Abstract
Escherichia coli FB-04(pta1), a recombinant l-tryptophan production strain, was constructed in our laboratory. However, the conversion rate (l-tryptophan yield per glucose) of this strain is somewhat low. In this study, additional genes have been deleted in an effort to increase the conversion rate of E. coli FB-04(pta1). Initially, the pykF gene, which encodes pyruvate kinase I (PYKI), was inactivated to increase the accumulation of phosphoenolpyruvate, a key l-tryptophan precursor. The resulting strain, E. coli FB-04(pta1)ΔpykF, showed a slightly higher l-tryptophan yield and a higher conversion rate in fermentation processes. To further improve the conversion rate, the phosphoenolpyruvate:glucose phosphotransferase system (PTS) was disrupted by deleting the ptsH gene, which encodes the phosphocarrier protein (HPr). The levels of biomass, l-tryptophan yield, and conversion rate of this strain, E. coli FB-04(pta1)ΔpykF/ptsH, were especially low during fed-batch fermentation process, even though it achieved a significant increase in conversion rate during shake-flask fermentation. To resolve this issue, four HPr mutations (N12S, N12A, S46A, and S46N) were introduced into the genomic background of E. coli FB-04(pta1)ΔpykF/ptsH, respectively. Among them, the strain harboring the N12S mutation (E. coli FB-04(pta1)ΔpykF-ptsHN12S) showed a prominently increased conversion rate of 0.178 g g−1 during fed-batch fermentation; an increase of 38.0% compared with parent strain E. coli FB-04(pta1). Thus, mutation of the genomic of ptsH gene provided an alternative method to weaken the PTS and improve the efficiency of carbon source utilization.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
l-Tryptophan, an important amino acid for humans and other animals, is extensively used as a supplement in food and animal feed, as well as for medicinal purposes [12, 15]. Given that the l-tryptophan biosynthetic pathway in microorganisms is fairly long and its regulation is intricate, the underlying regulatory networks have been modified in a variety of ways to improve l-tryptophan yield [1, 17, 28, 34]. l-Tryptophan production through Escherichia coli fermentation has been studied extensively, and l-tryptophan production strains with completely defined genetic traits and high yields have been constructed [4, 10, 13, 16]. In recent efforts to reduce productions costs during fermentation, effective utilization of carbon source has attracted considerable attention [6, 18, 30, 31].
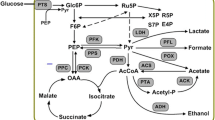
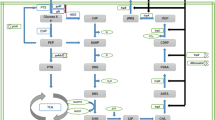
Phosphoenolpyruvate, which is a product of glycolysis, is an essential l-tryptophan precursor [18, 33]. In glycolysis, pyruvate kinases (PYKI and PYKII, EC 2.7.1.40, which are encoded by the pykF and pykA genes, respectively) catalyze the formation of pyruvate from phosphoenolpyruvate [31]. It was well known that knocking out either pykA or pykF increases the accumulation of phosphoenolpyruvate, which contributes to the production of aromatic amino acids [14]. The phosphoenolpyruvate:glucose phosphotransferase system (PTS), which is primarily responsible for the uptake and phosphorylation of sugars, consumes phosphoenolpyruvate. The PTS generally contains two central phosphoryl carriers, EI and HPr, which are encoded by ptsI and ptsH, respectively [9, 27]. Given that the PTS consumes phosphoenolpyruvate during glucose transfer, the PTS is extensively distributed in eubacteria, forming non-PTS glucose uptake mutants [3, 8]. Moreover, the PTS plays an important role in the regulation of carbon metabolism, which has a close relationship with normal cellular physiology [5, 22, 23, 27]. Remarkably, the effects of genomic mutations (other than deletion) in ptsI or ptsH on glucose consumption have gone under-reported, even though this strategy may increase the conversion rate without the deleterious effects of PTS disruption.
E. coli FB-04(pta1), a recombinant l-tryptophan production strain, was recently constructed and stored in our laboratory [16]. Unfortunately, this strain displays a relatively low conversion rate (l-tryptophan per glucose). Two series of experiments were performed to improve the conversion rate of this strain. In the first, a pykF deletion strain (ΔpykF), a pykA deletion strain (ΔpykA), and a pykF/pykA double deletion strain (ΔpykF/pykA) were constructed, and their ability to produce l-tryptophan was evaluated in the shake-flask fermentations. These results led to a construction of four strains containing the pykF deletion, but also harboring a mutation in the ptsH gene. The most promising of these strains, E. coli FB-04(pta1) ΔpykF-ptsHN12S, showed a remarkably increased conversion rate.
Materials and methods
Bacterial strains and plasmids
All bacterial strains and plasmids used or constructed in this study are listed in Table 1. E. coli strain JM109, obtained from Novagen (Madison, USA), was used for gene cloning and plasmid construction. Previously, recombinant E. coli FB-04(pta1) harboring plasmid pSTV-03 was constructed and reserved in our laboratory [16].
The pMD18-T simple vector was obtained from Takara (Dalian, China). Plasmids pKD13, pKD46, and pCP20, which were purchased from the E. coli Genetic Stock Center (Yale University, New Haven, CT, USA), were used for gene knockout procedures. Plasmid pMD-SK, which contains the kanamycin resistance gene (kan) and the levansucrase gene (sacB), was previously constructed and reserved in our laboratory [16].
Plasmid construction and gene mutation
The sequences of the primers designed for PCR amplification are shown in Table 2. Plasmid and chromosomal DNA were prepared using the Plasmid Mini-Prep Kit and Genomic DNA Isolation Kit, respectively, obtained from BIO Basic Inc. The preparation of competent cells and their transformation with plasmids were carried out using criterion processes [24].
To clone ptsH, the ptsH gene (GenBank accession number: gi 946886) was first amplified from E. coli FB-04(pta1) genomic DNA using V-ptsH primers (Table 2). The resulting DNA fragment was ligated into the pMD18-T simple vector to form plasmid pMD-ptsH. Plasmids pMD-ptsHN12S, pMD-ptsHN12A, pMD-ptsHS46A, and pMD-ptsHS46N were constructed using PCR-mediated site-directed mutagenesis, with plasmid pMD-ptsH as the template and primers C-ptsHN12S, C-ptsHN12A, C-ptsHS46A, and C-ptsHS46N, respectively (Table 2).
Construction of single- or multi-gene knockout mutants was performed using the λ Red recombination method [2]. To delete the pykF gene (GenBank accession number gi 946179) or the pykA gene (GenBank accession number: gi 946527), a DNA fragment including a kan fragment plus homologous sequences was amplified by PCR from plasmid pKD13, using primers P- pykF or P- pykA (Table 2). To delete the ptsH gene, a DNA fragment containing a kan-sacB fragment plus homologous sequences was amplified from plasmid pMD-SK using P-ptsH primers (Table 2). The homologous sequences used for homologous recombination are represented by the underlined letters. The resulting kan- or kan-sacB-containing fragment was transduced into competent cells harboring plasmid pKD46. The screening of mutant gene was carried out using kanamycin resistance. PCR using primers V- pykF, V- pykA, and V-ptsH primers was employed to verify the pykF, pykA, and ptsH gene knockout mutations, respectively (Table 2). Finally, the kan fragment in the pykF or pykA deletion strain was eliminated using helper plasmid pCP20. For the scar-less insertion of ptsH mutants into the genome, the corresponding ptsH mutant fragment, which was amplified from plasmid pMD-ptsHN12S, pMD-ptsHN12A, pMD-ptsHS46A, or pMD-ptsHS46N, respectively, using primers V-ptsH (Table 2), was transduced into the ptsH single-gene knockout mutant harboring plasmid pKD46, and sucrose (15% w/v) was as a selection marker. DNA sequencing was also carried out to verify all genetic mutations.
Media and culture conditions
For shake-flask fermentation, seed cultures of strain FB-04(pta1) and its derivatives were grown in Luria–Bertani medium (10.0 g L−1 tryptone, 5.0 g L−1 yeast extract, and 10.0 g L−1 NaCl) supplemented with 30 μg mL−1 kanamycin. These cultures were incubated at 37 °C for 10 h in a rotary shaker (200 rpm). Aliquots (200 μL) of these seed cultures were added into flask fermentation medium (100 mL), which contained (per L): 24 g K2HPO4, 9.6 g KH2PO4, 10 g glucose, 1 g MgSO4·7H2O, 2 g citric acid, 5 g (NH4)2SO4, 15 g yeast, and 3 mL of trace element solution [17]. The above-mentioned cultures were incubated in 500-mL flasks supplemented with 30 μg mL−1 kanamycin for 48 h at 37 °C with shaking (200 rpm). During the fermentation, 10 g L−1 glucose and 20% NH4OH were added to the medium every 8 h to provide carbon source and maintain the pH (6.5–7.2), respectively.
For fed-batch cultivation in a 3-L fermentor, seed cultures of strain FB-04(pta1) and its derivatives were initiated by inoculating LB medium (100 mL supplementing with 30 μg mL−1 kanamycin) in a 500-mL shake flask with 200 μL of glycerol stock (kept frozen at −80 °C). The resulting culture was incubated at 37 °C with shaking at 200 rpm for 10 h. These seed cultures (100 mL) were then transferred to a 3-L fermentor (BioFlo 110, New Brunswick Scientific Co., Edison, NJ) charged with 0.9 L of fermentation medium at pH 6.5 containing (per L): 15 g K2HPO4, 2 g yeast, 2 g MgSO4·7H2O, 2 g citric acid, 1.6 g (NH4)2SO4, 7.5 g glucose, 0.0129 g CaCl2, 0.075 g FeSO4·7H2O per liter, and 3 mL trace element solution [17], and added with kanamycin at a final concentration of 30 μg mL−1. The 3-L fermentations were performed at 37 °C for about 54 h with the pH at 6.5. The pH and dissolved oxygen (20%) were maintained by automatically adding NH4OH and adjusting the agitation speed, respectively. Samples were collected every 3–4 h. After the initial glucose was exhausted, additional glucose (800 g L−1) was added to the fermentation medium using an exponential feeding program, which maintained a cell-specific growth (OD600) rate of 0.15 h−1. The term glucose consumption refers to the total amount of glucose consumed by cells per liter, and the ratio of l-tryptophan yield divided by the glucose consumption is termed conversion rate.
Analytical methods
The concentration of cell was determined using the optical density at 600 nm (OD600) after proper dilution. To determine the levels of extracellular metabolites, the fermentation broth was clarified using centrifugation at 14,000×g for 10 min, and then reserved at −30 °C until it was analyzed by employing high-performance liquid chromatography (HPLC). The levels of glucose and acetate were measured at 50 °C with an Aminex HPX-87H column (300 mm × 7.8 mm; Bio-Rid, Hercules, CA). The mobile phase consisted of 50 mM H2SO4 and the flow rate was 0.5 mL min−1. The l-tryptophan concentration was determined using a previously described HPLC method [34].
Statistical analyses
All fermentation experiments were repeated at least three times. The fermentation parameters are displayed as the mean ± standard deviation of these replicates. Statistically significant (P < 0.01) differences were identified using Student’s t test.
Results
Inactivation of pyruvate kinase (PYK)
To investigate the effect of pyruvate kinase (PYK) inactivation on l-tryptophan production, the individual genes pykF and pykA, as well as the combination of pykF and pykA were deleted from the genome of E. coli FB-04(pta1) using λ Red recombination. We will refer to the resulting strains as ΔpykF, ΔpykA, and ΔpykF/pykA, respectively. Then, the fermentation performance of these new strains was compared with that of E. coli FB-04(pta1) in a series of shake-flask cultures. As shown in Fig. 1a, strains ΔpykF and ΔpykA displayed growth curves similar to that of parent strain FB-04(pta1). Among all the strains tested, ΔpykF achieved the highest total biomass, while the growth of strain ΔpykF/pykA was restricted (Table 4).
Shake-flask fed-batch fermentation of strains FB-04(pta1), ΔpykF, ΔpykA, andΔpykF/pykA. a Biomass levels (OD600); b l-tryptophan levels; c acetate levels; d glucose consumption levels and conversion rate. FB-04(pta1) (filled square); ΔpykF (filled circle); ΔpykA (upright open triangle); ΔpykF/pykA (inverted open triangle)
Throughout the fermentation process, the l-tryptophan levels of strains ΔpykF and ΔpykA were higher than that of FB-04(pta1), while strain ΔpykF/pykA showed a significant decrease (Fig. 1b). The final l-tryptophan titers of strains ΔpykF and ΔpykA were 2.43 and 2.25 g L−1, leading to significant increases of 16 and 8% (P < 0.01, Student’s t test), respectively, compared with FB-04(pta1) (Table 3). Acetate levels were also determined (Fig. 1c). Strains ΔpykF, ΔpykA, and ΔpykF/pykA showed 19, 10, and 49% decreases in acetate formation, respectively, compared with that of FB-04(pta1) (Table 4). The final glucose consumption of strains ΔpykF, ΔpykA, and ΔpykF/pykA was lower than that of parent strain FB-04(pta1), resulting in decreases of 13, 11, and 37%, respectively (Fig. 1d). In addition, the conversion rate (l-tryptophan yield per glucose) of strain ΔpykF was higher than that of strains FB-04(pta1), ΔpykA, and ΔpykF/pykA (Fig. 1d), leading to significant increase of 33, 16, and 27% (P < 0.01, Student’s t test), respectively.
Deletion of gene ptsH
To abolish the consumption of phosphoenolpyruvate in the glucose uptake process, the phosphoenolpyruvate:glucose phosphotransferase system (PTS) of strain ΔpykF was disrupted by genomic deletion of the ptsH gene, forming strain FB-04(pta1)ΔpykF/ptsH, which we will refer to as ΔpykF/ptsH. During shake-flask fermentation, the biomass level, l-tryptophan production, and glucose consumption of strain ΔpykF/ptsH were significantly lower than those of ΔpykF (Fig. 2a–c). The final glucose consumption of strain ΔpykF/ptsH was significantly lower than that of strain ΔpykF (Fig. 2d). However, the conversion rate of strain ΔpykF/ptsH was about twice that of the strain ΔpykF (Table 4).
Effect of ptsH modification on l-tryptophan production. a Biomass levels (OD600); b l-tryptophan levels; c acetate levels; d glucose consumption levels and conversion rate. ΔpykF (filled circle); ΔpykF/ptsH (open circle); ΔpykF-ptsHN12S (filled rhombus); ΔpykF-ptsHN12A (open rhombus); ΔpykF-ptsHS46A (filled pentagon); ΔpykF-ptsHS46N (open pentagon)
Genomic substitution of ptsH mutants
The phosphocarrier protein (HPr) of the PTS has been thoroughly studied, and a series of HPr mutants which led to different kinetic properties of enzyme I (EI) were found [19, 20]. In this study, HPr mutants N12A, N12S, S46A, and S46N were introduced into the genomic background of ΔpykF/ptsH to restore the growth and glucose uptake restrictions of ΔpykF/ptsH caused by its ptsH deletion. This process produced strains FB-04(pta1)ΔpykF-ptsHN12A, FB-04(pta1)ΔpykF-ptsHN12S, FB-04(pta1)ΔpykF-ptsHN46A, and FB-04(pta1)ΔpykF-ptsHN46S, which we will refer to as ΔpykF-ptsHN12A, ΔpykF-ptsHN12S, ΔpykF-ptsHN46A, and ΔpykF-ptsHN46S, respectively. The different kinetic properties of EI caused by HPr mutants N12A, N12S, S46A, and S46N are displayed in Table 3. To determine the performance of the mutant strains, they were cultivated in shake flasks. ΔpykF-ptsHN12S grew slightly faster than ΔpykF during the entire fermentation process (Fig. 3a). The final biomass of strain ΔpykF-ptsHN12A was the highest among all the strains. Strain ΔpykF-ptsHS46A achieved the second-highest biomass, even though the growth of ΔpykF-ptsHN12A and ΔpykF-ptsHS46A was lower than that of ΔpykF at the first 24 h. The growth of ΔpykF-ptsHS46N was restricted, although it was slightly higher than that of ΔpykF/ptsH during the entire fermentation process.
The l-tryptophan titers of all strains increased with time during the entire shake-flask fermentation process (Fig. 2b). Compared with strain ΔpykF, the final l-tryptophan titers of ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were 2.65, 3.25, and 2.84 g L−1, corresponding to significant increases of 9, 34, and 17% (P < 0.01, Student’s t test), respectively, compared with ΔpykF. The final l-tryptophan titer of ΔpykF-ptsHS46N was extremely low and did not represent a significant improvement over ΔpykF/ptsH (Table 4). The final acetate levels of ΔpykF-ptsHN12S, ΔpykF-ptsHS46A, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46N were successively lower than that of ΔpykF (Table 4). The glucose consumption of strains ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A was lower than that of ΔpykF (Fig. 2d), while the conversion rates of ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A increased by 18, 45, and 25% (P < 0.01, Student’s t test), respectively (Table 4). However, the glucose consumption and conversion rate of ΔpykF-ptsHS46N were extremely low and did not represent a significant improvement over ΔpykF/ptsH (Table 4).
3-L fed-batch fermentation
To further investigate l-tryptophan production by the strains containing ptsH mutations, ΔpykF, ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were cultivated in separate 3-L fermentations, and the results were compared with those of the parent FB-04(pta1) and ΔpykF/ptsH (Fig. 3). The growth curves of FB-04(pta1) and ΔpykF were similar, while the growth curves of ΔpykF/ptsH, ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were similar (Fig. 3a). Among all strains, ΔpykF achieved the highest total biomass, while the growth of ΔpykF/ptsH was extremely impaired.
l-Tryptophan production levels were determined (Fig. 3b). l-Tryptophan production by FB-04(pta1), ΔpykF, ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A increased with time, while the l-tryptophan yield of ΔpykF/ptsH increased continuously over the first 24 h of fermentation, then remained almost unchanged. Throughout the entire fermentation process, the l-tryptophan yield of ΔpykF was slightly higher than that of the parent strain FB-04(pta1), while the l-tryptophan yields of ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were lower than that of the parent strain, and the l-tryptophan yield of ΔpykF/ptsH was the lowest. The final l-tryptophan titers of ΔpykF, ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were 45.5, 26.0, 9.6, and 14.1 g L−1, which do not represent a significant improvement over that of FB-04(pta1) (44.0 g L−1) (Table 4). Even so, the conversion rates of ΔpykF and ΔpykF-ptsHN12S were significantly higher than that of FB-04(pta1) (Fig. 3c), leading to increases of 8.5 and 38.0% (P < 0.01, Student’s t test), respectively. The conversion rates of ΔpykF-ptsHN12A, ΔpykF-ptsHS46A, and ΔpykF/ptsH were lower (Table 4). Moreover, compared with parent strain FB-04(pta1), the final acetate levels of all of the mutants were diminished (Table 4).
Discussion
The l-tryptophan biosynthetic pathway and its complicated regulation mechanism have been studied for a long time, and various l-tryptophan-producing strains with defined genetic modifications have been constructed [4, 10, 12, 13, 28, 29, 34]. In a previous effort to produce a useful l-tryptophan production strain, feedback-resistant variants of the enzymes of the l-tryptophan biosynthetic pathway were overexpressed in the l-tryptophan production strain E. coli FB-04(pta1), which harbors plasmid pSTV-03 [16]. However, this strain has a lowered conversion rate (l-tryptophan yield per glucose), which diminishes its utility. Increasing the supply of the crucial l-tryptophan precursor phosphoenolpyruvate and erythrose-4-phosphate is an additional strategy commonly used to boost l-tryptophan production [18].
To overcome possible limitations in the supply of phosphoenolpyruvate, a pykF deletion strain (ΔpykF), a pykA deletion strain (ΔpykA), and a pykF/pykA double deletion strain (ΔpykF/pykA) were constructed, and their ability to produce l-tryptophan was evaluated in the shake-flask fermentations. The results showed that strain ΔpykF achieved a higher l-tryptophan yield and conversion rate than that of strains FB-04(pta1), ΔpykA, and ΔpykF/pykA (Table 3). Of the two PYK isoenzymes, PYKI, which is encoded by pykF gene, plays the major role in catalyzing the conversion of phosphoenolpyruvate to pyruvate [31]. In one pykF knockout mutant, the metabolic flux through glycolysis was reduced by 45% and the flux through the pentose phosphate pathway was increased by 45%, compared with a wild-type parent [25]. Moreover, the inactivation of PYK can decrease the metabolic flow through the TCA cycle and diminish the formation of acetate, a by-product which could inhibit the growth of cells [11, 26, 35]. The results obtained with strain ΔpykF in this study are consistent with these previous reports. Double deletion of pykF and pykA may have a significant impact on the normal cellular physiology; thus, the l-tryptophan yield and biomass production of strain ΔpykF/pykA were lower than that of its parent.
Given that the PTS consumes phosphoenolpyruvate for glucose uptake and the constitutively expressed galactose transporter also enables glucose transport [29], the PTS encoded within the genome of ΔpykF was inactivated by constructing the strain ΔpykF/ptsH. The conversion rate of ΔpykF/ptsH was remarkably higher than that of strain ΔpykF in shake-flask fermentation (Table 3), but the deletion of ptsH caused significant decreases in biomass, glucose consumption, and l-tryptophan production (Table 3). It has been reported that, in addition to its role in the transport of sugars, the PTS also plays an important role in the regulation of carbon metabolism [5, 22, 23, 27]. Thus, disruption of the PTS may have a detrimental effect on normal cellular physiology. To increase the l-tryptophan conversion rate without having this detrimental effect on cellular physiology, the PTS activity should, therefore, be weakened, not completely disrupted. When this study was initiated, it was known that HPr mutation could impair the phosphohydrolysis activity of EI and decrease PTS activity, and a series of mutants that altered the kinetic properties of EI had been reported [19, 20]. In this study, HPr mutants N12S, N12A, S46A, and S46N were introduced into the strain ΔpykF/ptsH to weaken, but not disrupt, PTS activity. l-Tryptophan production by the resulting strains ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, ΔpykF-ptsHS46A, and ΔpykF-ptsHS46N was first investigated in shake-flask fermentations. The results showed that the l-tryptophan yields and conversion rates of strains ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were higher than that of ΔpykF (Table 4). The levels of acetate produced by strains ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A were significantly lower than that produced by ΔpykF, which is consistent with the higher biomass they achieved (Fig. 2a). In shake-flask fermentations, the biomass, acetate, l-tryptophan production, and conversion rate of strain ΔpykF-ptsHS46N were not significantly different than that of strain ΔpykF/ptsH (Table 4). However, since the HPr mutants N12S, N12A, S46A, and S46N lead to different EI kinetic properties, these mutations may affect the fermentation performance of strains ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, ΔpykF-ptsHS46A, and ΔpykF-ptsHS46N differently. The weakened PTS could lower the glucose transport rate, lowering the acetate formation rate. In shake-flask cultivation, the dissolved oxygen level was lower, which favors the formation of acetate, a by-product that hampers cell growth and l-tryptophan production. Thus, the significantly decreased acetate levels, compared with the parent stain, of PTS-modified strains in shake-flask cultivation contributed to cell growth and l-tryptophan production.
It was worth noting that the fermentation performances of strains ΔpykF-ptsHN12S, ΔpykF-ptsHN12A, and ΔpykF-ptsHS46A in fed-batch fermentation were different from the fermentation performances observed for these strains in shake flasks (Fig. 3). In fed-batch cultivation, the dissolved oxygen levels of all strains were maintained at 20%. Under these conditions, the acetate levels of all strains were low enough throughout fed-batch fermentation process (Fig. 3c) that there was no effect on cell growth or l-tryptophan production. The conversion rate of ΔpykF-ptsHN12S, which reached 0.178, was 27.1% greater than that of ΔpykF, even though its l-tryptophan yield was lower (Table 4). The l-tryptophan yields and conversion rates of strains ΔpykF-ptsHN12A and ΔpykF-ptsHS46A were significantly lower (Table 4). HPr mutations N12A, S46A, and S46N increase K m of EI levels that are 2, 2.5, and 11 times that of the wild type, respectively (Table 3). Given that the phosphoenolpyruvate content of these cells is relatively low, the altered EI K m s may have remarkable effects on normal physiological functions. This is consistent with fermentation performances of strains ΔpykF-ptsHN12A, ΔpykF-ptsHS46A, and ΔpykF-ptsHS46N observed in this study (Fig. 3). Moreover, mutant N12S had no effect on K m of EI, and only decreased V max of EI to 65% that of the wild type (Table 3). The altered EI V max may influence the consumption of glucose and the growth rate, which is consistent with the observed fermentation performance of ΔpykF-ptsHN12S (Fig. 3).
Measuring the conversion rate is a standard way to evaluate the efficiency industrial fermentation processes. Increasing the conversion rate reduces production costs and improves the efficiency of carbon source utilization. Various strategies have been used to increase the carbon flux through the aromatic pathway in efforts to improve the production and conversion rate of aromatic amino acids. PEP synthase and transketolase were overexpressed to enhance the availability of the precursor substrates PEP and E4P, which improved aromatic amino acid production and the conversion rate [18, 21]. Carbon storage regulator (Csr) was overexpressed to increase PEP levels and channel carbon flux into the aromatic pathway [32]. In addition, fructose repressor (FruR) has been inactivated to enhance the carbon flux through glycolysis and the pentose phosphate pathway, improving the conversion rate by 52.3% [17]. However, inactivating FruR also led to increased levels of acetate [17]. In this study, introducing mutations into the ptsH gene provided a novel way to increase the conversion rate as well as decreasing acetate formation by E. coli FB-04(pta1) during the l-tryptophan fermentation process, even though the l-tryptophan yield of the final strain, ΔpykF-ptsHN12S, decreased. One potential method to increase the l-tryptophan yield and conversion rate at the same time would be to introduce mutations into the ptsH gene using error-prone PCR and screening the resulting strains for superior l-tryptophan fermentative capability. Alternatively, given that both erythrose-4-phosphate, another crucial l-tryptophan precursor [18, 32,33,34], and NADPH, a substrate in the common aromatic pathway [7, 16], come from the pentose phosphate pathway, l-tryptophan production, and the conversion rate should be enhanced by increasing the metabolic flow through the pentose phosphate pathway using genetic engineering methods. In addition, considering that the glucose feeding rate and fermentation temperature can be used as optimization factors [6, 33], a series of optimization strategies based on these two factors should be undertaken to improve l-tryptophan production. Finally, it was well known that glycerol was not transported by PTS; thus, glycerol could be used as a supplementary carbon source during the fermentation process to try to increase l-tryptophan production.
Conclusions
In efforts to increase the conversion rate of l-tryptophan production by E. coli FB-04(pta1), the pykF gene encoding pyruvate kinase I (PYKI) was deleted. The resulting strain (ΔpykF) showed a slightly higher l-tryptophan yield and conversion rate in fermentation processes. To further improve the conversion rate, the PTS was disrupted by deleting the ptsH gene, which encodes phosphocarrier protein HPr. The resulting strain (ΔpykF/ptsH) achieved a significant increase in conversion rate during shake-flask fermentations. However, the biomass level, l-tryptophan yield, and conversion rate of ΔpykF/ptsH were especially low during the fed-batch fermentation process. Finally, HPr mutant N12S was introduced in the genomic background of ΔpykF/ptsH. The resulting strain (ΔpykF-ptsHN12S), which showed prominent fermentation performance, displayed a conversion rate 38.0% higher than that of the parent strain E. coli FB-04(pta1). The strategies employed in this study improved the efficiency of carbon source utilization and are of great value for industrial l-tryptophan production.
References
Azuma S, Tsunekawa H, Okabe M, Okamoto R, Aiba S (1993) Hyper-production of l-trytophan via fermentation with crystallization. Appl Microbiol Biotechnol 39:471–476
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, Datsenko KA, Tomita M, Wanner BL, Mori H (2006) Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol 2:1–11
Baez JL, Bolivar F, Gosset G (2001) Determination of 3-deoxy-d-arabino-heptulosonate 7-phosphate productivity and yield from glucose in Escherichia coli devoid of the glucose phosphotransferase transport system. Biotechnol Bioeng 73:530–535
Chan EC, Tsai HL, Chen SL, Due-Gang Mou (1993) Amplification of the tryptophan operon gene in Escherichia coli chromosome to increase l-tryptophan biosynthesis. Appl Microbiol Biotechnol 40:301–305
Deutscher J, Francke C, Postma PW (2006) How phosphotransferase system-related protein phosphorylation regulates carbohydrate metabolism in bacteria. Microbiol Mol Biol Rev 70:939–1031
Dodge TC, Gerstner JM (2002) Optimization of the glucose feed rate profile for the production of tryptophan from recombinant E. coli. J Chem Technol Biotechnol 77:1238–1245
Dong X, Chen X, Qian Y, Wang Y, Wang L, Qiao W, Liu L (2016) Metabolic engineering of Escherichia coli W3110 to produce l-malate. Biotechnol Bioeng. doi:10.1002/bit.26190
Escalante A, Calderon R, Valdivia A, de Anda R, Hernandez G, Ramirez OT, Gosset G, Bolivar F (2010) Metabolic engineering for the production of shikimic acid in an evolved Escherichia coli strain lacking the phosphoenolpyruvate: carbohydrate phosphotransferase system. Microb Cell Fact. doi:10.1186/1475-2859-9-21
Garcia-Alles LF, Flukiger K, Hewel J, Gutknecht R, Siebold C, Schurch S, Erni B (2002) Mechanism-based inhibition of enzyme I of the Escherichia coli phosphotransferase system. Cysteine 502 is an essential residue. J Biol Chem 277:6934–6942
Gu PF, Yang F, Li FF, Liang QF, Qi QS (2013) Knocking out analysis of tryptophan permeases in Escherichia coli for improving l-tryptophan production. Appl Microbiol Biotechnol 97:6677–6683
Han K, Lim HC, Hong J (1992) Acetic acid formation in Escherichia coli fermentation. Biotechnol Bioeng 39:663–671
Ikeda M (2006) Towards bacterial strains overproducing l-tryptophan and other aromatics by metabolic engineering. Appl Microbiol Biotechnol 69:615–626
Ikeda M, Katsumata R (1999) Hyperproduction of tryptophan by Corynebacterium glutamicum with the modified pentose phosphate pathway. Appl Environ Microbiol 65:2497–2502
Kedar P, Colah R, Shimizu K (2007) Proteomic investigation on the pyk-F gene knockout Escherichia coli for aromatic amino acid production. Enzyme Microb Tech 41:455–465
Leuchtenberger W, Huthmacher K, Drauz K (2005) Biotechnological production of amino acids and derivatives: current status and prospects. Appl Microbiol Biotechnol 69:1–8
Liu LN, Duan XG, Wu J (2016) l-Tryptophan Production in Escherichia coli improved by weakening the Pta-AckA pathway. PLoS One. doi:10.1371/journal.pone.0158200
Liu LN, Duan XG, Wu J (2016) Modulating the direction of carbon flow in Escherichia coli to improve L-tryptophan production by inactivating the global regulator FruR. J Biotechnol 231:141–148
Lutke-Eversloh T, Stephanopoulos G (2007) l-Tyrosine production by deregulated strains of Escherichia coli. Appl Microbiol Biotechnol 75:103–110
Napper S, Anderson JW, Georges F, Quail JW, Delbaere LTJ, Waygood EB (1996) Mutation of serine-46 to aspartate in the histidine-containing protein of Escherichia coli mimics the inactivation by phosphorylation of serine-46 in HPrs from gram-positive bacteria. Biochemistry 35:11260–11267
Napper S, Delbaere LTJ, Waygood EB (1999) The aspartyl replacement of the active site histidine in histidine-containing protein, HPr, of the Escherichia coli phosphoenolpyruvate: sugar phosphotransferase system can accept and donate a phosphoryl group. Spontaneous dephosphorylation of acyl-phosphate autocatalyzes an internal cyclization. J Biol Chem 274:21776–21782
Patnaik R, Liao JC (1994) Engineering of Escherichia coli central metabolism for aromatic metabolite production with near theoretical yield. Appl Environ Microbiol 60:3903–3908
Saier MH Jr (1989) Protein phosphorylation and allosteric control of inducer exclusion and catabolite repression by the bacterial phosphoenolpyruvate: sugar phosphotransferase system. Microbiol Rev 53:109–120
Saier MH, Reizer J (1994) The Bacterial phosphotransferase system—new frontiers 30 years later. Mol Microbiol 13:755–764
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, vol 6, 3rd edn. Cold Spring Harbor Laboratory Press, New York, pp 4–6
Siddiquee KA, Arauzo-Bravo MJ, Shimizu K (2004) Effect of a pyruvate kinase (pykF-gene) knockout mutation on the control of gene expression and metabolic fluxes in Escherichia coli. FEMS Microbiol Lett 235:25–33
Siddiquee KA, Arauzo-Bravo MJ, Shimizu K (2004) Metabolic flux analysis of pykF gene knockout Escherichia coli based on C-13-labeling experiments together with measurements of enzyme activities and intracellular metabolite concentrations. Appl Microbiol Biotechnol 63:407–417
Steinsiek S, Bettenbrock K (2012) Glucose transport in Escherichia coli mutant strains with defects in sugar transport systems. J Bacteriol 194:5897–5908
Tribe DE, Pittard J (1979) Hyperproduction of tryptophan by Escherichia coli: genetic manipulation of the pathways leading to tryptophan formation. Appl Environ Microbiol 38:181–190
Wang QZ, Wu CY, Chen T, Chen X, Zhao XM (2006) Expression of galactose permease and pyruvate carboxylase in Escherichia coli ptsG mutant increases the growth rate and succinate yield under anaerobic conditions. Biotechnol Lett 28:203
Wang J, Cheng LK, Wang J, Liu Q, Shen T, Chen N (2013) Genetic engineering of Escherichia coli to enhance production of l-tryptophan. Appl Microbiol Biotechnol 97:7587–7596
Xie XX, Liang Y, Liu HL, Liu Y, Xu QY, Zhang CL, Chen N (2014) Modification of glycolysis and its effect on the production of threonine in Escherichia coli. J Ind Microbiol Biotechnol 41:1007–1015
Yakandawala N, Romeo T, Friesen AD, Madhyastha S (2008) Metabolic engineering of Escherichia coli to enhance phenylalanine production. Appl Microbiol Biotechnol 78:283–291
Zhao CJ, Xie XX, Cheng LK, Xu QY, Chen N (2009) Effect of temperature on the process of Escherichia coli l-tryptophan fermentation. Lett Biotechnol 20(4):534–537
Zhao ZJ, Zou C, Zhu YX, Dai J, Chen S, Wu D, Wu J, Chen J (2011) Development of l-tryptophan production strains by defined genetic modification in Escherichia coli. J Ind Microbiol Biotechnol 38:1921–1929
Zhu T, Phalakornkule C, Koepsel RR, Domach MM, Ataai MM (2001) Cell growth and by-product formation in a pyruvate kinase mutant of E. coli. Biotechnol Prog 17:624–628
Acknowledgements
This work was supported by grants from the National Science Fund for Distinguished Young Scholars (31425020), the project of outstanding scientific and technological innovation group of Jiangsu Province (Jing Wu), the 111 Project (No. 111-2-06), and the Research and Innovation Project for College Graduates of Jiangsu Province (No. KYLX15-1143).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Liu, L., Chen, S. & Wu, J. Phosphoenolpyruvate:glucose phosphotransferase system modification increases the conversion rate during l-tryptophan production in Escherichia coli . J Ind Microbiol Biotechnol 44, 1385–1395 (2017). https://doi.org/10.1007/s10295-017-1959-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-017-1959-3