Abstract
Environmental microbiology investigation was performed to determine the molecular diversity of β-lactamase genes among ampicillin-resistant bacteria from Jiaozhou Bay. β-lactamase genes were detected in 93.8% of the bacterial isolates identified as Enterobacteriaceae. The most frequently detected gene was bla TEM, followed by bla SHV, bla OAX-1, bla MOX and bla CMY. Most of the isolates (68.8%) were positive for the intI1 integrase gene, and two isolates were also found for the intI2 gene. The dfr and aadA gene cassettes were predominant. Anthropogenic contamination from onshore sewage processing plants might contribute predominantly to the β-lactamase gene reservoir in the studied coastal waters. Environmental antibiotic-resistant bacteria and resistance genes may serve as bioindicators of coastal environmental quality or biotracers of the potential contamination sources. This is the first report of the prevalence and characterization of β-lactamase genes and integrons in coastal Enterobacteriaceae from China.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Extensive and intensive use and misuse of antibiotics in medication, veterinary, agriculture and aquaculture have resulted in new selective pressures on environmental antibiotic-resistant bacteria (Costanzo et al. 2005; Cabello 2006). The emergence of antibiotic resistance among bacterial pathogens has become a serious global health and economical problem (French 2005). Resistance genetic determinants have also been detected in a large number of microbial communities from natural environments (Alonso et al. 2001). Antibiotic resistance in Enterobacteriaceae, partially bacterial opportunist pathogens, has increased dramatically in aquatic environments (Schwartz et al. 2003; Henriques et al. 2006; Mukherjee and Chakraborty 2006). The prevalence of diverse antibiotic-resistant bacteria and resistance genes makes coastal ecosystems a reservoir of antibiotic resistance (Young 1993; Biyela et al. 2004).
Bacterial resistance to β-lactam antibiotics caused by the spread of the extended-spectrum β-lactamases (ESBLs) and plasmid-mediated AmpC β-lactamases is becoming more and more important globally for clinical and environmental concerns (Bradford 2001; Bonfiglio et al. 2002; Philippon et al. 2002; Walther-Rasmussen and Hoiby 2002; Lee et al. 2006). Most ESBLs genes are usually derivatives of the common TEM, SHV or OXA genes (Bradford 2001). The transfer of ESBL genes in the Enterobacteriaceae are usually plasmid-mediated, too (Rupp and Fey 2003; Shah et al. 2004; Pfaller and Segreti 2006), facilitating the spread of resistance within and between species. These genes might also be strongly correlated with multidrug resistance in Enterobacteriaceae (Nathisuwan et al. 2001; Philippon et al. 2002). Moreover, the coexistence of ESBLs and AmpC β-lactamases in certain Enterobacteriaceae strains might provide a broader spectrum of resistance (Alvarez et al. 2004). The β-lactamase gene analyses may provide an understanding of the complexity of the environmental resistance determinants and their origins.
Transfer of antibiotic resistance can be facilitated by mobile genetic elements, such as plasmids, transposons and integrons (Olsen 1999; White and McDermott 2001). Recent studies have indicated that integrons play a particularly important role in the spread of multidrug resistance in Gram-negative bacteria due to their ability to capture and exchange resistance genes via site-specific recombination (Ploy et al. 2000; White et al. 2001; Weldhagen 2004). The detection and characterization of integrons and gene cassettes may be critical in understanding the resistance dissemination pathways and in evaluating the potential of an environment to act as an antibiotic resistance reservoir (Henriques et al. 2006).
Although the studies of resistance genetic determinants in aquatic environments are important, there are few reports from China. Jiaozhou Bay, located on the western coast of the Yellow Sea, is a semi-enclosed waterbody with significant anthropogenic perturbations. Surrounding small rivers used to form the major sources of discharged sewage. Improperly processed hospital effluent and civic wastewater could be a source of antibiotics and antibiotic-resistant bacteria (Kim and Aga 2007). Excessive usage of antibiotics in mariculture may also stimulate the propagation of resistant bacteria (Cabello 2006). Our previous studies indicated that the Jiaozhou Bay was contaminated with tetracycline- and chloramphenicol-resistant Enterobacteriaceae, especially at the eastern coast stations close to the onshore sewage processing plants of the Qingdao city (Dang et al. 2008a, b). In the current work, we studied the molecular diversity of the β-lactamase genes (especially those of the plasmid-mediated AmpC or other bla genes), integrons and gene cassettes of the environmental multidrug-resistant bacteria isolates, in order to gain a basic understanding of the potential influence of anthropogenic activities, such as wastewater discharging, on the clinically and environmentally important Enterobacteriaceae population in Jiaozhou Bay.
Materials and methods
Bacterial isolates and antimicrobial susceptibility tests
A total of 99 isolates resistant to OTC100 (100 μg/ml oxytetracycline), including 42 from A5 station, 42 from C4 station, 11 from Y1 station and 4 from D6 station (Fig. 1), were selected from a previous study (Dang et al. 2008a). Resistance to ampicillin might be a predictor of the presence of integrons (Leverstein-van Hall et al. 2003), thus this antibiotic was used as the starting reagent for multidrug-resistance screening. Isolates were inoculated onto tryptic soy agar (TSA, Difco formula) plates supplemented with 3% (w/v) NaCl and 100 μg/ml ampicillin (AMP100). The positively growing resistant isolates were selected for further study.
Antibiotic susceptibility tests were determined based on the method described by Dang et al. (2007). The TSA plates were supplemented with 3% (w/v) NaCl and one of the following antimicrobials: streptomycin (30 μg/ml), chloramphenicol (30 μg/ml), kanamycin (30 μg/ml), trimethoprime-sulfamethoxazole (30 μg/ml), erythromycin (15 μg/ml) and nalidixic acid (30 μg/ml), for multidrug resistance assay.
β-lactamase-specific PCRs
Primers used to amplify the plasmid-mediated AmpC (Hong et al. 2005) and other major members of the β-lactamase genes (Colom et al. 2003) are shown in Table 1. A boiling method was used for extraction of the total DNA from AMP100-resistant strains, including genomic and plasmid DNA (Dang et al. 2007). PCR products were analyzed by electrophoresis on 1% agarose gels stained with ethidium bromide.
Molecular determination of phylogenetic affiliations
The isolates harboring the β-lactamase genes determined in the above step were analyzed of 16S rRNA gene (rDNA) sequences of the isolates to determine their phylogenetic affiliations. The primers 27F and 1387R, and the PCR conditions were described previously (Dang and Lovell 2000). When an isolate’ phylogenetic affiliation could not be unambiguously determined with 16S rDNA sequence analysis, phenotypic characterization, including colony morphology, Gram stain, lactose fermentation, glucose oxidation and acetate reduction, was determined.
PCR amplification for integron detection
PCR amplification of specific integrase gene fragments was used for integron identifications of bacteria carrying β-lactamase genes (Table 1). PCR conditions for the detection of the class 1 and class 2 integrons and also the gene cassettes have been described elsewhere (Kraft et al. 1986; Levesque et al. 1995; Goldstein et al. 2001; White et al. 2001). Amplicons of gene cassettes were digested with MspI and HhaI, respectively. Restriction fragments were resolved by electrophoresis on 4% agarose gels in 0.5× TBE, and digitally photographed with an ImageMaster VDS imaging system (Pharmacia Biotech, USA). The band patterns of the RFLP analysis were compared to identify potential identical sequences. PCR products of typical RFLP profiles were purified and ligated into pMD19-T simple vectors (Takara, Japan), transformed into E. coli TOP10 competent cells. The transformants were selected by using X-Gal-IPTG Luria-Bertani (LB) indicator plates supplemented with 100 μg/ml ampicillin.
Sequencing and sequence analyses
Primers for amplification of each gene fragment in the above steps were used for sequencing, except for the gene cassette DNA fragments, which were sequenced using the clone vector primers M13-D (5′-AGGGTTTTCCCAGTCACGACG-3′) and RV-M (5′-GAGCGGATAACAATTTCACACAGG-3′). Bioinformatic determination of the sequence phylogenetic affiliations followed standard methods (Dang and Lovell 2000). NCBI GenBank database was queried for initial determinations of the nearest neighbor sequences via the BLAST program (Altschul et al. 1997). 16S rDNA sequences were aligned using the ClustalX program (version 1.8) (Thompson et al. 1994). Phylogenetic trees were constructed using the DNADIST and NEIGHBOR programs within the PHYLIP package (version 3.67) (Felsenstein 1989).
Nucleotide sequence accession numbers
The β-lactamase gene and the 16S rDNA sequences determined in the current study have been deposited in the GenBank database under accession nos. EU420893 to EU420956, and the representative class 1 and class 2 integrons and the gene cassettes sequences determined under accession nos. EU523051 to EU523064.
Results
Antibiotic resistance profiles
A total of 66 isolates (30 from C4 station, 28 from A5 station, 7 from Y1 station and 1 from D6 station) showed resistance to 100 μg/ml ampicillin. All these isolates were also multidrug-resistant (resistant to more than two classes of antibiotics). Most frequently those additional resistances were to trimethoprime-sulfamethoxazole (86.4%) and nalidixic acid (83.3%). Particularly among the resistant strains, 7 isolates from C4 station, 1 isolate from A5 station and 1 isolate from Y1 station were resistant to all the antibiotics tested (Table 2).
Occurrence and diversity of β-lactamase genes
A total of 66 AMP100-resistant isolates were selected to screen the β-lactamase genes by multiplex PCR methods. About 32 isolates were found to carry at least one of the β-lactamase genes (Table 2). Based on 16S rDNA analyses (Fig. 2), these isolates included 30 Enterobacteriaceae strains, 1 Acinetobacter strain and 1 Aeromonas strain. The predominant resistance gene was bla TEM, occurring in 17 isolates (53.1%), affiliated with Escherichia coli (10 isolates), Klebsiella oxytoca (5 isolates), Enterobacter sp. (1 isolates) and Kluyvera intermedia (1 isolate). The resistance gene bla SHV was detected in 9 isolates, affiliated with Klebsiella pneumoniae (8 isolates) and Citrobacter freundii (1 isolate). The resistance gene bla OXA-1 was detected in 1 Citrobacter freundii isolate and 1 Acinetobacter seohaensis isolate. For the plasmid-mediated AmpC β-lactamase genes, bla CMY was observed in 2 Citrobacter freundii isolates, and bla MOX-2 was observed in 1 Acinetobacter isolate and 1 Citrobacter freundii isolate. All the E. coli and K. oxytoca isolates carried the bla TEM gene, while all the K. pneumoniae isolates carried the bla SHV gene. Nucleotide sequence analysis revealed that all the sequences of the screened genes had 98% or higher similarity to their best-match known genes retrieved from the GenBank database, but it is difficult to distinguish from these fragments sequenced whether the bla TEM and bla SHV correspond to ESBLs or not, except that bla OXA-1 gene can be identified as ESBLs. Because the amplification fragments of bla TEM and bla SHV with the primers are too short to contain the point mutations. More specific techniques should be used to discriminate wild-type bla TEM and bla SHV genes.
Phylogenetic tree constructed based on partial 16S rRNA genes sequences using neighbor-joining method for the 32 β-lactamase gene-harboring bacterial isolates. Bootstrap values greater than 70% were indicated at the branch points as ‘*’, generated by 100 replicates of neighbor-joining and parsimony analysis. The β-lactamase genes detected are labeled in parentheses for the corresponding isolates
Occurrence and diversity of integrons and gene cassettes
A total of 22 out of the 32 β-lactamase gene-carrying isolates (68.8%) had the class 1 integrase gene intI1. Two strains, SA-C4-59 and SA-A5-61, were also found to possess the class 2 integrase gene intI2. The class 1 and class 2 regions of the gene cassettes of the integrase positive isolates were analysed. About 12 isolates were found to carry the gene cassette fragments amplified with 5′_CS/3′_CS (Table 2). Sequence analysis of the cassette arrays revealed a predominance of dfr and aadA cassettes that conferred resistance to trimethoprim, and streptomycin and spectinomycin, respectively. Additional gene cassette arrays were also detected, including catB8 and cmlA encoding resistance to chloramphenicol, aacA4 encoding resistance to aminoglycosides, arr-3 encoding resistance to rifampicin, and orfF with unknown function. Seven isolates (3 Klebsiella oxytoca, 3 Escherichia coli and 1 Kluyvera intermedia) carried the same class 1 integron gene cassette arrays (dfr17, aadA5).
Discussion
Enterobacteriaceae remain important causes of both nosocomial and community-acquired infections. The emergence and wide dissemination of multidrug resistant Enterobacteriaceae environmental strains represent a potential risk for human health. Our previous studies showed that antibiotic-resistant Enterobacteriaceae strains might be a serious contamination in Jiaozhou Bay (Dang et al. 2008a, b). In the current study, 30 out of 32 (93.8%) bacterial isolates with β-lactamase genes were identified as Enterobacteriaceae strains, with Klebsiella sp. and Escherichia coli being the most predominant strains. Our data indicate that the Jiaozhou Bay provides a good model for studying the molecular diversity and potential sources of antibiotic-resistant bacteria and resistance genes. Similar situations from other areas of the world coastal environments about antibiotic-resistance contamination from potential commensal bacteria worth being further studied.
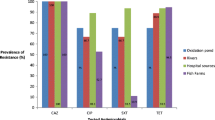
The distribution of lactam-resistant bacteria was quite heterogeneous and area-specific in Jiaozhou Bay. More than 60% of the OTC100-resistant bacteria isolated from the eastern coast stations, such as A5, C4 and Y1, were found to be AMP100 resistant, while only 25% were found from station D6. Station A5 is located near the river mouth of Licun and station C4 is located near the river mouth of Haipo (Fig. 1), where the nearby sewage processing plants discharge a huge amount of processed wastewater into the bay (Liu et al. 2005). Station Y1 is located at the river mouths of Loushan and Moshui near the Hongdao maricultural area. Station D6 is located at the outside of Jiaozhou Bay, possibly with less impact from terrestrial or maricultural contamination. Although the seawater of Jiaozhou Bay is dynamic and water mixing happens frequently due to currents, tides and other hydrological factors (Liu et al. 2004), confined distribution of resistant bacteria, especially the Enterobacteriaceae strains, still could be identified along the eastern coast, indicating a strong terrestrial or anthropogenic impact. Our current environmental microbiology study indicates that microbial spatial distribution might be closely related to geographical characteristics and anthropogenic activities, and sewage contamination might be a serious environmental problem of Jiaozhou Bay. The prevalence and dissemination of antibiotic-resistant commensal bacteria may serve as a bioindicator or biotracer, potentially useful in revealing the coastal environmental quality or contamination source (Dang et al. 2008a, b).
To date, although a variety of β-lactamase gene types have been described, bla TEM and bla SHV are the major types found in members of the family Enterobacteriaceae (Cantón et al. 2008; Villegas et al. 2008). In coastal waters of Jiaozhou Bay, the most frequently detected gene was bla TEM, followed by bla SHV, bla OXA-1, bla MOX and bla CMY. Our result is in agreement with other reports on the diversity and dominance of the lactam-resistant bacteria and β-lactamase genes in aquatic environments (Henriques et al. 2006; Alpay-Karaoglu et al. 2007), indicating that it might be a global phenomenon for the prevalence and dominance of Enterobacteriaceae bacteria and their lactam-resistant genes in aquatic environments with significant anthropogenic contamination impact.
In the current study, class 1 integrons were detected in 68.8% of the β-lactamase gene-carrying bacterial isolates from Jiaozhou Bay. This proportion is slightly higher than what was found in some aquatic environments (Lin and Biyela 2005), but much higher than other aquatic environments (Rosser and Young 1999; Roe et al. 2003; Henriques et al. 2006; Ozgumus et al. 2007). It has been found that there is a strong correlation between the presence of integrons and multidrug resistance (White et al. 2001). According to our susceptibility test result (Table 2), integron-positive isolates were more likely to be multidrug resistant than integron-negative isolates. These data indicated that integrons were prevalent and might play an important role in multidrug resistance in the environmental β-lactamase-producing Enterobacteriaceae strains.
Gene cassettes were found to be associated with resistance to several major classes of antibiotics. The most frequently found resistance gene cassettes dfr17 and aadA5 in the current study might be widespread in Enterobacteriaceae strains of aquatic environments (Ozgumus et al. 2007), similar to clinical settings (White et al. 2001). In our study, only aminoglycoside, trimethoprim and chloramphenicol could be directly related to the presence of the gene cassettes (Table 2). The association of the other antibiotics, such as ampicillin, erythromycin and tetracycline, with the presence of an integron is likely to be due to genetic linkage between integrons and conjugative plasmids or transposons (White et al. 2001). Our result showed that 11 isolates containing the intI1 gene were not detected with gene cassette regions. This was probably due to the lack of a 3′ conserved region or the variable region was too long to be amplified in these isolates (Barlow et al. 2004; Yao et al. 2007).
Coastal environment deterioration is becoming more complicated due to contaminations of antimicrobials and antibiotic-resistant bacteria. Discharged civic and hospital wastewater, even after processing, might contribute to the resistance bacteria and gene reservoir in aquatic ecosystems. Our study indicates the possibility that antibiotic-resistant bacteria and resistance genes may serve as bioindicators of coastal environmental quality or biotracers of major contamination sources. Efficient and timely surveillance is crucial for high-quality environmental management and for prevention and control of microbial diseases and infections caused by environmental antibiotic-resistant bacteria.
References
Alonso A, Sanchez P, Martinez JL (2001) Environmental selection of antibiotic resistance genes. Environ Microbiol 3:1–9. doi:10.1046/j.1462-2920.2001.00161.x
Alpay-Karaoglu S, Ozgumus OB, Sevim E, Kolayli F, Sevim A, Yesilgil P (2007) Investigation of antibiotic resistance profiles and TEM-type β-lactamase gene carriage of ampicillin-resistant Escherichia coli strains isolated from drinking water. Ann Microbiol 57:281–288
Altschul S, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. doi:10.1093/nar/25.17.3389
Alvarez M, Tran JH, Chow N, Jacoby GA (2004) Epidemiology of conjugative plasmid-mediated AmpC β-lactamases in the United States. Antimicrob Agents Chemother 48:533–537. doi:10.1128/AAC.48.2.533-537.2004
Barlow RS, Pemberton JM, Desmarchelier PM, Gobius KS (2004) Isolation and characterization of integron-containing bacteria without antibiotic selection. Antimicrob Agents Chemother 48:838–842. doi:10.1128/AAC.48.3.838-842.2004
Biyela PT, Lin J, Bezuidenhout CC (2004) The role of aquatic ecosystems as reservoirs of antibiotic resistant bacteria and antibiotic resistance genes. Water Sci Technol 50:45–50
Bonfiglio G, Perilli M, Stefani S, Amicosante G, Nicoletti G (2002) Prevalence of extended spectrum β-lactamases among Enterobacteriaceae: an Italian survey. Int J Antimicrob Agents 19:213–217. doi:10.1016/S0924-8579(01)00497-6
Bradford PA (2001) Extended-spectrum β-lactamase in the 21st century: characterization, epidemiology and detection of this important resistance threat. Clin Microbiol Rev 14:933–951. doi:10.1128/CMR.14.4.933-951.2001
Cabello FC (2006) Heavy use of prophylactic antibiotics in aquaculture: a growing problem for human and animal health and for the environment. Environ Microbiol 8:1137–1144. doi:10.1111/j.1462-2920.2006.01054.x
Cantón R, Novais A, Valverde A, Machado E, Peixe L, Baquero F et al (2008) Prevalence and spread of extended-spectrum β-lactamase-producing Enterobacteriaceae in Europe. Clin Microbiol Infect 14:144–153. doi:10.1111/j.1469-0691.2007.01850.x
Colom K, Pérez J, Alonso R, Fernández-Aranguiz A, Lariño E, Cisterna R (2003) Simple and reliable multiplex PCR assay for detection of bla TEM, bla SHV and bla OXA-1 genes in Enterobacteriaceae. FEMS Microbiol Lett 223:147–151. doi:10.1016/S0378-1097(03)00306-9
Costanzo SD, Murby J, Bates J (2005) Ecosystem response to antibiotics entering the aquatic environment. Mar Pollut Bull 51:218–223. doi:10.1016/j.marpolbul.2004.10.038
Dang HY, Lovell CR (2000) Bacterial primary colonization and early succession on surfaces in marine waters as determined by amplified rRNA gene restriction analysis and sequence analysis of 16S rRNA genes. Appl Environ Microbiol 66:467–475. doi:10.1128/AEM.66.2.467-475.2000
Dang H, Zhang X, Song L, Chang Y, Yang G (2007) Molecular determination of oxytetracycline resistant bacteria and their resistance genes from mariculture environments of China. J Appl Microbiol 103:2580–2592. doi:10.1111/j.1365-2672.2007.03494.x
Dang HY, Ren J, Song LS, Sun S, An LG (2008a) Diverse tetracycline resistant bacteria and resistance genes from coastal waters of Jiaozhou Bay. Microb Ecol 55:237–246. doi:10.1007/s00248-007-9271-9
Dang HY, Ren J, Song LS, Sun S, An LG (2008b) Dominant chloramphenicol-resistant bacteria and resistance genes in coastal marine waters of Jiaozhou Bay, China. World J Microbiol Biotechnol 24:209–217. doi:10.1007/s11274-007-9458-8
Felsenstein J (1989) PHYLIP—phylogeny inference package (version 3.2). Cladistics 5:164–166
French GL (2005) Clinical impact and relevance of antibiotic resistance. Adv Drug Deliv Rev 57:1514–1527. doi:10.1016/j.addr.2005.04.005
Goldstein C, Lee MD, Sanchez S, Hudson C, Philips B, Register B (2001) Incidence of class 1 and 2 integrases in clinical and commensal bacteria from livestock, companion animals, and exotics. Antimicrob Agents Chemother 45:723–726. doi:10.1128/AAC.45.3.723-726.2001
Henriques I, Moura A, Alves A, Saavedra MJ, Correia A (2006) Occurrence and diversity of integrons and β-lactamase genes among ampicillin-resistant isolates from estuarine waters. Res Microbiol 157:938–947. doi:10.1016/j.resmic.2006.09.003
Hong Y, Park H, Lee W, Song J, Jeong D (2005) Evaluation of phenotypic screening methods for detecting plasmid-mediated AmpC β-lactamases-producing isolates of Escherichia coli and Klebsiella pneumoniae. Diagn Microbiol Infect Dis 53:319–323. doi:10.1016/j.diagmicrobio.2005.07.004
Kim S, Aga DS (2007) Potential ecological and human health impacts of antibiotics and antibiotic-resistant bacteria from wastewater treatment plants. J Toxicol Environ Health B Crit Rev 10:559–573. doi:10.1080/15287390600975137
Kraft CA, Timbury MC, Platt DJ (1986) Distribution and genetic location of Tn7 in trimethoprim-resistant Escherichia coli. J Med Microbiol 22:125–131
Lee K, Lee M, Shin JH, Lee MH, Kang SH, Park AJ et al (2006) Prevalence of plasmid-mediated AmpC β-lactamases in Escherichia coli and Klebsiella pneumoniae in Korea. Microb Drug Resist 12:44–49. doi:10.1089/mdr.2006.12.44
Leverstein-van Hall MA, Blok HEM, Donders ART, Paauw A, Fluit AC, Verhoef J (2003) Multidrug resistance among Enterobacteriaceae is strongly associated with the presence of integrons and is independent of species or isolate origin. J Infect Dis 187:251–259. doi:10.1086/345880
Levesque C, Piche L, Larose C, Roy PH (1995) PCR mapping of integrons reveals several novel combinations of resistance genes. Antimicrob Agents Chemother 39:185–191
Lin J, Biyela PT (2005) Convergent acquisition of antibiotic resistance determinants amongst the Enterobacteriaceae isolates of the Mhlathuze River, KwaZulu-Natal (RSA). Water SA 31:257–260
Liu Z, Wei H, Liu G, Zhang J (2004) Simulation of water exchange in Jiaozhou Bay by average residence time approach. Estuar Coast Shelf Sci 61:25–35. doi:10.1016/j.ecss.2004.04.009
Liu SM, Zhang J, Chen HT, Zhang GS (2005) Factors influencing nutrient dynamics in the eutrophic Jiaozhou Bay, North China. Prog Oceanogr 66:66–85. doi:10.1016/j.pocean.2005.03.009
Mukherjee S, Chakraborty R (2006) Incidence of class 1 integrons in multiple antibiotic-resistant gram-negative copiotrophic bacteria from the River Torsa in India. Res Microbiol 157:220–226. doi:10.1016/j.resmic.2005.08.003
Nathisuwan S, Burgess DS, Lewis JS 2nd (2001) Extended-spectrum β-lactamases: epidemiology, detection, and treatment. Pharmacotherapy 21:920–928. doi:10.1592/phco.21.11.920.34529
Olsen JE (1999) Antibiotic resistance: genetic mechanisms and mobility. Acta Vet Scand Supp 92:15–22
Ozgumus OB, Celik-Sevim E, Alpay-Karaoglu S, Sandalli C, Sevim A (2007) Molecular characterization of antibiotic resistant Escherichia coli strains isolated from tap and spring waters in a coastal region in Turkey. J Microbiol 45:379–387
Pfaller MA, Segreti J (2006) Overview of the epidemiological profile and laboratory detection of extended-spectrum β-lactamases. Clin Infect Dis 42:S153–S163. doi:10.1086/500662
Philippon A, Arlet G, Jacoby GA (2002) Plasmid-determined AmpC type β-lactamases. Antimicrob Agents Chemother 46:1–11. doi:10.1128/AAC.46.1.1-11.2002
Ploy MC, Lambert T, Couty JP, Denis F (2000) Integrons: an antibiotic resistance gene capture and expression system. Clin Chem Lab Med 38:483–487. doi:10.1515/CCLM.2000.070
Roe MT, Vega E, Pillai SD (2003) Antimicrobial resistance markers of class 1 and class 2 integron-bearing Escherichia coli from irrigation waters and sediments. Emerg Infect Dis 9:822–826
Rosser SJ, Young HK (1999) Identification and characterization of class 1 integrons in bacteria from an aquatic environment. J Antimicrob Chemother 44:8–11. doi:10.1093/jac/44.1.11
Rupp ME, Fey PD (2003) Extended spectrum β-lactamase (ESBL)-producing Enterobacteriaceae: considerations for diagnosis, prevention and drug treatment. Drugs 63:353–365. doi:10.2165/00003495-200363040-00002
Schwartz T, Kohnen W, Janses B, Obst U (2003) Detection of antibiotic-resistant bacteria and their resistance genes in wastewater, surface water, and drinking water biofilms. FEMS Microbiol Ecol 43:325–335. doi:10.1111/j.1574-6941.2003.tb01073.x
Shah AA, Hasan F, Ahmed S, Hameed A (2004) Extended-spectrum β-lactamases (ESBLs): characterization, epidemiology and detection. Crit Rev Microbiol 30:25–32. doi:10.1080/10408410490266429
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 9:3251–3270
Villegas MV, Kattan JN, Quinteros MG, Casellas JM (2008) Prevalence of extended-spectrum β-lactamases in South America. Clin Microbiol Infect 14:154–158. doi:10.1111/j.1469-0691.2007.01869.x
Walther-Rasmussen J, Hoiby N (2002) Plasmid-borne AmpC β-lactamases. Can J Microbiol 48:479–493. doi:10.1139/w02-039
Weldhagen GF (2004) Integrons and β-lactamases—a novel perspective on resistance. Int J Antimicrob Agents 23:556–562. doi:10.1016/j.ijantimicag.2004.03.007
White DG, McDermott PF (2001) Biocides, drug resistance and microbial evolution. Curr Opin Microbiol 4:313–317. doi:10.1016/S1369-5274(00)00209-5
White PA, MacIver CJ, Rawlinson WD (2001) Integrons and gene cassettes in the Enterobacteriaceae. Antimicrob Agents Chemother 45:2658–2661. doi:10.1128/AAC.45.9.2658-2661.2001
Yao F, Qian YS, Chen SZ, Wang PF, Huan YC (2007) Incidence of extended-spectrum β-lactamases and characterization of integrons in extended-spectrum β-lactamase-producing Klebsiella pneumoniae isolated in Shantou, China. Acta Biochim Biophys Sin (Shanghai) 39:527–532. doi:10.1111/j.1745-7270.2007.00304.x
Young HK (1993) Antimicrobial resistance spread in aquatic environments. J Antimicrob Chemother 31:627–635. doi:10.1093/jac/31.5.627
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China grants (NSFC Grant Nos. 40576069 and 40476058), the Hi-Tech Research and Development Program of China grant 2007AA091903, the China Ocean Mineral Resources R&D Association grants DYXM-115-02-2-6 and DYXM-115-02-2-20.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, C., Dang, H. & Ding, Y. Incidence of diverse integrons and β-lactamase genes in environmental Enterobacteriaceae isolates from Jiaozhou Bay, China. World J Microbiol Biotechnol 24, 2889–2896 (2008). https://doi.org/10.1007/s11274-008-9827-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-008-9827-y