Abstract
In forests worldwide, Quercus is a major genus; however, the boundaries between the constituent species are relatively weak, and hybridization is reported frequently. In this study, we examined Quercus acuta and Quercus sessilifolia (subgenus Cyclobalanopsis), which have a putative hybrid—Q. x takaoyamensis. We investigated leaf morphological traits and microsatellites of Q. acuta and Q. sessilifolia in the area where the two species are both found. Although the leaf traits overlapped, the two species could be distinguished morphologically as demonstrated by principal component analysis based on a range of these traits. They were also genetically differentiated, with F ST = 0.104. However, they shared most of the alleles at all eight loci examined, and considerable genetic admixing was detected. Admixture analysis demonstrated that Q. acuta and Q. sessilifolia, respectively, contained 11 and 24 % of individuals with a probability of less than 0.9 of being correctly assigned to their species. Model-based testing showed that this admixing was created by not only shared ancestral polymorphism but also by hybridization. Effective population size and migration rate were estimated using the coalescent approach. We estimated 8.843 and 71.98 effective numbers of migrants per generation to Q. acuta and Q. sessilifolia, respectively. Theoretically, one to ten migrants per generation are required to prevent complete genetic differentiation. Based on the results of this study, it appears that genetic admixing, with sharing of most alleles, is probably common in the two species and is maintained by interspecific introgressive hybridization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
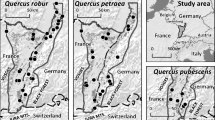
Quercus is a widely distributed tree genus. Its species boundaries are as weak as those of Populus (Martinsen et al. 2001; Stettler et al. 1996), Betula (Johnsson 1945; Palme et al. 2004), and Eucalyptus (Field et al. 2011; Potts and Dungey 2004), and genetic admixtures, considered to be due to interspecific hybridization, have been reported frequently all over the world (Cavender-Bares and Pahlich 2009; Guichoux et al. 2013; Lepais et al. 2009; Matsumoto et al. 2009; Moran et al. 2012; Zeng et al. 2011). Thus, it can be used as a model genus for studying hybridization. However, some pairs of Quercus species, although they are morphologically and genetically differentiated from each other, share almost all their alleles (Moran et al. 2012; Muir and Schlotterer 2005; Valbuena-Carabana et al. 2005). In these cases, it is possible that a shared ancestral polymorphism rather than hybridization is responsible for the genetic admixture between the species. In the case of Quercus robur and Quercus petraea, this possibility has been discussed (Muir and Schlotterer 2005), and finally, it has been concluded that hybridizations have truly occurred (Abadie et al. 2012; Guichoux et al. 2013; Lexer et al. 2006; Scotti-Saintagne et al. 2004). As hybridization is common in the genus Quercus, there is no doubt that the evolutionary dynamics of the genus, including speciation and range expansion, have been affected by hybridizations (Petit et al. 2003; Zeng et al. 2011).
The genus Quercus can be separated into two subgenera—Quercus and Cyclobalanopsis—based on whether the cupule is imbricate-scaled or lamellate. Subgenus Quercus is found across the northern hemisphere and contains ca. 300 species, while Cyclobalanopsis is found from the Himalayas and Southeast Asia to China and Japan and contains ca. 150 species (Huang et al. 1999). The leaf morphologies of subgenus Cyclobalanopsis trees resemble each other more than those of subgenus Quercus (Yan and Zhe-Kun 2002). Unlike subgenus Quercus, there have, to our knowledge, been no studies of hybridization using genetic makers in subgenus Cyclobalanopsis.
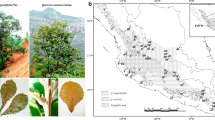
Our study species, Quercus acuta Thunb. and Quercus sessilifolia Blume, both belong to subgenus Cyclobalanopsis and are late successional species with evergreen leaves; they are found in warm temperate forest. Q. acuta, with oblong or elliptic leaves, 2–4 cm long petiole, a long acuminate leaf tip, cuneate or rounded leaf base, and an entire leaf margin, occurs in Korea and Japan (Ohashi et al. 2006; Ohba 2006). Q. sessilifolia, with narrowly oblong or elliptic-oblanceolate leaves, 0.4–1.2 cm long petiole, acute leaf tip, cuneate or attenuate leaf base, and nearly entire leaf margin with sparse small teeth, is found in Taiwan, China, and Japan (Ohba 2006). These two species grow in a somewhat higher elevation zone than other Cyclobalanopsis species; Q. acuta prefers upper and Q. sessilifolia lower slopes (Ito et al. 2007). Previous studies that investigated the cpDNA haplotypes of species of the subgenus Cyclobaranopsis in Japan found that Q. acuta and Q. sessilifolia exhibited the same haplotype (Ohyama et al. 1999, 2001). Based on leaf morphology, Quercus x takaoyamensis Makino, which is the putative hybrid between Q. acuta and Q. sessilifolia, has been identified in a mixed stand of the two parental species (Kobayashi and Midorikawa 1959; Makino 1920; Yamashita et al. 1999). These previous studies reported that the leaf morphology of Q. x takaoyamensis was intermediate between Q. acuta and Q. sessilifolia and highly variable. However, no studies investigating the details of its morphological and genetic traits have been published, and it is still unclear whether there actually is hybridization between Q. acuta and Q. sessilifolia.
The main objective of this study was to clarify the existence and the degree of interspecific hybridization between Q. acuta and Q. sessilifolia in the wild. First, we investigated morphological and genetic differences between Q. acuta and Q. sessilifolia using five leaf morphological traits and eight microsatellite markers. Second, the degree of genetic admixing of the two species was examined by means of Bayesian admixture analysis. Third, we investigated whether the genetic admixture is only the result of shared ancestral polymorphism or a combination of shared ancestral polymorphism and interspecific hybridization. Finally, in order to quantify the degree of interspecific hybridization, the migration rate between the two species was estimated.
Materials and methods
Study sites and sampling
We thoroughly searched habitats where Q. acuta and Q. sessilifolia occur in Gifu Prefecture, central Japan, and selected 15 sites where at least two individuals of one or both species were found (Fig. 1 and Table 1). Numbers of individuals were very low because the species are rare in this region. All trees taller than 2 m were selected for examination at each site, with the exception of Tsurusato (site number 14), where we examined only 30 individuals randomly selected from both species because they were so abundant. We sampled 3 to 4 shoots from 75 Q. acuta and 178 Q. sessilifolia individuals. Leaf specimens were collected for species identification and so that we could record the morphological traits. Three leaves picked from the sampled shoots were dried with silica gel and stored at room temperature while awaiting DNA extraction.
Measurement of morphological traits and data analysis
Initially, species statuses of individuals were identified subjectively based on the detailed observation of the specimens collected. Next, in order to confirm them objectively, leaf shapes, the length, and width of leaves (mm), petiole length (mm), shape of the leaf tip and base, and the absence or presence of leaf teeth were recorded for ten leaves per tree using the specimens collected. Leaf ratio was calculated by dividing length by width. The shape of the leaf tip was scored 0, 0.5, or 1, representing acute, intermediate, or acuminate, respectively. The shape of the leaf base was scored 0, 0.5, or 1, representing cuneate, intermediate, or obtuse, respectively. The status of leaf teeth was scored 0 or 1, representing without or with teeth, respectively. Five morphological traits, leaf ratio, petiole length, leaf tip, leaf base, and leaf teeth, were averaged for each individual tree. Principal component analysis (PCA) was conducted based on the five morphological traits, using the princomp function in R version 2.15.1 (R Development Core Team 2012). A correlation matrix was used for calculating eigenvalues to normalize the variables.
DNA extraction and microsatellite analysis
We selected seven genomic [QpZAG9, QpZAG110, and QpZAG119 (Steinkellner et al. 1997); QrZAG7, QrZAG20, and QrZAG101 (Kampfer et al. 1998); QM69-2M1 (Isagi and Suhandono 1997)] and seven EST microsatellite markers [QmC00141, QmC00898, QmC00963, QmC01133, QmC01368, and QmC02241 (Ueno et al. 2008); CR627959 (Ueno and Tsumura 2008)] developed for the subgenus Quercus. All were successfully amplified in our study species. Multiplex PCR was conducted using a Type-it Mnicrosatellite PCR Kit (QIAGEN, Hilden, Germany), following the manufacturers’ instructions, with a TaKaRa PCR Thermal Cycler (TaKaRa, Shiga, Japan). The length of the PCR products was scored with an ABI PRISMTM 310 Genetic Analyzer (Applied Biosystems, CA, USA), and allele size was determined using GENESCAN analysis software version 3.7 (Applied Biosystems, CA, USA). Quality of loci, deviation from Hardy-Weinberg equilibrium (HWE), genotyping error, and the fraction of null alleles for the 14 loci were checked using MicroChecker version 2.2.3 (Van Oosterhout et al. 2004). We also tested whether the loci were non-neutral, showing higher levels of interspecific genetic differentiation than expected if there was neutrality, using fdist2 (Beaumont and Nichols 1996). Finally, we used eight loci (QpZAG9, QrZAG7, QrZAG20, QrZAG101, QmC00141, QmC00963, QmC01368, and QmC02241; Table 2), which did not deviate from the assumption of HWE in both species, for which there was no evidence of genotype error or null alleles in both species, which satisfied the assumptions indicating selective neutrality and which contained enough genetic diversity to distinguish between the two species (gene diversity >0.1; see details in Supplementary material).
Genetic data analysis
Number of alleles and gene diversity of the eight loci were calculated using FSTAT version 2.9.3 (Goudet 2001). Genetic differentiations between the two species at the eight loci were tested by permutation tests not assuming HWE, implemented in FSTAT.
Bayesian admixture analysis was performed using STRUCTURE version 2.3.4 for estimating the genetic admixture rates of individuals (Falush et al. 2003; Pritchard et al. 2000). To determine the optimal number of clusters (K), ten independent runs were replicated for each value of K from 1 to 12, and the ΔK statistic was calculated using the CorrSieve package in R (Campana et al. 2011; Evanno et al. 2005). Independent runs were performed with a burn-in period of 500,000 and 500,000 subsequent Markov chain Monte Carlo (MCMC) steps. The ten independent results for each K value were merged into one using CLUMPP version 1.1.2 (Jakobusson and Rosenberg 2007).
In order to determine whether the observed genetic admixture is just the result of genetic drift or a combination of genetic drift and gene flow between the two species, relative roles of drift and gene flow were assessed using the 2MOD program that was based on coalescent theory (Ciofi et al. 1999). The QrZAG101 locus was excluded because it consisted of compound microsatellite repeats, the evolutionary processes of which are complex (Bull et al. 1999), and so, it was not ideal for the coalescent analysis. Seven loci were thus used in this analysis. 2MOD evaluates (1) the drift model, which assumes that drift is the sole reason for population divergence since a certain time, i.e., the observed genetic admixture was created by only shared ancestral polymorphisms, and (2) the gene flow model, which assumes immigration-drift equilibrium of gene frequencies within populations, i.e., the observed genetic admixture was created not only by shared ancestral polymorphisms but also by interspecific hybridization. Three independent MCMC simulations were conducted with 100,000 steps; 10,000 steps were discarded as the burn-in period and every fifth step was sampled. Ultimately, we used 54,000 samples, and the probabilities of each model [P (drift) and P (gene flow)] were calculated.
In order to quantify the interspecific gene flow, BayesAss version 3.0 (Wilson and Rannala 2003) and Migrate-n version 3.3.2 (Beerli 2006; Beerli and Felsenstein 2001) were used. Both programs are able to detect asymmetric migration events. The former can estimate the recent migration rate (over the last several generations), and the latter can estimate the historical migration rate based on coalescent theory. In the BayesAss analysis, asymmetric migration rates were estimated by Bayesian methods. Three replicate runs consisting of 1 million burn-in steps and a further 2 million MCMC steps were performed with different random number seeds, and posteriors were sampled every 2,000 steps. We confirmed that approximately the same posterior distributions were obtained from each of the three replicate runs. Trace plots for all parameters were produced, and their convergences were checked. The 95 % higher posterior density (95 % HPD) was calculated using the coda package in R (Plummer et al. 2006). In the Migrate-n analysis, because this is based on coalescent theory, we used seven loci, excluding the QrZAG101 locus, as was the case with the 2MOD analysis. The mutation-scaled effective population size (θ = 4N eμ) and migration rate (M = m/μ) were estimated by Bayesian methods. N e, m, and μ represent the effective population size, migration rate and mutation rate, respectively. Prior uniform distributions ranging from 0 to 100 and from 0 to 120 were used for θ and M, respectively, with starting values estimated from F ST. Relative mutation rates were estimated from the data, and the Brownian motion model was employed because the exact stepwise mutation model was extremely time-consuming and did not converge. The Metropolis-Hasting algorithm was used to generate posterior distributions. Three replicate runs consisting of 1 million burn-in steps and a further 200,000 MCMC steps were performed with different random number seeds, and posteriors were sampled every 100 steps. We confirmed that approximately the same posterior distributions were obtained from each of the three replicate runs. Trace plots for all parameters were produced, and their convergences were checked. A static heating scheme using 32 chains with the exponential increase heating term option was employed. Posterior distributions of parameters over all loci were calculated by averaging the randomly drawn values from the posterior distributions of each of the seven loci as empirical distributions, adjusting mutation rates among the loci using estimated relative mutation rates. The 95 % HPD was calculated, as was the case with the BayesAss analysis. Symmetric migration (migration rates were symmetrical), one-way migration, and no migration models were compared to the full model (asymmetric migration) to determine the direction of interspecific gene flow. The Bezier approximation and harmonic mean method were used to marginalize the likelihood, and we calculated log Bayes factors against the most parsimonious model [log (marginal likelihood of the most parsimonious model)–log (marginal likelihood of the model)] and determined which model was the best fit for the data (Beerli and Palczewski 2010).
Results
Morphological data
Although the distributions of each of the five leaf traits differed between the two species, there was some overlap for all of them (Fig. 2). However, using a combination of the first two principal components (PC) derived on the basis of the five leaf traits, the two species could be distinguished (Fig. 3). Although there was little overlap for PC 1, the distributions for the two species were very close. Distributions for PC 2 overlapped completely.
Distribution of principal components (PC) based on the five leaf morphological traits. Circles and squares represent individuals of Quercus acuta and Q. sessilifolia, respectively. Genetic statuses were estimated by the program STRUCTURE, based on the genotypes of eight microsatellite markers. P Qac and P Qse are the probabilities of belonging to Q. acuta and Q. sessilifolia, respectively
Genetic differentiation and genetic admixing
For the eight loci, although the ranges of the alleles overlapped and the two species shared most of the alleles (Fig. 4), genetic differentiations were highly significant for all eight loci (Table 2). Numbers of alleles of Q. acuta were lower than those of Q. sessilifolia for all eight loci. Gene diversities of Q. acuta were also lower than those of Q. sessilifolia for six of the eight loci.
In the Bayesian admixture analysis, the average value of the log probability of data plateaued at K = 4 to 8, then decreased with increasing values of K (Fig. 5). ΔK peaked at K = 2 and K = 4, indicating that these K values were appropriate to consider (Fig. 6), as secondary peak of the ΔK statistic may indicate the existence of within-species genetic structure. If such genetic structure is ignored, it can reduce the potential for species assignment. Thus, we used the results of K = 4 in the following analyses. Because the admixture coefficients q 41 and q 42, and q 43 and q 44 roughly correspond to Q. acuta and Q. sessilifolia, the sum of q 41 and q 42, and the sum of q 43 and q 44 could be considered to represent the probability of an individual belonging to Q. acuta (P Qac = q 41 + q 42) or Q. sessilifolia (P Qse = q 43 + q 44), respectively (Fig. 6). The values of P Qac and P Qse varied greatly within each species (Fig. 7). The average values of P Qac and P Qse for Q. acuta were 0.938 and 0.062, while those for Q. sessilifolia were 0.131 and 0.869, respectively. Of the individuals Q. acuta and Q. sessilifolia, 11 and 24 %, respectively, had a probability of less than 0.9 of being correctly assigned to their species. These two species thus showed similar levels of genetic admixing. P Qac and P Qse values for individual trees of the two species were compared with the results of the PCA (Fig. 3). However, individuals that showed genetic admixing were uniformly distributed across PC 1 and PC 2, and no clear relationships could be detected.
Population history and migration
The 2MOD population history test strongly supported a gene flow model rather than a drift model [P (gene flow) = 1.000, P (drift) = 0.000], indicating that interspecific hybridization significantly contributed to the observed genetic admixture between the two species.
In the detection of recent migration events by BayesAss, the medians (95 % HPD) of the posterior distributions of the migration rates from Q. sessilifolia to Q. acuta and from Q. acuta to Q. sessilifolia were 0.140 (0.099–0.187) and 0.086 (0.060–0.116), and their 95 % HPDs overlapped.
In the detection of historical migration events by Migrate-n, the most parsimonious model was that based on symmetric migration; models with different migration parameters were compared against the most parsimonious model using the log Bayes factor (LBF; Table 3). For all the models, LBFs were larger than 21.3 and 128.5 for the Bezier approximation and harmonic mean methods, respectively. When the Bayes factor for each model compared to the most parsimonious model is >150 (LBF against the most parsimonious model was greater than log (150) ≈ 5.0), support for the most parsimonious model is overwhelming (Link and Barker 2010). In our analysis, since LBF > > 5 in both methods, the symmetric migration model was strongly supported.
Medians (95 % HPD) of the posterior distribution of mutation-scaled population size (θ = 4N eμ) in Q. acuta and Q. sessilifolia were 2.433 (2.070–2.958) and 18.52 (15.30–21.49), respectively (Fig. 8a). The median (95 % HPD) of the posterior distribution of mutation-scaled migration rate (M = m/μ) was 14.19 (12.86–15.84; Fig. 8b). Posterior distributions of the effective number of migrants per generation (N e m) were calculated by θ × M/4, and medians (95 % HPD) of the values from Q. sessilifolia to Q. acuta and from Q. acuta to Q. sessilifolia were 8.843 (7.362–11.15) and 71.98 (56.64–84.72), respectively (Fig. 8c).
Discussion
What caused the genetic admixture?
Although Q. acuta and Q. sessilifolia were genetically and morphologically differentiated, they were genetically admixed in the selectively neutral genomic regions studied. Interspecific hybridization is one of the possible causes of this. In the genus Quercus, genetic admixing has often been reported over its distribution range and is considered to be the result of hybridization (Cavender-Bares and Pahlich 2009; Guichoux et al. 2013; Lepais et al. 2009; Matsumoto et al. 2009; Moran et al. 2012; Zeng et al. 2011). In the studied species, most of the alleles at each locus were shared between the two species, and the ranges of the alleles were almost entirely overlapping. These distributions of alleles suggest that this admixture may be the result of a shared ancestral polymorphism (Muir and Schlotterer 2005). If this admixture was created by only shared ancestral polymorphism, the underlying mechanism could be considered to be pure genetic drift after speciation. We compared pure drift model with a gene flow model (drift + gene flow), and the gene flow model was strongly supported. Despite the fact that these two species are phylogenetically very close to each other (Ohyama et al. 1999, 2001) and, therefore, the possibility that the shared ancestral polymorphism is the main cause of this genetic admixture that cannot be completely rejected, the result of the model comparison indicated that the interspecific gene flow significantly affected the genetic admixing of the two species. As with the species in our study, the European oaks Q. robur and Q. petraea exhibit genetic admixing, with most of their alleles shared, and this has been attributed to shared ancestral polymorphism at one time (Muir and Schlotterer 2005). However, increasing evidence based on genome scans and detailed artificial pollination experiments (Abadie et al. 2012; Guichoux et al. 2013; Lepais et al. 2013; Lexer et al. 2006; Scotti-Saintagne et al. 2004) supports the possibility that hybridization accounts for shared polymorphisms between these oak species. The results of previous studies and our study suggest that genetic admixing with the sharing of many alleles as a result of hybridization can be considered a common occurrence in the genus Quercus (Moran et al. 2012; Muir and Schlotterer 2005; Valbuena-Carabana et al. 2005).
Characteristics of hybridization between the two oaks
In both species, admixed individuals exhibited a very variable admixture rate. Moreover, genetic admixtures were observed even at the site where only Q. sessilifolia grew. As indicated by the results of the BayesAss and Migrate-n analyses, the genetic admixture observed in this study could have been the result not only of recent hybridization but also of several occurrences of historical hybridization and, thus, introgression. Admixed individuals exhibited a range of morphological traits. This could be because no loci used in this study violate the assumption of selective neutrality since, although introgression can occur between the selectively neutral genomic regions as a result of hybridization, this cannot occur in the genomic regions affected by natural selection that produced functional differences, even when interspecific gene flow occurs frequently (Butlin 2010; Via 2009). Thus, the two species could remain morphologically distinct. Most of the populations in the study region consisted of small numbers of individuals. This may have limited intraspecific pollination, and interspecific hybridization can be an effective way to ensure the reproduction. If this is so, the estimated admixture rates in this study may differ from rates in other regions where there are enough individuals of each species to minimize levels of hybridization.
A putative hybrid of the two species, Q. x takaoyamensis, is known, identified by its intermediate leaf morphology (Makino 1920). It has been reported that the leaf morphology of Q. x takaoyamensis is not stable and is highly variable among individuals (Kobayashi and Midorikawa 1959; Yamashita et al. 1999). Not all individuals with the intermediate genetic admixture rate exhibited intermediate morphological traits (PC 1), and thus, there was no clear congruence between the leaf morphology and genetic status. Although hybrid individuals between Q. acuta and Q. sessilifolia, i.e., Q. x takaoyamensis, are sure to exist in the wild as shown by the result of our genetic analyses, it is difficult to distinguish them from pure Q. acuta and Q. sessilifolia individuals on the basis of leaf morphology.
No significant asymmetric migration between the two species was detected in association with either recent or historical migration events. Generally, there are some differences in the strength of barriers to interspecific gene flow between species, and therefore, asymmetric hybridization occurs (Arnold 1997; Lewis and Crowe 1958; Tiffin et al. 2001). In the genus Quercus, some cases of asymmetric hybridization have been reported: For example, in Q. robur and Q. petraea, there were differences in fertilization success (Steinhoff 1993), and in Q. robur, Q. petraea, Quercus pubescens, and Quercus pyrenaica, post-zygotic barriers were the key determinant of the direction of hybridization (Lepais et al. 2013). In our study, although migration rates were symmetric across the whole region, at some sites (1, 6, and 14), the two species exhibited different levels of admixing. A more detailed study is required to reveal the direction of hybridization between Q. acuta and Q. sessilifolia.
In the coalescent-based historical migration analysis, although the mutation-scaled migration rate (M = m/μ) was fixed between the two species, the mutation-scaled effective population sizes (θ = 4N eμ) were different, and the value of θ for Q. sessilifolia was ca. 8 times larger than that for Q. acuta. As a result, effective numbers of migrants per generation (N e m) were different, and the values from Q. sessilifolia to Q. acuta and from Q. acuta to Q. sessilifolia were 8.843 and 71.98, respectively. Sewall Wright’s one migrant per generation rule states that, in ideal populations, one migrant per generation is enough to prevent complete population differentiation (Wright 1931). However, in real populations with factors in play other than those accounted by genetic theory, one to ten migrants per generation are considered necessary (Mills and Allendorf 1996; Wang 2004). The values of N e m observed in this study are equal to (N e m into Q. acuta) or larger than (N e m into Q. sessilifolia) 10, suggesting that there is sufficient hybridization to prevent complete genetic differentiation between the species and that the genetic condition of most alleles being shared between the species has been maintained. In order to confirm hybridization between the two species, artificial pollination experiments or paternity and parentage analyses to directly detect recent hybridizations are required.
Conclusions
Q. acuta and Q. sessilifolia exhibited genetic admixing, and we consider that the cause is not simply shared ancestral polymorphism but also interspecific hybridization. Hybridizations have occurred several times, and as a result, there has been introgression. Although these two species are genetically and morphologically differentiated, genetic admixture rates do not correspond to the observed morphology. This suggests that genomic regions related to morphology have been strongly affected by natural selection. N e m has been sufficient to prevent complete genetic differentiation, and the sharing of most of the alleles has been maintained by introgressive interspecific gene flow.
References
Abadie P, Roussel G, Dencausse B, Bonnet C, Bertocchi E, Louvet J-M, Kremer A, Garnier-Gere P (2012) Strength, diversity and plasticity of postmating reproductive barriers between two hybridizing oak species (Quercus robur L. and Quercus petraea (Matt) Liebl.). J Evol Biol 25:157–173
Arnold ML (1997) Natural hybridization and evolution. Oxford University Press, New York
Beaumont MA, Nichols RA (1996) Evaluating loci for use in the genetic analysis of population structure. Proc R Soc B 263:1619–1626
Beerli P (2006) Comparison of Bayesian and maximum-likelihood inference of population genetic parameters. Bioinformatics 22:341–345
Beerli P, Felsenstein J (2001) Maximum likelihood estimation of a migration matrix and effective population sizes in n subpopulations by using a coalescent approach. Proc Natl Acad Sci 98:4563–4568
Beerli P, Palczewski M (2010) Unified framework to evaluate panmixia and migration direction among multiple sampling locations. Genetics 185:313–326
Bull LN, Pabon-Pena CR, Freimer NB (1999) Compound microsatellite repeats: practical and theoretical features. Genome Res 9:830–838
Butlin RK (2010) Population genomics and speciation. Genetica 138:409–418
Campana MG, Hunt HV, Jones H, White J (2011) Corrsieve: software for summarizing and evaluating Structure output. Mol Ecol Resour 11:349–352
Cavender-Bares J, Pahlich A (2009) Molecular, morphological, and ecological niche differentiation of sympatric sister oak species, Quercus virginiana and Q. geminata (Fagaceae). Am J Bot 96:1690–1702
Ciofi C, Beaumont MA, Swingland IR, Bruford MW (1999) Genetic divergence and units for conservation in the Komodo dragon Varanus komodoensis. Proc R Soc B 266:2269–2274
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of cluster of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Field DL, Ayre DJ, Whelan RJ, Young AG (2011) Patterns of hybridization and asymmetrical gene flow in hybrid zones of the rare Eucalyptus aggregata and common E. rubida. Heredity 106:841–853
Goudet J (2001) FSTAT, a program to estimate and test genetic diversities and fixation indices (version 2.9.3). Available from <http://www2.unil.ch/popgen/softwares/fstat.htm>.
Guichoux E, Garnier-Gere P, Lagache L, Lang T, Boury C (2013) Outlier loci highlight the direction of introgression in oaks. Mol Ecol 22:450–462
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59:1633–1638
Huang C, Zhang Y, Bartholomew B (1999) Fagaceae. In: Wu Z-Y, Raven PH (eds) Flora of China, vol 4. Science Press, Beijing, pp 314–400
Isagi Y, Suhandono S (1997) PCR primers amplifying microsatellite loci of Quercus myrsinifolia Blume and their conservation between oak species. Mol Ecol 6:897–899
Ito S, Ohtsuka K, Yamashita T (2007) Ecological distribution of seven evergreen Quercus species in southern and eastern Kyushu, Japan. Veg Sci 24:53–63
Jakobusson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Johnsson H (1945) Interspecific hybridization within the genus Betula. Hereditas 31:163–176
Jost L (2008) G ST and its relatives do not measure differentiation. Mol Ecol 17:4015–4026
Kampfer S, Lexer C, Glossl J, Steinkellner H (1998) Characterization of (GA)n microsatellite loci from Quercus robur. Hereditas 129:183–186
Kobayashi Y, Midorikawa T (1959) Dendrological studies of the Japanese Fagaceae: On the ripening term of the fruits of Quercus, Castanopsis and Pasania. Bull Gov For Exp Station 117:11–42 (in Japanese)
Lepais O, Petit RJ, Guichoux E, Lavabre JE, Alberto F, Kremer A, Gerber S (2009) Species relative abundance and direction of introgression in oaks. Mol Ecol 18:2228–2242
Lepais O, Roussel G, Hubert F, Kremer A, Gerber S (2013) Strength and variability of postmating reproductive isolating barriers between four European white oak species. Tree Genet Genomes 9:841–853
Lewis D, Crowe LK (1958) Unilateral interspecific incompatibility in flowering plants. Heredity 12:233–256
Lexer C, Kremer A, Petit RJ (2006) Shared alleles in sympatric oaks: recurrent gene flow is a more parsimonious explanation than ancestral polymorphism. Mol Ecol 15:2007–2012
Link WA, Barker RJ (2010) Bayesian inference with ecological applications. Academic Press, London
Makino T (1920) A contributions to the knowledge of the flora of Japan. J Jpn Bot 2:13–16
Martinsen GD, Whitham TG, Turek RJ, Keim P (2001) Hybrid populations selectively filter gene introgression between species. Evolution 55:1325–1335
Matsumoto A, Kawahara T, Kanazashi A, Yoshimaru H, Takahashi M, Tsumura Y (2009) Differentiation of three closely related Japanese oak species and detection of interspecific hybrids using AFLP markers. Botany 87:145–153
Mills LS, Allendorf FW (1996) The one-migrant-per-generation rule in conservation and management. Conserv Biol 10:1509–1518
Moran EV, Willis J, Clark JS (2012) Genetic evidence for hybridization in red oaks (Quercus sect. Lobatae, Fagaceae). Am J Bot 99:92–100
Muir G, Schlotterer C (2005) Evidence for shared ancestral polymorphism rather than recurrent gene flow at microsatellite loci differentiating two hybridizing oaks (Quercus spp.). Mol Ecol 14:549–561
Ohashi H, Ohashi K, Takahashi K (2006) Identify of Quercus acuta Thunb. (Fagaceae) recorded from Taiwan and China. Jpn J Bot 81:268–274 (in Japanese)
Ohba H (2006) Fagaceae. In: Iwatsuki K, Boufford DE, Ohba H (eds) Flora of Japan, vol IIa. Kodansha Scientific, Tokyo, pp 42–60
Ohyama M, Baba K, Itoh T (1999) Possibility of grouping of Cyclobalanopsis species (Fagaceae) grown in Japan based on an analysis of several regions of chloroplast DNA. J Wood Sci 45:498–501
Ohyama M, Baba K, Itoh T (2001) Wood identification of Japanese Cyclobalanopsis species (Fagaceae) based on DNA polymorphism of the intergenic spacer between trnT and trnL 5' exon. J Wood Sci 47:81–86
Palme AE, Su Q, Palsson S, Lascoux M (2004) Extensive sharing of chloroplast haplotypes among European birches indicates hybridization among Betula pendula, B. pubescens and B. nana. Mol Ecol 13:167–178
Petit RJ, Bodenes C, Ducousso A, Roussel G, Kremer A (2003) Hybridization as a mechanism of invasion in oaks. New Phytol 161:151–164
Plummer M, Best N, Cowles K, Vines K (2006) CODA: convergence diagnosis and output analysis for MCMC. R News 6:7–11
Potts BM, Dungey HS (2004) Interspecific hybridization of Eucalyptus: key issues for breeders and geneticists. New For 27:115–138
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
R Development Core Team (2012) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Scotti-Saintagne C, Mariette S, Porth I, Goicoechea PG, Barreneche T, Bodenes C, Burg K, Kremer A (2004) Genome scanning for interspecific differentiation between two closely related oak species [Quercus robur L. and Q. petraea (Matt.) Liebl.]. Genetics 168:1615–1626
Steinhoff S (1993) Results of species hybridizaition with Quercus robur L and Quercus petraea (Matt) Liebl. Ann For Sci 50:137s–143s
Steinkellner H, Fluch S, Turetschek E, Lexer C, Streiff A, Kremer A, Burg K, Glossl J (1997) Identification and characterization of (GA/CT) n -microsatellite loci from Quercus petraea. Plant Mol Biol 33:1093–1096
Stettler RF, Zsuffa L, Wu R (1996) The role of hybridization in the genetic manipulation of Populus. In: Stettler RF, Bradshaw HD, Heilman PE, Hinckley TM (eds) Biology of Populus: and its implications for management and conservation, vol I. NRC Research Press, Ontario, pp 87–112
Tiffin P, Olson MS, Moyle LC (2001) Asymmetrical crossing barriers in angiosperms. Proc R Soc B 268:861–867
Ueno S, Taguchi Y, Tsumura Y (2008) Microsatellite markers derived from Quercus mongolica var. crispula (Fagaceae) inner bark expressed sequence tags. Genes Genet Syst 83:179–187
Ueno S, Tsumura Y (2008) Development of ten microsatellite markers for Quercus mongolica var. crispula by database mining. Conserv Genet 9:1083–1085
Valbuena-Carabana M, Gonzalez-Martinez SC, Sork VL, Soto A, Goicoechea PG (2005) Gene flow and hybridization in a mixed oak forest (Quercus pyrenaica Willd. and Quercus petraea (Matts.) Liebl.) in central Spain. Heredity 95:457–465
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Via S (2009) Natural selection in action during speciation. Proc Natl Acad Sci 106:9939–9946
Wang J (2004) Application of the one-migrant-per-generation rule to conservation and management. Conserv Biol 18:332–343
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191
Wright S (1931) Evolution in Mendelian populations. Genetics 16:97–159
Yamashita T, Nagai Y, Shoji T (1999) Quercus takaoyamensis Makino and Q. sessilifolia Blume (Fagaceae) at Jourakuji in Fuchu-machi, Toyama Prefecture. Bull Bot Gard Toyama 4:43–46 (in Japanese with English abstract)
Yan L, Zhe-Kun Z (2002) Leaf architecture in Quercus subgenus Cyclobalanopsis (Fagaceae) from China. Bot J Linn Soc 140:283–295
Zeng Y-F, Liao W-J, Petit RJ, Zhang D-Y (2011) Geographic variation in the structure of oak hybrid zones provides insights into the dynamics of speciation. Mol Ecol 20:4995–5011
Acknowledgments
We are grateful to N. Tomaru, S. Setsuko, N. Nakanishi, and Y. Hasegawa for their support with the genetic experiment; M. Tsuruta, T. Okada, Y. Watanabe, and H. Tabata for useful discussions; and N. Yanagisawa, K. Tsuda, and the members of the Laboratory of Satoyama for their help with the sample collections. We also thank the two anonymous reviewers for their helpful comments on the previous manuscript.
Conflict of interest
The authors declare that they have no conflict of interest.
Data archiving statement
We follow standard Tree Genetics and Genomes policy, and all geotype and phenotype data are deposited in the Dryad Repository doi:10.5061/dryad.c959s.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by I. J. Chybicki
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 166 kb)
Rights and permissions
About this article
Cite this article
Tamaki, I., Okada, M. Genetic admixing of two evergreen oaks, Quercus acuta and Q. sessilifolia (subgenus Cyclobalanopsis), is the result of interspecific introgressive hybridization. Tree Genetics & Genomes 10, 989–999 (2014). https://doi.org/10.1007/s11295-014-0737-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-014-0737-x