Abstract
Linkages among bioreactor operation and performance and microbial community structure were investigated for a fixed-bed biofilm system designed to remove perchlorate from drinking water. Perchlorate removal was monitored to evaluate reactor performance during and after the frequency and intensity of the backwash procedure were changed, while the microbial community structure was studied using clone libraries and quantitative PCR targeting the 16S rRNA gene. When backwash frequency was increased from once per month to once per day, perchlorate removal initially deteriorated and then recovered, and the relative abundance of perchlorate-reducing bacteria (PRB) initially increased and then decreased. This apparent discrepancy suggested that bacterial populations other than PRB played an indirect role in perchlorate removal, likely by consuming dissolved oxygen, a competing electron acceptor. When backwash intensity was increased, the reactor gradually lost its ability to remove perchlorate, and concurrently the relative abundance of PRB decreased. The results indicated that changes in reactor operation had a profound impact on reactor performance through altering the microbial community structure. Backwashing is an important yet poorly characterized procedure when operating fixed-bed biofilm reactors. Compared to backwash intensity, changes in backwash frequency exerted less disturbance on the microbial community in the current study. If this finding can be confirmed in future work, backwash frequency may serve as the primary parameter when optimizing backwash procedures.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Biofilm reactor technology provides an important avenue for drinking water treatment and has demonstrated to be effective in removing a range of contaminants from source waters (Nerenberg and Rittmann 2004; Bouwer and Crowe 1988; Choi et al. 2008; Upadhyaya et al. 2010; Ding and Yokota 2010). A key component in further advancing the technology is to elucidate the linkages among reactor operation, reactor performance, and microbial community (Briones and Raskin 2003; Curtis et al. 2003). In this study, a fixed-bed biofilm reactor designed to remove perchlorate from drinking water was studied to illustrate these relationships. Perchlorate is an endocrine disruptor with several adverse health effects (Greer et al. 2002) and has been detected in many drinking water sources (Urbansky 2002). Therefore, perchlorate is on U.S. EPA's drinking water contaminant candidate list (US-EPA 2008) and is regulated by a number of states in the U.S. at various levels ranging from 1 to 18 μg/L (CA-DHS 2005; MA-DEP 2006; US-EPA 2005).
In recent years, fixed-bed biofilm reactors have been evaluated for the removal of perchlorate from drinking water (Miller and Logan 2000; Brown et al. 2003; Choi et al. 2007; Li et al. 2010). During normal operation, the biofilm support medium in fixed-bed reactors remains stationary, as opposed to fluidized-bed reactors in which the support medium is fluidized. As water passes through the stationary bed in a downflow mode, particulates including microbial cell aggregates accumulate in void space and cause clogging. Clogging can be prevented by regular reactor backwashing, which is necessary for continued operation of these bioreactors (Hozalski and Bouwer 1998). Backwashing, the passing of a flow of water or a mixture of water and air in an upflow mode, removes particulate matter and resolves bioreactor clogging (Ahmad et al. 1998). Backwashing removes most loosely attached biofilms (i.e., cell aggregates in the pores of support media) and a fraction of tightly attached biofilms (Servais et al. 1991), depending on the backwash intensity. Backwashing with high intensity may cause too much biomass loss and impair reactor performance (Lahav et al. 2001). Backwashing with low intensity may result in insufficient removal of accumulated biomass and quick pressure buildup during operation, and consequently, frequent backwashes will be needed. Backwash frequency is usually established by monitoring pressure buildup caused by clogging across the reactor. In general, backwash procedures are experimentally optimized for each individual facility, which creates challenges to systematically study this important procedure and explains why studies on backwashing have been scarce.
In an earlier study, the effects of backwash procedures on reactor performance were investigated using two fixed-bed biofilm reactors operated to remove perchlorate from water (Choi et al. 2007). The two reactors had 3- and 1-mm diameter glass beads as support media, respectively, and were operated with various influent dissolved oxygen levels. Results showed that after the backwash frequency was increased from monthly to daily, reactor performance first decreased and then recovered when the backwashes were performed with low intensity. In contrast, daily strong backwashes caused a failure of perchlorate removal, particularly when influent DO level was high. The difference in performance responses to the various reactor operating conditions likely was rooted in the way microbial communities, especially perchlorate reducing bacterial populations, responded to the changes in backwash procedure.
Perchlorate reducing bacteria (PRB) can utilize perchlorate as an electron acceptor and convert it to non-toxic chloride (Coates and Achenbach 2004). PRB described so far mainly belong to three genera Dechloromonas, Azospira, and Dechlorospirillum (Coates et al. 1999; Waller et al. 2004; Bardiya and Bae 2008; Zhang et al. 2005). For most PRB, oxygen is the preferred electron acceptor over perchlorate (Coates and Achenbach 2004). Hence, to allow effective perchlorate removal in bioreactors, oxygen needs to be removed by PRB or other aerobic bacteria before biological perchlorate reduction can occur (Xu et al. 2003). Therefore, simultaneous studies of PRB and other aerobic bacteria are crucial in studying engineered systems designed for perchlorate removal.
In this study, linkages among reactor operating conditions (backwash frequency and intensity), reactor performance or microbial community function (removal of dissolved oxygen and perchlorate), and microbial community structure (relative abundance of PRB and other aerobic bacteria) were investigated for a fixed-bed biofilm reactor designed to remove perchlorate from drinking water. The microbial community structure was first determined using clone library analyses targeting the 16S rRNA gene and then major bacterial populations were monitored using quantitative polymerase chain reaction (qPCR). Microbial community responses to various backwash parameters were reported, which provide insights in designing backwash strategies for fixed-bed biofilm systems.
Materials and methods
Fixed-bed biofilm reactor and reactor operation
The reactor configuration, influent composition, operating conditions, and backwash procedures were similar to the ones reported in an earlier study (Choi et al. 2007). In brief, a bench-scale fixed-bed biofilm reactor with an inner diameter of 2.4 cm and a height of 24 cm was filled with 1-mm diameter glass beads, resulting in an empty bed volume of 63.4 mm3. The reactor was inoculated with biomass from a biologically active carbon filter previously operated to treat groundwater amended with 50 μg/L perchlorate using acetate as the electron donor (Lin 2004). Influent was pumped into the bottom of the reactor at a flow rate (Q influent) of 2.7 mL/min, resulting in an empty bed contact time of 23.4 min. A recirculation loop (Q recirculation = 5 × Q influent) was installed to the reactor to reduce the plug-flow characteristics of the reactor. The influent contained two electron acceptors, 3 mg/L DO and 50 μg/L perchlorate. A phosphate buffer solution with a final concentration of 0.5 mM maintained the pH inside the reactor at 7.5, and served as phosphorus source along with 0.01 mM NH4Cl as nitrogen source. Based on stoichiometric calculations (Rittmann and McCarty 2001) with an assumed net yield value of 0.2 g CODbiomass/g CODacetate, 2 mg C/L of acetate was added to reactor influent to serve as the sole electron donor. These operating parameters were defined as the baseline operating condition for this system. In a “weak backwash”, reactor content was stirred in 100-mL of previously collected reactor effluent in a 600-mL beaker with a 7.5-cm long magnetic stir bar at 75 revolutions per min (RPM) for 1 min, and then the supernatant was decanted. In a “strong backwash”, the reactor content was stirred in 125 mL previously collected effluent at 150 RPM twice for 1 min each time, with supernatant decanting and new effluent replenishment between these periods. The reactor was backwashed about once a month before the daily weak backwash experiment started. Six days after the daily weak backwash experiment, the daily strong backwash experiment was conducted.
Influent and effluent DO concentrations were measured using WTW multi340 m with CellOx325 sensors in WTW D201 flow cells (WTW Inc., Weilheim, Germany) connected to the inlet and outlet of the reactor. All other chemical parameters were measured according to Standard Methods (American Public Health Association (APHA) 1992). Perchlorate was measured using an ion chromatograph system (Dionex ICS-2000, Sunnyvale, CA) according to EPA Standard Method 314.1. An AS-16 analytical column and an AG-16 guard column were used, while the eluent was 65 mM KOH. The detection limits for DO and perchlorate were 0.01 mg/L and 1 μg/L, respectively.
Data analyses on reactor performance
Mass balance calculations on chemical concentrations (i.e., DO and perchlorate) were performed for the biofilm reactor. The governing equation was
where C represents the concentration of either DO or perchlorate in the reactor, C in and C out are the influent and effluent concentrations, V is the volume of the reactor, Q is the flow rate, and r represents the DO or perchlorate removal rate inside the reactor due to microbial activities. For the calculations presented here, completely stirred tank reactor conditions were assumed and therefore:
As a result, the governing equation was converted to:
where T is the empty bed contact time of 23.4 min or 0.01625 day. The initial removal rate r, for both DO and perchlorate, after a backwash was calculated using the last equation. The value of \( \frac{{d{C_{\text{out}}}}}{{dt}} \) was estimated by fitting measured C out vs. t using a second-order polynomial trendline and calculating the derivative of the trendline at the time of backwash. C in was 3 mg/L for DO or 50 μg/L for perchlorate. The C out at the time immediately after backwash was obtained from the corresponding trendline equations.
eDNA and iDNA extractions and protein assay
For clone library construction, biomass samples were collected from the biofilm reactor on day 0 of the daily weak backwash experiment and day 5 of the strong backwash experiment. Total DNA was extracted from these two biomass samples using FastDNA SPIN Kit (Qbiogene Inc., Irvine, CA). For qPCR measurements, biomass samples were collected from backwash water at all 11 backwash events. The biomass remaining in the reactor after each backwash was not sampled because the size of the reactor was small and daily sampling over the course of the experiment would have significantly changed the amount of biofilm in the reactor. eDNA was separated from cellular materials and extracted by following a published protocol (Corinaldesi et al. 2005). iDNA was extracted from the eDNA-free cellular materials by following a published protocol (Griffiths et al. 2000). All DNA extracts were quantified using a NanoDrop ND1000 (Thermo Scientific, Waltham, MA), and their qualities were evaluated using electrophoresis on a 1% agarose gel. Protein measurements were conducted using a protein assay kit (Bio-Rad, Hercules, CA).
Clone library
The bacterial 16S rRNA gene was amplified in triplicate PCR reactions on a Mastercycler (Eppendorf International, Hamburg, Germany) with the forward primer 8F (5′-AGA GTT TGA TCC TGG CTC AG-3′) and the reverse primer 1492R (5′-GG[C/T] TAC CTT GTT ACG ACT T-3′) (Dojka et al. 1998; Richardson et al. 2002). The composition of the PCR solutions and the conditions of the PCR reactions were adopted from the literature (Dojka et al. 1998). The 30-cycle PCR reaction started with 12 min of denaturation at 94°C and ended with a final extension at 72°C for 12 min. Each cycle consisted of denaturation at 94°C for 1 min, annealing at 50°C for 45 s, and extension at 72°C for 2 min. Pooled PCR products from triplicates were purified using electrophoresis on a 1% agarose gel and extracted using a MinElute Gel Extraction Kit (QIAGEN Inc., Valencia, CA). Purified PCR products were cloned into pCR4-TOPO vectors (Invitrogen Inc., Carlsbad, CA), and transformed into chemically competent Escherichia coli. The transformed E. coli cells were plated on Luria–Bertani agar that contained 50 μg/mL kanamycin and were incubated at 37°C overnight. Colonies were randomly picked and used to inoculate three 96-well microplates. Two of the three microplates in glycerol stocks were sent to the Genomic Center at Washington University (St. Louis, MO) for sequencing.
Phylogenetic analyses
A total of 384 clones (four 96-well microplates) were sequenced using vector primers T3 and T7. Nucleotide sequences were analyzed and edited using BioEdit (Hall 1999). Closely related sequences (i.e., Dechloromonas-, Zoogloea-, and Curvibacter-like clones in both clone libraries) identified via the Ribosomal Database Project (RDP) (Cole et al. 2007) were aligned using ClustalW (Chenna et al. 2003) for bacterial 16S rRNA genes. The differences between the corresponding genera in the two clone libraries were evaluated using the Library Comparison function on RDP. Sequences identified as chimeras using Mallard (Ashelford et al. 2006) were excluded from further analyses. The phylogenetic tree was based on a region of the 16S rRNA gene of ca. 600 bp starting at the 8F primer region and was created based on the evolutionary history inferred using the neighbor-joining method (Saitou and Nei 1987) and the evolutionary distance computed using the Jukes–Cantor method (Jukes and Cantor 1969) incorporated in the software program MEGA (Tamura et al. 2007). The percentages of replicate trees in which the associated taxa clustered together in the bootstrap test (1,000 replicates) were shown next to the branches (Felsenstein 1985). There were a total of 500 sites in the final dataset used to build the phylogenetic tree. Operational taxonomic units (OTUs) were defined as clones that shared 95% identity in the 16S rRNA gene (Fields et al. 2005). OTUs, diversity statistics, and rarefaction curves were calculated using DOTUR (Schloss and Handelsman 2005). The similarities of the bacterial 16S rRNA gene clone libraries were evaluated using ∫-LIBSHUFF (Schloss et al. 2004).
qPCR primer design and reaction conditions
qPCR primer sets were designed based on representative sequences of the clones of interests (i.e., Dechloromonas, Zoogloea, and Curvibacter) using the Primer3 program (Rozen and Skaletsky 2000) made available by Integrated DNA Technologies (www.idtdna.com) and were synthesized by Invitrogen (Carlsbad, CA). The lengths of the amplicons using the three primer sets ranged from 106 to 155 bp. The specificities of the designed primer sets were manually verified using the Probe Match function of RDP (Cole et al. 2007), while the coverage of the designed primer sets were evaluated against the clones of interest in relevant clone libraries using OligoReport (http://www.bioinformatics-toolkit.org/Cowslip/index.html) (Table 1). The specificities of the designed primer sets were further characterized with various ranges of annealing temperatures using the gradient PCR function of a real-time PCR thermocycler Mastercycler realplex system (Eppendorf International, Hamburg, Germany) (Fig. S1). The selected annealing temperatures for the three designed primer sets (Table 1) could differentiate the fluorescence signals between equal amounts of target and non-target templates (i.e., 106 copies/μL) by at least 15 C threshold units (Fig. S1). The PCR amplicons from the characterization experiments using the primer set Dchm0991F/1146R were also tested by electrophoresis on a 1% agarose gel (Fig. S2). The gel image showed the correlation between fluorescence signals from qPCR and band intensity on an agarose gel. The specificity of the designed primer sets were further tested by monitoring the melting temperatures from the melting curves on target and non-target templates (Fig. S3). The qPCR primer set Bact1369F/1492R for quantifying total bacteria was adopted from Suzuki et al. (2000).
All qPCR experiments were performed using a RealMasterMix SYBR Green Kit (Eppendorf International, Hamburg, Germany), which had a self-adjusting chelating mechanism to control Mg2+ concentrations. The reaction mixtures in a 25-μL final volume contained 11.25 μL of 2.5× RealMasterMix SYBR Green solution (including 0.05 U/μL HotMaster Taq DNA polymerase, 10 mM magnesium acetate, 1.0 mM dNTPs, and 2.5× SYBR Green solution), 150 nM of forward and reverse primers, Sigma water (Sigma-Aldrich, St. Louis, MO), and target/non-target template of known concentrations or 10 ng of DNA template from environmental samples. All qPCR experiments included Sigma water as a non-template control to assure identification of false positives. As suggested in the manual of the RealMasterMix SYBR Green kit, amplification involved one cycle of 95°C for 10 min for initial denaturation and then 40 cycles of 95°C for 15 s followed by annealing at the temperatures reported in Table 1 for 20 s and extension at 68°C for 30 s. Detection of SYBR Green fluorescence was set at the extension step of each cycle. Melting profiles were collected after 40 cycles of amplification to check the specificity of the amplification.
qPCR standard curves
Target and non-target templates were plasmid DNA extracted from clones obtained from the clone libraries constructed in the current study using QIAprep Miniprep Kit (QIAGEN Inc., Valencia, CA) and quantified using NanoDrop ND1000. The target templates contained the representative sequences based on which corresponding primer sets were designed, while the non-target templates contained sequences with the least number of mismatches with the designed primer sets in the relevant clone libraries (Table S1). Dilutions of purified E. coli plasmid DNA containing the 16S rRNA genes of Dechloromonas, Zoogloea, and Curvibacter were used as standards for qPCR in the quantification of specific populations. The plasmid DNA concentrations (copies/microliter) were calculated using the following equation
where 3,956 and 1,484 are the lengths of the pCR®4-TOPO vector (Invitrogen Corporation, Carlsbad, CA) and PCR insert, the constants of 607.4 and 157.9 were obtained for the calculation of the molecular weight of double stranded DNA (www.ambion.com).
The standard curve constructed from triplicate measurements for each qPCR primer set was linear between “C threshold” and “Log Target Concentration (copy # per microliter)” (Fig. S4). The R 2 values ranged from 0.993 to 0.995 (Table S1). Among the four primer sets used, Bact1369F/1492R had the narrowest linear range: the C threshold values for the standards of 101 and 102 copies per microliter fell outside the linear range and thus were not included in the standard curve. The linear ranges in Fig. S4 covered all the measurements from unknown samples. The high y-intercept value for Bact1369F/1492R reported in Table S1 indicates a lower detection limit at a given cycle number (Ritalahti et al. 2006). The amplification efficiencies (E) were calculated using the equation, \( E = {10^{{( - 1/{\text{slope}})}}} - 1 \) , where the slope is the slope of a standard curve. The amplification efficiencies for the four primer sets ranged from 0.627 (Bact1369F/1492R) to 0.990 (Zoog0487F/0627R, Table S1). To enumerate target copy numbers in environmental samples, standard curves were measured in triplicates on each 96-well microplates to avoid discrepancies across microplates.
Sequence submission
The sequences included in the phylogenetic tree have been submitted to the DDBJ/EMBL/GenBank databases under accession numbers FJ610473-FJ610667.
Results
Extracellular DNA
Because extracellular DNA (eDNA) is deemed important for biofilm studies due to its high abundance in certain natural environments (Corinaldesi et al. 2005; Steinberger et al. 2002) and its sequence similarity to intracellular DNA (iDNA) in certain biofilms (Steinberger and Holden 2005), we investigated whether the presence of eDNA would affect the 16S rRNA gene sequence analyses and the quantification of total DNA extracted for the fixed-bed biofilm reactor. The mass ratios of eDNA to total DNA for the biomass samples, collected in two sets of backwash experiments (i.e., daily weak and daily strong backwash experiments, see below), are presented in Fig. 1. The average percentages were 20.6 ± 4.6% (n = 6) and 24.8 ± 13.7% (n = 4, excluding the sample on day 1 of the daily strong backwash experiment) for the daily weak and daily strong backwash experiments, respectively. Statistical analyses showed no significant difference between the two sets (p = 0.610), indicating that the microbial community inside the biofilm reactor contained similar percentages of eDNA under different backwash intensities.
The ratios of eDNA to total DNA in biomass samples collected from the daily weak backwash (gray) and daily strong backwash experiments (dark). Due to contamination, the ratio for day 1 of the daily strong backwash experiment (hatched pattern) was not included in the reported average percentages in the text. The insert shows the relative abundances of the three dominant genera in the iDNA and “eDNA” extracts of that particular sample, suggesting iDNA contamination in the eDNA extract
qPCR results showed that eDNA extracts from the biofilm reactor did not contain 16S rRNA gene sequences that were amplifiable by the four primer sets used in this study (data not shown). Among the four primer sets, three were designed in this study to target the three dominant bacterial genera in the reactor (Table 1), and the other one was a published primer set targeting the bacterial domain (Suzuki et al. 2000). The only exception was the eDNA extract from day 1 of the daily strong backwash experiment, which was amplified by all four primer sets (insert in Fig. 1). This eDNA extract likely was contaminated with iDNA, because qPCR results showed the relative abundances of the three targeted genera (i.e., Dechloromonas, Zoogloea, and Curvibacter) in this eDNA extract to be very similar to those in the corresponding iDNA extract. Since the eDNA samples did not contain sequences that were amplifiable with the four PCR primer sets, separate analyses of eDNA and iDNA were not pursued in subsequent experiments.
Microbial community
To study the linkages among reactor operation, reactor performance, and microbial community structure, two sets of backwash experiments were conducted in sequence: a daily weak backwash experiment and a daily strong backwash experiment. Two 16S rRNA gene clone libraries were constructed to study the microbial community in the biofilm reactor: one for the biomass sample collected on day 0 of the daily weak backwash experiment, and one for the biomass sample collected on day 5 of the daily strong backwash experiment (Table 2). In both libraries, Betaproteobacteria was the most abundant class. In the first clone library, the two most abundant bacterial genera were Dechloromonas and Zoogloea. The most abundant genus in the second library was Curvibacter. Therefore, the relative abundances of these three genera were monitored using qPCR throughout the two backwash experiments. A phylogenetic tree was constructed for the clones that belonged to these three genera in the two clone libraries (Fig. 2). Clones associated with Dechloromonas in both clone libraries clustered together and were closely related to Dechloromonas sp. HZ and Dechloromonas sp. JM. The clones associated with Zoogloea and Curvibacter in both libraries also clustered together.
Phylogenetic tree of the 16S rRNA gene clones that were closely associated with the three major bacterial genera present in the biofilm reactor. “Day 0” and “Day 5” refer to day 0 of the daily weak backwash experiment and day 5 of the daily strong backwash experiment, respectively. The numbers in parentheses are the numbers of clones
Genus level analyses revealed changes in the overall community structure before and after the two daily backwash experiments. In Fig. 3, the Chao1 and ACE indices increased from 60 and 75 to 81 and 92, respectively, indicating the microbial richness increased during the backwash experiments. The same trend was supported also by the rarefaction curves in Fig. 3. In contrast, the Shannon–Weiner and Inverse Simpson's microbial diversity indices decreased from 2.77 and 11.51 to 2.40 and 4.73, respectively, due to the decrease in evenness. In addition, clone library results showed that the structures of the microbial communities before and after the backwash experiments were different (Table 2). This observation was statistically supported using ∫-LIBSHUFF (p < 0.05).
Rarefaction curves indicating bacterial 16S rRNA gene richness within the two clone libraries. The dashed line represents 1:1, indicating infinite diversity. The table lists the bacterial 16S rRNA gene sequence diversity indices. OTUs were defined as groups of sequences sharing 95% 16S rRNA gene sequence identity. The estimates of phylotype richness were calculated according to the abundance-based coverage estimate (ACE) and the bias-corrected Chao1 estimator. The Shannon–Weiner diversity index and the Inverse Simpson's diversity index, which consider both richness and evenness, were also calculated
Daily weak backwash experiment
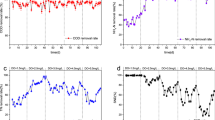
Before the daily weak backwash experiment started, the fixed-bed biofilm reactor had been operated using the baseline operating conditions for 388 days. The reactor was backwashed once every month and was able to consistently remove dissolved oxygen (DO) and perchlorate to below their respective detection limits of 0.01 mg/L and 1 μg/L (data not shown). The daily weak backwash initially caused disturbances in reactor performance, but a gradual recovery was observed (Fig. 4a, b). Immediately after the weak backwash on day 1, effluent DO started at about 0.5 mg/L and then dropped to 0.04 mg/L at the end of the day. During the next 4 days, the effluent DO concentration immediately after each backwash decreased from about 0.5 mg/L on day 2 to about 0.05 mg/L on day 5. To better illustrate the improvement of reactor performance in lowering effluent DO concentration during the course of the experiment, the initial DO removal rate after each weak backwash was plotted in Fig. 4a.
Effluent DO (top row) and perchlorate (bottom row) concentrations during the daily weak backwash experiment (left column) and the daily strong backwash experiment (right column). (Filled diamond) Effluent DO concentration; (empty diamond) initial DO removal rate; (filled circle) effluent perchlorate concentration; (empty circle) initial perchlorate removal rate; the solid lines are second-order polynomial trendlines to help visualize the improvements of DO and perchlorate removal rates immediately after backwash events. The arrows point to the time when backwash events occurred
A similar trend was noted for the effluent perchlorate profiles (Fig. 4b). On day 1, the effluent perchlorate concentration immediately after the weak backwash was close to the influent perchlorate concentration (i.e., 50 μg/L), and then decreased throughout the day. After each subsequent daily weak backwash, the effluent perchlorate concentration dropped faster than the day before. To better illustrate the improvement in perchlorate removal, the initial perchlorate removal rate after each weak backwash was calculated and plotted in Fig. 4b.
The amount of biomass removed was the highest for day 0 of the daily weak backwash experiment, because the reactor had been backwashed only once a month before this backwash experiment and had accumulated a large amount of biomass (Fig. 5a). Assuming 55% of bacterial dry weight is made of protein and excluding the datum point from day 0 of the daily weak backwash experiment, protein assay results suggested the average amounts of biomass washed out were 2.32 and 2.71 mg dry weight in weak and strong backwashes, respectively. With an assumed net yield of 0.2 g CODbiomass/g CODacetate, the dry weight of the biomass generated every day was calculated to be 2.66 mg, indicating that the daily biomass growth in the reactor approximately equaled the daily biomass loss through backwash.
Using qPCR, it was determined that the three most abundant bacterial populations responded differently to the daily weak backwashes. The relative abundance of Dechloromonas first decreased from 27.3% on day 0 to 2.5% on day 3, and then increased to 12.1% on day 5 (Fig. 5b). The relative abundance of Zoogloea first increased from 26.7% on day 0 to 36.1% on day 2, and then decreased to 15.6% on day 5. The relative abundance of Curvibacter increased from 19.8% on day 0 to 40.3% on day 5.
Daily strong backwash experiment
The bioreactor was not backwashed for 6 days after the daily weak backwash experiment. At the end of the 6 days, the reactor exhibited consistent and complete removal of DO and perchlorate (data not shown). Then, the reactor was subjected to strong backwashes for five consecutive days. The increase in backwash intensity dramatically affected reactor performance (Fig. 4c, d). Immediately after the strong backwash on day 1, the effluent DO concentration was 0.01 mg/L and remained low throughout the day. During the next few days, the effluent DO concentration immediately after each backwash increased as the experiment proceeded, except for day 4. Since the results of a replicate experiment with the same reactor (Fig. S5) also indicated a continuously increasing trend of effluent DO similar to that in Fig. 4c, the effluent DO concentration on day 4 was not included when calculating the trendline for the initial DO removal rate after each strong backwash. This trendline demonstrated a deteriorating ability of the microbial community to lower DO.
A decreasing ability to remove perchlorate was also observed during the daily strong backwash experiment. The effluent perchlorate concentration immediately after each strong backwash was close to the influent perchlorate concentration, 50 μg/L (Fig. 4d). The reactor was able to achieve complete perchlorate removal at the end of day 1. Starting on day 2, the ability of the reactor to remove perchlorate deteriorated. On day 5, the effluent perchlorate concentration remained at a level close to the influent perchlorate concentration throughout the day. The trendline for the initial perchlorate removal rate after each strong backwash showed that the ability of the reactor to remove perchlorate had decreased.
Similar to the daily weak backwash experiment, the three major bacterial genera responded differently to the strong backwashes as determined using qPCR. The relative abundance of Dechloromonas decreased from 7.0% on day 1 to 0.5% on day 5 (Fig. 5d). The relative abundance of Zoogloea first increased from 9.0% to 14.3% during the first 2 days, and then decreased to 2.4% on day 5. The relative abundance of Curvibacter increased from 60.9% on day 1 to 97.5% on day 5.
Discussion
Three qPCR primer sets, targeting strains that belong to the genera Dechloromonas, Zoogloea, and Curvibacter, were designed based on the sequences of the corresponding clones in two 16S rRNA gene clone libraries, instead of using all sequences for the relevant genera available in pubic databases. Thus, the designed primers are not strictly genus-specific primers. Such design demands lower stringency on primer coverage and warrants higher specificity. Thus, it is not surprising that even the closest non-targets for some designed primers had more than seven mismatches (Table S1). The primer sets were characterized to obtain optimized annealing temperatures for qPCR experiments using known concentrations of plasmids that were obtained from the clone libraries and contained target or non-target sequences.
Dechloromonas spp. and Azospira spp. were the only known PRB detected in the reactor. The dominance of Dechloromonas in perchlorate reducing systems has been shown by other studies in which acetate (Zhang et al. 2005) or hydrogen (Nerenberg et al. 2008) was used as the sole electron donor. In the biofilm reactor studied by Zhang and coworkers, about 23% of the microbes on the surface of the biofilm were detected to be Dechloromonas strains. In the hydrogen-fed membrane biofilm reactors (MBfR) studied by Nerenberg and coworkers, the relative abundance of Dechloromonas comprised 49% of the bacterial community. The Dechloromonas-like clones recovered in this study clustered together and were found to be closely related to Dechloromonas sp. HZ and Dechloromonas sp. JM, both of which are PRB and can utilize acetate as an electron donor (Zhang et al. 2002; Logan et al. 2001). Other PRB species, such as Dechlorospirillum spp. and Azospirillum spp., which were isolated from various natural and engineered systems (Bardiya and Bae 2008; Waller et al. 2004), were not detected in the current study.
The presence of Zoogloea spp. in the biofilm reactor is not surprising, as members of this genus have been characterized as slime formers in engineered systems (Rossellomora et al. 1995). Zoogloea spp. are aerobic (Xie and Yokota 2006; Unz 1984) and thus must have utilized DO as their electron acceptor in the current system. The genus Curvibacter has been described only relatively recently (Ding and Yokota 2004), and knowledge about the ecological function of Curvibacter spp. is limited, except that they have been detected in freshwater (Ding and Yokota 2004; Ding and Yokota 2010), Antarctic coastal waters (Gentile et al. 2006), and activated sludge (Thomsen et al. 2007).
Both reactor performance and PRB population abundance recovered from the disturbance imposed by the higher backwash frequency. Before the backwash frequency was increased from once a month to once a day, effluent perchlorate concentration was constantly below the detection limit. After the change, effluent perchlorate was high after each backwash event and then decreased for the rest of the day. Interestingly, across different days, perchlorate removal rate continuously improved (Fig. 4b); however, the relative abundance of Dechloromonas spp. showed a down-and-up trend (Fig. 5b). The discrepancy in the trends of the relative abundance of PRB and perchlorate removal suggests that bacterial population(s) other than Dechloromonas spp. impacted perchlorate removal efficiency. Such bacterial population(s) likely affected perchlorate removal by consuming DO, the competing electron acceptor, in the bioreactor. When the bulk DO level dropped towards the end of the daily weak backwash experiment, the DO level in the biofilm must have decreased accordingly. The decreased DO level in the biofilm could have been responsible for the increase in the relative abundance of Dechloromonas during that time period. In contrast, neither the reactor performance nor the PRB population recovered from the increase in backwash intensity. Reactor performance in terms of perchlorate removal and the relative abundance of Dechloromonas spp. continuously dropped during the daily strong backwash experiment (Figs. 4d and 5). Although the amount of biomass within the reactor could not be monitored experimentally, it was expected to be relatively constant over the course of the study, as the daily biomass growth approximately equaled the biomass loss through daily backwash.
In conclusion, after a minor disturbance in reactor operation, i.e., an increase in backwash frequency, the major PRB population (Dechloromonas spp.) was able to recover, and reactor performance (perchlorate removal) recovered gradually. In comparison, after a major disturbance in reactor operation, i.e., an increase in backwash intensity, Dechloromonas spp. failed to remain competitive in the microbial community, Curvibacter spp. became the dominant population, and consequently reactor performance deteriorated. To our knowledge, this is the first study designed to systematically investigate how two backwash parameters, frequency and intensity, affect the structure and function of microbial communities in biofilm reactors. Although different from those used in full-scale facilities, the backwash procedure used in this study allowed us to carefully control the two backwash parameters and to ensure high reproducibility between individual backwash events. The conclusions from this study, particularly the finding that increasing the backwash frequency disturbed the microbial community less compared to raising backwash intensity, are expected to be helpful to the development and optimization of backwash procedures at full-scale facilities.
References
Ahmad R, Amirtharajah A, Al-Shawwa A, Huck PM (1998) Effects of backwashing on biological filters. J Am Water Works Ass 90(12):62–73
Alm EW, Oerther DB, Larsen N, Stahl DA, Raskin L (1996) The oligonucleotide probe database. Appl Environ Microbiol 62(10):3557–3559
American Public Health Association (APHA) AWWAA, Water Environment Federation (WEF) (1992) Standard methods for the examination of water and wastewater, 18th edn. American Public Health Association, Washington, DC
Ashelford KE, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ (2006) New screening software shows that most recent large 16S rRNA gene clone libraries contain chimeras. Appl Environ Microbiol 72(9):5734–5741
Bardiya N, Bae JH (2008) Isolation and characterization of Dechlorospirillum anomalous strain JB116 from a sewage treatment plant. Microbiol Res 163(2):182–191
Bouwer EJ, Crowe PB (1988) Biological processes in drinking water treatment. J Am Water Works Ass 80(9):82–93
Briones A, Raskin L (2003) Diversity and dynamics of microbial communities in engineered environments and their implications for process stability. Curr Opin Biotechnol 14(3):270–276
Brosius J, Dull TJ, Sleeter DD, Noller HF (1981) Gene organization and primary structure of a ribosomal-RNA operon from Escherichia coli. J Mol Biol 148(2):107–127
Brown JC, Snoeyink VL, Raskin L, Lin R (2003) The sensitivity of fixed-bed biological perchlorate removal to changes in operating conditions and water quality characteristics. Water Res 37(1):206–214
CA-DHS (2005) Perchlorate drinking water action level and regulation. California Department of Health Services, http://www.dhs.ca.gov/ps/ddwem/chemicals/perchl/perchlindex.htm
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res 31(13):3497–3500
Choi YC, Li X, Raskin L, Morgenroth E (2007) Effect of backwashing on perchlorate removal in fixed bed biofilm reactors. Water Res 41(9):1949–1959
Choi YC, Li X, Raskin L, Morgenroth E (2008) Chemisorption of oxygen onto activated carbon can enhance the stability of biological perchlorate reduction in fixed bed biofilm reactors. Water Res 42(13):3425–3434
Coates JD, Achenbach LA (2004) Microbial perchlorate reduction: rocket-fuelled metabolism. Nat Rev Microbiol 2(7):569–580
Coates JD, Michaelidou U, Bruce RA, O'Connor SM, Crespi JN, Achenbach LA (1999) Ubiquity and diversity of dissimilatory (per)chlorate-reducing bacteria. Appl Environ Microbiol 65(12):5234–5241
Cole JR, Chai B, Farris RJ, Wang Q, Kulam-Syed-Mohideen AS, McGarrell DM, Bandela AM, Cardenas E, Garrity GM, Tiedje JM (2007) The ribosomal database project (RDP-II): introducing myRDP space and quality controlled public data. Nucleic Acids Res 35:D169–D172
Corinaldesi C, Danovaro R, Dell'Anno A (2005) Simultaneous recovery of extracellular and intracellular DNA suitable for molecular studies from marine sediments. Appl Environ Microbiol 71(1):46–50
Curtis TP, Head IM, Graham DW (2003) Theoretical ecology for engineering biology. Environ Sci Technol 37(3):64a–70a
Ding L, Yokota A (2010) Curvibacter fontana sp. nov., a microaerobic bacteria isolated from well water. J Gen Appl Microbiol 56:267–271
Ding LX, Yokota A (2004) Proposals of Curvibacter gracilis gen. nov., sp nov and Herbaspirillum putei sp nov for bacterial strains isolated from well water and reclassification of [Pseudomonas] huttiensis, [Pseudomonas] lanceolata, [Aquaspirillum] delicatum and [Aquaspirillum] autotrophicum as Herbaspirillum huttiense comb. nov., Curvibacter lanceolatus comb. nov., Curvibacter delicatus comb. nov and Herbaspirillum autotrophicum comb. nov. Int J Syst Evol Microbiol 54:2223–2230
Dojka MA, Hugenholtz P, Haack SK, Pace NR (1998) Microbial diversity in a hydrocarbon- and chlorinated-solvent-contaminated aquifer undergoing intrinsic bioremediation. Appl Environ Microbiol 64(10):3869–3877
Felsenstein J (1985) Confidence-limits on phylogenies—an approach using the bootstrap. Evolution 39(4):783–791
Fields MW, Yan TF, Rhee SK, Carroll SL, Jardine PM, Watson DB, Criddle CS, Zhou JZ (2005) Impacts on microbial communities and cultivable isolates from groundwater contaminated with high levels of nitric acid-uranium waste. FEMS Microbiol Ecol 53(3):417–428
Gentile G, Giuliano L, D'Auria G, Smedile F, Azzaro M, De Domenico M, Yakimov MM (2006) Study of bacterial communities in Antarctic coastal waters by a combination of 16S rRNA and 16S rDNA sequencing. Environ Microbiol 8(12):2150–2161
Greer MA, Goodman G, Pleus RC, Greer SE (2002) Health effects assessment for environmental perchlorate contamination: the dose response for inhibition of thyroidal radioiodine uptake in humans. Environ Health Perspect 110(9):927–937
Griffiths RI, Whiteley AS, O'Donnell AG, Bailey MJ (2000) Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA- and rRNA-based microbial community composition. Appl Environ Microbiol 66(12):5488–5491
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hozalski RM, Bouwer EJ (1998) Deposition and retention of bacteria in backwashed filters. J Am Water Works Ass 90(1):71–85
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian Protein Metabolism. Academic, New York, pp 21–132
Lahav O, Artzi E, Tarre S, Green M (2001) Ammonium removal using a novel unsaturated flow biological filter with passive aeration. Water Res 35(2):397–404
Li X, Upadhyaya G, Yuen W, Brown J, Morgenroth E, Raskin L (2010) Changes in the structure and function of microbial communities in drinking water treatment bioreactors upon addition of phosphorus. Appl Environ Microbiol 76(22):7473–7481
Lin R, M.S. thesis, (2004) Bacterial community analysis and optimization of biologically active carbon filters used to remove perchlorate from groundwater. University of Illinois, Urbana-Champaign
Logan B, Zhang H, Mulvaney P, Milner M, Head I, Unz R (2001) Kinetics of perchlorate- and chlorate-respiring bacteria. Appl Environ Microbiol 67(6):2499–2506
MA-DEP (2006) Drinking Water Status on Perchlorate by MA Department of Environmental Protection. http://www.mass.gov/dep/water/drinking/percinfo.htm#stds
Miller JP, Logan BE (2000) Sustained perchlorate degradation in an autotrophic, gas-phase, packed-bed bioreactor. Environ Sci Technol 34(14):3018–3022
Nerenberg R, Kawagoshi Y, Rittmann BE (2008) Microbial ecology of a perchlorate-reducing, hydrogen-based membrane biofilm reactor. Water Res 42(4–5):1151–1159
Nerenberg R, Rittmann BE (2004) Hydrogen-based, hollow-fiber membrane biofilm reactor for reduction of perchlorate and other oxidized contaminants. Water Sci Technol 49(11–12):223–230
Richardson RE, Bhupathiraju VK, Song DL, Goulet TA, Alvarez-Cohen L (2002) Phylogenetic characterization of microbial communities that reductively dechlorinate TCE based upon a combination of molecular techniques. Environ Sci Technol 36(12):2652–2662
Ritalahti KM, Amos BK, Sung Y, Wu QZ, Koenigsberg SS, Loffler FE (2006) Quantitative PCR targeting 16S rRNA and reductive dehalogenase genes simultaneously monitors multiple Dehalococcoides strains. Appl Environ Microbiol 72(4):2765–2774
Rittmann BE, McCarty PL (2001) Environmental biotechnology: principles and applications. McGraw-Hill, New York, NY
Rossellomora RA, Wagner M, Amann R, Schleifer KH (1995) The abundance of Zoogloea ramigera in sewage treatment plants. Appl Environ Microbiol 61(2):702–707
Rozen S, Skaletsky H (2000) Primer3 on the WWW for general users and for biologist programmers. Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, NJ
Saitou N, Nei M (1987) The neighbor-joining method—a new method for reconstructing phylogenetic trees. Mol Biol Evol 4(4):406–425
Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71(3):1501–1506
Schloss PD, Larget BR, Handelsman J (2004) Integration of microbial ecology and statistics: a test to compare gene libraries. Appl Environ Microbiol 70(9):5485–5492
Servais P, Billen G, Ventresque C, Bablon GP (1991) Microbial activity in GAC filters at the Choisy-le-Roi treatment plant. J Am Water Works Ass 83(2):62–68
Steinberger RE, Allen AR, Hansma HG, Holden PA (2002) Elongation correlates with nutrient deprivation in Pseudomonas aeruginosa-unsaturated biofilms. Microb Ecol 43(4):416–423
Steinberger RE, Holden PA (2005) Extracellular DNA in single- and multiple-species unsaturated biofilms. Appl Environ Microbiol 71(9):5404–5410
Suzuki MT, Taylor LT, DeLong EF (2000) Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5′-nuclease assays. Appl Environ Microbiol 66(11):4605–4614
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24(8):1596–1599
Thomsen TR, Kong Y, Nielsen PH (2007) Ecophysiology of abundant denitrifying bacteria in activated sludge. FEMS Microbiol Ecol 60(3):370–382
Unz RF (1984) Genus IV. Zoogloea Itzigsohn 1868, 30AL. In: Krieg NR, Holt JG (eds) Bergey's manual of systematic bacteriology, vol 1. Williams & Wilkins, Baltimore, pp 214–219
Upadhyaya G, Jackson J, Clancy TM, Hyun SP, Brown J, Hayes KF, Raskin L (2010) Simultaneous removal of nitrate and arsenic from drinking water sources utilizing a fixed-bed bioreactor system. Water Res 44(17):4958–4969
Urbansky ET (2002) Perchlorate as an environmental contaminant. Environmental Science and Pollution Research 9(3):187–192
US-EPA (2005) State Perchlorate Advisory Levels and Other Resources. http://www.epa.gov/fedfac/documents/perchlorate_links.htm
US-EPA (2008) Drinking Water Contaminant Candidate List 3 http://www.epa.gov/safewater/ccl/ccl3.html
Waller A, Cox E, Edwards E (2004) Perchlorate-reducing microorganisms isolated from contaminated sites. Environmental Microbiology 6(5):517–527
Xie CH, Yokota A (2006) Zoogloea oryzae sp nov., a nitrogen-fixing bacterium isolated from rice paddy soil, and reclassification of the strain ATCC 19623 as Crabtreella saccharophila gen. nov., sp nov. Int J Syst Evol Microbiol 56:619–624
Xu J, Song Y, Min B, Steinberg L, Logan B (2003) Microbial degradation of perchlorate: principles and applications. Environmental engineering Science 20(5):405–422
Zhang H, Bruns M, Logan B (2002) Perchlorate reduction by a novel chemolithoautotrophic, hydrogen-oxidizing bacterium. Environ Microbiol 4(10):570–576
Zhang H, Logan BE, Regan JM, Achenbach LA, Bruns MA (2005) Molecular assessment of inoculated and indigenous bacteria in biofilms from a pilot-scale perchlorate-reducing bioreactor. Microb Ecol 49(3):388–398
Acknowledgements
The project was supported by the National Science Foundation (BES-0123342) and the Department of Defense (ESTCP: ER-0544). The authors would like to thank Young Chul Choi and Jess Brown for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, X., Yuen, W., Morgenroth, E. et al. Backwash intensity and frequency impact the microbial community structure and function in a fixed-bed biofilm reactor. Appl Microbiol Biotechnol 96, 815–827 (2012). https://doi.org/10.1007/s00253-011-3838-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-011-3838-6