was the slowest in the autotrophic system (HYP). The pcrA gene accumulated in SI and played a key role in initiating the accelerated degradation of ClO4 − when its abundance reached a peak. Degradation in SII was primarily maintained by the cld gene. Acetate inhibited the growth of perchlorate-reducing bacteria (PRB), but its effect was weakened by nitrate (NO3 −), which promoted the growth of PRB in SI, and therefore, accelerated the ClO4 − degradation rate. In addition, ammonia (NH4 +), as nitrogen sources, accelerated the growth of PRB. The bacterial communities’ structure and diversity were significantly affected by electron donors and co-contaminants. Under heterotrophic conditions, both ammonia and nitrate promoted Azospira as the most dominant genera, a fact that might significantly influence the rate of ClO4 − natural attenuation by degradation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Perchlorate (ClO4 −) contamination has been widely detected in soil, groundwater, and even drinking water due to its extensive use over the past decade in chemical industries such as aerospace fuels, explosive chemicals, and firecrackers (Urbansky 1998). ClO4 − is of concern due to its harmful influence on the thyroid hormones production and inhibition of normal iodine absorption in human physiology (Charnley 2008; Tran et al. 2008). Furthermore, as ClO4 − is weakly absorbed by the soil, is highly soluble in water, and is not efficiently reduced due to its low reactivity, it persists in the environment. The ClO4 − concentration can be as high as 3,700 mg/L in ground water, 120 mg/L in surface water, and 811 μg/L in drinking water (US EPA, 2005).
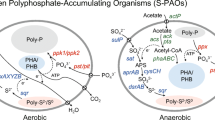
Currently, biological reduction of ClO4 − has been shown to be the most promising and effective approach for ClO4 − remediation because ClO4 − can be completely reduced to chloride (Cl−) by perchlorate-reducing bacteria (PRB) and chlorate reduction bacteria (CRB; Bardiya and Bae 2011; Nozawa-Inoue et al. 2011). PRB have been identified in the Proteobacteria classes Alphaproteobacteria, Betaproteobacteria, Gammaproteobacteria, and Deltaproteobacteria, among which members of the Betaproteobacteria, such as Azospira and Dechloromonas, are the most commonly detected PRB (Achenbach et al. 2001; Coates and Achenbach 2004; Coates et al. 1999; Michaelidou et al. 2000; Waller et al. 2004). In addition, denitrifying bacteria, such as Pseudomonas sp. strain PK, has also been shown to grow using ClO4 − or chlorate as an electron acceptor (Coates and Achenbach 2004). A variation in microbial communities and their degradative capabilities was detected in ClO4 − contaminated soils and the deep unsaturated zone (Gal et al. 2008). In addition, the examination of the microbial function and structure in a membrane biofilm reactor revealed the occurrence of the dominant class Betaproteobacteria in the presence of insufficient hydrogen during the degradation of ClO4 − (Zhao et al. 2011). To further analyze the function of microbial communities in the reduction of ClO4 − and nitrate (NO3 −) as well as degradation of ammonia (NH4 +), the pcrA gene (encoding perchlorate reductase) and cld gene (encoding chlorite dismutase) had been used to detect PRB and CRB, and the nirS gene (encoding copper and cytochrome cd1 nitrite reductase) had been used to detect denitrifying bacteria (De Long et al. 2010; De Long et al. 2012; Nozawa-Inoue et al. 2008; Zhao et al. 2011). However, our knowledge on the changes in the structure of microbial communities and function during ClO4 − degradation is limited, especially when different electron donors are supplied.
Additional electron donors such as hydrogen and acetate have been used to promote microbial reduction of ClO4 − in autotrophic and heterotrophic conditions, respectively. It has been reported that the rate of ClO4 − reduction in industrial soil was enhanced by hydrogen and inhibited by acetate, but the duration of ClO4 − reduction was shortened by acetate in Yolo loam soil (Nozawa-Inoue et al. 2011). Furthermore, aquifer materials from 12 different locations also degraded ClO4 − to an undetectable level in an environment with added acetate (Waller et al. 2004), and it is suggested that various microbial populations will respond differently to acetate during ClO4 − reduction. Previous studies have also shown that ClO4 − reduction is influenced by many physiochemical factors, including pH, temperature, dissolved oxygen (DO), and nitrate (Bruce et al. 1999; Chaudhuri et al. 2002; Coates et al. 1999; Logan 2001; Michaelidou et al. 2000; Wang et al. 2008). Nitrate and ClO4 − have similar physical and chemical characteristics including their reduction potential and very weak soil adsorption. Therefore, nitrate is a competing electron acceptor when present with ClO4 − during the reduction process (Herman and Frankenberger 1999). Nitrate also inhibited ClO4 − reduction in an autohydrogenotrophic treatment system and in heterotrophic fixed biofilm reactors (Choi and Silverstein 2008). However, it has been demonstrated that a higher nitrate concentration accelerated ClO4 − reduction (Choi and Silverstein 2008; Gal et al. 2008; London et al. 2011; Nerenberg and Rittmann 2004), and the degradation of ClO4 − was enhanced once NO3 − was completely removed via an elevated growth ratio of PRB (Chaudhuri et al. 2002; Giblin and Frankenberger 2001; Tipton et al. 2003; Waller et al. 2004; Xu et al. 2004). Furthermore, ammonium nitrate is a frequent component observed along with ClO4 − in water or soil in rocket fuel and explosives (Nozawa-Inoue et al. 2011). Ammonium ions (>0.4 %) have been shown to regenerate with perchlorate-laden resins during the ion-exchange process (Gingras and Batista 2002). In some natural sites, high concentrations of iron and manganese have also been noted to create a reducing environment for the coexistence of ammonia-perchlorate and ammonia-nitrate-perchlorate due to weak nitrification processes (Ye et al. 2013). Thus, ammonium ions are another ClO4 − co-contaminant that significantly affects perchlorate reduction (Gingras and Batista 2002). Nevertheless, the variances in the microbial communities’ composition during ammonia with ClO4 − and/or nitrate degradation have been rarely investigated.
In the present study, we aimed to answer the following four questions: (1) How do biodegradation rates differ when donors (hydrogen and acetate) are supplied and co-contaminants (nitrate-perchlorate, ammonia-perchlorate) are added? (2) How does the abundance of target genes vary in the above-mentioned degradative processes? (3) How does the bacterial community change in the lag stage (SI) and accelerated stage (SII) during different pretreatments? (4) Can microbiological properties explain the changes in the biodegradation rate caused by the addition of different donors and co-contaminants? To better understand the biodegradation mechanisms of ClO4 −, quantitative PCR (qPCR) and pyrosequencing were employed.
Materials and methods
Experimental setup
To analyze the effect of electron donors (acetate and hydrogen) and co-contaminants (nitrate and/or ammonium) on ClO4 − degradation, one control system and six different degradation systems were employed (NP, HYP, ACP, PNI, PAM, and PNA; Table 1). In 580-mL serum bottles, 500 mL of nutrient solution containing (per liter) 30 mg KH2PO4, 30 mg (NH4)2SO4, 10 mg NaHCO3, 10 mg MgSO4 · 7H2O, 5 mg H3BO3, 5 mg (NH4)6Mo7O24 · 4H2O, 2 mg FeSO4 · 7H2O, 2 mg ZnSO4 · 7H2O, 2 mg CuSO4 · 5H2O, 2 mg MnSO4 · H2O, 1 mg KAl(SO4)2 · 12H2O, 1 mg NiSO4 · 6H2O, and 1 mg CoSO4 · 7H2O were added and sealed with aluminum, and then subsequently sparged with N2 (99 %) for 30 min to remove DO and to achieve anoxic conditions. Finally, sterilization was performed for 25 min at 121 °C and 0.1 mpa. The microcosms were sampled from an anaerobic reactor in our laboratory and cultured at 30 °C with DO below 0.5 mg/L under constant shaking at 120 rpm on a rotary shaker. The microorganisms were inoculated at a dilution of 1:100 from the NP system into the prepared solutions in serum bottles to create five new systems: HYP, ACP, PNI, PAM, and PNA. In the all experiments, the ClO4 − concentration was 10 mg/L. The degradation of ClO4 − from 10 mg/L to the detection limit was considered to be a complete removal cycle. Triplicate batch-scale parallel control experiments were employed for each system. All the treatments and replicates were incubated at 30 °C, DO was maintained below 2 mg/L, and the pH was maintained between 7.0 and 8.0.
Analytical method
For each sample, 5 mL of the cultures was taken from the triplicate degradation systems with a sterile syringe and needle. All the samples were filtered through 0.22-μm filters prior to analysis. The ClO4 − concentrations were measured using an ion chromatograph ICS Dionex 90 (Dionex, Sunnyvale, CA, USA) equipped with an analytical column IonPAc AS 20 (4 × 250 mm) and guard column AG20 (4 × 50 mm; Xu et al. 2010), and 100 mM NaOH was used as a rinse solution for ClO4 − analysis and 10 mM H2SO4 was employed as the ClO4 − regeneration solution. The eluent used for ClO4 − was 65 mM NaOH. The sample injection volume was 100 μL and the flow rate was 1.0 mL/min. The levels of NO3 −-N, NO2 −-N, and NH4 +-N were assayed using the UV–visible spectrophotometer HP 8453 (Hewlett-Packard, Palo Alto, CA, USA) at the wavelengths of 220–275, 540, and 420 nm (London et al. 2011). The detection limit of ClO4 −, NO3 −-N, NO2 −-N, and NH4 +-N was 0.1 mg/L. Total organic carbon (TOC) was measured using TOC-VCPN (Shimadzu, Japan).
Biological samples and DNA extraction
The ClO4 − degradation rate of one period was calculated via two adjacent time point measurements for each ClO4 − degradation system. When the degradation rate between two adjacent periods increased more than 5 %, the degradation process was in accelerated stage (SII). The period between the start of incubation and beginning of the accelerated stage was the lag stage (SI). Therefore, all six perchlorate degradation systems were divided into two stages (SI and SII). Samples from each degradation system in SI and SII were collected. Twelve microbial samples in total were subjected to pyrosequencing.
Microbial samples for biological analysis were collected during a complete degradation cycle for NP, HYP, ACP, PNI, PAM, and PNA. The incubation bottles were shaken at 200 rpm for 30 min to mix the suspension thoroughly, and then 60 mL of mixed culture was transferred to a centrifuge tube and centrifuged at 12,000 rpm for 10 min. Subsequently, the supernatant was discarded and the samples were stored at −20 °C. Bacterial genomic DNA from 10 samples from each of the HYP, ACP, PNI, PAM, and PNA systems was extracted using a DNA isolation kit (Tiangen, China), whereas bacterial genomic DNA from 0.5 g of the samples from the NP system was extracted using a FastDNA SPIN kit (Mobio, Carlsbad, CA, USA). The extracted DNA was dissolved in 50 μL of nuclease-free water, and its quality was tested using a 1.5 % agarose gel. The quantity and quality of the isolated DNA were evaluated using a NanoDrop ND-2000 spectrophotometer (Thermo Fisher Scientific, Wilmington, DE, USA) and agarose gel electrophoresis (Bio-Rad, Hercules, CA, USA), respectively.
Quantification of pcrA, cld, nirS, and 16S rRNA genes
We estimated the number of pcrA, cld, nirS, and 16S rRNA gene copies using real-time qPCR. Plasmids containing target-gene fragments were constructed to create standard curves. Standard curves for the pcrA, cld, and nirS genes were based on serial dilutions of target genes (between 109 and 102 copies), and the 16S rRNA gene (between 1012 and 105 copies). These numbers were calculated from the concentration of extracted plasmids measured with Nano Drop ND-2000 spectrophotometer. qPCR was performed on an ABI Prism 7300 (Applied Biosystems, Foster City, CA, USA) with a reaction mixture containing 10 μL of SYBR FAST Universal 2X qPCR Master Mix from the KAPA SYBR FAST qPCR kit (Kapa Biosystems, Woburn, MA, USA), 1 μM of each primer, 2 μL of diluted DNA template corresponding to ∼10 ng of total DNA, and appropriate amount of RNase-free water to obtain a 20-μL volume. Thermal cycling conditions used for each reaction were shown in Table S1. Negative controls contained water instead of template DNA in the PCR mix. Triplicate PCR reactions for all samples and negative controls. The slopes of the plasmid standard curves and efficiency values for quantification by qPCR were as follows: −3.662 and 0.99 for pcrA, −3.672 and 0.99 for cld, −3.783 and 0.99 for nirS, and 3.182 and 0.99 for 16S rDNA, respectively. Pearson correlation between abundance of four genes and ClO4 − degradation rate (Table S2; Zhao et al. 2011). The cell ratios were estimated from the gene copy numbers using the following assumptions: one pcrA copy and two nirS copies per cell, as carried in the genome of ClO4 − and nitrate-reducing Dechloromonas aromatica RCB (Bender et al. 2005); one cld copy per cell carried in Dechloromonas agitata (De Long et al. 2010) and an average of seven copies of 16S rRNA genes per bacterial cell carried from Escherichia coli (Zhao et al. 2011).
Processing of pyrosequencing data
PCR was performed with the primers 338F (5′-ACTCCTACGGGAGGCAGCAG-3′; Lane 1991) and 907R (5-CCGTCAATTCMTTTGAGTTT-3; Muyzer 2004) to amplify the region V3–V5 from the 16S rRNA gene. Purified PCR products were subjected to high-throughput sequencing on the 454 GS FLX Sequencer using standard protocols (454 Life Sciences) at BGI, Beijing. All the 16S rRNA amplicons sequences were processed using the QIIME pipeline (the Quantitative Insights into Microbial Ecology, version 1.2; Caporaso et al. 2010b; Wang et al. 2012). The sequences were denoised with the Denoiser algorithm (Reeder and Knight 2010), including sequences that contained ambiguous base calls, were less than 300 nt or more than 450 nt in length and contained more than 20 homopolymers. Sequences were clustered into OTUs using a 97 % identity threshold with the seed-based uclust algorithm (Edgar 2010). Chimeras were removed using Chimera Slayer (Haas et al 2011). The most abundant sequence from each OTU was selected as a representative sequence, and then the representative sequences were aligned to the Greengenes imputed core reference alignment (DeSantis et al. 2006) using PyNAST (Caporaso et al. 2010a). The alignments were subsequently used to construct an approximate, maximum-likelihood phylogenetic tree with Jukes–Cantor distance using FastTree (Price et al. 2010) after removing gaps and hypervariable regions using a Lane mask. Taxonomic identity of each representative sequence was determined using a randomly selected representative sequence with the classifier platform within the Ribosomal Database Project (http://rdp.cme.msu.edu) reference alignment (Wang et al. 2007). We accounted for differences in sampling effort among samples by randomly subsampling 7,000 sequences per sample 1,000 times.
Statistical analyses
After normalizing sequence counts of each taxonomic group by the total number of reads, statistical analyses were performed on bacterial composition and abundance. All analyses were conducted with the statistical platform R (version 2.15.1). Taxonomic alpha diversity was calculated as estimated richness by using Chao1-richness and Pielou’s evenness. Chao1-richness (Chao 1984) and Pielou’s evenness (Pielou 1966) were used to estimate the community diversity. Chao1-richness is a non-parametric estimator of richness (Chao 1984) that is computed as Chao1-richness = Sobs + [a 2/(2 × b)], where Sobs is the number of species observed and a or b is the number of species that are observed just once or twice. Furthermore, phylogenetic diversity was calculated as Faith’s phylogenetic diversity (PD; Faith 1992). Faith’s PD measures the total phylogenetic branch length that joins the basal node to the tips of all the species in the sample (Faith 1992).
To quantify the taxonomic community similarity between the given samples, we used unweighted Unifrac (Lozupone and Knight 2005). We performed non-metric multidimensional scaling based on unweighted Unifrac to depict community composition in two dimensions.
Results
Perchlorate reduction under autotrophic and heterotrophic conditions
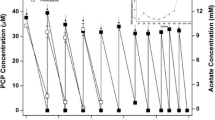
The biodegradation in NP system and NP with acetate/hydrogen as electron donor (ACP/HYP) systems were compared. The ClO4 − concentration in the control system did not change (not shown). In the NP, ACP, and HYP systems, 10 mg/L of ClO4 − was completely degraded within 10, 3.8, and 66 days, respectively (Fig. 1). The SI of NP accounted for about 1/10 of the total degradation cycle, and this ratio increased to 1/4 and 1/6 for ACP and HYP systems, respectively. The zero-order reaction rate equations of NP, ACP, and HYP were y = 0.048x − 0.0658 (R 2 = 0.989), y = 0.379x − 0.0147 (R 2 = 0.974), and y = 0.009x + 0.2443 (R 2 = 0.920), respectively.
Quantification of the pcrA, cld, nirS, and 16S rRNA genes and degradation of ClO4 − in the NP (a), ACP (b), and HYP (c) systems. NP only perchlorate contamination system, ACP NP system with acetate, HYP NP system with hydrogen, PNI ACP system with nitrate, PAM ACP system with ammonia, PNA ACP system with nitrate and ammonia
Effects of nitrate and/or ammonia on perchlorate reduction
In the PNI system, 10 mg/L of ClO4 − was completely degraded within about 70 h (Fig. 2a). The concentration of NO3 −-N decreased rapidly in the first 33 h (SI), and there was no significant change in the concentration of ClO4 −. The degradation rate of ClO4 − increased until NO3 −-N under detected limit, but the presence of NO3 −-N significantly shortened the ClO4 − degradation period about 20 h, when compared with that noted in the ACP system. The zero-order reaction rate equation in SII for the PNI system was y = 0.366x − 0.0099 (R 2 = 0.95). The biodegradation of ClO4 − was faster in the PAM system than that in the PNI. In the PAM system, 10 mg/L ClO4 − was completely removed within 60 h (Fig. 2b). The biodegradation of ClO4 − was accelerated in the presence of NH4 +-N, and 10 mg/L ClO4 − was degraded within 33 h in PAM system. The zero-order reaction rate equation in SII for the PAM system was y = 0.395x − 0.2004 (R 2 = 0.95). The degradation of ClO4 − in the PNA system was faster than that in the PNI or PAM systems, and 10 mg/L ClO4 − was degraded within 45 h (Fig. 2c). The zero-order reaction rate equation in SII for the PNA system was y = 0.195x − 0.0015 (R 2 = 0.998).
Abundance of functional and bacterial 16S rRNA genes
The gene abundance of the pcrA and cld in the contaminant systems were 104–106 and 106–109 copies per μg DNA, respectively (Figs. 1 and 3). The gene abundance of pcrA in the NP system rose to a peak at 1.2 day when the degradation proceeded to SII, while that of cld reached a peak at 4 day (Fig. 1a). In the ACP system, the cld and pcrA genes reached peaks in 1.2 and 1.5 day, respectively, and the gene abundance of cld about 107 copies per microgram DNA, which was higher than that noted in the NP and HYP systems (Fig. 1b). Furthermore, the gene abundance of the pcrA in the HYP system was slightly higher than that in the ACP system, while the abundances of cld and nirS genes were significantly lower than those noted in the ACP system (Fig. 1c). The abundance of 16S rRNA in the ACP system was higher than that in the HYP system. After 12 days, the degradation of ClO4 − was accelerated in the HYP system, and the abundance of the pcrA and cld genes began to increase simultaneously. When the degradation continued till the end of SII (the ClO4 − concentration dropped close to 4 mg/L), the peak of pcrA gene level was close to that noted in the PNA system and those observed in the other four systems.
In systems PNI, PAM, and PNA, the gene abundance of pcrA was similar to that noted in ACP and HYP system, but the abundance of the cld and nirS genes was 102–103 and 103–104 higher than that found in the ACP and HYP systems and HYP system (Fig. 3). In the PNI system, degradation of ClO4 − entered SII after 27 h, and the abundance of pcrA, cld, and nirS genes peaked at 56, 45, and 56 h, respectively. The gene abundance of pcrA did not exhibit a significant downward trend after the peak, and the average gene abundance of cld was 3–4 times higher than that of pcrA. In the PAM system, all the four genes reached a peak within 33 h, and the abundances of pcrA, cld, and nirS genes were close to that of the PNI system; however, the abundance of nirS gene significantly declined when compared with that noted in the PNI system. In the PNA system, three genes, 16S rRNA, cld, and nirS, reached peaks within a short time of about 3 h, while the pcrA gene reached a peak within 21 h, and the abundance of these genes were the highest observed in this study. The degradation of ClO4 − was accelerated at 21 h when the pcrA gene abundance peaked. The Pearson’s correlation coefficients between ClO4 − degradation rate and abundance of four genes are shown in Table S2, and none of the genes exhibited good correlation with ClO4 − degradation.
The bacterial community in the six contaminant systems
With pyrosequencing, 290,224 sequences and 5,871 OTUs (97 % cutoff) were obtained for 12 samples. The two principal components explained 45.66 % of the total variance (35.18 % for PC1 and 10.48 % for PC2) (Fig. 4). The bacterial community in the NP system was most similar to that in the ACP system and that in the HYP system was different from the communities in the other heterotrophic systems. In both the stages (SI and SII), the bacterial communities were grouped together in PAM and PNI systems (Fig. 4). However, in the PNA system, the bacterial communities were clearly separated from others. The alpha diversity of the bacterial communities in the NP system was the highest, whereas that of the bacterial communities in the PNA system was the lowest among the six contaminant systems (Table S3). With regard to the diversity of the other five systems inoculated from the NP system, ACP system exhibited higher diversity than the HYP system and three nitrogen and ClO4 − co-contaminant systems, and SI was higher than SII in each system.
The top five dominant phyla in the NP system were Proteobacteria (22.4 %), Bacteroidetes (11.1 %), Firmicutes (3.1 %), Chloroflexi (2.3 %), and Thermotogae (2.1 %). Among Proteobacteria, the most predominant family was Rhodocyclaceae, in which Syntrophus, Chlorobaculum, and Azospira were the dominant genera. In the ACP system, Pseudomonas, Azospira, Acinetobacter, Syntrophus, and Sterolibacterium were the dominant genera. In the HYP system, the top five genera were Azospira, Acinetobacter, Curvibacter, Limnohabitans, and Comamonas. Azospira was the most dominant genus, which decreased in number as the degradation proceeded, while the number of autotrophic genera Limnohabitans, Novosphingobium, and Pseudomonas increased. In the three co-contaminant systems, the relative proportions of Azospira and Acinetobacter were more than 90 %. As shown in Table S3, in the PNA system, Azospira was the most dominant genus accounting for 90 %. In addition, Dechloromonas, Zoogloea, Acidovorax, Pelomonas, Novosphingobium, and Stenotrophomonas were found in the PNA system (Table 2). In the heterotrophic ACP, PNI, and PNA systems, the abundance of Azospira increased in SII; whereas in the PAM system, the abundance of Azospira dramatically decreased in SII. The abundance of Azospira in these systems exhibited the following order: PNA, PNI, PAM, and ACP.
Discussion
The degradation of ClO4 − by microbes is regarded as the most promising method (Chung et al. 2007; London et al. 2011; Matos et al. 2006; Xiao et al. 2010). However, few studies employed different electron donors or co-contaminations (especially ammonia) to examine ClO4 − degradation. This study was performed to analyze the variations in the functional genes and microbial diversity when using acetate as an electron donor to elucidate the mechanisms of ClO4 − degradation with co-contaminating nitrate and/or ammonia.
Effect of electron donors and co-contaminants on ClO4 − degradation
In the NP system, insufficient TOC (∼9.21 mg/L) was available to completely reduce 10 mg/L of ClO4 −, and thus, the degradation rate was slower than that observed in the ACP system, which used acetate as an electron donor (Farhan and Hatzinger 2009). Excessive acetate in the ACP system was first used for microbial growth, whereas the decrease in acetate reduced the inhibition on ClO4 − degradation (Nozawa-Inoue et al. 2011; Oh and Silverstein 1999). In accordance with the research of Nozawa-Inoue et al. (2005) who used microbial communities in vadose soil with hydrogen as an electron donor, the HYP system using hydrogen as an electron donor exhibited the longest degradation period and lag time noted in the present study. Because of the low solubility of hydrogen in water during SI (0.08 mg/L at 5 % hydrogen), the growth of autotrophic bacterium was restricted by the limited hydrogen available in the culture medium. However, hydrogen solubility in the culture would increase as degradation proceeded, according to Henry’s law, and rapid growth of PRB was accompanied by an increase in the apparent ClO4 − degradation rate during the degradation phase (Fig. 1). In the PNI system, ClO4 − was rapidly reduced once NO3 −-N was degraded below the detection limit. The degradation rate of ClO4 − in the PAM system was slightly faster than that in the PNI system. The concentration of NH4 +-N declined slowly in the PAM and PNA systems, and DO was always at a low level (Fig. 2). At the same time, NH4 +-N could be detected even when the levels of ClO4 − and NO3 −-N had already decreased below the test limit. We presumed that NH4 +-N was used as the nitrogen source for microbial growth rather than for nitrification to NO3 −. The rate of ClO4 − reduction in the PAM and PNA systems was faster than that in the ACP system, which is consistent with the hypothesis that ClO4 − degradation could be inhibited by adding acetate, while NO3 −-N could weaken the inhibition of acetate on ClO4 − degradation in the co-contamination system (Nozawa-Inoue et al. 2011; Raj and Muruganandam 2012).
Relationship between the abundance of the four genes and ClO4 − degradation
Although the degradation rate of ClO4 − had different levels of correlation with the abundance of the four genes (Table S2), in the NP system, the acceleration period of ClO4 − reduction occurred simultaneously with the peak level of pcrA gene detection, and the abundance of the pcrA gene dropped rapidly, suggesting that the peak of pcrA expression was linked to the beginning of ClO4 − reduction. The reduction of ClO3 − to ClO2 − and the subsequent reduction to Cl− and O2 were promoted by a significant increase in the cld gene expression following SI. In the ACP system, the abundance of the pcrA gene was lower than that in the HYP and NP systems, indicating that acetate might have inhibited PRB growth. SI in the NP system was similar to that in the ACP system, whereas SII in the ACP system was more rapid than that in the NP and HYP systems. At the same time, the abundance of the cld gene in the ACP system was much higher than that in the NP or HYP system, suggesting that the expression of pcrA acted as a switch for the ClO4 − degradation process. Moreover, we presumed that the degradation rate in SII was determined by the expression of the cld gene. As a result of expanding the nirS-harboring denitrifier population in the presence of acetate, high nirS expression levels were observed, and this population was shown to participate in ClO4 − removal, shortening the ClO4 − degradation cycle. PRB that preferentially used hydrogen as an electron donor grew better in the HYP system, when compared with denitrifying bacteria. Consequently, the abundance of the pcrA gene in the HYP system was 10 and 70 times higher than that in the NP and ACP systems, respectively. Thus, we demonstrated that PRB predominantly influenced ClO4 − reduction with hydrogen as an electron donor.
The peak value of the pcrA and cld genes in the PNI system appeared later than that noted in the ACP system, in which acetate was added as the sole electron donor. It is suggested that acetate not only extended the ClO4 − degradation lag time, but also promoted the growth of denitrifying bacteria capable of ClO4 − degradation, thereby, accelerating the degradation of ClO4 − in SII and shortening the ClO4 − degradation cycle (Figs. 1b and 3). The peak of pcrA abundance in the PAM system appeared earlier than that in the PNI system, and ClO4 − degradation was immediately accelerated (Fig. 3). The DO in the PAM system remained at a low level (<2 mg/L), and under this condition, nitrification was inhibited such that ammonia was weakly converted to nitrate. The initial NH4 +-N concentration was 6 mg/L and remained in the 2–3 mg/L range before the level of ClO4 − fell below the detection limit. The abundance of the pcrA gene in the PAM system was higher than that in the PNI system, suggesting that the ammonia nitrogen source was used primarily for the growth of PRB, which accelerated ClO4 − degradation, thus shortening the degradation cycle in the PNI system. In the PNA system, the abundance of the pcrA, cld, nirS, and 16S rRNA genes were the highest among the systems examined in this study, and the pcrA gene peaked when the degradation of ClO4 − entered SII. Irrespective of whether or not nitrogen pollution was present, the pcrA gene declined rapidly at the end of SI. The abundance of the pcrA gene also declined rapidly in both the PAM systems, while that of the pcrA gene was stable with a slight increase in SII in the PNI system. However, the abundance of the cld gene increased rapidly in SI and remained at a high and almost same level in SII in these three systems. It is indicated that the pcrA gene might initiate and accelerate ClO4 − degradation, while the cld gene might maintain degradation in SII, which favored the conversion of ClO2 − to Cl− and O2. Moreover, with a concentration of 10 mg/L ClO4 −, degradation did not require excessive perchlorate reductase, which only needed to reach a threshold to initiate degradation. Some competing electron acceptors, such as nitrate, coexisted in the PNA system, and these competing electron acceptors promoted denitrifier growth. The changes in the abundance of the pcrA and nirS genes differed from those of the cld gene, which also maintained a relatively high abundance, indicating that PRB and nirS-harboring bacteria fully developed in SI. The subsequent reduction process was dominated by a series of CRB when ClO4 − reduction began. Ammonia and nitrate coexisted in the PNA system and the degradation of cycle was shortest for this system (Fig. 3), indicating that nitrate promoted the growth of the denitrifying bacteria capable of degrading ClO4 −, and thus, accelerated the degradation rate of ClO4 −. Ammonia can be used as an assimilable nitrogen source for the growth of bacteria in the natural environment. In addition, ammonia may be also used by PRB to promote growth, and thus may be the most important cause of accelerated ClO4 − degradation.
Effect of microbial community composition on ClO4 − degradation
Microbial community composition and function significantly influenced the degradation of pollutants, even in a complex pollution system. Indeed, these microbial populations play a decisive role in the synergistic processes and occurrence of degradation (Nozawa-Inoue et al. 2011). Microbial community composition changed with degradation in each of our experiments. The types and abundance of dominant bacteria reflected the process of pollution degradation (Li et al. 2012; Zhao et al. 2011). The microbial diversity of the ClO4 − and nitrogen contamination system decreased significantly, when compared with the three systems that only had ClO4 − as the pollutant (Table S3). The microbial community composition was similar in SI and SII in both the PNI and PAM systems. It was suggested that the co-contaminant system accelerated the dominance of PRB, and ammonia or nitrate and ammonia and nitrate together had little effect on the diversity of the bacterial communities relative to each other. In the autotrophic HYP system, the proportions of different bacteria in heterotrophic microbial communities became simpler, primarily because autotrophic microorganisms grew more slowly, and thus, became less abundant. The PCA results showed that the autotrophic system HYP was distinct from the other systems where hydrogen was provided as the electron donor (Fig. 4). The microbial community composition of the co-contaminated system formed one cluster, and was distinguished from the ClO4 −-alone contaminant system. In the PNA system, nitrate promoted the growth of PRB, while ammonia provided as a nitrogen source accelerated the growth of PRB, such that the PRB became dominant in a short time, and the microbial community composition of this system became distinct from the other systems examined in this study.
Azospira thrived in the heterotrophic ClO4 − contaminant system, and grew using ClO4 − and nitrate as an electron donor (Farhan and Hatzinger 2009; Sun et al. 2009). Azospira was undoubtedly the most dominant bacteria, especially with the co-contaminants ClO4 −, nitrate and ammonia. The relative abundance of Azospira significantly influenced the rate of ClO4 − degradation. Acinetobacter was the dominant bacteria in a wetland bioreactor used to remove ClO4 − and nitrate from contaminated groundwater (Krauter et al. 2005). Although both Acinetobacter and Azospira are capable of aerobic or anaerobic respiration with nitrate as the terminal electron acceptor, we presumed that Azospira outcompeted Acinetobacter in the presence of sufficient acetate and nitrate, while Acinetobacter became more dominant than Azospira in SII in the PAM system, where less acetate and ammonia were available as nitrogen sources. Pseudomonas is a relatively unusual chlorate-respiring bacterium due to its ability to completely degrade chlorate in the presence of dissolved oxygen. With ClO4 − and nitrogen as co-contaminants, Azospira still had a competitive advantage over Pseudomonas, but Pseudomonas was more dominant than Azospira in heterotrophic conditions with only ClO4 −. Although the relative abundance of certain PRB in the six degradation systems illustrated their competitive relationships, simple genus competition experiments is still needed for validation. Dechloromonas, Zoogloea, and Alicycliphilus can use ClO4 −, nitrate, and oxygen as electron acceptors for growth under both aerobic and anaerobic denitrifying conditions (Coates and Achenbach 2004). Although these bacteria have been observed as abundant bacteria in different environments, such as a fixed-bed biofilm reactor (Heylen et al. 2008; Li et al. 2012; Oosterkamp et al. 2013), they did not overcome Azospira as the abundant bacteria in the systems examined in the present study. In this analysis of bacterial community diversity and structure, the diversity of the bacterial community decreased quickly because of selection pressure caused by pollutants, and the dominant bacteria was primarily Azospira, a strain capable of using both perchlorate and nitrate as electron donors. However, bacterial competitiveness for electron donors varied in the six systems and autotrophic or heterotrophic nutritional conditions impacted the abundance of dominant bacteria, and thus, affected the degradation rate.
Conclusion
Six perchlorate degradation systems with different electron donors (hydrogen and acetate) and co-contaminants (nitrate and ammonium) were employed to study their influences on ClO4 − degradation. Acetate was more easily used as an electron donor than hydrogen in ClO4 − degradation, and nitrate weakened the inhibition of acetate and increased ClO4 − degradation by accelerating PRB growth. Furthermore, ammonia might have been used as an assimilable nitrogen source to promote the growth of PRB. The results of bacterial communities’ composition and function indicated that the pcrA gene probably acted as a switch for ClO4 − degradation in SI and that the cld gene sustained the degradation in SII. The bacterial community in the autotrophic system was obviously different from that in the five heterotrophic systems. Among the six systems, Azospira finally became the most dominant bacteria, and its relative abundance was the main factor influencing the rate of ClO4 − degradation. Thus, this study helped in achieving a better understanding of the biodegradation mechanism of ClO4 − reduction with nitrate and/or ammonia.
In the future studies, the N2/NO x produced by denitrification and anaerobic ammonia oxidation should be analyzed using GC-MS by replacing nitrogen with Ar2 and He. Furthermore, the threshold of the functional genes such as pcrA and cld for degradation in SI and SII should also be examined.
References
Achenbach LA, Michaelidou U, Bruce RA, Fryman J, Coates JD (2001) Dechloromonas agitata gen. nov., sp. nov. and Dechlorosoma suillum gen. nov., sp. nov., two novel environmentally dominant (per)chlorate-reducing bacteria and their phylogenetic position. Int J Syst Evol Microbiol 51:527–533
Bardiya N, Bae JH (2011) Dissimilatory perchlorate reduction: a review. Microbiol Res 166:237–254
Bender KS, Shang C, Chakraborty R, Belchik SM, Coates JD, Achenbach LA (2005) Identification, characterization, and classification of genes encoding perchlorate reductase. J Bacteriol 187:5090–5096
Bruce RA, Achenbach LA, Coates JD (1999) Reduction of (per)chlorate by a novel organism isolated from paper mill waste. Environ Microbiol 1:319–329
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010a) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Caporaso JG et al (2010b) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Stat 11:265–270
Charnley G (2008) Perchlorate: overview of risks and regulation. Food Chem Toxicol 46:2307–2315
Chaudhuri SK, O’Connor SM, Gustavson RL, Achenbach LA, Coates JD (2002) Environmental factors that control microbial perchlorate reduction. Appl Environ Microbiol 68:4425–4430
Choi H, Silverstein J (2008) Inhibition of perchlorate reduction by nitrate in a fixed biofilm reactor. J Hazard Mater 159:440–445
Chung J, Nerenberg R, Rittmann BE (2007) Evaluation for biological reduction of nitrate and perchlorate in brine water using the hydrogen-based membrane biofilm reactor. J Environ Eng-Asce 133:157–164
Coates JD, Achenbach LA (2004) Microbial perchlorate reduction: rocket-fuelled metabolism. Nat Rev Microbiol 2:569–580
Coates JD, Michaelidou U, Bruce RA, O’Connor SM, Crespi JN, Achenbach LA (1999) Ubiquity and diversity of dissimilatory (per)chlorate-reducing bacteria. Appl Environ Microbiol 65:5234–5241
De Long SK, Kinney KA, Kirisits MJ (2010) qPCR assays to quantify genes and gene expression associated with microbial perchlorate reduction. J Microbiol Meth 83:270–274
De Long SK, Li X, Bae S, Brown JC, Raskin L, Kinney KA, Kirisits MJ (2012) Quantification of genes and gene transcripts for microbial perchlorate reduction in fixed-bed bioreactors. J Appl Microbiol 112:579–592
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Faith DP (1992) Conservation evaluation and phylogenetic diversity. Biol Conserv 61:1–10
Farhan YH, Hatzinger PB (2009) Modeling the biodegradation kinetics of perchlorate in the presence of oxygen and nitrate as competing electron acceptors. Bioremediat J 13:65–78
Giblin T, Frankenberger WT (2001) Perchlorate and nitrate reductase activity in the perchlorate-respiring bacterium perclace. Microbiol Res 156:311–315
Gingras TM, Batista JR (2002) Biological reduction of perchlorate in ion exchange regenerant solutions containing high salinity and ammonium levels. J Environ Monitor 4:96–101
Haas BJ et al (2011) Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21:494–504
Gal H, Ronen Z, Weisbrod N, Dahan O, Nativ R (2008) Perchlorate biodegradation in contaminated soils and the deep unsaturated zone. Soil Biol Biochem 40:1751–1757
Herman DC, Frankenberger WT (1999) Bacterial reduction of perchlorate and nitrate in water. J Environ Qual 28:1018–1024
Heylen K, Lebbe L, De Vos P (2008) Acidovorax caeni sp. nov., a denitrifying species with genetically diverse isolates from activated sludge. Int J Syst Evol Microbiol 58:73–77
Krauter P, Daily B Jr, Dibley V, Pinkart H, Legler T (2005) Perchlorate and nitrate remediation efficiency and microbial diversity in a containerized wetland bioreactor. Int J Phytoremediation 7:113–128
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, West Sussex, United Kingdom, pp 115–175
Li X, Yuen W, Morgenroth E, Raskin L (2012) Backwash intensity and frequency impact the microbial community structure and function in a fixed-bed biofilm reactor. Appl Microbiol Biotechnol 96:815–827
Logan BE (2001) Assessing the outlook for perchlorate remediation. Environ Sci Technol 35:482–487
London MR, De Long SK, Strahota MD, Katz LE, Speitel GE Jr (2011) Autohydrogenotrophic perchlorate reduction kinetics of a microbial consortium in the presence and absence of nitrate. Water Res 45:6593–6601
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235
Matos CT, Velizarov S, Crespo JG, Reis MAM (2006) Simultaneous removal of perchlorate and nitrate from drinking water using the ion exchange membrane bioreactor concept. Water Res 40:231–240
Michaelidou U, Coates JD, Achenbach LA (2000) Isolation and characterization of two novel (per)chlorate-reducing bacteria from swine waste lagoons. Perchlorate in the Environment. Springer US, 271-283
Muyzer G (2004) Denaturing gradient gel electrophoresis (DGGE) in microbial ecology. In: G.A. Kowalchuk (ed.) Molecular microbial ecology manual. Dordrecht, Kluwer, pp 743–769
Nerenberg R, Rittmann BE (2004) Hydrogen-based, hollow-fiber membrane biofilm reactor for reduction of perchlorate and other oxidized contaminants. Water Sci Technol 49:223–230
Nozawa-Inoue M, Jien M, Hamilton NS, Stewart V, Scow KM, Hristova KR (2008) Quantitative detection of perchlorate-reducing bacteria by real-time PCR targeting the perchlorate reductase gene. Appl Environ Microbiol 74:1941–1944
Nozawa-Inoue M, Jien M, Yang K, Rolston DE, Hristova KR, Scow KM (2011) Effect of nitrate, acetate, and hydrogen on native perchlorate-reducing microbial communities and their activity in vadose soil. Fems Microbiol Ecol 76:278–288
Nozawa-Inoue M, Scow KM, Rolston DE (2005) Reduction of perchlorate and nitrate by microbial communities in vadose soil. Appl Environ Microbiol 71:3928–3934
Oh J, Silverstein J (1999) Acetate limitation and nitrite accumulation during denitrification. J Environ Eng-Asce 125:234–242
Oosterkamp MJ, Boeren S, Plugge CM, Schaap PJ, Stams AJ (2013) Metabolic response of Alicycliphilus denitrificans strain BC towards electron acceptor variation. Proteomics 13:2886–2894
Pielou E (1966) The measurement of diversity in different types of biological collections. Theor Biol 13:131–144
Price AL, Kryukov GV, de Bakker PI, Purcell SM, Staples J, Wei LJ, Sunyaev SR (2010) Pooled association tests for rare variants in exon-resequencing studies. Am J Hum Genet 86:832–838
Raj JRA, Muruganandam L (2012) Batch studies and effect of environmental parameters on microbial degradation of perchlorate using acclimatized effluent sludge research. J Chem Environ 16:44–50
Reeder J, Knight R (2010) Rapidly denoising pyrosequencing amplicon reads by exploiting rank-abundance distributions. Nat Methods 7:668–669
Sun Y, Gustavson RL, Ali N, Weber KA, Westphal LL, Coates JD (2009) Behavioral response of dissimilatory perchlorate-reducing bacteria to different electron acceptors. Appl Microbiol Biotechnol 84:955–963
Tipton DK, Rolston DE, Scow KM (2003) Transport and biodegradation of perchlorate in soils. J Environ Qual 32:40–46
Tran N et al (2008) Thyroid-stimulating hormone increases active transport of perchlorate into thyroid cells. Am J Physiol Endocrinol Metab 294:802–806
Urbansky ET (1998) Perchlorate chemistry: implications for analysis and remediation. Bioremediat J 2:81–95
Waller AS, Cox EE, Edwards EA (2004) Perchlorate-reducing microorganisms isolated from contaminated sites. Environ Microbiol 6:517–527
Wang C, Lippincott L, Meng X (2008) Kinetics of biological perchlorate reduction and pH effect. J Hazard Mater 153:663–669
Wang J, Soininen J, He J, Shen J (2012) Phylogenetic clustering increases with elevation for microbes. Environ Microbiol Rep 4:217–226
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Xiao Y, Roberts DJ, Zuo G, Badruzzaman M, Lehman GS (2010) Characterization of microbial populations in pilot-scale fluidized-bed reactors treating perchlorate- and nitrate-laden brine. Water Res 44:4029–4036
Xu J, Gao N, Tang Y, Deng Y, Sui M (2010) Perchlorate removal using granular activated carbon supported iron compounds: Synthesis, characterization and reactivity. J Environ Sci-China 22:1807–1813
Xu J, Trimble JJ, Steinberg L, Logan BE (2004) Chlorate and nitrate reduction pathways are separately induced in the perchlorate-respiring bacterium Dechlorosoma sp. KJ and the chlorate-respiring bacterium Pseudomonas sp. PDA. Water Res 38:673–680
Ye L, You H, Yao J, Kang X, Tang L (2013) Seasonal variation and factors influencing perchlorate in water, snow, soil and corns in Northeastern China. Chemosphere 90:2493–2498
Zhao HP, Van Ginkel S, Tang Y, Kang DW, Rittmann B, Krajmalnik-Brown R (2011) Interactions between perchlorate and nitrate reductions in the biofilm of a hydrogen-based membrane biofilm reactor. Environ Sci Technol 45:10155–10162
Acknowledgments
This research was supported by National Natural Science Foundation of China (40972162, granted to FL), Beijing Higher Education Young Elite Teacher Project (granted to XYG), and The Fundamental Research Funds for the Central Universities (2012071, granted to XYG).
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Gerald Thouand
Xiangyu Guan and Yuxuan Xie equally contributed to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 24 kb)
Rights and permissions
About this article
Cite this article
Guan, X., Xie, Y., Wang, J. et al. Electron donors and co-contaminants affect microbial community composition and activity in perchlorate degradation. Environ Sci Pollut Res 22, 6057–6067 (2015). https://doi.org/10.1007/s11356-014-3792-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-014-3792-9