Abstract
In northern Japan, coastal oak forests consist of Quercus dentata (Qd) on the coastal side and Q. mongolica var. crispula (Qc) on the inland side. In the forests of northern Hokkaido, Qd is rare, and a coastal ecotype of Qc with some Qd-like traits grows on the coastal side. To reveal the genetic background of this ecotype, nuclear microsatellite genotypes in closely related oak taxa were obtained from the Eurasian continent, Sakhalin, and Hokkaido. The clustering of these genotypes suggests an admixture of Qd in the coastal ecotype of Qc. Next, we evaluated the effects of admixture and coastal stress on the leaf and shoot traits of Qc and Qd along coastal–inland gradients in northern Hokkaido. The admixture of Qd in Qc was quantified by the Qd ancestry proportions. Coastal stress causes bud mortality in the upper parts of shoots and was quantified by the survival patterns of buds in shoots. The genetic and environmental effects on the traits at Qd-abundant and Qd-rare sites were estimated using linear mixed models. The genetic effect was detected in all traits. Both genetic and environmental effects were detected in most traits. Some traits differed between Qd-abundant and Qd-rare sites in addition to these effects, indicating more Qd-like traits at Qd-rare sites. The findings suggest that an admixture of Qd characterizes the genetic background of the coastal ecotype of Qc and that not only the coastal stress but also the genetic background is responsible for the leaf and shoot traits of Qc and Qd in northern Hokkaido.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coastal vegetation changes along a gradient from beaches to inland areas, exhibiting a zonation of different plant communities (Ciccarelli 2015). Behind the foredune vegetation of grasses, sedges, and pioneer herbs, coastal forests are often established with stress-tolerant trees, such as pines, laurels, and oaks (Doing 1985). Environmental factors that affect this zonation are stress and disturbance, such as wind, salinity, drought, heat, substrate instability, and nutrient scarcity (Ciccarelli 2015). Oak trees in coastal forests suffer from strong winds and salt spray in winter (Shinmura et al. 1990). This stress causes bud mortality in the upper parts of shoots (Asai et al. 1986). As a consequence of this mortality, new shoots are regenerated from surviving buds in the lower parts of old shoots, resulting in the frequent dieback of shoots, slow elongation of stems, and low canopy height of coastal oak forests (Nagasaka et al. 1991; Usui et al. 1988).
In northern Japan, the zonation of Quercus dentata Thunberg (Qd) on the coastal side and Q. mongolica Fischer ex Ledebour var. crispula (Blume) H. Ohashi (synonym: Q. crispula Blume) (Qc) on the inland side characterizes coastal oak forests (Hasegawa 1984). In northern Hokkaido, the northernmost part of Japan (Fig. 1), Qd trees are rare in coastal oak forests (Shimizu 1997). In these forests, a coastal ecotype of Qc with unique traits, which are similar to Qd traits related to tolerance against coastal stress, grows on the coastal side (Shimizu et al. 1992, 1994, 1995). To explore how the unique traits of the coastal ecotype of Qc arise, we revealed the genetic background of this ecotype and examined the effects of this genetic background on its unique traits in the following two steps.
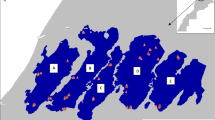
Sampling locations and study sites. aQuercus mongolica var. mongolica on the Eurasian continent (black squares), Q. mongolica var. crispula (Qc) on Sakhalin Island (gray squares), Qc in coastal Hokkaido (blue circles), Qc in inland Hokkaido (blue triangles), Q. dentata (Qd) in coastal Hokkaido (red circles), and Q. serrata in inland Hokkaido (green triangles). Inset shows northern Hokkaido. bQc in coastal habitats (blue circles) and inland habitats (blue triangles) and Qd in coastal habitats (red circles) at five Qd-abundant sites (red letters) and seven Qd-rare sites (blue letters) in northern Hokkaido
In the first step, we hypothesized that the genetic background of coastal Qc is (1) the same as that of inland Qc, (2) an admixture of Qd, and (3) an admixture of taxa other than Qd. Closely related oak species are interfertile (Matsumoto et al. 2009; Petit et al. 2004), and their hybrids usually have intermediate traits, resulting in an indistinct species boundary (Ishida et al. 2003; Ito 2009). Thus, an admixture of closely related oak taxa may alter the genetic background of coastal Qc, resulting in its unique traits. Some botanists have regarded coastal Qc as a putative hybrid between Qc and Qd (Ohba 2006), while others have believed that a part of coastal Qc belongs to Q. mongolica Fischer ex Ledebour var. mongolica (Qm) on the Eurasian continent (Miyabe and Kudo 1925). Because there is another oak species, Q. serrata Murray (Qs), in Hokkaido, this species also may be related to coastal Qc. Chloroplast DNA haplotypes are shared among Qc, Qd, and Qs (Kanno et al. 2004; Okaura et al. 2007) and between Qc and Qm (Aizawa et al. 2018), suggesting the admixtures among these taxa. On the other hand, nuclear microsatellites have indicated a similarity between inland and coastal Qc (Ohsawa et al. 2011), suggesting that inland and coastal Qc have the same genetic background.
In the second step, we examined whether the genetic background revealed in the first step affected leaf and shoot traits. In common gardens, some traits, such as the density and size of stellate hairs on lower leaf surfaces, the leaf mass per area, and the size and shape of leaves, are different between Qc and Qd and intermediate in their hybrids, suggesting genetic differentiation in these traits (Ubukata et al. 1996). On the other hand, these traits may change in response to environmental conditions along coastal–inland gradients due to phenotypic plasticity (Ramírez-Valiente et al. 2010). A unique trait of the coastal ecotype of Qc is an increased number of axillary buds at bud-scale scars in the lower parts of shoots (Shimizu et al. 1992, 1994, 1995). This trait seems to convey tolerance to coastal stress because more buds in the lower parts of shoots can compensate for bud mortality caused by coastal stress (Shimizu et al. 1992, 1994, 1995). With regard to such genetic and environmental effects on these traits, interactions between these effects, which are called reaction norms, may vary between inland and coastal Qc.
In this study, we aimed to clarify how the unique traits of the coastal ecotype of Qc arise. First, we investigated nuclear microsatellites in inland and coastal Qc and their related taxa to reveal the genetic background of coastal Qc. Second, we quantified their genetic ancestry using nuclear microsatellites and coastal stress based on the survival patterns of buds and measured leaf and shoot traits. Finally, we estimated both genetic and environmental effects on the traits and verified whether the genetic background affected the unique traits of the coastal ecotype of Qc.
Materials and methods
Study sites and samples
Samples of four oak taxa: Qm, Qc, Qd, and Qs, which potentially hybridize with each other (Okaura et al. 2007; Ubukata et al. 1999), were obtained from the Eurasian continent, Sakhalin Island, and Hokkaido Island (Fig. 1a) as follows.
Leaves and shoots of wild Qc and Qd trees were collected along coastal–inland gradients at 12 sites in northern Hokkaido from July to August in 2016 and 2017 (Fig. 1b). Among these sites, Qd is abundant at five southern sites and rare at seven northern sites (Fig. 1b). The gradient was categorized into five habitats: the (1) front, (2) middle, and (3) back of forests on coastal dunes or cliffs; (4) forests on coastal hills behind the dunes or cliffs; and (5) inland forests at the foot of mountains. At some sites, a few habitats were not investigated due to the lack of forests. Several trees were sampled from each habitat at each site. The sampled trees were more than 20 m apart from each other to avoid sampling closely related trees. The sampling locations were recorded using GPSMAP 64 (Garmin, Olathe, KS, USA). The collected leaves and shoots were dried and preserved in the Hokkaido Research Center of the Forestry and Forest Products Research Institute. We identified samples with hairy shoots as Qd and those with hairless shoots as Qc.
Additionally, we collected leaves of wild Qc trees from three inland locations in central Hokkaido. We also collected leaves of Qd trees that originated from 12 coastal locations in Hokkaido, which had been planted in the Bibai Experimental Forest of the Forestry Research Institute, Hokkaido Research Organization, in central Hokkaido. Furthermore, we collected leaves of Qm trees, which originated from three locations in northeastern China and then were planted in the Uryu Experimental Forest of Hokkaido University in northern Hokkaido (Kadomatsu 1997).
DNA was extracted from leaves dried with silica gel using the DNeasy Plant Mini Kit (Qiagen, Hilden, Germany) or the Maxwell RSC Plant DNA Kit (Promega, Madison, WI, USA). In addition, we obtained DNA samples of Qm from three locations in southern Primorsky Krai (Liu and Harada 2014), those of Qc from three locations in southern Sakhalin (Liu and Harada 2014; Okaura et al. 2007), and those of Qs from three inland locations in central Hokkaido (Kitamura et al. 2017) from these authors.
We arranged two sample sets, one from northeastern Asia (40–50˚N, 125–150˚E; Fig. 1a) and the other from northern Hokkaido (44–46˚N, 141–144˚E; Fig. 1b). The first sample set (number of samples, n = 198) consisted of six groups: (1) Qm on the Eurasian continent (n = 26), (2) Qc on the Sakhalin Island (n = 29), (3) Qc in coastal Hokkaido (i.e., the coastal ecotype of Qc), which were samples from the fore edge of coastal forests (habitat 1) at seven Qd-rare sites in northern Hokkaido (n = 32), (4) Qc in inland Hokkaido, which included samples from inland forests (habitat 5) at the 12 sites in northern Hokkaido and from the three inland locations in central Hokkaido (n = 48), (5) Qd in coastal Hokkaido, which included samples from habitat 1 at five Qd-abundant sites in northern Hokkaido and from the 12 coastal locations in Hokkaido (n = 31), and (6) Qs in inland Hokkaido (n = 32; Fig. 1a). The second sample set (n = 256) consisted of 182 Qc and 74 Qd samples from the 12 sites in northern Hokkaido (Fig. 1b).
Genetic analyses
We determined the genotypes of the two DNA sample sets at 22 loci of nuclear microsatellites (Table S1). These microsatellites were isolated from Qc (Mishima et al. 2006; Ueno et al. 2008; Ueno and Tsumura 2008), Castanopsis sieboldii (Ueno et al. 2009a), and Fagus crenata (Ueno et al. 2009b). Three unpublished loci developed by Ueno et al. (2009a, b) were also used (Table S1). We followed the methods used in these references to conduct polymerase chain reactions (PCRs). To determine multilocus genotypes, the base-pair length of PCR products was measured using an ABI PRISM 3100-xl Genetic Analyzer with POP-7 in 36 cm capillaries and GeneMapper 4.0 (Life Technologies, Foster City, CA, USA).
Bayesian clustering for the multilocus genotypes was conducted for the first sample set consisting of the six groups of Qm, Qc, Qd, and Qs in northeastern Asia to reveal their genetic backgrounds using Structure 2.3.1 (Falush et al. 2003). Next, clustering was performed for the second sample set of Qc and Qd in northern Hokkaido to examine their genetic variation along coastal–inland gradients at five Qd-abundant and seven Qd-rare sites. For each number of clusters ranging from 1 to 8, 20 independent runs were conducted. Each run included a burn-in length of 10,000 iterations and a sampling length of 20,000 iterations. We assumed clusters under the Hardy–Weinberg equilibrium, correlated allele frequencies, and asymmetric admixtures. An ancestry proportion of each cluster in each sample was obtained from a run with the maximum log likelihood for each number of clusters. To clarify the genetic background of the coastal ecotype of Qc, the ancestry proportions of individual clusters were compared between coastal and inland Qc in Hokkaido. The differences were tested with the Kruskal–Wallis rank-sum test using the kruskal.test function in R 3.3.2 (R Core Team 2016). A principal component analysis of all alleles at the 22 loci was also conducted for the six groups using the eigen function in R 3.3.2 (Patterson et al. 2006). Eigenvectors with much higher eigenvalues were also compared between coastal and inland Qc in Hokkaido, and the differences were tested in the same way.
We generated four populations, each of which consisted of samples with high (> 0.9) ancestry proportions of each cluster assigned to each taxon. To confirm whether the loci were independent with rare null alleles, the deviation from the Hardy–Weinberg equilibrium at each locus and linkage disequilibrium in every locus pair were tested in each population using GENEPOP 4.2 (Rousset 2008). To examine the genetic diversity of populations assigned to the clusters and their genetic divergence, the expected heterozygosity (HS) and fixation index (FIS) in each population and the genetic differentiation (FST) between them were estimated at each locus and across all loci using GENEPOP 4.2.
Morphological measurements
We selected three leaves of shoots collected from each tree sampled at the 12 sites in northern Hokkaido. For each leaf, the leaf length (mm) was measured, and the lateral veins were counted (Fig. 2). The dry weight (mg) of a leaf disc with a 9 mm diameter was also measured after dehydration at 60 °C for 48 h (Fig. 2). The stellate hairs on the lower surface were counted in a leaf area with a 3.1 mm diameter using a stereo microscope (Fig. 2). The angle (˚) of a tooth apex on a serration at the central part of a leaf was measured (Fig. 2).
We selected three 1-year-old shoots from each sampled tree. For each shoot, the diameter (mm) of the upper part was measured (Fig. 2). Buds of shoots were classified as a terminal bud and as axillary buds at upper stipule scars, at leaf scars, at lower stipule scars, and at bud-scale scars (Fig. 2).
Coastal stress was quantified as the proportion of flushing buds in the lower part of a shoot, m(m + n)−1, where m and n are the numbers of flushing buds in the lower and upper parts of a shoot, respectively. Buds in the upper part include the terminal bud, axillary buds at upper stipule scars, and the upper half of those at leaf scars. Buds in the lower part consist of axillary buds at bud-scale scars and at lower stipule scars and the lower half of those at leaf scars (Fig. 2). When there was an odd number of axillary buds at leaf scars, a living bud at the central position was divided into the upper and lower parts, and 0.5 was added to each value of m and n (Fig. 2). The lower flushing-bud proportion for each tree was the mean of values for three selected shoots.
Six morphological traits, stellate hair density, shoot diameter (mm), leaf mass per area, tooth apex angle (˚), lateral vein interval, and number of buds at bud-scale scars, were obtained from the measurements of leaves and shoots. The stellate hair density was calculated from the number of stellate hairs, h, in the observed area with the formula, (4/3.12)hπ−1 (mm−2). The leaf mass per area was calculated from the dry weight, w (mg), of the leaf disc with the formula, (4/0.92)wπ−1 (mg cm−2). The lateral vein interval was calculated from the number of lateral veins, v, and the leaf length, l (mm), with the formula, l(v + 1)−1 (mm). The trait value for each tree was the mean of values for three selected leaves or shoots.
Genetic and environmental effects on traits
The Kendall rank-correlation coefficients (τ) among the ancestry proportions of admixed clusters, the lower flushing-bud proportions, and the six morphological trait values were calculated and tested using the cor.test function in R 3.3.2.
To examine the genetic and environmental effects on each trait, we used a linear mixed model with an intercept, fixed effects of the ancestry proportion, the lower flushing-bud proportion, and their interaction, and random effects of sites. Models with the intercept and possible combinations of the fixed effects were prepared. The random effects were applied to possible combinations of the intercept and the fixed effects in each model. These models were fitted to the trait values of individual trees, which were assumed to follow a normal distribution, using the lmer function of the lme4 1.1–12 package (De Boeck et al. 2011) in R 3.3.2. Because the values of the five traits, except for the leaf mass per area, were skewed (Fig. S1), the values of the five traits were loge-transformed. Because the stellate hair density, x, and the number of buds at bud-scale scars, y, frequently were zero (Fig. S1), they were transformed with the formulae, loge(x + 1) and loge(y + 0.2), respectively.
A model with the minimum Akaike information criterion (AIC) value and some models with similar (< 2 differences) AIC values were selected from the examined models. Coefficients of the intercept, fixed effects, and random effects in the selected models were estimated using the lmer function. The regression lines of each trait in the observed ranges of ancestry proportions and lower flushing-bud proportions were obtained from the estimated coefficients at individual sites. These lines on the ancestry proportions were drawn in two cases: when the lower flushing-bud proportions were low (0.1) and high (0.9). The lines on the lower flushing-bud proportions were also drawn in two cases: when the ancestry proportions were low (0.1) and high (0.9). To test whether the random effects differ between Qd-abundant and Qd-rare sites, the Kruskal–Wallis rank-sum test was applied to the estimated coefficients of the random effects of sites using the kruskal.test function in R 3.3.2.
Qc samples were few in coastal habitats at Qd-abundant sites, and Qd samples were few at Qd-rare sites (Fig. S2). Thus, samples were biased on the coordinates of the ancestry proportions and lower flushing-bud proportions. To reduce this bias and examine the genetic and environmental effects on the traits within Qc samples in both coastal and inland habitats, we fitted the linear mixed models to not only all samples but also partial samples with < 0.5 ancestry proportions at Qd-rare sites (Fig. S2).
Results
Genetic background
Genotypes at 22 nuclear microsatellite loci were determined for the 198 DNA samples of the four oak taxa from the Eurasian continent, Sakhalin Island, and Hokkaido Island. Bayesian clustering for the multilocus genotypes resulted in an increasing median of log likelihood (L) with an increasing number of clusters from one (L = − 14,224) to five (L = − 12,855; Fig. S3a). An increase in the log likelihood was negligible from five to six clusters (L = − 12,853; Fig. S3a). When the number of clusters was higher than six, the log likelihood decreased (L ≤ − 12,929; Fig. S3a). At two clusters (L = − 13,547), the clusters were assigned to Qs and others, and admixtures between them were rare (Fig. 3). At three clusters (L = − 13,115), the clusters were assigned to Qs, Qd, and others, and admixtures of Qd in Qc in coastal Hokkaido were frequent (Fig. 3). At four clusters (L = − 12,929), the clusters corresponded to the four taxa, and admixtures of Qm in Qc were observed (Fig. 3). Admixtures of Qm in Qc in Sakhalin were more frequent than those in Hokkaido (Fig. 3). At five clusters, a cluster assigned to Qc was divided into two clusters corresponding to Qc in Sakhalin and Hokkaido, between which admixtures were frequent (Fig. 3). A principal component analysis for 233 alleles at the 22 loci of the 198 samples provided similar results (Fig. S4).
Ancestry proportions of clusters assigned to Q. mongolica var. mongolica (Qm, black), Quercus mongolica var. crispula (Qc) in Sakhalin (gray), Qc in Hokkaido (blue), Q. dentata (Qd, red), and Q. serrata (Qs, green) in samples of six groups: Qm on the Eurasian continent, Qc on Sakhalin Island, Qc in coastal Hokkaido, Qc in inland Hokkaido, Qd in coastal Hokkaido, and Qs in inland Hokkaido
In Hokkaido, the ancestry proportion of the Qd cluster was higher in coastal Qc than in inland Qc when the number of clusters (k) ranged from three to five (P < 0.001). On the other hand, no differences were detected in the ancestry proportions of clusters assigned to Qs (2 ≤ k ≤ 5), Qm (4 ≤ k ≤ 5), or Qc in Sakhalin (k = 5) between coastal and inland Qc (0.217 < P < 0.767). Eigenvectors of the principal component analysis showed similar results (Fig. S4b, c).
Populations corresponding to the four taxa were organized from samples with > 0.9 ancestry proportions of the four clusters assigned to Qm, Qc, Qd, and Qs (sample sizes: 23, 67, 24, and 30, respectively). Four of the 22 loci deviated from the Hardy–Weinberg equilibrium in one or two of the four populations (P < 0.049; Table S1). Linkage disequilibrium was detected in 11 of the 231 locus pairs in at least one population (P < 0.042). Thus, most of the loci were regarded as independent loci with rare null alleles. The four populations had low fixation indices across all loci (0.012 ≤ FIS ≤ 0.040; Table S1). Among the populations, genetic diversity was highest in Qc (HS = 0.697) and lowest in Qd (HS = 0.583; Table S1). Genetic differentiation was largest between Qd and Qs (FST = 0.211) and smallest between Qm and Qc (FST = 0.074; Table S1).
Distributions of Qc and Qd differed in coastal–inland gradients at the 12 sites in northern Hokkaido (Fig. 1b). At five Qd-abundant sites (01–03 and 11–12), most samples (65 of 76) in coastal forests (habitats 1–3) were identified as Qd. At seven Qd-rare sites (04–10), a few samples (7 of 94) in coastal forests (habitats 1–3) were identified as Qd. At the 12 sites, most samples (25 of 27) in forests on coastal hills (habitat 4) and all samples (59) in inland forests (habitat 5) were identified as Qc. The clustering for genotypes of the 256 samples at the 22 loci resulted in the maximum median of log likelihood at two clusters (L = − 17,052; Fig. S3b). When the number of clusters was greater than two, the log likelihood decreased (L ≤ − 17,191; Fig. S3b). The two clusters corresponded to Qc and Qd. The ancestry proportions of the Qd cluster varied within Qd-abundant sites and was low at Qd-rare sites (Fig. 4a). The Qd ancestry proportions were higher in coastal habitats than in inland habitats, the differences between which were much larger at Qd-abundant sites (Fig. 4b) than at Qd-rare sites (Fig. 4c).
Ancestry proportions of Quercus dentata (Qd) cluster (upper panels) and proportions of flushing buds in the lower parts of shoots (lower panels) at five Qd-abundant sites (red letters) and seven Qd-rare sites (blue letters) (a, d) and in five habitats along a coastal–inland gradient at Qd-abundant sites (b, e) and seven Qd-rare sites (c, f). In each plot, central lines indicate medians, boxes indicate first and third quartiles, and whiskers indicate ranges
Coastal stress and traits
We measured the leaf and shoot morphology of the 256 genotyped samples in northern Hokkaido. The proportions of flushing buds in the lower parts of shoots and the values of the six morphological traits were obtained from the samples (Fig. S1).
The lower flushing-bud proportion varied within sites and overlapped among the 12 sites (Fig. 4d). Along coastal–inland gradients, the lower flushing-bud proportion was often highest in the fore edge of coastal forests (habitat 1) and tended to decrease toward inland forests (habitat 5), although the range overlapped among habitats (Fig. 4e, f). The lower flushing-bud proportion was positively correlated with the Qd ancestry proportion (P < 0.001) due to the sampling bias (Fig. S2). However, the correlation coefficient between them was relatively low (τ = 0.151; Fig. S1).
The six morphological traits were unimodal, although the Qd ancestry proportion and lower flushing-bud proportion were bimodal (Fig. S1). The six traits were positively correlated with each other (0.119 ≤ τ ≤ 0.480, 0 < P < 0.005), with the Qd ancestry proportion (0.206 ≤ τ ≤ 0.412, 0 < P < 0.001), and with the lower flushing-bud proportion (0.094 ≤ τ ≤ 0.355, 0 < P < 0.028; Fig. S1).
Genetic and environmental effects on traits
Linear mixed models with possible combinations of genetic (the Qd ancestry proportion) and environmental (the lower flushing-bud proportion) effects, their interactions, and random site effects were applied to each of the six morphological traits of all samples. As a result, three types of models were selected (Table 1).
First, a model with site-specific intercepts and a positive genetic effect was selected for the stellate hair density (Table 1). This trait was mainly determined genetically, and the density increased as Qd ancestry increased (Fig. 5a). The coefficients of the intercepts were higher at Qd-rare sites than at Qd-abundant sites (P = 0.042; Table 1), indicating more stellate hairs at Qd-rare sites (Fig. 5a). A model with site-specific intercepts and site-specific genetic effects was also selected for this trait (Table 1).
Regressions of six morphological traits (a stellate hair density, b shoot diameter, c tooth apex angle, d lateral vein interval, e leaf mass per area, and f number of buds at bud-scale scars) against ancestry proportions of Quercus dentata (Qd) cluster and proportions of flushing buds in the lower part of shoots. Regression coefficients are estimated from selected models with the lowest AIC values. Solid and dotted lines on Qd ancestry proportions indicate regressions when lower flushing-bud proportions are low (0.1) and high (0.9), respectively. Solid and dotted lines on lower flushing-bud proportions indicate regressions when Qd ancestry proportions are low (0.1) and high (0.9), respectively. Lines are shown in ranges of Qd ancestry proportions and lower flushing-bud proportions observed at individual sites. Red and blue lines indicate regressions at Qd-abundant and Qd-rare sites, respectively
Second, a model with site-specific intercepts, a genetic effect, and an environmental effect was selected for the shoot diameter and the tooth apex angle (Table 1). These traits were determined both genetically and environmentally. The positive coefficients of these effects indicated that the trait values increased as both Qd ancestry and coastal stress increased (Table 1). A model with site-specific intercepts, a positive genetic effect, and site-specific environmental effects was also selected for the lateral vein interval (Table 1; Fig. 5d).
Third, a model with site-specific intercepts, a positive genetic effect, a positive environmental effect, and an interaction between these effects was selected for the shoot diameter, the tooth apex angle, and the number of buds at bud-scale scars (Table 1). The coefficients of the intercepts for the number of buds at bud-scale scars were higher at Qd-rare sites than at Qd-abundant sites (P = 0.028; Table 1; Fig. 5f). A model with site-specific intercepts, a positive genetic effect, site-specific environmental effects, and an interaction between these effects was also selected for the leaf mass per area (Table 1; Fig. 5e). The negative coefficients of the interactions (Table 1) indicated that the genetic and environmental effects were antagonistic, i.e., the effect of Qd ancestry was smaller when coastal stress was higher, and vice versa (Fig. 5b, c, e, f).
The linear mixed models were also applied to the traits of partial samples with low (< 0.5) Qd ancestry in both coastal and inland habitats at Qd-rare sites. As with the selected models for all samples, models with positive genetic effects were selected for all six traits (Table S2). Environmental effects were positive in selected models for the shoot diameter, the tooth apex angle, the lateral vein interval, and the number of buds at bud-scale scars (Table S2). The coefficients of the interactions between genetic and environmental effects were negative in the four traits and the leaf mass per area (Table S2).
Discussion
Genetic background
We hypothesized that the genetic background of the coastal ecotype of Qc in northern Hokkaido is (1) the same as that of inland Qc, (2) an admixture of Qd, and (3) an admixture of taxa other than Qd. The results support the second hypothesis and reject the first one because the Qd ancestry proportion was higher in coastal Qc than in inland Qc. Although the admixture of taxa other than Qd, such as Qm and Qs, was found in coastal Qc, their ancestry proportions did not differ between coastal and inland Qc. Therefore, the results do not support the third hypothesis. Some botanists have regarded the coastal ecotype of Qc as a putative hybrid between Qc and Qd (Ohba 2006). The results generally agree with this opinion and additionally suggest that most trees of this ecotype are not first generation (F1) hybrids because of their low (< 0.5) Qd ancestry proportions. A previous study using nuclear microsatellites indicated a genetic similarity between inland and coastal Qc (Ohsawa et al. 2011). Our study using more taxa and more loci than Ohsawa et al. (2011) successfully detected a difference in their genetic backgrounds.
Other botanists have posited that a part of coastal Qc belongs to Qm on the Eurasian continent (Miyabe and Kudo 1925) because of the geographic proximity of northern Hokkaido to continental Asia through Sakhalin and the formation of land bridges among them during the glacial periods. Although Qm and coastal Qc have similar traits, such as rounded serrations, some of their traits are distinctive. For example, stellate hairs on the lower leaf surface are present in coastal Qc but absent in Qm (Aizawa et al. 2018). In Japan, there is a taxon with characters similar to those of Qm, Q. mongolicoides (H. Ohba) Hiroki, in central Honshu (Hiroki 2017). In this taxon, an admixture of Qm was suggested by nuclear microsatellites (Aizawa et al. 2018). Thus, some taxa of Qc in Japan are likely to have various genetic backgrounds. We found a genetic cline with a declining Qm ancestry from the Eurasian continent to Hokkaido Island through Sakhalin Island, which is consistent with a previous study (Aizawa et al. 2018). Thus, historical migration among these regions may have resulted in this genetic cline. However, this cline is irrelevant to the genetic background of the coastal ecotype of Qc in northern Hokkaido.
The admixture of Qd in coastal Qc can result from both gene flow between these species and incomplete lineage sorting of polymorphisms in their ancestral species (Lexer et al. 2006; Muir and Schlötterer 2005). The former, involving hybridization and backcross between species after their divergence, is thought to be common in oaks (Lexer et al. 2006). Introgression has been detected in various oak taxa (Eaton et al. 2015; Ortego et al. 2018), even between deeply divergent lineages (McVay et al. 2017). Gene flow between Qc and Qd seems relatively frequent near the northern limit of the distributional range of Qd because the flowering synchrony between them increases at higher latitudes in Hokkaido (Shimizu et al. 1992, 1994, 1995). In spite of these facts, we cannot determine the contribution of gene flow and incomplete lineage sorting to the admixture.
The nuclear microsatellites examined in this study seem reliable because they properly revealed the characteristics of four investigated taxa. In Hokkaido, the distributional ranges of Qd and Qs are more restricted than that of Qc (Kanazashi and Kanazashi 2009; Ubukata 2009; Yokoi 2009). Consistently, the genetic diversity of Qd and Qs was lower than that of Qc. Among the four taxa, the genetic differentiation between Qm and Qc is lowest, agreeing with their taxonomic treatments, in which these taxa belong to the same species (Ohashi 1988).
Genetic and environmental effects on traits
The admixture between Qc and Qd characterized the genetic variation along coastal–inland gradients in northern Hokkaido. F1 hybrids between Qc and Qd and their descendants are often found between the coastal and inland sides of oak forests (Matsumoto et al. 2009). Similarly, we observed a decline in Qd ancestry from the fore edge of coastal forests to inland forests at Qd-abundant sites. In the northernmost part of Hokkaido, Qd is rare (Hasegawa 1984), and the coastal ecotype of Qc is abundant (Shimizu et al. 1992, 1994, 1995; Shimizu 1997). Consistently, Qd ancestry was relatively low, even in the fore edge of coastal forests at Qd-rare sites.
Many traits differ between Qc and Qd (Ishida et al. 2003; Ito 2009). Some morphological traits of leaves and shoots differ among Qc, Qd, and their hybrids in common gardens (Ubukata et al. 1996), suggesting genetic effects on these traits. We consistently detected genetic effects on all six examined traits under the same coastal stress. The consistent genetic effects found in the partial samples with low (< 0.5) Qd ancestry indicates that the trait differences between coastal and inland Qc depend on their genetic backgrounds. Thus, the genetic background of the coastal ecotype of Qc, which is characterized by an admixture of Qd, results in Qd-like traits.
In addition to the genetic effects, we detected environmental effects on five of the six traits, which changed with coastal stress in trees with the same Qd ancestry. Because the correlation coefficient between Qd ancestry and coastal stress was relatively low, indicating weak collinearity between the genetic and environmental effects, these effects could be distinguished in our mixed linear models. Furthermore, those genetic and environmental effects were also found in the partial samples with low (< 0.5) Qd ancestry at Qd-rare sites. Thus, the traits of coastal and inland Qc depend on both their genetic backgrounds and environmental conditions. Because coastal stress caused by strong winds and salt spray in winter results in bud mortality in the upper parts of shoots (Asai et al. 1986; Shinmura et al. 1990), the lower flushing-bud proportion can be a surrogate measure for environmental conditions along a coastal–inland gradient. These conditions may acclimate the shape (tooth apex angle and lateral vein interval) and thickness (leaf mass per area) of leaves, shoot diameter, and bud production in the lowest parts of shoots (number of buds at bud-scale scars) to coastal stress. Similar phenotypic plasticity in leaf morphology has been found in Mediterranean oaks with different water availabilities (Ramírez-Valiente et al. 2010).
Negative interactions between the genetic and environmental effects were detected in several traits. This result suggests different reaction norms to coastal stress among inland Qc, coastal Qc, and Qd and lower phenotypic plasticity in Qd and coastal Qc than in inland Qc. Thus, Qd and coastal Qc trees tend to exhibit stress-tolerant traits irrespective of environmental conditions. As compared to Qc, Qd tends to be more stress tolerant, shade intolerant, and slow growing and to survive in more stressful habitats (Kanazashi and Kanazashi 2009; Ubukata 2009). These features of Qd correspond to the stress-tolerant strategy in stably unproductive habitats (Grime and Mackey 2002), resulting in reduced phenotypic plasticity (Valladares et al. 2007). Although reaction norms should be estimated from clones exposed to different environments (Callaway et al. 2003), we estimated trait variation from trees with the same Qd ancestry obtained from nuclear microsatellites. Thus, the trait variation regarded as phenotypic plasticity may include genetic variation at loci other than nuclear microsatellites.
Geographic variation in trait differences between Qc and Qd has been observed among several locations in Hokkaido (Ito 2009). Leaf hairiness and bud production in the lowest parts of shoots were higher at Qd-rare sites than at Qd-abundant sites in trees with the same Qd ancestry under the same coastal stress. This result indicates more Qd-like traits of the coastal ecotype of Qc without genetic or environmental effects. There are three explanations for this phenomenon. First, the specific environments other than coastal stress in the northernmost part of Hokkaido may induce the Qd-like traits in coastal Qc. Second, the introgression of trait-associated alleles from Qd to coastal Qc may lead to the Qd-like traits. The backcrossing of hybrids between Qc and Qd to Qc results in the introgression of Qd alleles to Qc. Because Qd alleles at loci associated with tolerance to coastal stress are likely selected in coastal habitats, coastal Qc seems to have such alleles more frequently at the trait-associated loci than at selectively neutral loci in the genetic background. This phenomenon, an environment-dependent introgression of adaptive alleles (adaptive introgression) (Goulet et al. 2017), has been suggested in oaks along a dry–wet gradient (Khodwekar and Gailing 2017). Third, the novel mutations at the trait-associated loci may result in the Qd-like traits of coastal Qc.
Conclusions
Oaks along coastal–inland gradients in northern Hokkaido are suitable for studying adaptation to coastal environments because of their clear habitat heterogeneity and substantial selection from coastal stress. To explore how the unique traits of the coastal ecotype of Qc arise, we revealed the genetic background of this ecotype and examined whether it affected these traits. The results suggest that not only coastal stress but also the genetic background admixed with Qd is responsible for the traits. Furthermore, additional factors, such as the adaptive introgression of trait-associated alleles from Qd, may affect the traits of the coastal ecotype of Qc.
References
Aizawa M, Maekawa K, Mochizuki H et al (2018) Unveiling the origin of Quercus serrata subsp. mongolicoides found in Honshu, Japan, by using genetic and morphological analyses. Plant Species Biol 33:174–190. https://doi.org/10.1111/1442-1984.12207
Asai T, Shinmura Y, Usui G (1986) Mortality factors of the winter buds of Quercus dentata and Quercus mongolica var. grosseserrata in natural coastal forests of northern Hokkaido. J Jpn For Soc 68:368–374
Callaway RM, Pennings SC, Richards CL (2003) Phenotypic plasticity and interactions among plants. Ecology 84:1115–1128. https://doi.org/10.1890/0012-9658(2003)084[1115:ppaiap]2.0.co;2
Ciccarelli D (2015) Mediterranean coastal dune vegetation: Are disturbance and stress the key selective forces that drive the psammophilous succession? Estuar Coast Shelf Sci 165:247–253. https://doi.org/10.1016/j.ecss.2015.05.023
De Boeck P, Bakker M, Zwitser R et al (2011) The estimation of item response models with the lmer function from the lme4 package in R. J Stat Softw 39:1–28. https://doi.org/10.18637/jss.v039.i12
Doing H (1985) Coastal fore-dune zonation and succession in various parts of the world. Vegetatio 61:65–75. https://doi.org/10.1007/BF00039811
Eaton DAR, Hipp AL, González-Rodríguez A, Cavender-Bares J (2015) Historical introgression among the American live oaks and the comparative nature of tests for introgression. Evolution 69:2587–2601. https://doi.org/10.1111/evo.12758
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: Linked loci and correlated allele frequencies. Genetics 164:1567–1587
Goulet BE, Roda F, Hopkins R (2017) Hybridization in plants: Old ideas, new techniques. Plant Physiol. https://doi.org/10.1104/pp.16.01340
Grime JP, Mackey JML (2002) The role of plasticity in resource capture by plants. Evol Ecol 16:299–307. https://doi.org/10.1023/A:1019640813676
Hasegawa S (1984) Basic studies on the conservation of the natural coastal forests in Hokkaido: The structure and regeneration of Quercus dentata Thunb. forest in Ishikari. Res Bull Hokkaido Univ For 41:313–422
Hiroki S (2017) A new scientific name proposed for a unique deciduous Quercus in Chubu and Kanto districts, Japan. J Phytogeogr Taxon 64:73–76
Ishida TA, Hattori K, Sato H, Kimura MT (2003) Differentiation and hybridization between Quercus crispula and Q. dentata (Fagaceae): Insights from morphological traits, amplified fragment length polymorphism markers, and leafminer composition. Am J Bot 90:769–776. https://doi.org/10.3732/ajb.90.5.769
Ito M (2009) Variation in leaf morphology of Quercus crispula and Quercus dentata assemblages among contact zones: a method for detection of probable hybridization. J For Res 14:240–244. https://doi.org/10.1007/s10310-009-0121-0
Kadomatsu M (1997) Differences in phenology of Quercus collected from northeast China, eastern Hokkaido and western Honshu. Res Bull Hokkaido Univ For 54:188–201
Kanazashi A, Kanazashi T (2009) Quercus mongolica Fischer ex Ledeb. var. crispula (Blume) Ohashi. In: Suzuki W, Ohsumi K, Yamanaka N (eds) Silvics of Japan, vol 1. Japan Forestry Investigation Committee, Tokyo, pp 635–667
Kanno M, Yokoyama J, Suyama Y et al (2004) Geographical distribution of two haplotypes of chloroplast DNA in four oak species (Quercus) in Japan. J Plant Res 117:311–317. https://doi.org/10.1007/s10265-004-0160-8
Khodwekar S, Gailing O (2017) Evidence for environment-dependent introgression of adaptive genes between two red oak species with different drought adaptations. Am J Bot 104:1088–1098. https://doi.org/10.3732/ajb.1700060
Kitamura K, Namikawa K, Kawahara T et al (2017) Genetic structure of remnant Quercus serrata populations at the northernmost limit of their distribution in Japan. Acta Phytotaxon Geobot 68:1–15
Lexer C, Kremer A, Petit RJ (2006) Shared alleles in sympatric oaks: Recurrent gene flow is a more parsimonious explanation than ancestral polymorphism. Mol Ecol 15:2007–2012. https://doi.org/10.1111/j.1365-294X.2006.02896.x
Liu H-Z, Harada K (2014) Geographic distribution and origin of the chloroplast T/C-type in Quercus mongolica var. crispula in northeastern Japan. Plant Species Biol 29:207–211
Matsumoto A, Kawahara T, Kanazashi A et al (2009) Differentiation of three closely related Japanese oak species and detection of interspecific hybrids using AFLP markers. Botany 87:145–153. https://doi.org/10.1139/B08-121
McVay JD, Hipp AL, Manos PS (2017) A genetic legacy of introgression confounds phylogeny and biogeography in oaks. Proc R Soc B Biol Sci 284:20170300. https://doi.org/10.1098/rspb.2017.0300
Mishima K, Watanabe A, Isoda K et al (2006) Isolation and characterization of microsatellite loci from Quercus mongolica var. crispula. Mol Ecol Notes 6:695–697. https://doi.org/10.1111/j.1471-8286.2006.01313.x
Miyabe K, Kudo Y (1925) Icones of the essential forest trees of Hokkaido, vol II. XII). Hokkaido Government, Sapporo
Muir G, Schlötterer C (2005) Evidence for shared ancestral polymorphism rather than recurrent gene flow at microsatellite loci differentiating two hybridizing oaks (Quercus spp.). Mol Ecol 14:549–561. https://doi.org/10.1111/j.1365-294X.2004.02418.x
Nagasaka Y, Kikuchi K, Yamazaki Y (1991) Growth pattern of natural coastal forest in North-Hokkaido: The structure of Quercus dentata forest in Teshio. Trans Hokkaido Branch Jpn For Soc 39:153–155
Ohashi H (1988) The new name instead of Quercus mongolica Fisch. var. grosseserrata (Bl.) Rehd. Wilis. (Fagaceae). J Jpn Bot 63:13–14
Ohba H (2006) Fagaceae. In: Iwatsuki K, Boufford DE, Ohba H (eds) Flora of Japan. Volume IIa. Kodansha, Tokyo, pp 42–60
Ohsawa T, Tsuda Y, Saito Y, Ide Y (2011) The genetic structure of Quercus crispula in northeastern Japan as revealed by nuclear simple sequence repeat loci. J Plant Res 124:645–654. https://doi.org/10.1007/s10265-010-0402-x
Okaura T, Quang ND, Ubukata M, Harada K (2007) Phylogeographic structure and late Quaternary population history of the Japanese oak Quercus mongolica var. crispula and related species revealed by chloroplast DNA variation. Genes Genet Syst 82:465–477. https://doi.org/10.1266/ggs.82.465
Ortego J, Gugger PF, Sork VL (2018) Genomic data reveal cryptic lineage diversification and introgression in Californian golden cup oaks (section Protobalanus). New Phytol 218:804–818. https://doi.org/10.1111/nph.14951
Patterson N, Price AL, Reich D (2006) Population structure and eigenanalysis. PLoS Genet 2:2074–2093. https://doi.org/10.1371/journal.pgen.0020190
Petit RJ, Bodénès C, Ducousso A et al (2004) Hybridization as a mechanism of invasion in oaks. New Phytol 161:151–164. https://doi.org/10.1046/j.1469-8137.2003.00944.x
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ramírez-Valiente JA, Sánchez-Gómez D, Aranda I, Valladares F (2010) Phenotypic plasticity and local adaptation in leaf ecophysiological traits of 13 contrasting cork oak populations under different water availabilities. Tree Physiol 30:618–627. https://doi.org/10.1093/treephys/tpq013
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8:103–106. https://doi.org/10.1111/j.1471-8286.2007.01931.x
Shimizu H (1997) Effects of geomorphological evolution on distribution of Quercus around the Sarobetsu peatland. Trans Hokkaido Branch Jpn For Soc 45:41–44
Shimizu H, Nagasaka Y, Yamazaki Y (1992) Local variation of bud number on current shoot of Quercus in North-Hokkaido. Trans Hokkaido Branch Jpn For Soc 40:220–222
Shimizu H, Kikuchi K, Yamada K (1994) Relationship between number of axillary buds of bud-scales and salt-spray resistance Quercus mongolica var. grosseserrata. Trans Hokkaido Branch Jpn For Soc 42:171–173
Shimizu H, Kikuchi K, Yamada K (1995) Local variation of bud number on axillary buds of bud-scales of Quercus dentata in coastal forest along Japan sea of Hokkaido. Trans Hokkaido Branch Jpn For Soc 43:140–142
Shinmura Y, Asai T, Usui G (1990) The relationships between environment and budbursting patterns of Quercus dentata and Quercus mongolica var. grosseserrata in natural coastal forests of northern Hokkaido. J Jpn For Soc 72:478–487
Ubukata M (2009) Quercus dentata Thunb. In: Suzuki W, Ohsumi K, Yamanaka N (eds) Silvics of Japan, vol 1. Japan Forestry Investigation Committee, Tokyo, pp 195–213
Ubukata M, Kohno K, Iizuka K (1996) Morphological characteristics of Quercus crispula × dentata hybrids. Trans Hokkaido Branch Jpn For Soc 44:113–116
Ubukata M, Itahana N, Kohno K (1999) Cross-compatibility between Quercus mongolica var. grosseserrata and Quercus dentata and both the reproductive ability and flowering time of their interspecific hybrids. J Jpn For Soc 81:286–290
Ueno S, Tsumura Y (2008) Development of ten microsatellite markers for Quercus mongolica var. crispula by database mining. Conserv Genet 9:1083–1085. https://doi.org/10.1007/s10592-007-9462-4
Ueno S, Taguchi Y, Tsumura Y (2008) Microsatellite markers derived from Quercus mongolica var. crispula (Fagaceae) inner bark expressed sequence tags. Genes Genet Syst 83:179–187. https://doi.org/10.1266/ggs.83.179
Ueno S, Aoki K, Tsumura Y (2009a) Generation of expressed sequence tags and development of microsatellite markers for Castanopsis sieboldii var. sieboldii (Fagaceae). Ann For Sci 66:509
Ueno S, Taguchi Y, Tomaru N, Tsumura Y (2009b) Development of EST-SSR markers from an inner bark cDNA library of Fagus crenata (Fagaceae). Conserv Genet 10:1477–1485. https://doi.org/10.1007/s10592-008-9764-1
Usui G, Shimizu H, Shinmura Y, Asai T (1988) A transformation model of asymmetric canopies of coastal forests. J Jpn For Soc 70:309–314
Valladares F, Gialoni E, Gómez JM (2007) Ecological limits to plant phenotypic plasticity. New Phytol 176:749–763. https://doi.org/10.1111/j.1469-8137.2007.02275.x
Yokoi S (2009) Quercus serrata Thunb. In: Suzuki W, Ohsumi N, Yamanaka N (eds) Silvics of Japan, vol 1. Japan Forestry Investigation Committee, Tokyo, pp 287–341
Acknowledgements
We thank the towns of Obira and Omu, as well as the Hokkaido Regional Forest Offices and the Soya and Kitami District Forest Offices, for their permission to collect samples; Ko Harada and Keiko Kitamura for providing DNA samples; Akiko Takazawa for her assistance in the laboratory work; Hiroki Itoh for his advice regarding statistical analyses; and Oliver Gailing, Ichiro Tamaki, and Kentaro Uchiyama for their suggestions for improving the manuscript. This study was supported by JSPS KAKENHI Grant Number 17K07859 to TN and Research Grant Number 201610 of the Forestry and Forest Products Research Institute to AN.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nagamitsu, T., Shimizu, H., Aizawa, M. et al. An admixture of Quercus dentata in the coastal ecotype of Q. mongolica var. crispula in northern Hokkaido and genetic and environmental effects on their traits. J Plant Res 132, 211–222 (2019). https://doi.org/10.1007/s10265-018-01079-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-018-01079-2