Abstract
The phosphoenolpyruvate-dependent glucose phosphotransferase system (PTSGlc) is the major uptake system responsible for transporting glucose, and is involved in glucose translocation and phosphorylation in Corynebacterium glutamicum. For the longest time, the PTSGlc was considered as the only uptake system for glucose. However, some PTS-independent glucose uptake systems (non-PTSGlc) were discovered in recent years, such as the coupling system of inositol permeases and glucokinases (IPGS) and the coupling system of β-glucoside-PTS permease and glucokinases (GPGS). The products (e.g. lysine, phenylalanine and leucine) will be increased because of the increasing intracellular level of phosphoenolpyruvate (PEP), while some by-products (e.g. lactic acid, alanine and acetic acid) will be reduced when this system become the main uptake pathway for glucose. In this review, we survey the uptake systems for glucose in C. glutamicum and their composition. Furthermore, we summarize the latest research of the regulatory mechanisms among these glucose uptake systems. Detailed strategies to manipulate glucose uptake system are addressed based on this knowledge.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Corynebacterium glutamicum is a non-pathogenic Gram-positive bacterium, which is widely used as workhorse for producing various amino acids in industry (Wang et al. 2019). Since 1984, the research on genetics and metabolism of C. glutamicum has accelerated because of the successful application of tools for genetically modifying C. glutamicum and the DNA sequencing of entire genome of two model strains (i.e., C. glutamicum ATCC13032, Brevibacterium flavum ATCC14067) (Xu et al. 2020). The metabolic engineering strategy for C. glutamicum has been extended from core biosynthetic pathways to central metabolism route, cofactor regeneration systems, energy metabolism, global regulation and material transmembrane transports (Ikeda 2012; Xu et al. 2018). On the other hand, Michiko and Shiio (1987) firstly discovered a system for glucose uptake and phosphorylation in C. glutamicum, and it was named as phosphoenolpyruvate-dependent sugar phosphotransferase system (i.e., PTS). At present, the physiological significance and mechanism of glucose uptake systems have been a hot topic in breeding high-productive strain of target products.
Glucose has been widely used as raw materials in fermentation industry because of the low price and the ubiquitous effect. The uptake and phosphorylation of glucose in C. glutamicum is mainly through PTSGlc (Blombach and Seibold 2010; Xu et al. 2019). PTS is composed of a membrane-bound carbohydrate-specific EIIABC component (EII) and two cytoplasmic components [i.e., enzyme I (EI) and histidine protein (HPr)]. PTS is widely found in bacteria but not in archaea and fungi (Ikeda 2012), which can efficiently transport and phosphorylate a variety of sugars at the same time. For example, glucose is transported and phosphorylated by PTSGlc to generate glucose-6-phosphate (Xu et al. 2019). However, the assimilation of glucose by PTSGlc requires phosphoenolpyruvate (PEP) as phosphoryl group donor, resulting in a reduction in intracellular PEP level and in production of PEP-derived products, such as l-lysine, succinic acid, aromatic compounds (Becker et al. 2011; Xu et al. 2019). In addition, the high efficiency of glucose utilization by PTSGlc causes the unbalance between glycolytic pathway and tricarboxylic acid cycle, resulting in overflow metabolism and deficient cell growth (Becker et al. 2011; Lara et al. 2008; Xu et al. 2020). Therefore, researchers have been looking for PTS-independent glucose uptake systems (i.e., non-PTSGlc) to replace PTSGlc to increase glucose utilization and intracellular PEP levels.
It has long been thought that PTSGlc is the only way to transport and phosphorylate glucose, but in fact there are a series of non-PTSGlc in C. glutamicum. For example, Kumar et al. (2019) reported that glucose can be transported into cells by glucose permease. In addition, Ikeda et al. (2011) pointed out that HPr-deficient strain also uses glucose as the sole carbon source and resists to 2-deoxyglucose, indicating that there is non-PTSGlc in C. glutamicum. Further studies confirmed that the coupling system of inositol permeases and glucokinases (designed as IPGS) participates in glucose uptake and phosphorylation in C. glutamicum. Moreover, Ikeda et al. (2015) found that the recombinant strain with double inactivation of PTSGlc and IPGS also grows on a medium with glucose as the sole carbon source, indicating that there is a third glucose uptake system in C. glutamicum. It is now established that the third glucose uptake system is the coupling system of β-glucoside-PTS permease and glucokinases (designed as GPGS).
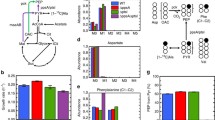
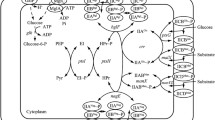
As mentioned above, there are three glucose uptake systems in C. glutamicum (Fig. 1). Therefore, many strategies can be used to improve the effective use of glucose. This review summarizes the current status and application prospects of glucose uptake systems in C. glutamicum. Moreover, regulatory mechanisms of different uptake systems were highlighted, which will provide theoretical bases for genetically modifying strains to improve glucose uptake rate. Finally, this review covers strategies for manipulating glucose uptake systems to construct high-productive strain of target products.
Three variant glucose uptake systems in Corynebacterium glutamicum. Black, yellow and green lines indicate three variant pathways for glucose transfer and transformation in C. glutamicum, i.e., PTSGlc, IPGS and GPGS. The green route (i.e., GPGS) is only found in C. glutamicum ATCC31833. PTSGlc is negatively affected by DeoR-type transcriptional regulators, e.g., SugR and FruR. IPGS is negatively affected by RpiR-Like transcriptional regulators, e.g., IolR. GPGS is negatively affected by glucose, but it is activated during introducing mutation G134T at the upstream of bglF, i.e., bglFup134. PYK Pyruvate kinase, PYC Pyruvate car boxylase, PPC Phosphoenolpyruvate carboxylase
The glucose uptake systems in C. glutamicum
The phosphoenolpyruvate-dependent glucose phosphotransferase system (i.e., PTSGlc)
Glucose is an important carbon source for C. glutamicum, which is mainly transported and phosphorylated by PTSGlc. PTSGlc is a glucose-specific uptake system consisting of two cytoplasmic protein components (i.e., EI and HPr; encoded by ptsI and ptsH, respectively) and one membrane-bound glucose-specifc EIIBCA component (i.e., EIIGlc; encoded by ptsG). EI is a highly conserved protein with a molecular weight of about 64 kDa, and it contains two functional domains, i.e., phosphopyruvate kinase and PEP synthetase (Kotrba et al. 2001b). The conserved sequence of EI contains a histidine active site to ensure that PEP or ATP can be used as a phosphate donor for phosphorylation (Kotrba et al. 2001b). HPr has a molecular weight of about 9 kDa and contains a single domain. The histidine located at position 15 of HPr is the highly conserved site, in which HPr will be phosphorylated (Ikeda 2012). Unlike EI and HPr, EIIBCA components (i.e., EIIs) can specifically recognize one or more specific carbohydrates (Aboulwafa and Saier 2002). In general, EIIs are composed of two hydrophilic cytoplasmic domains (i.e., IIA and IIB) and a transmembrane domain (i.e., IIC)(Moon et al. 2005). IIA, containing 100 to 160 amino acid residues, adopts phosphate groups and then phosphorylates the active site to form IIA-P. IIA-P will promote the phosphorylation of IIB at active site to form IIB-P. IIC, consisting of 6 or 8 transmembrane helixes and containing about 350 amino acid residues, phosphorylates carbohydrates and then transports them into cytoplasm (Moon et al. 2007). However, the phosphorylation and transport of carbohydrates by IIC are regulated by IIB-P, resulting in rapid dissociation of phosphorylated carbohydrates from EIIs (Lee et al. 1994).
The tandem phosphorylation reaction by PTS is completed by five steps (Fig. 2). These five steps are catalysed by EI, HPr and EIIs, and reversibly transfer phosphate groups (Kotrba et al. 2001a). Firstly, EI adopts the high-energy phosphate groups from PEP to auto-phosphorylate to form EI-P. Then, the phosphate group from EI-P is transported to HPr and subsequently participates in the reactions catalyzed by EIIs (i.e., IIA, IIB, IIC). In these catalyzed reactions, IIA is phosphorylated by HPr, and then to phosphorylates IIB, thus opening the permease channel of IIC. Finally, the phosphate groups from PEP are transported to carbohydrates (Moon et al. 2007).
PTS-independent glucose uptake systems (i.e., non-PTSGlc)
The coupling system of inositol permeases and glucokinases (i.e., IPGS)
In 2011, Ikeda et al. (2011) found that HPr-deficient C. glutamicum strain SPH2 also uses glucose as the sole carbon source and resists to 2-deoxyglucose indicating that this recombinant strain does not rely on the PTSGlc system to assimilate glucose. Further studies confirmed that the coupling system of inositol permeases and glucokinases (designed as IPGS, similarly hereinafter) participates in glucose uptake and phosphorylation, which belongs to PTS-independent glucose uptake systems (i.e., non-PTSGlc). IPGS consists of myo-inositol permeases and glucokinases. In IPGS, carbohydrates are firstly transported into intracellular by myo-inositol permeases, and then phosphorylated by glucokinases (Lindner et al. 2013). IolT1 and IolT2 are two of myo-inositol permeases, which are responsible for transporting myo-inositol in C. glutamicum (Krings et al. 2006). Unlike Bacillus subtilis (Morinaga et al. 2010), IolT1 and IolT2 in C. glutamicum have sequence identity of up to 55%, and have similar gene expression levels and kinetic characteristics (Krings et al. 2006). Glk and PpgK are two of glucokinases in C. glutamicum. Glk and PpgK have sequence identity of 28% and have typical ATP binding sites (Lindner et al. 2010). However, IPGS will be activated only at the unavailability of PTSGlc or at the excess glucose.
The coupling system of β-glucoside-PTS permease and glucokinases (i.e., GPGS)
The coupling system of β-glucoside-PTS permease and glucokinases (designed as GPGS, similarly hereinafter) was confirmed in 2015. Ikeda et al. (2015) found that the recombinant strain with double inactivation of PTSGlc and IPGS also grow on a medium with glucose as the sole carbon source, indicating that GPGS is the third glucose uptake system in C. glutamicum except PTSGlc and IPGS. GPGS consists of β-glucoside-PTS permease (i.e., EIIBgl) and glucokinases. EIIBgl, encoded by gene bglF, has been identified as bglF-specified EII component of β-glucoside-PTS (Kotrba et al. 2003). EIIBgl has a molecular weight of about 65 kDa and contains three domains, i.e., IIA, IIB, and IIC (Chen and Amster-Choder 1998). In GPGS, carbohydrate is firstly transported into intracellular by EIIBgl, and then phosphorylated by glucokinases. The glucokinases in GPGS are the same as in IPGS (Ikeda et al. 2015). Since GPGS was only found in mutant strain derived from C. glutamicum ATCC 31833, its application remains to be explored.
Regulatory mechanisms of genes in glucose uptake systems
The regulatory mechanisms in PTSGlc
Regulations of the DeoR-type transcriptional regulators
When C. glutamicum was cultured in medium with a mixed carbon source, the consumption rate of glucose was decreased (Wendisch et al. 2000). This is because SugR, an acetic acid-mediated deoxyribonucleotide repressor (DeoR)-type transcriptional regulator, inhibits gene expression in the PTSGlc (Gaigalat et al. 2007). SugR is encoded by the gene sugR, which inhibits the expression of many genes in C. glutamicum, such as 6-phosphate fructokinase, fructose-1, 6-bisphosphate aldolase, enolase, pyruvate kinases and l-lactate dehydrogenase (Engels et al. 2008). In the PTS, SugR binds to a specific region in the promoter of the key genes ptsH and ptsI, thereby suppressing the expression of these genes (Tanaka et al. 2008a). The specific region is a GTTGCACA sequence or a TG(T)2-5G sequence (Tanaka et al. 2008b). In addition, a 21 bp fragment of the ptsI–fruR promoter region is also binded by SugR (Gaigalat et al. 2007).
FruR is another DeoR-type transcription regulator to regulate the PTS, and its coding gene fruR is located at the upstream of the ptsF gene (Liu et al. 2016). Its expression level is increased with the increase of intracellular PTS sugar concentration, thus inhibiting the expression of the genes ptsI and ptsH (Tanaka et al. 2008b). In E. coli, FruR also plays an important role in regulating a large number of key genes in carbon metabolism, such as pckA (encoding PEP carboxykinase), edd (encoding 6-phosphogluconate dehydratase) (Liu et al. 2017). However, no reports indicated that FruR in C. glutamicum inhibits the expression of genes except the PTS-related genes. These results indicated that FruR in C. glutamicum seems to play a role in regulating the expression of genes involved in the PTS, thereby preventing the excessive intake of sugar by PTS system.
Regulations of the GntR-type transcriptional regulators
GntR-type transcriptional regulators are a family of important transcriptional regulators, which are widespread in bacteria and actinomycetes (Lambrecht et al. 2018). The structure of GntRs generally consists of a conserved N-terminal helix-turn-helix motif and a variable C-terminal effector binding/oligomerization domain (Taw et al. 2015). There are two GntR-type transcription regulators in C. glutamicum, called GntR1 and GntR2, respectively (Tanaka et al. 2012). Similar in E. coli (Porco et al. 1997) and B. subtilis (Reizer et al. 1991), GntR-type transcriptional regulators in C. glutamicum strongly inhibit the expression of genes involved in the regulation of gluconic acid catabolism, i.e., gntP (encoding glycosyl peroxidase), gntK (encoding gluconate kinase) and gnd (encoding gluconate dehydrogenase) (Tanaka et al. 2012). However, the difference is that these transcriptional regulators in C. glutamicum activate the expression of the key gene ptsG in the PTSGlc, thus increasing the glucose uptake rate (Frunzke et al. 2008; Teramoto et al. 2011). One possible reason is that GntR1/2 binding site of the genes gntP and gntK is similar to the starting position of the ptsG gene (Teramoto et al. 2011). This may be why C. glutamicum can co-utilize glucose and other carbon sources but E. coli or B. Subtilis can not do it (Stansen et al. 2005).
The regulatory mechanisms in non-PTSGlc
Regulations of the RpiR-like transcriptional regulators
As mentioned above, the IPGS is consisting of myo-inositol permeases (i.e., IolT1 and IolT2) and endogenous glucokinase (i.e., GlK, PpgK). Klaffl et al. (2013) found that a RpiR-Like transcriptional regulatory factor IolR locates at the upstream of the coding genes of IolT1 and IolT2 (i.e., iolT1 and iolT2, respectively) in the opposite direction. In addition, the binding site of the IolR was found in the promoter region of iolT1 and iolT2 (Klaffl et al. 2013). Previous researches indicated that that the mRNA level of the iol gene in the IolR-mutated strain was significantly increased (> 100-fold). These results indicated that IolR suppresses the expression of iol gene and presumably regulated by negative autoregulation. Moreover, Zhou et al. (2015) found that IolR may inhibits the expression of two glucokinase genes (i.e., glk and ppgk). However, the inhibition mechanism of IolR on glucokinase needs to be further explored.
Regulations of the EII component of the β-glucoside-PTS
As the key component of the β-glucoside-PTS, EIIbgl was encoded by gene bglF, which located in the gene cluster bglF-bglA-bglG (Tanaka et al. 2009). This gene cluster is responsible for encoding genes related to β-glucoside transport. Similar in Streptococcus mutans, β-glucoside-PTS-related genes in C. glutamicum are sensitive to the presence of glucose (Cote et al. 2000). However, the transcription level of bglA has no significant effect on bglF expression (Tanaka et al. 2011). Experimental results showed that a ribonucleic antiterminator (RAT) sequence exists in the upstream of bglF (Tanaka et al. 2009). The role of the RAT sequence was activated during the presence of glucose, and then to disable the anti-termination mechanism of the gene cluster bglF-bglA-bglG, thus inhibiting bglF expression (Kotrba et al. 2003). It should be noted that the regulatory effect of the RAT sequence always exists, though C. glutamicum depends on PTSGlc to transport glucose. Interestingly, glucose also has a weak inhibitory effect on bglF expression in the strain with a deletion of the RAT sequence. It may be that glucose is able to weakly block the initiation of bglF gene transcription. In contrast, an antiterminator protein BglG, encoded by gene bglG, can relieve the inhibition of glucose on bglF gene expression (Tanaka et al. 2011).
Manipulation strategy for improving the glucose conversion ratio in glucose uptake systems
The strategies based on manipulated PTSGlc
Introducing the exogenous substances to increase the glucose uptake rate
Maltose is an incomplete hydrolysis product of starch liquefaction and saccharification. Previous researches indicated that C. glutamicum can use maltose as the sole carbon source to produce amino acids (Xu et al. 2016). Maltose was generally catalyzed by 4-α-glucan transferase to produce maltodextrin and glucose (Kalebina et al. 2008). Then, the glucose enters in the glycolytic pathway catalyzed by glucokinases, whereas the maltodextrin is degraded to produce glucose-1-phosphate catalyzed by maltodextrin phosphorylase (MalP) (Seibold et al. 2009). Unlike E. coli, C. glutamicum doses not exhibit a diauxic growth during use of maltose and glucose as a mixed carbon source (Xu et al. 2016). Interestingly, the growth rate of bacteria was significantly accelerated during growth on mixture of maltose and glucose as compared with on glucose as a single carbon source, though the content of total carbon source in culture medium is unchanged (Henrich et al. 2013). In addition, Krause et al. (2010) pointed out that the transcription level of ptsG was increased during addition of maltose, indicating that maltose promotes glucose metabolism in C. glutamicum. Further studies conclusively showed that maltose was transported into cells by the MusEFGK2I ABC transport system, resulting in increasing the transcription level of ptsG (Henrich et al. 2013). Therefore, addition of maltose to medium is an effective method to increase the glucose uptake rate in bacteria. However, maltose is expensive, thus limiting its application in industry. To solve this problem, addition of starch may be a good choice during use of a recombinant strain with heterologous expression of α-amylase. Previous research indicated that a recombinant C. glutamicum strain with overexpression of AmyA from Streptococcus bovis 148 on its cell surface can hydrolyze starch to maltose, thus promoting the expression of ptsG (Tateno et al. 2007). This may provide an interesting idea for improving the glucose uptake rate of strain. However, this strategy was merely a technological speculation, it takes a long time to realize industrial production manner. Generally speaking, the strategy based on addition of exogenous substance to increase the glucose uptake rate may not be a optimum approach in industry because of the high-cost production.
Increasing the activity of enzymes/proteins in PTS
PTSGlc consists of two cytoplasmic protein components (i.e., EI and HPr) and one membrane-bound glucose-specific EIIBCA component (i.e., EIIGlc) (Xu et al. 2019). In theory, the glucose uptake rate of strain can be increased by overexpression of key genes in PTSGlc. For example, Krause et al. (2010) indicated that the glucose consumption rate of recombinant strain C. glutamicum (pBB1-ptsG) with overexpression of ptsG gene was significantly increased as compared with the origin strain. The similar results were also found in other research reported by Lindner et al. (2013), in which overexpression of ptsG gene in lysine-producing strain C. glutamicum DM1729 Δpgi increased glucose consumption rate, cell growth rate and l-lysine production on glucose medium. The above-mentioned researches show that improvement of the enzymes/proteins in PTSGlc is beneficial to increase the glucose uptake rate of strain. However, the positive effects are very limited because the genes in the PTS pathway are regulated by many factors, such as the transcriptional regulator SugR (Engels et al. 2008; Lindner et al. 2013). Therefore, it is difficult to significantly improve the glucose uptake rate by simple overexpression of genes in PTSGlc.
Eliminating the negative transcriptional regulators in PTS
SugR is mediated by acetic acid and inhibits the expression of genes in PTS, especially the key genes ptsG, ptsI and ptsH in PTSGlc (Gaigalat et al. 2007). Therefore, eliminating or weakening the effect of SugR is an effective strategy to increase the expression of genes in PTS, thereby increasing the glucose uptake rate in bacteria. And researchers have made preliminary attempts in this regard. For example, Engels and Wendisch (2007) deleted SugR-coding gene sugR in C. glutamicum and found that the growth rate of this recombinant strain was increased during growth on PTS sugar (e.g., glucose, fructose and sucrose), because the expression level of genes in PTS was increased. In addition, Gaigalat et al. (2007) found that the negative effect of SugR on ptsG gene was weakened when fructose-1-phosphate (F-1-P), 1,6-fructose diphosphate (F-1,6-P) or glucose 6-phosphate (G-6-P) are present, because F-1-P, F-1,6-P and G-6-P keep SugR from binding with the promoter of ptsG. Interestingly, the transcription level of genes in PTS were increased in SugR-deficient C. glutamicum strain, both on PTS sugar and non-PTS sugar medium (Tanaka et al. 2008a). There results indicated that the regulatory relationships between SugR and PTS are very complex. Maybe the third part factors intervene in the process of regulation. However, we don’t know what the total number of factors and what they would be.
It should be noted that improvement of PTSGlc helps to increase the glucose consumption rate, but it also causes the unbalance between glycolytic pathway and tricarboxylic acid cycle, resulting in overflow metabolism and deficient cell growth (Becker et al. 2011; Lara et al. 2008; Xu et al. 2020). The strain with high efficient of PTSGlc not only improve the yield of the target product, but also increase the content of by-products. These results implied that increase of the glucose conversion ratio is not satisfied with simply improving the PTSGlc.
The strategies based on manipulated non-PTSGlc
Increasing the expression level of the enzymes/proteins in non-PTSGlc
Aside from engineering PTS to increase the glucose uptake rate, many studies pointed out the important role of non-PTSGlc for cell growth and yield of target products. Up to now, there are two types of non-PTSGlc, i.e., IPGS and GPGS (Xu et al. 2019). Glucose assimilation through non-PTSGlc is regulated by permeases (i.e., IolT1, IolT2 and EIIBgl) and glucokinase (i.e., GlK, PpgK and Cgl2647) (Fig. 1). Therefore, changing the expression level of these protein-coding genes in non-PTSGlc can regulate the glucose uptake rate in this system. Zhou et al. (2015) pointed out that co-overexpression of ppgk and iolT1 and knock-out of iolR in a PTS-deficient C. glutamicum strain completely restore the glucose uptake rate and increase the yield of succinic acid. Similar results was also found in another study reported by Zhang et al. (2015), in which the strain with insertion of the iolT2-ppgk cassette at the ptsI locus showed the similar glucose uptake rate as original strain, but showed a high l-phenylalanine production and a low by-products. In addition, Xu et al. (2019) have pointed out that the recombinant strain C. glutamicum ZL-92 with introducing two point mutations in the promoter of iolT1 and replacing the nature promoter of iolT2 and ppgK by the tuf promoter showed a quick growth and a high glucose consumption rate and l-lysine production. As compared with overexpression of genes in PTSGlc, increasing the expression level of genes in non-PTSGlc exhibited the best glucose uptake system for producing l-lysine. These studied have a common point that the recombinant strain showed the increase of the intracellular PEP concentration because of inactivation of PTSGlc.
Optimizing the configuration of non-PTSGlc
Although non-PTSGlc participates the glucose assimilation, PTSGlc is the main glucose-specific uptake system in C. glutamicum (Xu et al. 2020). This is because permease, a key compenent in non-PTSGlc, is not the ubiquitous protein in C. glutamicum (Ikeda 2012). In addition, these permeases normally show a low affinity to glucose (Lindner et al. 2011) and a low expression level (Ikeda 2012). In addition, different types of glucokinase use different substrates (i.e., ATP, ADP and inorganic polyphosphates) as phosphoryl donor (Lindner et al. 2011). Therefore, optimizing the configuration of non-PTSGlc is also a good strategy besides overexpression of permease-coding genes. The expression of IolT1 is repressed by IolR (Klaffl et al. 2013), but knock-out of IolR causes some negative effects on strain (Brusseler et al. 2018). Fortunately, introduction of two point mutations at position − 113 (A→G) and − 112 (C→G) in the nature promoter of iolT1 reliefs the repression of IolR and avoids the negative effects on strain (Brusseler et al. 2018). In addition, Ikeda et al. (2015) indicated that introduction of a point mutation at position 134 (G→T) in the upstream of bglF gene significantly activates the expression of EIIBgl. Moreover, co-overexpression of ADP-GlK/PFK from Methanococcus maripaludis and ADP-GlK from Bacillus subtilis in l-lysine producer effectively improves glucose consumption rate, cell growth, l-lysine production and by-products accumulation either in shake-flasks or in fed-batch fermentation (Xu et al. 2020).
Knocking-out or site-directed mutating the negative transcriptional regulators in non-PTSGlc
As a DeoR transcriptional regulator, IolR inhibits the expression of glucokinases (i.e., GlK and PpgK) and IolT1 (Klaffl et al. 2013; Zhou et al. 2015). In order to increase participation of non-PTSGlc in glucose assimilation, many researches focus on genetically modifying the IolR. For example, Zhou et al. (2015) found that the transcription levels of key genes in non-PTSGlc were significantly increased during knock-out of iolR gene, resulting in completely restoring the glucose consumption rate and increasing succinic acid production. Similar results were also found in other previous researches (Michiko and Shiio 1987), in which the glucose uptake rate and the production of target products were increased during knock-out of iolR gene. However, knock-out of IolR causes some negative effects on strain (Brusseler et al. 2018). So could we relief the repression of IolR and avoid the negative effects on strain by introducing point mutations in iolR gene? Therefore, further research is necessary.
In addition, the expression of the gene bglF is affected by the RAT sequence and the anti-termination protein BglG. Tanaka et al. (2011) pointed out that the inhibition of bglF expression by glucose will be able to somewhat relieved by over-expressing bglG. However, the most important limiting factor of bglF expression level is the negative effect of glucose on RAT sequence function. Ikeda et al. (2015) found that a mutant strain of C. glutamicum could grow on a medium with glucose as the sole carbon source, even there were no PTS and IPGS. Further study found that the expression level of bglF was increased by about 11 times than that of wild type because it had a mutation (T to G) at 134 bp upstream of bglF (Ikeda et al. 2015). It shoud be noted that the mutation T134Up of bglFG was located in the RAT sequence, precisely so this mutation hinders the binding of the RAT sequence to glucose. This also provides ideas for the strengthening of the GPGS approach.
Conclusions and future prospects
Glucose is mainly transported by PTSGlc in C. glutamicum, but PTSGlc uses PEP as phosphoryl donor, which affects the production of the PEP-derived products and by-products accumulation (Lindner et al. 2013; Xu et al. 2019). Therefore, researchers have been searching for other glucose uptake systems that do not depend on PEP as phosphoryl donor. Up to now, two types of non-PTSGlc, i.e., IPGS and GPGS, were found in C. glutamicum. Many researches have proved that increasing the activity of enzymes/proteins in non-PTSGlc not only increase the intracellular PEP content to improve the production of target products, but also reduce the accumulation of by-products (e.g., lactic acid, alanine, acetic acid) (Lara et al. 2008; Xu et al. 2019). Therefore, enhancing the participate of non-PTSGlc in glucose assimilation is one of the best strategy to develop a high target product producing strain with high productivity and glucose conversion rate. However, it should be noted that the glucose transporting efficiency by non-PTSGlc is not significantly increased and even slightly decreased as compared with PTSGlc at present. Therefore, how to improve the glucose transporting efficiency by non-PTSGlc is the important aspect to invest in the future.
References
Aboulwafa M, Saier MH Jr (2002) Dependency of sugar transport and phosphorylation by the phosphoenolpyruvate-dependent phosphotransferase system on membranous phosphatidyl glycerol in Escherichia coli: studies with a pgsA mutant lacking phosphatidyl glycerophosphate synthase. Res Microbiol 153:667–677. https://doi.org/10.1016/s0923-2508(02)01376-1
Becker J, Zelder O, Hafner S, Schroder H, Wittmann C (2011) From zero to hero–design-based systems metabolic engineering of Corynebacterium glutamicum for l-lysine production. Metab Eng 13:159–168. https://doi.org/10.1016/j.ymben.2011.01.003
Blombach B, Seibold GM (2010) Carbohydrate metabolism in Corynebacterium glutamicum and applications for the metabolic engineering of l-lysine production strains. Appl Microbiol Biotechnol 86:1313–1322. https://doi.org/10.1007/s00253-010-2537-z
Brusseler C, Radek A, Tenhaef N, Krumbach K, Noack S, Marienhagen J (2018) The myo-inositol/proton symporter IolT1 contributes to d-xylose uptake in Corynebacterium glutamicum. Bioresour Technol 249:953–961. https://doi.org/10.1016/j.biortech.2017.10.098
Chen Q, Amster-Choder O (1998) The different functions of BglF, the E. coli beta-glucoside permease and sensor of the bgl system, have different structural requirements. Biochemistry US 37:17040–17047. https://doi.org/10.1021/bi980067n
Cote CK, Cvitkovitch D, Bleiweis AS, Honeyman AL (2000) A novel beta-glucoside-specific PTS locus from Streptococcus mutans that is not inhibited by glucose. Microbiology 146(Pt 7):1555–1563. https://doi.org/10.1099/00221287-146-7-1555
Engels V, Wendisch VF (2007) The DeoR-type regulator SugR represses expression of ptsG in Corynebacterium glutamicum. J Bacteriol 189:2955–2966. https://doi.org/10.1128/JB.01596-06
Engels V, Lindner SN, Wendisch VF (2008) The global repressor SugR controls expression of genes of glycolytic and of the L-lactate dehydrogenase LdhA in Corynebacterium glutamicum. J Bacteriol 190:8033–8044. https://doi.org/10.1128/JB.00705-08
Frunzke J, Engels V, Hasenbein S, Gatgens C, Bott M (2008) Co-ordinated regulation of gluconate catabolism and glucose uptake in Corynebacterium glutamicum by two functionally equivalent transcriptional regulators, GntR1 and GntR2. Mol Microbiol 67:305–322. https://doi.org/10.1111/j.1365-2958.2007.06020.x
Gaigalat L, Schluter JP, Hartmann M, Mormann S, Tauch A, Puhler A, Kalinowski J (2007) The DeoR-type transcriptional regulator SugR acts as a repressor for genes encoding the phosphoenolpyruvate:sugar phosphotransferase system (PTS) in Corynebacterium glutamicum. BMC Mol Biol 8:104. https://doi.org/10.1186/1471-2199-8-104
Henrich A, Kuhlmann N, Eck AW, Kramer R, Seibold GM (2013) Maltose uptake by the novel ABC transport system MusEFGK2I causes increased expression of ptsG in Corynebacterium glutamicum. J Bacteriol 195:2573–2584. https://doi.org/10.1128/jb.01629-12
Ikeda M (2012) Sugar transport systems in Corynebacterium glutamicum: features and applications to strain development. Appl Microbiol Biotechnol 96:1191–1200. https://doi.org/10.1007/s00253-012-4488-z
Ikeda M, Mizuno Y, Awane S, Hayashi M, Mitsuhashi S, Takeno S (2011) Identification and application of a different glucose uptake system that functions as an alternative to the phosphotransferase system in Corynebacterium glutamicum. Appl Microbiol Biotechnol 90:1443–1451. https://doi.org/10.1007/s00253-011-3210-x
Ikeda M, Noguchi N, Ohshita M, Senoo A, Mitsuhashi S, Takeno S (2015) A third glucose uptake bypass in Corynebacterium glutamicum ATCC 31833. Appl Microbiol Biotechnol 99:2741–2750. https://doi.org/10.1007/s00253-014-6323-1
Kalebina TS, Egorov SN, Arbatskii NP, Bezsonov EE, Gorkovskii AA, Kulaev IS (2008) The role of high-molecular-weight polyphosphates in activation of glucan transferase Bgl2p from Saccharomyces cerevisiae cell wall. Doklady Biochem Biophys 420:142–145. https://doi.org/10.1134/s1607672908030125
Klaffl S, Brocker M, Kalinowski J, Eikmanns BJ, Bott M (2013) Complex regulation of the phosphoenolpyruvate carboxykinase gene pck and characterization of its GntR-type regulator IolR as a repressor of myo-inositol utilization genes in Corynebacterium glutamicum. J Bacteriol 195:4283–4296. https://doi.org/10.1128/JB.00265-13
Kotrba P, Inui M, Yukawa H (2001a) Bacterial phosphotransferase system (PTS) in carbohydrate uptake and control of carbon metabolism. J Biosci Bioeng 92:502–517. https://doi.org/10.1263/jbb.92.502
Kotrba P, Inui M, Yukawa H (2001b) The ptsI gene encoding enzyme I of the phosphotransferase system of Corynebacterium glutamicum. Biochem Biophys Res Commun 289:1307–1313. https://doi.org/10.1006/bbrc.2001.6116
Kotrba P, Inui M, Yukawa H (2003) A single V317A or V317M substitution in Enzyme II of a newly identified beta-glucoside phosphotransferase and utilization system of Corynebacterium glutamicum R extends its specificity towards cellobiose. Microbiology 149:1569–1580. https://doi.org/10.1099/mic.0.26053-0
Krause FS, Henrich A, Blombach B, Kramer R, Eikmanns BJ, Seibold GM (2010) Increased glucose utilization in Corynebacterium glutamicum by use of maltose, and its application for the improvement of l-valine productivity. Appl Environ Microbiol 76:370–374. https://doi.org/10.1128/aem.01553-09
Krings E, Krumbach K, Bathe B, Kelle R, Wendisch VF, Sahm H, Eggeling L (2006) Characterization of myo-inositol utilization by Corynebacterium glutamicum: the stimulon, identification of transporters, and influence on l-lysine formation. J Bacteriol 188:8054–8061. https://doi.org/10.1128/jb.00935-06
Kumar S, Narayan KS, Shandilya S, Sood SK, Kapila S (2019) Role of non-PTS dependent glucose permease (GlcU) in maintaining the fitness cost during acquisition of nisin resistance by Enterococcus faecalis. Fems Microbiol Lett. https://doi.org/10.1093/femsle/fnz230
Lambrecht SJ, Wahlig JML, Steglich C (2018) The GntR family transcriptional regulator PMM1637 regulates the highly conserved cyanobacterial sRNA Yfr2 in marine picocyanobacteria. DNA Res 25:489–497. https://doi.org/10.1093/dnares/dsy019
Lara AR, Caspeta L, Gosset G, Bolivar F, Ramirez OT (2008) Utility of an Escherichia coli strain engineered in the substrate uptake system for improved culture performance at high glucose and cell concentrations: an alternative to fed-batch cultures. Biotechnol Bioeng 99:893–901. https://doi.org/10.1002/bit.21664
Lee JK, Sung MH, Yoon KH, Yu JH, Oh TK (1994) Nucleotide sequence of the gene encoding the Corynebacterium glutamicum mannose enzyme II and analyses of the deduced protein sequence. Fems Microbiol Lett 119:137–145. https://doi.org/10.1111/j.1574-6968.1994.tb06880.x
Lindner SN, Knebel S, Pallerla SR, Schoberth SM, Wendisch VF (2010) Cg2091 encodes a polyphosphate/ATP-dependent glucokinase of Corynebacterium glutamicum. Appl Microbiol Biotechnol 87:703–713. https://doi.org/10.1007/s00253-010-2568-5
Lindner SN, Seibold GM, Henrich A, Kramer R, Wendisch VF (2011) Phosphotransferase system-independent glucose utilization in Corynebacterium glutamicum by inositol permeases and glucokinases. Appl Environ Microbiol 77:3571–3581. https://doi.org/10.1128/AEM.02713-10
Lindner SN et al (2013) Phosphotransferase system-mediated glucose uptake is repressed in phosphoglucoisomerase-deficient Corynebacterium glutamicum strains. Appl Environ Microbiol 79:2588–2595. https://doi.org/10.1128/aem.03231-12
Liu L, Duan X, Wu J (2016) Modulating the direction of carbon flow in Escherichia coli to improve l-tryptophan production by inactivating the global regulator FruR. J Biotechnol 231:141–148. https://doi.org/10.1016/j.jbiotec.2016.06.008
Liu L, Chen S, Wu J (2017) Phosphoenolpyruvate:glucose phosphotransferase system modification increases the conversion rate during l-tryptophan production in Escherichia coli. J Ind Microbiol Biotechnol 44:1385–1395. https://doi.org/10.1007/s10295-017-1959-3
Michiko M, Shiio I (1987) Phosphoenolpyruvate: sugar phosphotransferase systems and sugar metabolism in Brevibacterium flavum. Agric Biol Chem 51:8. https://doi.org/10.1080/00021369.1987.10868474
Moon MW, Kim HJ, Oh TK, Shin CS, Lee JS, Kim SJ, Lee JK (2005) Analyses of enzyme II gene mutants for sugar transport and heterologous expression of fructokinase gene in Corynebacterium glutamicum ATCC 13032. Fems Microbiol Lett 244:259–266. https://doi.org/10.1016/j.femsle.2005.01.053
Moon MW, Park SY, Choi SK, Lee JK (2007) The phosphotransferase system of Corynebacterium glutamicum: features of sugar transport and carbon regulation. J Mol Microbiol Biotechnol 12:43–50. https://doi.org/10.1159/000096458
Morinaga T, Matsuse T, Ashida H, Yoshida K (2010) Differential substrate specificity of two inositol transporters of Bacillus subtilis. Biosci Biotechnol Biochem 74:1312–1314. https://doi.org/10.1271/bbb.100125
Porco A, Peekhaus N, Bausch C, Tong S, Isturiz T, Conway T (1997) Molecular genetic characterization of the Escherichia coli gntT gene of GntI, the main system for gluconate metabolism. J Bacteriol 179:1584–1590. https://doi.org/10.1128/jb.179.5.1584-1590.1997
Reizer A, Deutscher J, Saier MH Jr, Reizer J (1991) Analysis of the gluconate (gnt) operon of Bacillus subtilis. Mol Microbiol 5:1081–1089. https://doi.org/10.1111/j.1365-2958.1991.tb01880.x
Seibold GM, Wurst M, Eikmanns BJ (2009) Roles of maltodextrin and glycogen phosphorylases in maltose utilization and glycogen metabolism in Corynebacterium glutamicum. Microbiology 155:347–358. https://doi.org/10.1099/mic.0.023614-0
Stansen C, Uy D, Delaunay S, Eggeling L, Goergen JL, Wendisch VF (2005) Characterization of a Corynebacterium glutamicum lactate utilization operon induced during temperature-triggered glutamate production. Appl Environ Microbiol 71:5920–5928. https://doi.org/10.1128/AEM.71.10.5920-5928.2005
Tanaka Y, Okai N, Teramoto H, Inui M, Yukawa H (2008a) Regulation of the expression of phosphoenolpyruvate: carbohydrate phosphotransferase system (PTS) genes in Corynebacterium glutamicum R. Microbiology 154:264–274. https://doi.org/10.1099/mic.0.2007/008862-0
Tanaka Y, Teramoto H, Inui M, Yukawa H (2008b) Regulation of expression of general components of the phosphoenolpyruvate: carbohydrate phosphotransferase system (PTS) by the global regulator SugR in Corynebacterium glutamicum. Appl Microbiol Biotechnol 78:309–318. https://doi.org/10.1007/s00253-007-1313-1
Tanaka Y, Teramoto H, Inui M, Yukawa H (2009) Identification of a second beta-glucoside phosphoenolpyruvate: carbohydrate phosphotransferase system in Corynebacterium glutamicum R. Microbiology 155:3652–3660. https://doi.org/10.1099/mic.0.029496-0
Tanaka Y, Teramoto H, Inui M, Yukawa H (2011) Translation efficiency of antiterminator proteins is a determinant for the difference in glucose repression of two beta-glucoside phosphotransferase system gene clusters in Corynebacterium glutamicum R. J Bacteriol 193:349–357. https://doi.org/10.1128/JB.01123-10
Tanaka Y, Ehira S, Teramoto H, Inui M, Yukawa H (2012) Coordinated regulation of gnd, which encodes 6-phosphogluconate dehydrogenase, by the two transcriptional regulators GntR1 and RamA in Corynebacterium glutamicum. J Bacteriol 194:6527–6536. https://doi.org/10.1128/jb.01635-12
Tateno T, Fukuda H, Kondo A (2007) Production of l-Lysine from starch by Corynebacterium glutamicum displaying alpha-amylase on its cell surface. Appl Microbiol Biotechnol 74:1213–1220. https://doi.org/10.1007/s00253-006-0766-y
Taw MN, Lee HI, Lee SH, Chang WS (2015) Characterization of MocR, a GntR-like transcriptional regulator, in Bradyrhizobium japonicum: its impact on motility, biofilm formation, and soybean nodulation. J Microbiol 53:518–525. https://doi.org/10.1007/s12275-015-5313-z
Teramoto H, Inui M, Yukawa H (2011) Transcriptional regulators of multiple genes involved in carbon metabolism in Corynebacterium glutamicum. J Biotechnol 154:114–125. https://doi.org/10.1016/j.jbiotec.2011.01.016
Wang YY, Xu JZ, Zhang WG (2019) Metabolic engineering of l-leucine production in Escherichia coli and Corynebacterium glutamicum: a review. Crit Rev Biotechnol 39:633–647. https://doi.org/10.1080/07388551.2019.1577214
Wendisch VF, de Graaf AA, Sahm H, Eikmanns BJ (2000) Quantitative determination of metabolic fluxes during coutilization of two carbon sources: comparative analyses with Corynebacterium glutamicum during growth on acetate and/or glucose. J Bacteriol 182:3088–3096. https://doi.org/10.1128/jb.182.11.3088-3096.2000
Xu J, Zhang J, Liu D, Zhang W (2016) Increased glucose utilization and cell growth of Corynebacterium glutamicum by modifying the glucose-specific phosphotransferase system (PTS(Glc)) genes. Can J Microbiol 62:983–992. https://doi.org/10.1139/cjm-2016-0027
Xu JZ, Yang HK, Zhang WG (2018) NADPH metabolism: a survey of its theoretical characteristics and manipulation strategies in amino acid biosynthesis. Crit Rev Biotechnol 38:1061–1076. https://doi.org/10.1080/07388551.2018.1437387
Xu JZ, Yu HB, Han M, Liu LM, Zhang WG (2019) Metabolic engineering of glucose uptake systems in Corynebacterium glutamicum for improving the efficiency of l-lysine production. J Ind Microbiol Biotechnol 46:937–949. https://doi.org/10.1007/s10295-019-02170-w
Xu JZ, Ruan HZ, Yu HB, Liu LM, Zhang W (2020) Metabolic engineering of carbohydrate metabolism systems in Corynebacterium glutamicum for improving the efficiency of l-lysine production from mixed sugar. Microb Cell Fact 19:39. https://doi.org/10.1186/s12934-020-1294-7
Zhang C, Zhang J, Kang Z, Du G, Chen J (2015) Rational engineering of multiple module pathways for the production of l-phenylalanine in Corynebacterium glutamicum. J Ind Microbiol Biotechnol 42:787–797. https://doi.org/10.1007/s10295-015-1593-x
Zhou Z, Wang C, Xu H, Chen Z, Cai H (2015) Increasing succinic acid production using the PTS-independent glucose transport system in a Corynebacterium glutamicum PTS-defective mutant. J Ind Microbiol Biotechnol 42:1073–1082. https://doi.org/10.1007/s10295-015-1630-9
Acknowledgements
We thank Prof. Weiguo Zhang from School of Biotechnology at Jiangnan University for assistance in manuscript writing. This work was supported by the National Natural Science Foundation of China (Grant number 31601459); the China Postdoctoral Science Foundation (Grant number 2016M590410); and the Top-Notch Academic Programs Project of Jiangsu Higher Education Institutions, the 111 project (Grant number 111-2-06).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ruan, H., Yu, H. & Xu, J. The glucose uptake systems in Corynebacterium glutamicum: a review. World J Microbiol Biotechnol 36, 126 (2020). https://doi.org/10.1007/s11274-020-02898-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-020-02898-z