Abstract
Glycine max (soybean) production can be dramatically affected by frogeye leaf spot (FLS) caused by Cercospora sojina Hara. The inoculation of biocontrol agents may be an alternative strategy for C. sojina control. The native biocontrol bacterium Bacillus sp. CHEP5 reduced the severity of FLS in soybean by inducing systemic resistance. We suggest that the defense response was primed since the expression of the defense related gene GmAOS was enhanced in induced plants treated with both methyl jasmonate and C. sojina. Furthermore, as GmAOS is related to jasmonic acid biosynthesis, we assume that this phytohormone is involved in induced systemic resistance signaling defense pathway in soybean against C. sojina.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glycine max (soybean) is an economically very important crop throughout the word. It is susceptible to many pathogens, with great damage being caused by fungi. Frogeye leaf spot (FLS) caused by the necrotrophic fungi Cercospora sojina Hara can dramatically affect this crop production. FLS symptoms usually appear at V3-V4 phenological stages and led to a premature defoliation (Fehr and Caviness 1977; Mian et al. 2008).
Effective control of FLS disease can be reached by supplying a recommended fungicide, a strategy that could contribute to environmental pollution and to greater production cost, or by using resistant cultivars, which is not always practical since cultivar selection depends on the characteristics of the agricultural area. Biological control of phytopathogenic fungi is eco-friendly and a potential component of integrated disease management.
In nature, plants form beneficial associations with soilborne microbes. Plant growth-promoting bacteria (PGPB) stimulate either directly or indirectly the plant growth. Some PGPB lessen or prevent the deleterious effect of phytopathogenic organisms by antibiosis, competition for space and nutrients, production of siderophores or induction of systemic resistance in plants against a broad-spectrum of root and foliar pathogens (Podile and Kishore 2006; Lugtenberg and Kamilova 2009).
The induced systemic resistance (ISR) is an indirenct biocontrol mechanism that consists in the activation of plant defense state only upon pathogen attack (Kloepper et al. 1992). ISR involves a cascade of defense reactions that spread from the site of induction to distant parts of the plant and encompass signal transduction mediated by phytohormones, generation of phytoalexins, oxidative stress protection, synthesis of enzymes related to plant defense (phenylalanine ammonia-lyase, peroxidase, β-1, 3-glucanase) and formation of structural barriers such as wall thickening, callose deposition and accumulation of phenolics (Reymond and Farmer 1998; Verhagen et al. 2004). A common feature of the resistance responses induced by beneficial microorganisms is priming. This phenomenon is characterized by a faster and/or stronger activation of cellular defense responses upon pathogen attack, resulting in a major resistance to its deleterious effect. Primed plants are sensitized for enhanced defense expression and thus, there is little cost in yield lost to the priming process in the absence of pathogens (Paré et al. 2005; Conrath et al. 2006; Pieterse et al. 2000; Pozo et al. 2008; Van der Ent et al. 2009; Buensanteai et al. 2009).
Although several researchers revealed that beneficial microorganisms triggered the salicylic acid-dependent defense response in the signaling pathway activated by systemic resistance inducing microorganisms (Buensanteai et al. 2009; De Mayer and Höfte 1997), the phytohormones jasmonic acid (JA) and ethylene (ET) are central players in the regulation of ISR (Van der Ent et al. 2009). Interestingly, ISR induced by PGPB is associated with an enhanced sensitivity to these hormones, instead of an augmented production of them (Pieterse et al. 2000).
JA-signaling has also been described as the main pathway in ISR against aboveground herbivores and is stimulated by root-associated microbes (Van der Ent et al. 2009).
Allene oxide synthase (AOS) is a major intermediate enzyme in the oxylipin pathway to JA biosynthesis affecting the synthesis of all JA-related compounds in plants, and therefore playing a significant role in plant defense.
It has been demonstrated that constitutive GmAOS overexpression conferred an enhanced tolerance to cotton worm in transgenic tobacco, by increasing activities of allene oxide synthase, peroxidase, chymotrypsin inhabitor, and the trichome. Moreover, in soybean plants highly resistant to cotton worm this gene expression was faster and greater than in susceptible cultivars.
Previous studies in our laboratory demonstrated that the native isolate Bacillus sp. CHEP5 induces systemic resistance in peanut against the pathogen Sclerotium rolfsii (Tonelli et al. 2011). In this research work, we evaluated the ability of Bacillus sp. CHEP5 to induce soybean systemic defense response against C. sojina, and determined whether it is related to the priming of JA signaling pathway by measuring GmAOS gene expression in induced plants.
Materials and methods
Bacterial strain, pathogen and culture conditions
The native biocontrol bacterium Bacillus sp. CHEP5 was cultured at 28 °C on Luria–Bertani broth (LB) or Agar medium. The strain was kept in 20 % glycerol at −80 °C for long-term storage and in 40 % glycerol at −20 °C for short-term storage.
The fungal pathogen C. sojina CCC 172-09 (Carmona et al. 2009) was obtained from Centro de Referencia en Micología (CEREMIC), Universidad Nacional de Rosario, Argentina. It was grown on potato dextrose agar (PDA) supplemented with streptomycin 100 µg ml−1, at room temperature for 10 days. The phytopathogen was kept in 15 % glycerol at −20 °C for long-term storage or in PDA tubes covered with vaseline.
Plant material and growth conditions
Soybean seeds cv. DM 4676 susceptible to C. sojina, were surface sterilized. Briefly, the seeds were soaked in 96 % ethanol for 20 s followed by 20 % bleach for 20 min, and then washed six times with sterile distilled water (Buensanteai et al. 2009). The surface sterilized seeds were germinated at 28 °C in sterilized Petri dishes with one layer of Whatman #1 filter paper and moist cotton, until the radicle reached approximately 2 cm. Seedlings were sown in plastic cups filled with sterilized quartz sand, watered regularly and supplied once a week with Hoagland medium (Hoagland and Arnon 1950). Plants were grown under controlled environment (light intensity of 200 mmol m−2 s−1, 16-h day/8-h night cycle, at a constant temperature of 28 °C and a relative humidity of 50 %).
Bacterial inoculum preparation
Bacillus sp. CHEP5 was cultured on LB broth for 24 h at 28 °C, until the culture reached an OD620nm = 1, and the number of viable cells was determined (108–109 cfu ml−1 approximately) (Somasegaran and Hoben 1994). The cultures were centrifuged at 4,000 g for 12 min at room temperature and the cells were re-suspended in 10 mM MgSO4 sterile solution.
Phytopathogen inoculum preparation
Cercospora sojina spores were suspended in a sterile aqueous solution containing 0.015 % of the surfactant Tween 20 to a final concentration of 106 spores ml−1.
Bioassays to evaluate the induction of systemic resistance
Soybean seedlings were obtained as described above. Two days after the seedlings were sown in the cups filled with sterilized quartz sand, their roots were inoculated with 4 ml of Bacillus sp. CHEP5 culture to obtain a final concentration of 107 cfu g−1 of quartz sand. At V3-V4 phenological growth stage (10 days after bacterial inoculation), trifoliate leaves were challenged with C. sojina (106 spores ml−1) (Carmona et al. 2009). Twenty-one days after challenge inoculation, disease severity was assessed by determining the disease level and the percentage of damaged leaf area (Carmona et al. 2009). Disease rating was expressed on the basis of symptom severity and damaged leaf area measured in challenged trifoliate leaves: (1) healthy leaves without spots, (2) spots covering 5 % of leaves area approximately, (3) spots covering 15 % of leaves area approximately, (4) spots covering 35 % of leaves area approximately, (5) 50 % or more of their surfaces affected by the FLS. The shoot dry weight and total chlorophyll content were also determined.
Nonpathogenized and non-bacterized control plants were also included. Plants were grown under controlled environment and regularly supplied with Hoagland medium.
The experiment was repeated three times with six replicates for each treatment.
Total chlorophyll determination
The amount of total chlorophyll was determined. Briefly, 0.10–0.15 g of fresh weight soybean leaves was placed into a mortar and the tissue was ground to fine pulp after the addition of 80 % acetone. The extract was transferred to a Buchner funnel containing a pad of Whatman filter paper. While filtering the extract, the leaves pulp was grounded and the final volume of the filtrate was adjusted to 10 ml (Arnon 1949). The chlorophyll content was spectrophotometically determined at 650 and 665 nm. The amount of total chlorophyll was calculated on the basis of µg of chlorophyll gram−1 of fresh leaf tissue, according to the following equation (Mc Kinney 1938):
Total chlorophyll = 6.45 (Absorbance at 665 nm) + 17.72 (Absorbance at 650 nm).
Phytopahtogen and methyl jasmonate treatments
Soybean seedlings were obtained and Bacillus sp. CHEP5 was inoculated as described before. At V3-V4 phenological growth stage, trifoliate leaves were challenged with C. sojina (106 spores ml−1) and hervested at 0 and 1 h after the phytopathogen treatment. To obtain methyl jasmonate (MeJA) treated plants, trifoliate leaves were moistened with an aqueous solution containing 50 μM MeJA and 0.015 % of the surfactant Tween 20 (Pozo et al. 2008). Trifoliate leaves were harvested at 0, 1 and 3 h after MeJA treatment.
Immediately after the leaves were harvested, they were frozen in liquid nitrogen. Samples were conserved at −80 °C until RNA was extracted. Plants that were not induced were used as controls.
The experiment was repeated two times with three replicates for each treatment.
RNA isolation and obtention of soybean cDNA
RNA was isolated from three replicates samples that were pooled to reduce noise arising from biological variation. Leaf samples were ground with sterile mortars and pestles under liquid nitrogen. Total RNA was isolated using the NucleoSpin RNA Clean-Up Kit (Macherey–Nagel) according to the manufacture´s protocol. Traces of genomic DNA were eliminated by DNase digestion (Macherey–Nagel). First strand cDNA synthesis was carried out using MMLV Reverse Transcriptase 1st Strand cDNA Synthesis Kit (Epicentre) according to the manufacture´s specifications with random primers. The reaction conditions for cDNA synthesis were 2 min at 65 °C, then 60 min at 37 °C and finally 5 min at 85 °C.
Determination of GmAOS transcript level
Real Time-PCR was performed in 20 μl reaction mixtures containing 2X Brillant SYBR green QPCR master mix (Stratagene), 1:10 dilution of synthesized cDNA and 400 nM of the primers GmAOSRTF (5′ GGGCCTCCCTCAATTCTTAG 3′) and GmAOSRTR (5′ CGCTCCGCTTCGTCTAATAC 3′). The soybean Actin gene was used as an endogenous control for sample normalization. Actin gene was amplified by using 100 nM primers GmACTRTf (5′ TCGTATGAGCAAGGAAATTGG 3′) and GmACTRTr (5′ TAGAGCCACCAATCCAGACAC 3′). The primers were designed based on soybean Actin sequence (accesion number V00450) and AOS sequence (accession number EU366252) (Wu et al. 2008), using Primer3′v4.0 software (Rozen and Skaletsky 2000).
Real-Time PCR determinations were carried out with RNA extracted from two independent biological samples and the threshold cycle (Ct) was determined in triplicate. The relative levels of transcription were calculated by using the 2−ΔΔCt method (Livak and Schmittgen 2001). Melting curves were recorded after cycle 40 by heating from 55 to 95 °C.
Statistical analysis
It was performed by subjecting the data to ANOVA and analyzed by LSD´s tests for statistical significance at p < 0.05, using Infostat software (1.0, FCA, UNC, Argentina). Student´s t test with α = 0.05 was performed subjecting the data to SigmaStat 3.5 software.
Results
Bacillus sp. CHEP5 systemically protects soybean plants against C. sojina
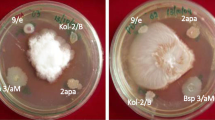
The ability of Bacillus sp. CHEP5 to systemically protect soybean plants against C. sojina was evaluated. Plants inoculated with Bacillus sp. CHEP5 previously to pathogen challenge, showed a reduced disease level, a decrease of approximately 40 % of the damaged leaf area and an elevated content of chlorophyll compared to non induced-challenged plants (Fig. 1 and Table 1).
Shoot dry weight increase, although not statistically significant, was also observed in induced plants compared to challenged ones (Table 1).
Bacillus sp. CHEP5 induced resistance is dependent on JA
It is known that JA has a main role in ISR signaling regulation and in plant defense response against necrotrophic fungi. Moreover, plant systemic defense induced by PGPB is usually associated with an enhanced sensitivity to this phytohormone upon pathogen attack.
To determine whether resistance to C. sojina induced by Bacillus sp. CHEP5 is associated with the JA signaling pathway, we analyzed GmAOS transcript level at 1 h post pathogen challenge. In response to C. sojina, an increase in the relative expression of GmAOS gene was observed in plants inoculated with Bacillus sp CHEP5 compared with the uninoculated plants (Fig. 2). Therefore, the Bacillus sp CHEP5-mediated resistance to C. sojina in soybean occurs in a JA-dependent manner.
Bacillus sp. CHEP5 primes the defense response associated to JA
Methyl jasmonate application induces the expression of all of the genes involved in JA biosynthesis, including AOS (Sasaki et al. 2001). To determine whether GmAOS gene is primed in the resistance response to C. sojina induced by Bacillus sp. CHEP5, we examined the GmAOS transcript level in Bacillus sp. CHEP5 inoculated or uninoculated soybean plants exposed to exogenous MeJA.
Inoculation of Bacillus sp. CHEP5 in MeJA exposed plants, led to increased GmAOS transcript level at 3 h after treatment (Fig. 3). These results suggest that Bacillus sp. CHEP5 inoculation induced in soybean plants a defense-priming mechanism that may facilitate more potent response against C. sojina. Noteworthy, at 0 and 1 h post MeJA treatment, GmAOS transcript level was lower in Bacillus sp. CHEP5-induced plants than in MeJA-treated plants, probably related with a decrease in the JA signaling pathway to suppress early activated defense response (Stein et al. 2008; Zamioudis and Pieterse 2012).
Discussion
Some root colonizing nonpathogenic bacteria may trigger disease resistance in the host plant, a phenomenon that has been termed ISR and that is effective against different types of plant pathogens. The ISR is generally regarded as a widespread and conserved trait, since the phenomenon is known for many plant species (Van Loon et al.1998, 2008; Verhagen et al. 2010). Thus, one of the objectives of this work was to determine if Bacillus sp. CHEP5, previously reported as ISR-inducing bacteria in peanut (Tonelli et al. 2011), protects soybean from FLS foliar disease by activating systemic responses.
Plants inoculated with Bacillus sp. CHEP5 previously to pathogen challenge showed a reduce disease level with healthier, greener leaves compared to non inoculated-challenged plants.
Bacterial induction of resistance in the plant is systemic, as Bacillus sp. CHEP5 is applied onto the roots and the response is detected in the shoot system, where the pathogen C. sojina is applied. Hence, it may be attributed to ISR.
Induction of systemic resistance in soybean plants by gram positive strains has been previously described. Buensanteai et al. (2009) reported that the inoculation of soybean seeds with the strain Bacillus amyloliquefaciens KPS46 induced systemic protection against the leaf disease caused by Xanthomonas axonopodis pv. Glycines, reducing the percentage of infected leaf area. This protection encompassed primed and incremented activity of defense related proteins and phenolic content. It was also determined that both SA- and JA-dependent signaling pathways are systemically triggered by KPS46 seed treatment.
It is known that the oxylipin JA and its derivatives play an important role in the regulation of plant-beneficial microbe interactions being central players in the regulation of ISR (Van der Ent et al. 2009). Moreover, JA is the major signaling molecule in plant defense response against herbivores and necrotrophic pathogens (Froehlich et al. 2001; Pieterse et al. 2012; Wasternack and Hause 2013; Ye et al. 2013).
Allene oxide synthase is one of the first enzymes of the oxylipin pathway to JA biosynthesis. The second objective of this work was to determine if the mechanism to induce systemic resistance of Bacillus sp. CHEP5 involves the priming of JA-dependent pathway. The increased GmAOS transcript level in Bacillus sp. CHEP5 and pathogen challenged plants strongly suggest that the enhanced soybean resistance to C. sojina attack induced by this bacterium occurs in a JA-dependent manner.
Although induced defense responses allow plants to conserve energy in the absence of pathogen challenge, plants may suffer damage during the lag phase required to mount defenses once an attack occurs (Frost et al. 2008). To compensate for this vulnerability, some plants have evolved defense-priming mechanisms that facilitate more rapid and effective responses to pathogen challenge, with low metabolic requirements. Defense priming can lead to an enhanced state of readiness for the induction of defense responses, allowing plants to be physiologically prepared to induce more potent defense responses upon subsequent pathogen attack (Ye et al. 2013).
Taking into account that the resistance to C. sojina enhanced by Bacillus sp. CHEP5 involves the JA-signaling pathway, we further examined whether this bacterium induces defense-priming mechanisms by measuring GmAOS expression in inoculated or uninoculated plants exposed to exogenous MeJA application. The fact that Bacillus sp. CHEP5 inoculation induced a more effective defense response than MeJA alone after 3 h treatment indicates that it is serving as an apparent priming agent for JA response.
Moreover, the transcript level of GmAOS was higher in plants induced with Bacillus sp. CHEP5 after 1 h C. sojina challenge than in noninduced plants, suggesting that the defense response upon phytopathogen attack is primed faster than with MeJA treatment.
In conclusion, results from this study demonstrated that Bacillus sp. CHEP5 induced the systemic resistance in soybean and reduced the severity of FLS. In addition, it is shown that JA-dependent pathway is involved in the response triggers by Bacillus sp. CHEP5. Irrespective of the pathway involved, this strain is a putative biocontrol agent to be used in intensive soybean production to decrease chemical inputs, although this ability needs to be proven in field assays.
References
Arnon DI (1949) Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol 24:1–15
Buensanteai N, Yuen G, Prathuangwong S (2009) Priming, signaling, and protein production associated with induced resistance by Bacillus amyloliquefaciens KPS46. World J Microbiol Biotechnol 25:1275–1286
Carmona MA, Scandiani M, Luque A (2009) Severe outbreaks of soybean frogeye leaf spot caused by Cercospora sojina in the pampean region, Argentina. Plant Dis 93:966
Conrath U, Beckers G, Flors V, García-Agustín P, Jakab G, Mauch F, Newman MA, Pieterse CMJ, Poinssot B, Pozo MJ, Pugin A, Schaffrath U, Ton J, Wendehenne D, Zimmerli L, Mauch-Mani B (2006) Priming: getting ready for battle. Mol Plant Microbe Interact 19:1062–1071
De Mayer G, Höfte M (1997) Salicylic acid produced by the rhizobacterium Pseudomonas aeruginosa 7NSK2 induces resistance to leaf infection by Botrytis cinerea on bean. Phytopathology 87:588–593
Fehr WR, Caviness CE (1977) Stages of soybean development. Iowa St. Univ Special Report 80, 11p
Froehlich JE, Itoh A, Howe GA (2001) Tomato allene oxide synthase and fatty acid hydroperoxide lyase, two cytochrome P450s involved in oxylipin metabolism, are targeted to different membranes of chloroplast envelope. Plant Physiol 125:306–317
Frost CJ, Mescher MC, Carlson JE, De Moraes CM (2008) Plant defense priming against herbivores: getting ready for a different battle. Plant Physiol 146:818–824
Hoagland DR, Arnon DI (1950) The water culture method for growing plants without soil. Calif Agric Expt Stn Circ 347:1–39
Kloepper J, Tuzun S, Kúc J (1992) Proposed definitions related to induced disease resistance. Biocontrol Sci Technol 2:349–351
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using Real-Time quantitative PCR and the 2−ΔΔCt method. Methods 25:402–408
Lugtenberg B, Kamilova F (2009) Plant-growth-promoting Rhizobacteria. Annu Rev Microbiol 63:541–556
Mc Kinney G (1938) Some absorption spectra of leaf extract. Plant Physiol 13:128–140
Mian MAR, Missaoui AM, Walker DR, Phillips DV, Boerma HR (2008) Frogeye leaf spot of soybean: a review and proposed race designations for isolates of Cercospora sojina Hara. Crop Sci 48:14–24
Paré PW, Farag MA, Krishnamachari V, Zhang H, Ryu CM, Kloepper JW (2005) Elicitors and priming agents initiate plant defense responses. Photosynth Res 85:149–159
Pieterse C, Van Pelt J, Ton J, Parchmann S, Mueller M, Buchala A, Meâ Traux JP, Van Loon L (2000) Rhizobacteria-mediated induced systemic resistance (ISR) in Arabidopsis thaliana requires sensitivity to jasmonate and ethylene but is not accompanied by an increase in their production. Physiol Mol Plant Pathol 57:123–134
Pieterse CMJ, Van der Does D, Zamioudis C, Leon-Reyes A, Van Wees S (2012) Hormonal modulation of plant immunity. Annu Rev Cell Dev Biol 28:489–521
Podile AR, Kishore K (2006) Plant growth-promoting rhizabacteria. In: Gnanamanickam SS (ed) Plant-associated bacteria. Springer, Berlin, pp 195–230
Pozo MJ, Van Der Ent S, Van Loon LC, Pieterse CMJ (2008) Transcription factor MYC2 is involved in priming for enhanced defense during rhizobacteria-induced systemic resistance in Arabidopsis thaliana. New Phytol 180:511–523
Reymond P, Farmer E (1998) Jasmonate and salicylate as global signals for defense gene expression. Curr Opin Plant Biol 5:404–411
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386. http://fokker.wi.mit.edu/primer3/
Sasaki Y, Asamizu E, Shibat D, Nakamura Y, Kaneko T, Awai K, Amagai C, Tsugane T, Masuda T, Shimada H, Takamiya K, Ohta H, Tabata S (2001) Monitoring of Methyl jasmonate-responsive genes in Arabidopsis by cDNA macroarray: self-activation of jasmonic acid biosynthesis and crosstalk with other phytohormone signaling pathways. DNA Res 8:153–161
Somasegaran P, Hoben H (1994) Quantifying the growth of rhizobia. In: Robert C. Garber (ed) Handbook for rhizobia: methods in legume rhizobia technology. Springer, New York, Section 3, pp 382–390
Stein E, Molitor A, Kogel KH, Waller F (2008) Systemic resistance in Arabidopsis conferred by the mycorrhizal fungus Piriformospora indica requires jasmonic acid signaling and the cytoplasmic function of NPR1. Plant Cell Physiol 49:1747–1751
Tonelli ML, Furlán A, Taurian T, Castro S, Fabra A (2011) Peanut priming induced by biocontrol agents. Physiol Mol Plant Pathol 75:100–105
Van der Ent S, Van Wees SCM, Pieterse CMJ (2009) Jasmonate signaling in plant interactions with resistance-inducing beneficial microbes. Phytochemestry 70:1581–1588
Van Loon LC, Bakker PAHM, Pieterse CMJ (1998) Systemic resistance induced by rhizosphere bacteria. Annu Rev Phytopathol 36:453–483
Van Loon LC, Bakker PAHM, Van der Heijdt WHW, Wendehenne D, Pugin A (2008) Early responses of tobacco suspension cells to rhizobacterial elicitors of induced systemic resistance. Mol Plant Microbe Interact 21:1609–1621
Verhagen BWM, Glazebrook J, Zhu T, Chang HS, Van Loon LC, Pieterse CMJ (2004) The transcriptome of rhizobacteria-induced systemic resistance in Arabidopsis. Mol Plant Microbe Interact 17:895–908
Verhagen BWM, Trotel-Aziz P, Couderchet M, Couderchet M, Höfte M, Aziz A (2010) Pseudomonas spp.-induced systemic resistance to Botrytis cinerea is associated with induction and priming of defense responses in grapevine. J Exp Bot 61:249–260
Wasternack C, Hause B (2013) Jasmonates: biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Anna Bot 111:1021–1058
Wu J, Wu Q, Wu Q, Gai J, Yu D (2008) Constitutive overexpression of AOS-like gene from soybean enhanced tolerance to insect attack in transgenic tobacco. Biotechnol Lett 30:1693–1698
Ye M, Song Y, Long J, Wang R, Baerson S, Pan Z, Zhu-Salzman K, Xie J, Cai K, Luo S, Zeng R (2013) Priming of jasmonate-mediated antiherbivore defense responses in rice by silicon. Proc Nat Acad Sci 110:3631–3639
Zamioudis C, Pieterse CMJ (2012) Modulation of host immunity by beneficial microbes. Mol Plant Microbe Interact 25:139–150
Acknowledgments
This study was financially supported by the SECyT-UNRC, CONICET, Ministerio de Ciencia y Tecnología de Córdoba, ANPCyT. M. L. Tonelli and A. Fabra are members of the Research Career from CONICET.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tonelli, M.L., Fabra, A. The biocontrol agent Bacillus sp. CHEP5 primes the defense response against Cercospora sojina . World J Microbiol Biotechnol 30, 2503–2509 (2014). https://doi.org/10.1007/s11274-014-1675-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-014-1675-3